Abstract
Sequence-related amplified polymorphism (SRAP) analysis was used to uncover genetic polymorphisms among alfalfa populations recurrently selected for superior tolerance to freezing (TF populations). Bulk DNA samples (45 plants/bulk) from each of the cultivar Apica (ATF0), and populations ATF2, ATF4, ATF5, and ATF6 were evaluated with 42 different SRAP primer pairs. Several polymorphisms that progressively intensified or decreased with the number of recurrent cycles were identified. Four positive polymorphisms (F10-R14, Me4-R8, F10-R8 and F11-R9) that, respectively, yielded 540-, 359-, 213-, and 180-bp fragments were selected for further analysis. SRAP amplifications with genotypes within ATF populations confirmed that the polymorphisms identified with bulk DNA samples were reflecting changes in the frequency of their occurrence in response to selection. In addition, the number of genotypes cumulating multiple polymorphisms markedly increased in response to recurrent selection. Independent segregation of the four SRAP polymorphisms suggests location at unlinked loci. Homology search gave matches with BAC clones from syntenic Medicago truncatula for the four SRAP fragments. Analysis of the relationship with low temperature tolerance showed that multiple SRAP polymorphisms are more frequent in genotypes that maintain superior regrowth after freezing. These results show that SRAP analysis of bulk DNA samples from recurrent selections is an effective approach for the identification of genetic polymorphisms associated with quantitative traits in allogamous species. These polymorphisms could be useful tools for indirect selection of freezing tolerance in alfalfa.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Alfalfa is an open-pollinated autotetraploid (2n = 4x = 32) that shows extensive genetic variability within its gene pool (Jones and Bingham 1995). Cultivars currently available are not well adapted to harsh winter conditions that occur in the midwest United States, Canada and northern Europe (Volenec et al. 2002). Improvement of winter hardiness has essentially been achieved by selection within field nurseries of plants that survive test winters. However, the unpredictability of the occurrence of adequate screening conditions along with genotype by environment (G × E) interactions requires the costly maintenance of nurseries for many years and, in some cases, at several locations (Limin and Fowler 1991). We recently applied a recurrent selection protocol entirely performed indoor to accelerate the development of alfalfa populations selectively improved for their tolerance to freezing (TF populations). These TF populations showed significant increases in freezing tolerance and survival to harsh winter conditions in response to selection (Castonguay et al. 2009).
Bulked segregant analysis (BSA; Michelmore et al. 1991) is a simple and very effective approach to identify genetic polymorphisms linked to a specific trait by comparing pools of DNA from plant siblings with contrasted phenotypes (Toppino et al. 2008). BSA of alfalfa TF populations that show a progressive increase in freezing tolerance could be particularly useful to uncover variations in genomic DNA closely linked to loci controlling freezing tolerance in alfalfa. The sequence-related amplified polymorphism (SRAP) technique of Li and Quiros (2001) is a highly reproducible technique for tagging and mapping neutral markers in plants (Jones et al. 2009). These polymorphisms result mainly from differences in length of introns, promoters and spacers among genotypes or populations. SRAP was shown to be more informative for detecting genetic diversity than other PCR-based techniques (Budak et al. 2004a). SRAPs have been successfully used to study genetic relationships and diversity and to uncover molecular markers in several species (Bertrand et al. 2009; Budak et al. 2004b; Ferriol et al. 2003; Gulsen et al. 2007; Liu et al. 2007; Sun et al. 2006; Yi et al. 2008), including alfalfa (Ariss and Vandemark 2007; Vandemark et al. 2006).
The objective of this work was to identify and validate DNA polymorphisms associated with superior freezing tolerance in alfalfa. Bulk DNA samples from populations recurrently selected for superior tolerance to freezing (TF populations) were used to uncover SRAP polymorphisms linked to tolerance loci. The co-inheritance of these polymorphisms with the freezing tolerance trait in segregating genotypes within TF populations was also assessed.
Materials and methods
Plant materials
Genotypes of alfalfa (Medicago sativa spp. sativa), cultivar Apica (ATF0) (Michaud et al. 1983), and populations (ATF2, ATF4, ATF5, and ATF6) derived from this cultivar after, respectively, 2, 4, 5, and 6 cycles of recurrent selection for superior freezing tolerance (Castonguay et al. 2009) were seeded individually in Ray Leach Cone-tainers™ (SC-10 Super Cell, Stuwe & Sons Inc) filled with a mixture of (10:3, v:v) of top soil/peat moss (Pro-mix BX, Premier Peat Moss, Rivière-du-Loup, QC, Canada) supplemented with a controlled release fertilizer [N: 17% (w:w); P: 7.31% (w:w); K: 14.1% (w:w); 250 g/35 l; Muticote 4, Haifa Chemicals Ltd, Haifa Bay, Israel]. Plants were maintained in an environmentally controlled chamber set to: photoperiod 16 h, day-time temperature 22°C, and night-time temperature 17°C. Artificial lighting was provided by a mixture of high pressure sodium and metal halide 400 W lamps (PL light systems, Beamsville, ON, Canada) with photosynthetic photon flux density of 600–800 μmol photons m−2 s−1. Plants were kept well watered and fertilized twice a week with a 1 g l−1 of a commercial fertilizer (20–20–20 plus micronutrients, Plant-Prod, Brampton, ON, Canada). Micronutrients percent composition of the fertilizer was 1.0 mg l−1 Mn, 0.5 mg l−1 Cu, 0.2 mg l−1 B, and 0.005 mg l−1 Mo.
Evaluation of freezing tolerance
In the summer of 2004, several cuttings were harvested from newly developed internodes from each of ~45 genotypes within each of ATF0, ATF2 and ATF5 populations and placed in wet vermiculite beds for rooting. Cuttings were maintained under the environmentally controlled conditions described previously for a period of 3–4 weeks until roots were initiated. Clonal propagules were then individually transplanted in 10 cm pots filled with the top soil/peat moss mixture described above, and grown 6 weeks. After this establishment period, shoots were trimmed to stimulate new stems development and promote plant vigor. Plants were then allowed to grow for three more weeks before their transfer to cold acclimation conditions.
At the beginning of October 2004, the clonal propagules were transferred to an unheated greenhouse located at a site near Québec City, Canada (latitude 46°47′15″, longitude 71°12′00″, altitude ≈45 m asl) for their acclimation to natural hardening conditions. The unheated greenhouse was continuously ventilated during the day to keep the inside temperature close to that of the outside. When the inside air temperature remained permanently below freezing, plants were covered with a layer of Astro-Foil™ reflective insulator (Innovative Energy Inc., Lowell, IN) to simulate snow cover. Air temperature outside and inside the greenhouse and soil temperature in pots were monitored at 30-min intervals and recorded from the end of October 2004 to mid-March 2005 using stand alone data loggers (RD-temp). Plants were assessed twice (25 January and 23 February 2005) for their freezing tolerance.
Freezing tests were conducted in a large walk-in freezer equipped with a programmable ramp temperature controller. Temperatures were decreased to −12°C by steps of 2°C using a stepwise decline described by Castonguay et al. (2009). Briefly, after a 24-h equilibration period at −2°C, temperatures were lowered by 2°C decrements using a 30-min decline period followed by a 90-min plateau at each temperature until −12°C was reached. Plants were then maintained 90 min at the −12°C test temperature and were subsequently thawed 24 h at 4°C. A group of plants used as non-stressed controls were transferred directly from the unheated greenhouse to 4°C without being exposed to the freezing stress. After thawing, shoots were clipped and plants were transferred to the environmentally controlled growth chamber conditions for regrowth. Three weeks later, shoots were individually harvested and dried at 65°C for the determination of the dry matter of regrowth. Freezing tolerance of each genotype was estimated as the ratio of regrowth of six clonal propagules exposed to a single test temperature of −12°C (T) over that of six unstressed controls (C). A ratio (T/C) value near 0 reflects almost complete sensitivity to the freezing test whereas a ratio of ~1 indicates full tolerance.
DNA extraction
Genomic DNA was extracted from individual and pooled samples from 45 genotypes from each of the cultivar Apica (ATF0) and populations (ATF2, ATF4, ATF5, and ATF6) derived from this cultivar through recurrent cycles of selection for superior freezing tolerance. Total genomic DNA was extracted using the CTAB procedure of Rogers and Bendich (1988). DNA was quantified by visual assessment by comparison with the molecular weight marker II (Roche Diagnostics, Canada) of known concentration, and diluted to 10 ng μl−1 for use in SRAP reactions.
SRAP reactions
All SRAP reactions were performed in a total volume of 25 μl in 0.2 ml PCR strips containing 2.5 μl of 10× PCR buffer, 1 μl each of 5 μM primers, 0.5 μl of 10 mM dNTP (Roche Diagnostics, Indianapolis, IN), 0.5 μl of 5′ Taq polymerase 5 U μl−1 (Inter Medico, Markham, ON, Canada) and 5 μl of 10 ng μl−1 genomic DNA (50 ng). The conditions for PCR were as follows: an initial denaturing step was performed at 94°C for 3 min followed by five cycles at 94°C for 1 min, 35°C for 1 min, 72°C for 1 min, 35 cycles at 94°C for 1 min, 50°C for 1 min, 72°C for 1 min and a final extension of 7 min at 72°C. All the reactions were performed on an Eppendorf Mastercycler ep System (Eppendorf Canada, Mississauga, ON, Canada). Twenty microliters of each reaction was loaded and run for 3 h at 70 V on a 2% agarose gel stained with ethidium bromide. DNA fragments were visualized using a UVP BioDoc-It system (UVP, Upland, CA). The nucleotide sequences of the forward and reverse primers used in this study are listed in Table 1.
Cloning and sequencing of amplified fragments
DNA fragments were recovered from agarose gels using the QIAquick gel extraction kit (QIAGEN Inc., Mississauga, ON, Canada) according to the manufacturer’s recommendation. Purified DNA was cloned into the pGEM®-T Easy Vector (Promega, Madison, WI) as recommended by the manufacturer’s instructions. Positive transformants were recovered, grown on liquid medium and submitted to a plasmid purification protocol using the QIAprep Spin Miniprep Kit (QIAGEN Inc., Mississauga, ON, Canada). Plasmid preparations were sent for bidirectional sequencing using M13 forward and reverse primers. BLASTn and BLASTx sequence homology searches were performed using the BLAST algorithm at http://www.ncbi.nlm.nih.gov of National Center for Biotechnology Information, with the program BLASTN and BLASTX.
Stastistical analyses
Chi-squared tests for differences in the distribution of cumulative SRAP polymorphisms between ATF populations and Cochran-Mantel–Haenszel Chi-square statistic for independent assortment for pairwise combinations of SRAP polymorphisms were performed with the FREQ procedure of SAS (1999). Means of groups of genotypes with contrasted freezing tolerance were compared using the Student’s t test procedure of SAS (1999).
Results
SRAP polymorphism among ATF populations
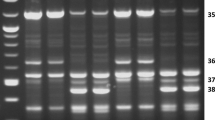
A total of 42 SRAP primer pairs combinations were used for the amplification of DNA bulk samples (genotypes) of ATF populations (Table 1). The PCR profiles were highly consistent among the recurrently selected populations with the noticeable exception of several polymorphisms that either increased or decreased in intensity in response to selection for superior tolerance to freezing as illustrated in Fig. 1a, b. Four polymorphisms that showed a clear positive response to selection were retained for further analyses (Supplementary Fig. 1). Assessment of the occurrence of the F10-R14 polymorphism within 45 genotypes of each ATF population revealed that the increase in intensity of this polymorphic fragment detected with bulk samples reflects differences in genotypic frequency within these populations (Supplementary Fig. 2). Four genotypes of ATF0 had the specific F10-R14 fragment while the number of positive genotypes increased to 7 and 13 in the ATF2 and ATF5 populations, respectively. Similar results were obtained for the three other primer pair combinations (Table 2). It is noteworthy that except for the F10-R14 polymorphism, increase in genotypic frequency of the SRAP polymorphisms was noticeable only in the later cycle of selection (ATF5) which is consistent with the observations made with bulks. Analysis of the distribution of genotypes in five categories from zero up to four polymorphisms revealed that several ATF0 genotypes did not have any of the four SRAP polymorphisms and very few genotypes combined more than one polymorphism (Table 3). Conversely, the proportion of ATF5 genotypes that did not have a SRAP polymorphism was very low while the number of those that had two or more polymorphisms markedly increased from ATF2 to ATF5. This resulted in a highly significant difference in the distribution of genotypes in groups of cumulative polymorphisms between ATF0 and ATF5. However, the distribution of the ATF2 genotypes among the five classes of polymorphisms did not significantly differ from that of ATF0.
SRAPs of bulk samples (45 genotypes/bulk) from the cultivar Apica (ATF0) and from populations ATF2, ATF4, ATF5, and ATF6, recurrently selected for superior tolerance to freezing (TF) within this cultivar. a and b show, respectively, a positive (black arrow) and a negative (open arrow) polymorphism in response to selection for superior freezing tolerance. Primer pair combinations are indicated
DNA sequence analysis of polymorphic fragment
Sequence analysis of the SRAP fragments revealed GC contents exceeding 40% in three of the four sequences. Homology search performed by the Blast program of the National Center for Biotechnology Information did not reveal any significant hit with coding sequences in gene data banks. However, it did find matches with BAC clones sequences from Medicago truncatula for the four SRAP polymorphisms (Table 4; Supplementary Table 1). Three of these homologous sequences from M. truncatula were putatively located on distinct linkage groups (data not shown).
Relationship between SRAP polymorphisms and freezing tolerance
To evaluate the link between SRAP polymorphisms and freezing tolerance, genotypes that were screened for DNA polymorphisms within each of ATF0, ATF2, and ATF5 populations were also evaluated for their tolerance to freezing. A quantitative response ranging from almost complete sensitivity to almost complete tolerance to exposure to −12°C was observed in the ATF0 and ATF5 populations (Fig. 2). However, the average ratio of regrowth in ATF5 (0.58) was greater than that of ATF0 (0.50). This was attributed to differences in the number of genotypes in the bottom quartile of tolerance (≤0.25) which decreased from ten sensitive genotypes in ATF0 down to four in ATF5. Conversely, the number of genotypes in the upper quartile of tolerance (≥0.75) increased from 7 in ATF0 up to 16 in ATF5. Based on the ranking of the two freezing tests, nine genotypes from ATF0 that showed a high sensitivity to freezing and nine genotypes from ATF5 that showed high tolerance to freezing were selected. These two groups showed highly significant differences in freezing tolerance in both tests (Table 5). These genotypes were scored for the presence of the polymorphic fragments initially identified with bulk samples from ATF populations. A highly contrasted occurrence of the polymorphisms was observed between the two groups of genotypes (Fig 3a–d). Whereas only four polymorphisms were found among the highly freezing sensitive genotypes, this number increased to 23 in the group of highly freezing tolerant genotypes (Fig. 4). Within that latter group, several genotypes had three polymorphisms or more and all genotypes had at least one polymorphism. The occurrence of the four SRAP polymorphisms shown in Fig. 3 appears to be unrelated based on their independent segregation. This was confirmed by the analysis of independent assortment of all pairwise combinations of the polymorphisms among 136 genotypes from ATF0 which showed a lack of correlation for all pairwise combinations except for the Me4R8 and F10-R14 combination (Table 6).
Genotypic distribution of freezing tolerance expressed as the mean ratio of regrowth of clones of ATF0 (N = 44) and ATF5 (N = 42) genotypes that were exposed to −12°C over respective controls that were immediately transferred to growth chambers. A ratio near 1 indicates complete tolerance while a ratio near zero indicates complete sensitivity. Each bar represents a single genotype. Mean ratio of each population is indicated
Occurrence of four SRAP polymorphisms in nine highly freezing sensitive genotypes from ATF0 and nine highly freezing tolerant genotypes from ATF5. Polymorphisms detected with bulks (45 plants) from both populations are identified in boxes. Genotypes with positive amplifications are marked with dots below each panel. Molecular weights (bp) of the four polymorphic fragments and primer pair combinations are indicated in a–d
Discussion
Bulk segregant analysis of recurrent selections
Although some improvement in winter hardiness has been achieved in cultivars released in the US in late twentieth century, the variability for winter stress tolerance remains largely untapped by plant breeding programs (Volenec et al. 2002). Tolerance to low subfreezing temperatures is the single most determinant factor that affects winter survival across a wide range of environments (Castonguay et al. 2006). The identification of linked markers that target loci with genes that confer superior adaptation to cold would greatly facilitate the selective improvement of freezing tolerance without affecting other agronomic traits. In that perspective, alfalfa TF populations that are significantly improved in their tolerance to freezing temperatures and winter survival under field conditions constitute a unique resource to identify the genetic bases of superior freezing tolerance in alfalfa.
The genetic bases of abiotic stress resistance are difficult to determine due to the quantitative nature of inheritance and large environmental effects (Miklas et al. 2006). Recurrent selections are powerful tools to study quantitative genetics relevant to agricultural traits. They are however seldom exploited in genetic studies because of the large number of cycles that are sometimes required to evoke phenotypic changes (Briggs and Goldman 2006). In the current report, we present evidence that SRAP analysis of bulk DNA samples from recurrent selections is a very effective strategy to search for DNA polymorphisms associated to quantitative traits in populations of allogamous species. BSA is an efficient genotyping approach that has been frequently used to identify markers linked to loci affecting qualitative traits but that has been seldom applied for the study of quantitative traits (Abberton et al. 2003; Quarrie et al. 1999; Serraj et al. 2009; Shashidar et al. 2005). Recently, Venuprasad et al. (2009) used BSA to identify loci associated with grain yield under stress at considerable savings of time, cost and efforts. Their success was also partly attributed to the strategic nature of the genetic material they used.
Using BSA of alfalfa TF populations, we identified several positive and negative DNA polymorphisms that occured in response to selection. This indicates that the improvement of freezing tolerance relies on both the increase in frequency of favorable alleles and the elimination of unfavorable ones within populations of alfalfa. These DNA polymorphisms likely arise from changes in allele frequency at loci affecting freezing tolerance since selection was performed within a close genetic background and was specifically targeted toward the improvement of that trait under highly controlled conditions (Castonguay et al. 2009). We thus conclude that BSA of heterogeneous populations selectively improved through several cycles of recurrent selection can facilitate the identification of markers for complex traits.
SRAP polymorphisms and freezing tolerance
We uncovered several polymorphisms between ATF populations that were indicative of selection. This is in agreement with reports that SRAP markers combine reliability and genomic abundance with high levels of polymorphisms (Yi et al. 2008). Variations in the intensity of the amplification of the polymorphic fragments between bulk samples reflected differences in genotypic frequency between ATF populations. This is an important observation that suggests that SRAP analysis of bulk DNA samples provides a near quantitative assessment of allele frequency within heterogeneous populations. Although further analyses are needed to validate this preliminary evidence, this could mean that BSA identification of DNA polymorphisms could allow inferences on a population genetic response to selection. Given the heterogeneous nature of alfalfa cultivars, Vandemark et al. (2006) suggested that real-time PCR could be used to quantify the amount of target SRAP markers present in bulk plant samples from different genetic backgrounds.
Although the SRAP method of Li and Quiros (2001) has been successfully used with BSA to narrow the search for markers tightly linked to qualitative traits (He et al. 2009; Mutlu et al. 2008; Yi et al. 2008; Zhang et al. 2009), this combination had yet to be applied to quantitative traits. In species such as alfalfa lacking genome information, anonymous SRAP markers could provide a starting point for the identification of functional genes through the search for homologous regions in model species. This is supported by the existence of significant homologies between the polymorphic SRAP fragments and BAC sequences from M. truncatula, a model species with reported synteny with diploid and tetraploid M. sativa (Choi et al. 2004; Endre et al. 2002). The M. truncatula genomic resources have been previously used to identify the physical position of functional genes in Medicago sativa (Dalmadi et al. 2008). Pursuit of a similar approach with SRAP polymorphisms associated to freezing tolerance could help find genes affecting this trait.
Independent assortment of SRAP polymorphisms is consistent with their homologies with sequences located on different chromosomes of M. truncatula (Table 4). This suggests that these polymorphisms are associated with distinct regions of the genome that affect freezing tolerance. These polymorphisms could thus be used to increase the frequency of desirable alleles and pyramid adaptive genes in alfalfa populations of high agronomic value using marker-assisted selection. The observation of a markedly higher frequency of the four SRAP polymorphisms in ATF5 as compared to ATF0 or in cold tolerant as compared to cold-sensitive genotypes strongly suggests a link between these variations in the genome and tolerance to low subfreezing temperatures. It is noteworthy that Brouwer et al. (2000), using two backcross populations of tetraploid alfalfa, identified QTLs affecting winter injury that showed additive gene action. The lack of response or slow increase in the frequency of the four SRAP polymorphisms observed in ATF2 could be due to low allele frequency in the initial population, the need to break up undesirable linkages and to recombine favorable alleles in the early cycles of selection. Regardless of the underlying genetic causes, it highlights the importance to pursue several cycles of selection in order to increase the response to selection and increase chances to identify allelic selection associated to the trait of interest.
Conclusions
The large genomes and complex genetics of open-pollinated species such as alfalfa pose enormous challenge to the identification of genes with major effects on quantitative traits like freezing tolerance (Brummer 2004). Approaches that exploit gene diversity among populations could facilitate the identification of polymorphisms that correlate with phenotypic variation (Buckler and Thornsberry 2002; Takeda and Matsuoka 2008). In this report, we show that long-term investment in the development of recurrent selections performed within heterogeneous populations of allogamous species and the subsequent probing of their genome with BSA can lead to the identification of DNA variants associated to the improvement of quantitative traits. Using this approach we identified DNA polymorphisms with potential linkage with loci that affect freezing tolerance in alfalfa. A closer analysis of four SRAP polymorphisms supports an association with cold tolerance and potential usefulness in marker-assisted selection (MAS). Evaluation of the phenotypic impact of selection based on these SRAP markers is currently underway. MAS could increase selection efficiency not only by allowing earlier selection but also by reducing population size during breeding. This would be particularly advantageous in perennial crops such as alfalfa by reducing years of field assessment (Castonguay et al. 2006). Whether these SRAP polymorphisms are present in other initial backgrounds and if they show a similar association with freezing tolerance is an important question that is currently under investigation.
References
Abberton MT, Michaelson YTPT, Bowen C, Marshall AH, Prewer W, Carlile E (2003) Bulked segregant AFLP analysis to identify markers for the introduction of the rhizomatous habit from Trifolium ambiguum into T. repens (white clover). Euphytica 134:217–222
Ariss JJ, Vandemark GJ (2007) Assessment of genetic diversity among nondormant and semidormant alfalfa populations using sequence-related amplified polymorphisms. Crop Sci 47:2274–2284
Bertrand A, Castonguay Y, Cloutier J, Couture L, Hsiang T, Dionne J, Laberge S (2009) Genetic diversity for pink snow mold resistance in greens-type annual bluegrass. Crop Sci 49:589–599
Briggs WH, Goldman IL (2006) Genetic variation and selection response in model breeding populations of Brassica rapa following a diversity bottleneck. Genetics 172:457–465
Brouwer DJ, Duke SH, Osborn TC (2000) Mapping genetic factors associated with winter hardiness, fall growth, and freezing injury in autotetraploid alfalfa. Crop Sci 40:1387–1396
Brummer EC (2004) Applying genomics to alfalfa breeding programs. Crop Sci 44:1904–1907
Buckler EI, Thornsberry JM (2002) Plant molecular diversity and applications to genomics. Curr Opin Plant Biol 5:107–111
Budak H, Shearman RC, Parmaksiz I, Dweikat I (2004a) Comparative analysis of seeded and vegetative biotypes buffalo grasses based on phylogenetic relationship using ISSRs, SSRs, RAPDs and SRAPs. Theor Appl Genet 109:280–288
Budak H, Shearman RC, Parmaksiz I, Gaussoin RE, Riordan TP, Dweikat I (2004b) Molecular characterization of Buffalograss germplasm using sequence-related amplified polymorphism markers. Theor Appl Genet 108:328–334
Castonguay Y, Laberge S, Brummer EC, Volenec JJ (2006) Alfalfa winter hardiness: a research retrospective and integrated perspective. Adv Agron 90:203–265
Castonguay Y, Michaud R, Nadeau P, Bertrand A (2009) An indoor screening method for improvement of freezing tolerance in alfalfa. Crop Sci 49:809–818
Choi HK, Kim D, Uhm T, Limpens E, Lim H, Mun JH, Kalo P, Penmetsa RV, Seres A, Kulikova O, Roe BA, Bisseling T, Kiss GB, Cook DR (2004) A sequence-based genetic map of Medicago truncatula and comparison of marker colinearity with M. sativa. Genetics 166:1463–1502
Dalmadi A, Kalo P, Jakab J, Saskoi A, Petrovics T, Deak G, Kiss GB (2008) Dwarf plants of diploid Medicago sativa carry a mutation in the gibberellin 3-β-hydroxylase gene. Plant Cell Rep 27:1271–1279
Endre G, Kereszt A, Kevei Z, Mihacea S, Kalo P, Kiss GB (2002) A receptor kinase gene regulating symbiotic nodule development. Nature 417:962–966
Ferriol M, Pico B, Nuez F (2003) Genetic diversity of a germplasm collection of Cucurbita pepo using SRAP and AFLP markers. Theor Appl Genet 107:271–282
Gulsen O, Karagul S, Abak K (2007) Diversity and relationships among Turkish okra germplasm by SRAP and phenotypic marker polymorphism. Biologia 62:41–45
He YH, Ning GG, Sun YL, Qi YC, Bao MZ (2009) Identification of a SCAR marker linked to a recessive male sterile gene (Tems) and its application in breeding of marigold (Tagetes erecta). Plant Breed 128:92–96
Jones JS, Bingham ET (1995) Inbreeding depression in alfalfa and open-pollinated crops. Plant Breed Rev 13:209–233
Jones N, Ougham H, Thomas H, Pasakinskiene I (2009) Markers and mapping revisited: finding your gene. New Phytol 183:935–966
Li G, Quiros CF (2001) Sequence-related amplified polymorphism (SRAP), a new marker system based on a simple PCR reaction: its application to mapping and gene tagging in Brassica. Theor Appl Genet 103:455–461
Limin AE, Fowler DB (1991) Breeding for cold hardiness in winter wheat: problems, progress and alien gene expression. Field Crops Res 27:201–218
Liu L, Liu G, Gong Y, Dai W, Wang Y, Yu F, Ren Y (2007) Evaluation of genetic purity of F1 hybrid ISSR, seeds in cabbage with RAPD, SRAP, and SSR markers. Hortscience 42:724–727
Michaud R, Richard C, Willemot C, Gasser H (1983) Apica alfalfa. Can J Plant Sci 63:547–549
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Nat Acad Sci USA 88:9828–9832
Miklas P, Kelly J, Beebe S, Blair M (2006) Common bean breeding for resistance against biotic and abiotic stresses: From classical to MAS breeding. Euphytica 147:105–131
Mutlu N, Boyacı F, Göçmen M, Abak K (2008) Development of SRAP, SRAP-RGA, RAPD and SCAR markers linked with a Fusarium wilt resistance gene in eggplant. Theor Appl Genet 117:1303–1312
Quarrie SA, Lazic JV, Kovacevic D, Steed A, Pekic S (1999) Bulk segregant analysis with molecular markers and its use for improving drought resistance in maize. J Exp Bot 50:1299–1306
Rogers SO, Bendich AJ (1988) Extraction of DNA from plant tissues. In: Gelvin S, Schilperoort RA (eds) Plant molecular biology manual A6. Kluwer, Dordrecht, pp 1–10
SAS Institute Inc (1999) SAS/Stat user’s guide. Cary, NC
Serraj R, Kumar A, McNally KL, Slamet-Loedin I, Bruskiewich R, Mauleon R, Cairns J, Hijmans RJ, Donald LS (2009) Improvement of drought resistance in rice, Chap 2. Adv Agron 103:41–99
Shashidar HE, Vinod MS, Sudhir GV, Sharma N, Krishnamurthy K (2005) Markers linked to grain yield using bulked segregant analysis approach in rice (Oryza sativa L.). Rice Genet Newsl 22:69–71
Sun SJ, Gao W, Lin SQ, Zhu J, Xie BG, Lin ZB (2006) Analysis of genetic diversity in Ganoderma population with a novel molecular marker SRAP. Appl Microbiol Biotechnol 72:537–543
Takeda S, Matsuoka M (2008) Genetic approaches to crop improvement: responding to environmental and population changes. Nat Rev Genet 9:444–457
Toppino L, Valc G, Rotino GL (2008) Inheritance of Fusarium wilt resistance introgressed from Solanum aethiopicum Gilo and Aculeatum groups into cultivated eggplant (S. melongena) and development of associated PCR-based markers. Mol Breed 22:237–250
Vandemark GJ, Ariss JJ, Bauchan GA, Larsen RC, Hughes TJ (2006) Estimating genetic relationships among historical sources of alfalfa germplasm and selected cultivars with sequence related amplified polymorphisms. Euphytica 152:9–16
Venuprasad R, Dalid CO, Del Valle M, Zhao D, Espiritu M, Sta Cruz MT, Amante M, Kumar A, Atlin GN (2009) Identification and characterization of large-effect quantitative trait loci grain yield under lowland drought stress in rice using bulk-segregant analysis. Theor Appl Genet. doi:10.1007/s00122-009-1168-1
Volenec JJ, Cunningham SM, Haagenson DM, Berg WK, Joern BC, Wiersma DW (2002) Physiological genetics of alfalfa improvement: past failures, future prospects. Field Crops Res 75:97–110
Yi YJ, Liu HY, Huang XQ, An LZ, Wang F, Wang XL (2008) Development of molecular markers linked to the wheat powdery mildew resistance gene Pm4b and marker validation for molecular breeding. Plant Breed 127:116–120
Zhang W, He H, Guan Y, Du H, Yuan L, Li Z, Yao D, Pan J, Cai R (2009) Identification and mapping of molecular markers linked to the tuberculate fruit gene in the cucumber (Cucumis sativus L.). Theor Appl Genet. doi:10.1007/s00122-009-1182-3
Acknowledgments
The authors acknowledge the contribution of Mrs. Josée Bourassa and Josée Michaud for their technical assistance and David Gagné for the bioinformatics search.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by C. Quiros.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Castonguay, Y., Cloutier, J., Bertrand, A. et al. SRAP polymorphisms associated with superior freezing tolerance in alfalfa (Medicago sativa spp. sativa). Theor Appl Genet 120, 1611–1619 (2010). https://doi.org/10.1007/s00122-010-1280-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-010-1280-2