Abstract
The main techniques, approaches, methods, and information products used in recent years for the identification of chemical compounds are summarized. The methodology used in target analysis has largely remained unchanged; only the identification criteria have undergone some adjustments. The scope of research in non-target analysis has been significantly expanded. In this case, the main problems lie in revealing candidates for identification. These versions are tested against typical criteria of target analysis. Effective search for suitable candidate compounds has become possible with the apearance of modern high-resolution chromatography–mass spectrometers and progress in informatics. The latter includes the development of algorithms and programs for processing chromatographic and mass spectrometric data; comparing them with reference values; and predicting mass spectra, retention parameters, and other quantities. Chemical databases enable the assessment of the prevalence of chemical compounds and, correspondingly, their potential as candidates for identification.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
The procedure of the identification of specific compounds is an integral part of chemical analysis. The fundamental principles, rules, and criteria for chemical identification were formulated in the first decade of the 21st century ([1–3] and references in these publications). Meanwhile, the methodology for target analysis (TA, determination of specified analytes according to respective analysis methods) has changed a little to the present day, with only, for instance, some adjustments made to the identification criteria [4].
A significantly more complex task is the identification of previously unknown chemical compounds within the framework of non-target analysis (NTA) [5]. The ability to address this challenge, associated with increased needs in in-depth substance analysis, has significantly expanded over the last 10−15 years. Firstly, this is due to the widespread adoption of new analytical instruments (high and ultra-high resolution chromatographs and mass spectrometers) and, secondly, the rapid advancements in informatics, demonstrated by the emergence of new algorithms, programs, databases (DB), and networks. In many cases, these circumstances contribute to a relatively rapid and reliable search for candidates for identification and gathering of evidence regarding the specific identity of analytes.
The methodology of non-target analysis is rapidly evolving, which, in turn, necessitates the regular review and preparation of new overviews in this field. In this article, we briefly summarized the results of the most significant recent works related to methods and approaches of chemical identification, primarily focusing on low-molecular-weight compounds and predominantly using mass spectrometry (MS), chromatography, and chromatography–mass spectrometry (CMS). Other relevant information, particularly related to the analysis of biological samples, environmental samples, food products, and natural compounds, can be found in the reviews concerning modern identification procedures [6–12].
GENERAL ISSUES
The primary methods of identification based on the application of (a) mass spectrometry, including tandem mass spectrometry (MS2) and high-resolution mass spectrometry (HRMS, HRMS2), (b) gas chromatography (GC) and liquid chromatography (LC), including high-performance liquid chromatography (HPLC) and ultra-high performance liquid chromatography (UPLC), and (c) ion mobility spectrometry (IMS) are listed in Table 1. Unambiguous and true identification is achieved using the first method of identification (the simultaneous determination of the analyte and the standard) [1–3]. Common reagents and chemicals are often used as analytical standards, though their purity is not always regulated. The vast market of commercially available chemical compounds supports the substantial potential of this identification method (see below). The use of authentic analytical standards (certified reference materials) would be the most reliable identification option, but the nomenclature of the standards [14] is several orders of magnitude smaller.
In metabolomics [15] and environmental analysis [16], positive results from using identification method no. 1 (Table 1) are associated with its first/highest level. Other methods only lead to candidates for identification (second level and below). However, more reliable results, termed “putatively annotated compounds” [15] or “probable structure” [16], are obtained using the second method of identification (Table 1). At the close recording conditions of the experimental and reference data, a possibility of correct identification is presumed. Therefore, this approach is frequently used and practically justified, but its rigorous substantiation requires validation for many analytes and matrices.
In recent scientific literature, particularly in the field of metabolomics, the term “annotation” [9, 15] is frequently used along with the term “identification”. Annotation refers to the process of assigning a putative formula/structure or other characteristics of a chemical compound to an analyte (analytical signal) and implies a notion of “putative identification.”
Quantitative measures of trueness (reliability) of identification can be established. In the case of reliable identification (method no. 1) of a particular analyte, it is logical to consider the corresponding trueness as 100%. Other identification methods typically yield lower values, which can be assessed using various probabilistic models of the discussed procedure [1–3]. The trueness of a specific method or an identification procedure for a series of analyses or a group of analytes is measured by the fraction of truely identified positive (TP) results (a conventional indicator of qualitative analysis [2]).
Earlier, the concept of an identification point (IP) was proposed—a single characteristic (a single value of a measured quantity, a single property) of an analyte used for identification by comparing it with the properties of a known reference compound [2, 3]. For reliable target identification, several IPs are required (matching chromatographic retention parameters, ion masses, and intensities of mass peaks), some of which (exact masses) are more significant. The IP concept has recently been extended to non-target analysis carried out by various HPLC–HRMS methods [6, 17]. The identification points are aggregated as integer scores [6]. A fractional scale for estimating the match of mass spectra and chromatograms has also been introduced, including a comparison of the experimental data with the corresponding calculated values of ion masses and retention times (RTs) [17].
TARGET ANALYSIS: REGULATORY IDENTIFICATION CRITERIA
The identification of predefined analytes is performed using analytical methods, many of which were developed under guidelines from international or national regulatory bodies. Updated summaries of normative chromatographic and mass spectrometric identification criteria have been published recently [4]. Subsequently, new guidelines have emerged from the U.S. Food and Drug Administration (FDA, 2020 [18]), the European Commission (EC, pesticides, 2021 [19]), and the World Anti-Doping Agency (2023 [20]). The latter two documents do not contain substantial changes in criteria compared to their previous editions [4].
In the updated FDA recommendations [18], similar to the previous EC document [4], universal maximum deviations in relative peak intensities (±30%) from the corresponding values for reference mass spectra have been introduced. This deviation is irrespective of the type of the mass spectrum (single mass analysis, MS1, with a minimum of three ions; MS2, with two or more fragment ions) and scan type (recording individual ions/transitions or full scans). For HRMS, the criteria include a mass window (±5 ppm) for two ions. Similarity in intensities is not required, as matching exact masses holds much greater importance in identification than peak intensity similarity [21], as in previous criteria. Interestingly, the explicit inclusion of a commonly used chromatographic criterion, such as the maximum deviation from the known retention times, seems to have disappeared from the discussed guideline. However, there is a requirement for the analyte peaks to match across all mass chromatograms within a given chromatographic run [18].
The mentioned regulatory/standardized criteria, tested in specialized studies, have generally proven effective but may still be improved [22]. For instance, 169 veterinary drugs were detected in egg products using UPLC–MS/MS with a triple quadrupole mass analyzer (QqQ) and the influence of the number of recorded fragment ion peaks on the rates of false positive (FP) and false negative (FN) identification results was revealed. The account of two recommended fragments resulted in a low level of FP (observed in <10% of analyses) for 95% of the analytes, while recording three fragments, feasible in modern instruments, ensured this low FP level for 99% of the analytes. Similarly, a relatively high FN rate (more than 10% of analyses) was significantly reduced (from 25–45 to 10–12% of analytes) when three transitions were recorded instead of two ones [22].
NON-TARGET ANALYSIS
A pronounced current trend is the shift of chemical analytical studies towards non-target (NTA) [5]. Identifying unknown components of a substance in the context of NTA, even when these components are detected, poses a much more challenging task than confirming or disproving the identity of the predefined analytes in target analysis (as discussed above). Unlike target analysis, which usually involves the first, and more rarely the second method of identification (Table 1), all the discussed methods are applicable within NTA. However, their implementation is rather complex and often leads to ambiguous results.
Co-analysis with analytical standards (method no. 1, Table 1) is feasible but unproductive if little is known about the nature of the analyte, making it challenging to select reference substances. Reference data (method no. 2) might be absent or might pertain to dissimilar experimental conditions. Wide application of HRMS (method no. 3) contributes to addressing challenges in NTA. However, as the molecular mass of the analyte increases, the determination of its sole molecular formula, moreover its structure, becomes more complex. Another area of progress in NTA involves the rapid development of algorithms predicting mass spectra and other characteristics (method no. 4) and also predicting structures based on this data (method no. 5). The results produced by these software programs compensate for the lack of reference data from experimental sources and eliminate the need for the “manual” interpretation of spectra.
The identification methods in NTA are illustrated in Fig. 1 and are considered in detail below, starting with the concept of chemical space, which represents the set (subset) of chemical compounds as potential results of NTA. The details and interpretations not covered in the article can be found in publications [6, 9, 11, 24–27].
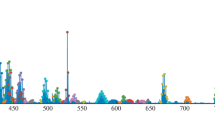
Identification of compound A in the context of non-target analysis. Satisfactory matching of the tandem mass spectrum (bottom left corner) corresponding to peak 15 in the chromatogram with the reference spectra of compound A (top row), including the in silico spectrum, is observed. The distinction from the second candidate compound (B, top right corner) is much more significant. The retention time t is also much closer to the reference value for compound A (tA). Using high-resolution mass spectrometry, the empirical formula of the unknown analyte CxHyNzOt was determined, and significantly more information was discovered for analyte A than for compound B; the Annotation Record Count is the size of the information dossier in the PubChem database [23] (see below).
Chemical space. In general, within the test samples, an unknown component can potentially be practically any known (“unknown known”) or even unknown (“unknown unknown”) compound with varying probability. The existence of an enormous number (1011 and much more, Table 2) of individual compounds, the synthesis pathways for which are theoretically known, is conceivable. More defined information is available for compounds that could be synthesized on demand. When combined with reagents and chemicals supplied from storage, the total number of the available compounds seems to approach around 1 billion (Table 2), although this is challenging to verify.
The actual chemical space appears to be limited to approximately ~200 million individual compounds and their simple mixtures (substances) (Table 2). Within this, there exist numerous rare compounds, likely not present in common analytical samples, such as human and animal biosamples, food products, plant materials, and environmental samples. Various subsets/subspaces of the most prevalent compounds (up to ~1–2 million) are included in specialized databases and also listed in Table 2. Under otherwise equal conditions, focus is typically directed toward the presence of prevalent/popular and adequately studied compounds in the analyzed samples, such as primary components of biological samples (metabolites), chemical products, frequently used reagents, and products derived from their transformations in the environment, among others. Thus, among the analytes, compounds that have previously been detected in the analyzed samples are predominant or may predominate. The mathematical equivalent of this assertion is Bayesian statistics, which is good for simple tests [2, 3] but is also applicable to modern identification methods [38].
The measures of prevalence/popularity essentially represent the relative volume of prior information (metadata) about a given compound and the frequency of references to it. Consequently, they are effectively estimated using the available chemical databases (Table 3). Thus, if two versions of an analyte, exhibiting similar mass spectra and other characteristics, significantly differ in their information indicators, the more popular compound should be considered the primary candidate for identification, and the corresponding hypothesis should be tested first. It is logical to consider any non-zero value of such an indicator as a threshold when considering a possibility of the compound presence in an analyzed sample. Similar conclusions can be drawn based on the simple presence or absence of a compound in a specialized database.
The measures of prevalence/popularity of chemical compounds are considered in addition to the results of chemical analysis. The fact of the prevalence of a particular suitable analyte by itself cannot be regarded a strict evidence of its presence in the sample. However, it is highly crucial in supporting or rejecting an identification hypothesis and in the primary selection of analytical standards in obtaining comprehensive evidence for a specific identity.
Table 3 shows that, among the considered indicators of popularity, larger databases like ChemSpider and PubChem are more frequently used than the other ones, and they are freely accessible, which is particularly important in practice. However, it is challenging to categorize them as curated databases, because they are compiled by computational means from other electronic data sets, which might make them incomplete and prone to inaccuracies. It is regrettable that the most comprehensive curated database, Chemical Abstracts, which holds verified chemical information and had no competitors in the era of paper-based informatics, is currently only accessible through commercial online channels.
The presence of a specific compound in a specialized database (lower part of Table 2) implies the potential for its detection in samples or matrices of a corresponding type. When there is information about the potential composition of the matrix, NTA takes the status of the determination of “anticipated/suspected compounds” (ASC) [5]. Therefore, compiling summaries of the suspected analytes [32, 56], which have certain chances of being detected and identified, significantly simplifies the work of analysts.
Mass spectral libraries. The use of computerized mass spectral libraries is one of the earliest and still the most prevalent methods for the automated identification [2, 3, 57, 58]. Identification is especially effective for volatile analytes and electron ionization mass spectra (EI-MS1), achieving a trueness of approximately 80%. The two largest libraries are NIST 20 (with 350 643 spectra and 306 869 compounds) and Wiley Registry 12th (with 817 290 spectra and 668 435 compounds) [5]. These data sets also include retention index (RI) guides for gas chromatography (GC-RI). Many issues regarding the creation and use of such libraries have been resolved, but certain problems still remain open [59–61].
The history of tandem mass spectral libraries (MS/MS spectra) is considerably shorter, and they are far from being comprehensive [57, 58]. It is believed that their use allows the identification of only a few percent of metabolites and other low-molecular-weight compounds [62, 63]. The type of MS/MS spectra varies widely and depends on the instrument type (QqQ; triple quadrupole mass analyzer, Q-TOF; quadrupole-time-of-flight, Orbitrap), collision energies, and some other factors [3, 62, 64]. This significantly affects the true positive rate (TP) when searching these libraries. Therefore, alongside expanding these libraries, standardization of conditions for their creation and enhancement of spectral data quality are necessary [58, 62]. This is particularly crucial because spectral libraries serve as a basis for selecting training sets of spectra in in silico prediction procedures (in silico spectra, see below).
The largest MS2 spectral libraries are listed in Table 4; information about other collections can be found in [57, 58]. These libraries can be conveniently divided into commercial and freely accessible ones, where the latter may be open for users to contribute and annotate their spectra (repositories, such as GNPS, see Table 4). When laboratories specialize in particular analytes, it is convenient to use libraries (in-house or inter-laboratory) focused on specific classes and groups of compounds [58, 68–70]. Some libraries of tandem mass spectra contain reference information about retention times and collision cross sections (see below) [58, 71].
Several ways to enhancing libraries and leading to improved identification have been noted in the literature. For example, it has been proposed to expand libraries by including spectra corresponding to new collision energies; these data are obtained by interpolating the existing spectra assigned to other collision energy values [72]. Another perspective involves using fragments from all precursor ions of a given analyte [71].
Modernization of library searches. A comparison of experimental mass spectra with reference spectra results in ranking the latters based on their similarity values. The most similar reference spectra (first-ranked computer responses) correspond to the primary candidates for identification. Candidates in the first rank are more often considered in statistics for correct results (as a percentage of true positive matches).
Conventionally, the cosine function (dot product function) is used as a measure of similarity [2, 3], although it does not exclude the appearance of false identification candidates. One way to increase the efficiency of library searches and expand their use is to introduce improved measures of similarity between the experimental and reference mass spectra. This is important for tandem mass spectra because of their insufficient reproducibility, the presence of background peaks, the multiple nature of precursor ions, the absence of some fragment ions, and their limited quantity. Several similarity measures, which complement or substitute the cosine function, have been discussed in publications [12, 73].
In recent studies, attention has been directed towards not only ion mass similarities but also towards matching mass differences (neutral losses), enabling searches for structurally similar compounds, for example in [12]. The corresponding similarity metric is a modified cosine function that additionally takes into account precursor and fragment mass differences in the compared spectra [74]. Another new promising approach involves using spectral entropy in measures of tandem mass spectra similarity [75]. There was also a proposal to consider variability in high-resolution mass spectra when comparing them [76].
Enumerating various innovations, we mention that the use of deep neural networks helps to improve the results of conventional searches in electron ionization mass spectra EI-MS1 libraries [77]. This approach is also beneficial for comparing tandem mass spectra [78].
Molecular networks [79, 80] refer to graphs representing subsets of chemical compounds linked together, usually based on the similarity of their MS2 spectra (Fig. 2). In the nodes of the graph, there are masses of ions (precursor ions) from different compounds, and each of these nodes may correspond to one or several similar spectra. The edges of the graph eventually connect pairs of structurally similar compounds whose mass spectra are very close. The molecular network coexists with a mass spectral library, allowing for the identification of “unknown knowns.” If a node embedded in the network relates to a new compound for the information system, it can be identified based on the mass differences between the ions of that analyte and a linked known compound, considering the structure of the latter.
Fragment of the molecular network (adapted from [80]). The numbers indicate the masses (Da) of the precursor ions of each compound. The mass difference of 611.161 and 595.166 corresponds to an oxygen atom, which, in consideration of the similarity of the mass spectra, allows the unknown analyte (“Unknown”) to be regarded as an oxidation product of the known compound (“Known”).
The most well-known molecular network in the field of natural compounds is the Global Natural Products Social Molecular Networking (GNPS, see Table 4), which represents a part of a comprehensive information system. This system comprises mass spectral libraries, a repository for these spectra, and various software tools for data processing and user–system interaction, among other components. The methodology of networks is combined with the potentialities of group identification and the use of predicted mass spectra [81, 82]. It is applicable to electron ionization mass spectrometry as well [83]. The methods of constructing networks and their variations are continually being improved [12, 84–87].
Prediction of mass spectra [10, 12, 63, 88]. Computer methods for predicting mass spectra (in silico spectra) enable obtaining data that substitutes experimental reference mass spectra, which is particularly crucial for MS2. Various prediction algorithms are typically based on machine learning using a training set of known mass spectra and/or fragmentation rules (heuristic information). Well-known programs include CFM-ID, MetFrag, and MS-FINDER. Predicting tandem mass spectra of compounds with regular structures, such as lipids, whose ions undergo fragmentation according to the same rules, is easier. New prediction methods and corresponding computer programs are continuously emerging (see, for instance, [36, 89–91]), some of which enable the assessment of the correctness level of a conditional identification (annotation) [92].
In silico mass spectra, serving as reference comparison spectra alone, are far from being a panacea, and the results of corresponding library searches depend on various factors [93]. The proportion of true first-rank answers obtained in most significant instances can reach ~50–70% and sometimes be higher than this level [48, 63, 94, 95]. Higher success rates are observed in combination with other identification methods and with incorporating prior information, such as compound prevalence indicators (as mentioned above).
A possibility of predicting mass spectra, including tandem spectra, using quantum chemical calculations, is being investigated [96, 97].
Structural interpretation of mass spectra [10, 12, 63, 88]. Corresponding programs based on machine learning solve the task opposite to predicting mass spectra: they derive structural fragment variants and overall candidate structures based on the mass spectrum of an unknown analyte. Consider, for example, the widely known program CSI:FingerID, which has been integrated into the SIRIUS software. The latter, based on precursor ion masses, isotopic patterns, and MC2 spectra, selects the most probable candidate compounds from those included in chemical databases [98]. Subsequently, the MAD HATTER software was created for the same purposes, enabling over 70% accurate identifications in a test dataset of tandem spectra [55]. Other new programs for the automatic extraction of structural data from mass spectra are developed [36, 99, 100]. Overall, there is high significance attributed to deep learning algorithms applied to solving this task [11, 101]. However, the potentialities of the manual interpretation of tandem mass spectra are far from being exhausted [102, 103].
Retention parameters in chromatography. The combined use of mass spectrometry and chromatography is a requirement of regulatory documents in TA and a condition for reliable identification in NTA.
Experimental retention indices (RIs) have been long and successfully used in gas chromatography (GC). They are now substantially represented in databases such as NIST (139 693 compounds [5]) and other data summaries [2, 3, 104]. Most practically significant volatile compounds are covered by this data. Nevertheless, new compounds are emerging suitable for determination by GC, such as volatile derivatives of low-volatile compounds, which necessitates information about their reference RIs. Corresponding predictions are performed by various machine-learning approaches and some other methods [104–106].
The concept of retention indices (RIs) is less important for liquid chromatography (HPLC/UPLC); in this context, retention is more often characterized by retention times (RTs). There are increasingly large collections of experimental retention times available, for instance, for 80 000 compounds in the METLIN information system [107]. These characteristics are determined under reversed-phase liquid chromatography conditions.
The experimental data sets serve as training samples for predicting retention times using machine-learning methods (as reviewed in [6, 10, 108]). Molecular descriptors are used as structural variables correlated with retention times (“quantitative structure-retention relationship”). Reversed-phase and hydrophilic interaction liquid chromatography (HILIC) columns are considered separately [108, 109]. Various errors in modern retention time prediction mostly fall within the range of ~10 s–1 min [10, 108–112]. Such accuracy of predictions might not be sufficient to capture subtle differences in the analyte structure, but it is suitable for eliminating many identification candidates. For example, 68% of candidate compounds predicted from the mass spectra of blood plasma components did not match the predicted retention time estimations: they were outside the range (retention time ±1 min) [109]. Due to the relatively low reproducibility of retention times and their strong dependence on the experimental conditions, recalculating the reference sets of retention times is crucial in adapting them to the current experiment [10, 113].
Ion mobility spectrometry (IMS) [114] is a method that involves the separation of ions based on their mobility in the gas phase and can be combined with MS and LC–MS. IMS–MS and LC–IMS–MS instruments with various technical (physical) implementations of ion mobility are manufactured by major mass spectrometer manufacturers.
The ion mobility stage enhances the separation of analytes, which is particularly evident for mixtures of isomers or isobaric compounds. Separation depends on the collision cross-section (CCS), which is an individual characteristic of chemical compounds and is suitable for their identification. Values of this parameter, both experimentally determined and calculated, are tabulated in several databases [12, 112, 114, 115], and can be used as reference data in combination with mass spectrometric and chromatographic information [6, 12].
A combination of identification techniques is necessary to enhance its reliability. The simultaneous application of MS1, MS2, U/HPLC, and ion mobility is referred to as a “four-dimensional” approach to identification [12]. In an ideal situation, with high-quality (accurate) experimental and reference data, the recognition criteria for an analyte—slight differences in precursor ion masses, retention values (RTs), collision cross-sections (CCSs), and high similarity scores of MS2 spectra—should be met by only one candidate compound. In reality, different reference data can lead to conflicting and ambiguous results. To select the best version and further verify it, one can assess the degree of data matching by counting the identification points in their numerical representation [6, 17].
Program packages (fusion tools). It is convenient for analysts if different software required for the identification of nontarget analytes is integrated into a single software package (either on the instrument’s computer or an accessible website). This integration allows combine searches in mass spectral libraries, electronic collections of retention indices (RIs) and collision cross-sections (CCSs), and chemical databases with efficient evaluations of these characteristics. In a review [10], eight such packages were mentioned, the components of which have been partially discussed earlier. In the analysis of complex samples, data processing software for chromatography and mass spectrometry and the extraction of corresponding analytical signals are critically important [116]. Overall, there are hundreds of software products described in the literature aimed at transforming experimental data and identifying analytes based on mass spectra and chromatographic parameters. Specialized programs were developed to navigate this field [117].
Comparisons. Conventional practices in analytical chemistry, including chemical identification [2, 3], involve comparative experiments assessing the professional level and quality of laboratory work, the efficiency of different methods and analytical approaches, the suitability of analytical standards, and more. The key inter- and intra-laboratory comparisons in recent years related to the identification of components in provided samples as unknown samples or the analysis of the data are summarized in Table 5. The main conclusion drawn from these works indicates the far from complete reproducibility of the results across different laboratories (nos. 1 and 8) and inferences made using different LC–MS configurations and data acquisition modes within a single laboratory (no. 7, as well as [123]), the results of applying different computer programs (nos. 2 and 4), and different measures of similarity between the mass spectra (no. 10). However, the identification outputs proved to be quite satisfactory (nos. 6 and 9), and they improve with additional information about the expected sample composition and the prevalence of analytes (nos. 1, 3, 5).
CONCLUSIONS: PARADIGM SHIFT AND GENERATIONAL CHANGE
Summarizing the above, we can argue that the main trend in research in recent years involves the development of methodologies for searching candidate compounds (conditionally identified or annotated compounds as a result) within the realm of NTA. If a definitive identification according to typical regulatory criteria is required, having a list of candidate compounds, it is advisable to (a) use analytical standards or, less reliably and requiring validation, (b) record mass spectra and chromatograms (as well as ion mobility spectra) under the conditions as close as possible to those in which the most high-quality/accurate reference data were determined. The thematic transition from the development of principles, rules, and criteria for identification to the methodology of searching for candidate compounds represents, in our view, a paradigm shift in the objectives and content of chemical-analytical research.
Changes of this kind are manifested, firstly, in the emergence and widespread use of new analytical instruments (such as high-resolution chromatography-mass spectrometers, among others) and, secondly, in the substantial amplification of the role of informatics. The latter encompasses the development of numerous machine learning-based automated search programs for candidate identification, expansion of mass spectral libraries, electronic collections of other reference data, and chemical databases. This trend reflects the integration of a new generation of information technology experts into this field. Thus, the transition of generations of instruments and experts and the paradigm shift are interdependent and respond to the broad requirements for detailed substance analysis.
The changing era is not without its challenges. Specifically, certain publications, notably Chemical Abstracts, which previously contained verified information about most known chemical compounds, did not gain open access upon transition into an electronic form. This can be regarded as a loss in the field of informatics. Moreover, many experienced chemists from recent generations have directly worked with substances and can provide significant insights about a sample composition by its color, odor, crystal form, solubility, transparency, basic tests, etc. The results are crucial in the analysis of complex unknown substances. The identification of individual compounds by chromatography–mass spectrometry (CMS) techniques is another component of such work. As demonstrated earlier, experts well-versed in informatics play an important role in these endeavors. In our opinion, the best results are achieved when their activities are combined with the work of traditional chemists.
REFERENCES
Milman, B.L., TrAC, Trends Anal. Chem., 2005, vol. 24, no. 6, p. 493. https://doi.org/10.1016/j.trac.2005.03.013
Milman, B.L., Vvedenie v khimicheskuyu identifikatsiyu (Introduction into Chemical Identification), St. Petersburg: VVM, 2008.
Milman, B.L., Chemical Identification and Its Quality Assurance, Berlin: Springer, 2011.
Milman, B.L. and Zhurkovich, I.K., Anal. Kontrol’, 2020, vol. 24, no. 3, p. 164. https://doi.org/10.15826/analitika.2020.24.3.003
Milman, B.L. and Zhurkovich, I.K., J. Anal. Chem., 2022, vol. 77, no. 5, p. 537.
Monge, M.E., Dodds, J.N., Baker, E.S., Edison, A.S., and Fernandez, F.M., Annu. Rev. Anal. Chem., 2019, vol. 12, p. 177.
Molyneux, R.J., Beck, J.J., Colegate, S.M., Edgar, J.A., Gaffield, W., Gilbert, J., Hofmann, T., McConnell, L.L., and Schieberle, P., Pure Appl. Chem., 2019, vol. 91, no. 8, p. 1417. https://doi.org/10.1515/pac-2017-1204
Nash, W.J. and Dunn, W.B., TrAC, Trends Anal. Chem., 2019, vol. 120, p. 115324. https://doi.org/10.1016/j.trac.2018.11.022
Place, B.J., Ulrich, E.M., Challis, J.K., Chao, A., Du, B., Favela, K., Feng, Y.L., Fisher, C.M., Gardinali, P., Hood, A., Knolhoff, A.M., McEachran, A.D., Nason, S.L., Newton, S.R., Ng, B., Nuñez, J., Peter, K.T., Phillips, A.L., Quinete, N., Renslow, R., Sobus, J.R., Sussman, E.M., Warth, B., Wickramasekara, S., and Williams, A.J., Anal. Chem., 2021, vol. 93, no. 49, p. 16289. https://doi.org/10.1021/acs.analchem.1c02660
Tian, Z., Liu, F., Li, D., Fernie, A.R., and Chen, W., Comput. Struct. Biotechnol. J., 2022, vol. 20, p. 5085. https://doi.org/10.1016/j.csbj.2022.09.004
De Jonge, N.F., Mildau, K., Meijer, D., Louwen, J.J., Bueschl, C., Huber, F., and Van der Hooft, J.J., Metabolomics, 2022, vol. 18, no. 12, p. 103. https://doi.org/10.1007/s11306-022-01963-y
Cai, Y., Zhou, Z., and Zhu, Z.J., TrAC, Trends Anal. Chem., 2023, vol. 158, 116903. https://doi.org/10.1016/j.trac.2022.116903
ZINC20. https://zinc20.docking.org. Accessed March 18, 2023.
Reference Materials. http://www.sigmaaldrich.com/RU/en/products/analytical-chemistry/reference-materials. Accessed March 18, 2023.
Sumner, L.W., Amberg, A., Barrett, D., Beale, M.H., Beger, R., Daykin, C.A., Fan, T.W.M., Fiehn, O., Goodacre, R., Griffin, J.L., Hankemeier, T., Hardy, N., Harnly, J., Higashi, R., Kopka, J., Lane, A.N., Lindon, J.C., Marriott, P., Nicholls, A.W., Reily, M.D., Thaden, J.J., and Viant, M.R., Metabolomics, 2007, vol. 3, p. 211. https://doi.org/10.1007/s11306-007-0082-2
Schymanski, E.L., Jeon, J., Gulde, R., Fenner, K., Ruf, M., Singer, H.P., and Hollender, J., Environ. Sci. Technol., 2014, vol. 48, no. 4, p. 2097. https://doi.org/10.1021/es5002105
Alygizakis, N., Lestremau, F., Gago-Ferrero, P., Gil-Solsona, R., Arturi, K., Hollender, J., Schymanski, E.L., Dulio, V., Slobodnik, J., and Thomaidis, N.S., TrAC, Trends Anal. Chem., 2023, vol. 159, p. 116944. https://doi.org/10.1016/j.trac.2023.116944
Methods, Method Verification and Validation. https://www.fda.gov/media/73920/download. Accessed March 19, 2023.
Analytical Quality Control and Method Validation Procedures for Pesticide Residues Analysis in Food and Feed. https://www.eurl-pesticides.eu/userfiles/file/EurlALL/SANTE_11312_2021.pdf. Accessed March 19, 2023.
Minimum Criteria for Chromatographic–Mass Spectrometric Confirmation of the Identity of Analytes for Doping Control Purposes. http://www.wada-ama.org/sites/default/files/2023-02/td2023idcrv1.1_eng_final.pdf. Accessed March 19, 2023.
Milman, B.L. and Zhurkovich, I.K., Mass Spectrom. Lett., 2018, vol. 9, no. 3, p. 73. https://doi.org/10.5478/MSL.2018.9.3.73
Lehotay, S.J., Anal. Bioanal. Chem., 2022, vol. 414, no. 1, p. 287. https://doi.org/10.1007/s00216-021-03380-x
PubChem. https://pubchem.ncbi.nlm.nih.gov. Accessed March 20, 2023.
Duhrkop, K., Nothias, L.F., Fleischauer, M., Reher, R., Ludwig, M., Hoffmann, M.A., Petras, D., Gerwick, W.H., Rousu, J., Dorrestein, P.C., and Bocker, S., Nat. Biotechnol., 2021, vol. 39, no. 4, p. 462. https://doi.org/10.1038/s41587-020-0740-8
Rey-Stolle, F., Dudzik, D., Gonzalez-Riano, C., Fernandez-Garcia, M., Alonso-Herranz, V., Rojo, D., Barbas, C., and Garcia, A., Anal. Chim. Acta, 2022, vol. 1210, p. 339043. https://doi.org/10.1016/j.aca.2021.339043
Caballero-Casero, N., Belova, L., Vervliet, P., Antignac, J.P., Castano, A., Debrauwer, L., Lopez, M.E., Huber, C., Klanova, J., Krauss, M., Lommen, A., Mol, H.G.J., Oberacher, H., Pardo, O., Price, E.J., Reinstadler, V., Vitale, C.M., Van Nuijs, A.L.N., and Covaci, A., TrAC, Trends Anal. Chem., 2021, vol. 136, p. 116201. https://doi.org/10.1016/j.trac.2021.116201
Misra, B.B., New software tools, databases, and resources in metabolomics: updates from 2020, Metabolomics, 2021, vol. 17, no. 5, p. 49. https://doi.org/10.1007/s11306-021-01796-1
Milman, B.L. and Zhurkovich, I.K., TrAC, Trends Anal. Chem., 2017, vol. 97, p. 179. https://doi.org/10.1016/j.trac.2017.09.013
CAS. https://www.cas.org/about/cas-content. Accessed March 20, 2023.
ChemSpider. http://www.chemspider.com. Accessed March 20, 2023.
CompTox Chemistry Dashboard. https://comptox.epa.gov/dashboard. Accessed March 20, 2023.
NORMAN-SLE. http://www.norman-network.com/?q=node/236. Accessed March 20, 2023.
The Human Metabolome Database (HMDB). https://hmdb.ca. Accessed March 20, 2023.
Sorokina, M. and Steinbeck, C., J. Cheminf., 2020, vol. 12, no. 1, p. 20. https://doi.org/10.1186/s13321-020-00424-9
FooDB. https://foodb.ca/compounds. Accessed March 20, 2023.
O’Shea, K. and Misra, B.B., Metabolomics, 2020, vol. 16, no. 3, p. 35. https://doi.org/10.1007/s11306-020-01657-3
Banimfreg, B.H., Shamayleh, A., and Alshraideh, H., Metabolites, 2022, vol. 12, no. 10, p. 1002. https://doi.org/10.3390/metabo12101002
Ludwig, M., Doctoral Dissertation, Jena: Friedrich-Schiller-Univ., 2020. http://www.db-thueringen.de/ servlets/MCRFileNodeServlet/dbt_derivate_00050369/dissludwig.pdf. Accessed March 20, 2023.
Milman, B.L. and Konopelko, L.A., Fresenius’ J. Anal. Chem., 2000, vol. 367, p. 621. https://doi.org/10.1007/s002160000426
Milman, B.L. and Kovrizhnych, M.A., Fresenius’ J. Anal. Chem., 2000, vol. 367, p. 629. https://doi.org/10.1007/s002160000427
Milman, B.L., Anal. Chem., 2002, vol. 74, no. 7, p. 1484. https://doi.org/10.1021/ac010611p
Milman, B.L., J. Chem. Inf. Model., 2005, vol. 45, no. 5, p. 1153. https://doi.org/10.1021/ci049716u
Little, J.L., Cleven, C.D., and Brown, S.D., J. Am. Soc. Mass Spectrom., 2011, vol. 22, no. 2, p. 348.https://doi.org/10.1007/s13361-010-0034-3
Little, J.L., Williams, A.J., Pshenichnov, A., and Tk-achenko, V., J. Am. Soc. Mass Spectrom., 2012, vol. 23, no. 1, p. 179. https://doi.org/10.1007/s13361-011-0265-y
Ridder, L., Van der Hooft, J.J.J., and Verhoeven, S., Mass Spectrom., 2014, vol. 3, no. 2, p. 0033. https://doi.org/10.5702/massspectrometry.S0033
Woldegebriel, M. and Vivo-Truyols, G., Anal. Chem., 2016, vol. 88, no. 19, p. 9843. https://doi.org/10.1021/acs.analchem.6b03026
Ruttkies, C., Schymanski, E.L., Wolf, S., Hollender, J., and Neumann, S., J. Cheminf., 2016, vol. 8, no. 1, p. 3. https://doi.org/10.1186/s13321-016-0115-9
Blaženović, I., Kind, T., Torbašinović, H., Obrenović, S., Mehta, S.S., Tsugawa, H., Wermuth, T., Schauer, N., Jahn, M., Biedendieck, R., Jahn, D., and Fiehn, O., J. Cheminf., 2017, vol. 9, p. 32. https://doi.org/10.1186/s13321-017-0219-x
McEachran, A.D., Chao, A., Al-Ghoul, H., Lowe, C., Grulke, C., Sobus, J.R., and Williams, A.J., Metabolites, 2020, vol. 10, no. 6, p. 260. https://doi.org/10.3390/metabo10060260
Milman, B.L., Ostrovidova, E.V., and Zhurkovich, I.K., J. Anal. Chem., 2021, vol. 76, p. 1477. https://doi.org/10.1134/S1061934821130086
Milman, B.L. and Zhurkovich, I.K., Analitika, 2020, vol. 10, no. 6, p. 464. https://doi.org/10.22184/2227-572X.2020.10.6.464.469
Milman, B.L. and Zhurkovich, I.K., Molecules, 2021, vol. 26, no. 8, p. 2394. https://doi.org/10.3390/molecules26082394
Schymanski, E.L., Kondic, T., Neumann, S., Thiessen, P.A., Zhang, J., and Bolton, E.E., J. Cheminf., 2021, vol. 13, no. 1, p. 19. https://doi.org/10.1186/s13321-021-00489-0
Milman, B.L. and Zhurkovich, I.K., Analitika, 2023, vol. 13, no. 1, p. 56. https://doi.org/10.22184/2227-572X.2023.13.1.56.59
Hoffmann, M.A., Kretschmer, F., Ludwig, M., and Bocker, S., Metabolites, 2023, vol. 13, no. 3, p. 314. https://doi.org/10.3390/metabo13030314
Cave, J.R., Parker, E., Lebrilla, C., and Waterhouse, A.L., J. Agric. Food Chem., 2019, vol. 67, no. 48, p. Ñ. 13318. https://doi.org/10.1021/acs.jafc.9b04384
Milman, B.L. and Zhurkovich, I.K., TrAC, Trends Anal. Chem., 2016, vol. 80, p. 636. https://doi.org/10.1016/j.trac.2016.04.024
Bittremieux, W., Wang, M., and Dorrestein, P.C., Metabolomics, 2022, vol. 18, no. 12, p. 94. https://doi.org/10.1007/s11306-022-01947-y
Samokhin, A., Sotnezova, K., and Revelsky, I., Eur. J. Mass Spectrom., 2019, vol. 25, no. 6, p. 439. https://doi.org/10.1177/1469066719855503
Chua, C.K., Lv, Y., Zhao, W., Ren, Y., and Zhang, H.J., Int. J. Mass Spectrom. Ion Processes, 2020, vol. 451, 116321. https://doi.org/10.1016/j.ijms.2020.116321
Samokhin, A.S. and Matyushin, D.D., Rapid Commun. Mass Spectrom., 2023, vol. 37, no. 3, e9437. https://doi.org/10.1002/rcm.9437
Oberacher, H., Sasse, M., Antignac, J.P., Guitton, Y., Debrauwer, L., Jamin, E.L., Schulze, T., Krauss, M., Covaci, A., Caballero-Casero, N., Rousseau, K., Damont, A., Fenaille, F., Lamoree, M., and Schymanski, E.L., Environ. Sci. Eur., 2020, vol. 32, p. 43. https://doi.org/10.1186/s12302-020-00314-9
Krettler, C.A. and Thallinger, G.G., Briefings Bioinf., 2021, vol. 22, no. 6, bbab073. https://doi.org/10.1093/bib/bbab073
Milman, B.L., TrAC, Trends Anal. Chem., 2015, vol. 69, p. 24. https://doi.org/10.1016/j.trac.2014.12.009
Montenegro-Burke, J.R., Guijas, C., and Siuzdak, G., in Computational Methods and Data Analysis for Metabolomics, Li, S., Ed., New York: Humana, 2020, p. 149. https://doi.org/10.1007/978-1-0716-0239-3_9
m/zCloud. https://www.mzcloud.org. Accessed March 22, 2023.
MassBank. https://massbank.eu/MassBank/Contents. Accessed March 22, 2023.
Lee, S., Hwang, S., Seo, M., Shin, K.B., Kim, K.H., Park, G.W., Kim, J.Y., Yoo, J.S., and No, K.T., Phytochemistry, 2020, vol. 177, 112427. https://doi.org/10.1016/j.phytochem.2020.112427
Davidsen, A., Mardal, M., Linnet, K., and Dalsgaard, P.W., PloS One, 2020, vol. 15, no. 11, p. e0242224. https://doi.org/10.1371/journal.pone.0242224
Li, Y., Zhu, W., Xiang, Q., Kim, J., Dufresne, C., Liu, Y., Li, T., and Chen, S., Int. J. Mol. Sci., 2023, vol. 24, no. 3, p. 2249. https://doi.org/10.3390/ijms24032249
Tada, I., Tsugawa, H., Meister, I., Zhang, P., Shu, R., Katsumi, R., Wheelock, C.E., Arita, M., and Chaleckis, R., Metabolites, 2019, vol. 9, no. 11, p. 251. https://doi.org/10.3390/metabo9110251
King, E., Overstreet, R., Nguyen, J., and Ciesielski, D., J. Chem. Inf. Model., 2022, vol. 62, no. 16, p. 3724. https://doi.org/10.1021/acs.jcim.2c00620
Kim, S., Kato, I., and Zhang, X., Metabolites, 2022, vol. 12, no. 8, p. 694. https://doi.org/10.3390/metabo12080694
Bittremieux, W., Schmid, R., Huber, F., Van der Hooft, J.J., Wang, M., and Dorrestein, P.C., J. Am. Soc. Mass Spectrom., 2022, vol. 33, no. 9, p. 1733. https://doi.org/10.1021/jasms.2c00153
Li, Y., Kind, T., Folz, J., Vaniya, A., Mehta, S.S., and Fiehn, O., Nat. Methods, 2021, vol. 18, no. 12, p. 1524. https://doi.org/10.1038/s41592-021-01331-z
Roberts, M.J., Moorthy, A.S., Sisco, E., and Kearsley, A.J., Anal. Chim. Acta, 2022, vol. 1230. https://doi.org/10.1016/j.aca.2022.340247
Matyushin, D.D., Sholokhova, A.Y., and Buryak, A.K., Anal. Chem., 2020, vol. 92, no. 17, p. 11818. https://doi.org/10.1021/acs.analchem.0c02082
Huber, F., Van der Burg, S., Van der Hooft, J.J., and Ridder, L., J. Cheminf., 2021, vol. 13, no. 1, p. 84. https://doi.org/10.1186/s13321-021-00558-4
GNPS. https://gnps.ucsd.edu/ProteoSAFe/static/gnps-splash.jsp. Accessed March 23, 2023.
Aksenov, A.A., Lab. Proizvod., 2019, no. 6, p. 8. https://doi.org/10.32757/2619-0923.2019.6.10.8.15
Quinlan, Z.A., Koester, I., Aron, A.T., Petras, D., Aluwihare, L.I., Dorrestein, P.C., Nelson, C.E., and Kelly, L.W., Metabolites, 2022, vol. 12, no. 12, p. 1275. https://doi.org/10.3390/metabo12121275
Neto, F.C. and Raftery, D., Anal. Chem., 2021, vol. 93, no. 35, p. 12001. https://doi.org/10.1021/acs.analchem.1c02041
Elie, N., Santerre, C., and Touboul, D., Anal. Chem., 2019, vol. 91, no. 18, p. 11489. https://doi.org/10.1021/acs.analchem.9b02802
Olivon, F., Elie, N., Grelier, G., Roussi, F., Litaudon, M., and Touboul, D., Anal. Chem., 2018, vol. 90, no. 23, p. 13900. https://doi.org/10.1021/acs.analchem.8b03099
Chen, L., Lu, W., Wang, L., Xing, X., Chen, Z., Teng, X., Zeng, X., Muscarella, A.D., Shen, Y., Cowan, A., McReynolds, M.R., Kennedy, B.J., Lato, A.M., Campagna, S.R., Singh, M., and Rabinowitz, J.D., Nat. Methods, 2021, vol. 18, no. 11, p. 1377. https://doi.org/10.1038/s41592-021-01303-3
Zhou, Z., Luo, M., Zhang, H., Yin, Y., Cai, Y., and Zhu, Z.J., Nat. Commun., 2022, vol. 13, p. 6656. https://doi.org/10.1038/s41467-022-34537-6
Treen, D.G., Wan, M., Xing, S., Louie, K.B., Huan, T., Dorrestein, P.C., Northen, T.R., and Bowen, B.P., Nat. Commun., 2022, vol. 13, p. 2510. https://doi.org/10.1038/s41467-022-30118-9
Ljoncheva, M., Stepisnik, T., Dzeroski, S., and Kosjek, T., Trends Environ. Anal. Chem., 2020, vol. 28, e00099. https://doi.org/10.1016/j.teac.2020.e00099
Fan, Z., Alley, A., Ghaffari, K., and Ressom, H.W., Metabolomics, 2020, vol. 16, p. 104. https://doi.org/10.1007/s11306-020-01726-7
Young, A., Wang, B., and Rost, H., arXiv:2111.04824, 2021. https://doi.org/10.48550/arXiv.2111.04824
Murphy, M., Jegelka, S., Fraenkel, E., Kind, T., Healey, D., and Butler, T., arXiv:2301.11419, 2023. https://doi.org/10.48550/arXiv.2301.11419
Hoffmann, M.A., Nothias, L.F., Ludwig, M., Fleischauer, M., Gentry, E.C., Witting, M., Dorrestein, P.C., Duhrkop, K., and Bocker, S., Nat. Biotechnol., 2022, vol. 40, no. 3, p. 411. https://doi.org/10.1038/s41587-021-01045-9
Bremer, P.L., Vaniya, A., Kind, T., Wang, S., and Fiehn, O., J. Chem. Inf. Model., 2022, vol. 62, no. 17, p. 4049. https://doi.org/10.1021/acs.jcim.2c00936
Milman, B.L., Ostrovidova, E.V., and Zhurkovich, I.K., Mass Spectrom. Lett., 2019, vol. 10, no. 3, p. 93. https://doi.org/10.5478/msl.2019.10.3.93
Wang, F., Liigand, J., Tian, S., Arndt, D., Greiner, R., and Wishart, D.S., Anal. Chem., 2021, vol. 93, no. 34, p. 11692. https://doi.org/10.1021/acs.analchem.1c01465
Koopman, J. and Grimme, S., J. Am. Soc. Mass Spectrom., 2021, vol. 32, no. 7, p. 1735. https://doi.org/10.1021/jasms.1c00098
Schnegotzki, R., Koopman, J., Grimme, S., and Sussmuth, R.D., Chem.—Eur. J., 2022, vol. 28, no. 27. https://doi.org/10.1002/chem.202200318
Duhrkop, K., Fleischauer, M., Ludwig, M., Aksenov, A.A., Melnik, A.V., Meusel, M., Dorrestein, P.C., Rousu, J., and Bocker, S., Nat. Methods, 2019, vol. 16, no. 4, p. 299. https://doi.org/10.1038/s41592-019-0344-8
Stravs, M.A., Duhrkop, K., Bocker, S., and Zamboni, N., Nat. Methods, 2022, vol. 19, no. 7, p. 865. https://doi.org/10.1038/s41592-022-01486-3
Zulfiqar, M., Gadelha, L., Steinbeck, C., Sorokina, M., and Peters, K., J. Cheminf., 2023, vol. 15, p. 32. https://doi.org/10.1186/s13321-023-00695-y
Liu, Y., De Vijlder, T., Bittremieux, W., Laukens, K., and Heyndrickx, W., Rapid Commun. Mass Spectrom., 2021, p. e9120. https://doi.org/10.1002/rcm.9120
Niessen, W.M.A. and Correa, C.R.A., Interpretation of MS-MS Mass Spectra of Drugs and Pesticides, Hoboken: Wiley, 2017. https://toc.library.ethz.ch/objects/pdf03/e01_978-1-118-50018-7_01.pdf. Accessed March 24, 2023.
Steckel, A. and Schlosser, G., Molecules, 2019, vol. 24, no. 3, p. 611. https://doi.org/10.3390/molecules24030611
Matyushin, D.D. and Buryak, A.K., IEEE Access, 2020, vol. 8, p. 223140. https://doi.org/10.1109/access.2020.3045047
Matyushin, D.D., Sholokhova, A.Y., Karnaeva, A.E., and Buryak, A.K., Chemom. Intell. Lab. Syst., 2020, vol. 202, p. 104042. https://doi.org/10.1016/j.chemolab.2020.104042
Kireev, A., Osipenko, S., Mallard, G., Nikolaev, E., and Kostyukevich, Y., Separations, 2022, vol. 9, no. 10, p. 265. https://doi.org/10.3390/separations9100265
Domingo-Almenara, X., Guijas, C., Billings, E., Montenegro-Burke, J.R., Uritboonthai, W., Aisporna, A.E., Chen, E., Benton, H.P., and Siuzdak, G., Nat. Commun., 2019, vol. 10, no. 1, p. 5811. https://doi.org/10.1038/s41467-019-13680-7
Witting, M. and Bocker, S., J. Sep. Sci., 2020, vol. 43, nos. 9–10, p. 1746. https://doi.org/10.1002/jssc.202000060
Bonini, P., Kind, T., Tsugawa, H., Barupal, D.K., and Fiehn, O., Anal. Chem., 2020, vol. 92, no. 11, p. 7515. https://doi.org/10.1021/acs.analchem.9b05765
Fedorova, E.S., Matyushin, D.D., Plyushchenko, I.V., Stavrianidi, A.N., and Buryak, A.K., J. Chromatogr. A, 2022, vol. 1664, 462792. https://doi.org/10.1016/j.chroma.2021.462792
Osipenko, S., Nikolaev, E., and Kostyukevich, Y., Separations, 2022, vol. 9, no. 10, p. 291. https://doi.org/10.3390/separations9100291
Lenski, M., Maallem, S., Zarcone, G., Garcon, G., Lo-Guidice, J.M., Antherieu, S., and Allorge, D., Metabolites, 2023, vol. 13, no. 2, p. 282. https://doi.org/10.3390/metabo13020282
Bouwmeester, R., Martens, L., and Degroeve, S., Anal. Chem., 2020, vol. 92, no. 9, p. 6571. https://doi.org/10.1021/acs.analchem.0c00233
Paglia, G., Smith, A.J., and Astarita, G., Mass Spectrom. Rev., 2022, vol. 41, no. 5, p. 722. https://doi.org/10.1002/mas.21686
Belova, L., Caballero-Casero, N., Van Nuijs, A.L., and Covaci, A., Anal. Chem., 2021, vol. 93, no. 16, p. 6428. https://doi.org/10.1021/acs.analchem.1c00142
Hohrenk, L., Itzel, F., Baetz, N., Tuerk, J., Vosough, M., and Schmidt, T.C., Anal. Chem., 2019, vol. 92, no. 2, p. 1898. https://doi.org/10.1021/acs.analchem.9b04095
Dekermanjian, J., Labeikovsky, W., Ghosh, D., and Kechris, K., Metabolites, 2021, vol. 11, no. 10, p. 678. https://doi.org/10.3390/metabo11100678
Schymanski, E.L., Singer, H.P., Slobodnik, J., Ipolyi, I.M., Oswald, P., Krauss, M., Schulze, T., Haglund, P., Letzel, T., Grosse, S., Thomaidis, N.S., Bletsou, A., Zwiener, C., Ibanez, M., Portoles, T., De Boer, R., Reid, M.J., Onghena, M., Kunkel, U., Schulz, W., Guillon, A., Noyon, N., Leroy, G., Bados, P., Bogialli, S., Stipanicev, D., Rostkowski, P., and Hollender, J., Anal. Bioanal. Chem., 2015, vol. 407, p. 6237. https://doi.org/10.1007/s00216-015-8681-7
CASMI. http://www.casmi-contest.org/2022/index.shtml. Accessed March 25, 2023.
Pezzatti, J., Gonzalez-Ruiz, V., Boccard, J., Guillarme, D., and Rudaz, S., Metabolites, 2020, vol. 10, no. 11, p. 464. https://doi.org/10.3390/metabo10110464
Clark, T.N., Houriet, J., Vidar, W.S., Kellogg, J.J., Todd, D.A., Cech, N.B., and Linington, R.G., J. Nat. Prod., 2021, vol. 84, no. 3, p. 824. https://doi.org/10.1021/acs.jnatprod.0c01376
Wong, J.W., Wang, J., Chang, J.S., Chow, W., Carlson, R., Rajski, L., Fernandez-Alba, A.R., Self, R., Cooke, W.K., Lock, C.M., Mercer, G.E., Mastovska, K., Schmitz, J., Vaclavik, L., Li, L., Panawennage, D., Pang, G.F., Zhou, H., Miao, S., Ho, C., Lam, T.C.H., To, Y.B.S., Zomer, P., Hung, Y.C., Lin, S.W., Liao, C.D., Culberson, D., Taylor, T., Wu, Y., Yu, D., Lim, P.L., Wu, Q., Schirle-Keller, J.P.X., Williams, S.M., Johnson, Y.S., Nason, S.L., Ammirata, M., Eitzer, B.D., Willis, M., Wyatt, S., Kwon, S.Y., Udawatte, N., Priyasantha, K., Wan, P., Filigenzi, M.S., Bakota, E.L., Sumarah, M.W., Renaud, J.B., Parinet, J., Bire, R., Hort, V., Prakash, S., Conway, M., Pyke, J.S., Yang, D.H.D., Jia, W., Zhang, K., and Hayward, D.G., J. Agric. Food Chem., 2021, vol. 69, no. 44, p. 13200. https://doi.org/10.1021/acs.jafc.1c04437
Anderson, B.G., Raskind, A., Habra, H., Kennedy, R.T., and Evans, C.R., Anal. Chem., 2021, vol. 93, no. 48, p. 15840. https://doi.org/10.1021/acs.analchem.1c02149
Funding
The work was carried out in accordance with the thematic plan of applied and scientific research and development in the field of healthcare within the State Assignment of the Federal Medical Biological Agency of Russia (code 64.001.23.800).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
The authors of this work declare that they have no conflicts of interest.
Additional information
Translated by O. Zhukova
Publisher’s Note.
Pleiades Publishing remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Milman, B.L., Zhurkovich, I.K. New Trends in Chemical Identification Methodology. J Anal Chem 79, 119–133 (2024). https://doi.org/10.1134/S1061934824020126
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1134/S1061934824020126