Abstract
Caspofungin and other echinocandins have been used for the treatment of human infections by the opportunistic yeast pathogen, Candida albicans. There has been an increase in infections by non-albicans Candida species such as Candida glabrata, Candida parapsilosis, Candida tropicalis, Candida krusei, and Candida auris in clinical or hospital settings. This is problematic to public health due to the increasing prevalence of echinocandin resistant species/strains. This review will present a summary on various studies that investigated the inhibitory action of caspofungin on 1,3-β-d-glucan synthesis, on cell wall structure, and biofilm formation of C. albicans. It will highlight some of the issues linked to caspofungin resistance or reduced caspofungin sensitivity in various Candida species and the potential benefits of antimicrobial peptides and other compounds in synergy with caspofungin.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Caspofungin ((MK-0991; L-743,872) is a fungicidal, water-soluble semisynthetic echinocandin that inhibits synthesis of β-1,3-d-glucan, a main structural component of the fungal cell wall (Fig. 1) [1]. Apart from caspofungin, micafungin and anidulafungin are two additional echinocandins approved for used by the US Food and Drug Administration (FDA) (Fig. 1) [2, 3]. Though these echinocandins have different side chains, there have three common components essential for their activities [2]. They are as follows: (a) a homotyrosine amino acid residue essential for the antifungal activity and for the inhibition of the glucan synthase enzyme; (b) proline residues which enhances the antifungal potency of the echinocandin drugs; and (c) the hydroxyl groups in the echinocandin B nucleus which improve their stability and increase their water solubility [2]. Currently, a novel echinocandin derived from anidulafungin, rezafungin (also known as CD101), is undergoing phase-III trials [4,5,6].
Caspofungin and β-(1,3)-glucan synthase
The model yeast Saccharomyces cerevisiae
Saccharomyces cerevisiae cell wall comprises an inner layer containing the polysaccharides β-1,3-glucan, β-1,6-glucan, and chitin and while mannoproteins act as “fillers” affecting cell wall porosity and are in the outer layer the yeast cell wall (Fig. 2) [7, 8]. In S. cerevisiae, β-1,3-glucan synthase has been shown to catalyze the formation of a β-1,3-glucan polymer, a major component of the fungal cell wall (Fig. 2). In yeast and many fungal species, the β-1,3-D-glucan chains form a solid three-dimensional matrix, which gives the cell wall its shape and mechanical strength. Kollár et al. [9, 10] demonstrated that chitin (a linear polymer composed of β-(1,4)-linked N-acetylglucosamine subunits) is glycosidically linked to nonreducing branches of the β1,3-glucan and β1,6-glucan in S. cerevisiae. Later work by Cabib et al. [11] found that β-1,3-D-glucan formed a noncovalent complex with chitin. β-1,6-glucan plays a role in the organization of the yeast cell wall by interconnecting all other wall components into a lattice by attaching mannoproteins via their glycosylphosphatidylinositol (GPI) glycan remnant to β-1,3-glucan and chitin [12, 13].
Caspofungin’s target in Saccharomyces cerevisiae and Candida albicans. a Wild-type strains Saccharomyces cerevisiae BY4741 and Candida albicans DAY185 were stained with fluorochrome Aniline Blue (AB; Biosupplies Australia PTY Ltd) for β-1,3-glucan detection in yeast cell wall (in yellow) under UV fluorescence. b Schematic diagram of the yeast cell wall membrane highlighting the inhibitory action of caspofungin on β-1,3-glucan synthesis (red lightning bolt)via noncompetitive inhibition of the β-1,3-glucan synthase complex (Fks1p and Fks2p; green cylinder)
β-1,3-glucan synthesis in S. cerevisiae involves the integral membrane proteins Fks1p and Fks2p which act as subunits of the β-1,3 glucan-synthase enzyme complex (Fig. 2) [14,15,16,17] and the regulatory subunit encoded by RHO1 [18,19,20]. Another FKS homolog, FKS3, was required for normal spore wall formation while FKS2 (GSC2) was the primary β-1,3-glucan synthase in S. cerevisiae sporulation [21]. Interestingly, work on FKS1 and FKS2 by Mazur and colleagues [17] showed that not only FKS1 was affected by the echinocandin but that FKS2 was also sensitive to the echinocandin L-733,560, due to the increased sensitivity of fks1 null mutants to this drug. However, work by Douglas et al. [22] and El-Sherbeini and Clemas [23] presented evidence of mutations within the FKS1 gene that could affect the sensitivity to the semisynthetic pneumocandin B, L-733,560 in S. cerevisiae. Screening of the S. cerevisiae deletion mutant collection for altered sensitivity to the drug found that deletions in 52 genes led to caspofungin hypersensitivity and those in 39 genes to resistance [24]. Use of a genomic approach to identify genes involved in caspofungin susceptibility in S. cerevisiae showed that the disruption of 20 genes involved in key functions such as in cell wall and membrane function, chitin and mannan biosynthesis, vacuole, and transport functions led to increased caspofungin sensitivity [25]. For example, the loss of ERG3, a C-5 sterol desaturase which catalyzes the introduction of a C-5(6) double bond into episterol, a precursor in ergosterol biosynthesis, led to increased caspofungin resistance in S. cerevisiae [25]. Furthermore, Carolus et al. [26], Rybak et al. [27], and Spettel et al. [28] identified that the disruption/mutation in ERG3 resulted in increased resistance to azole and echinocandin antifungals in C. albicans, Candida auris, and Candida parapsilosis.
Candida albicans
Since identifying the FKS1 homolog in C. albicans [29] and demonstrating that the non-competitive binding ability of echinocandins to FKS1 gene in C. albicans [30], echinocandins including caspofungin have been used in the treatment of Candida spp. and other fungal infections [1]. FKS2 and FKS3 are also found in C. albicans and in C. albicans mutants lacking either FKS2 or FKS3 that FKS1 expression was upregulated suggesting that FKS2 and FKS3 act as negative regulators of FKS1 [31].
However, there have been many reports documenting echinocandin resistance in Candida species [32,33,34,35]. Such echinocandin resistance in Candida spp. is due to point mutations in 2 highly conserved “hot spot” regions, i.e., HS1 and HS2 of the FKS1 gene [32, 33, 36]. Previous studies by Park and colleagues [37] demonstrated that substitutions in the Fks1p subunit of GS in S. cerevisiae and four clinical C. albicans isolates and a Candida krusei isolate were sufficient to confer reduced susceptibility to echinocandins.
The haploid Candida glabrata, an evolutionarily close relative of S. cerevisiae, causes mucosal and systemic infections especially in the human immunodeficiency virus-infected population [38]. Its genome has also three GS homologs, FKS1, FKS2, and FKS3 [39]. In this particular species, mutations in both FKS1 and FKS2 but not FKS3 have been associated with echinocandin resistance [40, 41]. Clinical C. glabrata isolates displaying reduced susceptibility or resistance to anidulafungin, caspofungin, and micafungin were not only due to FKS1 modification but to point mutations in the FKS2 [42]. Similarly, FKS1 hot spot 1 (HS1) and FKS2 HS1 have been identified in clinical C. auris isolates with reduced caspofungin susceptibility [43]. Genetic engineering for full-length replacement of the FKS1 gene, containing FKS1 hotspot (HS) regions HS1 or HS2 mutations from C. albicans, the F659 deletion in the FKS2 allele of C. glabrata and the naturally occurring P660A substitution in FKS1 of C. parapsilosis respectively into Candida lusitaniae confirmed the role of FKS mutations associated with in vitro caspofungin resistance or reduced echinocandin susceptibility [44]. For additional information, Arendrup [45], Arendrup and Perlin [46], and Lackner et al. [47] listed mutations in FKS1 and FKS2 known to contribute to resistance in various Candida isolates. Another study using fluorescently labeling caspofungin, Jaber and colleagues [48] observed enhanced caspofungin uptake in the vacuoles of echinocandin-resistant C. albicans and C. glabrata strains with point mutations in the FKS genes compared to echinocandin-sensitive isogenic strains.
Other fungal pathogens displaying decreased echinocandin susceptibility have been reported such as FKS1 mutation (E671Q) in Aspergillus fumigatus following anidulafungin exposure [49]. Mutations studies in FKS1 and FKS2 have helped in our understanding in the intrinsic resistance to echinocandins in Scedosporium prolificans and Scedosporium apiospermum [50], and Fusarium solani [51].
Another explanation for tolerance to caspofungin independent of FKS1 (located on chromosome 1; Ch1) in C. albicans was observed in strains adapted in vitro to lethal doses of caspofungin [52]. Similarly in the previous study investigating C. albicans adaptation to toxic levels of the sugar L-sorbose [53], monosomy of chromosome 5 (Ch5) played a role in tolerance of C. albicans to caspofungin [52]. In addition, monosomy of the left arm and trisomy of the right arm of Ch5 were also detected in caspofungin-adapted C. albicans cells [52]. In such mutants, there was a downregulation of FKS genes hence a decreased amount of β-1,3-D-glucan content and an increase in chitin content in the cell wall [52]. Further analyses of Ch5 found genes encoding positive regulators of caspofungin susceptibility, CNB1 (the regulatory subunit of calcineurin B), and MID1 (a putative stretch-activated Ca2+ channel) [52]. Negative regulators of caspofungin susceptibility CHT2 (a GPI-dependent chitinase), PGA4 (a GPI-anchored cell surface 1,3-β-D-glucanosyltransferase), and CSU51 (a putative GPI-anchored protein) were also found in the same chromosome [52].
Effect of caspofungin on C. albicans cell surface
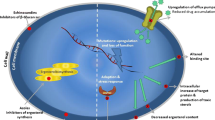
Caspofungin and other echinocandins not only disrupt β-1,3-glucan synthesis in C. albicans (Fig. 2), which can compromise cell wall integrity, but can also kill C. albicans in a dose-dependent manner [54] by causing metacaspase-dependent apoptosis [55, 56] and necrosis [55]. However, many studies have been focused on cell wall remodeling/mechanical strength and the paradoxical growth (PG) of C. albicans and various Candida species in response to caspofungin (Fig. 3) and how it could be linked to caspofungin or echinocandin resistance in Candida spp.
Paradoxical growth of Candida albicans DAY185 cells treated with caspofungin at different concentrations stained with calcofluor white (CFW). C. albicans DAY185 grown in ½ strength PDB liquid 30 °C supplemented with caspofungin at 0, 2, 4, 10, and 100 ng/mL respectively were examined with an Olympus BX50 upright microscope with UPlanApo × 100/1.35 oil objective equipped with a filter cube U-MWU2 (excitation 330–385 nm/emission 420 nm/dichromatic mirror 400 nm). Images were acquired with SPOT Camera using Spot RT analysis software. Black and red arrows highlight the presence of enlarged and elongated yeast cells (putative pseudohyphae). Note the increase in flocculation and CFW fluorescence emitted by cells treated with an increasing concentration of caspofungin. All images were acquired after 8 ms
In four Candida species (C. albicans, Candida orthopsilosis, C. parapsilosis, and Candida tropicalis) exhibiting PG at approximately 16 μg mL−1 of caspofungin, PG cells had a decrease in β-1,3-glucan and an elevated chitin cell wall content compared to control untreated cells [57] (Fig. 3). In the same study, PG cells were altered in their morphology where they formed clumps of enlarged cells, had abnormal septa, and lacked filamentation [57] (Fig. 3). Another feature was that the control (WT) C. albicans 1399 cells had the characteristic two layered cell walls, i.e., an electron-dense outer layer and an inner layer of low electron density, while caspofungin-treated C. albicans 1399 PG cells had a decreased inner cell wall layer and a predominant more-electron-dense layer [57]. Prior work by Nishiyama and colleagues [58] observed similar morphological changes induced by 0.1 μg mL−1 or above of micafungin post 24 h in C. albicans ATCC 99,028, a strain known to be susceptible to micafungin. Later work by Rueda et al. [59] suggested that the increase in chitin in caspofungin-treated C. albicans may contribute to the protection of the cells from the fungicidal effect of the drug. Interestingly, paradoxical growth can be induced by the application of 4 μg mL−1 caspofungin for 24 h in A. fumigatus [60]. In this case, Wagener and Loiko [61] proposed that an increase in chitin in fungi in response to echinocandins could facilitate their survival upon the inhibition of β-1,3 glucan synthesis. However, paradoxical growth was not observed in a laboratory A. fumigatus strain Af293 and 7 clinical A. fumigatus strains in the presence of micafungin or anidulafungin [62].
Candida cell wall remodeling—elevated chitin
An atomic force microscopy (AFM) study investigating the effect of caspofungin (at 50 ng ml−1 for 2 h) on C. albicans cell wall surface demonstrated that there was a decrease in mechanical cell wall strength, i.e., caspofungin-induced softening of the cell wall due to a decrease in β-1,3 glucan content [63]. This would affect C. albicans’ cell shape, mechanical rigidity, and resistance to osmotic pressure leading to the induction of osmotically fragile cells or swollen cells [63]. Another study using the AFM approach combined with Fourier transform infrared spectroscopy in attenuated total reflection mode (ATR-FTIR) investigated the effect in caspofungin on Candida lusitaniae CBS 6936 and its caspofungin-resistant mutant (bearing the mutation S645P in FKS1) [64]. They demonstrated that cell wall stiffening occurred only at low concentration of caspofungin (∼0.06 μg mL−1; 0.5 MIC) for WT cells, whereas it was observed at high concentration (∼6.25 μg mL−1; 50 MIC) for resistant strains [64].
To restore cell wall integrity, C. albicans stimulates chitin synthesis to enable cells to survive lethal concentrations of echinocandins in vitro [65]. Lee et al. [66] observed that high chitin C. albicans cells were less susceptible to caspofungin in mice infection and that C. tropicalis, C. parapsilosis, Candida guilliermondii, and C. krusei elevated their chitin cell wall content in response to caspofungin treatment [67]. Similar observations were documented in C. auris in response to caspofungin [43]. Protein kinase C (PKC), high osmolarity glycerol (HOG) mitogen-activated protein (MAP) kinase, and Ca2+/calcineurin signaling pathways have been shown to regulate chitin synthases, CHS1, CHS2, CHS3, and CHS8 gene expression and chitin synthesis in C. albicans and various mutants grown in YPD supplemented with different agents such as caffeine, cyclosporin A, the calcineurin inhibitor, FK506, and A23187 (Calcimycin) [68]. Recent work by Han et al. [69] on C. albicans SC5314 and its deleted mutants of β-1,6-glucan synthesis, KRE6, and SKN1 found that cell wall chitin levels increased through the post-transcriptional regulation of the chitin synthase Chs3 leading to the cell viability maintenance via Ca2+/calcineurin and PKC signaling pathways. Furthermore, β-1,3-glucan had no role in compensating β-1,6-glucan synthesis in C. albicans kre6Δ/Δ skn1Δ/Δ cells as both the WT and mutants grew on YPD plates containing 0.064 μg mL−1 of caspofungin [69].
In S. cerevisiae, deletion of FKS1 induced a compensatory mechanism, i.e., high rates of chitin synthesis due to a significant increase in CHS3 activity [70]. Calcofluor white staining of a β-1,3-glucan synthase knockout fks1::URA3 strain of S. cerevisiae displayed an elevated fluorescence signal (i.e., elevated chitin content) compared to the wild type [71]. Lesage et al. [24] suggested that there was functional link between chitin and glucan synthesis where an increased in chitin synthesis could compensate for defective β-1,3-glucan assembly for survival in the presence of caspofungin such as the deletion of CHS3 or CHS4–7 leading to caspofungin hypersensitivity in S. cerevisiae.
Cell–cell interactions
A review by Heredia et al. [72] highlighted that there are three transcription factors Sko1, Rlm1, and Cas5 that coordinate and regulate the caspofungin-induced cell wall damage response in C. albicans. The transcription factor Sko1 (ORF 19.1032) and its upstream regulator, the PAS-domain protein, and protein kinase Psk1 (ORF19.7451) were shown to be involved in C. albicans’ wall regulatory pathway [73]. Deletion mutants sko1 Δ/Δ and psk1 Δ/Δ were hypersensitive to 125 ng mL−1 caspofungin compared to its wild-type strain C. albicans DAY185 [73]. The same authors demonstrated that up to 79 caspofungin-responsive genes were regulated by Sko1 including key genes involved in cell wall biosynthesis CRH11, MNN2, and SKN1 and in cell wall damage, PGA13 [73]. The latter encodes a GPI protein and pga13Δ mutants exhibited a higher surface hydrophobicity, and increased adherence and flocculation (cell–cell interactions) [74]. Later work by Alonso et al. [75] highlighted that Sko1 mediated the hyphal formation in C. albicans by repressing two genes, HWP1 (Hyphal Wall Protein 1), and ECE1 (Extent of Cell Elongation 1) known to be involved in yeast-to-hyphal transition [76] as well as oxidative stress response via HOG1.
The role of the transcription factor RLM1 in the maintenance of the cell wall integrity was shown by susceptibility assays of C. albicans Δ/Δ rlm1 mutants to 30 ng mL−1 caspofungin and various compounds such as calcofluor white [77]. In this study, the authors found that the caspofungin susceptible rlm1 deletion mutants which in the presence of 1 M sorbitol reverted to a wild-type phenotype had an elevated chitin and a reduced mannan cell wall content compared to the wild type [77]. Microarray analysis of the rlm1 deletion mutants in the absence of stress showed an upregulation of genes linked to cell adhesion like ECE1, HWP1, and two genes that belong to agglutinin-like sequence (ALS) family, ALS1 and ALS3 [77]. Within this family, eight genes (ALS1, ALS2, ALS3, ALS4, ALS5, ALS6, ALS7, and ALS9) encode cell-surface glycoproteins that play a role in adhesion, biofilm formation, hydrophobicity, and pathogenesis in C. albicans [78,79,80]. Interestingly, ALS1 has been linked to caspofungin-induced cell flocculation/aggregation of C. albicans as yeast cells flocculated in growth media supplemented with 10 ng mL−1 and 100 ng mL−1 caspofungin, respectively [81]. Furthermore, compared to the wild-type strains, the als1Δ/Δ mutant cells had diminished flocculation in the presence of 100 ng mL−1 caspofungin [81]. In the same study, the authors linked flocculation to the regulator of morphogenesis, EFG1, a Candida homolog of PHD1 from S. cerevisiae as efg1 Δ/Δ deletion mutants were susceptible to caspofungin and impaired in flocculation compared to C. albicans wild-type strains [81]. Similarly, efg1 knockouts of C. parapsilosis were sensitive to caspofungin compared to the wild-type strain CLIB214 and the efg1/ACT1-EFG1 complemented strain [82]. EFG1 and various transcription factors (TFs) have been linked to hyphal morphogenesis (the switch from a unicellular budding yeast to multicellular filamentous hyphal growth) thus allowing C. albicans’ hyphae to attach and to penetrate through the epithelial cell layers of an infected host [83]. Past work by Noffz et al. [84] and Stoldt et al. [85] demonstrated that the EFG1 overexpression in C. albicans led to pseudohyphae development. Another TF is C. albicans CaSFL1 (suppressor for flocculation gene) which acts as a negative regulator of hyphal development and flocculation in C. albicans [86, 87]. Interestingly, a recent study demonstrated a link with SFL1 and EFG1 in negatively and positively regulating hyphal morphogenesis and microcolony formations [88].
Moreover, efg1 interacts with cas5, a transcription factor involved in stress responses, cell cycle regulation, and drug resistance [89, 90] in vivo and both regulators are critical for the induction of caspofungin-responsive genes such as ALS1 in C. albicans [91]. Apart from ALS1 and PGA13, additional GPI-anchored proteins were found to be affected by caspofungin in C. albicans [92].
One of the consequences of caspofungin inducing cell wall changes in various Candida species is that the additional modification to cell wall GPI-anchored proteins and the increase in chitin/glucan exposure decreased phagocytosis of C. albicans, C. tropicalis, C. dubliniensis, C. lusitaniae, and C. guilliermondii by J774 macrophages [93]. In addition, there was no change in phagocytosis by J774 macrophages with C. glabrata and C. parapsilosis in the presence and absence of caspofungin as there were no changes in glucan exposure in response to caspofungin treatment in these species [93]. It has been shown that chitin blocked the recognition of live C. albicans yeast cells by human peripheral blood mononuclear cells (PBMCs), leading to significant reduction in the stimulation of TNF-α, IL-6, and IL-1β [94]. In contrast, an increased elicitation of TNF-α from macrophages in a Dectin-1-dependent manner has been linked to the unmasking (or exposure) of β-1,3-glucan in C. albicans and its cho1Δ/Δ mutant which is unable to synthesize phosphatidylserine [95]. Similar observations have been made due to unmasking of β-1,3-glucan in C. albicans kre5Δ/Δ to hyperelicit TNFα from macrophages [96]. KRE5 encodes a UDP-glucose:glycoprotein glucosyltransferase localized in the endoplasmic reticulum in C. albicans and S. cerevisiae [97]. Work by Herrero et al. [97] demonstrated that the lack of Kre5p in C. albicans reduced adherence to human epithelial cells and the KRE5 homozygous mutant strains were avirulent in a BALB/c mouse model of systemic infection. A recent investigation in C. glabrata also identified that a functional homolog of KRE5, CgKRE5, and C. glabrata cells with the tetracycline-dependent system to repress CgKRE5 in the presence of 20 μg mL−1 doxycycline (DOX) had enhanced sensitivity to micafungin (at 3 μg mL−1) compared to those grown in the absence of DOX [98]. AFM work on C. albicans SC5314 (WT), the cho1Δ/Δ mutant, the kre5Δ/Δ mutant, and the caspofungin-treated WT confirmed that by inhibiting key steps in cell wall synthesis increased cell wall roughness and decreased cell wall elasticity [99]. By using AFM tips functionalized with sDectin-1-Fc, which is highly specific for glucans with a pure (1 → 3)-β-linked backbone structure [100], the authors demonstrated that the kre5Δ/Δ mutant had the highest frequency of binding (or peak adhesion frequency) followed by caspofungin-treated WT cells, the cho1Δ/Δ mutant, and almost no binding with the WT [101]. Thus, the differences in β-1,3-glucan layer exposure could contribute to Candida’s pathogenicity, virulence, and the immune system evasion and/or survival.
C. albicans biofilms and caspofungin
Another aspect of C. albicans contributing to its pathogenicity and virulence is its ability to form biofilms, i.e., where densely packed communities yeast cells adhere to surfaces [101,102,103]. In C. albicans, biofilm formation involve adherence, the formation of microcolonies, and of hyphae surrounded by an extracellular matrix (ECM) of polysaccharides to the subsequent dispersal of planktonic cells after reaching maturation [104,105,106]. C. tropicalis biofilms are similar in structure to C. albicans while C. parapsilosis form pseudo-hyphae while C. auris and C. glabrata biofilms are made up of blastospores within an ECM [104, 105, 107]. Many genes involved in biofilm formation in C. albicans and other Candida species have been studied (refer to reviews [104, 105, 107,108,109]). Bachmann et al. [110] spotlighted the benefits of caspofungin against C. albicans biofilms in vitro. Later investigations by Ferreira et al. [111] and Melo et al. [112] observed paradoxical growth in caspofungin-treated biofilms, i.e., enlarged, globose cells, and a resurgence of growth at drug concentrations above the MIC in clinical Candida species.
Candida spp. biofilm formation poses a clinical problem in transplant, oncology, and intensive care medicine and echinocandins are still used in its management [113,114,115]. An in vitro study on the novel echinocandin, rezafungin (CD101), on C. albicans biofilms suggests that it could be useful in preventing and treating biofilm-associated nosocomial infections [113].
Synergistic activity of caspofungin with other compounds
Due to increased resistance of C. albicans to caspofungin, there are have been various studies in the use of combination therapies which would result in synergistic action and greater potency than the constituent drugs used in monotherapy [116, 117].
Work by Troskie et al. [118] have shown the benefits of combining tyrocidines, a type of cationic cyclodecapeptides with potent antibacterial and antimalarial activities from Bacillus aneurinolyticus, with caspofungin against C. albicans strain SC5314, which is known to form robust biofilms. In combination, the three major tyrocidines, TrcA, TrcB, and TrcC, significantly increased the C. albicans biofilm eradication activities of caspofungin [118]. In addition, the fractional inhibitory concentration index (FICI) values of TrcA with caspofungin were more promising than TrcB and TrcC with caspofungin combination respectively against 24-h-old C. albicans biofilms [118]. Further testing in the C. elegans infection model, 5 days posttreatment of a single dose of 3.0 μM TrcA and 0.19 μM caspofungin almost doubled the nematode survival rate of C. albicans-infected nematodes compared to C. albicans-infected nematodes treated with a single dose of 0.19 μM caspofungin only [118].
A recent review by Oshiro and colleagues [119] highlighted the benefits of antifungal peptides (AFPs) or antimicrobial peptides (AMPs) such as plant defensins, cathelicidins, and histatins in the inhibition and eradication of Candida spp. biofilms. It has been shown that the use of AMPs with caspofungin had an enhanced antifungal activity against C. albicans in vitro and in vivo [117]. One AMP of human origin, hMUC7–12 [120], and one of amphibian origin, DsS3(1–16) [121], when combined with caspofungin respectively were shown to improve the survival of wax moth larvae (Galleria mellonella) infected with C. albicans SC5314 compared to those infected wax moth larvae treated with PBS or hMUC7–12 (25 mg kg−1), DsS3(1–16) (25 mg kg−1), and caspofungin (0.5 mg kg−1) only [117]. The same authors observed similar results in infected wax moth larvae when treated with the cyclic peptide, colistin sulfate (10 mg kg−1) in combination with caspofungin [117].
Plant defensins have also been investigated for their synergistical efficacy with caspofungin against C. albicans. Of the two radish defensins, RsAFP1 and RsAFP2, the later induced mislocalization of septins in C. albicans CAI4 cells expressing SEP7-GFP-tagged allele and blocked the yeast-to-hypha transition in a dose-dependent manner in C. albicans CAI4 cells [122]. Further work by Vriens et al. [123], using the recombinant (r)RsAFP2, heterologously expressed in Pichia pastoris, demonstrated that RsAFP2 prevented C. albicans biofilm formation and acted synergistically with caspofungin and amphotericin B in the prevention and eradication of C. albicans biofilms. Recent work by Cools et al. [124] demonstrated that a truncated peptide variant of the plant defensin HsAFP, isolated from Heuchera sanguinea, HsLin06_18, when combined with caspofungin reduced in vitro biofilm formation of C. albicans SC5314 WT on polyurethane catheters as well as a caspofungin-resistant C. albicans mutant strain M177 and, the use of a subcutaneous rat catheter model in immunosuppressed female Sprague–Dawley rats, the combination reduced biofilm formation of C. albicans in vivo. Furthermore, the authors observed that the caspofungin facilitated the internalization and the membrane permabilization of HsLin06_18 into planktonic C. albicans SC5314 WT cells [124].
Work by Sun and colleagues [125] demonstrated the benefits of combining polyphenols such as caffeic acid phenethyl ester (CAPE) with caspofungin against C. albicans. CAPE not only deprived iron and increased ROS production in C. albicans YEM30 cells but when used in combination with caspofungin, there was a significant 16-fold decrease for the minimum inhibitory concentrations (MICs) of CAPE and caspofungin compared to the MIC values of individual drugs [125].
The natural plant metabolite, poacic acid (diferulate, 8–5-DC; PA), was shown to bind to cell wall β-1,3-glucan in S. cerevisiae and inhibit β-1,3-glucan synthase activity in vitro [126]. PA also inhibited the growth of plant fungal pathogens Sclerotinia sclerotiorum and Alternaria solani and the oomycete Phytophthora sojae [126]. Work by Lee et al. [127] explored the effects of PA against human pathogenic Candida species. C. guilliermondii, C. orthopsilosis, and C. parapsilosis were more sensitive to PA than C. albicans, C. dubliniensis, C. glabrata, and C. tropicalis [127]. Furthermore, C. albicans strains containing an amino acid substitution in Fks1 Hotspot1 (S645Y or S645P) not only had decreased sensitivity to caspofungin but increased sensitivity to PA suggesting that there is a difference in the mode of action of PA and caspofungin [127].
Conclusion: what else could we learn about caspofungin?
Recent cryo-electron tomography work by Jiménez-Ortigosa et al. [128] has provided a preliminary structure of the putative C. glabrata GS complex, i.e., as clusters of hexamers, each subunit with two notable cytosolic domains, the N-terminal and central catalytic domains. The mechanism of action for echinocandins is its ability to inhibit β-1,3-glucan synthesis by non-competitive binding to GS. Fluorescence microscopy work by Utsugi et al. [129] demonstrated with the movement of Fks1p tagged with the green fluorescence protein was colocalized with cortical actin on the S. cerevisiae cell surface. Therefore, it would be interesting to see how various mutations in the FKS1/FKS2 and the application of caspofungin (or other echinocandins) in synergy with other drugs could affect the assembly of GS on the cell plasma membrane and β-1,3-glucan synthesis in C. albicans and other pathogenic yeasts.
References
Letscher-Bru V, Herbrecht R (2003) Caspofungin: the first representative of a new antifungal class. J Antimicrob Chemother 51:513–521. https://doi.org/10.1093/jac/dkg117
Patil A, Majumdar S (2017) Echinocandins in antifungal pharmacotherapy. J Pharm Pharmacol 69:1635–1660. https://doi.org/10.1111/jphp.12780
Sucher AJ, Chahine EB, Balcer HE (2009) Echinocandins: the newest class of antifungals. Ann Pharmacother 43:1647–1657. https://doi.org/10.1345/aph.1M237
Garcia-Effron G (2020) Rezafungin-mechanisms of action, susceptibility and resistance: similarities and differences with the other echinocandins. J Fungi 6:262. https://doi.org/10.3390/jof6040262
Sofjan AK, Mitchell A, Shah DN, Nguyen T, Sim M, Trojcak A, Beyda ND, Garey KW (2018) Rezafungin (CD101), a next-generation echinocandin: a systematic literature review and assessment of possible place in therapy. J Glob Antimicrob Resist 14:58–64. https://doi.org/10.1016/j.jgar.2018.02.013
Stewart AG, Paterson DL (2021) How urgent is the need for new antifungals? Expert Opin Pharmacother. https://doi.org/10.1080/14656566.2021.1935868
Lenardon MD, Sood P, Dorfmueller HC, Brown AJ, Gow NA (2020) Scalar nanostructure of the Candida albicans cell wall: a molecular, cellular and ultrastructural analysis and interpretation. Cell Surf 6:100047. https://doi.org/10.1016/j.tcsw.2020.100047
Zlotnik H, Fernandez MP, Bowers B, Cabib E (1984) Saccharomyces cerevisiae mannoproteins form an external cell wall layer that determines wall porosity. J Bacteriol 159:1018–1026. https://doi.org/10.1128/jb.159.3.1018-1026.1984
Kollár R, Petráková E, Ashwell G, Robbins PW, Cabib E (1995) Architecture of the yeast cell wall. The linkage between chitin and β (1→3)-glucan. J Biol Chem 270:1170–1178. https://doi.org/10.1074/jbc.270.3.117
Kollár R, Reinhold BB, Petráková E, Yeh HJ, Ashwell G, Drgonová J, Kapteyn JC, Klis FM, Cabib E (1997) Architecture of the yeast cell wall. β (1→6)-glucan interconnects mannoprotein, β (1→3)-glucan, and chitin. J Biol Chem 272:17762–17775. https://doi.org/10.1074/jbc.272.28.17762
Cabib E, Blanco N, Arroyo J (2012) Presence of a large β-(1–3)-glucan linked to chitin at the Saccharomyces cerevisiae mother-bud neck suggests involvement in localized growth control. Eukaryot Cell 11:388–400. https://doi.org/10.1128/EC.05328-11
Shahinian S, Bussey H (2000) β-1,6-glucan synthesis in Saccharomyces cerevisiae. Mol Microbiol 35:477–489. https://doi.org/10.1046/j.1365-2958.2000.01713.x
Shahinian S, Dijkgraaf GJ, Sdicu AM, Thomas DY, Jakob CA, Aebi M, Bussey H (1998) Involvement of protein N-glycosyl chain glucosylation and processing in the biosynthesis of cell wall β-1,6-glucan of Saccharomyces cerevisiae. Genetics 149:843–856
Inoue SB, Takewakt N, Takasuka T, Mio T, Adachi M, Fujii Y, Miyamoto C, Arisawa M, Furuichi Y, Watanabe T (1995) Characterization and gene cloning of 1,3-β-D-glucan synthase from Saccharomyces cerevisiae. Eur J Biochem 231:845–854. https://doi.org/10.1111/j.1432-1033.1995.0845d.x
Dijkgraaf GJP, Abe M, Ohya Y, Bussey H (2002) Mutations in Fks1p affect the cell wall content of β-1,3- and β-1,6-glucan in Saccharomyces cerevisiae. Yeast 19:671–690. https://doi.org/10.1002/yea.866
Douglas CM, Foor F, Marrinan JA, Morin N, Nielsen JB, Dahl AM, Mazur P, Baginsky W, Li W, El-Sherbeini M (1994) The Saccharomyces cerevisiae FKS1 (ETG1) gene encodes an integral membrane protein which is a subunit of 1,3-β-D-glucan synthase. Proc Nat Acad Sci USA 91:12907–12911. https://doi.org/10.1073/pnas.91.26.12907
Mazur P, Morin N, Baginsky W, el-Sherbeini M, Clemas JA, Nielsen JB, Foor F (1995) Differential expression and function of two homologous subunits of yeast 1,3–β-D-glucan synthase. Mol Cell Biol 15:5671–5681. https://doi.org/10.1128/MCB.15.10.5671
Drgonová J, Drgon T, Tanaka K, Kollár R, Chen GC, Ford RA, Chan CS, Takai Y, Cabib E (1996) Rho1p, a yeast protein at the interface between cell polarization and morphogenesis. Science 272:277–279. https://doi.org/10.1126/science.272.5259.277
Mazur P, Baginsky W (1996) In vitro activity of 1,3-β-D-glucan synthase requires the GTP-binding protein Rho1. J Biol Chem 271:14604–14609. https://doi.org/10.1074/jbc.271.24.14604
Qadota H, Python CP, Inoue SB, Arisawa M, Anraku Y, Zheng Y, Watanabe T, Levin DE, Ohya Y (1996) Identification of yeast Rho1p GTPase as a regulatory subunit of 1,3-β-glucan synthase. Science 272:279–281. https://doi.org/10.1126/science.272.5259.279
Ishihara S, Hirata A, Nogami S, Beauvais A, Latge JP, Ohya Y (2007) Homologous subunits of 1,3-β-glucan synthase are important for spore wall assembly in Saccharomyces cerevisiae. Eukaryot Cell 6:143–156. https://doi.org/10.1128/EC.00200-06
Douglas CM, Marrinan JA, Li W, Kurtz MB (1994) A Saccharomyces cerevisiae mutant with echinocandin-resistant 1,3-β-D-glucan synthase. J Bacteriol 176:5686–5696. https://doi.org/10.1128/jb.176.18.5686-5696.1994
El-Sherbeini M, Clemas JA (1995) Nikkomycin Z supersensitivity of an echinocandin-resistant mutant of Saccharomyces cerevisiae. Antimicrob Agents Chemother 39:200–207. https://doi.org/10.1128/AAC.39.1.200
Lesage G, Sdicu AM, Ménard P, Shapiro J, Hussein S, Bussey H (2004) Analysis of β-1,3-glucan assembly in Saccharomyces cerevisiae using a synthetic interaction network and altered sensitivity to caspofungin. Genetics 167:35–49. https://doi.org/10.1534/genetics.167.1.35
Markovich S, Yekutiel A, Shalit I, Shadkchan Y, Osherov N (2004) Genomic approach to identification of mutations affecting caspofungin susceptibility in Saccharomyces cerevisiae. Antimicrob Agents Chemother 48:3871–3876. https://doi.org/10.1128/AAC.48.10.3871-3876.2004
Carolus H, Pierson S, Muñoz JF, Subotić A, Cruz RB, Cuomo CA, Van Dijck P (2021) Genome-wide analysis of experimentally evolved Candida auris reveals multiple novel mechanisms of multidrug resistance. mBio 12:e03333-20. https://doi.org/10.1128/mBio.03333-20
Rybak JM, Dickens CM, Parker JE, Caudle KE, Manigaba K, Whaley SG, Nishimoto AT, Luna-Tapia A, Roy S, Zhang Q, Barker KS, Palmer GE, Sutter TR, Homayouni R, Wiederhold NP, Kelly SL, Rogers PD (2017) Loss of C-5 sterol desaturase activity results in increased resistance to azole and echinocandin antifungals in a clinical isolate of Candida parapsilosis. Antimicrob Agents Chemother 61:e00651-e717. https://doi.org/10.1128/AAC.00651-17
Spettel K, Barousch W, Makristathis A, Zeller I, Nehr M, Selitsch B, Lackner M, Rath PM, Steinmann J, Willinger B (2019) Analysis of antifungal resistance genes in Candida albicans and Candida glabrata using next generation sequencing. PLoS ONE 14:e0210397. https://doi.org/10.1371/journal.pone.0210397
Mio T, Adachi-Shimizu M, Tachibana Y, Tabuchi H, Inoue SB, Yabe T, Yamada-Okabe T, Arisawa M, Watanabe T, Yamada-Okabe H (1997) Cloning of the Candida albicans homolog of Saccharomyces cerevisiae GSC1/FKS1 and its involvement in β-1,3-glucan synthesis. J Bacteriol 179:4096–4105. https://doi.org/10.1128/jb.179.13.4096-4105.1997
Douglas CM, D’Ippolito JA, Shei GJ, Meinz M, Onishi J, Marrinan JA, Li W, Abruzzo GK, Flattery A, Bartizal K, Mitchell A, Kurtz MB (1997) Identification of the FKS1 gene of Candida albicans as the essential target of 1,3-β-D-glucan synthase inhibitors. Antimicrob Agents Chemother 41:2471–2479. https://doi.org/10.1128/AAC.41.11.2471
Suwunnakorn S, Wakabayashi H, Kordalewska M, Perlin DS, Rustchenko E (2018) FKS2 and FKS3 genes of opportunistic human pathogen Candida albicans influence echinocandin susceptibility. Antimicrob Agents Chemother 62:e02299-e2317. https://doi.org/10.1128/AAC.02299-17
Balashov SV, Park S, Perlin DS (2006) Assessing resistance to the echinocandin antifungal drug caspofungin in Candida albicans by profiling mutations in FKS1. Antimicrob Agents Chemother 50:2058–2063. https://doi.org/10.1128/AAC.01653-05
Hori Y, Shibuya K (2018) Role of FKS gene in the susceptibility of pathogenic fungi to echinocandins. Med Mycol J 59:E31–E40. https://doi.org/10.3314/mmj.18.004
Pristov K, Ghannoum M (2019) Resistance of Candida to azoles and echinocandins worldwide. Clin Microbiol Infect 25:792–798. https://doi.org/10.1016/j.cmi.2019.03.028
Walker LA, Gow NA, Munro CA (2010) Fungal echinocandin resistance. Fungal Genet Biol 47:117–126. https://doi.org/10.1016/j.fgb.2009.09.003
Perlin DS (2015) Echinocandin resistance in Candida. Clin Infect Dis 61:S612–S617. https://doi.org/10.1093/cid/civ791
Park S, Kelly R, Kahn JN, Robles J, Hsu MJ, Register E, Li W, Vyas V, Fan H, Abruzzo G, Flattery A, Gill C, Chrebet G, Parent SA, Kurtz M, Teppler H, Douglas CM, Perlin DS (2005) Specific substitutions in the echinocandin target Fks1p account for reduced susceptibility of rare laboratory and clinical Candida sp. isolates. Antimicrob Agents Chemother 49:3264–3273. https://doi.org/10.1128/AAC.49.8.3264-3273.2005
Fidel PL Jr, Vazquez JA, Sobel JD (1999) Candida glabrata: review of epidemiology, pathogenesis, and clinical disease with comparison to C. albicans. Clin Microbiol Rev 12:80–96. https://doi.org/10.1128/CMR.12.1.80
Katiyar S, Pfaller M, Edlind T (2006) Candida albicans and Candida glabrata clinical isolates exhibiting reduced echinocandin susceptibility. Antimicrob Agents Chemother 50:2892–2894. https://doi.org/10.1128/AAC.00349-06
Garcia-Effron G, Lee S, Park S, Cleary JD, Perlin DS (2009) Effect of Candida glabrata FKS1 and FKS2 mutations on echinocandin sensitivity and kinetics of 1,3-β-D-glucan synthase: implication for the existing susceptibility breakpoint. Antimicrob Agents Chemother 53:3690–3699. https://doi.org/10.1128/AAC.00443-09
Katiyar SK, Alastruey-Izquierdo A, Healey KR, Johnson ME, Perlin DS, Edlind TD (2012) Fks1 and Fks2 are functionally redundant but differentially regulated in Candida glabrata: implications for echinocandin resistance. Antimicrob Agents Chemother 56:6304–6309. https://doi.org/10.1128/AAC.00813-12
Hou X, Healey KR, Shor E, Kordalewska M, Ortigosa CJ, Paderu P, Xiao M, Wang H, Zhao Y, Lin LY, Zhang YH, Li YZ, Xu YC, Perlin DS, Zhao Y (2019) Novel FKS1 and FKS2 modifications in a high-level echinocandin resistant clinical isolate of Candida glabrata. Emerg Microbes Infecti 8:1619–1625. https://doi.org/10.1080/22221751.2019.1684209
Lara-Aguilar V, Rueda C, García-Barbazán I, Varona S, Monzón S, Jiménez P, Cuesta I, Zaballos Á, Zaragoza Ó (2021) Adaptation of the emerging pathogenic yeast Candida auris to high caspofungin concentrations correlates with cell wall changes. Virulence 12:1400–1417. https://doi.org/10.1080/21505594.2021.1927609
Accoceberry I, Couzigou C, Fitton-Ouhabi V, Biteau N, Noël T (2019) Challenging SNP impact on caspofungin resistance by full-length FKS1 allele replacement in Candida lusitaniae. J Antimicrob Chemother 74:618–624. https://doi.org/10.1093/jac/dky475
Arendrup MC (2013) Candida and candidaemia. Susceptibility and epidemiology. Dan Med J 60:B4698
Arendrup MC, Perlin DS (2014) Echinocandin resistance: an emerging clinical problem? Curr Opin Infect Dis 27:484–492. https://doi.org/10.1097/QCO.0000000000000111
Lackner M, Tscherner M, Schaller M, Kuchler K, Mair C, Sartori B, Istel F, Arendrup MC, Lass-Flörl C (2014) Positions and numbers of FKS mutations in Candida albicans selectively influence in vitro and in vivo susceptibilities to echinocandin treatment. Antimicrob Agents Chemother 58:3626–3635. https://doi.org/10.1128/AAC.00123-14
Jaber QZ, Bibi M, Ksiezopolska E, Gabaldon T, Berman J, Fridman M (2020) Elevated vacuolar uptake of fluorescently labeled antifungal drug caspofungin predicts echinocandin resistance in pathogenic yeast. ACS Cent Sci 6:1698–1712. https://doi.org/10.1021/acscentsci.0c00813
E Silva AP, Miranda IM, Branco J, Oliveira P, Faria-Ramos I, Silva RM, Rodrigues AG, Costa-de-Oliveira S (2020) FKS1 mutation associated with decreased echinocandin susceptibility of Aspergillus fumigatus following anidulafungin exposure. Sci Rep 10:11976. https://doi.org/10.1038/s41598-020-68706-8
Johnson ME, Katiyar SK, Edlind TD (2011) New Fks hot spot for acquired echinocandin resistance in Saccharomyces cerevisiae and its contribution to intrinsic resistance of Scedosporium species. Antimicrob Agents Chemother 55:3774–3781. https://doi.org/10.1128/AAC.01811-10
Katiyar SK, Edlind TD (2009) Role for Fks1 in the intrinsic echinocandin resistance of Fusarium solani as evidenced by hybrid expression in Saccharomyces cerevisiae. Antimicrob Agents Chemother 53:1772–1778. https://doi.org/10.1128/AAC.00020-09
Yang F, Zhang L, Wakabayashi H, Myers J, Jiang Y, Cao Y, Jimenez-Ortigosa C, Perlin DS, Rustchenko E (2017) Tolerance to caspofungin in Candida albicans is associated with at least three distinctive mechanisms that govern expression of FKS genes and cell wall remodeling. Antimicrob Agents Chemother 61:e00071-e117. https://doi.org/10.1128/AAC.00071-17
Yang F, Kravets A, Bethlendy G, Welle S, Rustchenko E (2013) Chromosome 5 monosomy of Candida albicans controls susceptibility to various toxic agents, including major antifungals. Antimicrob Agents Chemother 57:5026–5036. https://doi.org/10.1128/AAC.00516-13
Badrane H, Nguyen MH, Clancy CJ (2016) Highly dynamic and specific phosphatidylinositol 4,5-bisphosphate, septin, and cell wall integrity pathway responses correlate with caspofungin activity against Candida albicans. Antimicrob Agents Chemother 60:3591–3600. https://doi.org/10.1128/AAC.02711-15
Hao B, Cheng S, Clancy CJ, Nguyen MH (2013) Caspofungin kills Candida albicans by causing both cellular apoptosis and necrosis. Antimicrob Agents Chemother 57:326–332. https://doi.org/10.1128/AAC.01366-12
Shirazi F, Kontoyiannis DP (2015) Micafungin triggers caspase-dependent apoptosis in Candida albicans and Candida parapsilosis biofilms, including caspofungin non-susceptible isolates. Virulence 6:385–394. https://doi.org/10.1080/21505594.2015.1027479
Bizerra FC, Melo AS, Katchburian E, Freymüller E, Straus AH, Takahashi HK, Colombo AL (2011) Changes in cell wall synthesis and ultrastructure during paradoxical growth effect of caspofungin on four different Candida species. Antimicrob Agents Chemother 55:302–310. https://doi.org/10.1128/AAC.00633-10
Nishiyama Y, Uchida K, Yamaguchi H (2002) Morphological changes of Candida albicans induced by micafungin (FK463), a water-soluble echinocandin-like lipopeptide. J Electron Microsc 51:247–255. https://doi.org/10.1093/jmicro/51.4.247
Rueda C, Cuenca-Estrella M, Zaragoza O (2014) Paradoxical growth of Candida albicans in the presence of caspofungin is associated with multiple cell wall rearrangements and decreased virulence. Antimicrob Agents Chemother 58:1071–1083. https://doi.org/10.1128/AAC.00946-13
Moreno-Velásquez SD, Seidel C, Juvvadi PR, Steinbach WJ, Read ND (2017) Caspofungin-mediated growth inhibition and paradoxical growth in Aspergillus fumigatus involve fungicidal hyphal tip lysis coupled with regenerative intrahyphal growth and dynamic changes in β-1,3-glucan synthase localization. Antimicrob Agents Chemother 61:e00710-e717. https://doi.org/10.1128/AAC.00710-17
Wagener J, Loiko V (2017) Recent insights into the paradoxical effect of echinocandins. J Fungi (Basel) 4:5. https://doi.org/10.3390/jof4010005
Fortwendel JR, Juvvadi PR, Perfect BZ, Rogg LE, Perfect JR, Steinbach WJ (2010) Transcriptional regulation of chitin synthases by calcineurin controls paradoxical growth of Aspergillus fumigatus in response to caspofungin. Antimicrob Agents Chemother 54:1555–1563. https://doi.org/10.1128/AAC.00854-09
El-Kirat-Chatel S, Beaussart A, Alsteens D, Jackson DN, Lipke PN, Dufrêne YF (2013) Nanoscale analysis of caspofungin-induced cell surface remodelling in Candida albicans. Nanoscale 5:1105–1115. https://doi.org/10.1039/c2nr33215a
Quilès F, Accoceberry I, Couzigou C, Francius G, Noël T, El-Kirat-Chatel S (2017) AFM combined to ATR-FTIR reveals Candida cell wall changes under caspofungin treatment. Nanoscale 9:13731–13738. https://doi.org/10.1039/c7nr02170d
Walker LA, Munro CA, de Bruijn I, Lenardon MD, McKinnon A, Gow NA (2008) Stimulation of chitin synthesis rescues Candida albicans from echinocandins. PLoS Pathog 4:e1000040. https://doi.org/10.1371/journal.ppat.1000040
Lee KK, MacCallum DM, Jacobsen MD, Walker LA, Odds FC, Gow NA, Munro CA (2012) Elevated cell wall chitin in Candida albicans confers echinocandin resistance in vivo. Antimicrob Agents Chemother 56:208–217. https://doi.org/10.1128/AAC.00683-11
Walker LA, Gow NA, Munro CA (2013) Elevated chitin content reduces the susceptibility of Candida species to caspofungin. Antimicrob Agents Chemother 57:146–154. https://doi.org/10.1128/AAC.01486-12
Munro CA, Selvaggini S, de Bruijn I, Walker L, Lenardon MD, Gerssen B, Milne S, Brown AJ, Gow NA (2007) The PKC, HOG and Ca2+ signalling pathways co-ordinately regulate chitin synthesis in Candida albicans. Mol Microbiol 63:1399–1413. https://doi.org/10.1111/j.1365-2958.2007.05588.x
Han Q, Wang N, Pan C, Wang Y, Sang J (2019) Elevation of cell wall chitin via Ca2+–calcineurin-mediated PKC signaling pathway maintains the viability of Candida albicans in the absence of β-1,6-glucan synthesis. Mol Microbiol 112:960–972. https://doi.org/10.1111/mmi.14335
García-Rodriguez LJ, Trilla JA, Castro C, Valdivieso MH, Durán A, Roncero C (2000) Characterization of the chitin biosynthesis process as a compensatory mechanism in the fks1 mutant of Saccharomyces cerevisiae. FEBS Lett 478:84–88. https://doi.org/10.1016/s0014-5793(00)01835-4
Perrine-Walker F, Payne J (2021) Rapid screening method of Saccharomyces cerevisiae mutants using calcofluor white and aniline blue. Braz J Microbiol 52:1077–1086. https://doi.org/10.1007/s42770-021-00515-1
Heredia MY, Gunasekaran D, Ikeh M, Nobile CJ, Rauceo JM (2020) Transcriptional regulation of the caspofungin-induced cell wall damage response in Candida albicans. Curr Genet 66:1059–1068. https://doi.org/10.1007/s00294-020-01105-8
Rauceo JM, Blankenship JR, Fanning S, Hamaker JJ, Deneault JS, Smith FJ, Nantel A, Mitchell AP (2008) Regulation of the Candida albicans cell wall damage response by transcription factor Sko1 and PAS kinase Psk1. Mol Biol Cell 19:2741–2751. https://doi.org/10.1091/mbc.e08-02-0191
Gelis S, de Groot PW, Castillo L, Moragues MD, Sentandreu R, Gómez MM, Valentín E (2012) Pga13 in Candida albicans is localized in the cell wall and influences cell surface properties, morphogenesis and virulence. Fungal Genet Biol 49:322–331. https://doi.org/10.1016/j.fgb.2012.01.010
Alonso-Monge R, Román E, Arana DM, Prieto D, Urrialde V, Nombela C, Pla J (2010) The Sko1 protein represses the yeast-to-hypha transition and regulates the oxidative stress response in Candida albicans. Fungal Genet Biol 47:587–601. https://doi.org/10.1016/j.fgb.2010.03.009
Fan Y, He H, Dong Y, Pan H (2013) Hyphae-specific genes HGC1, ALS3, HWP1, and ECE1 and relevant signaling pathways in Candida albicans. Mycopathologia 176:329–335. https://doi.org/10.1007/s11046-013-9684-6
Delgado-Silva Y, Vaz C, Carvalho-Pereira J, Carneiro C, Nogueira E, Correia A, Carreto L, Silva S, Faustino A, Pais C, Oliveira R, Sampaio P (2014) Participation of Candida albicans transcription factor RLM1 in cell wall biogenesis and virulence. PLoS ONE 9:e86270. https://doi.org/10.1371/journal.pone.0086270
Hoyer LL, Green CB, Oh S-H, Zhao X (2008) Discovering the secrets of the Candida albicans agglutinin-like sequence (ALS) gene family - a sticky pursuit. Med Mycol 46:1–15. https://doi.org/10.1080/13693780701435317
Hoyer LL, Cota E (2016) Candida albicans agglutinin-like sequence (Als) family vignettes: a review of als protein structure and function. Front Microbiol 7:280. https://doi.org/10.3389/fmicb.2016.00280
Willaert RG (2018) Adhesins of yeasts: protein structure and interactions. J Fungi 4:119. https://doi.org/10.3390/jof4040119
Gregori C, Glaser W, Frohner IE, Reinoso-Martín C, Rupp S, Schüller C, Kuchler K (2011) Efg1 Controls caspofungin-induced cell aggregation of Candida albicans through the adhesin Als1. Eukaryot Cell 10:1694–1704. https://doi.org/10.1128/EC.05187-11
Connolly LA, Riccombeni A, Grózer Z, Holland LM, Lynch DB, Andes DR, Gácser A, Butler G (2013) The APSES transcription factor Efg1 is a global regulator that controls morphogenesis and biofilm formation in Candida parapsilosis. Mol Microbiol 90:36–53. https://doi.org/10.1111/mmi.12345
Villa S, Hamideh M, Weinstock A, Qasim MN, Hazbun TR, Sellam A, Hernday AD, Thangamani S (2020) Transcriptional control of hyphal morphogenesis in Candida albicans. FEMS Yeast Res 20:foaa005. https://doi.org/10.1093/femsyr/foaa005
Noffz CS, Liedschulte V, Lengeler K, Ernst JF (2008) Functional mapping of the Candida albicans Efg1 regulator. Eukaryot Cell 7:881–893. https://doi.org/10.1128/EC.00033-08
Stoldt VR, Sonneborn A, Leuker CE, Ernst JF (1997) Efg1p, an essential regulator of morphogenesis of the human pathogen Candida albicans, is a member of a conserved class of bHLH proteins regulating morphogenetic processes in fungi. EMBO J 16:1982–1991. https://doi.org/10.1093/emboj/16.8.1982
Bauer J, Wendland J (2007) Candida albicans Sfl1 suppresses flocculation and filamentation. Eukaryot Cell 6:1736–1744. https://doi.org/10.1128/EC.00236-07
Li Y, Su C, Mao X, Cao F, Chen J (2007) Roles of Candida albicans Sfl1 in hyphal development. Eukaryot Cell 6:2112–2121. https://doi.org/10.1128/EC.00199-07
McCall AD, Kumar R, Edgerton M (2018) Candida albicans Sfl1/Sfl2 regulatory network drives the formation of pathogenic microcolonies. PLOS Pathog 14:e1007316. https://doi.org/10.1371/journal.ppat.1007316
Bruno VM, Kalachikov S, Subaran R, Nobile CJ, Kyratsous C, Mitchell AP (2006) Control of the C. albicans cell wall damage response by transcriptional regulator Cas5. PLoS Pathog 2:e21. https://doi.org/10.1371/journal.ppat.0020021
Xie JL, Qin L, Miao Z, Grys BT, Diaz JC, Ting K, Krieger JR, Tong J, Tan K, Leach MD, Ketela T, Moran MF, Krysan DJ, Boone C, Andrews BJ, Selmecki A, Ho Wong K, Robbins N, Cowen LE (2017) The Candida albicans transcription factor Cas5 couples stress responses, drug resistance and cell cycle regulation. Nature Commun 8:499. https://doi.org/10.1038/s41467-017-00547-y
Xiong K, Su C, Sun Q, Lu Y (2021) Efg1 and Cas5 orchestrate cell wall damage response to caspofungin in Candida albicans. Antimicrob Agents Chemother 65:e01584-e1620. https://doi.org/10.1128/AAC.01584-20
Plaine A, Walker L, Da Costa G, Mora-Montes HM, McKinnon A, Gow NA, Gaillardin C, Munro CA, Richard ML (2008) Functional analysis of Candida albicans GPI-anchored proteins: roles in cell wall integrity and caspofungin sensitivity. Fungal Genet Biol 45:1404–1414. https://doi.org/10.1016/j.fgb.2008.08.003
Walker LA, Munro CA (2020) Caspofungin induced cell wall changes of Candida species influences macrophage interactions. Front Cell Infect Microbiol 10:164. https://doi.org/10.3389/fcimb.2020.00164
Mora-Montes HM, Netea MG, Ferwerda G, Lenardon MD, Brown GD, Mistry AR, Kullberg BJ, O’Callaghan CA, Sheth CC, Odds FC, Brown AJ, Munro CA, Gow NA (2011) Recognition and blocking of innate immunity cells by Candida albicans chitin. Infect Immun 79:1961–1970. https://doi.org/10.1128/IAI.01282-10
Davis SE, Hopke A, Minkin SC Jr, Montedonico AE, Wheeler RT, Reynolds TB (2014) Masking of β(1–3)-glucan in the cell wall of Candida albicans from detection by innate immune cells depends on phosphatidylserine. Infect Immun 82:4405–4413. https://doi.org/10.1128/IAI.01612-14
Wheeler RT, Fink GR (2006) A drug-sensitive genetic network masks fungi from the immune system. PLoS Pathog 2:e35. https://doi.org/10.1371/journal.ppat.0020035
Herrero AB, Magnelli P, Mansour MK, Levitz SM, Bussey H, Abeijon C (2004) KRE5 gene null mutant strains of Candida albicans are avirulent and have altered cell wall composition and hypha formation properties. Eukaryot Cell 3:1423–1432. https://doi.org/10.1128/EC.3.6.1423-1432.2004
Tanaka Y, Sasaki M, Ito F, Aoyama T, Sato-Okamoto M, Takahashi-Nakaguchi A, Chibana H, Shibata N (2016) KRE5 suppression induces cell wall stress and alternative ER stress response required for maintaining cell wall Integrity in Candida glabrata. PLoS ONE 11:e0161371. https://doi.org/10.1371/journal.pone.0161371
Hasim S, Allison DP, Retterer ST, Hopke A, Wheeler RT, Doktycz MJ, Reynolds TB (2016) β-(1,3)-Glucan unmasking in Some Candida albicans mutants correlates with increases in cell wall surface roughness and decreases in cell wall elasticity. Infect Immun 85:e00601-e616. https://doi.org/10.1128/IAI.00601-16
Adams EL, Rice PJ, Graves B, Ensley HE, Yu H, Brown GD, Gordon S, Monteiro MA, Papp-Szabo E, Lowman DW, Power TD, Wempe MF, Williams DL (2008) Differential high-affinity interaction of dectin-1 with natural or synthetic glucans is dependent upon primary structure and is influenced by polymer chain length and side-chain branching. J Pharmacol Exp Ther 325:115–123. https://doi.org/10.1124/jpet.107.133124
Gulati M, Nobile CJ (2016) Candida albicans biofilms: development, regulation, and molecular mechanisms. Microbes Infect 18:310–321. https://doi.org/10.1016/j.micinf.2016.01.002
Mayer FL, Wilson D, Hube B (2013) Candida albicans pathogenicity mechanisms. Virulence 4:119–128. https://doi.org/10.4161/viru.22913
Pereira R, dos Santos FR, de Brito E, de Morais S (2021) Biofilm of Candida albicans: formation, regulation and resistance. J Appl Microbiol 131:11–22. https://doi.org/10.1111/jam.14949
Araújo D, Henriques M, Silva S (2016) Portrait of Candida species biofilm regulatory network genes. Trends Microbiol 25:62–75. https://doi.org/10.1016/j.tim.2016.09.004
Cavalheiro M, Teixeira MC (2018) Candida biofilms: threats, challenges, and promising strategies. Front Med 5:28. https://doi.org/10.3389/fmed.2018.00028
Chandra J, Kuhn DM, Mukherjee PK, Hoyer LL, McCormick T, Ghannoum MA (2001) Biofilm formation by the fungal pathogen Candida albicans: development, architecture, and drug resistance. J Bacteriol 183:5385–5394. https://doi.org/10.1128/JB.183.18.5385-5394.2001
Rodríguez-Cerdeira C, Martínez-Herrera E, Carnero-Gregorio M, López-Barcenas A, Fabbrocini G, Fida M, El-Samahy M, González-Cespón JL (2020) Pathogenesis and clinical relevance of Candida biofilms in vulvovaginal candidiasis. Front Microbiol 11:544480. https://doi.org/10.3389/fmicb.2020.544480
Alim D, Sircaik S, Panwar SL (2018) The significance of lipids to biofilm formation in Candida albicans: an emerging perspective. J Fungi 4:140. https://doi.org/10.3390/jof4040140
Finkel JS, Mitchell A (2011) Genetic control of Candida albicans biofilm development. Nature Rev Microbiol 9:109–118. https://doi.org/10.1038/nrmicro2475
Bachmann SP, VandeWalle K, Ramage G, Patterson TF, Wickes BL, Graybill JR, López-Ribot JL (2002) In vitro activity of caspofungin against Candida albicans biofilms. Antimicrob Agents Chemother 46:3591–3596. https://doi.org/10.1128/AAC.46.11.3591-3596.2002
Ferreira JA, Carr JH, Starling CE, de Resende MA, Donlan RM (2009) Biofilm formation and effect of caspofungin on biofilm structure of Candida species bloodstream isolates. Antimicrob Agents Chemother 53:4377–4384. https://doi.org/10.1128/AAC.00316-09
Melo AS, Colombo AL, Arthington-Skaggs BA (2007) Paradoxical growth effect of caspofungin observed on biofilms and planktonic cells of five different Candida species. Antimicrob Agents Chemother 51:3081–3088. https://doi.org/10.1128/AAC.00676-07
Chandra J, Ghannoum MA (2018) CD101, a novel echinocandin, possesses potent antibiofilm activity against early and mature Candida albicans biofilms. Antimicrob Agents Chemother 62:e01750-e1817. https://doi.org/10.1128/AAC.01750-17
Larkin EL, Dharmaiah S, Ghannoum MA (2018) Biofilms and beyond: expanding echinocandin utility. J Antimicrob Chemother 73:i73–i81. https://doi.org/10.1093/jac/dkx451
Swaminathan S, Kamat S, Pinto NA (2018) Echinocandins: their role in the management of Candida biofilms. Indian J Med Microbiol 36:87–92. https://doi.org/10.4103/ijmm.IJMM_17_400
Delattin N, De Brucker K, Vandamme K, Meert E, Marchand A, Chaltin P, Cammue BP, Thevissen K (2014) Repurposing as a means to increase the activity of amphotericin B and caspofungin against Candida albicans biofilms. J Antimicrob Chemother 69:1035–1044. https://doi.org/10.1093/jac/dkt449
MacCallum DM, Desbois AP, Coote PJ (2013) Enhanced efficacy of synergistic combinations of antimicrobial peptides with caspofungin versus Candida albicans in insect and murine models of systemic infection. Eur J Clin Microbiol Infect Dis 32:1055–1062. https://doi.org/10.1007/s10096-013-1850-8
Troskie AM, Rautenbach M, Delattin N, Vosloo JA, Dathe M, Cammue BPA, Thevissen K (2014) Synergistic activity of the tyrocidines, antimicrobial cyclodecapeptides from Bacillus aneurinolyticus, with amphotericin B and caspofungin against Candida albicans biofilms. Antimicrob Agents Chemother 58:3697–3707. https://doi.org/10.1128/AAC.02381-14
Oshiro KGN, Rodrigues G, Monges BED, Cardoso MH, Franco OL (2019) Bioactive peptides against fungal biofilms. Front Microbiol 10:2169. https://doi.org/10.3389/fmicb.2019.02169
Wei G-X, Bobek LA (2005) Human salivary mucin MUC7 12-mer-L and 12-mer-D peptides: antifungal activity in saliva, enhancement of activity with protease inhibitor cocktail or EDTA, and cytotoxicity to human cells. Antimicrob Agents Chemother 49:2336–2342. https://doi.org/10.1128/AAC.49.6.2336-2342.2005
Mor A, Hani K, Nicolas P (1994) The vertebrate peptide antibiotics dermaseptins have overlapping structural features but target specific microorganisms. J Biol Chem 269:31635–31641
Thevissen K, de Mello TP, Xu D, Blankenship J, Vandenbosch D, Idkowiak-Baldys J, Govaert G, Bink A, Rozental S, de Groot PWJ, Davis TR, Kumamoto CA, Vargas G, Nimrichter L, Coenye T, Mitchell A, Roemer T, Hannun YA, Cammue BPA (2012) The plant defensin RsAFP2 induces cell wall stress, septin mislocalization and accumulation of ceramides in Candida albicans. Mol Microbiol 84:166–180. https://doi.org/10.1111/j.1365-2958.2012.08017.x
Vriens K, Cools TL, Harvey PJ, Craik DJ, Spincemaille P, Cassiman D, Braem A, Vleugels J, Nibbering PH, Drijfhout JW, De Coninck B, Cammue BP, Thevissen K (2015) Synergistic activity of the plant defensin HsAFP1 and caspofungin against Candida albicans biofilms and planktonic cultures. PLoS ONE 10:e0132701. https://doi.org/10.1371/journal.pone.0132701
Cools TL, Struyfs C, Drijfhout JW, Kucharíková S, Romero CL, Dijck PV, Ramada MHS, Bloch C, Cammue BP, Thevissen K (2017) A linear 19-mer plant defensin-derived peptide acts synergistically with caspofungin against Candida albicans biofilms. Front Microbiol 8:2051. https://doi.org/10.3389/fmicb.2017.02051
Sun L, Hang C, Liao K (2018) Synergistic effect of caffeic acid phenethyl ester with caspofungin against Candida albicans is mediated by disrupting iron homeostasis. Food Chem Toxicol 116:51–58. https://doi.org/10.1016/j.fct.2018.04.014
Piotrowski JS, Okada H, Lu F, Li SC, Hinchman L, Ranjan A, Smith DL, Higbee AJ, Ulbrich A, Coon JJ, Deshpande R, Bukhman YV, McIlwain S, Ong IM, Myers CL, Boone C, Landick R, Ralph J, Kabbage M, Ohya Y (2015) Plant-derived antifungal agent poacic acid targets β-1,3-glucan. Proc Nat Acad Sci USA 112:E1490–E1497. https://doi.org/10.1073/pnas.1410400112
Lee KK, Kubo K, Abdelaziz JA, Cunningham I, de Silva DA, Chen X, Okada H, Ohya Y, Gow N (2018) Yeast species-specific, differential inhibition of β-1,3-glucan synthesis by poacic acid and caspofungin. Cell Surf 3:12–25. https://doi.org/10.1016/j.tcsw.2018.09.001
Jiménez-Ortigosa C, Jiang J, Chen M, Kuang X, Healey KR, Castellano P, Boparai N, Ludtke SJ, Perlin DS, Dai W (2021) Preliminary structural elucidation of β-(1,3)- glucan synthase from Candida glabrata using cryo-electron tomography. J Fungi 7:120. https://doi.org/10.3390/jof7020120
Utsugi T, Minemura M, Hirata A, Abe M, Watanabe D, Ohya Y (2002) Movement of yeast 1,3-β-glucan synthase is essential for uniform cell wall synthesis. Genes Cells 7:1–9. https://doi.org/10.1046/j.1356-9597.2001.00495.x
Acknowledgements
Special acknowledgement is given to the BioImaging Platform at La Trobe University and Peter Lock for the helpful advice and training on various microscopes. The author would like to thank Marilyn Anderson for her support and Charles Hocart (Australian National University, Australia) for additional advice.
Author information
Authors and Affiliations
Contributions
F.P-W conceptualized the article, performed the literature search and data analysis, and drafted the manuscript and figures.
Corresponding author
Ethics declarations
Conflict of interest
The author declares no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Perrine-Walker, F. Caspofungin resistance in Candida albicans: genetic factors and synergistic compounds for combination therapies. Braz J Microbiol 53, 1101–1113 (2022). https://doi.org/10.1007/s42770-022-00739-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42770-022-00739-9