Abstract
Plant growth-promoting rhizobacteria are known to have stimulating effects on plant growth and have become an emerging field of research in sustainable agriculture. The present study aims to select potent strains with a diverse set of in vitro plant growth-promoting (PGP) traits from the rhizosphere of Curcuma longa L cv. Lakadong and evaluates their growth-promoting ability in greenhouse experiment. A total of 33 isolates were obtained, purified, and subjected to preliminary in vitro screening for the production of IAA, ammonia, siderophore and phosphate solubilization. The isolates were further evaluated quantitatively for the production of IAA, solubilized phosphate, ammonia, and siderophore. Molecular identification of four selected isolates (BS1, BS4, IJ2, and IJ10) which showed promising in vitro PGP activity was done based on 16S rRNA gene analysis and their plant growth promotion was evaluated in greenhouse conditions. The isolates produced 2.67–52.88 μg ml−1 IAA, 31.56–320.50 μg ml−1 solubilized phosphate, 0.52–2.07 µmol ml−1 ammonia, and 16.99–43.30% of siderophore. The partial 16S rRNA sequence analysis identified the selected isolates as Arthrobacter sp. BS1, Pseudarthrobacter chlorophenolicus BS4, Pseudomonas sp. IJ2, and Bacillus subtilis IJ10. All the plants inoculated with Pseudomonas sp. IJ2 and B. subtilis IJ10 showed a significant difference at p ≤ 0.05 in all the growth parameters. These isolates, therefore, may provide a viable alternative to chemical inputs for improved growth and yield of C. longa.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Turmeric (Curcuma longa L.), a rhizomatous perennial herb belonging to the family Zingiberaceae is widely grown in India. Because of its high curcumin content (6.8–7.3%) [1], Lakadong may be considered as one of India’s finest varieties of turmeric. In addition to flavoring food, it is also used in medicines, cosmetics and as a dyeing agent [2]. Aggarwal and Sung [3] reported that curcumin has antioxidant, antimicrobial, anti-inflammatory, and anti-mutagenic properties.
The use of plant growth-promoting rhizobacteria (PGPR) covers a small yet growing niche in organic farming and is expected to increase in the future [4]. PGPR may induce plant growth and yield by either direct or indirect modes of action [5]. Direct mechanisms include the production of phytohormones, fixation of nitrogen, solubilization of inorganic phosphate, production of siderophores, regulating ethylene concentration, etc. [6], and indirect modes include the production of antibiotics, hydrogen cyanide, lytic enzymes such as chitinase, antifungal metabolites, competition, and systemic resistance induction [7].
Chemical fertilizers have been extensively used for many years and are detrimental to soil and human health, impacting soil fertility and altering the microbial population [8]. As an alternative solution to these chemicals, different agricultural practices have employed new biotechnological methods, which not only increase crop production but also preserve soil quality [9]. The use of PGPR as bioinoculants would be an ideal alternative for minimizing the indiscriminate use of chemical fertilizers [10].
The present study aims to select potent strains with a diverse set of in vitro plant growth-promoting (PGP) traits from the rhizosphere of C. longa and tests them under greenhouse conditions for their growth-promoting ability toward the crop.
Material and Methods
Sample Collection and Isolation of Rhizobacteria
Rhizospheric soil sample of C. longa cv. Lakadong was collected from Shangpung village, West Jaintia Hills District of Meghalaya (25°18’ N latitudes and 92°22’ E longitudes). Briefly, the soil attaching the root system was removed by shaking and subsequent brushing of the remaining soil particles in autoclaved plastic bags. The soil sample was then brought to the laboratory, labeled, kept at 4 °C in the refrigerator, and processed within 48 h after collection.
Rhizobacteria were isolated by serial dilution technique [11]. For isolation, 1 g of soil was transferred to a culture tube containing 10 ml of sterile distilled water. The tube was vortexed for 5 min. Dilutions of up to 10–6 grades were made. Hundred microliters each of 10–4, 10–5, and 10–6 dilution was spread on nutrient agar plates and incubated at 30 °C for 24–48 h. Bacterial isolates with different morphological appearances were purified, maintained and stored for further studies.
In vitro Evaluation of PGP Traits
Indole Acetic Acid (IAA) Production
The production of IAA by bacterial isolates was determined using Salkowski’s reagent based spectrophotometric method [12]. Bacterial isolates were grown in Luria Bertani broth supplemented with Tryptophan (100 μg ml−1) at 30 ± 2 °C for 72 h. Culture supernatants from each of these isolates were mixed with Salkowski’s reagent (49 ml of 35% of perchloric acid, 1 ml 0.5 M FeCl3 solution) in the ratio of 1:2. The pink color development indicates positive production of IAA and its optical density was measured at 530 nm using Lambda 35 UV–VIS spectrophotometer (PerkinElmer, USA). The concentration of IAA produced was estimated against the standard curve of IAA (Himedia, India).
Phosphate Solubilization
For determining phosphate solubilizing activity, rhizobacterial isolates were spot inoculated on Pikovskaya’s agar medium containing 0.04% Bromo Cresol Green as a pH indicator (Himedia, India). Isolates that developed a clear zone around it were considered positive after 3–5 days of incubation at 30 ± 2 °C. Quantitative estimation of solubilized P was determined by the method described by Subba Rao [13] in Pikovskaya broth containing 1000 μg ml−1 tricalcium phosphate. Using the Lambda 35 UV–VIS spectrophotometer (PerkinElmer, USA), absorbance for each isolate was measured at 430 nm and the concentration of solubilized P was extrapolated from the standard curve of potassium dihydrogen phosphate.
Ammonia Production
Ammonia production was determined both qualitatively and quantitatively using the method of Cappuccino et al. [14]. Bacterial isolates were inoculated in test tubes containing 5 ml of peptone water and incubated for 48 h at 30 ± 2 °C. The culture was centrifuged at 10,000 rpm for 5 min after incubation and 1 ml of Nessler's reagent was added to the supernatant in each tube. The formation of a yellow to brownish color is positive for the production of ammonia. The production of ammonia was measured spectrophotometrically at 450 nm using the Lambda 35 UV–VIS spectrophotometer, and the amount of ammonia produced was extrapolated from the standard curve of ammonium sulfate.
Siderophore Production
Siderophore production was analyzed using Chrome Azurol S (CAS) agar plate assay [15]. Each rhizobacterial isolate was spot inoculated on CAS agar plates and incubated at 30 ± 2 °C. After 48–72 h of incubation, an orange halo around the colonies indicates the production of siderophores. Quantitatively, siderophore content was estimated by CAS-Shuttle assay [16], in which 0.5 ml of culture supernatant was mixed with 0.5 ml of CAS reagent and the absorbance was recorded at 630 nm against a reference consisting of 0.5 ml of uninoculated broth and 0.5 ml of CAS reagent. The quantitative production of siderophores was calculated using the formula:
where Areference = absorbance of the reference, and Asample = absorbance of the sample.
Screening of Hydrolytic Enzymes
Cellulase activity: The production of cellulase was assessed using the method described by Cattelan et al. [17]. Pure culture isolates were inoculated in nutrient agar amended with 1% carboxymethylcellulose (CMC) and incubated at 30 ± 2 °C for 48–72 h. Extracellular cellulase was detected by flooding the plates with 0.1% congo red solution for 15 min and the plates were destained with 1 M NaCl solution for 15 min. The plates were then observed for positive cellulase production.
Protease activity: Protease activity was detected according to Smibert et al. [18]. Skim milk agar was used as the medium containing 0.04% of Bromo Cresol Green as a pH indicator (Himedia, India). Isolates forming a halo zone around the colonies specify positive protease activity after 48–72 h of growth at 30 ± 2 °C.
Molecular Identification of Selected PGPR
For molecular identification, DNA was extracted using the lysozyme method. Briefly, 2 ml overnight culture was taken in a 2 ml centrifuge tube and centrifuged at 13,000 rpm for 5 min. The pellet was washed twice with 0.5 ml sterile distilled water. To the pellet, 3.3 µl of 3 mg ml−1 lysozyme and 96.7 µl sterile water were added and mixed by vortexing. The tube was incubated at 37 °C for 1 h and 95 °C for 5 min. Centrifugation was done at 13,000 rpm for 5 min and the supernatant was transferred to a fresh tube. The 16S rRNA gene PCR amplification was performed using universal primers 27F 5' AGAGTTTGATCMTGGCTCAG 3' and 1492R 5' TACGGYTACCTTGTTACGACTT 3' [19]. The PCR mixture of 25 μl final volume consists of 1X PCR buffer with 1.5 mM MgCl2, 200 μM dNTPs, 0.2 μM of each primer, 3 U Taq DNA polymerase and 50 ng template DNA was performed in a thermal cycler (Bio-Rad C1000 touch, USA). The amplicons were separated on 0.8% agarose gel containing ethidium bromide (10 mg ml−1) in 1X TBE buffer. The amplified products were sequenced at Macrogens Inc., Korea. The sequences obtained were identified by using NCBI nucleotide BLAST analysis.
Preparation of PGPR Inoculum and Treatment
Four potent isolates were used as PGPR inoculum. Briefly, each bacterial isolate was cultured in a 500ml Erlenmeyer flask containing 250 ml of nutrient broth (Himedia, India) and incubated in an orbital rotary incubator at 150 rpm for 48–72 h at 30ºC. For the treatment, bacterial inoculum load was adjusted to 108 CFU ml−1 using sterile distilled water. The sprouted rhizomes were surface sterilized using 2% sodium hypochlorite solution for 1 min and soaked in the bacterial solution for 12 h. The treated rhizomes were air-dried under a sterile airflow. The rhizomes were then planted in polythene bags containing 5 kg of autoclaved soil. 250 ml of each PGPR inoculum (108 CFU ml−1) was added 30 and 60 days after planting (DAP) as a booster dosage. The growth parameters were analyzed after 180 DAP.
Statistical Analysis
The statistical analysis was performed using SPSS, v.25 (Chicago, IL, USA) and MS-Excel v.2007 (Microsoft, Washington, DC, USA). The data obtained were subjected to analysis of variance (ANOVA) and Tukey’s HSD test at p ≤ 0.05 using replicates to evaluate the effects of PGPR on the growth of C. longa.
Results and Discussion
Isolation of Rhizobacteria
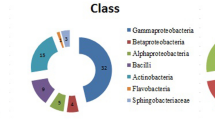
A total of 33 isolates were obtained from the rhizosphere of C. longa. The number of rhizobacteria isolated is summarized in Table 1.
In vitro Evaluation of PGP Traits
Preliminary Screening of PGP Traits
All the isolates were screened for different PGP traits and it was found that 22 isolates were positive for IAA production, 14 for phosphate solubilization, 29 for ammonia production and 21 for siderophore production. Moreover, eight isolates were positive to all the PGP traits tested while one isolate did not exhibit any of the traits. All the PGP traits tested are summarized in Table 1.
Quantitative Estimation of PGP Traits
The 32 isolates which were positive to at least one of the PGP traits were further quantified for the production of IAA, solubilized phosphate, ammonia, and siderophore. The isolates produced 2.67–52.88 μg ml−1 IAA, 31.56–320.50 μg ml−1 solubilized phosphate, 0.52–2.07 µmol ml−1 ammonia, and 16.99–43.30% of siderophore (Table 1, Fig. 1).
The production of IAA by PGPR is responsible for an increased number of adventitious roots, length and root volume through which roots can provide the plant with a large number of nutrients [20]. In our study, Pseudarthrobacter chlorophenolicus BS4 exhibits the highest IAA production (52.88 μg ml−1). Forni et al. [21] documented that, using tryptophan as a precursor, all Arthrobacter species were able to produce IAA. Another important PGP trait is inorganic phosphate solubilization in which bacteria produced several organic acids that lower the pH and released the phosphates that are attached to inorganic calcium complexes. Pseudomonas sp. IJ2 was found to solubilize the highest inorganic phosphate (320.50 μg ml−1). Different free-living rhizospheric bacteria like Bacillus, Pseudomonas and Azotobacter have been documented as excellent phosphate solubilizers and siderophore producers [7]. Siderophores are low molecular weight molecules which bind to Fe3+ and chelate Fe3+ making it less accessible to other microorganisms. Bacillus subtilis IJ10 produced the highest (43.30%) siderophore among all the isolates. O'Sullivan and O'Gara [22] showed that by depriving the pathogen of iron nutrition, efficient siderophore producers can regulate several plant diseases, resulting in increased crop yield. In the study, Arthrobacter sp. BS1 produced the highest ammonia (2.07 µmol ml−1) content. The production of ammonia by PGPR also contributes to an increase in the root, shoot growth and biomass production through nitrogen accumulation [23].
Screening for the Production of Hydrolytic Enzymes
All the 33 isolates were screened for protease and cellulase enzyme activity, of which 16 isolates were protease positive and 15 were cellulase positive (Table 1, Supplementary Fig. 1). The production of hydrolytic enzymes such as cellulase and protease by PGPR may play an important role in the decomposition of organic matter and nutrient mineralization and may also help to promote the entry of microorganisms into host tissues [24].
Molecular Identification of Selected PGPR
Molecular identification of four selected isolates was performed by amplifying the 16S rRNA gene and the sequences were obtained from Macrogens Inc., Korea. The sequences obtained were identified using NCBI BLAST analysis. The analysis identified the isolates as Arthrobacter sp. BS1, Pseudarthrobacter chlorophenolicus BS4, Pseudomonas sp. IJ2, Bacillus subtilis IJ10, and the sequences were deposited in GenBank database (NCBI) under the respective accession numbers MT192804, MT192880, MT193010, and MT219296.
Effects of PGPR Treatment on Plant Growth
Four selected isolates (BS1, BS4, IJ2, and IJ10) which showed promising in vitro PGP activity were considered for the pot trial experiment. Plants inoculated with the selected isolates exhibit better growth performance than uninoculated (control) plants (Fig. 2 and Supplementary Fig. 2). All the plants inoculated with Pseudomonas sp. IJ2 and B. subtilis IJ10 showed a significant difference at p ≤ 0.05 in all the growth parameters relative to control (Table 2, Fig. 2). This may be due to their different PGP traits. The primary factors in enhancing plant growth and yield have been attributed to these multiple PGP characteristics [25]. In a similar study, Kumar et al. [26] found that P. fluorescens CL12 inoculation resulted in better growth and yield of turmeric rhizome. Upon inoculation with Pseudomonas sp., chickpea and green gram resulted in an enhanced growth [27]. The species complex of B. subtilis is well known for its ability to promote plant growth, especially B. subtilis and B. amyloliquefaciens [28]. Such organisms encourage plant growth directly through the production of phytohormones, siderophores, organic acids involved in P-solubilization, and/or nitrogen fixation [29]. The root-colonizing B. subtilis subsp. subtilis PTS-394G is capable of persisting on the roots and of stimulating tomato plant growth [30].
Conclusion
The study established the value of isolating and screening rhizobacteria for various PGP traits and evaluating them for their ability to promote plant growth toward the crop in greenhouse environment. On the basis of greenhouse experiment, Pseudomonas sp. IJ2 and B. subtilis IJ10 were able to significantly enhance the growth of C. longa. These isolates, therefore, may provide a viable alternative to chemical inputs for improved growth and yield of C. longa.
References
Kanjilal PB, Kotoky R, Singh RS (2002) Morphological and chemical parameter of certain cultivars of chilli, ginger and turmeric grown in Meghalaya. Adv Plant Sci 15(1):225–229
Chattopadhyay I, Kaushik B, Bandyopadhyay U, Banerjee RK (2004) Turmeric and curcumin: biological actions and medicinal applications. Curr Sci 87(1):44–51
Aggarwal BB, Sung B (2009) Pharmacological basis for the role of curcumin in chronic diseases: an age-old spice with modern targets trends. Trends Pharmacol Sci 30:85–94. https://doi.org/10.1016/j.tips.2008.11.002
Lanong S, Kharshandi F, Kayang H, Syiem D (2018) Report on isolation of plant growth promoting bacteria (PGPB) from the gut of lesser horseshoe bat collected from the North-Eastern part of India. Int J Adv Sci Res Manag 3(12):78–89
Glick BR (1995) The enhancement of plant growth by free-living bacteria. Can J Microbiol 41:109–117. https://doi.org/10.1139/m95-095
Glick BR, Patten CL, Holguin G, Penrose DM (1999) Biochemical and Genetic mechanisms used by plant growth promoting bacteria. Imperial College Press, London
Ahmad F, Ahmad I, Khan MS (2008) Screening of free-living rhizospheric bacteria for their multiple plant growth promoting activities. Microbiol Res 163:173–181. https://doi.org/10.1016/j.micres.2006.04.001
Islam MR, Sultana T, Joe MM, Yim W, Cho JC, Sa T (2013) Nitrogen-fixing bacteria with multiple plant growth-promoting activities enhance growth of tomato and red pepper. J Basic Microbiol 53:1004–1015. https://doi.org/10.1002/jobm.201200141
Fernando WGD, Nakkeeran S, Zhang Y (2005) Biosynthesis of antibiotics by PGPR and its relation in biocontrol of plant diseases. In: Siddiqui ZA (ed) PGPR biocontrol and biofertilization. Springer, Dordrecht, pp 67–109
Ali B, Sabri AN, Hasnain S (2010) Rhizobacterial potential to alter auxin content and growth of Vigna radiata L. World J Microb Biot 26(5):1379–1384. https://doi.org/10.1007/s11274-010-0310-1
Johnson LF, Curl AE (1972) In: Method for the research on ecology of soil borne plant pathogens. Minneapolis Burgess Publishing Company, pp 247
Brick JM, Bostock RM, Silverstone SE (1991) Rapid in situ assay for indole acetic acid production by bacteria immobilized on nitrocellulose membrane. Appl Environ Microbiol 57:535–538
Subbarao NS (1988) Phosphate solubilizing microorganism. In: Biofertilizer in agriculture and forestry. Regional Biofert. Dev. Centre, Hissar, India, pp 133–142
Cappuccino JC, Sherman N (1992) A Laboratory Manual. In: Microbiology. New York, pp 125–179
Schwyn B, Neilands JB (1987) Universal chemical assay for the detection and determination of siderophores. Anal Biochem 160:47–56. https://doi.org/10.1016/0003-2697(87)90612-9
Payne SM (1994) Detection, isolation and characterization of siderophore. Methods Enzymol 235:329–344. https://doi.org/10.1016/0076-6879(94)35151-1
Cattelan AJ, Hartel PG, Fuhrmann JJ (1999) Screening for plant growth-promoting rhizobacteria to promote early soybean growth. Soil Sci Soc Am J 63:1670–1680. https://doi.org/10.2136/sssaj1999.6361670x
Smibert RM, Krieg NR (1994) Phenotypic characterization. In: Gerhardt P, Murray RGE, Wood WA, Krieg NR (eds) Methods for general and molecular bacteriology. American Society of Microbiology, Washington DC, pp 607–654
Heuer H, Krsek M, Baker P, Smalla K, Wellington EM (1997) Analysis of actinomycete communities by specific amplification of genes encoding 16S rRNA and gel-electrophoretic separation in denaturing gradients. Appl Environ Microbiol 63:3233–3241
Ramos SB, Barriuso J, Gutierrez MFJ (2008) Physiological and molecular mechanisms of plant growth promoting rhizobacteria (PGPR). In: Ahmad I, Pichtel J, Hayat S (eds) Plant-bacteria interactions: strategies and techniques to promote plant growth. Wiley-VCH, Weinheim, pp 41–54
Forni C, Riov J, GrillicCaiola M, Tel-Or E (1992) Indole-3-acetic acid (IAA) production by Arthrobacter species isolated from Azolla. J Gen Microbiol 138:377–381. https://doi.org/10.1099/00221287-138-2-377
O’Sullivan DJ, O’Gara F (1992) Traits of fluorescent Pseudomonas sp. involved in suppression of plant root pathogens. Microbiol Rev 56:662–676. https://doi.org/10.1128/MMBR.56.4.662-676.1992
Marques APGC, Pires C, Moreira H, Rangel AOSS, Castro PML (2010) Assessment of the plant growth promotion abilities of six bacterial isolates using Zea mays as indicator plant. Soil Biol Biochem 42:1229–1235. https://doi.org/10.1016/j.soilbio.2010.04.014
Lima LHC, Marco JL, Felix JR (1998) Enzimas hidroliticas envolvidas no controle biologic por miciparasitisma. In: Melo IS, Azvedo JL (eds) Controle Biologic. EMBRAPA-Meio Ambiente 11, Jaguraiuna, pp 263–304
Bashan Y, De-Bashan LE (2010) How the plant growth-promoting bacterium Azospirillum promotes plant growth-a critical assessment. Adv Agron 108:77–136. https://doi.org/10.1016/S0065-2113(10)08002-8
Kumar A, Vandana RS, Singh M, Singh PP, Singh SK, Singh PK, Pandey KD (2016) Isolation of plant growth promoting rhizobacteria and their impact on growth and curcumin content in Curcuma longa L. Biocatal Agric Biotechnol 8:1–7. https://doi.org/10.1016/j.bcab.2016.07.002
Goswami D, Vaghela H, Parmar S, Dhandhukia P, Thakker JN (2013) Plant growth promoting potential of Pseudomonas sp. OG isolated from marine water. J plant Interact 8(4):281–290. https://doi.org/10.1080/17429145.2013.768360
Fritze D (2004) Taxonomy of the genus bacillus and related genera: the aerobic endospore-forming bacteria. Phytopathology 94(11):1245–1248. https://doi.org/10.1094/PHYTO.2004.94.11.1245
Kumar A, Guleria S, Mehta P, Walia A, Chauhan A, Shirkot CK (2015) Plant growth-promoting traits of phosphate solubilizing bacteria isolated from Hippophae rhamnoides L. (Sea-buckthorn) growing in cold desert Trans-Himalayan Lahul and Spiti regions of India. Acta Physiol Plant 37(3):1–12. https://doi.org/10.1007/s11738-015-1793-z
Qiao J, Yu X, Liang X, Liu Y, Borriss R, Liu Y (2017) Addition of plant-growth-promoting Bacillus subtilis PTS-394 on tomato rhizosphere has no durable impact on composition of root microbiome. BMC Microbiol 17:131. https://doi.org/10.1186/s12866-017-1039-x
Acknowledgements
The first author is thankful to University Grants Commission, India, under the NET-JRF Scheme for financial support.
Author information
Authors and Affiliations
Contributions
All the authors contributed to the study's conception and design. The first author (FK) performed sample collection, laboratory experiments, data analysis and writing of the manuscript. The second author (AK) assists in experiments related to molecular identification of the isolates. The third author (HK) supervised the research work and experimental design. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have declare no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Significance Statement The study found that Pseudomonas sp. IJ2 and B. subtilis IJ10 were able to significantly enhance the growth of C. longa. These isolates, therefore, may provide a viable alternative to chemical inputs for improved growth and yield of C. longa.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Kharshandi, F., Khyllep, A. & Kayang, H. Plant Growth-Promoting Rhizobacteria of Curcuma longa L. and Their Impact on its Growth. Proc. Natl. Acad. Sci., India, Sect. B Biol. Sci. 91, 769–776 (2021). https://doi.org/10.1007/s40011-021-01268-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40011-021-01268-5