Abstract
Pseudomonas isolates have frequently been isolated from the rhizosphere of plants, and several of them have been reported as plant growth-promoting rhizobacteria. In the present work, tomato (Solanum lycopersicum) seeds were germinated in greenhouse conditions, and the seedling height, length of plants, collar diameter and number of leaves were measured from plants grown in soil inoculated by bacterial isolates. Pseudomonas isolates were isolated from the rhizosphere. We used the Newman-Keuls test to ascertain pairwise differences. Isolates were identified as a new Pseudomonas species by rpoD gene sequencing. The results showed that isolates of Pseudomonas sp. (Q6B) increased seed germination (P = 0.01); Pseudomonas sp. (Q6B, Q14B, Q7B, Q1B and Q13B) also promoted seedling height (P = 0.01). All five isolates promoted plant length and enlarged the collar diameter (P = 0.01). Pseudomonas sp. (Q1B) also increased leaf number (P = 0.01). The investigation found that Pseudomonas isolates were able to solubilize phosphate, produce siderophores, ammonia, and indole-3-acetic acid and colonize the roots of tomato plants. This study shows that these five novel Pseudomonas sp. isolates can be effective new plant growth-promoting rhizobacteria.
Similar content being viewed by others
Introduction
Plant growth-promoting rhizobacteria (PGPR)1 are an indispensable part of the rhizospheric biota and are beneficial bacteria that colonize plant roots2,3. PGPR increase the growth, yield, and stress tolerance of crop plants4. They improve plant growth through enhanced nutrient uptake from soil5 and a wide variety of mechanisms such as phosphate solubilization, siderophore production, biological nitrogen fixation, phytohormone production, antifungal activity, and systemic resistance induction. Such mechanisms make PGPR potentially usable biofertilizers6. Their emergence as a potent alternative has come in response to the overuse of agrochemical products such as fertilizers and pesticides, which lead to contamination of soil, fruits and vegetables5. This threat has prompted field workers to seek viable alternatives to reduce the use of chemical products.
According to studies conducted on this issue, PGPR can be included in biofertilizers and biopesticide applications, which makes them the most effective organic alternative6,7,8. PGPR comprise 2–5% of the total rhizobacterial community1 and facilitate its application. In line with the efficacy of PGPR, additional qualities also aid in increasing seed germination9, seedling vigor and plant growth10. PGPR are beneficial microorganisms that may be used in place of synthetic chemicals; furthermore, they improve plant growth by increasing nutrient availability. Overall, PGPR help to sustain environmental health and soil productivity11,12,13. In the analysis of PGPR, many studies confirm that the Pseudomonas genus represents the core of PGPR for many crops1,14,15,16,17,18,19.
Studies detail a twofold action in PGPR function. Direct mechanisms operate to produce metabolites. Kloepper et al.20 reported the production of metabolites, such as siderophores, by PGPR, contributing to enhanced growth. PGPR are also characterized by their ability to produce plant hormones, such as indole-3-acetic acid (I-3-AA)21,22,23, gibberellic acid24 and cytokinins25. Furthermore, PGPR solubilize phosphate26,27,28. Mishra et al.29 described Pseudomonas sp. MA-4 as the most efficient producer of ammonia that significantly increased biomass of the medicinal plant Geranium. Indirectly, PGPR promote plant growth by colonizing plant roots30,31 and suppressing diseases caused by pathogens32. Weller33 himself found that specific strains have the capacity to colonize the whole root system and survive for several weeks in the presence of the natural microflora. The combination of processes that PGPR contribute to make them an alternative to synthetic chemicals.
Our objective was to investigate the growth-promoting effects of laboratory fluorescent Pseudomonas isolates on tomato growth. Experiments were carried out to precisely evaluate the capacity to produce compounds such as those involved in regulation (I-3-AA), growth promotion (siderophore) and nutrient uptake assistance (phosphate solubilization). We also investigated their direct effect on plant height, collar diameter and leaf number in greenhouse conditions.
Results
Isolation of Pseudomonas sp
As already described above, bacterial populations were quantified from the samples collected from the tomato greenhouse of an experimental farm. The results obtained (Supplement Material Fig. S1) were mainly based on the method of Amkraz et al.34. They showed that the rhizospheric soil of tomatoes is rich in bacteria. The detected bacterial flora was superior to fluorescent Pseudomonas for all samples of roots. The fluorescent Pseudomonas were more abundant in the rhizoplane than the rhizosphere or endorhizosphere. They were characterized by discrete classes of 4.1 × 105, 2.9 × 105and 4 × 104 cfu in rhizoplane, rhizospheric soil and endorhizosphere, respectively. They represented 4.24%of the bacteria in the rhizoplane. In our investigation of the 19 fluorescent Pseudomonas isolates from all samples, 11 isolates (Q029B, Q052B, Q063B, Q021B, Q13B, Q056B, Q066B, Q059B, Q017B, Q034B and Q14B) were isolated from the rhizoplane, 5 isolates (Q6B, Q003B, Q054B, Q1B and Q7B) from the endorhizosphere and 3 isolates (Q015B, Q007B and Q027B) from the rhizospheric soil.
Greenhouse assay measuring seed germination and seedling height
Tomato seeds bacterized with fluorescent Pseudomonas were distinguished by their potential efficacious effects on seed germination and plant growth. Among the 19 isolates tested, Q6B was characterized by the highest significant effect on germination percentage (96%) compared to that of the control (Fig. 1). Twenty days after germination, the seedling height was enhanced significantly by five Pseudomonas isolates (Q14B, Q13B, Q1B, Q6B and Q7B) ranging from 8 cm to 8.75 cm for and Q7B and Q14B, respectively (Fig. 2). Due to such efficacious contributions to seedling height for Q14B, Q13B, Q1B, Q6B and Q7B, it was reasonable to test their effects on plant growth after transplantation.
Characterization of five fluorescent Pseudomonas isolates
The five isolates (Q14B, Q13B, Q1B, Q6B and Q7B) showed fluorescence production with diffusible yellowish-green pigment in KB medium under ultraviolet light (360 nm). The tested isolates were positive for motility and oxidase and arginine dehydrogenase activity. They were gram-negative. The isolates Q6B, Q13B and Q1B were positive for catalase activity and levan production, whereas Q7B and Q14B were negative (Table 1). Q7B and Q1B were positive for nitrate reduction, while Q6B, Q13B and Q14B were negative. Q13B was positive for gelatin liquefaction, but the others were negative (Table 1). Pure cultures of each isolate were used to confirm their identity on a species level using a molecular approach. To identify the isolated bacteria, a partial rpoD gene sequence of each isolate was amplified and sequenced. The obtained sequence was submitted to GenBank (MN091814 to MN091824, supplement material Table S1) and used for searching against the GenBank database using BLAST (http://www.blast.ncbi.nlm.nih.gov). The BLAST analysis results showed alignment of the sequence with several species of Pseudomonas, where the sequence exhibited maximum identity with Pseudomonas sp. Therefore, to characterize each isolate more precisely, the most closely related sequences were used to reconstruct a phylogenetic tree using maximum likelihood analysis. For the Moroccan Pseudomonas isolates, maximum likelihood phylogenetic trees inferred from the aligned partial rpoD nucleotide sequences from our study and from NCBI provided key information about their relationship within Pseudomonas spp. Diverse group. We could reconstruct the entire Pseudomonas phylogenetic landscape with all Pseudomonas Moroccan isolates grouped into two genetically close clusters. One cluster containing sequences of 5 Moroccan isolates closely related (98% nt similarity) to GenBank sequences of Pseudomonas type isolates from USA (LMG_2257T, accession number D86020) and France (DSM14164T, accession number FN554488) (Fig. 3). The other set of sequences from six Moroccan isolates clustered separately in a distant homogenous clade (99% nt similarity) sharing 99% identity with Chinese Pseudomonas sp. stains (MH758786). The inferred phylogenetic tree confirmed that the isolated bacteria lie in the genus Pseudomonas, are related to newly annotated Pseudomonas species, and are constituting a genetically independent branch (Fig. 3). Thus, taken all together, these results strongly indicate that the isolated organisms may belong to a new species in the genus Pseudomonas that have not yet been described.
Evolutionary tree representing Maximum likelihood phylogenetic analysis inferred from partial rpoD nucleotide sequences of the Moroccan (green triangles) and representative Pseudomonas strains. The GenBank accession numbers of the sequences are indicated together with the strain name. BioNumerics analysis included most of the available rpoD sequences. The phylogenetic clusters are delineated with vertical bars. Branch lengths on the phylogenetic tree represent the genetic distance, the numbers at the branches represent the percentage of replicates in which the topology of the branch was observed after 500 bootstrap replicates.
Plant growth
The performance of Pseudomonas isolates on tomato plant growth parameters was observed under greenhouse conditions 20 days after transplantation. The number of leaves, plant length (aerial region) and collar diameter were recorded. The results of this study showed that all of the bacteria significantly promote (p < 0.01) plant length and collar diameter. The longest significant plant length was obtained with Q13B, with a 31.5% increase in plant length (statistical supplementary file) and a 48.57% increase in collar diameter compared to those of the control. Pseudomonas sp. (Q1B) significantly enhanced leaf number by 83.33% (P < 0. 01) compared to that of the control (Table 2).
PGPR mechanisms
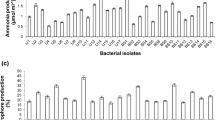
The five Pseudomonas isolates (Q14B, Q13B, Q1B, Q6B and Q7B) were tested for PGPR activity as follows: phosphate solubilization ability, I-3-AA production, siderophore production, ammonia production and root colonization ability (Table 3). All these isolates solubilized phosphate in solid NBRIP medium. Phosphate solubilization is manifested in the clear halo around the colony. Solubilization is shown for Q13B (11 to 75 mm halo diameter). In liquid medium, these isolates showed successful phosphate solubilization. The best solubilization was shown by Q6B (3.80 mg/ml). The five isolates produce siderophores in both solid and liquid medium. Production is visualized in the orange halo around the colony. The highest level of production was by Q13B (53.8%). The five isolates produce ammonia, with the best production shown by Q1B (3.55 mol/ml). The production of I-3-AA was detected in the five isolates in both solid and liquid media. The presence of I-3-AA was indicated by red spots on Whatman paper (No. 2) after treatment with Salkowski reagent. As far as liquid medium is concerned, the best production was shown by Q1B (2.26 µg/ml). All five isolates are characterized by their capacity to completely colonize the roots of seedlings.
Discussion
We achieved the objective of confirming that the Pseudomonas isolates focused on in this study facilitate siderophore, I-3-AA, and ammonia production, phosphate solubilization and the growth of tomatoes in greenhouse conditions.
The results showed that the rhizospheric soil of tomatoes is rich in bacteria. The number of aerobic bacteria was superior to fluorescent Pseudomonas for all root samples. These results were similar to those reported by Amkraz et al.34 but with an apparent difference in terms of the percentage of Pseudomonas in each sample of roots. The study performed by Amkraz et al.34 showed that Pseudomonas proportions were 7% in rhizospheric soil, 20% in the rhizoplane and 0.21% in the endorhizospheric component compared to 10%, 4.24% and 9.72%, respectively, in our survey. This disparity is attributed to the soil type, plant age, the season in which the samples were collected and the use of pesticides. Bacterial presence is variable, and the heightened sensitivity/susceptibility of bacteria is due to the heterogeneous distribution of bacteria in the soil in relation to farming practices, the use of chemical products, crop type and soil type19,35. Under our conditions, Pseudomonas was highly represented in comparison to the cultivable bacteria detected.
This study additionally demonstrated that among 19 isolates, five are effective PGPR isolates; they increased seed germination and plant growth of tomatoes under organic growing conditions. Growth stimulation mechanisms, including the production of phytohormones, phosphate solubilization, ammonia production and colonization of plant roots, are the most efficacious mechanisms that explain PGPR effects35,36. Chin-A-Woeng et al.37 reported that the ability of Pseudomonas isolates to suppress disease relies mainly on their ability to colonize roots. The results revealed a significant increase in seed germination due to the mixed bacteria formulation compared to the control. This effect is due to the increased synthesis of hormones linked to growth such as I-3-AA and gibberellins, which triggered the activity of specific enzymes that promote early germination38.
The molecular characterization showed that the five selected bacteria lie in the genus Pseudomonas, and they are a new strain of Pseudomonas species that has not yet been described. Phylogenetic relationship of the Moroccan isolates was revealed based on polymorphism in rpoD gene region, used earlier as an ecological marker39. The sequence of Pseudomonas-specific single-copy gene rpoD gene from 11 fluorescent pseudomonads was analyzed with maximum likelihood algorithm and compared with the corresponding sequence of a broad range of rpoD sequences from Pseudomonas representative type strains available in Genbank. This placed the Moroccan isolates in two separate well supported clusters suggesting that these lineages are genotypically heterogeneous and might belong to a new species within Pseudomonas sensu stricto. Comparative phylogenetic analyses with rpoD gene sequences was performed to assign the Moroccan isolates with high resolving power to Pseudomonas species. The phylogenetic interrelatedness among Pseudomonas taxa inferred by rpoD gene-based phylogeny was congruent with that inferred by other phylogenies. Our robust maximum likelihood phylogenetic analysis permitted fine differentiation of these Pseudomonas isolates, suggesting our rpoD gene-based phylogeny is valuable for fine differentiation and efficient classification of closely related Pseudomonas isolates, as it was found in other hallmark studies, especially for resolving closely related isolates in microbial ecology40,41. This study highlights also the necessity for complete genomic sequencing of these isolates for strong phylogenomic taxonomy to facilitate robust assignment of these species and to avoid potential taxonomic inconsistencies.
This study showed that the new Pseudomonas sp. isolates are able to produce indole-3-acetic acid, ammonia, and siderophores and solubilize phosphate.
Phosphate solubilizing is explained by the production of various organic acids and enzymes42,43. These factors transform insoluble phosphates into substances that can be easily assimilated by plants44,45,46. Among these organic acids are gluconic, tartaric, and oxalic acids42,43,47. Gluconic acid is the main component during solubilization48,49 Indole-3-acetic acid50 is the most important auxin produced by bacteria, plants and fungi. I-3-AA initiates root, leaf and flower development51. Its importance lies in its central role in cell division, elongation, fruit development and senescence52. Siderophores themselves are useful for phytostabilization. They facilitate plant growth and coalescence of metals and reduce metal bioavailability in soil53; as such they are of use in bioremediation of soils. These characteristics determine plant growth in general. As a result, the five novel isolates of Pseudomonas selected can be considered as a new PGPR that have an important role as biological fertilizers with bioremediation qualities6. Pseudomonas enhances nutrient bioavailability and bioassimilation. The use of these bacterial complements may lead to reduced application rates or even elimination of chemical fertilizers52. PGPR increase chlorophyll formation in plant leaves, which enhances photosynthesis54. Photosynthesis is the primary metabolic process by which plants grow55. PGPR affect healthy plant growth and have a pesticidal role in protecting plants from soil pathogens.
Such diverse benefits have made PGPR a potential resource in themselves. Further work should use this new species as a new candidate for biofertilizers on other plants, aiming at preventing damage to ecosystem structure since “the tapestry becomes threadbare and begins to fall into tatters, becoming irreparable even for the most capable weaver, the last remnants serving essentially no function whatsoever, without all the fibers in their proper places, without all the right words, without all the integral elements of the fabric of life”56.
Future investigation is recommended with both biochemical and molecular approaches
We should elucidate the pathways of the mechanisms involved in this approach such as auxins57, siderophores and phosphate solubilization58. In addition, we suggest that future studies make use of Bayesian approaches to unify the different subjective areas to form a predictive function for the effectiveness and application of PGPR mechanisms in alternative crop species.
Methods
To achieve our objectives, we followed the protocols shown in sections 4.1–4.9.
Isolation of Pseudomonas spp
The samples were collected from an experimental farm of the National Institute for Agricultural Research, Agadir, southwestern Morocco (30°02′42.2“N 9°33′13.4“W) in 2016 from tomato roots and rhizospheric soil. For each sample, 500 g was collected from healthy tomato plants in a greenhouse. Samples were kept in the laboratory at 4 °C before analysis.
The bacterial communities of the rhizosphere (RS), the rhizoplane (RH) and the endorhizosphere (ER) were isolated as defined by Dommergues and Mangenot59 through the following steps:
-
(i).
To isolate bacteria from the rhizosphere, the roots were carefully shaken. Then, 1 g of rhizospheric soil was added to 9 ml of sterile physiological water, and the mixture was agitated at 120 rpm for 2 min.
-
(ii).
To isolate bacteria from the rhizoplane, the rhizospheric soil was dislodged from the fresh roots, and 1 g of root segments was agitated for 2 hours in 9 ml of sterile physiological water at 120 rpm.
-
(iii).
To isolate bacteria from the endorhizosphere, the surface of root segments was subsequently disinfected using 2.5% sodium hypochlorite solution for 3 min60, rinsed three times with sterile distilled water and 1 g was blended in 9 ml of sterile physiologic water.
-
(iv).
Serial dilutions were separately prepared from the extracts (RS, RH and ER), and 0.1 ml of each dilution was seeded onto King B medium to isolate and quantify fluorescent Pseudomonas spp.61. Three replicates were made for each extract, and the plates were incubated at 28 °C for 48 hours. Fluorescent colonies on King B medium under UV light were subcultured twice before storage at 4 °C on yeast dextrose carbonate (YDC) agar and at −80 °C in 40% glycerol62,63.
Greenhouse seed germination and seedling height measurement
Tomato seeds (Campbell 33) were disinfected with 2.5% sodium hypochlorite solution for 3 min57, rinsed three times with sterile distilled water and air-dried. These seeds were inoculated by soaking for 30 min in a suspension of bacteria (108cfu/ml) amended with 2% carboxymethylcellulose (CMC) and air-dried for 12 hours. The treated seeds were sown in disinfected cell trays (two seeds per cell) containing a mixture of sterilized sand and peat (1:2 v/v). Controls for the experiment were treated with a mixture of sterile distilled water and 2% CMC. The trays were placed in an experimental greenhouse set with a photoperiod of 12 hours, a temperature of 25 ± 2 °C, and a relative humidity (RH) of 80 ± 5% and were watered every 2 days. The rate of germination compared to that of the control was determined five days after sowing. Seedling height measurements were made 20 days after germination. Three replicates were carried out for all treatments with 10 seeds.
Bacterial characterization
The five selected bacteria were characterized for the following biochemical traits: Gram test, motility, oxidase, catalase, glucose fermentation, arginine-dihydrolase, nitrate reduction, gelatin hydrolysis, levan production, manitol utilization, and carbon source utilization14,18,64,65.
Bacterial isolates were cultured for 24 hours, and DNA was extracted using a Bioline ISOLATE II Genomic DNA Kit (Bioline, Australia) following the manufacturer’s guidelines for cell culture extractions with the following adaptation: three to four colonies from fresh culture were suspended in sterile water as the starting material. Extracted DNA was visualized using gel electrophoresis on a 1% agarose gel prestained with ethidium bromide electrophoresed at 100 V for 40–60 min and viewed under UV light. DNA was stored at −20 °C for further use.
Molecular detection and partial rpoD sequencing
A fragment of the rpoD gene was PCR-amplified with the primers PsrpoD FNP1 (5′-TGAAGGCGARATCGAAATCGCCAA-3′) and PsrpoDnprpcr1 (5′-YGCMGWCAGCTTYTGCTGGCA-3′)66. PCR was performed in a 25 µl volume containing 1 µlof DNA (25 ng/µl), 2.5 µlof 10 × PCR buffer with 20 mM MgCl2 (Roche), 2.5 µlof dNTPs (2 mM each), 0.75 µlof forward primer (10 μM), 1.5 µlof reverse primer (10 μM) and 0.125 µlof FastStart polymerase (5 U/µl; Roche). The PCR temperature profile was performed according to Parkinson et al.66. All amplification products were purified with the GeneJET PCR Purification Kit (Thermo Fisher Scientific Inc.) and sequenced with both forward and reverse primers using a commercial service (Macrogen Inc.). The rpoD amplicons were trimmed to 578 nucleotides according to Parkinson et al.66 with BioNumerics 7.1 software (Applied Maths, Belgium). The 578 nt sequence was used to classify the bacterial isolates at the species level by a BlastN query against the nucleotide database of NCBI. All Pseudomonas strains and isolates used in this study for sequencing and phylogenetic analysis are detailed in supplement material (Table S1).
rpoD sequences and phylogenetic analysis
The obtained Sanger sequences from the rpoD genomic region were assembled, analyzed and aligned using the BioNumerics 7 (Applied Math version 7.6.1). Sequence identity of the Moroccan Pseudomonas isolates was confirmed by similarity search using the BLASTn program in GenBank (https://blast.ncbi.nlm.nih.gov/Blast.cgi). In addition to Moroccan rpoD partial sequences, a selection of nucleotide sequences of the rpoD all representative Pseudomonas isolates from different countries and hosts were retrieved from GenBank, aligned and used for phylogenetic analyses and molecular evolutionary genetics analysis. The evolutionary history and phylogenetic tree was inferred by using the Maximum Likelihood method and Tamura-Nei model67. The percentage of trees in which the associated taxa clustered together is shown next to the branches. Initial tree(s) for the heuristic search were obtained automatically by applying Neighbor-Join and BioNJ algorithms to a matrix of pairwise distances estimated using the Maximum Composite Likelihood (MCL) approach, and then selecting the topology with superior log likelihood value. The robustness of the internal branches was tested with assessment of the confidence of branching patterns by bootstrap analysis with 500 pseudo-random iterations. The tree was drawn to scale, with branch lengths measured in the number of substitutions per site. This analysis involved 117 nucleotide sequences. Codon positions included were 1st + 2nd + 3rd + Noncoding. There were a total of 584 positions in the final dataset. Evolutionary analyses were conducted in MEGA X68.
Plant growth assay
Testing the effects of the five Pseudomonas isolates on tomato plant growth, substrate that was a mixture of sand and peat (1:2 v/v) was inoculated by strain suspensions (108cfu/g)34 and was distributed in plastic pots (4 l). Twenty days after sowing, 10 seedlings of the same height that grew out from inoculated seeds were transplanted to the pre-prepared substrate. Plants were grown under typical greenhouse growing conditions (min and max temperatures:18°Cand 26 °C, respectively, 75 ± 5%relative humidity)69 and were watered every 2 days and fertilized each week with a commercial nutrient solution (NPK 20-20-20)34. The control plants grown from untreated seeds were transplanted into soil substrate without inoculants. All treatments were replicated three times. The pots containing plants were arranged in a completely randomized design. After 20 days, the plant length (aerial region), collar diameter and number of leaves were measured.
Phosphate solubilization
Phosphate solubilization by the isolated fluorescent Pseudomonas isolates was tested in solid media by the method described by Nautiyal70. Ten microliters of the culture of each strain was spotted on the surface of the solid media in Petri dishes. The solubilization capacity was assessed by the transparent area formed around the colony. Ten days after incubation at 30 °C, the diameter of the solubilization halo (DSH) was determined by the following formula (Eq. 1):
where THD is the total halo diameter, and CD is the colony diameter.
Quantitative estimation of phosphate solubilization in broth was carried out in Erlenmeyer flasks (250 ml) containing 100 ml of NBRIP medium70. The flasks were incubated at 28 °C for 5 days at 120 rpm. Then, 5 ml of each isolate culture was centrifuged at 3000 rpm for 20 min. Phosphate concentration in culture supernatant was estimated as described by Olsen and Sommers71. Three replicates were used, and a standard calibration curve was made with KH2PO4 solution (Sigma-Aldrich).
Indole-3-Acetic acid production
I-3-AA production was detected as described by Bric et al.72. Cultures were maintained on Luria-Bertani (LB) agar medium (Sigma-Aldrich Chemical Co., St. Louis, MO). The pH was adjusted to 7.5 before autoclaving. The LB medium was supplemented with 5 mM L-tryptophan. Agar plates (9-cm diameter) were inoculated using sterile toothpicks; each inoculated Petri dish was overlaid with an 82-mm diameter disk of Whatman paper (No.2). The plates were transferred into an incubator at 27 °C for 3 days. Once the diameter of colonies on LB medium was 2 mm, I-3-AA was present. Whatman disks were treated with Salkowski reagent (2% 0. 5 MFeCl3 in 35% perchloric acid). Tests were carried out at room temperature. Bacteria producing I-3-AA were identified by the formation of a characteristic red halo. Three replicates were performed for each isolate.
The production of I-3-AA by the five fluorescent Pseudomonas isolates was evaluated spectrophotometrically, with the presence being identified at 535 nm. Liquid cultures were prepared in 250-ml flasks containing 100 ml of 50% TSB, with 200 mg/ml L-tryptophan (Sigma-Aldrich). The flasks were inoculated with 100 µl of each isolate culture (108cfu/ml) and incubated overnight (approx. 12 hours) at 27 °C23. Following inoculation, we observed an incubation period of 72 hours on a rotary shaker (150 rev/min, 28 °C). Bacterial cells were removed by centrifugation (4000 g, 10 min). One milliliter of each supernatant was mixed vigorously with 2 ml of Salkowski reagent. The mixture was incubated at room temperature (24 °C) for 20 min. I-3-AA production was observed as a pink-red color, and the absorbance was measured at 535 nm using an Optizen 3220UV Double Beam UV-Vis spectrophotometer (Mecasys, Korea). The concentration of I-3-AA was determined using a standard curve prepared using serial dilutions of a 50 mg/ml I-3-AA (Sigma-Aldrich) solution in 50% TSB. Four replicates were used for each treatment23.
Root colonization
The following steps were carried out: (i) spontaneous antibiotic-resistant mutants were obtained by transferring bacteria to King B medium containing rifampicin (180 µg/ml) (Sigma-Aldrich)73. To prevent contamination by fungi, cycloheximide (100 µg/ml) (Sigma-Aldrich) was added to the medium. Colonies growing in this medium were serially transferred three times to the same medium and stored at 4 °C; (ii) tomato seeds (Campbell 33) were disinfected, treated with rifampicin-resistant bacteria and sown in a mixture of sand and peat (1:2, v/v); (iii) After 2 weeks, plants and the soil adhering to the roots were carefully removed. Then, roots were placed on King B medium supplemented with rifampicin (180 µg/ml) and cycloheximide (100 µg/ml). The plates were incubated at 28 °C for 48 hours, and root colonization was evaluated on the basis of the growth of the introduced fluorescent Pseudomonas along the roots34.
Production of siderophores
Siderophore production was tested qualitatively using Chrome Azurol S medium (CAS-medium)74. Each fluorescent Pseudomonas isolate was streaked on the surface of CAS agar medium and incubated at 28 °C for 3 days. Siderophore production was confirmed by observation of an orange halo around the colonies after incubation. Three replicates were performed.
Quantitative analysis of siderophore production was performed in King B liquid medium inoculated with 100 µl of Pseudomonas isolate culture (108cfu/ml) and incubated at 28 °C for 72 hours. Cultures were centrifuged at 5000 rpm for 30 min, and 500 µl of the supernatant was mixed with 500 µlof CAS solution. The color changed from blue to orange, indicating siderophore production. After 20 min of incubation, optical density was measured by an Optizen 3220UV Double Beam UV-Vis spectrophotometer (Mecasys, Korea) at 630 nm. The percentage of siderophores was calculated using the following formula (Eq. 2):
where RA represents the absorbance of the reference (CAS reagent), and SA represents the absorbance of the sample75.
Ammonia production
Ammonia production was measured by growing cultures in peptone water with Nessler’s reagent76. A color change from brown to yellow indicates ammonia production; optical density was measured using an Optizen 3220UV Double Beam UV-Vis spectrophotometer at 450 nm77. The concentration of ammonia was estimated based on a standard curve of ammonium sulfate (Sigma-Aldrich) ranging from 0.1 to 1 µmol/ml. Sterile peptone water served as a negative control.
Data analysis
Data were subjected to ANOVA. Data for plant growth experiments are presented as the means ± standard deviation. Any difference mentioned is significant at p < 0.01 using the Newman–Keuls test78 (details are provided in a statistics supplementary file).
References
Kloepper, JW. & Schroth MN. Plant growth-promoting rhizobacteria on radishes. In: Proceedings of the 4th international conference on plant pathogenic bacteria, vol 2. Station de Pathologie Végétale et de Phytobactériologie, INRA, Angers, France, pp 879–882 (1978).
Buddrus-Schiemann, K., Schmid, M., Schreiner, K., Welzl, G. & Hartmann, A. Root Colonization by Pseudomonas Sp. Dsmz 13134 and Impact on the Indigenous Rhizosphere Bacterial Community of Barley. Microb Ecol. 2, 381–393 (2010).
Fließbach, A., Winkler, M., Lutz, M. P., Oberholzer, H.-R. & Mäder, P. Soil Amendment with Pseudomonas Fluorescens Cha0: Lasting Effects on Soil Biological Properties in Soils Low in Microbial Biomass and Activity. Microb Ecol. 4, 611–623 (2009).
Jorquera, M. A. et al. Plant Growth-Promoting Rhizobacteria Associated with Ancient Clones of Creosote Bush (Larrea Tridentata). Microb Ecol. 4, 1008–1017 (2012).
Adesemoye, A., Torbert, H. & Kloepper, J. Plant Growth-Promoting Rhizobacteria Allow Reduced Application Rates of Chemical Fertilizers. Microb Ecol. 4, 921–929 (2009).
Vessey, J. K. Plant Growth Promoting Rhizobacteria as Biofertilizers. Plant soil. 2, 571–586 (2003).
Bharti, N., Barnawal, D., Maji, D. & Kalra, A. Halotolerant Pgprs Prevent Major Shifts in Indigenous Microbial Community Structure under Salinity Stress. Microb Ecol. 1, 196–208 (2015).
Bloemberg, G. V. & Lugtenberg, B. J. Molecular Basis of Plant Growth Promotion and Biocontrol by Rhizobacteria. Curr Opin Plant Biol. 4, 343–350 (2001).
Sarma, R. K. & Saikia, R. Alleviation of Drought Stress in Mung Bean by Strain Pseudomonas Aeruginosa Ggrj21. Plant soil. 1-2, 111–126 (2014).
Rahmoune, B. et al. Isolation and Characterization of Three New Pgpr and Their Effects on the Growth of Arabidopsis and Datura. Plants. J Plant Inter. 1, 1–6 (2017).
Esitken, A., Ercisli, S., Karlidag, H. & Sahin, F. Potential Use of Plant Growth Promoting Rhizobacteria (Pgpr) in Organic Apricot Production. in Proceedings of the international scientific conference: Environmentally friendly fruit growing, Polli, Estonia, 7–9 September, 2005: Tartu Univ Press (2005).
O’connell, P. F. Sustainable Agriculture-a Valid Alternative. Outlook on Agriculture. 1, 5–12 (1992).
Singh, R. K., Malik, N. & Singh, S. Improved Nutrient Use Efficiency Increases Plant Growth of Rice with the Use of Iaa-Overproducing Strains of Endophytic Burkholderia Cepacia Strain Rre25. Microb Ecol. 2, 375–384 (2013).
Bossis, E., Lemanceau, P., Latour, X. & Gardan, L. The Taxonomy of Pseudomonas Fluorescens and Pseudomonas Putida: Current Status and Need for Revision. Agronomie. 1, 51–63 (2000).
Burr, T., Schroth, M. & Suslow, T. Increased Potato Yields by Treatment of Seed Pieces with Specific Strains of Pseudomonas Fluorescens and Pseudomonas Putida [Bacterization]. Phytopathology 11, 1078–1082 (1978).
Chabot, R., Antoun, H. & Cescas, M. P. Stimulation De La Croissance Du Maïs Et De La Laitue Romaine Par Des Microorganismes Dissolvant Le Phosphore Inorganique. Can J Microbiol. 10, 941–947 (1993).
Schippers, B., Bakker, A. W. & Bakker, P. A. Interactions of Deleterious and Beneficial Rhizosphere Microorganisms and the Effect of Cropping Practices. Annu Rev Phytopathol. 1, 339–358 (1987).
Stanier, R. Y., Palleroni, N. J. & Doudoroff, M. The Aerobic Pseudomonads a Taxonomic Study. Microbiology. 2, 159–271 (1966).
Sutra, L., Risede, J. & Gardan, L. Isolation of Fluorescent Pseudomonads from the Rhizosphere of Banana Plants Antagonistic Towards Root Necrosing Fungi. Lett Appl Microbiol. 4, 289–293 (2000).
Kloepper, J. W., Leong, J., Teintze, M. & Schroth, M. N. Enhanced Plant Growth by Siderophores Produced by Plant Growth-Promoting Rhizobacteria. Nature. 5776, 885–886 (1980).
Abbass, Z. & Okon, Y. Plant Growth Promotion by Azotobacter Paspali in the Rhizosphere. Soil Biol Biochem. 8, 1075–1083 (1993).
Egamberdiyeva, D. Plant‐Growth‐Promoting Rhizobacteria Isolated from a Calcisol in a Semi‐Arid Region of Uzbekistan: Biochemical Characterization and Effectiveness. J Plant Nutr soil Sci. 1, 94–99 (2005).
Gravel, V., Antoun, H. & Tweddell, R. J. Growth Stimulation and Fruit Yield Improvement of Greenhouse Tomato Plants by Inoculation with Pseudomonas Putida or Trichoderma Atroviride: Possible Role of Indole Acetic Acid (Iaa). Soil Biol Biochem. 8, 1968–1977 (2007).
Gutiérrez-Mañero, F. J. et al. The Plant-Growth-Promoting Rhizobacteria Bacillus Pumilus and Bacillus Licheniformis Produce High Amounts of Physiologically Active Gibberellins. Physiol Plant. 2, 206–211 (2001).
Garcia de Salamone, I. E., Hynes, R. K. & Nelson, L. M. Cytokinin Production by Plant Growth Promoting Rhizobacteria and Selected Mutants. Can J Microbiol. 5, 404–411 (2001).
Jeon, J.-S., Lee, S.-S., Kim, H.-Y., Ahn, T.-S. & Song, H.-G. Plant Growth Promotion in Soil by Some Inoculated Microorganisms. J Bacteriol Soil. 4, 271–276 (2003).
Kushwaha, C., Tripathi, S. & Singh, K. Soil Organic Matter and Water-Stable Aggregates under Different Tillage and Residue Conditions in a Tropical Dryland Agroecosystem. Appl Soil Ecol. 3, 229–241 (2001).
Naz, I. & Bano, A. Biochemical, Molecular Characterization and Growth Promoting Effects of Phosphate Solubilizing Pseudomonas Sp. Isolated from Weeds Grown in Salt Range of Pakistan. Plant soil. 1-2, 199–207 (2010).
Mishra, R. K., Prakash, O., Alam, M. & Dikshit, A. Influence of Plant Growth Promoting Rhizobacteria (Pgpr) on the Productivity of Pelargonium Graveolens L. Herit. Recent res. sci. technol. 5 (2010).
Raaijmakers, J. M. & Weller, D. M. Exploiting Genotypic Diversity of 2, 4-Diacetylphloroglucinol-Producing Pseudomonas Spp.: Characterization of Superior Root-Colonizing Pseudomonas sp strain Q8r1-96. Appl Environ Microbiol. 6, 2545–2554 (2001).
Weller, D. M., Raaijmakers, J. M., Gardener, B. B. M. & Thomashow, L. S. Microbial Populations Responsible for Specific Soil Suppressiveness to Plant Pathogens 1. Annu Rev Phytopathol. 1, 309–348 (2002).
Van Loon, L. C. & Glick, B. R. Increased Plant Fitness by Rhizobacteria, in Molecular Ecotoxicology of Plants (2004).
Weller, D. M. Biological Control of Soilborne Plant Pathogens in the Rhizosphere with Bacteria. Annu Rev Phytopathol. 1, 379–407 (1988).
Amkraz, N., Boudyach, E., Boubaker, H., Bouizgarne, B. & Aoumar, A. A. B. Screening for Fluorescent Pseudomonades, Isolated from the Rhizosphere of Tomato, for Antagonistic Activity toward Clavibacter Michiganensis Subsp. Michiganensis. World J Microbiol Biotechnol. 6, 1059–1065 (2010).
Suslow, T. V. & Schroth, M. N. Rhizobacteria of sugar beets: effects of seed application and root colonization on yield. Phytopathology 72, 199–206 (1982).
Bruto, M., Prigent-Combaret, C., Muller, D. & Moënne-Loccoz, Y. Analysis of genes contributing to plant-beneficial functions in plant growth-promoting rhizobacteria and related Proteobacteria. Sci repo 4, 6261 (2014).
Chin-A-Woeng, T. F., Bloemberg, G. V., Mulders, I. H., Dekkers, L. C. & Lugtenberg, B. J. Root Colonization by Phenazine-1-Carboxamide-Producing Bacterium Pseudomonas Chlororaphis Pcl1391 Is Essential for Biocontrol of Tomato Foot and Root Rot. Mol Plant Microbe Interact. 12, 1340–1345 (2000).
Bharathi, R., Vivekananthan, R., Harish, S., Ramanathan, A. & Samiyappan, R. Rhizobacteria-Based Bio-Formulations for the Management of Fruit Rot Infection in Chillies. Crop Prot. 9, 835–843 (2004).
Rajwar, A. & Sahgal, M. Phylogenetic Relationships of Fluorescent Pseudomonads Deduced From the Sequence Analysis of 16S rRNA, Pseudomonas-specific and rpoD genes. 3 Biotech 6, 80 (2016).
Ghyselinck, J., Coorevits, A., Van Landschoot, A., Samyn E Heylen, K. & De Vos, P. An rpoD Gene Sequence Based Evaluation of Cultured Pseudomonas diversity On Different Growth Media. Microbiology 159, 2097–2108 (2013).
Mulet, M., Bennasar, A., Lalucat, J. & Garcia-Valdes, E. An rpoD based PCR Procedure For the Identification of Pseudomonas Species and For Their Detection In Environmental Samples. Mol Cell Probes 23, 140–147 (2009).
Chen, Y. et al. Phosphate Solubilizing Bacteria from Subtropical Soil and Their Tricalcium Phosphate Solubilizing Abilities. Appl Soil Ecol. 1, 33–41 (2006).
Patel, D. K., Archana, G. & Kumar, G. N. Variation in the Nature of Organic Acid Secretion and Mineral Phosphate Solubilization by Citrobacter Sp. Dhrss in the Presence of Different Sugars. Curr Microbiol. 2, 168–174 (2008).
Brígido, C., Glick, B. R. & Oliveira, S. Survey of Plant Growth-Promoting Mechanisms in Native Portuguese Chickpea Mesorhizobium Isolates. Microb Ecol. 4, 900–915 (2017).
Rodrı́guez, H. & Fraga, R. Phosphate Solubilizing Bacteria and Their Role in Plant Growth Promotion. Biotech Adv. 4, 319–339 (1999).
Rodríguez, H., Fraga, R., Gonzalez, T. & Bashan, Y. Genetics of Phosphate Solubilization and Its Potential Applications for Improving Plant Growth-Promoting Bacteria. Plant soil. 1, 15–21 (2006).
Zeng, Q., Wu, X. & Wen, X. Effects of Soluble Phosphate on Phosphate-Solubilizing Characteristics and Expression of Gcd Gene in Pseudomonas Frederiksbergensis Jw-Sd2. Curr Microbiol. 2, 198–206 (2016).
Goldstein, A. H. Recent Progress in Understanding the Molecular Genetics and Biochemistry of Calcium Phosphate Solubilization by Gram Negative Bacteria. Biol Agric Hort. 2, 185–193 (1995).
Vyas, P. & Gulati, A. Organic Acid Production in Vitro and Plant Growth Promotion in Maize under Controlled Environment by Phosphate-Solubilizing Fluorescent Pseudomonas. BMC Microbiol. 1, 174 (2009).
Sadfi-Zouaoui, N., Essghaier, B., Hannachi, I., Hajlaoui, M. R. & Boudabous, A. First Report on the Use of Moderately Halophilic Bacteria against Stem Canker of Greenhouse Tomatoes Caused By Botrytis Cinerea. Ann Microbiol. 3, 337–339 (2007).
Phillips, K. A. et al. Vanishing Tassel2 Encodes a Grass-Specific Tryptophan Aminotransferase Required for Vegetative and Reproductive Development in Maize. Plant Cell. 2, 550–566 (2011).
Duca, D., Lorv, J., Patten, C. L., Rose, D. & Glick, B. R. Indole-3-Acetic Acid in Plant–Microbe Interactions. Antonie van Leeuwenhoek. 1, 85–125 (2014).
Chen, Y. et al. Metal-Tolerant Enterobacter Sp. Strain Eg16 Enhanced Phytoremediation Using Hibiscus Cannabinus Via Siderophore-Mediated Plant Growth Promotion under Metal Contamination. Plant soil. 1-2, 203–216 (2017).
Nadeem, S. M., Zahir, Z. A., Naveed, M. & Arshad, M. Rhizobacteria Containing Acc-Deaminase Confer Salt Tolerance in Maize Grown on Salt-Affected Fields. Can J Microbiol. 11, 1302–1309 (2009).
Furze, J. N., Zhu, Q., Qiao, F. & Hill, J. Utopian Exploration of Global Patterns of Plant Metabolism. in Modelling, Identification & Control (ICMIC), 2013 Proc Inte Conf on: IEEE (2013).
Swing, K. The Vocabulary of Nature,. Mathematical Advances Towards Sustainable Environmental Systems. Springer.: vii-ix (2017).
Defez, R., Andreozzi, A. & Bianco, C. The Overproduction of Indole-3-Acetic Acid (Iaa) in Endophytes Upregulates Nitrogen Fixation in Both Bacterial Cultures and Inoculated Rice Plants. Microb Ecol.: 1–12 (2017).
Vyas, P., Rahi, P. & Gulati, A. Stress Tolerance and Genetic Variability of Phosphate-Solubilizing Fluorescent Pseudomonas from the Cold Deserts of the Trans-Himalayas. Microb Ecol. 2, 425–434 (2009).
Dommergues, Y., Mangenot, F. & Yvon Dommergues, F. M. Ecologie Microbienne Du Sol, Masson (1970).
Mew, T. & Rosales, A. Bacterization of Rice Plants for Control of Sheath Blight Caused by Rhizoctonia Solani. Phytopathology. 11, 1260–1264 (1986).
King, E. O., Ward, M. K. & Raney, D. E. Two Simple Media for the Demonstration of Pyocyanin and Fluorescin. J Lab Clin Med. 301–307 (1954).
Kaur, R., Singh, R. & Alabouvette, C. Antagonistic Activity of Selected Isolates of Fluorescent Pseudomonas against Fusarium Oxysporum F. Sp. Ciceri. Asian J Plant Sci. 1682–3974 (2007).
Parke, J., Moen, R., Rovira, A. & Bowen, G. Soil Water Flow Affects the Rhizosphere Distribution of a Seed-Borne Biological Control Agent, Pseudomonas Fluorescens. Soil Biol Biochem. 6, 583–588 (1986).
Falkow, S. Activity of Lysine Decarboxylase as an Aid in the Identification of Salmonellae and Shigellae. Am J Clin Path. 6, 598–600 (1958).
Hugh, R. & Leifson, E. The Taxonomic Significance of Fermentative Versus Oxidative Metabolism of Carbohydrates by Various Gram Negative Bacteria. J Bacteriol. 1, 24 (1953).
Parkinson, N., Bryant, R., Bew, J. & Elphinstone, J. Rapid phylogenetic identification of members of the Pseudomonassyringae species complex using the rpoD locus. Plant Pathol. 60, 338–344 (2011).
Tamura, K. & Nei, M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Molecular Biology and Evolution 10, 512–526 (1993).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Molecular Biology and Evolution 35, 1547–1549 (2018).
Ekesi, S. D. M. et al. Taxonomy, Ecology, and Management of Native and Exotic Fruit Fly Species in Africa. Annu Rev Entomol. 219–238 (2016).
Nautiyal, C. S. An Efficient Microbiological Growth Medium for Screening Phosphate Solubilizing Microorganisms. FEMS Microbiol Lett. 1, 265–270 (1999).
Olsen, S., Sommers, L. & Page, A. Methods of Soil Analysis. Part 2. Chemical and microbiological properties of Phosphorus. ASA Monogr. 9, 403–430 (1982).
Bric, J. M., Bostock, R. M. & Silverstone, S. E. Rapid in Situ Assay for Indoleacetic Acid Production by Bacteria Immobilized on a Nitrocellulose Membrane. Appl Environ Microbiol. 2, 535–538 (1991).
Shahid, M. et al. Root Colonization and Growth Promotion of Sunflower (Helianthus Annuus L.) by Phosphate Solubilizing Enterobacter Sp. Fs-11. World J Microbiol Biotechnol. 8, 2749–2758 (2012).
Schwyn, B. & Neilands, J. Universal Chemical Assay for the Detection and Determination of Siderophores. Anal Biochem. 1, 47–56 (1987).
Pastor, N., Rosas, S., Luna, V. & Rovera, M. Inoculation with Pseudomonas Putida Pci2, a Phosphate Solubilizing Rhizobacterium, Stimulates the Growth of Tomato Plants. Symbiosis. 3, 157–167 (2014).
Goswami, D. et al. Elucidating Multifaceted Urease Producing Marine Pseudomonas Aeruginosa Bg as a Cogent Pgpr and Bio-Control Agent. Plant Growth Regul. 1, 253–263 (2015).
Cappucino, J. & Sherman, N. Microbiology: A Laboratory Manual. 3rd edn Benjamin/Cumming Pub. 125–179 (1992).
De Muth, J. E. Basic Statistics and Pharmaceutical Statistical Applications. CRC Press (2014).
Acknowledgements
Authors express their deep gratitude to Steve Bayen (ILVO) for his excellent support in the database verifications and bioinformatics. Brigitte De Paepe and Cinzia Van Malderghem (ILVO) are gratefully acknowledged for their help in the molecular work.
Author information
Authors and Affiliations
Contributions
Redouan Qessaoui and Rachid Bouharroud conceived the experiments. Redouan Qessaoui, Rachid Bouharroud, Mohamed El Aalaoui, Hayat Akroud, Abderahim Amarraque, Rachid Tahzima and Johan Van Vaerenbergh conducted the experiments. Redouan Qessaoui, Rachid Bouharroud, Rachid Tahzima, Johan Van Vaerenbergh, El hassan Mayad, James N. Furze and Bouchra Chebli analyzed the results. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Qessaoui, R., Bouharroud, R., Furze, J.N. et al. Applications of New Rhizobacteria Pseudomonas Isolates in Agroecology via Fundamental Processes Complementing Plant Growth. Sci Rep 9, 12832 (2019). https://doi.org/10.1038/s41598-019-49216-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-49216-8
- Springer Nature Limited
This article is cited by
-
As(III)-oxidizing and plant growth-promoting bacteria increase the starch biosynthesis-related enzyme activity, 2-AP levels, and grain quality of arsenic-stressed rice plants
BMC Plant Biology (2024)
-
Uncovering the antifungal activities of wild apple-associated bacteria against two canker-causing fungi, Cytospora mali and C. parasitica
Scientific Reports (2024)
-
Physiological and Genomic Elucidation of Cold-Resilient Rhizobacteria Reveals Plant Growth Promotion by Siderophore Optimization and Enhanced Biocontrol Potential Against Fungal Pathogens
Journal of Plant Growth Regulation (2024)
-
Impact of Drought Stress on Plant Growth and Its Management Using Plant Growth Promoting Rhizobacteria
Indian Journal of Microbiology (2024)
-
Trunk injection of oxytetracycline improves plant performance and alters the active bark and rhizosphere microbiomes in huanglongbing-affected citrus trees
Biology and Fertility of Soils (2024)