Abstract
Crop genetic resources are vital inputs in crop genetic improvement. In this study, genetic diversity, population structure, and similarity centers for 131 safflower accessions obtained from 28 countries were investigated using 12 ISSR markers. A sum of 201 ISSR bands were obtained among which 188 (93.844%) were found polymorphic. Mean polymorphism information content (0.448) and diversity parameters including mean effective number of alleles (1.655), mean Shannon’s information index (0.557), mean expected heterozygosity (0.354), and mean overall gene diversity (0.377) showed a good level of genetic diversity in the studied safflower materials. Model-based structure, unweighted pair-group method with arithmetic means, and principal coordinate analysis clustered all accessions into three main populations; A, B, and C and an unclassified population. Accessions originated from Asian countries like Pakistan and Israel were found most diverse. Three accessions, Pakistan-11, Israel-1, and Pakistan-26, were found most genetically distant and might be used as parental sources for genetic combinations in safflower breeding activities. Analysis of molecular variance revealed highly significant differentiation among the identified populations and population × country combinations. The results presented in this work most probably supported the hypothesis of seven similarity centers of safflower but need to be validated with further confirmed investigations. The information provided herein is expected to be helpful for the scientific community interested in safflower breeding.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
The fast growth of the world’s population requires sufficient availability of food in terms of calories and other nutrients (Long et al. 2015). Daily average of per capita available calories across the world was 2789 kcal in the year 2000, and it is believed that it will become 3130 kcal by year 2050 (FAO 2019), highlighting a steady increase in food demand paralleling the growth of world’s population. A steady increase in the production of safflower has been observed during the last two decades to mainly meet the vegetable oil shortage. However, according to FAO’s 2018–2027 predictions, the increase in oilseed production will amount 1.5% annually, implying a great drop relative to the last decade. Vegetable oil has one of the highest trade shares (41%) of production of all agricultural commodities. This share is expected to remain stable throughout the outlook period, with global vegetable oil exports reaching 96 Mt by 2027 (http://www.fao.org/docrep/i9166e/i9166e_Chapter4_Oilseeds.pdf). Scientists agree on the importance of increasing the crop yield instead of increasing area under the cultivation in the process of sustainable agriculture (Godfray et al. 2010). Therefore, in order to mitigate the vegetable oilseed shortage and avoid future demand problems, there is a need to focus on the breeding activities.
Safflower (Carthamus tinctorius L.) belongs to family Asteraceae and is self-pollinated having haploid genome size of about 1.4 GB with 2n = 24 chromosomes (Kumari et al. 2017). Safflower is a multipurpose crop cultivated on wide geographical zones all over the world for different purposes like dyes production, edible oil extraction, and several pharmaceutical utilizations (Weiss 2000; Ali et al. 2019a). Safflower oil production is lower compared to other oilseed crops, but still it is cultivated due to its potential of better adaptability to saline and drought conditions (Weiss 1983). Safflower has popularized due to its huge potential as biofuel crop in the recent years (Dordas and Sioulas 2009). Safflower has been used since pre-historic time, while archeological remains of Carthamus species were found 7500 years BC at sites of Syria (Marinova and Riehl 2009), indicating that safflower was distributed from these locations to connected geographies, i.e., Egypt, the Aegean, and southern Europe.
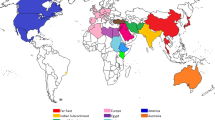
Safflower accessions revealing similarity based on various agro-morphological traits from different regions of the world are referred to as safflower similarity centers. Different researchers proposed various safflower similarity centers across the world, but still the number of actual similarity centers is ambiguous as ascertained by different molecular systems (Chapman et al. 2010). Seven safflower similarity centers were suggested by Knowles (1969) (1: Far East, 2: India-Pakistan, 3: Middle East, 4: Egypt, 5: Sudan, 6: Ethiopia, and 7: Europe). Similarly, ten similarity centers (1: Near East, 2: Iran/Afghanistan, 3: Turkey, 4: Egypt, 5: Ethiopia, 6: Sudan, 7: Far East, 8: India/Pakistan, 9: Europe, and 10: Kenya) were proposed by Ashri (1975); however, five similarity centers (1: Near East, 2: Iran and Afghanistan, Turkey, 3: Egypt, Ethiopia, (Sudan), 4: Far East, India/Pakistan, (Sudan), 5: Europe) were identified by Chapman et al. (2010). Still, this debate is ongoing and scientist did not come to single hypothesis about the similarity centers of safflower.
Safflower is known as an underutilized oilseed crop compared to other crops like soybean, rapeseed, and sunflower due to low oil content and seed yield as well as susceptibility to different diseases and insect pest attack limiting its quality and productivity (Zeinali 1999; Ali et al. 2019a). Current safflower local and traditional varieties comprise narrow genetic base; thus, assessment of the genetic diversity present in the safflower germplasm from different geographical zones will aid to provide valuable information necessary for the conservation and future utilization of safflower breeding programs (Yang et al. 2007). Safflower genetic diversity went to bottleneck during the process of domestication, which resulted in the significant decrease in its capability against the threatening environments, especially the environmental stresses (Yang et al. 2007; Mayerhofer et al. 2011). Genetic diversity serves as an effective tool and provides a rich source of variations (Baloch et al. 2017; Nadeem et al. 2018b). Diversity analyses in the germplasm possess great importance to plan an efficient and successful safflower breeding program (Mary and Gopalan 2006; Guliyev et al. 2018; Yildiz et al. 2019). Crop germplasm that consists of diverse desirable traits may be integrated in a better way during breeding programs in order to develop superior cultivars (Arystanbekkyzy et al. 2018; Yaldiz et al. 2018). Assessment of the crop genetic diversity is important for the germplasm conservation and also food security. Decreasing genetic diversity in the crop plants is one of the most important environmental concerns as outlined by the Food and Agriculture Organization (Castañeda-Álvarez et al. 2016). Limited information about genetic diversity within C. tinctorius and lack of efficient genomic tools has hampered the pace to improve economical traits during various safflower breeding programs. Need of the safflower genome molecular characterization and development of the efficient molecular markers has been recognized by a number of research groups (Amiri et al. 2001; Johnson et al. 2007).
Molecular markers overcome limitations present in the morphological and biochemical markers by detecting diversity at the DNA level (Erzurumlu et al. 2018; Nadeem et al. 2018a). It should be understood that different molecular markers have different characteristics, thus reflecting the different genetic diversity aspects (Talebi et al. 2012; Nadeem et al. 2018a). Safflower genetic diversity using several molecular markers like random amplified polymorphic DNA (RAPD), inter-simple sequence repeat (ISSR), amplified fragment length polymorphism (AFLP), simple sequence repeats (SSRs), iPBS-retrotransposon, and single nucleotide polymorphism (SNP) has been estimated (Johnson et al. 2007; Yang et al. 2007; Amini et al. 2008; Khan et al. 2009; Chapman et al. 2010; Lee et al. 2014; Pearl and Burke 2014; Ambreen et al. 2015, 2018; Kumar et al. 2015; Ali et al. 2019b). These researchers suggested the presence of good level of genetic diversity among different global safflower germplasm and also validate some of the similarity centers that were initially based on various morpho-agronomic traits (Johnson et al. 2007; Chapman et al. 2010; Pearl and Burke 2014; Kumar et al. 2015). Inter-simple sequence repeat (ISSR) primers based on di, tetra, or penta-nucleotide repeats are routinely used for various purposes (Zietkiewicz et al. 1994). The advantages of simple procedure, low cost, high reproducibility, and excellent stability made ISSR primers suitable for the determination of genetic diversity analysis (Rawat et al. 2016; Hadian et al. 2017; Ekincialp et al. 2019), mapping studies (Casaoli et al. 2001; Cekic et al. 2011; Tanyolac 2003), and germplasm identification (Nagaoka and Ogihara 1997; Potter et al. 2002). ISSR markers have been effectively employed for the determination of genetic diversity and molecular characterization of different crops including; cluster bean (Ansari et al. 2016), Chickpea (Gautam et al. 2016), and Brassica (Khalil and El zayat 2019). Besides these crops, ISSR markers were also used for the genetic diversity and molecular characterization of safflower by various researchers (Yang et al. 2007; Sabzalian et al. 2009; Golkar et al. 2011; Bagmohammadi et al. 2012; Majidi and Zadhoush 2014; Houmanat et al. 2016) revealing its efficient potential for the germplasm characterization. The current study was aimed to determine the genetic diversity, population structure, and safflower similarity centers at molecular level using ISSR markers, and the useful information will be then used for the future safflower breeding programs by the scientific community.
2 Materials and methods
Plant materials and DNA isolation
– One hundred and thirty-one safflower accessions collected from twenty-eight different countries were tested during the current study. The safflower accessions 94, 17, and 20 were provided by the United States Department of Agriculture (USDA), Plant Genetic Resources Institute (PGRI) Pakistan, and the Turkish Central Research Institute for Field Crops, respectively (Table 1). The safflower accessions provided by USDA (94 accessions) and PGRI (17 accessions) were landraces. The 20 Turkish safflower accessions were single plant selection among the international safflower germplasm obtained from the USDA and are candidate cultivars. Safflower seeds of each accession were sown at the research and experimental area of Bolu Abant Izzet Baysal University. The fresh, healthy, and young leaves from each accession were harvested for the DNA isolation and frozen at the temperature of − 80 °C in laboratory. Bulk of leaves from each accession was used for the DNA extraction following the CTAB protocol (Doyle and Doyle 1990) with slight modifications (Baloch et al. 2016). DNA concentration was estimated using agarose gel (0.8%) and was then confirmed with NanoDrop (DeNovix DS-11 FX, USA). Final DNA concentration of the 131 safflower accessions was adjusted to 5 ng μL−1 for the purpose of polymerase chain reactions. All samples were stored at the temperature of − 25 °C until PCR amplification.
ISSR PCR amplifications
– Ninety ISSR primers were initially screened using eight randomly selected accessions of safflower for PCR amplifications. Twelve ISSR primers were found most polymorphic by producing strong bands and were used for the amplification of PCR (Table 2). A total reaction volume of 25 µL for PCR amplifications were comprised of 4 ng ul−1 template DNA, 4 µL dNTPs (Thermo Scientific), 0.2 µL U Taq DNA polymerase (Thermo Scientific), 1 µL primer, 2.5 µL 1X PCR buffer (Thermo Scientific), 2 µL MgCl2, and 11.3 µL distilled water. Reactions were performed in the sequence of denaturation at 94 °C for 3 min, subsequently followed by 30 denaturation cycles at 94 °C for 1 min, annealing temperature of 48–54 °C for one minute depending upon the primer, and a final extension for 10 min at 72 °C. Agarose gel 1.8% (w/v) containing 0.5X TBE buffer was used for the electrophoreses of the amplified DNA fragments at a constant voltage of 120 V for 240 min. Ethidium bromide was used to perform the staining of the gel and then visualized using UV Imager Gel Doc XR + system (Bio-Rad, USA) light and photographed. A 100 bp + DNA ladder was used as a molecular weight marker.
Data analysis
– Strong, unambiguous, and clear bands were used for the purpose of scoring, while vague bands were not selected as they could not be easily detected. ISSR markers are dominant markers and scored in a binary matrix as 1 for present or 0 for absent, respectively, of all the bands with relative to 100 bp + DNA ladder. PopGene ver. 1.32 (Yeh et al. 2000) was applied to compute genetic diversity indices like effective alleles number (Ne), Shannon’s information index (I), and gene diversity (He) for individual ISSR markers (Table 3). Baloch et al. (2015) criteria were used to determine the polymorphism information content (PIC) for each ISSR marker. Pairwise genetic distance (GDj) was determined using R statistical software as measured by Jaccard’s coefficient (Jaccard 1908). Analysis of molecular variance (AMOVA) was investigated using R statistical software considering variation among structure populations and structure populations within country (Table 4). The population structure was assessed using the Bayesian clustering model-based STRUCTURE software. The UPGMA and principle coordinate analysis (PCoA) were performed using R software to explore the level of diversity among 131 safflower accessions (Team 2013). Evanno et al. (2005) protocol was used through STRUCTURE software to determine the most suitable number of clusters (K subpopulations). We plotted the clusters number (K) against logarithm probability relative to standard deviation (ΔK). Assignment of the individual safflower accessions to the separate population was based on the membership coefficient magnitude being greater than or equal to 50% as outlined by Habyarimana (2016).
3 Results
ISSR marker analysis and genetic diversity
– Twelve most polymorphic ISSR primers produced a total of 201 scorable bands having average of 16.75 bands per primer using 131 safflower accessions. Among 201 ISSR bands, 188 (93.844%) were identified polymorphic having average of 15.67 bands per primer (Table 3). Primer ISSR809 displayed the highest number of total (22) and polymorphic (21) bands, while a lowest number of total (11) and polymorphic (10) bands were found with primer ISSR868. Diversity parameters like mean polymorphism information content, mean effective number of alleles, mean Nei’s gene diversity, mean Shannon’s information index, and mean expected heterozygosity were, respectively, 0.448, 1.655, 0.377, 0.557, and 0.354 among the 12 ISSR primers using 131 safflower accessions (Table 3). The primer ISSR868 was the most informative by revealing a good amount of polymorphism information content (0.592), effective number of alleles (1.849), Nei’s gene diversity (0.454), Shannon’s information index (0.645), and expected heterozygosity (0.441), while the primer ISSR810 was least informative by exhibiting low values of polymorphism information content (0.274), effective number of alleles (1.458), Nei’s gene diversity (0.282), Shannon’s information index (0.436), and expected heterozygosity (0.253).
Pairwise genetic distance with the Jaccard coefficient was computed among the 131 safflower accessions in order to understand the picture of genetic diversity more clearly. The mean genetic distance among 131 accessions was found 0.336. Accessions Pakistan-11 and Israel-1 revealed highest genetic distance (0.816), while accessions USA-5 and Iran-10 showed lowest genetic distance (0.063). Analysis of molecular variance (AMOVA) resulted in highly significant effects of model-based structure (P = 0.001) and model-based structure × country combination (P = 0.003) on genetic differentiation (Table 4).
In accordance with the observed most suitable goodness of fit (K = 3), the Bayesian clustering model implemented in STRUCTURE software divided the evaluated safflower accessions into three main populations; 47 accessions (35.88%) in the population A (green), 19 accessions (14.50%) in the population B (red), 64 accessions (48.86%) in the population C (blue), and 1 accession (0.76%) in an unclassified population (Fig. 1). The UPGMA clustering divided 131 safflower accessions into three main populations and an unclassified population corresponding to the populations identified using the model-based structure (Fig. 2). PCoA divided all accessions into three populations; A, B, and C and an unclassified population which were similar to structure based clustering (Fig. 3).
4 Discussion
The knowledge on the partition of the genetic variation that existed in crop gene pools is helpful to describe the evolution of crop lineages and also disclose the unexplored sources of variation that enhance future crop improvement efforts (Tanksley and McCouch 1997; Yamasaki et al. 2005). Until now, population genetic analysis regarding safflower gene pool has not been fully exploited, and also the hypothesis of Knowles (1969) and Ashri (1975) about the safflower similarity centers is still unclear at the genetic level. Our data presented herein strongly supported the Knowles (1969) hypothesis proposing seven similarity centers. Very few attempts have been done to investigate the total spectrum of variation in global safflower germplasm at the DNA level. Genetic diversity characterization within safflower gene pools is vital for its development and improvement. Our results about mean polymorphism (93.844%) was higher to that of Houmanat et al. (2016), as they found mean polymorphism of 63.38% using ISSR markers evaluating a safflower set of 55 accessions. Similarly, Golkar et al. (2011) reported lower polymorphism (70%) than ours using ISSR markers. Polymorphism information content (PIC) is a widely used metric of the usefulness of molecular markers (Anderson et al. 1993). Higher PIC (0.448) was obtained in the current study in comparison to Talebi and Abhari (2016). They evaluated 25 safflower accessions using 13 ISSR markers. Moreover, Houmanat et al. (2016) revealed lower PIC value (0.23) than us using ISSR markers in safflower. The presence of higher number of effective alleles revealed the availability of maximum level of genetic diversity and is always desirable. We obtained higher effective number of alleles (1.458–1.849) than that of Sung et al. (2010) (1.02–1.09). Obtaining superiority of various diversity parameters in this study than the previous results might be due to the difference of the experimental materials used in the current assessment and also the difference of the ISSR markers used. Shannon’s information index usually distinguishes the level of available genetic diversity in a population, combining abundance and evenness. Kumar et al. (2015) observed lower Shannon’s information index (0.24–0.44) contrary to our observation (0.436–0.645) using AFLP markers. It is a clear indication of the presence of higher level of genetic diversity in the studied safflower accessions with genetic variants evenly distributed throughout the population. Wodajo et al. (2015) reported lower mean Shannon’s information index (0.46) than us (0.557) using ISSR markers. Our results about expected heterozygosity (0.354) are supported by Lee et al. (2014) as they revealed similar expected heterozygosity (0.386). Wodajo et al. (2015) studied 70 safflower accessions using ISSR markers and found Nei’s gene diversity of 0.30, which is lesser than the value (0.377) obtained in this study. Diversity parameters revealed the presence of higher genetic variability in the studied materials suggesting the studied safflower accessions can provide useful building blocks for future breeding programs to enhance safflower productivity. Also, the ISSR markers used in this evaluation should be used for the genetic diversity investigation as these markers exhibited higher diversity levels.
The evaluation of pairwise genetic distance showed a mean of 0.336, with the highest genetic distance between accessions Pakistan-11 and Israel-1, followed by Pakistan-26 and Israel-1 with respective distance values of 0.816 and 0.808. Greater similarity was found between USA-5 and Iran-10 accessions showing least genetic distance of 0.063. One understandable reason behind the presence of maximum genetic similarity might be due to their origin from the common parents. The three most diverse safflower accessions (Pakistan-11, Israel-1, and Pakistan-26) identified during the current study can be recommended as a candidate parents for future breeding programs. The analysis of molecular variance (AMOVA) was used to determine the pattern of the partition of the total gene diversity among and within populations and to assess genetic differentiation. AMOVA showed that most of genetic structure was explained by variations among populations and the genetic populations within countries (Table 4).
The model-based structure application proved more robust and informative in previous investigations (Bouchet et al. 2012; Nadeem et al. 2018b; Ali et al. 2019b). Structure was therefore used in this work as a benchmark for clustering algorithms. The studied 131 safflower accessions were clearly separated into three main populations: A, B, and C and an unclassified population using structure (Fig. 1). Population A consists of 47 accessions originated from Israel (3 accessions), Romania (1 accession), Morocco (2 accessions), Egypt (5 accessions), Pakistan (7 accessions), Spain (4 accessions), Portugal (5 accessions), Iraq (2 accessions), Syria (2 accession), Turkey (6 accessions), Iran (2 accessions), Jordan (1 accession), Afghanistan (2 accession), USA (1 accession), China (2 accessions), and India (2 accessions). Population B comprised 19 safflower accessions including; Iran (2 accessions), Turkey (2 accessions), Pakistan (3 accessions), China (5 accessions), France (1 accession), Austria (1 accession), Libya (1 accession), India (2 accessions), Bangladesh (1 accession), and Russia (1 accession). Clustering of safflower accessions from Mediterranean region with Europe and Asian countries identifies its origin from Mediterranean region and distribution to other parts of the world. The 64 safflower accessions clustered in population C were originated from Syria (1 accession), India (2 accessions), Jordan (4 accessions), Israel (1 accession), Turkey (8 accessions), Afghanistan (3 accessions), China (2 accessions), Canada (2 accession), USA (4 accessions), Iran (6 accession), Bangladesh (3 accessions), Uzbekistan (3 accessions), Australia (1 accession), Austria (1 accession), Pakistan (17 accessions), Hungary (1 accession), Kazakhstan (1 accession), Thailand (1 accession), Argentina (1 accession), Egypt (1 accession), and Portugal (1 accession). Clustering pattern of accessions and their distribution in population C was found similar to populations A and B. Distribution of safflower accessions from Mediterranean region to Asia took place through Turkey, being used as a bridge. According to Nadeem et al. (2018b), Turkey acts as bridge for the diffusion of various crops among the continents. One safflower accession originated from Pakistan (Pakistan-12) made up the unclassified population as its membership coefficient magnitude was less than 50% as proposed by Habyarimana (2016).
Population A included accessions from Asia (29 accessions), Europe (10 accessions), Africa (7 accessions), and American (1 accession) continents. Population B exhibited accessions from Asia (16 accessions), Europe (2 accessions), and Africa (1 accession). Population C revealed accessions from Asia (52 accessions), America (7 accessions), Oceania (1 accession), Europe (3 accessions), and Africa (1 accession). The unclassified population exhibited only one accession that is originated from Asian (Pakistan) continent. Besides sharing common parentage, accessions similarity in the same population during the clustering might also be due to convergent evolution and selection (Golkar et al. 2011). Population C stood the most diverse population as it comprised accessions from all the available continents.
Knowles (1969) suggested the presence of seven similarity centers for safflower throughout the world using various morpho-agronomic traits. Most of the accessions evaluated in this study follow the hypothesis of seven similarity centers at molecular level. But the data obtained from the ISSR markers in this study did not fully support the Knowles’s hypothesis of similarity centers. Safflower accessions from different similarity centers clustered together and highlighted the lack of importance of similarity centers at molecular level which was previously reported in the scientific literature (Chapman and Burke 2007). Safflower accessions from Israel, Iraq, Syria, Turkey, Iran, and Jordan were present in population A and can be assigned to the Middle East similarity center. Similarly, accessions from India and Pakistan were also present in population A comprising the India-Pakistan similarity center. Accessions from Pakistan, India, and Bangladesh were clustered in population B and made the India–Pakistan similarity center. Population C revealed the Middle East similarity center as it exhibited safflower accessions from Syria, Jordan, Israel, Turkey, Afghanistan, and Iran. Population C also exhibited safflower accessions from the India-Pakistan (India, Bangladesh, and Pakistan) and Europe (Australia, Austria, Hungary, Argentina, and Portugal) similarity centers. Very recently Ali et al. (2019b) aimed to evaluate the similar centers pattern at molecular level using 13 iPBS-retrotransposon markers and supported the Knowles (1969) hypothesis proposing seven similar centers. Our results are supported by their findings revealing similar safflower similarity centers patterns. Besides obtaining supportive results to the Knowles’s hypothesis of seven similarity centers, still there is a need to conduct more research at the molecular level by collecting and testing safflower accessions from the all proposed similarity centers.
The exploration of genetic relationships between the studied 131 safflower accessions using UPGMA resulted in a comparable clustering pattern to that of model-based algorithm with few exceptions as three accessions belonging to population B (Russia-1, India-3, and India-4) clustered with population C (Fig. 2). Seven accessions belonging to population C (Jordan-5, Portugal-5, Egypt-3, Pakistan-7, Iran-2, Uzbekistan-1, and Jordan-2) clustered with population A. Similarly, two accessions from population A (Turkey-3 and Afghanistan-2) clustered with population C. Unclassified safflower accession (Pakistan-12) clustered with population A. Accessions present in the same population revealed full membership coefficients in model-based Structure. The discrepancies displayed in UPGMA clustering might be described by its reduced resolution power relative to the model-based Structure (Bouchet et al. 2012).
Principal coordinate analysis (PCoA) confirmed the clustering based on structure algorithm of 131 safflower accessions into clearly distinguishable three main populations and an unclassified population using 12 ISSR primers (Fig. 3). The occurrence of some light differences between model-based structure and PCoA can derive from its differing clustering resolution, with more resolution revealed by the model-based structure analysis. Existence of the genomic admixture might be the reason for the misclassification in the principal coordinate space of the 131 safflower accessions. Also, same pattern of the similarity centers as obtained through structure based analysis was exhibited by PCoA analysis.
Overall, a reasonable level of genetic diversity was revealed by ISSR markers and the obtained diversity can be used for the improvement of safflower in future breeding programs. As ISSR marker system revealed competitive results in the current and previous studies, it is highly warranted that ISSR markers should be used as an important tool for the evaluation of the safflower germplasm at DNA level for productivity enhancement. To confirm the uncertainty about the safflower similarity centers, there is a need to include various robust sampling techniques like random sampling without replacement to be implemented in the accessions stored in the various world safflower seed repositories. It will be interesting to characterize those accessions through the various clustering algorithms such as those implemented in this work.
The presence of good level of genome diversity was observed among the studied materials. Clustering algorithms like model-based structure, UPGMA, and PCoA clustered safflower accessions according to collection countries and similarity centers. The current findings greatly support the Knowles hypothesis of seven similarity centers for safflower at DNA level with ISSR markers, but still need to be confirmed by conducting further research work. Analysis of molecular variance (AMOVA) exhibited highly significant differentiation among the identified populations, and population × country combinations. Safflower accessions from Asian continent revealed higher genetic diversity in comparison to other continents. On individual bases, safflower accessions; Pakistan-11, Israel-1, and Pakistan-26 were found most diverse at DNA level and might be suggested as parental germplasm resources for future breeding programs.
References
Ali F, Yilmaz A, Chaudhary HJ, Nadeem MA, Arslan Y, Nawaz MA et al (2019a) Investigation of morpho-agronomic performance and selection indices in the international safflower panel for breeding perspectives. Turk J Agric For. https://doi.org/10.3906/tar-1902-49
Ali F, Yılmaz A, Nadeem MA, Habyarimana E, Subaşı I, Nawaz MA et al (2019b) Mobile genomic element diversity in world collection of safflower (Carthamus tinctorius L.) panel using iPBS-retrotransposon markers. PLoS ONE 14:2
Ambreen H, Kumar S, Variath MT, Joshi G, Bali S, Agarwal M, Kumar A, Jagannath A, Goel S (2015) Development of genomic microsatellite markers in Carthamus tinctorius L. (Safflower) using next generation sequencing and assessment of their cross-species transferability and utility for diversity analysis. PLoS ONE 10:8
Ambreen H, Kumar S, Kumar A, Agarwal M, Jagannath A, Goel S (2018) Association mapping for important agronomic traits in safflower (Carthamus tinctorius L.) core collection using microsatellite markers. Front Plant Sci 402:9
Amini F, Saeidi G, Arzani A (2008) Study of genetic diversity in safflower genotypes using agro-morphological traits and RAPD markers. Euphytica 163:21–30
Amiri RM, Azdi SB, Ghanadha MR, Abd MC (2001) Detection of DNA polymorphism in landrace populations of safflower in Iran using RAPD-PCR technique. Iran J Agric Sci 32:737–745
Anderson JA, Churchill GA, Autrique JE, Tanksley SD, Sorrells ME (1993) Optimizing parental selection for genetic linkage maps. Genome 36:181–186
Ansari A, Sikarwar P, Lade S, Yadav H, Ranade S (2016) Genetic diversity clusters in germplasm of Cluster Bean (Cyamopsis tetragonoloba L., Taub), an important food and an industrial legume crop. J Agric Sci Technol 18:1407–1418
Arystanbekkyzy M, Nadeem MA, Aktas H, Yeken MZ, Zencirci N, Nawaz MA, Ali F, Haider MS, Tunc K, Chung G, Baloch FS (2018) Phylogenetic and taxonomic relationship of Turkish wild and cultivated emmer (Triticum turgidum ssp. dicoccoides) revealed by IPBS-retrotransposons markers. Int J Agric Biol 21:155–163
Ashri A (1975) Evaluation of the germ plasm collection of safflower, Carthamus tinctorius L. V. distribution and regional divergence for morphological characters. Euphytica 24:651–659
Bagmohammadi H, Pahlevani M, Ahmadikhah A, Razavi SE (2012) Genetic variation of safflower (Carthamus tinctorius L.) and related species revealed by ISSR analysis. Plant Breed Seed Sci 66:139–150
Baloch FS, Alsaleh A, de Miera LES, Hatipoglu R, Ciftci V, Karakoy T, Yıldız M, Ozkan H (2015) DNA based iPBS-retrotransposon markers for investigating the population structure of pea (Pisum sativum) germplasm from Turkey. Biochem Syst Ecol 61:244–252
Baloch FS, Alsaleh A, Andeden EE, Hatipoglu R, Nachit M, Ozkan H (2016) High levels of segregation distortion in the molecular linkage map of bread wheat representing West Asia and North Africa region. Turk J Agric For 40:352–364
Baloch FS, Alsaleh A, Shahid MQ, Çiftçi V, de Miera LES, Aasim M et al (2017) A whole genome DArTseq and SNP analysis for genetic diversity assessment in Durum Wheat from Central Fertile Crescent. PLoS ONE 12:1
Bouchet S, Pot D, Deu M, Rami JF, Billot C, Perrier X et al (2012) Genetic structure, linkage disequilibrium and signature of selection in sorghum: lessons from physically anchored DArT markers. PLoS ONE 7:3
Casaoli M, Mattion C, Cherubini M, Villani FA (2001) Genetic linkage map of European chestnut (Castanea sativa Mill.) based on RAPD, ISSR and isozyme markers. Theor Appl Genet 102:1190–1199
Castañeda-Álvarez NP, Khoury CK, Achicanoy HA et al (2016) Global conservation priorities for crop wild relatives. Nat Plants 2:4
Cekic C, Battey NH, Wilkinson MJ (2011) The potential of ISSR-PCR primer-pair combinations for genetic. Theor Appl Genet 103:540–546
Chapman MA, Burke JM (2007) DNA sequence diversity and the origin of cultivated safflower (Carthamus tinctorius L.; Asteraceae). BMC Plant Biol 7:60
Chapman MA, Hvala J, Strever J, Burke JM (2010) Population genetic analysis of safflower (Carthamus tinctorius L.; Asteraceae) reveals a near Eastern origin and five centers of diversity. Am J Bot 97:831–840
Dordas CA, Sioulas C (2009) Dry matter and nitrogen accumulation, partitioning, and retranslocation in safflower (Carthamus tinctorius L.) as affected by nitrogen fertilization. Field Crop Res 110:35–43
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:13–15
Ekincialp A, Erdinc C, Turan S, Cakmakci O, Nadeem MA, Baloch FS et al (2019) Genetic characterization of Rheum ribes (Wild Rhubarb) genotypes in lake Van Basin of Turkey through ISSR and SSR Markers. IJAB 21:795–802
Erzurumlu GS, Sultana N, Vural M, Serce S (2018) Genetic and phenotypic variation among Turkish terrestrial orchid species as revealed by RAPD and morphological characteristics. Turk J Agric For 42:227–236
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
FAO (2018) FAO Statistical Yearbook 2012. http://www.fao.org/docrep/017/i3138e/i3138e.pdf. Accessed 5 Sept 2018
FAO (2019) Oilseeds and oilseed products. Agricultural Outlook 2018–2027. http://www.fao.org/docrep/i9166e/i9166e_Chapter4_Oilseeds.pdf. Accessed 7 April 2019
Gautam AK, Gupta N, Bhadkariya R, Srivastava N, Bhagyawant SS (2016) Genetic diversity analysis in chickpea employing ISSR markers. Agrotech 5:2
Godfray HC, Beddington JR, Crute IR, Haddad L, Lawrence D, Muir JF, Pretty J, Robinson S, Thomas SM, Toulmin C (2010) Food security: the challenge of feeding 9 billion people. Science 327:812–818
Golkar P, Arzani A, Rezaei AM (2011) Genetic variation in safflower (Carthamus tinctorious L) for seed quality-related traits and inter-simple sequence repeat (ISSR) markers. Int J Mol Sci 12:2664–2677
Guliyev N, Sharifova S, Ojaghi J, Abbasov M, Akparov Z (2018) Genetic diversity among melon (Cucumis melo L.) accessions revealed by morphological traits and ISSR markers. Turk J Agric For 42:393–401
Habyarimana E (2016) Genomic prediction for yield improvement and safeguarding of genetic diversity in CIMMYT spring wheat (Triticum aestivum L.). Aust J Crop Sci 10:127–136
Hadian J, Raeisi S, Azizi A, Pezhmanmehr M, Sarkhosh A (2017) Genetic diversity of natural populations of medicinally valuable plant Satureja khuzistanica Jamzad based on ISSR markers. Braz J Bot 40:771–781
Houmanat K, Charafi J, Mazouz H, El Fechtali M, Nabloussi A (2016) Genetic diversity analysis of safflower (Carthamus tinctorius L.) accessions from different geographic origins using ISSR markers. Int J Agric Biol 18:881–887
Jaccard P (1908) Nouvelles recherches sur la distribution florale. Bulletin de la Société vaudoise des sciences naturelles 144:223–270
Johnson RC, Kisha TJ, Evans MA (2007) Characterizing safflower germplasm with AFLP molecular markers. Crop Sci 47:1728–1736
Khalil RMA, El zayat MAS (2019) Molecular characterization of some Brassica species. Adv Plants Agric Res 9:112–119
Khan MA, Von Witzke-Ehbrecht S, Maass BL, Becker HC (2009) Relationships among different geographical groups, agro-morphology, fatty acid composition and RAPD marker diversity in safflower (Carthamus tinctorius L.). Genet Resour Crop Evol 56:19–30
Knowles PF (1969) Centers of plant diversity and conservation of crop germplasm: Safflower. Econ Bot 23:324–329
Kumar S, Ambreen H, Murali TV, Bali S, Agarwal M, Kumar A, Goel S, Jagannath A (2015) Assessment of genetic diversity and population structure in a global reference collection of 531 accessions of Carthamus tinctorius (Safflower) using AFLP markers. Plant Mol Biol Rep 33:1299–1313
Kumari S, Choudhary RC, Kumara Swamy RV, Saharan V, Joshi A, Munot J (2017) Assessment of genetic diversity in safflower (Carthamus tinctorius L.) genotypes through morphological and SSR marker. J Pharmacogn Phytochem 6:2723–2731
Lee GA, Sung JS, Lee SY, Chung JW, Yi JY, Kim YG, Lee MC (2014) Genetic assessment of safflower (Carthamus tinctorius L.) collection with microsatellite markers acquired via pyrosequencing method. Mol Ecol Resour 14:69–78
Long SP, Marshall-Colon A, Zhu XG (2015) Meeting the global food demand of the future by engineering crop photosynthesis and yield potential. Cell 161:56–66
Majidi MM, Zadhoush S (2014) Molecular and morphological variation in a world-wide collection of safflower. Crop Sci 54:2109–2119
Marinova E, Riehl S (2009) Carthamus species in the ancient Near East and south-eastern Europe: archaeobotanical evidence for their distribution and use as a source of oil. Veg Hist Archaeobot 18:341–349
Mary SS, Gopalan A (2006) Dissection of genetic attributes yield traits of fodder cowpea in F3 and F4. J Appl Sci Res 2:805–808
Mayerhofer M, Mayerhofer R, Topinka D, Christianson J, Good AG (2011) Introgression potential between safflower (Carthamus tinctorius L.) and wild relatives of the genus Carthamus. BMC Plant Biol 11:47
Nadeem MA, Nawaz MA, Shahid MQ, Dogan Y, Comertpay G, Yıldız M, Hatipoglu R, Ahmad F, Alsaleh A, Labhane N, Ozkan H, Chung G, Baloch FS (2018a) DNA molecular markers in plant breeding: current status and recent advancements in the genomic selection and genome editing. Biotechnol Biotechnol Equip 32:261–285
Nadeem MA, Habyarimana E, Çiftçi V, Nawaz MA, Karaköy T, Comertpay G, Shahid MQ, Hatipoğlu R, Yeken MZ, Ali F, Ercişli S (2018b) Characterization of genetic diversity in Turkish common bean gene pool using phenotypic and whole-genome DArTseq-generated silicoDArT marker information. PLoS ONE 13:10
Nagaoka T, Ogihara Y (1997) Applicability of inter-simple sequence repeat markers in wheat for use as DNA markers in comparison to RFLP and RAPD markers. Theor Appl Genet 94:597–602
Pearl SA, Burke JM (2014) Genetic diversity in Carthamus tinctorius (Asteraceae; safflower), an underutilized oilseed crop. Am J Bot 10:1640–1650
Potter D, Gao FY, Aliello G, Leslie C, McGranahan G (2002) Inter simple sequence repeat markers for fingerprinting and determining genetic relationships of walnut (Juglans regia) cultivars. J Am Soc Hortic Sci 127:75–81
Rawat S, Jugran AK, Bhatt ID, Rawal RS, Nandi SK (2016) Genetic diversity analysis in natural populations of Roscoea procera Wall. from West Himalaya, India. Braz J Bot 39:621–630
Sabzalian MR, Mirlohi A, Saeidi G, Rabbani MT (2009) Genetic variation among populations of wild safflower, Carthamus oxyacanthus analyzed by agro-morphological traits and ISSR markers. Genet Resour Crop Evol 56:1057–1064
Sung JS, Cho GT, Lee GA, Baek HJ, Hu MK (2010) Phylogenetic relationships and genetic diversity in collected resources of Carthamus tinctorius by Random Amplified Polymorphic DNA markers. J Life Sci 20:1764–1771
Talebi R, Abhari SA (2016) Evaluation of genetic diversity in safflower (Carthamus tinctorius L.) using agro-morphological, fatty acid composition and ISSR molecular markers. Res J Biotechnol 11:7
Talebi R, Fayaz F, Karami E (2012) Morphometric and amplified fragment length polymorphism marker analysis in some landrace wheat (Triticum aestivum) genotypes collected from north-west Iran. Environ Exp Biol 10:49–56
Tanksley SD, McCouch SR (1997) Seed banks and molecular maps: Unlocking genetic potential from the wild. Science 277:1063–1066
Tanyolac B (2003) Inter-simple sequence repeat (ISSR) and RAPD variation among wild barely (Hordeum vulgare subsp. spontaneum) populations from west Turkey. Genet Resour Crop Evol 50:611–614
Team RC (2013) R: a language and environment for statistical computing. R foundation for statistical computing. Vienna, Austria. 2013; ISBN 3-900051-07-0. http://www.R-project.org/
Weiss EA (1983) Oilseed crops. Longman Inc., Harlow, p 660
Weiss EA (2000) Safflower: oil seed crops. Blackwell Science, Oxford
Wodajo B, Mustefa FB, Tesfaye K (2015) Clustering analysis of Ethiopian safflower (Carthamus tinctorius L.) using ISSR Markers. Int J Sci Res Publ 5:434–440
Yeh FC, Yang R, Boyle TJ, Ye Z, Xiyan, JM (2000) PopGene32, Microsoft Windows-based freeware for population genetic analysis, Version 1.32. Molecular Biology and Biotechnology Centre, University of Alberta, Edmonton, Alberta, Canada
Yaldiz G, Camlica M, Nadeem MA, Nawaz MA, Baloch FS (2018) Genetic diversity assessment in Nicotiana tabacum L. with iPBS-retrotransposons. Turk J Agric For 42:154–164
Yamasaki M, Tenaillon MI, Bi IV, Schroeder SG, Sanchez-Villeda H, Doebley JF, Gaut BS, McMullen MD (2005) A large-scale screen for artificial selection in maize identifies candidate agronomic loci for domestication and crop improvement. Plant Cell 17:2859–2872
Yang YX, Wu W, Zheng YL, Chen L, Liu RJ, Huang CY (2007) Genetic diversity and relationships among safflower (Carthamus tinctorius L.) analyzed by inter-simple sequence repeats (ISSRs). Genet Resour Crop Evol 54:104–1051
Yildiz M, Koçak M, Nadeem MA, Cavagnaro P, Barboza K, Baloch FS et al (2019) Genetic diversity analysis in the Turkish pepper germplasm using iPBS retrotransposon-based markers. Turk J Agric For. https://doi.org/10.3906/tar-1902-10
Zeinali E (1999) Safflower, characteristics, productions and utilization. Gorgan University of Agricultural Sciences and Natural Resources Press, Gorgan, p 137
Zietkiewicz E, Rafalski A, Labuda D (1994) Genome fingerprinting by simple sequence repeat (SSR)-anchored polymerase chain reaction amplification. Genomics 20:176–183
Acknowledgements
The authors express their gratitude to TUBITAK (The Scientific and Technological Research Council of Turkey) for providing a research fellowship to Fawad Ali under the TUBITAK-2216 fellowship program for international researchers.
Author information
Authors and Affiliations
Contributions
HJC provided experimental materials. FA, MAN, and AY performed experiment and writing of the manuscript. FSB supervised the experiment. MAN, IHK, SE, GC, and HJC performed editing of the manuscript. EH and MAN supervised the molecular analysis and helped in structuring the manuscript.
Corresponding authors
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ali, F., Nadeem, M.A., Habyarimana, E. et al. Molecular characterization of genetic diversity and similarity centers of safflower accessions with ISSR markers. Braz. J. Bot 43, 109–121 (2020). https://doi.org/10.1007/s40415-019-00574-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40415-019-00574-7