Abstract
Proteomic studies were performed to identify proteins involved in the response of Oenothera glazioviana seedlings under Cu stress. Exposure of 28-d-old seedlings to 50 μM CuSO4 for 3 d led to inhibition of shoot and root growth as well as a considerable increase in the level of lipid peroxidation in the roots. Cu absorbed by O. glazioviana accumulated more easily in the root than in the shoot. Label-free proteomic analysis indicated 58 differentially abundant proteins (DAPs) of the total 3,149 proteins in the roots of O. glazioviana seedlings, of which 36 were upregulated and 22 were downregulated under Cu stress conditions. Gene Ontology analysis showed that most of the identified proteins could be annotated to signal transduction, detoxification, stress defence, carbohydrate, energy, and protein metabolism, development, and oxidoreduction. We also retrieved 13 proteins from the enriched Kyoto Encyclopaedia of Genes and Genomes and the protein-protein interaction databases related to various pathways, including the citric acid (CA) cycle. Application of exogenous CA to O. glazioviana seedlings exposed to Cu alleviated the stress symptoms. Overall, this study provided new insights into the molecular mechanisms of plant response to Cu at the protein level in relation to soil properties.
Similar content being viewed by others

Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Introduction
Soil pollution by heavy metals deteriorates due to anthropogenic activities (e.g., metallurgy industry and sewage water irrigation), and it is a major problem of global concern1. Excess heavy metals (e.g. Cd, As, Hg, Se and Mo) severely reduce crop yields and cause health problems in humans, since they enter the food chain due to bioaccumulation in the edible parts of the plants2. Copper (Cu), as an essential micronutrient for plants, plays key roles in the citric acid (CA) cycle, pyruvate metabolism, and cell wall metabolism3, 4. However, excess Cu induces phytotoxicity, leading to growth inhibition, stunting, leaf chlorosis, necrosis and lipid peroxidation in membrane5, 6. The toxic rationales of Cu are due to its combination with nucleic acids and enzyme active sites7, 8. In addition, Cu inhibits the absorption of other elements such as Fe9. Long-term exposure to Cu results in low vegetation coverage and density10, thus, it is necessary to develop new plant varieties to make full use of such soil.
A better understanding of plants responses to heavy metal stress might help to develop effective detoxification measures and identify stress-tolerant genes or proteins11. Although the tolerance to Cu stress has been studied extensively at the phenotypic, physiological, and genetic level, and many candidate genes associated with heavy metal detoxification, tolerance, and stress response have been identified12, 13, the underlying mechanisms remain unclear, since gene expression is regulated at the transcriptional, translational, and post-translational level1, 14. Proteins have direct stress-acclimation functions that lead to changes in plasma membrane, cell cytoplasm, and the intracellular compartment composition15. Consequently, the plant response to heavy metal stress at the protein level needs further investigation.
Proteomics is one of the most advanced high-throughput biotechnological approaches that are used to address the biological function of proteins in response to different biotic or abiotic stresses16, 17. Previous proteomic studies on plant responses to heavy metal stress have mainly focused on Cd, Hg, and As18,19,20,21,22,23, whereas that fouced on Cu have been carried out in Arabidopsis thaliana, Agrostis capillaris L., Cannabis sativa, Elsholtzia splendens, Triticum aestivum L., and Oryza sativa 10, 24,25,26,27,28. The effects of heavy metals on plants vary with metal concentration and type, and also populations within a plant species. Therefore, further proteomic studies are needed in various species to investgate the molecular mechanisms of plants under Cu stress.
Oenothera glazioviana is a dominant species in the mine tailings of Tongling City, Anhui Province, China, which can efficiently stabilize Cu in the root and reduce its mobility and bioavailability29. Thus, O. glazioviana has been suggested as a potential candidate for the phytoexclusion of Cu-contaminated soils. However, little is known about the response mechanisms of O. glazioviana to Cu stress, especially at the protein level. In this study, a label-free quantitative proteomic approach based on nanoscale ultra-performance liquid chromatography tandem mass spectrometry (nano-UPLC-MS/MS) was conducted to identify Cu-responsive differentially abundant proteins (DAPs) in O. glazioviana. Our results in combination with physiological data might enhance our understanding regarding the interactions between O. glazioviana and Cu.
Results
Effects of Cu Stress on Phenotype and Growth Parameters
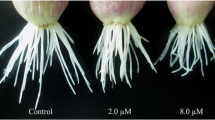
Oenothera glazioviana seedlings exposed to 50 μM CuSO4 for 3 d did not show any leaf chlorosis or withering symptoms. However, a considerable reduction in the shoot and root growth was observed compared with the control (Fig. 1). Quantitative analysis showed that the root length, root tip number, root surface area, root volume, and leaf surface area of Cu treated seedlings were lower decreased by 5.9%, 58.3%, 76.2%, 39.1%, and 4.4%, respectively, compared with those of the control (Table 1). In addition, the shoot fresh weight (SFW), root fresh weight (RFW), shoot dry weight (SDW), and root dry weight (RDW) of Cu treated seedlings were significantly reduced by 9.2%, 16.9%, 47.2%, and 14.8%, respectively, compared with those of the control. The magnitude of Cu stress was higher in the roots than in the shoots. As shown in Fig. 1, the root of Cu treated seedlings is normal with only a few slightly brown parts.
Levels of Thiobarbituric Acid Reactive Substances (TBARS) and Cu in Leaves and Roots
TBARS concentration in the shoot (4.53 ± 0.30 nmol g−1 FW) and the root (9.17 ± 0.43 nmol g−1 FW) of Cu treated seedlings was 1.15-fold and 2.03-fold higher, respectively, compared with that in the respective tissues of the control (3.77 ± 0.61 and 4.33 ± 1.04 nmol g-1 FW in the shoot and root, respectively) and also was 2.11-fold higher in the root than in the shoot (Fig. 2A). These results showed that the TBARS content in the root, but not in the shoot, was significantly affected by Cu stress. Similarly, the Cu concentration in the shoot (25.6 ± 11.7 μg g-1 DW) and the root (728.0 ± 223.7 μg∙g-1 DW) of Cu treated seedlings was 1.77-fold and 18.36-fold higher, respectively, compared with that in the respective tissues of the control (14.4 ± 5.1 and 39.6 ± 11.9 μg∙g-1 DW in the shoot and root, respectively) and also was 28.4-fold higher in the root than in the shoot (Fig. 2B).
Effects of copper stress on the levels of (A) thiobarbituric acid reactive substances (TBARS) and (B) copper (Cu) in the leaves and roots of Oenothera glazioviana seedlings. Different letters indicate significant differences at p < 0.05. Bars represent one standard error. Each experiment was conducted in triplicate.
Proteome in O. glazioviana Roots in Response to Cu Stress
Through label free-based shotgun quantification approach, a total of 3149 proteins was successfully identified in O. glazioviana seedlings that treated or not with Cu (Table S1). Of these, 58 proteins (1.8% of the total proteins) were classified as DAPs (Table S2); 36 proteins were upregulated and 22 proteins were downregulated in response to Cu stress (Table 2).
To gain a better understanding of the molecular functions and biological processes involved in O. glazioviana response to Cu stress, Gene Ontology (GO) analysis was performed and showed that DAPs were annotated to protein metabolism (18 DAPs), carbohydrate and energy metabolism (15 DAPs), signal transduction (eight DAPs), detoxification and stress defence (seven DAPs), development (five DAPs), oxidoreduction (three DAPs), and other unknown functions (two DAPs) (Fig. 3A).
(A) Gene Ontology (GO) and (B) Kyoto Encyclopaedia of Genes and Genomes (KEGG) analysis of 58 differentially expressed proteins (DAPs) in the roots of Oenothera glazioviana seedlings. Pie charts show the distribution of 58 DAPs on of the Cu-responsive proteins into their functional classes in percentage. Pathways are coloured from blue (lowest p value) to black (highest p value).
The Kyoto Encyclopaedia of Genes and Genomes (KEGG) analysis indicated that six pathways (involved 13 DAPs), including the CA cycle, carbon metabolism, pyruvate metabolism, fructose and mannose metabolism, glycolysis/gluconeogenesis, and amino sugar and nucleotide sugar metabolism, were significantly enriched (p < 0.01) (Fig. 3B; Table S3). The CA cycle was the most significantly enriched (p = 1.91e-05; Fig. 4; Table S4), and the citrate synthase was the most up-regulated among these 13 DAPs.
Interaction network of the Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathway and biological processes based on protein fold change at p < 0.01. Circle nodes refer to proteins (red, up-regulation; green, down-regulation). Rectangles refers to KEGG pathway or biological process (yellow, lowest p value; blue, highest p value).
Effect of Exogenous CA Application on Cu Tolerance
The application of exogenous CA to O. glazioviana seedlings exposed to 50 μM CuSO4 for 3 d greatly alleviated stress symptoms (Fig. 5). Quantitative analysis showed that the fresh and dry weights of Cu + CA treated seedlings were significantly higher than those of the Cu treated seedlings. However, the TBARS content in the root of Cu + CA treated seedlings was significantly lower than that in the root of the Cu treated seedlings (Table 3).
Discussion
Cu is an essential trace element in plants; however, in excess concentrations, it induces a wide range of biochemical effects and metabolic disturbances, which are responsible for a strong growth inhibition. The root growth is more susceptible to Cu toxicity than the shoot growth either the plant grows in the soil30 or in a culture solution31. In our study, O. glazioviana seedlings showed visible damage when exposed to 50 μM CuSO4 for 3 d. The roots became slightly brown (Fig. 1), and their growth was markedly inhibited. The root tip number, root surface area, root volume, and leaf surface area of Cu treated seedlings were significantly lower compared with those of the control (Table 1). The inhibitory effect of Cu on the root growth may be due to the reduced cell root meristem division and proliferation, damaged cell integrity in the root transition zone and retarded normal root cell growth32. These results were in agreement with those reported in findings in maize, hemp, and tobacco25, 33, 34.
Plants accumulate readily more Cu in the root than in other tissues such as leaves26. In Brassica napus, Cu accumulation increases significantly with Cu exposure and is higher in the root, followed by that in the stem and leaf35. In our study, the Cu content in the root and the shoot of Cu treated seedlings was 18.36-fold and 1.77-fold higher, respectively, compared with that in the respective tissues of the control (Fig. 2B), revealing the low translocation coefficient of Cu; thus, the shoot was less stressed. Excessive metal(loid) exposure, especially to Cu, generates reactive oxygen species (ROS) that damage the plant cells and inhibit plant growth36. TBARS, as a product of lipid peroxidation, is a sensitive biomaker of oxidative damage37. Here, the level of TBARS did not change significantly in the shoot, but significantly increased in the root of Cu treated seedlings (Fig. 2A), confirming previous studies in O. glazioviana 29 and suggesting that the tolerance/accumulation mechanism in the roots might restrict the root-to-shoot transfer of Cu.
The morphological and physiological changes exhibited in O. glazioviana seedlings exposed to 50 μM CuSO4 for 3 d suggested that the metabolic and biological processes are regulated by Cu application. Using the label free-based shotgun quantification method, we found that the abundance of 58 proteins significantly changed in Cu treated seedlings compared with the control. The Cu-responsive proteins were related to a wide range of molecular functions, including protein metabolism (31%), carbohydrate and energy metabolism (26%), signal transduction (14%), detoxification and stress defence (12%), development (9%), oxidoreduction (5%), and other unknown functions (3%). The observed diversity in the biological functions of DAPs suggested that the response of O. glazioviana to Cu stress might be a complex process, and some physiological and biochemical changes were altered to counteract the adverse conditions.
Protein Metabolism
Previous study in graph shown that Cu exposure markedly affects the protein metabolism and leads to protein reduction38. Here, 19 DAPs were identified in the roots of O. glazioviana seedlings exposed to Cu. Among these, the elongation factor Tu (No. 1; Fc = 6.69) catalyses the extension of the amino acid chain on the ribosome that further controls protein synthesis; heat shock proteins (No. 9, Fc = 1.55; No. 10, Fc = 2.43) increase in abundance under various abiotic stresses, since they prevent the aggregation of non-native proteins under normal and stress conditions39. Peptidyl-prolyl cis-trans isomerase and protein disulfide isomerase (No. 5, Fc = 0.55; No. 8, Fc = 0.64) play an important role in the maturation of newly synthesized proteins by correcting improper fold40; and ubiquitin-conjugating enzymes (UBCs, No. 7, Fc = 0.62) catalyse the second step in the ubiquitin-dependent proteolytic pathway that is one of the major protein degradation pathways in eukaryote. UBCs are induced under stress conditions and are responsible for the selective degradation of proteins with incorrect folding41. Under Cu stress conditions, we observed the up-regulation of No. 1, 9, and 10 that suggested the accumulation of damaged or misfolded proteins under Cu stress. Whereas the down-regulation of No. 5, 7, and 8 that indicated the synthesis of inappropriate proteins that led to the abnormal growth of O. glazioviana seedlings.
Carbohydrate and Energy Metabolism
The CA cycle is an important pathway in energy metabolism, responsible for the oxidation of respiratory substrates that lead to ATP synthesis and the adaptation to unfavourable environments42, 43. Here, we identified five proteins, citrate synthase 1 (No. 19, Fc = 3.17), succinyl-CA ligase subunit beta (No. 22, Fc = 1.55), pyruvate dehydrogenase E1 component subunit alpha (No. 20, Fc = 2.68), malate dehydrogenase, cytoplasmic 1 (No. 21, Fc = 2.28), dihydrolipoyl dehydrogenase 2, chloroplastic precursor (No. 28, Fc = 1.56), and 6-phosphogluconate dehydrogenase, decarboxylating 2, chloroplastic (No. 29, Fc = 1.79) that were involved in the CA cycle. Pyruvate dehydrogenase catalyzes the conversion of pyruvate to acetyl-CoA, and links the glycolysis pathway to the TCA cycle44. In previous studies, the expression of citrate synthase gene increased the citrate synthase activity and the citric acid content45. In the present study, we identified citrate synthase and used exogenous CA to experimentally verify its role in the alleviation of Cu stress symptoms. We also identified glucose-6-phosphate isomerase, cytosolic (No. 27, Fc = 1.57), which suggested that the glycolytic pathway might be involved in plant response to Cu stress. Shu et al. showed that enhanced glycolysis leads to the accumulation of acetyl-CoA in the CA cycle and the increased production of ATP to support stress resistance46. Here, most of the identified glycolysis-related proteins were up-regulated, indicating that O. glazioviana seedlings could maintain their essential respiration and provide more glycolytically generated ATP by reinforcing the CA cycle and glycolytic pathway under Cu stress conditions.
Signal Transduction
Many transporters, such as V-type proton ATPase subunit B1 (No. 36, Fc = 1.83), aquaporin (No. 37, Fc = 0.61), importin subunit alpha (No. 38, Fc = 0.57), and GDP dissociation inhibitor (No. 40, Fc = 2.40) were identified in the present study. V-type proton ATPase changes the H+ electrochemical gradient in the vacuole membrane47. Fukuda et al. shown that salt stress increases the transcription level of V-ATPase in the root of barley seedlings, which is beneficial for the ion accumulation in the vacuole48. Aquaporins are major water transporters that participate in the detoxification and compartmentalization of heavy metals49. The activity and expression of aquaporins and V-type proton ATPase can be affected by many external stimuli such as salinity50 and heavy metals51. Two small GTP-binding proteins, Ras-related protein RIC1 (No. 34, Fc = 0.34) and Ras-related protein RABH1b (No. 35, Fc = 0.38), play vital roles in signaling, the nuclear transportation of proteins and RNAs, and the regulation of cell cycle progression52. In the present, we found that Cu stress induced the up-regulation of Aquaporin that might influence the intracellular transport of Cu, as well as the up-regulation and activation of V-type proton ATPase that led to the excessive accumulation of Cu in the vacuole.
Detoxification and Stress Defence
We found several proteins related to cell detoxification, including Aldo-keto reductase (No. 44, Fc = 1.97) that is known to be effective in the detoxification of lipid peroxidation-derived reactive aldehydes53, 54. Transgenic tobacco plants overexpressing alfalfa AKR (MsALR) showed increased tolerance against a variety of oxidative stresses induced by methylviologen, heavy metals, and long-term drought53,54,55. Aldehyde dehydrogenase (No. 45, Fc = 1.71) is considered as a general detoxifying enzyme that eliminate toxic biogenic and xenobiotic aldehydes56. Cp-ALDH and Ath-ALDH3 from Craterostigma plantagineum and A. thaliana, respectively, respond to a variety of stress treatments57. Two ALDHs from barley were also shown to be up-regulated by drought stress58. Our proteomic analysis indicated ALDH might be associated with the removal of harmful substances under Cu stress in O. glazioviana seedlings.
Development
Translationally controlled tumour protein (No. 49, Fc = 0.38) is considered as a major regulator of cell growth in plants. Prohibitins (No. 50, Fc = 0.59) play an important role in root hair elongation, cell division, and development59. Methylmalonate-semialdehyde dehydrogenase (No. 51, Fc = 0.64) is a mitochondrial enzyme involved in the distal part of the valine and pyrimidine catabolic pathways. MMSDH was decreased in the seminal roots of slr1 mutants that were thinner compared with those of the wild type, supporting that MMSDH is a key factor in root development60. ADP-ribosylation factor (No. 53, Fc = 0.50) participates in membrane traffic, since it regulates the normal auxin efflux to exert a positive function in the cell polar localization61,62,63. The down-regulation of the ADP-ribosylation factor in A. thaliana results in severe growth inhibition64. In the present study, No. 49, 50, 51, and 53 were down-regulated in the roots of Cu treated seedlings, revealing that these proteins might be involved in growth inhibition.
Oxidoreduction
Cu, as a redox-active metal, can catalyse the formation of hydroxyl radicals to generate ROS that create oxidative stress and damage cellular macromolecules, resulting in cell death65. Plants have developed a vigorous antioxidant mechanism that is associated with enzymatic (peroxidase) and non-enzymatic components (glutathione). Here, phosphomannomutase (No. 54, Fc = 2.10), GDP-mannose 3,5-epimerase (No. 56, Fc = 1.90), and peroxidase 73 (No. 55, Fc = 1.86) that play crucial roles in ROS scavenging were accumulated in the roots of O. glazioviana seedlings exposed to Cu, suggesting that they might be associated with oxidative stress response.
Proteins do not perform their functions as single entities, but together in networks14. Meanwhile, signal molecules, usually help plants to recognize environmental factors, and regulate the expression of related genes in the signal pathways. When exogenous CA was applied to O. glazioviana seedlings exposed to 50 mM CuSO4, the stress symptoms were alleviated (Fig. S1, Table 3), indicating that CA might act as a signal molecule and regulate the expression of several proteins through a direct or indirect mechanism under Cu stress conditions.
Overall, our study showed that Cu stress inhibited the growth of O. glazioviana seedlings and increased the root Cu concentration. Our proteomic analysis identified 58 DAPs in the roots of O. glazioviana seedlings involved in protein metabolism, carbohydrate metabolism, signal transduction, detoxification and stress defence, development, and oxidoreduction. Using KEGG and PPI analysis, we identified 13 DAPs that were involved in different pathways. The CA cycle was the most significantly enriched, and then the citrate synthase exhibited most up-regulated among these 13 DAPs. These results suggested that CA might play a critical role in the overall plant response process to Cu. Subsequently, we applied exogenous CA to Cu treated seedlings in order to verify our assumption. We found that exogenous CA alleviated Cu stress symptoms, probably because it regulates the expression of proteins related to plant response to Cu stress. These results provided new insights into the molecular mechanisms of plant response to Cu.
Methods
Ethic Statement
No permissions were required for collecting O. glazioviana seeds from the Cu mine tailings in Tongling City, Anhui Province, China. O. glazioviana is not an endangered or protected plant species. The authors maintained the population at sustainable levels. The study was conducted following the national and international guidelines.
Plant Growth Conditions and Cu Treatments
The study design is shown in Fig. 6. O. glazioviana seeds were soaked in distilled water for 24 h and then, sown in plastic pots filled with vermiculite. The pots were placed in a growth chamber at a 12 h day/12 h night photoperiod, 20 °C day/25 °C night temperature, and light intensity of 250 μmol m−2 s−1. The cotyledons opened at approximately 7 d after sowing. The seedlings were fixed in cystose and transferred to vessels with 1 L of Hoagland’s nutrient solution, consisted of 5 mM Ca(NO3)2, 5 mM KNO3, 1 mM KH2PO4, 50 μM H3BO3, 1 mM MgSO4, 4.5 μM MnCl2, 3.8 μM ZnSO4, 0.32 μM CuSO4, 0.1 mM (NH4)6Mo7O24, and 10 μM Fe-ethylenediaminetetraacetic acid (EDTA). The nutrient solution was renewed every 3 d. After 21 d, the seedlings were exposed to 50 μM CuSO4 for 3 d. Each treatment (10 plants) was conducted in five replicates, and the control plants were grown in Hoagland’s nutrient solution without the addition of Cu. Plant roots and shoots were cut, pooled together, rinsed in deionized water, flash frozen in liquid nitrogen, and stored at −80 °C until analysis.
Growth Parameters
The maximum shoot length, SFW, and RFW were measured after 3 d of Cu exposure. Root and leaf samples were dried at 80 °C to constant weight for determining SDW and RDW. Root length, root tip number, root surface area, root volume, and leaf surface area were measured using a scanner-based image analysis system (WinRHIZO; Regent Instruments, Quebec, Canada)66. Prior to analysis, roots were preserved in 70% ethanol.
Determination of Cu Concentration
Root samples were collected and immerged in 25 mM EDTA-Na solution for 15 min to desorb metal ions on root surfaces. Next, root and leaf samples were washed thoroughly with tap water, rinsed with deionized water, cleaned with tissue paper, dried in an oven at 120 °C for 0.5 h to deactivate enzymes, and stored at 80 °C for 24 h. Next, these samples were ground to a fine powder, and 0.2 g was separately digested using an acid mixture of HNO3/HClO4 (87:13, v:v)67. The digests were dissolved in 5% HNO3 for Cu analysis using a NOVA 300 atomic absorption spectrophotometer (Analytik, Jena, Germany).
Determination of TBARS Levels
Lipid peroxidation was determined by estimating the levels of TBARS as described by Jin et al.37. Briefly, 0.5 g of fresh root tissues was homogenized in a mortar with 5 mL of 0.25% 2-thiobarbituric acid and 10% trichloroacetic acid. The mixture was heated at 95 °C for 30 min, quickly cooled in an ice bath, and centrifuged at 10,000 × g for 10 min. The absorbance of the supernatant was measured at 532 nm and corrected for unspecific absorbance at 600 nm.
Protein Extraction and Digestion
Root total proteins was extracted using a total protein extraction kit (Sigma-Aldrich, St. Louis, MO, USA), following the manufacturer’s instructions. Briefly, 250 mg of root tissue (10 plants pooled) was homogenized in liquid nitrogen. The homogenate was washed with methanol and acetone, and then, pelleted and dried with a SpeedVac (Thermo-Fisher Scientific, Waltham, MA, USA). The root tissue pellet was extracted with Type 4 Working Solution, containing 7 M urea, 2 M thiourea, 40 mM Trizma base, and 1% sodium dodecyl sulphate. After incubation for 15 min, the suspension was centrifuged at 14,000 × g for 30 min to remove the insoluble materials. The protein content in the supernatant was quantified using the Bradford assay (Bio-Rad, Hercules, CA, USA).
Protein samples (200 µg of bovine serum albumin equivalent) were digested using the filter-aided sample preparation method68. Briefly, the protein extracts were reduced by 10 mM dithiothreitol for 1 h at 56 °C, alkylated by 55 mM of iodoacetamide for 45 min at 25 °C in the dark, and buffer-exchanged with 100 mM NH4HCO3 (pH 8.5) using 10 KDa molecular weight cut-off Amicon Spin Tube (Millipore, Billerica, MA, USA). Subsequently, 4 µg of sequencing-grade modified trypsin (Promega, Madison, WA, USA) was added to each sample for protein digestion at 37 °C overnight (trypsin: protein, 1: 50). The digested peptides were desalted by Sep-Pak C18 cartridges (Waters, Milford, MA, USA) and quantified using a NanoDrop spectrophotometer (Thermo-Fisher Scientific).
Conditions of Nano-UPLC-MS
For label-free relative quantification analysis, five biological replicates of each treatment group were analysed by an on-line nano-LC system (Thermo-Fisher Scientific) coupled with a linear trap quadrupole mass spectrometer (LTQ-Orbitrap; Thermo Scientific). The resulting peptides (1.5 μg) were acidified with 0.1% formic acid and subsequently loaded into the nano trap column (Acclaim PepMap100 C18; 75 μm × 2 cm, 3 μm, 100 Å; Thermo-Fisher Scientific) at a flow rate of 4 μL min−1 in a loading buffer, containing 2% acetonitrile and 0.1% formic acid in high performance liquid chromatography-grade water. Chromatographic separation was carried out using an analytical column (Acclaim PepMap RSLC C18; 75 μm × 15 cm, 3 μm, 100 Å; Thermo-Fisher Scientific) with a linear gradient of 3–55% Buffer B (80% acetonitrile and 0.1% FA) at a flow rate of 0.25 μl min−1 over 112 min. Due to loading and washing steps, the total time for an LC-MS/MS run was approximately 160 min.
One scanning cycle included an MS1 scan (m/z 300–1800) at a resolution of 60,000, followed by 10 MS2 scans by LTQ. The 10 most abundant precursor ions were fragmented at 35%. The lock mass calibration was activated, and the dynamic exclusion time was 30 s.
Label-free Data Analysis
Raw MS files were processed by MaxQuant 1.5.2.5 employing the Andromeda algorithm and searched against the UniprotKB reference database for Viridiplantae (green plants) kingdom. In Andromeda search, the precursor and fragment ions mass tolerance was 6 ppm and 20 ppm, respectively. The maximum number of missed cleavages was two. The carbamidomethylation of cysteine was set as a fixed modification, with protein N-terminal oxidation of methionine as a variable modification. The false discovery rate (FDR) was set at 0.01. Protein abundances were calculated using the label-free quantitation algorithm69. Quantification was achieved using the label-free quantification (LFQ) with unique peptides. The match between runs option was enabled, allowing a time window of 2 min to search for already identified peptides in all obtained chromatograms. Protein abundance was calculated on the basis of the normalized spectral protein intensity (LFQ intensity), and proteins were quantified with a minimum of two ratio counts. The generated ‘proteingroups.txt’ table was filtered for contaminants, reverse hits, and number of unique peptides (≥1) using Perseus 1.5.3.2.
Bioinformatics Studies of DAPs
DAPs were characterized proteins with an average fold change in abundance (Cu/Control) more than 1.5 and a p value less than 0.05. GO annotations were retrieved from a large number of references, whereas KEGG70 and PPI analysis were performed using Omicsbean (http://www.omicsbean.cn). The strengths of the PPI network relationships were visualized by assigning line weights to the compiled scores. PPI analysis was done with minimum required interaction score set to medium confidence 0.40071.
Effect of Exogenous CA Application on O. glazioviana Seedlings Exposed to Cu
After 21 d in Hoagland’s nutrient solution, O. glazioviana seedlings were exposed to 50 μM Cu SO4 or 50 μM Cu SO4 and 50 μM CA for 3 d. Control plants were grown in Hoagland’s nutrient solution without Cu. FW, DW, and TBARS were determined as described above. Experiments were conducted in triplicate.
Statistical Analysis
One-way analysis of variance (ANOVA) in conjunction with Duncan’s test was performed to identify significant differences (p < 0.05) between the groups using SPSS 19.0 (IBM, Armonk, NY, USA). All data were expressed as mean ± standard deviation.
References
Ahsan, N., Renaut, J. & Komatsu, S. Recent developments in the application of proteomics to the analysis of plant responses to heavy metals. Proteomics 9, 2602–2621 (2009).
Cobbett, C. & Goldsbrough, P. Phytochelatins and metallothioneins: roles in heavy metal detoxification and homeostasis. Annu. Rev. Plant Biol. 53, 159–182 (2002).
Printz, B., Lutts, S., Hausman, J.-F. & Sergeant, K. Copper trafficking in plants and its implication on cell wall dynamics. Frontiers in plant science 7 (2016).
Yruela, I. Transition metals in plant photosynthesis. Metallomics 5, 1090–1109 (2013).
Nowicka, B., Pluciński, B., Kuczyńska, P. & Kruk, J. Physiological characterization of Chlamydomonas reinhardtii acclimated to chronic stress induced by Ag, Cd, Cr, Cu and Hg ions. Ecotoxicol. Environ. Saf. 130, 133–145 (2016).
Lequeux, H., Hermans, C., Lutts, S. & Verbruggen, N. Response to copper excess in Arabidopsis thaliana: impact on the root system architecture, hormone distribution, lignin accumulation and mineral profile. Plant Physiol. Biochem. 48, 673–682 (2010).
Hall, J. Cellular mechanisms for heavy metal detoxification and tolerance. J. Exp. Bot. 53, 1–11 (2002).
Himelblau, E. & Amasino, R. M. Delivering copper within plant cells. Curr. Opin. Plant Biol. 3, 205–210 (2000).
Marschner, H. & Rimmington, G. Mineral nutrition of higher plants. Plant Cell Environ. 11, 147–148 (1988).
Hego, E. et al. Differential accumulation of soluble proteins in roots of metallicolous and nonmetallicolous populations of Agrostis capillaris L. exposed to Cu. Proteomics 14, 1746–1758 (2014).
Benyó, D. et al. Physiological and molecular responses to heavy metal stresses suggest different detoxification mechanism of Populus deltoides and P. x canadensis. J. Plant Physiol. 201, 62–70 (2016).
Thapa, G., Sadhukhan, A., Panda, S. K. & Sahoo, L. Molecular mechanistic model of plant heavy metal tolerance. Biometals 25, 489–505 (2012).
Lin, C.-Y. et al. Comparison of early transcriptome responses to copper and cadmium in rice roots. Plant Mol. Biol. 81, 507–522 (2013).
Hao, J., Dong, C., Zhang, Z., Wang, X. & Shang, Q. Insights into salicylic acid responses in cucumber (Cucumis sativus L.) cotyledons based on a comparative proteomic analysis. Plant Sci. 187, 69–82 (2012).
Kosová, K., Vítámvás, P., Prášil, I. T. & Renaut, J. Plant proteome changes under abiotic stress-contribution of proteomics studies to understanding plant stress response. J. Proteomics 74, 1301–1322 (2011).
Chen, T. et al. iTRAQ-Based Quantitative Proteomic Analysis of Cotton Roots and Leaves Reveals Pathways Associated with Salt Stress. PLoS One 11, e0148487 (2016).
Lawrence II, S., Parker, J. & Chen, S. In Agricultural Proteomics Volume 2, 203–225 (Springer, 2016).
Wang, Y. et al. Proteomic Profiling of the Interactions of Cd/Zn in the Roots of Dwarf Polish Wheat (Triticum polonicum L.). Frontiers in Plant Science 7 (2016).
Zhang, Y.-W., Yan, L., Huang, L. & Huang, H.-Q. Cerebral ganglion ultrastructure and differential proteins revealed using proteomics in the aplysiid (Notarcus leachii cirrosus Stimpson) under cadmium and lead stress. Environ. Toxicol. Pharmacol. 46, 17–26 (2016).
Li, Y. et al. Comparative metalloproteomic approaches for the investigation proteins involved in the toxicity of inorganic and organic forms of mercury in rice (Oryza sativa L.) roots. Metallomics (2016).
Li, G. et al. Proteomic changes in maize as a response to heavy metal (lead) stress revealed by iTRAQ quantitative proteomics. Genetics and molecular research: GMR 15 (2016).
Tang, Z., Kang, Y., Wang, P. & Zhao, F.-J. Phytotoxicity and detoxification mechanism differ among inorganic and methylated arsenic species in Arabidopsis thaliana. Plant Soil 401, 243–257 (2016).
Ge, Y. et al. Quantitative proteomic analysis of Dunaliella salina upon acute arsenate exposure. Chemosphere 145, 112–118 (2016).
Gill, T., Dogra, V., Kumar, S., Ahuja, P. S. & Sreenivasulu, Y. Protein dynamics during seed germination under copper stress in Arabidopsis over-expressing Potentilla superoxide dismutase. J. Plant Res. 125, 165–172 (2012).
Bona, E., Marsano, F., Cavaletto, M. & Berta, G. Proteomic characterization of copper stress response in Cannabis sativa roots. Proteomics 7, 1121–1130 (2007).
Li, F. et al. Proteomic characterization of copper stress response in Elsholtzia splendens roots and leaves. Plant Mol. Biol. 71, 251–263 (2009).
Song, Y. et al. Proteomic analysis of copper stress responses in the roots of two rice (Oryza sativa L.) varieties differing in Cu tolerance. Plant Soil 366, 647–658 (2013).
Hego, E. et al. Copper stress-induced changes in leaf soluble proteome of Cu-sensitive and tolerant Agrostis capillaris L. populations. Proteomics 16, 1386–1397 (2016).
Guo, P. et al. Phytostabilization potential of evening primrose (Oenothera glazioviana) for copper-contaminated sites. Environmental Science and Pollution Research 21, 631–640 (2014).
Baszyński, T. et al. Photosynthetic apparatus of spinach exposed to excess copper. Zeitschrift für Pflanzenphysiologie 108, 385–395 (1982).
Stiborova, M., Doubravova, M., Brezinova, A. & Friedrich, A. Effect of heavy metal ions on growth and biochemical characteristics of photosynthesis of barley (Hordeum vulgare L.). Photosynthetica 20, 418–425 (1986).
Ernst, W. H., Nelissen, H. J. & Ten Bookum, W. M. Combination toxicology of metal-enriched soils: physiological responses of a Zn-and Cd-resistant ecotype of Silene vulgaris on polymetallic soils. Environ. Exp. Bot. 43, 55–71 (2000).
Aly, A. A. & Mohamed, A. A. The impact of copper ion on growth, thiol compounds and lipid peroxidation in two maize cultivars (Zea mays L.) grown in vitro. Australian Journal of Crop Science 6, 541 (2012).
Gori, P., Schiff, S., Santandrea, G. & Bennici, A. Response of shape in vitro cultures of shape Nicotiana tabacum L. to copper stress and selection of plants from Cu-tolerant callus. Plant Cell Tissue Organ Cult. 53, 161–169 (1998).
Herrero, E., López-Gonzálvez, A., Ruiz, M., Lucas-Garcia, J. & Barbas, C. Uptake and distribution of zinc, cadmium, lead and copper in Brassica napus var. oleifera and Helianthus annus grown in contaminated soils. Int. J. Phytoremediation 5, 153–167 (2003).
Hossain, M. A., Piyatida, P., da Silva, J. A. T. & Fujita, M. Molecular mechanism of heavy metal toxicity and tolerance in plants: central role of glutathione in detoxification of reactive oxygen species and methylglyoxal and in heavy metal chelation. Journal of Botany 2012 (2012).
Wu, Q., Su, N., Cai, J., Shen, Z. & Cui, J. Hydrogen-rich water enhances cadmium tolerance in Chinese cabbage by reducing cadmium uptake and increasing antioxidant capacities. J. Plant Physiol. 175, 174–182 (2015).
Llorens, N., Arola, L., Bladé, C. & Mas, A. Effects of copper exposure upon nitrogen metabolism in tissue cultured Vitis vinifera. Plant Sci. 160, 159–163 (2000).
Wang, W., Vinocur, B., Shoseyov, O. & Altman, A. Role of plant heat-shock proteins and molecular chaperones in the abiotic stress response. Trends Plant Sci. 9, 244–252 (2004).
Song, J. L. & Wang, C. C. Chaperone-like activity of protein disulfide-isomerase in the refolding of rhodanese. Eur. J. Biochem. 231, 312–316 (1995).
Garbarino, J. E., Oosumi, T. & Belknap, W. R. Isolation of a polyubiquitin promoter and its expression in transgenic potato plants. Plant Physiol. 109, 1371–1378 (1995).
Sweetlove, L. J., Beard, K. F., Nunes-Nesi, A., Fernie, A. R. & Ratcliffe, R. G. Not just a circle: flux modes in the plant TCA cycle. Trends Plant Sci. 15, 462–470 (2010).
Mailloux, R. J. et al. The tricarboxylic acid cycle, an ancient metabolic network with a novel twist. PLoS One 2, e690 (2007).
Jardine, K. J. et al. Gas phase measurements of pyruvic acid and its volatile metabolites. Environ. Sci. Technol. 44, 2454–2460 (2010).
Seki, M. et al. Monitoring the expression profiles of 7000 Arabidopsis genes under drought, cold and high-salinity stresses using a full-length cDNA microarray. The Plant Journal 31, 279–292 (2002).
Shu, L. et al. Genetic, proteomic and metabolic analysis of the regulation of energy storage in rice seedlings in response to drought. Proteomics 11, 4122–4138 (2011).
Fukuda, A. & Tanaka, Y. Effects of ABA, auxin, and gibberellin on the expression of genes for vacuolar H+-inorganic pyrophosphatase, H+-ATPase subunit A, and Na+/H+ antiporter in barley. Plant Physiol. Biochem. 44, 351–358 (2006).
Fukuda, A. et al. Effect of salt and osmotic stresses on the expression of genes for the vacuolar H+-pyrophosphatase, H+-ATPase subunit A, and Na+/H+ antiporter from barley. J. Exp. Bot. 55, 585–594 (2004).
Chrispeels, M. J. et al. Aquaporins of plants: structure, function, regulation, and role in plant water relations. Curr. Top. Membr 51, 277–334 (2001).
Muries, B., Carvajal, M. & del Carmen Martínez-Ballesta, M. Response of three broccoli cultivars to salt stress, in relation to water status and expression of two leaf aquaporins. Planta 237, 1297–1310 (2013).
Zhang, W.-H. & Tyerman, S. D. Inhibition of water channels by HgCl2 in intact wheat root cells. Plant Physiol. 120, 849–858 (1999).
Dasso, M. The Ran GTPase: theme and variations. Curr. Biol. 12, R502–R508 (2002).
Hideg, É., Nagy, T., Oberschall, A., Dudits, D. & Vass, I. Detoxification function of aldose/aldehyde reductase during drought and ultraviolet-B (280-320 nm) stresses. Plant Cell Environ. 26, 513–522 (2003).
Oberschall, A. et al. A novel aldose/aldehyde reductase protects transgenic plants against lipid peroxidation under chemical and drought stresses. The Plant Journal 24, 437–446 (2000).
Hegedüs, A. et al. Transgenic tobacco plants overproducing alfalfa aldose/aldehyde reductase show higher tolerance to low temperature and cadmium stress. Plant Sci. 166, 1329–1333 (2004).
Yoshida, A., Rzhetsky, A., Hsu, L. C. & Chang, C. Human aldehyde dehydrogenase gene family. Eur. J. Biochem. 251, 549–557 (1998).
Kirch, H. H., Nair, A. & Bartels, D. Novel ABA-and dehydration-inducible aldehyde dehydrogenase genes isolated from the resurrection plant Craterostigma plantagineum and Arabidopsis thaliana. The Plant Journal 28, 555–567 (2001).
Ozturk, Z. N. et al. Monitoring large-scale changes in transcript abundance in drought-and salt-stressed barley. Plant Mol. Biol. 48, 551–573 (2002).
Wang, Y., Ries, A., Wu, K., Yang, A. & Crawford, N. M. The Arabidopsis Prohibitin Gene PHB3 Functions in Nitric Oxide–Mediated Responses and in Hydrogen Peroxide–Induced Nitric Oxide Accumulation. The Plant Cell 22, 249–259 (2010).
Oguchi, K., Tanaka, N., Komatsu, S. & Akao, S. Methylmalonate-semialdehyde dehydrogenase is induced in auxin-stimulated and zinc-stimulated root formation in rice. Plant Cell Rep. 22, 848–858 (2004).
Koizumi, K. et al. VAN3 ARF–GAP-mediated vesicle transport is involved in leaf vascular network formation. Development 132, 1699–1711 (2005).
Robles, P. et al. The RON1/FRY1/SAL1 gene is required for leaf morphogenesis and venation patterning in Arabidopsis. Plant Physiol. 152, 1357–1372 (2010).
Sieburth, L. E. et al. SCARFACE encodes an ARF-GAP that is required for normal auxin efflux and vein patterning in Arabidopsis. The Plant Cell 18, 1396–1411 (2006).
Gebbie, L. K., Burn, J. E., Hocart, C. H. & Williamson, R. E. Genes encoding ADP-ribosylation factors in Arabidopsis thaliana L. Heyn.; genome analysis and antisense suppression. J. Exp. Bot. 56, 1079–1091 (2005).
Mattie, M. D. & Freedman, J. H. Copper-inducible transcription: regulation by metal-and oxidative stressresponsive pathways. American Journal of Physiology-Cell Physiology 286, C293–C301 (2004).
Wang, M.-B. & Zhang, Q. Issues in using the WinRHIZO system to determine physical characteristics of plant fine roots. Acta Ecologica Sinica 29, 136–138 (2009).
Fu, L. et al. Differences in copper absorption and accumulation between copper-exclusion and copper-enrichment plants: a comparison of structure and physiological responses. PLoS One 10, e0133424 (2015).
Wisniewski, J. R., Zougman, A., Nagaraj, N. & Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 6, 359 (2009).
Dislich, B. et al. Label-free quantitative proteomics of mouse cerebrospinal fluid detects β-site APP cleaving enzyme (BACE1) protease substrates in vivo. Mol. Cell. Proteomics 14, 2550–2563 (2015).
Kanehisa, M., Furumichi, M., Tanabe, M., Sato, Y. & Morishima, K. KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic acids research 45, D353–D361 (2017).
Brorsson, C. A. et al. Genetic risk score modelling for disease progression in new-onset type 1 diabetes patients: Increased genetic load of islet-expressed and cytokine-regulated candidate genes predicts poorer glycemic control. Journal of diabetes research 2016 (2016).
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2016YFD0800700), the National Natural Science Foundation of China (41571307; 31371545), the Science Foundation of Jiangsu Province, China (BE2016812; BE2015680; BE2016743).
Author information
Authors and Affiliations
Contributions
The authors have made the following declarations about their contributions: Y.C. and C.W. conceived the project. Y.C. and C.W. designed the study. C.W. performed the experiments. C.W., Y.C., Y.X., Z.S. and C.C. analyzed the data and C.W. wrote the article. C.W., J.W. and X.W. corrected the article.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wang, C., Wang, J., Wang, X. et al. Proteomic analysis on roots of Oenothera glazioviana under copper-stress conditions. Sci Rep 7, 10589 (2017). https://doi.org/10.1038/s41598-017-10370-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-017-10370-6
- Springer Nature Limited
This article is cited by
-
Phytotoxicity Evaluation of Five Proton-Pump Inhibitors Through Plant-Based Eukaryotic Test Models
Proceedings of the National Academy of Sciences, India Section B: Biological Sciences (2024)
-
Proteome insights of citric acid-mediated cadmium toxicity tolerance in Brassica napus L.
Environmental Science and Pollution Research (2023)