Abstract
Fruit ripening (FR) is attributed to the selective expression of several genes precisely governed by various specific transcription factors (TFs). The NAC (NAM, ATAF, and CUC) TF, MaNAC029, positively regulated banana ripening by directly inducing ethylene biosynthesis and transcription of fruit quality-related genes. However, its upstream regulatory mechanism still needs to be clarified. Herein, yeast one-hybrid screening revealed that a SQUAMOSA promoter binding protein-like (SPL) TF, MaSPL16, was a potentially upstream regulator of Musa acuminata NAC (NAM, ATAF, CUC) 029 (MaNAC029). Furthermore, gel mobility shift assay revealed that MaSPL16 can directly bound with the “GTAC” element of the MaNAC029 promoter. The gene expression and promoter activity assays demonstrated that Musa acuminata SPL (SQUAMOSA promoter binding protein-like) 16 (MaSPL16) expression was inducible by ethylene and ripening. MaSPL16 was localized to the nucleus, displayed a potenial capacity for transcriptional activation of MaNAC029. More critically, the transient expression of MaSPL16 in bananas accelerated FR via the upregulation of MaNAC029 and its downstream genes. Collectively, the mechanistic basis of a regulatory cascade involving MaSPL16-MaNAC029 that governed ethylene biosynthesis and fruit quality throughout the entire process of banana fruit ripening was unveiled. These outcomes increase the understanding of the gene-transcriptional regulatory mechanisms in FR. They are envisaged to help devise molecular techniques to regulate maturation and improve future fruit quality.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fruit ripening (FR) is a precisely orchestrated and irreversible process involving significant modifications in color, aroma, flavor, and texture, producing a palatable fruit that exhibits desirable quality-related characteristics (Wang et al. 2022). FR is modulated by the differential expression of several functionally essential genes, a mechanism tightly controlled by specific transcription factors (TFs) (Chen et al. 2020). In tomato, the non-ripening mutants, namely non-ripening (nor), ripening inhibitor (rin), and colorless non-ripening (cnr), have been studied extensively. Hence, the function of TFs in crucially regulating FR has increasingly garnered interest. The rin, nor, and cnr encode TFs that belong to MCM1, AG, DEF and SRF (MADS); SQUAMOSA promoter binding protein-like (SPL); and NAM, ATAF1/2, and CUC2 (NAC) families of TFs, respectively (Giovannoni 2007; Wang et al. 2020). Among these, RIN has been studied most thoroughly in various species. It has been well established that the targets of RIN are involved in multiple ripening-associated metabolic pathways, which include ethylene synthesis and perception, aromatic biosynthesis, metabolism of sucrose, and energy production (Qin et al. 2016; Ito et al. 2017; Li et al. 2020). The TFs, FUL2 and SlHY5, which belong to the MADS-box and bZIP families, respectively, favorably regulated ethylene production and improvement in fruit quality throughout the ripening process of tomatoes (Wang et al. 2014, 2021). Nevertheless, SlMADS1 and AP2a negatively influenced the maturation of fruits (Karlova et al. 2011; Dong et al. 2013). The other types of TFs, including bHLH, EIL and MYB, contributed to FR in apples, strawberries, kiwis, longans, and papayas (Yin et al. 2010; Tisza et al. 2010; Zhou et al. 2015; Kuang et al. 2012; Fu et al. 2019).
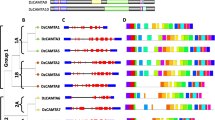
SPL is a highly influential family of TFs exclusive to the plant kingdom and regulates numerous biological processes, including development, senescence, and stress responses (Agarwal and Lahiri 2020; Jerome Jeyakumar et al. 2020). SBP is a conserved DNA-binding domain of SPLs with double zinc-finger structures (Lai et al. 2020; You et al. 2021). The contribution of SPLs to the modulation of FR was initially observed in the cnr mutant of tomato, characterized by a lack of color and ripening, exhibited a significant decrease in ethylene production, and lowered pigment accumulation due to the nor mutation. Consequently, the fruit remained colorless and failed to ripen (Thompson et al. 1999). The cnr mutation was linked to a transient epimutation that triggered DNA hypermethylation in the promoter of SQUAMOSA promoter binding protein-like-colorless non-ripening (SPL-CNR) (Zhong et al. 2013). Genome-encoded analyses revealed that ethylene insensitive 3 (EIN3) acts as a regulatory factor upstream of SPL-CNR, and EIN3-SPL-CNR constituted a positive feedback loop that governed ethylene biosynthesis (Lv et al. 2018; Gao et al. 2019). Additionally, the function of SPL-CNR in ripening is not only dependent on its nuclear localization signal (NLS) and zinc-finger motifs (ZFMs) but also regulated by phosphorylation catalyzed by the sucrose nonfermenting1-related protein kinase1 (SlSnRK1) (Lai et al. 2020). Ripening induced the differential expression of 15 other SPLs in tomatoes (Salinas et al. 2012). Several studies have demonstrated the evidence supporting the functional role of SPLs throughout the ripening of various fruits. For example, 14 Carica papaya L. SQUAMOSA promoter binding protein-like (CpSPL) and 12 Prunus avium L. SQUAMOSA promoter binding protein-like (PavSPLs) were systematically identified from the genomes of papaya and sweet cherry, respectively, and their expression patterns in several vegetative tissues and throughout FR were clearly described (Xu et al. 2020; Sun et al. 2023). Nevertheless, the specific regulatory mechanisms, especially the regulatory pathways, are still unclear.
Various TFs act synergistically or antagonistically to regulate gene expression (Bemer et al. 2017). In citrus fruits, CitWRKY1 cooperates with CitNAC62 to transactivate aconitase 3, ultimately promoting the degradation of citric acid throughout FR (Li et al. 2017). Conversely, LcR1MYB1 interacts with LcNAC13 to weaken the LcNAC13-based transrepression of the anthocyanin biosynthesis genes in FR in litchi (Jiang et al. 2019). In addition to protein–protein interactions, TFs compose a series of transcriptional cascades that control FR (Wang et al. 2020). For example, in apple, an induction in the expression of Malus domestica MYB (v-myb avian myeloblastosis viral) 9 (MdMYB9) and MdMYB11 was caused by the binding of MdNAC52 to their promoters, respectively, which in turn promoted the activating effect of MdMYB9 and MdMYB11 on proanthocyanidin and anthocyanidin biosynthesis pathways' genes in fruits (Sun et al. 2019). Similarly, PpNAC.A59, PpERF.A16, BLOOD, and PpMYB10.1 constitute a gene-transcriptional cascade that regulates fruit quality during ripening in peaches (Zhou et al. 2015; Guo et al. 2021). Thus, to construct a transcriptional regulatory network for FR, it is vital to determine the interrelationship between the ripening-related TFs.
Banana fruits (Musa acuminata, AAA group) are consumed globally and primarily cultivated in developing nations due to the specific climatic requirements for growth. Moreover, banana is a major commodity exported, generating billions of dollars in revenue, and is primarily consumed in developed countries (Aurore et al. 2009). Bananas are usually harvested at the mature green phase to enhance convenience during manufacturing and transportation. Subsequently, they are transported to the wholesale markets and ripened to achieve a desirable golden yellow color and an optimal edible quality. The ripening process was facilitated by applying ethylene, which contributed to determining the value of banana fruit as a commercial commodity (Pérez et al. 1997; Maduwanthi and Marapana, 2019). Therefore, much attention has been focused on understanding the regulatory mechanisms of postharvest FR in bananas. The involvement of various TFs, including bHLH (Xiao et al. 2018), EIL (Zhu et al. 2023) and MYB (Wei et al. 2023), in banana fruit ripening-associated transcriptional mechanisms has been elucidated. MaNAC029 directly impacted the expression of genes involved in ethylene biosynthesis and the degradation of cell walls, starch, and chlorophyll (Wei et al. 2022). Nevertheless, the upstream regulatory mechanisms governing the expression of MaNAC029 remain unclear. This study identified a specific SPL-encoding gene, MaSPL16, induced by ripening and ethylene. MaSPL16 bound putatively to the promoter of MaNAC029 and induced its transcriptional activation, which activated the genes associated with the biosynthesis of ethylene, determined fruit quality, promoted FR, and enhanced the quality of the bananas. These results identified a new cascade involved in transcriptional regulation, MaSPL16-MaNAC029, which governed the biosynthesis of ethylene and an improvement in fruit quality throughout the ripening process in bananas. These findings broadened the comprehension of the transcriptional modulation of FR.
Materials and methods
Plant material and treatments
The fruits of banana (Musa acuminata, AAA group, cv. Cavendish) were obtained from a plantation near Guangzhou, China, upon reaching a maturation level of 75% – 80% at 90 days after flowering. The bananas were separated randomly into untreated control (natural ripening), ethylene (100 μL/L), and 1-methylcyclopropene (1-MCP) treatment (0.5 μL/L) groups. Following treatment, all fruits were incubated at 20°C until complete ripening, as delineated in an earlier study (Zhu et al. 2020). The samples were immediately frozen in liquid N2 and cryopreserved at –80°C until subsequent utilization.
Gene expression analysis
The hot borate technique acquired the total RNA from the fruits. qRT-PCR was conducted using CFX96™ PCR (Bio-Rad, Hercules, CA, USA) and the Hieff® qPCR Kit (Yeasen, Shanghai, China). The primers were designed using the Primer software (ver 5.0) (Chen et al. 2011). MaRPS4 was employed as the internal reference gene.
Yeast one-hybrid (Y1-H) assay
The Matchmaker™ yeast one-hybrid system (Clontech, Mountain View, CA, USA) was employed to conduct the Y1-H screening. The pAbAi vector was utilized to construct the bait plasmid by inserting the short fragment, –750 bp upstream of “ATG” in the promoter of MaNAC029, followed by linearization. The construct was transposed into the Y1-H strain, resulting in the generation of a reporter strain specific to the bait, followed by the examination of a cDNA library of ethylene-treated bananas. The DNA–protein interactions were assessed by analyzing the maturation potential of co-transformants on an SD/-Leu medium fortified with Aureobasidin A (AbA), following the standardized protocols.
Electrophoretic mobility shift assay (EMSA)
The coding sequence (CDS) of MaSPL16 was cloned into the pGEX-4 T-1 vector with the appropriate reading frame and GST tag. Next, Glutathione S-transferase-Musa acuminata SPL (SQUAMOSA promoter binding protein-like) 16 (GST-MaSPL16) was expressed in the Escherichia coli strain, BM Rosetta and the fusion protein was purified using glutathione-sepharose 4B beads. The probes were derived from the fragments of the MaNAC029 promoter that were amplified and then labeled with biotin at the 5’ end. The EMSA kit (Thermo Scientific, Waltham, MA, USA) was employed, as published earlier (Wei et al. 2023). The recombinant GST-MaSPL16 protein was incubated with biotin-labeled probes, and the unbound probes were separated using a native acrylamide gel. The unlabeled probes were employed as the competitors, and GST protein as the negative control.
Assay of promoter activity and subcellular localization
The tobacco BY-2 protoplasts were attributed to conducting the assay of promoter activity and subcellular localization (Shan et al. 2020). The promoter of MaSPL16 was cloned into the pGreenII 0800-LUC reporter vector to activate the Firefly luciferase (LUC) expression. The protoplasts were transformed with the CaMV35S-REN/MaSPL16pro-LUC construct through the PEG-mediated approach and were treated with either 0 or 10 μL/L ethylene. A dual LUC assay kit (Promega, Madison, WI, USA) was utilized to detect the activities of LUC and renilla luciferase (REN).
The CDS of MaSPL16 was inserted into the pBI221-GFP vector to ascertain the subcellular location of MaSPL16 while employing the nucleus-targeted mCherry as a nuclear biomarker. The protoplasts were cotransfected with Musa acuminata SPL (SQUAMOSA promoter binding protein-like) 16-Green fluorescent protein (MaSPL16-GFP) and nuclear localization signal-monomer cherry red fluorescent protein (NLS-mCherry) based on a previously described method. An Axioskop fluorescence microscope (Zeiss, Oberkochen, Germany) was employed to study the fluorescence signals of GFP and mCherry.
Assay of transcriptional activation in yeast
A yeast two-hybrid (Y2-H) system (Clontech, Mountain View, CA, USA) was used to analyze the transcriptional activation of MaSPL16. The CDS of MaSPL16 was subcloned into the vector. The lithium acetate method was utilized to individually transform the cells of the Y2HGold yeast strain with the fusion (pGBKT7 + MaSPL16), positive control (pGBKT7-p53 + pGADT7-T-antigen), and negative control (pGBKT7) vector plasmids. The yeast cells were cultured on a selective medium, and the possible autoactivation was assessed by following the growth conditions and determining the α-galactosidase activity.
Dual-luciferase transient expression assay
The entire CDS of MaSPL16 was cloned into the pGreenII 62SK-BD vector (effector) to evaluate its capacity for transcriptional activation. The reporter vector was constructed with a dual-reporter system comprising the firefly LUC regulated by five GAL4-binding elements. Based on a previous report, the tobacco leaves were cotransformed with the effector and recombinant reporter constructs (Wei et al. 2023). The activities of LUC and REN were quantified after 60 h following Agrobacterium. tumefaciens injection as described in the promoter activity assay.
To evaluate the influence of MaSPL16 on the activity of the promoter of MaNAC029, the pGreenII 62-SK effector vector was utilized for the cloning of MaSPL16, and the pGreenII 0800-LUC reporter vector for the colonization of the promoter of MaNAC029 (Hellens et al. 2005). The A. tumefaciens strain, EHA105 (pSoup), was employed for the cotransformation of tobacco leaves with the effector and reporter plasmids. The ratio of LUC to REN reflected the transcriptional activation of the MaNAC029 promoter. Each pair was measured six times.
Transient overexpression in banana fruits
An analysis of the transient overexpression of MaSPL16 in banana fruits was performed as previously outlined (Shan et al. 2020). The open reading frame was cloned into the pCXUN-HA vector, which was used to transform the A. tumefaciens cells that were then used to inoculate the pulp of banana fruits. The transformed fruits were supplemented with 100 µL/L ethylene on the first-day post-inoculation and incubated at 20°C for seven days. Samples were collected to ascertain the gene expression levels. Fruit firmness, ethylene construction, color index, starch content, and chlorophyll levels were monitored at each predetermined sampling point, adopting a previously reported method (Wei et al. 2022).
Statistical analysis
The statistical analysis was performed using the SPSS software (ver. 19.0). The data were reported as the mean ± standard error (SE) from three or six replicates. The statistical significance of the variations between the specimens was evaluated through the Student’s t-test or the analysis of variance (ANOVA) as required.
Primers
Table S1 lists the primers used in this study.
Results
MaSPL16 directly targets the MaNAC029 promoter
MaNAC029 contributed to the biosynthesis of ethylene and determining fruit quality throughout the postharvest ripening period in banana fruits (Wei et al. 2022). The proteins interacting with the promoter of MaNAC029 were explored to determine upstream regulators by screening the Y1-H libraries. The cDNA fragment (Ma07_g24410) of an SPL-encoding gene was detected. A phylogenetic tree indicated that Ma07_g24410 was evolutionarily highly related to the rice OsSPL16 (Fig. S1a) and hence was designated MaSPL16. The Y1-H assay was employed to validate the interaction between MaSPL16 and the promoter of MaNAC029 by utilizing the complete CDS of MaSPL16 as prey. Even the basal activity of the MaNAC029 promoter could not be detected in yeast in the presence of AbA (Fig. 1a). In contrast, the yeast cells that express MaSPL16 exhibited enhanced resistance to AbA due to elevated gene expression attributable to the higher activity of the MaNAC029 promoter. They displayed strong growth on a medium containing AbA, confirming the physical interaction between MaSPL16 and the MaNAC029 promoter.
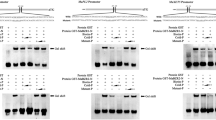
Musa acuminata SPL (SQUAMOSA promoter binding protein-like) 16 (MaSPL16) binds to the promoter of Musa acuminata NAC (NAM, ATAF, CUC) 029 (MaNAC029). (a) The effects of MaSPL16 on the MaNAC029 promoter ascertained through a Y1-H approach. Left: the MaNAC029 promoter was not expressed in yeast cultivated on SD media lacking Leu but supplemented with 700 ng/mL Aureobasidin A (AbA). Right: the yeast growth assay of Y1-H strains was performed after transformation with either the empty pGADT7 plasmid (negative control) or the one containing a cassette expressing the MaSPL16 effector. Their interaction was evaluated based on the capacity of transformed yeast cells to grow when cultivated on a Leu-deficient SD medium supplemented with AbA. (b) The binding of MaSPL16 to the MaNAC029 promoter was ascertained via EMSA. The GST or GST-MaSPL16 protein was incubated with the probes. Subsequently, the DNA–protein complexes obtained were segregated on native polyacrylamide gels. The competitors used comprised of unlabeled wild-type and mutated probes. Fig. S3 presents the uncropped and full-length blot for EMSA
An analysis of the sequence of the promoter of MaNAC029 indicated the presence of the “GTAC” core sequence (Text S1), a key element recognized by the SPLs (Kropat et al. 2005). Hence, an EMSA was conducted to ascertain the probable direct interaction between the MaSPL16 and the MaNAC029 promoter. The recombinant protein, GST-MaSPL16, was expressed in E. coli and purified (Fig. S2). The core “GTAC” sequence, which comprises the DNA segments derived from the promoter of MaNAC029, was employed as the probe. The results indicate that the recombinant protein, GST-MaSPL16 directly interacted with certain regions of the promoter of MaNAC029, thereby inducing evident changes in mobility (Fig. 1b). Additionally, the administration of increasing quantities of unlabeled probes with the same sequence caused a significant reduction in the displacement of bands. However, including mutants as competitors did not result in a similar decline, and no significant alterations in mobility were detected upon incubation with identical probes containing only GST. Collectively, these results suggest that MaSPL16 directly targets the MaNAC029 promoter via the “GTAC” core.
Ethylene and ripening induce the expression of MaSPL16
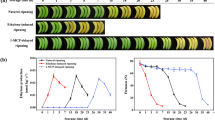
An analysis of the expression patterns throughout the ripening process was conducted to assess the possible association between MaSPL16 and FR. Bananas are climacteric fruits, and their ripening was triggered by ethylene, which was delayed by 1-MCP. A peak in ethylene production throughout the natural FR process occurred after 18 days of storage, with complete ripening on day 20. The ethylene treatment expedited FR, characterized by the appearance of an ethylene peak on day three and complete maturation on day five. Treatment with 1-MCP delayed the ethylene peak until day 30 and full ripening until day 35 (Zhu et al. 2020). Herein, the association between MaSPL16 and FR in bananas was explored by examining the expression patterns of MaSPL16 throughout the natural, ethylene-induced, and 1-MCP-delayed ripening processes. Figure 2a – c illustrated that the expression of MaSPL16 was remarkably induced by ethylene, with its transcript levels showing a tendency to rise markedly and then decrease during the three ripening processes, which was per ethylene production.
The expression of Musa acuminata SPL (SQUAMOSA promoter binding protein-like) 16 (MaSPL16) is induced by ethylene and ripening. (a) The expression of MaSPL16 in banana fruits was examined in natural ripening (b) Ethylene-induced ripening (c) Ripening delayed by 1-MCP treatment. The relative expression levels at each time point were referred to as the gene transcript levels in the control at the 0 day. The physiological data about fruit ripening and softening have been expounded upon in a prior investigation (Zhu et al. 2020). The open and closed triangles symbolized the initiation and culmination of ethylene production at different time points for each treatment, respectively. The data presented in this study represent the mean ± SE obtained from three independent replicates. The lowercase letters indicated the statistically significant differences observed among the different groups, as determined by a one-way ANOVA followed by Tukey’s test (P < 0.05). (d) The activity of the MaSPL16 promoter in response to ethylene was investigated. The reporter construct was transiently introduced into the tobacco BY-2 protoplasts and subsequently tested for induction by ethylene. The error bars in the graph indicate the SE calculated from six replicates (Student’s t-test, **P < 0.01). REN, Renilla luciferase; LUC, Firefly luciferase; CaMV-term, cauliflower mosaic virus terminator; Eth, ethylene
Additionally, a transient protoplast assay was employed to identify the impact of ethylene on the activity of the MaSPL16 promoter. The MaSPL16 promoter was isolated, fused with LUC, and cloned into the dual luciferase reporter vector. The 35S promoter was used to drive the expression of REN in the identical vector, which served as an internal control (Fig. 2d). The results revealed that the activity of the MaSPL16 promoter was induced by ethylene in tobacco BY-2 protoplasts transfected with CaMV35S-REN/MaSPL16 promoter-LUC, which was confirmed by a remarkable rise in the ratio of LUC/REN. In summary, these findings revealed that MaSPL16 was induced by ethylene and ripening.
Molecular characterization of MaSPL16
An analysis of the 154 – 169 amino-acid sequence of MaSPL16 indicated the existence of a nuclear localization signal (NLS) (Fig. S1b). The subcellular location of MaSPL16 was determined using a transient expression of GFP tagged with MaSPL16 in the tobacco BY-2 protoplasts. Subsequently, the nucleus-targeted mCherry (NLS-mCherry) was coexpressed as a control to observe the nucleus. The GFP-fluorescence was detected in both the cytoplasm and nucleus in the control group (Fig. 3a). The nuclear localization of MaSPL16 was confirmed by the colocalization of the green fluorescent signal emitted by MaSPL16-GFP and the red fluorescent signal emitted by NLS-mCherry (Fig. 3a).
Molecular characteristics of Musa acuminata SPL (SQUAMOSA promoter binding protein-like) 16 (MaSPL16). (a) The localization of MaSPL16 in the tobacco BY-2 protoplasts. The nuclear marker, the nucleus-targeted mCherry (NLS-mCherry) was employed in this study, bars = 25 μm. (b) Stimulation of the transcription of MaSPL16 in yeast cells. The pGBKT7 vector from BD was used as a negative control, while p53-BD + T-antigen-AD was used as a positive control. (c) The transcriptional activity of MaSPL16 in tobacco leaves. VP16 was employed as a positive control to establish a baseline. The ratio of LUC to REN activity in the empty pBD vector was normalized to 1, which served as the negative control. Error bars represent the SE from six replicates (Student’s t-test, **P < 0.01). GFP, Green fluorescent protein; LUC, Firefly luciferase; REN, Renilla luciferase
The capacity of MaSPL16 for transcriptional activation was evaluated through a yeast reporter system responsive to GAL4. MaSPL16 was fused with GAL4-binding domain (GAL4BD) and then expressed in Y2HGold yeast. The yeast cells that were successfully transformed with the DBD-p53, AD-T-antigen (positive control), and DBD-MaSPL16 vectors exhibited normal growth even in the absence of Trp (Fig. 3b). These cells exhibited α-Gal activity and were able to survive even in the absence of Trp, His and Ade. On the other hand, the yeast transformed with the negative control did not exhibit any growth (Fig. 3b). These findings suggest that MaSPL16 can activate self-transcription in yeast. Utilizing a dual-luciferase reporter system with 5 × GAL4 DNA-binding components, the transcriptional activation capacity of MaSPL16 was additionally investigated in planta. VP16, a robust transcriptional activator, was used as a positive control (Fig. 3c). In comparison to the empty BD vector, MaSPL16 enhanced the LUC/REN ratio significantly by 6.1-fold (Fig. 3c), thereby serving as additional evidence supporting the transcriptional activation potential of MaSPL16. The data presented suggested that MaSPL16 may possess the potential to act as a transcriptional activator contributing to FR in bananas.
MaSPL16 directly activates the transcription of MaNAC029
Since MaSPL16 exhibited the capability of transcriptional activation and was directly bound to the MaNAC029 promoter (Figs. 1, 3), a transient dual luciferase assay was conducted to examine the potential role of MaSPL16 in inducing the transcription of MaNAC029. The MaNAC029 promoter was fused with LUC, while the 35S promoter-driven REN was utilized as an internal control using the same vector for both (Fig. 4a). The effector construct contained the 35S promoter-driven MaSPL16 cDNA (Fig. 4a). The activity of the promoter of MaNAC029 enhanced remarkably upon cotransfection with the 35S::MaSPL16 effector, as compared to the empty effector (Fig. 4b), suggesting that MaSPL16 possessed the ability to transactivate the promoter of MaNAC029 as indicated by the relatively greater LUC/REN ratio.
Musa acuminata SPL (SQUAMOSA promoter binding protein-like) 16 (MaSPL16) activates the promoter of Musa acuminata NAC (NAM, ATAF, CUC) 029 MaNAC029. (a) Diagram illustrating the reporter and effector constructs. (b) MaSPL16 enhanced the activity of the MaNAC029 promoter. The Firefly luciferase (LUC) to Renilla luciferase (REN) ratio of an empty vector containing the MaNAC029 promoter was employed as a calibrator and designated 1. Error bars represent the SE from six replicates (Student’s t-test, **P < 0.01). CaMV-term, cauliflower mosaic virus terminator
Overexpression of MaSPL16 enhances the MaNAC029-based activation of ripening-related genes
MaNAC029 activated the genes involved in ethylene biosynthesis and the degradation of cell walls, starch, and chlorophyll throughout the FR process in bananas (Wei et al. 2022). As MaSPL16 bound directly to the promoter of MaNAC029 and induced its expression (Figs. 1, 4), it is reasonable to assume that MaSPL16 enhanced the MaNAC029-based transactivation of ripening-related genes. To verify this hypothesis, MaSPL16 was transiently overexpressed in banana fruits (Fig. 5a, b) which was confirmed by qRT-PCR (Fig. 5b). In contrast to the empty vector (control), the transient overexpression of MaSPL16 enhanced FR in banana, resulting in a faster yellowing (ripening) phenotype (Fig. 5a). In MaSPL16-over-expressing fruits, the decline in the color index was advanced by two days during maturation, and concomitantly, the ethylene production increased dramatically, while the firmness, chlorophyll content, and starch levels of fruits reduced throughout the ripening process (Fig. 5c). More importantly, the overexpression of MaSPL16, in turn, induced the enhanced expression of MaNAC029 in fruits throughout ripening (Fig. 5d). In parallel, the expression of target genes downstream of MaNAC029, were induced markedly in the MaSPL16-overexpressing fruits. These included Musa acuminata 1-amino-cyclopropane-1-carboxylic acid synthase 1 (MaACS1), Musa acuminata 1-amino-cyclopropane-1-carboxylic acid oxidase 1 (MaACO1) and MaACO13, involved in ethylene biosynthesis (Jourda et al. 2014); Musa acuminata Expansins 2 (MaEXP2), MaEXP15, Musa acuminata xyloglucan endotransglycosylase/hydrolases 28 (MaXTH28) and MaXTH30, associated with cell wall degradation (Han et al. 2016); Musa acuminata α-glucan water dikinase 1 (MaGWD1) related with starch degradation (Xiao et al. 2018); and Musa acuminata Stay-Green 1 (MaSGR1) and Musa acuminata Pheophytinase (MaPPH), involved in chlorophyll catabolism (Wei et al. 2023) (Fig. 6). These findings suggest that MaSPL16 induced the expression of MaNAC029, thereby enhancing the MaNAC029-based transcriptional activation of genes involved in ethylene biosynthesis, enhancement in fruit quality, promoting FR, and determination of quality in banana.
Transient upregulation of Musa acuminata SPL (SQUAMOSA promoter binding protein-like) 16 (MaSPL16) in banana fruits accelerates fruit ripening and determination of quality. (a) The appearance of banana fruits that transiently overexpressed MaSPL16 or the empty vector throughout ripening. (b) qRT-PCR indicating the levels of MaSPL16 mRNAs in these banana fruits. (c) Color index, ethylene production, firmness, starch levels, and total chlorophyll content in bananas as shown in (a). (d) Relative expression of Musa acuminata NAC (NAM, ATAF, CUC) 029 (MaNAC029) in bananas as shown in (a). Error bars in (b), (c) and (d) represent the SE from six and three replicates, respectively (Student’s t-test, **P < 0.01)
The expression of MaACS1, MaACO1, MaACO13, MaEXP2, MaEXP15, MaXTH28, MaXTH30, MaGWD1, MaSGR1 and MaPPH in the MaSPL16-overexpressing and control banana fruits. Error bars represent the SE from three replicates (Student’s t-test, **P < 0.01). MaACS1, Musa acuminata 1-amino-cyclopropane-1-carboxylic acid synthase 1; MaACO1, Musa acuminata 1-amino-cyclopropane-1-carboxylic acid oxidase 1; MaEXP2, Musa acuminata Expansins 2; MaXTH28, Musa acuminata xyloglucan endotransglycosylase/hydrolases 28; MaGWD1, Musa acuminata α-glucan water dikinase 1; MaSGR1, Musa acuminata Stay-Green 1; MaPPH, Musa acuminata Pheophytinase
Discussion
Typically, bananas are harvested upon reaching the mature green phase and ripened using ethylene. This process results in bananas attaining a desirable golden yellow color and optimal edible quality during the postharvest phase (Maduwanthi and Marapana 2019). Therefore, postharvest ripening determined the commodity value of banana fruits. The participation of several TFs precisely modulates the process of FR (Li et al. 2021), such as MaNAC029, which regulated the biosynthesis of ethylene and fruit quality in bananas throughout postharvest ripening (Wei et al. 2022). However, the regulatory mechanisms upstream of MaNAC029 are still not precise. Herein, MaSPL16, a transcription regulator upstream of MaNAC029, was identified, and the molecular mechanisms by which the MaSPL16-MaNAC029 transcriptional cascade module regulated banana postharvest ripening were discussed.
The three groups of TFs, namely, MADS, SPL, and NAC, have been acknowledged for their significant contribution to FR (Giovannoni et al. 2017; Li et al. 2019; Liu et al. 2022). An analysis of the genes encoding TFs revealed that NACs and MADSs form a dual-loop circuit, modulating FR in bananas (Lv et al. 2018). However, the correlation between the regulatory effects of SPLs and NACs is still not understood. The current investigation illustrated that MaSPL16 specifically interacted with the promoter of MaNAC029, resulting in the activation of its expression (Figs. 1, 4), indicating that MaSPL16 and MaNAC029 constituted a transcriptional cascade involved in banana ripening. These data enrich the knowledge of the interrelationship between ripening-correlated TFs and assist in building a transcriptional modulatory network for FR.
Banana, being a climacteric fruit, requires ethylene for its ripening. The investigation of the modulatory pathways of ethylene biosynthesis during FR in bananas has long been a prominent area of research (Tang et al. 2023). The direct modulation of the genes involved in ethylene biosynthesis throughout the FR process was attributed to the NAC TFs in several fruits, including tomatoes (Gao et al. 2018), kiwifruits (Wu et al. 2020), and peaches (Dai et al. 2023). The NAC TF, MaNAC029, induced by ripening and ethylene, directly influenced the expression of MaACS1, MaACO1 and MaACO13 (Wei et al. 2023). Nevertheless, the precise mechanism through which ethylene enhances the expression of MaNAC029 remains ambiguous. This study demonstrated that ethylene stimulated the expression of MaSPL16 by enhancing the activity of its promoter (Fig. 2). Moreover, MaSPL16 induced the MaNAC029-involved activation of MaACS1, MaACO1 and MaACO13, which in turn promoted ethylene production (Figs. 5, 6), that may serve as a feedback regulatory mechanism.
Fruit quality is a composite trait involving flavor, aroma, color, and texture that finally determines the commercial value of fruit (Gapper et al. 2013; Hu et al. 2019). Several TFs from various families have been identified to play a role in the modulation of FR in bananas via the direct control of the expression of genes associated with fruit quality. For instance, Musa acuminata basic helix-loop-helix 6 (MabHLH6) and Musa acuminata MYB (v-myb avian myeloblastosis viral) 3 (MaMYB3) directly regulated the genes correlated with starch degradation, where MabHLH6 promoted the degradation of starch during FR, while MaMYB3 inhibited it (Fan et al. 2018; Xiao et al. 2018). MaMYB60 and Musa acuminata ethylene insensitive 3-like 9 (MaEIL9) serve as upstream regulators of chlorophyll degradation and carotenoid biosynthesis, respectively, during the ripening of bananas (Wei et al. 2023; Zhu et al. 2023). Nevertheless, the upstream transcriptional modulators of the genes encoding these TFs and the transcriptional cascade-associated interconnections among them remain unidentified. MaNAC029 was recently reported to directly impact the expression of genes involved in the cell wall, starch, and chlorophyll degradation (Wei et al. 2022). In this study, MaSPL16 directly induced the expression of MaNAC029, which in turn upregulated the downstream target genes associated with the degradation of the cell wall, starch, and chlorophyll, ultimately promoting FR and enhancing the quality (Figs. 1, 4, 5 , 6).
Moreover, the possibility of MaSPL16 directly regulating the activation of the fruit quality-related genes requires further study. In summary, the MaSPL16-MaNAC029 transcriptional cascade acted as an upstream regulatory module for establishing quality in banana fruits. These results enrich the understanding of the regulatory mechanisms underlying fruit quality and provide excellent gene resources for breeding banana varieties with high-quality fruits.
Relying on the current and past research conducted by our research group, a functional model explaining the mechanism through which MaSPL16 operates in banana ripening was proposed (Fig. 7). The activation of MaSPL16 was observed throughout the process of postharvest maturation. MaSPL16 directly bound to the promoter of MaNAC029, resulting in the activation of its expression, which in turn promoted the transcriptional activation of ethylene biosynthesis-related genes, resulting in the feedback regulation of ethylene biosynthesis. In addition, the quality-related structural genes were also activated, stimulating the production of good-quality banana fruits throughout the FR processes.
A model proposed to explain the function of the transcriptional cascade, Musa acuminata SPL (SQUAMOSA promoter binding protein-like) 16-Musa acuminata NAC (NAM, ATAF, CUC) 029 (MaSPL16-MaNAC029) in banana ripening. MaSGR1, Musa acuminata Stay-Green 1; MaPPH, Musa acuminata Pheophytinase; MaGWD1, Musa acuminata α-glucan water dikinase 1; MaEXP2/15, Musa acuminata Expansins 2/15; MaXTH28/30, Musa acuminata xyloglucan endotransglycosylase/hydrolases 28/30; MaACS1, Musa acuminata 1-amino-cyclopropane-1-carboxylic acid synthase 1; MaACO1/13, Musa acuminata 1-amino-cyclopropane-1-carboxylic acid oxidase 1/13
Conclusions
In summary, a transcriptional regulatory cascade, specifically the MaSPL16-MaNAC029 pathway was demonstrated, which linked ethylene biosynthesis to fruit quality. These findings provide novel perspectives on the gene-transcriptional mechanisms behind FR. These factors could potentially influence the advancement of molecular techniques that seek to augment the quality and durability of fleshy fruits.
Availability of data and materials
Data will be available on request.
References
Agarwal PR, Lahiri A. Comparative study of the SBP-box gene family in rice siblings. J Biosci. 2020;45:83. https://doi.org/10.1007/s12038-020-00048-z.
Aurore G, Parfait B, Fahrasmane L. Bananas, raw materials for making processed food products. Trends Food Sci Technol. 2009;20:78–91. https://doi.org/10.1016/j.tifs.2008.10.003.
Bemer M, van Dijk ADJ, Immink RGH, Angenent GC. Cross-Family transcription factor interactions: an accitional layer of gene reculation. Trends Plant Sci. 2017;22:66–80. https://doi.org/10.1016/j.tplants.2016.10.007.
Chen L, Zhong HY, Kuang JF, Li JG, Lu WJ, Chen JY. Validation of reference genes for RT-qPCR studies of gene expression in banana fruit under different experimental conditions. Planta. 2011;234:377–90. https://doi.org/10.1007/s00425-011-1410-3.
Chen T, Qin G, Tian S. Regulatory network of fruit ripening: current understanding and future challenges. New Phytol. 2020;228:1219–26. https://doi.org/10.1111/nph.16822.
Dai JY, Xu Z, Xu YT, Fang ZH, Shah K, Kang TY, et al. A novel NAC transcription factor, PpNAP6, is involved in peach ripening by activating ethylene synthesis. Postharvest Biol Technol. 2023;201:112363. https://doi.org/10.1016/j.postharvbio.2023.112363.
Dong TT, Hu ZL, Deng L, Wang Y, Zhu MK, Zhang JL, et al. A tomato MADS-Box transcription factor, SlMADS1, acts as a negative regulator of fruit ripening. Plant Physiol. 2013;163:1026–36. https://doi.org/10.1104/pp.113.224436.
Fan ZQ, Ba LJ, Shan W, Xiao YY, Lu WJ, Kuang JF, et al. A banana R2R3-MYB transcription factor MaMYB3 is involved in fruit ripening through modulation of starch degradation by repressing starch degradation-related genes and MabHLH6. Plant J. 2018;96:1191–205. https://doi.org/10.1111/tpj.14099.
Fu CC, Han YC, Fan ZQ, Chen JY, Chen WX, Lu WJ, et al. The papaya transcription factor CpNAC1 modulates carotenoid biosynthesis through activating phytoene desaturase genes CpPDS2/4 during fruit ripening. J Agric Food Chem. 2019;64:5454–63. https://doi.org/10.1021/acs.jafc.6b01020.
Gao Y, Wei W, Zhao XD, Tan XL, Fan ZQ, Zhang YP, et al. A NAC transcription factor, NOR-like1, is a new positive regulator of tomato fruit ripening. Hortic Res. 2018;5:75. https://doi.org/10.1038/s41438-018-0111-5.
Gao Y, Zhu N, Zhu XF, Wu M, Jiang CZ, Grierson D, et al. Diversity and redundancy of the ripening regulatory networks revealed by the fruit ENCODE and the new CRISPR/Cas9 CNR and NOR mutants. Hortic Res. 2019;6:39. https://doi.org/10.1038/s41438-019-0122-x.
Gapper NE, McQuinn RP, Giovannoni JJ. Molecular and genetic regulation of fruit ripening. Plant Mol Biol. 2013;82:575–91. https://doi.org/10.1007/s11103-013-0050-3.
Giovannoni JJ. Fruit ripening mutants yield insights into ripening control. Curr Opin Plant Biol. 2007;10:283–9. https://doi.org/10.1016/j.pbi.2007.04.008.
Giovannoni J, Cuong N, Ampofo B, Zhong S, Fei Z. The epigenome and transcriptional dynamics of fruit ripening. Annu Rev Plant Bio. 2017;68:61–84. https://doi.org/10.1146/annurev-arplant-042916-040906.
Guo ZH, Zhang YJ, Yao JL, Xie ZH, Zhang YY, Zhang SL, et al. The NAM/ATAF1/2/CUC2 transcription factor PpNAC.A59 enhances PpERF.A16 expression to promote ethylene biosynthesis during peach fruit ripening. Hortic Res. 2021;8:209. https://doi.org/10.1038/s41438-021-00644-6.
Han YC, Kuang JF, Chen JY, Liu X, Xiao YY, Fu CC, et al. Banana transcription factor MaERF11 recruits histone deacetylase MaHDA1 and represses the expression of MaACO1 and Expansins during fruit ripening. Plant Physiol. 2016;171:1070–84. https://doi.org/10.1104/pp.16.00301.
Hellens RP, Allan AC, Friel EN, Bolitho K, Grafton K, Templeton MD, et al. Transient expression vectors for functional genomics, quantification of promoter activity and RNA silencing in plants. Plant Methods. 2005;1:13. https://doi.org/10.1186/1746-4811-1-13.
Hu BX, Sun DW, Pu HB, Wei QY. Recent advances in detecting and regulating ethylene concentrations for shelf-life extension and maturity control of fruit: A review. Trends Food Sci Technol. 2019;91:66–82. https://doi.org/10.1016/j.tifs.2019.06.010.
Ito Y, Nishizawa-Yokoi A, Endo M, Mikami M, Shima Y, Nakamura N, et al. Re-evaluation of the rin mutation and the role of RIN in the induction of tomato ripening. Nat Plants. 2017;3:866–74. https://doi.org/10.1038/s41477-017-0041-5.
Jerome Jeyakumar JM, Ali A, Wang WM, Thiruvengadam M. Characterizing the role of the miR156-SPL network in plant development and stress response. Plants. 2020;9:1206. https://doi.org/10.3390/plants9091206.
Jiang G, Li Z, Song Y, Zhu H, Lin S, Huang R, et al. LcNAC13 physically interacts with LcR1MYB1 to coregulate anthocyanin biosynthesis-related genes during litchi fruit ripening. Biomolecules. 2019;9:135. https://doi.org/10.3390/biom9040135.
Jourda C, Cardi C, Mbéguié-A-Mbéguié D, Bocs S, Garsmeur O, D’Hont A, et al. Expansion of banana (Musa acuminata) gene families involved in ethylene biosynthesis and signaling after lineage-specific whole-genome duplications. New Phytol. 2014;202:986–1000. https://doi.org/10.1111/nph.12710.
Karlova R, Rosin FM, Busscher-Lange J, Parapunova V, Do PT, Fernie AR, et al. Transcriptome and metabolite profiling show that APETALA2a is a major regulator of tomato fruit ripening. Plant Cell. 2011;23:923–41. https://doi.org/10.1105/tpc.110.081273.
Kropat J, Tottey S, Birkenbihl RP, Depege N, Huijser P, Merchant S. A regulator of nutritional copper signaling in chlamydomonas is an SBP domain protein that recognizes the GTAC core of copper response element. Proc Natl Acad Sci U S A. 2005;102:18730–5. https://doi.org/10.1073/pnas.0507693102.
Kuang JF, Chen JY, Luo M, Wu KQ, Sun W, Jiang YM, et al. Histone deacetylase HD2 interacts with ERF1 and is involved in longan fruit senescence. J Exp Bot. 2012;63:441–54. https://doi.org/10.1093/jxb/err290.
Lai TF, Wang XH, Ye BS, Jin MF, Chen WW, Wang Y, et al. Molecular and functional characterization of the SBP-box transcription factor SPL-CNR in tomato fruit ripening and cell death. J Exp Bot. 2020;71:2995–3011. https://doi.org/10.1093/jxb/eraa067.
Li SJ, Yin XR, Wang WL, Liu XF, Zhang B, Chen KS. Citrus CitNAC62 cooperates with CitWRKY1 to participate in citric acid degradation via upregulation of CitAco3. J Exp Bot. 2017;68:3419–26. https://doi.org/10.1093/jxb/erx187.
Li S, Chen K, Grierson D. A critical evaluation of the role of ethylene and MADS transcription factors in the network controlling fleshy fruit ripening. New Phytol. 2019;221:1724–41. https://doi.org/10.1111/nph.15545.
Li S, Zhu BZ, Pirrello J, Xu C, Zhang B, Bouzayen M, et al. Roles of RIN and ethylene in tomato fruit ripening and ripening-associated traits. New Phytol. 2020;226:460–75. https://doi.org/10.1111/nph.16362.
Li S, Chen K, Grierson D. Molecular and hormonal mechanisms regulating fleshy fruit ripening. Cells. 2021;10:1136. https://doi.org/10.3390/cells10051136.
Liu GS, Li HL, Grierson D, Fu DQ. NAC transcription factor family regulation of fruit ripening and quality: a review. Cells. 2022;11:525. https://doi.org/10.3390/cells11030525.
Lv PT, Yu S, Zhu N, Chen YR, Zhou BY, Pan Y, et al. Genome encode analyses reveal the basis of convergent evolution of fleshy fruit ripening. Nat Plants. 2018;4:784–91. https://doi.org/10.1038/s41477-018-0249-z.
Maduwanthi SDT, Marapana RAUJ. Induced ripening agents and their effect on fruit quality of banana. Int J Food Sci. 2019;2520179. https://doi.org/10.1155/2019/2520179.
Pérez AG, Cert A, Rios JJ, Olias JM. Free and glycosidically bound volatile compounds from two bananas cultivars: Valery and Pequeña Enna. J Agric Food Chem. 1997;45:4393–7. https://doi.org/10.1021/jf9704524.
Qin GZ, Zhu Z, Wang WH, Cai JH, Chen Y, Li L, et al. A tomato vacuolar invertase inhibitor mediates sucrose metabolism and influences fruit ripening. Plant Physiol. 2016;172:1596–611. https://doi.org/10.1104/pp.16.01269.
Salinas M, Xing S, Hoehmann S, Berndtgen R, Huijser P. Genomic organization, phylogenetic comparison and differential expression of the SBP-box family of transcription factors in tomato. Planta. 2012;235:1171–84. https://doi.org/10.1007/s00425-011-1565-y.
Shan W, Kuang JF, Wei W, Fan ZQ, Deng W, Li ZG, et al. MaXB3 Modulates MaNAC2, MaACS1, and MaACO1 stability to repress ethylene biosynthesis during banana fruit ripening. Plant Physiol. 2020;184:1153–71. https://doi.org/10.1104/pp.20.00313.
Sun QG, Jiang SH, Zhang TL, Xu HF, Fang HC, Zhang J, et al. Apple NAC transcription factor MdNAC52 regulates biosynthesis of anthocyanin and proanthocyanidin through MdMYB9 and MdMYB11. Plant Sci. 2019;289:110286. https://doi.org/10.1016/j.plantsci.2019.110286.
Sun YT, Wang YY, Xiao YQ, Zhang X, Du BY, Turupu M, et al. Genome-wide identification of the squamosa promoter-binding protein-like (SPL) transcription factor family in sweet cherry fruit. Int J Mol Sci. 2023;24:2880. https://doi.org/10.3390/ijms24032880.
Tang YQ, Yan Y, Tie WW, Ye XX, Zeng LW, Zeng LM, et al. Transcriptional regulation of MbACO2-mediated ethylene synthesis during postharvest banana ripening. Postharvest Biol Technol. 2023;200:112325. https://doi.org/10.1016/j.postharvbio.2023.112325.
Thompson, Tor, Barry, Vrebalov, Orfila, Jarvis, et al. Molecular and genetic characterization of a novel pleiotropic tomato-ripening mutant. Plant Physiol. 1999;120:383–90. https://doi.org/10.1104/pp.120.2.383.
Tisza V, Kovacs L, Balogh A, Heszky L, Kiss E. Characterization of FaSPT, a SPATULA gene encoding a bHLH transcription factor from the nonclimacteric strawberry fruit. Plant Physiol Biochem. 2010;48:822–6. https://doi.org/10.1016/j.plaphy.2010.08.001.
Wang SF, Lu G, Hou Z, Luo ZD, Wang TT, Li HX, et al. Members of the tomato FRUITFULL MADS-box family regulate style abscission and fruit ripening. J Exp Bot. 2014;65:3005–14. https://doi.org/10.1093/jxb/eru137.
Wang RF, Angenent GC, Seymour G, de Maagd RA. Revisiting the role of master regulators in tomato ripening. Trends Plant Sci. 2020;25:291–301. https://doi.org/10.1016/j.tplants.2019.11.005.
Wang WH, Wang PW, Li XJ, Wang YY, Tian SP, Qin GZ. The transcription factor SlHY5 regulates the ripening of tomato fruit at both the transcriptional and translational levels. Hortic Res. 2021;8:83. https://doi.org/10.1038/s41438-021-00523-0.
Wang WH, Wang YY, Chen T, Qin GZ, Tian SP. Current insights into posttranscriptional regulation of fleshy fruit ripening. Plant Physiol. 2022;192:1785–98. https://doi.org/10.1093/plphys/kiac483.
Wei W, Yang YY, Chen JY, Lakshmanan P, Kuang JF, Lu WJ, et al. MaNAC029 modulates ethylene biosynthesis and fruit quality and undergoes MaXB3-mediated proteasomal degradation during banana ripening. J Adv Res. 2022;S2090–1232:00270–3. https://doi.org/10.1016/j.jare.2022.12.004.
Wei W, Yang YY, Lakshmanan P, Kuang JF, Lu WJ, Pang XQ, et al. Proteasomal degradation of MaMYB60 mediated by the E3 ligase MaBAH1 causes high temperature-induced repression of chlorophyll catabolism and green ripening in banana. Plant Cell. 2023;35:1408–28. https://doi.org/10.1093/plcell/koad030.
Wu YY, Liu XF, Fu BL, Zhang QY, Tong Y, Wang J, Grierson D, Yin XR, et al. Methyl jasmonate enhances ethylene synthesis in kiwifruit by inducing NAC genes that activate ACS1. J Agric Food Chem. 2020;68:3267–76. https://doi.org/10.1021/acs.jafc.9b07379.
Xiao YY, Kuang JF, Qi XN, Ye YJ, Wu ZX, Chen JY, et al. A comprehensive investigation of starch degradation process and identification of a transcriptional activator MabHLH6 during banana fruit ripening. Plant Biotechnol J. 2018;16:151–64. https://doi.org/10.1111/pbi.12756.
Xu YJ, Xu HX, Wall MM, Yang JZ. Roles of transcription factor SQUAMOSA promoter binding protein -like gene family in papaya (Carica papaya) development and ripening. Genomics. 2020;112:2734–47. https://doi.org/10.1016/j.ygeno.2020.03.009.
Yin XR, Allan AC, Chen KS, Ferguson IB. Kiwifruit EIL and ERF genes involved in regulating fruit ripening. Plant Physiol. 2010;153:1280–92. https://doi.org/10.1104/pp.110.157081.
You Y, Zheng YT, Wang J, Chen G, Li SJ, Shao JF, et al. Molecular evolution and genome-wide analysis of the SBP-box family in cucumber (Cucumis sativas). Plant Growth Regul. 2021;93:175–87. https://doi.org/10.1007/s10725-020-00677-2.
Zhong SL, Fei ZJ, Chen YR, Zheng Y, Huang MY, Vrebalov JL, et al. Single-base resolution methylomes of tomato fruit development reveal epigenome modifications associated with ripening. Nat Biotechnol. 2013;31:154–9. https://doi.org/10.1038/nbt.2462.
Zhou H, Wang KL, Wang H, Gu C, Dare AP, Espley RV, et al. Molecular genetics of blood-fleshed peach reveals activation of anthocyanin biosynthesis by NAC transcription factors. Plant J. 2015;82:105–21. https://doi.org/10.1111/tpj.12792.
Zhu LS, Liang SM, Chen LL, Wu CJ, Wei W, Shan W, et al. Banana MaSPL16 modulates carotenoid biosynthesis during fruit ripening through activating the transcription of lycopene beta-cyclase genes. J Agric Food Chem. 2020;68:1286–96. https://doi.org/10.1021/acs.jafc.9b07134.
Zhu LS, Chen L, Wu CJ, Shan W, Cai DL, Lin ZX, et al. Methionine oxidation and reduction of the ethylene signaling component MaEIL9 are involved in banana fruit ripening. J Integr Plant Biol. 2023;65:150–66. https://doi.org/10.1111/jipb.13363.
Acknowledgements
Not applicable.
Funding
This study was supported by the National Natural Science Foundation of China (No. 32072279 and 32202124) and the China Agriculture Research System of MOF and MARA (No. CARS-31).
Author information
Authors and Affiliations
Contributions
The study was conceived and designed by WS and WW who participated as a collaborator in conducting the experiments. The process of data analysis was conducted by a collaborative team which was comprised by WW, Y-Y Y, C-J W, J-F K and WS. The process of manuscript composition and revision was undertaken by J-Y C, WS and WW. All authors actively engaged in discussions related to the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Additional file 1: Fig. S1.
Phylogenetic analysis of MaSPL16. Fig. S2. Coomassie blue-stained SDS-PAGE shows the prokaryote-expressed and purified GST-MaSPL16 recombinant protein. Fig. S3. Uncropped version of the blots presented in Fig. 1b. Text S1. The SPL TF recognition sequences (GTAC) in the nucleotide sequences of Musa acuminata NAC (NAM, ATAF, CUC) 029 (MaNAC029) promoter. Table S1. Summary of primers used in this study.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wei, W., Yang, Yy., Wu, Cj. et al. MaSPL16 positively regulates fruit ripening in bananas via the direct transcriptional induction of MaNAC029. HORTIC. ADV. 1, 10 (2023). https://doi.org/10.1007/s44281-023-00013-4
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s44281-023-00013-4