Abstract
Multidrug resistance in groundwater contaminants is becoming a major health concern in developing countries, especially the ampicillin-resistant Escherichia coli. This study aimed at profiling antibiotic resistance and possible detection of ampicillin-resistant genes in Escherichia coli obtained from forty groundwater samples (20 each of boreholes and wells) in Osogbo metropolis, Southwest Nigeria. Grab sampling was done using 1L sterile plastic bottles, isolation of E. coli was performed using pour plate technique on eosin methylene blue agar and their identity confirmed by polymerase chain reaction (PCR) using uidA gene. Antibiotics susceptibility test of the isolates to ten commercially available antibiotics was done following Kirby–Bauer disc diffusion technique. Multiple antibiotic resistant phenotypes (MARPs) and indexing (MARI) were estimated accordingly. The possible presence of Ctx-M, SHV, ampC, and TEM-H resistant genes in all ampicillin-resistant E. coli were checked for using PCR. All the 55 presumptive E. coli isolates, 35 from 10 boreholes and 20 from 7 wells, were uidA positive. Overall, 50 (91%) of the E. coli were resistant to ampicillin, followed by trimethoprim and ertapenem 43 (78%), doxycycline 40 (73%), ceftazidime 38 (69%), and tetracycline 37 (67%). Of the 55 E. coli isolates, only 1 was resistant to 2 drugs (AK-AMP), others were multi-resistant, ranging from 4 to 9 drugs, with the highest MARP (9) being AMP–CAZ–ETP–S–AK–DO–TE–W–C. MARI also between 0.4 and 0.9, above the 0.2 acceptable limit. Exactly 24 (48%) of the phenotypically ampicillin-resistant E. coli isolates harboured TEM-H only. The existence of multidrug-resistant E. coli and TEM-H resistant gene in the groundwater pose a huge threat to all and sundry who rely heavily on this source of water for diverse purposes. Hence, adequate monitoring and antimicrobial resistance surveillance of groundwater bodies is advocated to safeguard public health.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Water is a prerequisite for living. Its essentiality in the overall well-being of man cannot be overemphasized. Among diverse water sources, groundwater (borehole and hand-dug well) is considered the ideal candidate for drinking owing to its perceived cleanness and safety (Edokpayi et al. 2018a, b). It is presumed that the different earth’s strata filter out contaminants in surface water during percolation before reaching the underground (Keesari et al. 2015). Unfortunately, groundwaters are now recognised to be prone to microbial contamination. Over two decades ago, outbreaks of waterborne diseases were reported in some nations of the world following ingestion of contaminated groundwaters (Anderson and Bohan, 2001; Paruch et al. 2015; Vignesh et al. 2015). The location, type and degree of human activities, and environmental condition in and around groundwater play a significant role in influencing groundwater quality (Olasoji et al. 2019).
Pollution of groundwater arises when boreholes are drilled, and wells dug without adhering strictly to hygienic standards and/or consulting relevant regulatory agencies. Previous studies observed bacteria, including Escherichia coli, Clostridium perfringens, Staphylococcus aureus, Enterococci sp., somatic coliphages, species of Salmonella, Shigella, and Klebsiella, Pseudomonas aeruginosa, Vibrio parahaemolyticus, and Aeromonas hydrophila in groundwater supplies (Potgieter et al. 2006; Aydin 2007; Olaitan et al. 2013; Llopis-González et al. 2014; Owolabi et al. 2014; Keesari et al. 2015; Onyango et al. 2018; Odiyo and Makungo, 2018; Alsalme et al. 2021).
Microbiological assessment is vital to identifying contamination of water with pathogens capable of compromising human health and provides an avenue to avert potential health hazards including morbidity and mortality (Stein et al. 2010; Titilawo et al. 2020). Escherichia coli is commonly regarded as the indicator organism of microbial water contamination (USGS 2018). Albeit, some pathogenic strains of E. coli are responsible for several morbidities and mortalities in all ages (Kaper et al. 2004). E. coli is mostly significant owing to its ability to transmit water-related diseases such as diarrhoea, the second cause of mortality amidst children below 5 years of age (Chissaque et al. 2018). It accounts for approximately 9% of 5.6 million childhood deaths and 88% of all reported cases of diarrhoea in Black Africa and South Asia countries (Liu et al 2016; Chissaque et al. 2018).
Antimicrobials are key in reducing illnesses and deaths linked with veterinary and human infectious diseases. For E. coli infections, ampicillin has been the drug of choice for treatment but recently, its resistance rate has increased (Alanazi et al. 2018). In general, ampicillin prevents the synthesis of bacterial cell wall during replication in bacteria, however, resistant strains encode β-lactamase, changes the target protein in cell wall, reduces the permeability of outer membrane, and increases the expression of drug efflux pump (Kapoor et al. 2017).
The excessive and indiscriminate usage of antimicrobial in therapy, growth promotion, and prophylaxis in animal and human medicine are dominant drivers of the development and communicability of resistance traits among disease-causing and normal floral bacteria (Dadgostar 2019). In addition, complex socioeconomic and behavioural patterns traceable to individuals promote transmission of drug resistant microorganisms (Okeke et al. 1999; Medina and Pieper 2016). Aquatic milieu is known to play a key role in the dissemination of microbes among humans, animals and the environment (Amaya et al. 2012).
Poor water availability and inaccessibility to portable water by the people of Osogbo, Osun State, Nigeria is a great challenge. Available data reveal that about 844 million people do not have access to basic water service, portable water is not available for 2.1 billion people worldwide and the Goal 6 of sustainable development goals (SDGs) targeted at providing adequate supply and access to safe clean water by the year 2030 is gradually becoming a mirage (UN 2018). Thus, continuous supply of potable water remains a major challenge to millions of people around the world, especially in developing countries.
Osogbo township is blessed with abundance of natural water bodies but activities around them limit their usage for drinking purposes. Hence, majority of the inhabitants rely heavily on boreholes or hand-dug well for sustenance. Unfortunately, the water sources are rarely treated or untreated, exposed, or handled unhygienically. This is the scenario in areas, where they depend on a single source for daily living. In addition, some groundwaters are found near contaminated environments, including, municipal dumpsites, sewerage systems, amongst others, all of which pose health risks to human beings.
Reports on microbial assessments of groundwater used by the inhabitants of Osogbo have been documented (Olowe et al. 2005; Owolabi et al. 2014), with emphasis on presumptive identification of microorganisms in the water. To our knowledge, the antimicrobial susceptibility of PCR-confirmed E. coli from boreholes and hand-dug wells in Osogbo, Osun State Nigeria has not been investigated, let alone the detection of resistance determinants. The current study, therefore, elucidated the drug resistance pattern and ampicillin resistant genes in PCR-confirmed E. coli obtained from selected boreholes and hand-dug wells within Osogbo metropolis, Osun State, Nigeria.
Methodology
Sample collection
Forty water samples, comprising 20 each of boreholes and wells, were collected across Osogbo metropolis, Nigeria over a 3-month sampling period (September–November 2020). The geographic coordinates of the sampling locations were documented (Fig. 1). Water samples from borehole outlets were collected into a germ-free plastic of 1 L capacity. With the aid of a clean rope, sterilized plastic bottles were suspended into hand-dug wells, pulled up when filled and immediately covered. All samples were transported to the laboratory on ice and analysed within 6 h.
Isolation of Escherichia coli from the groundwater samples
Pour plate technique was employed for isolation of E. coli. Aliquot of water sample (1 ml) was dispensed into sterile Petri dishes in duplicates. To the Petri dishes, 20 ml of eosin methylene blue agar was dispensed, mixed gently, allowed to gel, and incubated at 37 °C for 18–24 h. Distinctive metallic-sheen colonies observed were enumerated as appropriate. The colonies were purified by streaking on nutrient agar plates until pure cultures were obtained and stored on agar slants until when needed.
Molecular identification of the E. coli isolates
DNA extraction
Genomic DNA extraction of the E. coli isolates was done by boiling method (Maugeri et al. 2004; Torres et al. 2005). Briefly, distinct 18–24 h old E. coli colonies on nutrient agar plates were aseptically picked and resuspended in 200 µl sterile distilled water, boiled at 100 °C for 10 min, and centrifuged at 12,000 rpm for 10 min. The supernatant containing DNA was carefully removed with a sterile pipette into germ-free Eppendorf tubes and kept at − 80 °C for PCR amplification.
PCR-based confirmation of E. coli isolates
The uidA gene (F: AAAACGGCAAGAAAAAGCAG; R: ACGCGTGGTTAACAGTCTTGCG) of the 55 E. coli DNA extracts was amplified using thermocycler. Briefly, PCR mixture (25 µl) consisting of 12.5 µl of PCR master mix (Thermo Scientific), 0.5 µl each of primer (Inqaba Biotech, SA), 5 µl of template DNA, and 6.5 µl of PCR grade water was amplified using SimpliAmp Thermal Cycler (Applied Biosystems, Life Technologies, Singapore). Controls, both positive and negative were also included. Cycling condition, i.e., initial denaturation at 94 °C for 5 min followed by 35 cycles of 30 s denaturation at 95 °C; annealing at 58 °C for 1 min; extension at 72 °C for 1 min, and a final extension step for 5 min at 72 °C, as previously described (Titilawo et al. 2015a).
Gel electrophoresis
Amplicons (5 µl) were loaded in a gel electrophoresis tank containing 1% SeaKem LE agarose gel (Lonza, USA) dissolved in 1X TAE electrophoresis buffer-stained ethidium bromide (Sigma-Aldrich, USA), and run at 100 V for 1 h. Viewing was done using the Bio-Rad ChemDoc (Hercules, USA).
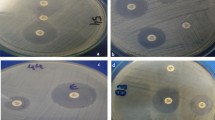
Antimicrobial susceptibility testing
Disc diffusion assay (Kirby-Bauer et al. 1966) was employed in estimating the susceptibility pattern of the isolates to ten commercially available antibiotic discs, belonging to seven classes, i.e., aminoglycosides–amikacin (30 μg), streptomycin (10 μg), carbapenem–ertapenem (10 μg), sulfonamides–trimethoprim (5 μg), phenicols–chloramphenicol (30 μg); tetracyclines–tetracycline (10 μg), penicillins–ampicillin (10 µg), tetracyclines–doxycycline (30 µg), cephalosporins–ceftazidime (30 µg), quinolones–ofloxacin (5 µg). Escherichia coli isolates (18–24 h) were suspended into physiological saline. Exactly 100 µl of 0.5 McFarland standard bacteria solution was inoculated and spread on Muller Hinton agar plates and allowed to dry. Antibiotic discs were gently placed on the plates using a disc dispenser (Oxoid), and incubated at 37 °C for 24 h. The diameter of zone of inhibition was measured and recorded in the nearest millimetre. Isolates were considered as susceptible (S), intermediate (I) and resistant (R) to test drugs based on the diameter of inhibition interpretation guideline of the Clinical and Laboratory Standards Institute (CLSI 2019). The frequency of antibiotic-resistant isolates was calculated using the equation: (X/Y) × 100. Where ‘X’ is the sum of isolates resistant to a drug and ‘Y’ is the sum of isolates from the sample.
Multiple antibiotic resistant phenotypes and indexing of the isolates
Multiple antibiotic-resistant phenotypes (MARPs) for each sampling site were generated for isolates exhibiting resistance to three or more antibiotics according to Wose et al. (2010). Multiple antibiotic resistance indexes (MARI) for each location were also estimated from the expression:
where ‘x’ represents the number of antimicrobial agents to which the isolate was resistant and ‘y’ is the total sum of antimicrobial agent tested against one isolate (Blasco et al. 2008; Titilawo et al. 2015a).
Also, the antibiotic resistance index (ARI) for each sampling location was evaluated using the formula:
where ‘A’ is the total number of resistant isolates recorded, ‘N’ is the number of isolates and ‘Y’ represents the total number of antibiotics tested (Tandra and Sudha 2014; Titilawo et al. 2015b).
Detection of ampicillin resistance determinants
PCR amplification and gel electrophoresis procedures (described above) were employed for the identification of resistant determinants among E. coli resistant to ampicillin. The details of primers used are shown in Table 1.
Results
Escherichia coli counts from the groundwater samples
A total of 55 distinct colonies showing characteristic greenish metallic sheen on EMB agar and presumptively identified as E. coli were obtained from the groundwater samples (Table 2). Borehole water had a higher mean count (35) from 10 samples compared to well water with 20 colonies recovered from 7 samples. Interestingly, not one E. coli was recovered from 10 and 13 borehole and well water samples, respectively (Table 2).
Molecular confirmation of the recovered E. coli isolates
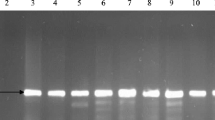
All the 55 phenotypically identified E. coli isolates yielded 147 bp on gel electrophoresis after PCR amplification of the uidA gene. The gel electrophoresis profile of representative PCR-confirmed E. coli is shown in Fig. 2.
Antibiotic resistance patterns of the E. coli isolates
Resistance of the isolates to the different test antibiotics ranged from 20% to 91%. Exactly 50 (91%) of the E. coli showed resistance to ampicillin, followed by trimethoprim and ertapenem 43 (78%), doxycycline 40 (73%), ceftazidime 38 (69%), tetracycline 37 (67%), streptomycin 22 (40%) and amikacin 18 (33%) (Fig. 3).
Susceptibilities of the isolates were in the order: ofloxacin 43 (78%), chloramphenicol 34 (62%), amikacin 33 (60%), streptomycin 25 (45%) and tetracycline 16 (29%), trimethoprim 12 (22%), ceftazidime and ertapenem 9 (16%), doxycycline 8 (15%) and ampicillin 5 (9%), respectively (Fig. 3).
Multiple antibiotic resistance phenotypes (MARPs) of the E. coli isolates
The MARPs generated for the E. coli isolates are presented in Table 3. Only 1 of the 55 E. coli isolates was resistant to 2 drugs (AK-AMP), while the remaining 54 were multi-resistant, ranging from 4 to 9 drugs (Table 3). None was resistant to the 10 drugs involved in the study. When this was expressed in terms of frequency, 2%, 11%, 13%, 21%, 23%, and 30% of the isolates exhibited multiple antibiotic resistance to 9, 5, 8, 7, 4, and 6 antimicrobials, respectively. The highest MARP (9) obtained across the sampling sites was AMP–CAZ–ETP–S–AK–DO–TE–W–C (Table 3). In the same vein, antibiotic resistance index (ARI) ranged from 0.08 to 0.80, whereas multiple antibiotic resistance index (MARI) was from 0.4 to 0.9 (Table 4).
Detection of ampicillin-resistant genes
All ampicillin-resistant E. coli (50) were examined for Ctx-M, SHV, ampC, and TEM-H resistance genes. Surprisingly, only 24 (48%) of the phenotypic ampicillin-resistant E. coli isolates harboured TEM-H (516 bp) and other genes were not detected. Figure 4 shows the representative gel picture of TEM-H amplification.
Discussion
Unsafe water is implicated in several waterborne illnesses (Nanfack et al. 2014), with contaminated groundwater fast becoming an emerging public health concern (Li et al. 2021). The current study elucidated the antibiotic resistance profile and ampicillin-resistant gene in PCR-confirmed E. coli isolates from selected boreholes and hand-dug wells within Osogbo metropolis, Osun State, Nigeria.
Escherichia coli is a specific indicator for faecal water pollution, and the presence of other waterborne pathogens (Takal and Quaye-Ballard 2018). In this study, E. coli was detected in 17 groundwater sources suggesting contamination with faecal material. More E. coli was recovered from borehole (35) compared to well water (20) (Table 2). This signifies the extent of pollution relating either to natural and/or anthropogenic activities around the water sources. The presumptive 55 E. coli isolates were uidA positive after PCR amplification. Earlier investigations also isolated E. coli from groundwater sources in India (Sharma et al. 2017), Indonesia (Dayanti et al. 2018), Nigeria (Titilawo et al. 2020), and Ghana (Takal and Quaye-Ballard 2018). Diseases such as diarrhoea, cholera, dysentery, typhoid, polio, and deaths have been associated with consumption of contaminated (WHO 2021). According to World Health Organization, water for drinking purposes should be devoid of E. coli (WHO 2011).
In the current investigation, the incidence of E. coli in some borehole and well water is not surprising as the water sources were unhygienically located and/or handled by users. Groundwater contamination occurs through seepage, fractured rock, holes and cracks, macropores root systems, and animal burrows (Romijn 2002; Mokuolu et al. 2017). Some of the water sources were sited close to contaminated flowing river, gutters, toilets facilities/sewerage systems, dumpsite, rarely or not treated, since construction and some of the wells were uncovered. The behaviour of users also determines the level of contamination of drinking water sources (Abdellah et al. 2012). During this study, some of the well water dependents were seen using dirty drawers to collect water, especially the shallow ones. This is unsafe and could transfer microbes on the bucket surface into the water.
The attainment of sustainable development goals (SDGs) associated with well-being, food security, safe water, and cleanliness is facing severe opposition by the widely acknowledged resistance to antimicrobial agents (Interagency Coordination Group on Antimicrobial Resistance 2019). This work reveals that resistance of our E. coli isolates to ampicillin was 50 (91%), trimethoprim and ertapenem 43 (78%), doxycycline 40 (73%), ceftazidime 38 (69%), tetracycline 37 (67%), streptomycin 22 (40%) and amikacin 18 (33%). The highest prevalence of ampicillin-resistant E. coli observed in this study agrees with Aslan et al. (2018), Praveenkumarreddy et al. (2020), and Singh et al. (2020). This is particularly worrisome, because ampicillin is the choice drug for treating livestock and human infections due to its effectiveness and low toxicity (FAO 2016; Pacifici 2021). Resistance to this drug, therefore, has possible consequences on increasing infections and deaths in humans and animals (Monger et al., 2021). Other researchers also reported high resistances to trimethoprim, ertapenem, doxycycline, ceftazidime, tetracycline, and streptomycin in aquatic environments, clinical and veterinary samples (Brolund et al. 2010; Hyle et al. 2010; Tadesse et al. 2012; Olorunmola et al. 2013; Mulder et al. 2019; Jahantigh et al. 2020; Mapanguy et al. 2021).
High-level susceptibility of the isolates to ofloxacin, chloramphenicol, and amikacin may be due to decreased exposure arising from low use of the antibiotics. In addition, the intravenous route of administration of ofloxacin and amikacin possibly restricts their indiscriminate use (Cheesbrough 2000). Significantly reduced resistances to fluoroquinolone and aminoglycoside antibiotics have been documented elsewhere (Titilawo et al. 2015a; Singh et al. 2020) Hence, the antimicrobials can be effective for treating E. coli infections.
In this work, antibiogram presented 100% resistance to ≥ 2 antimicrobials (Table 3). This finding is alarming and a serious health concern. Multiple drug resistance is a signal of abuse or excessive use of an antimicrobials for treatment of clinical and veterinary bacterial infections (Ramesh et al. 2010; Titilawo et al. 2015a). Previous investigation reported 75% multi-resistant E. coli from groundwater samples in India (Sharma et al. 2017), 46.9% from drinking water sources in Tanzania (Lyimo et al. 2016), and 100%, 90%, and 83% in soil, irrigated vegetable and wastewater, respectively, in Nsukka, Southeast Nigeria (Chigor et al. 2020). Olowe et al. (2008) also noted > 90% drug-resistant E. coli isolates to three or more commonly used antibiotics from clinical samples in Osogbo, Southwest Nigeria.
Multidrug-resistant Escherichia coli is a major threat to sustainable healthy living and livestock management. Altogether, the MDR E. coli in our investigation were not sensitive to a range of four to nine drugs, and mostly to ampicillin, ceftazidime, ertapenem, doxycycline and tetracycline. Previous finding of Lateef et al. (2005) revealed that a significant number of E. coli recovered from hospital, edible, and effluent samples in Southwest Nigeria were also not susceptible to different antibiotics, ranging between two and seven drugs, and mostly cotrimoxazole, tetracycline, and amoxicillin. Likewise, Titilawo et al. (2015a) noted E. coli resistance to five to nine drugs, majorly sulphamethoxazole, gentamycin, amoxycillin and ampicillin. Resistance of our isolates to ceftazidime and ertapenem is a public health concern as the drugs are known to be a viable therapeutic option to treat E. coli infections due to their effectiveness and low toxicity profile (Lartigue et al. 2007; Yuan et al. 2016). Ampicillin and tetracycline are commonly implicated in the therapy of airway and gastrointestinal infections, mastitis, etc. in humans and animals (Okeke et al. 1995; Hart and Ariuki 1998; Hidron et al. 2008; Willems and Schaik 2009; Zhang et al. 2012). The increased ineffectiveness of tetracyclines in the current work can be attributed to the availability and accessibility of the drug for self-prescription and medication in both humans and livestock farms in Nigeria (Chigor et al. 2010; Olatoye 2010).
Usually, drug-resistant bacteria spread to the environment via excreta, and water environments are recognised reservoirs and transmission paths for the spreading of antibiotic resistance (Karkman et al. 2018). In the natural environment, antimicrobials provide selective pressure which alters the behaviour and fitness of a bacterial, advancing antibiotic resistance (Amarasiri et al. 2020). Oftentimes, environments harbouring antibiotic-resistant bacteria signal the possibility of antimicrobial contamination of that area (Gunaseelan and Ruban 2011).
Location of the boreholes close to channels of polluted waterbody earlier reported to harbour MDR bacterial (Titilawo et al. 2020), toilet facilities, dumpsites, and heavy anthropogenic activities around the groundwaters may be the source of contamination with MDR E. coli. In addition, free-range animals and their wastes also sighted around W5 during sampling could be a source of contamination. Animals are often administered antibiotics during rearing and their faeces contains relatively high levels of antimicrobial resistant bacteria (He et al. 2020). All these possibly influenced a higher than the safe limit for MARI (0.2) as observed in this study. MARI between 0.4 and 0.9 signifies high-risk origin of the isolates, where antimicrobials are frequently employed for treatment. Other studies reported MARI value greater than 0.2 in surface waters of Osun state Nigeria (Titilawo et al. 2015a), water purification and supply lines in North–West Province of South Africa (Ateba et al. 2020), raw meats, ready-to-eat meat, and their related sample in Ghana (Adzitey et al. 2021).
Intrinsic mechanism of Enterobacteriaceae to confer resistance to beta-lactam antibiotics through inactivation of beta-lactam ring present a huge challenge to public health concern (Bush and Jacoby 2010; Bush 2018; Matloko et al. 2021). The detection and prevalence (24; 48%) of TEM-H gene in the E. coli suggests that the gene is responsible for the phenotypic resistance noticed in the isolates. Nonetheless, the null detection of ampC, CTX-M, and SHV indicates that the isolates did not habour the genes or they are plasmid-encoded in Assawatheptawee et al. (2017) and Titilawo et al. (2015a) detected ampC gene in E. coli from aquatic environments in Northern Thailand and Southwestern Nigeria, respectively. Generally in E. coli, ampC gene encoded in the chromosome is constitutively low or poorly expressed but non-inducible, and same determinants located on mobile genetic plasmid favours overproduction of beta-lactamases and are easily transmitted within and between different bacterial species and hosts (Hanson and Sanders 1999; Haenni et al. 2014).
Conclusion
The quality of groundwater, an important resource for drinking in Osogbo metropolis is currently threatened by contamination with multidrug-resistant microorganisms. The study investigated the prevalence of multidrug-resistant E. coli from 40 groundwater sources (20 each of borehole and well water), and detected ampicillin-resistant gene in phenotypic resistant isolates. A total of 17 groundwater samples were contaminated by E. coli, and borehole waters recorded higher counts compared to well water. This suggests unhygienic location and poor sanitary practices around the water sources. Interestingly, all the 55 (100%) isolates were confirmed E. coli using uidA gene and showed resistance to more than one antimicrobial, the most resistant being to ampicillin 53 (91%). Multiple antibiotic-resistant phenotypes and indices estimated indicate elevated levels of drug resistant E. coli in the groundwater samples and their occurrence can jeopardize human health especially in households and neighbourhoods that depend on the waters to meet their daily needs. The detection of TEM-H gene in the isolates poses a huge threat to public health, because the resistant gene is easily transferred horizontally within and between bacterial species and hosts. Thus, there is a need to regularly treat the groundwaters, prior consumption to reduce the risk of E. coli infection. Proper location and construction, and hygienic practices around groundwaters should be encouraged. The challenge of antibiotic resistance observed herein can be reduced by creating more awareness together with enforcement of law and policy that restrict illicit prescription and dispensing of antimicrobials. In addition, well-planned inspection programmes targeted at monitoring groundwater associated antimicrobial resistance patterns are essential. Further study on microbial water quality and profiling of antibiotic resistance aimed at mapping various waterborne diseases in Osogbo metropolis, Nigeria is recommended.
Data availability
The data supporting the findings of this study are available within the article.
References
Abdellah AM, Abdel-Maid HM, Yahia NA (2012) Assessment of drinking water microbial contamination in Al-Butana region of Sudan. J Appl Sci 12(9):856–862
Adzitey F, Huda N, Shariff AHM (2021) Phenotypic antimicrobial susceptibility of Escherichia coli from raw meats, ready-to-eat meats, and their related samples in one health context. Microorganisms 9:326
Alanazi MQ, Alqahtani FY, Aleanizy FS (2018) An evaluation of E. coli in urinary tract infection in emergency department at KAMC in Riyadh, Saudi Arabia: retrospective study. Ann Clin Microbiol Antimicrob 17:3
Alsalme A, Al-Zaqri N, Ullah R, Yaqub S (2021) Approximation of ground water quality for microbial and chemical contamination. Saudi J Biol Sci 28:1757–1762
Amarasiri M, Sano D, Suzuk S (2020) Understanding human health risks caused by antibiotic resistant bacteria (ARB) and antibiotic resistance genes (ARG) in water environments: current knowledge and questions to be answered. Crit Rev Environ Sci Technol 50(19):2016–2059
Amaya E, Reyes D, Paniagua M, Calderón S, Rashid M-U, Colque P, Kühn I, Möllby R, Weintraub A, Nord CE (2012) Antibiotic resistance patterns of Escherichia coli isolates from different aquatic environmental sources in Leon. Nicaragua Clin Microbiol Infect 18(9):E347–E354
Anderson Y, Bohan P (2001) Disease surveillance and water borne outbreaks. In: Fewtrell L, Bartram J (eds) Water quality. Guidelines, standards and health. IWA Publishing, London, pp 11–133
Aslan A, Cole Z, Bhattacharya A, Oyibo O (2018) Presence of antibiotic-resistant Escherichia coli in wastewater treatment plant effluents utilized as water reuse for irrigation. Water 10:805
Assawatheptawee K, Tansawai U, Kiddee A, Thongngen P, Punyadi P, Romgae T, Kongthai P, Sumpradit T, Niumsup PR (2017) Occurrence of extended-Spectrum and AmpC-Type β-Lactamase genes in Escherichia coli isolated from water environments in Northern Thailand. Microbes Environ 32(3):293–296
Ateba CN, Tabi NM, Fri J, Ebob M, Bissong A, Bezuidenhout CC (2020) Occurrence of antibiotic-resistant bacteria and genes in two drinking water treatment and distribution systems in the North-West province of South Africa. Antibiotics 9:745
Aydin D (2007) The microbiological and physico-chemical quality of groundwater in West Thrace. Turkey Polish J Environ Stud 16(3):377–383
Blasco MD, Esteve C, Alcaide E (2008) Multi-resistant waterborne pathogens isolated from water reservoirs and cooling systems. J Appl Microbiol 105:469–475
Brolund A, Sundqvist M, Kahlmeter G, Grape M (2010) Molecular characterization of trimethoprim resistance in Escherichia coli and Klebsiella pneumoniae during a two-year intervention on trimethoprim use. PLoS ONE 5(2):e9233
Bush K (2018) Past and present perspectives on β-lactamases. Antimicrob Agents Chemother 62(10):e01076-e1118
Bush K, Jacoby GA (2010) Updated functional classification of β-lactamases. Antimicrob Agents Chemother 54(3):969–976
Cheesbrough M (2000) District laboratory practice in tropical countries, 2nd edn. Cambridge University Press, Cambridge
Chigor VN, Umoh JV, Smith IS, Igbinosa OE, Okoh IA (2010) Multidrug resistance and plasmid patterns of Escherichia coli O157 and other E. coli isolated from diarrhoeal stools and surface waters from some selected sources in Zaria, Nigeria. Int J Environ Res Public Health 7:3831–3841
Chigor V, Ibangha IA, Chigor C, Titilawo Y (2020) Treated wastewater used in fresh produce irrigation in Nsukka, Southeast Nigeria is a reservoir of enterotoxigenic and multidrug-resistant Escherichia coli. Heliyon 6(4):e03780
Chissaque A, de Deus N, Vubil D, Mandomando I (2018) The epidemiology of diarrhea in children under 5 years of age in Mozambique. Curr Trop Med Rep 5:115–124
CLSI (2019) Performance standard for antimicrobial disk susceptibility test MO12-A12. Clinical and Laboratory Standard Institute, Wayne
Colom K, Perez J, Alonso R, Fernandez-Aranguiz A, Larino E, Cisterna R (2003) Simple and reliable multiplex PCR assay for detection of blaTEM, blaSHV and blaOXA - 1 genes in Enterobacteriaceae. FEMS Microbiol Lett 223:147–151
Dadgostar P (2019) Antimicrobial resistance: implications and costs. Infect Drug Resist 12:3903–3910
Dayanti MP, Fachrul MF, Wijayanti A (2018) Escherichia coli as bioindicator of the groundwater quality in Palmerah District, West Jakarta, Indonesia. IOP Conf Ser: Earth Environ Sci 106:012081
Edokpayi JN, Odiyo JO, Popoola EO, Msagati TM (2018a) Evaluation of microbiological and physicochemical parameters of alternative source of drinking water. A case study of Nzhelele River, South Africa. Open Microbiol J 12:18–27
Edokpayi JN, Rogawski ET, Kahler DM, Hill CL, Reynolds C, Nyathi E, Smith JA, Odiyo JO, Samie A, Bessong P (2018b) Challenges to sustainable safe drinking water: a case study of water quality and use across seasons in rural communities in Limpopo Province, South Africa. Water 10:159
FAO (2016) Drivers, dynamics and epidemiology of antimicrobial resistance in animal production. Food and Agriculture Organisation of the United Nations, Rome
Gunaseelan C, Ruban P (2011) Antibiotic resistance of bacteria from Krishna Godavari Basin, Bay of Bengal, India. Environ Exp Biol 9:133–136
Haenni M, Châtre P, Métayer V, Bour M, Signol E, Madec JY, Gay E (2014) Comparative prevalence and characterization of ESBL-producing Enterobacteriaceae in dominant versus subdominant enteric flora in veal calves at slaughterhouse. France Vet Microbiol 171(3–4):321–327
Hanson ND, Sanders CC (1999) Regulation of inducible AmpC beta-lactamase expression among Enterobacteriaceae. Curr Pharm Des 5(11):881–894
Hart A, Ariuki KS (1998) Antimicrobial resistance in developing countries. Br Med J 317:647–650
He Y, Yuan Q, Mathieu J, Stadler L, Senehi N, Sun R, Alvarez PJJ (2020) Antibiotic resistance genes from livestock waste: occurrence, and treatment. NPJ Clean Water 3:4
Hidron AI, Edwards JR, Patel J, Horan TC, Sievert DM, Pollock DA, Fridkin SK (2008) Antimicrobial-resistant pathogens associated with healthcare-associated infections: annual summary of data reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2006–2007. Infect Control Hosp Epidemiol 29(11):996–1011
Hyle E, Ferraro M, Silver M, Lee H, Hooper D (2010) Ertapenem-resistant Enterobacteriaceae risk factors for acquisition and outcomes. Infect Control Hosp Epidemiol 31(12):1242–1249
Interagency Coordination Group on Antimicrobial Resistance (2019) No time to wait: securing the future from drug-resistant infections report to the Secretary-General of the United Nations. Assessed 20 Aug 2021
Jahantigh M, Samadi K, Dizaji RE, Salari S (2020) Antimicrobial resistance and prevalence of tetracycline resistance genes in Escherichia coli isolated from lesions of colibacillosis in broiler chickens in Sistan, Iran. BMC Vet Res 16:267
Kaper J, Nataro J, Mobley H (2004) Pathogenic Escherichia coli. Nat Rev Microbiol 2:123–140
Kapoor G, Saigal S, Elongavan A (2017) Action and resistance mechanisms of antibiotics: a guide for clinicians. J Anaesthesiol Clin Pharmacol 33:300–305
Karkman A, Do TT, Walsh F, Virta MPJ (2018) Antibiotic-resistance genes in wastewater. Trends Microbiol 26(3):220–228
Keesari T, Ramakumar KL, Prasad MBK, Chidambaram S, Perumal P, Prakash D, Nawani N (2015) Microbial evaluation of groundwater and its implications on redox condition of a multi-layer sedimentary aquifer system. Environ Process 2:331–346
Kirby-Bauer WM, Sherris JC, Turck M (1966) Antibiotic susceptibility testing by single disc method. Am J Clin Pathol 45:4
Lartigue MF, Poirel L, Poyart C, Réglier-Poupet H, Nordmann P (2007) Ertapenem resistance of Escherichia coli. Emerg Infect Dis 13(2):315–317
Lateef AJ, Oloke K, Gueguimkana EB (2005) The prevalence of bacterial resistance in clinical, food, water and some environmental samples in Southwest Nigeria. Environ Monit Assess 100:59–69
Li P, Karunanidhi D, Subramani T, Srinivasamoorthy K (2021) Sources and consequences of groundwater contamination. Arch Environ Contamin Toxicol 80:1–10
Liu L, Oza S, Hogan D, Chu Y, Perin J, Zhu J, Lawn JE, Cousens S, Mathers C, Black RE (2016) Global, regional, and national causes of under-5 mortality in 2000–15: an updated systematic analysis with implications for the sustainable development goals. Lancet 388:3027–3035
Llopis-GonzálezA SAL, Requena PM, Suárez-Varela MM (2014) Assessment of the microbiological quality of groundwater in three regions of the Valencian community (Spain). Int J Environ Res Public Health 11:5527–5540
Lyimo B, Buza J, Subbiah M, Smith W, Call DR (2016) Comparison of antibiotic resistant Escherichia coli obtained from drinking water sources in northern Tanzania: a cross-sectional study. BMC Microbiol 16:254
Mapanguy CA, Adedoja A, Kecke LGV, Vouvoungui JC, Nguimbi E, Velavan TP, Ntounmi F (2021) High prevalence of antibiotics-resistant Escherichia coli in Congolese students. Int J Infect Dis 103:119–123
Matloko K, Fri J, Ateba TP, Molale-Tom LG, Ateba CN (2021) Evidence of potentially unrelated AmpC beta-lactamase producing Enterobacteriaceae from cattle, cattle products and hospital environments commonly harboring the blaACC resistance determinant. PLoS ONE 16(7):e0253647
Maugeri TL, Carbone M, Fera MT, Irrera GP, Gugliandolo C (2004) Distribution of potentially pathogenic bacteria as free living and plankton associated in a marine coastal zone. J Appl Microbiol 97:354–361
Medina E, Pieper DH (2016) Tackling threats and future problems of multidrug-resistant bacteria. Curr Top Microbiol Immunol 398:3–33
Mokuolu OA, Jacob SO, Ayanshola AM (2017) Groundwater quality assessment near a Nigerian dumpsite. Ethiopian J Environ Stud Manag 10(5):588–596
Monger XC, Gilbert A-A, Saucier L, Vincent AT (2021) Antibiotics resistance: from pig to meat. Anitibiotics 10:1209
Mulder M, Verbon A, Lous J, Goessens W, Stricker BH (2019) Use of other antimicrobial drugs is associated with trimethoprim resistance in patients with urinary tract infections caused by E. coli. Eur J Clin Microbiol Infect Dis 38:2283–2290
Nanfack NAC, Fonteh FA, Payne VK, Katte B, Fogoh JM (2014) Eaux non conventionnelles: un risqueouune solution aux problems d’eau pour les classes pauvres. Larhyss J 17:47–64
Odiyo JO, Makungo R (2018) Chemical and microbial quality of groundwater in Siloam village, implication to human health and sources of contamination. Int J Environ Res Public Health 15:317
Okeke N, Lamikana A, Edelman R (1995) Socioeconomic and behavioural factors leading to acquired bacterial resistance in developing countries. Emerg Infect Dis 5:18–27
Okeke IN, Lamikanra A, Edelman R (1999) Socioeconomic and behavioural factors leading to acquired bacterial resistance to antibiotics in developing countries. Emerg Infect Dis 5:18–27
Olaitan JO, Akinde SB, Salami AO, Akinyode OA (2013) Quality surveillance of surface water catchments in selected Obokun rural communities, in South-Western Nigeria. Afr J Microbiol Res 7(36):4491–4500
Olasoji SO, Oyewole NO, Abiola B, Edokpayi JN (2019) Water quality assessment of surface and groundwater sources using a water quality index method. A case study of a peri-urban town in Southwest Nigeria. Environments 6:23
Olatoye IO (2010) The incidence and antibiotics susceptibility of Escherichia coli O157:H7 from beef in Ibadan Municipal, Nigeria. Afr J Biotechnol 9(8):1196–1199
Olorunmola FO, Kolawole DO, Lamikanra A (2013) Antibiotic resistance and virulence properties in Escherichia coli strains from cases of urinary tract infections. Afr J Infect Dis 7(1):1–7
Olowe OA, Ojurongbe O, Opaleye OO, Adedosu OT, Olowe RA, Eniola KIT (2005) Bacteriological quality of water samples in Osogbo metropolis. Afr J Clin Exp Microbiol 6(3):219–222
Olowe OA, Okanlawon BM, Olowe RA, Olayemi AB (2008) Antimicrobial resistant pattern of Escherichia coli from human clinical samples in Osogbo, South-Western Nigeria. Afr J Microbiol Res 2(1):8–11
Onyango AE, Okoth MW, Kunyanga CN, Aliwa BO (2018) Microbiological quality and contamination level of water sources in Isiolo County in Kenya. J Environ Public Health. https://doi.org/10.1155/2018/2139867
Owolabi JB, Olaitan JO, Alao AA, Deile AK, Ige OO, Oloke JK (2014) Bacteriological and physicochemical assessment of water from student hostels of Osun State University, Main Campus, Osogbo, Southwest Nigeria. Covenant J Phys Life Sci 2(1):1–15
Pacifici GM (2021) Clinical pharmacology of ampicillin in infants and children. J Drug Des Res 8(2):1082
Paruch AM, Mæhlum T, Robertson L (2015) Changes in microbial quality of irrigation water under different weather conditions in Southeast Norway. Environ Process 2(1):115–124
Potgieter N, Mudau LS, Maluleke FRS (2006) Microbiological quality of groundwater sources used by rural communities in Limpopo province, South Africa. Water Sci Technol 54(11–12):371–377
Praveenkumarreddy Y, Akiba M, Guruge KS, Balakrishna K, Vandana KE, Kumar V (2020) Occurrence of antimicrobial-resistant Escherichia coli in sewage treatment plants of South India. J Water Sanit Hyg Dev 10(1):48–55
Ramesh S, Manivasagan P, Ashokkumar S, Rajaram G, Mayavu P (2010) Plasmid profiling and multiple antibiotic resistances of heterotrophic bacteria isolated from Muthupettai Mangrove Environment, Southeast Coast of India. Curr Res Bacteriol 3:227–237
Romijn E (2002) Groundwater quality and contamination, Groundwater contamination—a methodological guide. In: Zaporozec A (ed) IHP-VI, series on groundwater no. 2. UNESCO, London, pp 17–22
Saladin M, Cao VTB, Lambert T, Donay JL, Herrmann JL, Ould-Hocine Z, Verdet C, Delisle F, Philippon A, Arlet G (2002) Diversity of CTX-M β-lactamases and their promoter regions from Enterobacteriaceae isolated in three Parisian hospitals. FEMS Microbiol Lett 209:161–801
Sharma B, Parul VAK, Jain U, Yadav JK, Singh R, Mishra R (2017) Occurrence of multidrug resistant Escherichia coli in groundwater of Brij region (UP) and its public health implications. Vet World 10(3):293–301
Singh AK, Das S, Kumar S, Gajamer VR, Najar IN, Lepcha YD, Tiwari HK, Singh S (2020) Distribution of antibiotic-resistant Enterobacteriaceae pathogens in potable spring water of Eastern Indian Himalayas: emphasis on virulence gene and antibiotic resistance genes in Escherichia coli. Front Microbiol 11:581072
Stein H, Kellermann C, Schmidt SI, Brielmann H, Steube C, Berkhoff SE, Fuchs A, Hahn HJ, Thulind B, Griebler C (2010) The potential use of fauna and bacteria as ecological indicators for the assessment of groundwater quality. J Environ Monit 12:242–254
Tadesse DA, Zhao S, Tong E, Ayers S, Singh A, Bartholomew MJ, McDermott PF (2012) Antimicrobial drug resistance in Escherichia coli from humans and food animals, United States, 1950–2002. Emerg Infect Dis 18(5):741–749
Takal JK, Quaye-Ballard JA (2018) Bacteriological contamination of groundwater in relation to septic tanks location in Ashanti Region, Ghana. Cogent Environ Sci 4(1):1556197
Tandra M, Sudha G (2014) Prevalence of antibiotic-resistant bacteria in three different aquatic environments over three seasons. Environ Monit Assess 186:5089–5100
Titilawo Y, Obi L, Okoh A (2015a) Antimicrobial resistance determinants of Escherichia coli Isolates recovered from some rivers in Osun State, Southwestern Nigeria: implications for public health. Sci Total Environ 523:82–94
Titilawo Y, Sibanda T, Obi L, Okoh A (2015b) Multiple antibiotic resistance indexing of Escherichia coli to identify high-risk sources of faecal contamination of water. Environ Sci Pollut Res 22:10969–10980
Titilawo Y, Nwuakpa F, Bankole S, Nworie O, Okoro C, Titilawo M, Olaitan J (2020) Quality audit of drinking water sources in Ikwo rural setting of Ebonyi Southeastern Nigeria. Int J Energy Water Resour 4(3):321–334
Torres AG, Zhou X, Kaper JB (2005) Adherence of diarrheagenic Escherichia coli strains to epithelial cells. Infect Immun 73:18–29
United Nations (UN) (2018) Sustainable development goal 6. Synthesis report on water and sanitation United Nations New York, New York 10017, United States of America, p 195. Assessed 10 Aug 2021
USGS (2018) Bacteria and E. coli in water. https://www.usgs.gov/special-topic/water-science-school/science/bacteria-and-e-coli-water?qt-science_center_objects=0#qt-science_center_objects. Assessed Aug 2021
Vignesh S, Dahms HU, Kumarasamy P, Rajendran A, Kim BR, James RA (2015) Microbial effects on geochemical parameters in a tropical river basin. Environ Process 2:125–144
WHO (2011) Guidelines for drinking water quality (4th ed.). Geneva: WHO. https://apps.who.int/iris/bitstream/handle/10665/44584/9789241548151_eng.pdf;jsessionid=FF5283355E789441877C34680F82E433?sequence=1. Assessed 7 Sept 2021
WHO (2021) Drinking-water. https://www.who.int/news-room/fact-sheets/detail/drinking-water2019. Assessed 17 Nov 2021
Willems RJ, Schaik W (2009) Transition of Enterococcus faecium from commensal organism to nosocomial pathogen. Future Microbiol 4(9):1125–1135
Wose KCN, Ateba N, Kawadza TD (2010) Antibiotic resistance profiles of Escherichia coli isolated from different water sources in the Mmabatho locality, North-West Province, South Africa. Res Lett 106(1–2):44–49
Yuan XY, Yu DY, Qu XH, Xiao XQ, Bi B, Sun SB, Chang AY, Zhang QB (2016) Increased resistance rate to ceftazidime among blood culture isolates of ESBL-producing Escherichia coli in a university-affiliated hospital of China. J Antibiot 69:169–172
Zhang X, Paganelli FL, Bierschenk D, Kuipers A, Bonten MJM, Willems RJL, Schaik WV (2012) Genome-wide identification of ampicillin resistance determinants in Enterococcus faecium. PLoS Genet 8(6):e1002804
Zhu YG, Johnson TA, Su JQ, Qiao M, Guo GX, Stedtfeld RD, Hashsham SA, Tiedje JM (2013) Diverse and abundant antibiotic resistance genes in Chinese swine farms. Proc Natl Acad Sci 110:3435–3440
Acknowledgements
The authors appreciate individuals and communities who granted access to collect water samples.
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Titilawo, M.A., Fatoki, C.O., Titilawo, Y. et al. Assessment of multidrug-resistant phenotypes and detection of ampicillin-resistant determinants among Escherichia coli isolates of groundwater origin: case study—Osogbo, Southwest Nigeria. Sustain. Water Resour. Manag. 9, 8 (2023). https://doi.org/10.1007/s40899-022-00785-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s40899-022-00785-z