Abstract
The present study is undertaken to analyse the genetic diversity of Roscoea procera Wall. (an endemic Himalayan medicinal herb) collected from 11 different populations of West Himalaya using ISSR markers. Of the total 130 ISSR primers screened, only ten ISSR primers generated 94 clear and reproducible banding patterns, of which 89 bands (94.68 %) were polymorphic. Based on the percentage of polymorphic loci (94.68 %), Nei’s gene diversity (0.198) and Shannon’s information index (0.329), high genetic diversity was detected at the species level. Genetic differentiation (G ST = 0.202) was also recorded as moderate level due to high gene flow (Nm = 1.976) among populations. Analysis of molecular variance (AMOVA) indicated that most of the genetic variation (90 %) resided within the populations (Φ ST = 0.10). STRUCTURE analysis revealed two ancestral groups of the species, which were further supported by neighbour-joining cluster analysis. Among the different ecological habitats, high genetic diversity in populations under open grassy land showed suitability and preferred habitat conditions. High genetic diversity within population suggests in situ and ex situ conservation of selected populations for conserving and maintaining wide genetic pool of the species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Roscoea procera Wall. (Syn. R. purpurea Sm.) is one of the endemic Himalayan herbs found at an altitude of 1800–3000 m above mean sea level (amsl). It has been reported from Kashmir to Assam in India including the neighbouring countries Nepal and Bhutan (Ngamriabsakul et al. 2000; Mohandass et al. 2015). Tuberous roots of the species are used in many traditional medicines and substitute of ‘Safed musli’, which is one of the ingredients of Ayurvedic formulations, e.g. Muslipak, Chyvanprash and Chyvanprash aveleha (Rana et al. 2004). The species is also reported to be used in seminal debility, diabetes, leucorrhoea, diarrhoea, dysentery, burning, phthisis, malaria and in several other medicines. For example, root powder mixed with black pepper is reported to be applied on boils for quick healing, decoction of root used in jaundice treatment and boiled rhizome consumed as food in Nepal (Rana et al. 2004; Sahu et al. 2010; Rawat et al. 2014). The species is regularly harvested from wild to meet the demand of local and traditional healers. Further, unregulated harvesting along with natural calamities in the region has altered the habitat and consequently reduced the natural populations severely. Therefore, there is a need to develop conservation strategies of this species.
Conservation of the genetic resources of rare, endangered and endemic plants is crucial, and worldwide efforts are on to combat this issue to save species with restricted and small population size (Bauert et al. 1998). Genetic markers have extensively been used in conservation genetics, which gives insight into genetic processes in populations (e.g. mating system, gene flow, etc.), along with past demographical events (e.g. bottlenecks, genetic differentiation, etc.) to propose conservation measures (Milligan et al. 1994). In this context, molecular markers such as inter-simple sequence repeats (ISSR) have been widely used for population genetic studies of many endemic and medicinal plants (Xiao et al. 2004; Li and Xia 2005; Luan et al. 2006; Li and Jin 2007; Wang et al. 2008; Hu et al. 2010; Zhao et al. 2012). ISSR primers of repeat sequences (microsatellites) are reported to be dispersed throughout genomes anchored either at 5″ or 3″ end with one or few specific nucleotides and amplify the sequences between the two microsatellite loci (Hu et al. 2010). Because of the higher annealing temperature and longer sequence of ISSR primers, they can yield more reliable and reproducible bands than rapid amplified polymorphic DNA (RAPD; Goulao et al. 2001). Cost of the analyses is relatively lower in ISSR markers than that of some other markers, such as restriction fragment length polymorphism (RFLP), simple sequence repeats (SSRs) and amplified fragment length polymorphism (AFLP; Wang et al. 2008). Hence, in view of the importance of R. procera, this study was carried out to analyse genetic diversity of natural populations of R. procera from West Himalaya, India using ISSR markers. The results will be useful in designing conservation strategies for this important medicinal plant.
Materials and methods
Study sites and plant materials
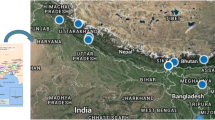
Eleven populations of Roscoea procera were sampled from Uttarakhand region of West Himalaya for DNA extraction. All the populations belong to two geographical regions: Garhwal and Kumaun. Each population was positioned by a Global Positioning System (GPS), and the location details are given in Fig. 1 and Supplementary Table 1. In each population, individuals were randomly collected and planted in herbal garden of G. B. Pant Institute of Himalayan Environment and Development, Kosi-Katarmal, Almora, Uttarakhand, India. Fresh young leaves were collected and kept in ziplock bags with silica gel and brought to laboratory for DNA extraction.
Geographical location of populations of R. procera in Uttarakhand; populations are presented as MK Manjkhali, CB Chaubatiya, LG Lamgarha, NT Nainital, DG Dunagiri, TK Thakurh, PK Pandukholi, MS Mussoorie, SK Suakholi, BK Buranshkhanda and SR Surkunda. Populations at right hand side (total 7 No.) represented Kumaun and left hand side (4 No.) Garhwal geographical region. Pie diagrams represent geographical distribution of fractions of Bayesian clusters (encoded by red and green colour) in combined populations of Garhwal and Kumaun geographical regions. (Color figure online)
DNA extraction
Total genomic DNA of 78 genotypes belonging to 11 different populations was extracted from young leaf tissue using CTAB method (Doyle and Doyle 1987). Briefly, 1 g leaf material was powdered by transferring into liquid nitrogen, then placed in 10-ml centrifuge tube containing 5 ml preheated CTAB buffer with 2-mercaptoethanol and incubated at 65 °C for 60 min. Subsequently, 4 ml cold chloroform:isoamyl alcohol (24:1, v/v) was added, followed by gentle shaking for 5 min and centrifugation at 10,000 rpm (10 min, 25 ± 2 °C). The supernatant was recovered and mixed with 3 ml of ice-cold isopropanol. DNA was then recovered as pellet after centrifugation (12,000 rpm; 10 min, −4 ± 2 °C), washed in 70 % ethanol, dried and resuspended in 250 μl Tris–EDTA buffer (TE buffer). The quality of DNA was checked in 1 % agarose gel and quantified using a spectrophotometer (U 2100, Hitachi, Japan).
ISSR-PCR amplification
PCR amplification reaction for ISSR analysis were conducted in a total volume of 20 µl in a 0.2 ml microfuge tube with 2 µl of 10 X reaction buffer, 2 mM MgCl2, 200 µM dNTPs, 10 pmol primer, 20 ng DNA template and 1 U Taq DNA polymerase (Genetix, Delhi, India). The programme for PCR was set for 5 min at 95 °C followed by 35 cycles of 94 °C for 1 min, annealing at different temperatures (40–55 °C) for different primers for 1 min, extension at 72 °C for 2 min, and final extension cycle of 7 min at 72 °C. The reactions were then placed at 4 °C until gel analysis. In total, 30 (Merck Biosciences Germany) and 100 primers (University of British Columbia primer set no. 9; purchased from Invitrogen, Bangalore) were screened, and based on the number and quality of polymorphic fragments, ten primers were selected for further analysis. The amplification was carried in a T-gradient PCR system (Biometra, Goettingen, Germany). The amplification products were separated by gel electrophoresis on 2 % agarose and visualised under the UVI Pro platinum gel imaging system (Version 11.9, Cambridge, UK). Gene Ruler TM DNA Ladder Mix (Genetix, Delhi, India) was used as the DNA fragment size marker.
Data analysis
The amplified DNA fragments were scored as the presence (1) or the absence (0), and only those consistently reproducible bands were scored. The resulting binary data matrix was analysed using POPGENE (Version 1.32). Percentage of polymorphic loci (Pp%), effective number of alleles (Ne), observed number of alleles (Na), Nei’s (1973) gene diversity (He) and Shannon’s information index (I) were calculated for genetic diversity assessment. At the species level, total genetic diversity (H T), genetic diversity within populations (H S) and Nei’s (1973) coefficient of genetic differentiation among populations (G ST, G ST = (H T − H S)/H T) were calculated. The corresponding estimates of gene flow (Nm) were subsequently calculated (McDermott and McDonald 1993). A dendrogram was also constructed by application of the unweighted pair group method with arithmetic average (UPGMA) cluster analysis using DARwin (version 5.0.23; Perrier and Jacquemoud-Collet 2006). Principal component analysis (PCoA) for genetic relationship of individuals was performed by the GenAlex (ver. 6 package) (Peakall and Smouse 2006).
A Bayesian clustering-based STRUCTURE analysis was performed on the entire data set using STRUCTURE programme (version 2.3.4), which uses Markov Chain Monte Carlo (MCMC) algorithm to detect population structure (Pritchard et al. 2000) and estimate individuals into subgroups (K) in a sample and to assign individuals to one or more of these populations (K). The “Structure Harvester” program was used (http://taylor0.biology.ucla.edu) to determine the final population structure. The most likely number of clusters was estimated according to the model value (ΔK) based on the second-order rate of change, with respect to K, of the likelihood function, following the procedure described by Evanno et al. (2005).
Analysis of molecular variance (AMOVA) was used to estimate the partitioning of genetic variance among and within populations using GenAlex6 (Peakall and Smouse 2006). In order to test the correlation between genetic and geographical distances (in kilometres), Mantel test (Mantel 1967) was performed among populations using GenAlex6 computing 999 permutations (Peakall and Smouse 2006).
Results
Genetic diversity
A total of 94 bands using 10 ISSR primers were recorded, of which 89 bands (94.68 %) were polymorphic (Table 1; Supplementary Fig. 1). The size of amplified bands ranged between 250 and 1800 bp from 78 individuals representing 11 populations of the species. Primer UBC-848 yielded the lowest number of bands (07), whereas primers UBC-842 the highest number of bands (14). The average number of bands per primer was 9.40. The percentage of polymorphic loci ranged from 40.43 (Manjkhali) to 72.34 (Pandukholi) (Table 2), the observed number of alleles ranged from 1.40 (Manjkhali) to 1.72 (Pandukholi) and the effective number of alleles ranged from 1.15 (Manjkhali) to 1.36 (Lamgarha). At species level, the observed number of alleles (1.95) and effective number of alleles 1.29 were found. Nei’s gene diversity values ranged from 0.102 (Manjkhali) to 0.217 (Lamgarha) among different populations and 0.198 at the species level. The value of Shannon information index ranged from 0.165 (Manjkhali) to 0.331 (Lamgarha) within populations and 0.329 at species levels.
Genetic differentiation and population structure
Total gene diversity (H T) and within-population gene diversity (H S) were recorded to be 0.198 and 0.166, respectively. The genetic differentiation (G ST) among the 11 sampled populations of this species was low (0.202). Estimated gene flow among the populations was 1.976 individuals per generation, which indicated a higher rate of migration of individuals between populations per generations (Table 3). Analysis of molecular variance (AMOVA) showed highly significant genetic differences among populations and revealed a higher proportion of genetic variation within the population (90 %) compared to among populations diversity (Φ ST = 10 %) (Table 3). The maximum Nei’s genetic distance (0.100) was found between Suakholi and Dunagiri, whereas Mussoorie population showed the least genetic distance (−0.005) to Lamgarha and Thakurh population (Table 4). Individually, neighbour-joining cluster analysis revealed mixing of individuals in clusters of different population and regions with each other. In cluster analysis, initially two groups were formed. The first group represents 25 individuals while the second 53, which was further subdivided into three more groups representing 12, 25 and 18 individuals (Fig. 2).
Unrooted tree obtained by the Unweighted Neighbour-Joining method illustrating genetic relationship of for 78 individuals from 11 populations of Roscoea procera using ISSR markers. Every population is represented in a separate colour. Population codes are as follows: MK Manjkhali, CB Chaubatiya, LG Lamgarha, NT Nainital, DG Dunagiri, TK Thakurh, PK Pandukholi, MS Mussoorie, SK Suakholi, BK Buranshkhanda, SR Surkunda. All the individuals clustered into two groups, A and B
Based on maximum likelihood and delta K (ΔK) values, two optimal groups or clusters (K) were identified (Fig. 3). By using a membership probability threshold (Q) of 0.60, majority of the individuals were clearly assigned into two specific groups. Among these, 52 individuals (66.67 %) formed the first group (cluster-I) with individuals mainly collected from Manjkhali, Nainital, Dunagiri and Pandukholi populations. On the other hand, 19 individuals (24.36 %) formed a second group (cluster-II) with individuals mainly collected from Surkunda, while 9 individuals were retained in the admixed group. This also indicates that all the 11 populations analysed in the present study represented two discrete genetic clusters showing strong admixture of individuals across the natural populations from the two regions (Garhwal and Kumaun) of the West Himalaya (Supplementary Table 1, Fig. 1).
A Mean log probability of the data for the 12 Structure runs at each K. Error bars are standard deviations; B Delta K, rate of change in the log probability of data between successive K values; Biplot detected the maximum peak at K = 2 (the optimal number of clusters) based on Evanno et al. (2005) prediction; C Population structure inferred by Bayesian cluster analyses (Structure) for 78 individuals from 11 populations. Results for K = 2 (number of clusters) are shown. Each individual (grouped by population) is represented by a vertical bar. The proportion of the bar in each of K colours corresponds to the average posterior likelihood that the individual is assigned to the cluster indicated by that colour. Populations are separated by black lines
The results of Mantel test showed a nonsignificant relation (y = −0.00031x + 0.043; R 2 = 0.0027) between geographical and genetic distance. Thus, these results reject the hypothesis of isolation-by-distance. While, correlating the genetic diversity of the species with altitude, significant relationship was not found with different diversity parameters, i.e. number of polymorphic loci (r = 0.331), percentage of polymorphic loci (r = 0.331), observed number of alleles (r = 0.343), effective number of alleles (r = 0.429), Nei’s gene diversity (0.448) and Shannon’s information index (r = 0.449), but showed increasing trend with altitude (data not shown). Among different habitat conditions, genetic diversity of the species in open grassy habitat (He—1.81) was comparatively better than the species under oak (He—1.67), oak mixed (He—1.63) and pine forest (He—0.127) habitat types.
Discussion
The present study offers the first detailed report on genetic diversity and population structure for R. procera. Information on genetic diversity on rare, endangered species is important for effective conservation strategies. The genetic diversity at population level has been reported in several species of family Zingiberaceae. Compared to the present study, lower levels of genetic diversity (Nei’s genetic diversity—He) were recorded in several Zingiber species (Zingiber zerumbet (L.) Smith—0.1407; Z. nimmonii (J.Graham) Dalzell—0.1506; Z. neesanum (J.Graham) Ramamoorthy—0.1637; and Z. officinale Roscoe—0.0969) in Western Ghats of India using AFLP markers (Kavitha et al. 2010). However, the levels of genetic diversity were more or less similar compared with Hedychium spicatum Buch.-Ham. ex. Smith using allozyme (Pp% = 73.08 %, He = 0.372, Ne = 1.73) (Jugran et al. 2011) and RAPD analysis (Pp% = 89.58, Na = 1.896, Ne = 1.496, I = 0.294 and He = 0.447) (Chaudhary et al. 2012). The level of genetic diversity of the species (He = 0.197) was found to be comparatively lower than the widespread or long-lived 37 perennial plants (He = 0.22) based on RAPD analysis (Nybom 2004). Further, Shannon’s diversity index (I) of target species was also lesser (0.329) than the reported mean value of out-crossing species (He = 0.447) (Bussell 1999). Several studies based on ISSR data have demonstrated that endangered and endemic species tend to possess lower levels of genetic diversity (Xiao et al. 2004; Li and Xia 2005); many others have, however, reported contrasting findings (Luan et al. 2006; Li and Jin 2007). Genetic diversity of a plant species can be affected by many factors such as distribution range, life form, breeding system and seed-dispersal mechanism. It has been reported that a species, which has a long life, high frequency of gene flow and more number of seeds, tends to possess high genetic diversity (Hamrick and Godt 1990). Therefore, it cannot be conclusively stated that the polymorphism is solely responsible for rarity of any plant species. The roles of other factors, such as anthropogenic pressure, changing climate, habitat destruction, and other factors, cannot be ignored.
The data on genetic structure of the present study showed that differentiation coefficients (G ST = 0.202 and Φ ST = 0.100) were low in the species, although it was higher or similar to the average coefficients of long-lived perennial (G ST = 0.19, n = 24) and out-crossing species (G ST = 0.22, n = 31) (Nybom 2004). R. procera can be included under moderate level of genetic differentiation (G ST = 0.202) compared to the reported values for different levels (Nei 1978). Although, high gene flow observed in this investigation (Nm = 1.976) might be a factor responsible for lowering the genetic differentiation among the populations, the value of gene flow higher than 1.0 individual per generation, however, has generally not been regarded as threshold quantity beyond which significance differentiation occurs (Slatkin 1987). R. procera is a possible out-crossing species in which the upper and lower lobes of flower make a good configuration for pollinator attraction. Clustering of the different populations of R. procera in two clusters as revealed in the STRUCTURE and neighbour-joining cluster analyses supported the admixture of individuals among the assumed geographical populations. Roscoea species with striking orchid-like flowers possess a long floral tube that apparently attracts insects for pollination, particularly the long-tongued insects (Fenster et al. 2004). Roscoea mostly grows as a clump of individuals, and wind may also play a part in the pollination (Ngamriabsakul 2005). Cytological examination showed that species is diploid and possesses 13 pairs of chromosomes (2n = 26) (Ngamriabsakul 2004). Lesser number of chromosomes and the absence of polyploidy reduced the capacity to hinder the gene exchange among populations (Wei and Wu 2012). The breeding system of flowering plant species greatly affects population genetic differentiation (Hamrick and Godt 1989). Similarly, recent studies on the genus Roscoea indicate that the plants of the genus breed through self-pollination under natural conditions (Zhang and Li 2008), which may delay pollination mechanism (Fan and Li 2012). However, it is not clear whether delayed flowering affects reproductive success. Earlier, it has been reported that long-tongued insects, which are considered pollinators of Roscoea, might be absent but the long-lived and self-compatible flowers coupled with the presence of generalist pollinators may reproduce and continue to persist in the alpine habitats (Zhang et al. 2011).
Gene flow was found to be high among the populations that historically represent genetic exchange, and are indicative of current migration rates. For rare and endangered species, it is of greater concern when gene flow estimates are high as they may reflect previous intermingling of populations; therefore, it cannot be interpreted as indication of the current state of population isolation (Godt et al. 2005). The present study estimated higher Nm values between populations, accompanied by high levels of genetic diversity within geographical regions, corresponds with a migrant-pool migratory model, which describes a migratory pattern with colonists recruited from a random sample of previously existing populations (Wade and McCauley 1998). Roscoea procera being a perennial rhizomatous herb maintains a high genetic diversity over the years.
The grouping of populations did not strictly follow the geographical distance in cluster analysis. Some populations such as Pandukholi and Dunagiri, which are geographically closer in Kumaun region of Uttarakhand fall in distant cluster. Wrights (1951) demonstrated that when the gene flow and genetic drift among the populations reach to equilibrium, then only positive correlation between genetic distance and geographical distance occurs. But when such equilibrium is disrupted, positive correlation may not be detected because either gene flow or genetic drift among populations can overwhelmingly influence the population structure (Guo et al. 2011). Thus, high gene flow in the present study might be a reason for insignificant relationship between genetic and geographical distance.
In order to correlate genetic diversity with altitudinal gradient, significant relationship was not found (P < 0.05). Generally, decreasing temperature associated with increasing altitude cause a decline in resource availability (Koo et al. 2015). Moreover, habitat suitability decreased at the upper edge of the altitudinal range and population size of other species was reported to decline (Haln et al. 2012; Jugran et al. 2013). It needs to be added that R. procera is well adapted to high altitude areas as it grows in exposed to drier and cooler environments rather than other plants of family Zingiberaceae. Moreover, unlike some other members of Zingiberaceae, the species has closed leaf sheaths (Spearing 1977). Plants of family Zingiberaceae usually form local populations of various sizes, exhibiting a marked ability for extended local persistence due to perenniality and/or clonality (Körner 2003). The characteristics of fragmented populations have profound consequences on the species genetic patterns, which is crucial to elucidate for the adequate management of endangered populations and species. The potential for altitudinal climatic gradients to influence the distribution of genetic variation within and among plant populations is highly complex and variable among species (Ohsawa and Ide 2008). Higher genetic diversity in higher altitude indicated good adaptability of species in higher elevation thus showing resistance against harsh climatic condition.
Higher diversity in open grassy land and oak forest compared to pine forest indicated its habitat preference and found to be helpful in maintaining diversity among populations. Generally, open grassy land and oak forest supports large number of flowering species (Jugran et al. 2013), providing forage resources for attracting more number of pollinators for cross pollination, which increases the level of genetic diversity (Potts et al. 2003). Thus, ecological niches can be an important tool to understand the process of evolution. Holderegger et al. (2010) focussed the need of addition of some landscape or environmental variable of interest (other geographical distance) at or in between sampling populations to understand micro-evolutionary processes such as gene flow and adaptation on the scale of natural landscapes. In hilly landscape of Himalaya, it becomes more important due to rapid changes in micro-environmental conditions which affect pollen and seed dispersal, and create geographical barrier for gene flow.
The goal of conservation is to ensure sustainable survival of populations and to preserve their evolutionary potential; hence any loss of genetic diversity could lead to decrease in the ability of species to survive under changing environmental conditions and demographical process (Ellstrand and Elam 1993; Milligan et al. 1994). Therefore, information on the extent and distribution of the genetic diversity is important for designing conservation strategies for threatened and endangered species (Hamrick and Godt 1989). High genetic diversity within and high gene flow among populations suggests ex situ conservation of populations like Pandukholi, Mussoorie and Buranskhanda; however, populations with low genetic diversity, such as Majkhali and Thakurh, also need to be conserved to widen the gene pool of the species. High genetic diversity under open grassy land and oak forest suggests that the same can be adopted for in situ conservation as well as understanding the evolutionary processes. Although, the species maintains relatively high level of genetic diversity as revealed by the study, continuous harvest from the wild would certainly be a threat to the target species; hence mass multiplication through conventional and in vitro methods should be carried out so that the propagules can be reintroduced in the wild. Besides, good harvesting practices should also be standardised along with cultivation practices to reduce the pressure on the wild.
High genetic diversity, genetic differentiation and gene flow indicated the need for promotion of in situ and ex situ conservation of R. procera. Among the 11 populations, Surkunda, Pandukholi and Suakholi populations harbour maximum diversity thereby deserving higher priority for in situ conservation. These populations can be utilised for obtaining propagules for cultivation so as to minimise the possibilities of inbreeding depression. The populations with low genetic diversity, i.e., Bhowali, Lamgarha, Ranikhet, Chaubatiya and Mussoorie, deserve equal attention on account of their higher risk of potential local extinction under changed scenario. Such populations, while requiring in situ conservation, would also need strong ex situ backup.
References
Bauert MR, Kalin M, Baltisberger M, Edwards PJ (1998) No genetic variation detected within isolated relic populations of Saxifraga cernua in the Alps using RAPD markers. Mol Ecol 7:1519–1527
Bussell JD (1999) The distribution of random amplified polymorphic DNA (RAPD) diversity amongst populations of Isotoma petraea (lobeliaceae). Mol Ecol 8:775–789
Chaudhary A, Dhar U, Ahmad A, Bhatt ID, Jugran A, Kaur G (2012) Analysis of genetic diversity in Hedychium spicatum Buch. Ham. ex. D. Don in West Himalaya, India. J Med Plant Res 6:3984–3991
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
Ellstrand NC, Elam DR (1993) Population genetic conservation of small population size: implication for plant conservation. Annu Rev Ecol Evol Syst 24:217–242
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Fan YL, Li QJ (2012) Stigmatic fluid aids self-pollination in Roscoea debilis (Zingiberaceae): a new delayed selfing mechanism. Ann Bot 110:969–975
Fenster CB, Armbruster WS, Wilson P, Thomson JD, Dudash MR (2004) Pollination syndromes and floral specialization. Annu Rev Ecol Evol Sys 35:375–403
Godt MJW, Caplow F, Hamrick JL (2005) Allozyme diversity in the federally threatened golden paintbrush, Castilleya levisecta (Scrophulariaceae). Conserv Genet 6:87–99
Goulao L, Valdiviesso T, Santana C, Oliveira CM (2001) Comparison between phenetic characterisation using RAPD and ISSR markers and phenotypic data of cultivated chestnut (Castanea sativa Mill.). Genet Resour Crop Evol 48:329–338
Guo W, Jeong J, Kim Z, Wang R, Kim E, Kim S (2011) Genetic diversity in Lillium tsingtauense in China and Korea revealed by ISSR markers and morphological characters. Biochem Syst Ecol 39:352–360
Haln T, Kettle CJ, Ghazoul J, Frei ER, Matter P, Pluess AR (2012) Pattern of genetic variation across altitude in three plant species of semi-dry grassland. PLoS One 7:e41608. doi:10.1371/journal.pone.0041608
Hamrick JL, Godt MJW (1989) Allozyme diversity in plants. In: Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant population genetics, breeding and genetic resources. Sinauer, Sunderland, pp 43–61
Hamrick JL, Godt MJW (1990) Allozyme diversity in plant species. In: Brown AHD, Clegg MT, Kahler AL, Weir BS (eds) Plant population genetics, breeding, and genetic resources. Sinauer Associates, Sunderland, pp 43–63
Holderegger R, Buehler D, Gugerli F, Manel S (2010) Landscape genetics of plants. Trends Plant Sci 15:175–683
Hu Y, Wang L, Xie W, Yang J, Li Y, Zhang H (2010) Genetic diversity of wild populations of Rheum tanguticum endemic to China as revealed by ISSR analysis. Biochem Syst Ecol 38:264–274
Jugran A, Bhatt ID, Rawat S, Giri L, Rawal RS, Dhar U (2011) Allozyme variation in natural populations of Hedychium spicatum. Biochem Genet 49:806–818
Jugran A, Bhatt ID, Rawal RS, Nandi SK, Pande V (2013) Pattern of morphological and genetic diversity of Valeriana jatamansi Jones in different habitats and altitudinal range of West Himalaya, India. Flora 208:13–21
Kavitha PG, Kiran AG, Raj RD, Sahu M, Thomas G (2010) Amplification fragment length polymorphism analysis unravel a striking difference in the four species of genus Zingiber Boehm. from the Western Ghats, south India. Curr Sci 98:242–247
Koo KA, Patten BC, Madden M (2015) Predicting effect of climate change on habitat suitability of red spruce (Picea rubens Sarg.) in the southern Appalachian mountain of the USA: understanding complex systems mechanism through modeling. Forest 6:1208–1226
Körner C (2003) Alpine plant life: functional plant ecology of high mountain ecosystems. Springer, Berlin
Li JM, Jin ZX (2007) Genetic variation and differentiation in Torreya jackii Chun. an endangered plant endemic to China. Plant Sci 172:1048–1053
Li FG, Xia NH (2005) Population structure and genetic diversity of an endangered species, Glyptostrobus pensilis (Cupressaceae). Bot Bull Acad Sin 46:155–162
Luan S, Chiang TY, Gong X (2006) High genetic diversity vs. low genetic differentiation in Nouelia insignis (Asteraceae), a narrowly distributed and endemic species in China. Revealed by ISSR fingerprinting. Ann Bot 98:583–589
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
McDermott J, McDonald B (1993) Gene flow in plant pathosystems. Annu Rev Phytopathol 31:353–373
Milligan BG, Leebens-Mack J, Strand AE (1994) Conservation genetics: beyond the maintenance of marker diversity. Mol Ecol 12:844–855
Mohandass D, Zhao JL, Xia YM, Campbell MJ (2015) Increasing temperature causes flowering onset time changes of alpine ginger Roscoea in the Central Himalayas. J Asia-Pacific Biodiver 8:191–198
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci USA 70:3321–3323
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89:583–590
Ngamriabsakul C (2004) A chromosomal study of Roscoea and Cautleya (Zingiberaceae): phylogenetic implications. Walailak J Sci Technol 1:70–86
Ngamriabsakul C (2005) Morphological study of the versatile anther group in the tribe Zingibereae (Zingiberaceae). Walailak J Sci Technol 2:11–21
Ngamriabsakul C, Newman MF, Cronk CB (2000) Phylogeny and disjunction in Roscoea (Zingiberaceae). Edinburgh J Bot 57:39–61
Nybom H (2004) Comparison of different nuclear DNA markers for estimating intraspecific genetic diversity in plants. Mol Ecol 13:1143–1155
Ohsawa T, Ide Y (2008) Global patterns of genetic variation in plant species along vertical and horizontal gradients on mountains. Glob Ecol Biogeogr 17:152–163
Peakall R, Smouse PE (2006) GenAlEx6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Perrier X, Jacquemoud-collet JP (2006) DARwin software, version 5.0.158. Department Systems Biologiques (BIOS), CIRAD, Montpellier, France. http://darwin.cirad.fr/darwin
Potts SG, Vulliamy B, Dafni A, Ne’Eman G, Willmer P (2003) Linking bees and flowers: how do floral communities structure pollinator communities? Ecology 84:2628–2642
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rana JC, Sharma BD, Jha BJ, Kumar M (2004) Cultivation of Hedychium spicatum (Kapoor Kachari), Valeriana wallichii and Roscoea purpurea in hill regions of India. Indian For 130:1008–1018
Rawat S, Andola H, Bhatt ID, Giri L, Dhyani P, Jugran A, Rawal RS (2014) Assessment of nutritional and antioxidant potential of selected vitality strengthening Himalayan medicinal plants. Int J Food Prop 17:03–712
Sahu MS, Mail PY, Waikar SB, Rangari VD (2010) Evaluation of immunomodulation potential of ethanolic extract of Roscoea procera rhizomes in mice. J Pharm Bioallied Sci 4:346–349
Slatkin M (1987) Gene flow and the geographic structure of natural population. Nature 236:787–792
Spearing JK (1977) A note on closed leaf-sheaths in Zingiberaceae- Zingiberoideae. Notes R Bot Gard Edinburgh 35:217–220
Wade M, McCauley DE (1998) Extinct and recolonization: their effects on the genetic differentiation of local population. Evolution 43:1349–1368
Wang C, Zhang H, Qian ZQ, Zhao GF (2008) Genetic differentiation in endangered Gynostemma pentaphyllum (Thunb.) Makino based on ISSR polymorphism and its implications for conservation. Biochem Systemat Ecol 36:699–705
Wei L, Wu SJ (2012) Genetic variation and population differentiation in a medicinal herb Houttuynia cordiats in China revealed by inter simple sequence repeats (ISSR). Int J Mol Sci 12:8159–8170
Wright S (1951) The genetic structure of population. Ann Eugen 15:323–354
Xiao LQ, Ge XJ, Gong X, Hao G, Zheng SX (2004) ISSR variation in the endemic and endangered plant Cycas guizhouensis (Cycadaceae). Ann Bot 94:133–138
Zhang ZQ, Li QJ (2008) Autonomous selfing provides reproductive assurance in an alpine ginger Roscoea schneideriana (Zingiberaceae). Ann Bot 102:531–538
Zhang ZQ, Kress WJ, Xie WJ et al (2011) Reproductive biology of two Himalayan alpine gingers (Roscoea spp., Zingiberaceae) in China: pollination syndrome and compensatory floral mechanisms. Plant Biol 13:582–589
Zhao X, Ma Y, Sun S, Wen X, Milne R (2012) High genetic diversity and low differentiation of Michelia coriacea (Magnoliaceae), a critically endangered endemic in southeast Yunnan, China. Int J Mol Sci 13:4396–4411
Acknowledgments
The authors thank the Director, GBPIHED for providing the facilities and encouragement. All colleagues of the Biodiversity Conservation and Management Thematic section are thanked for their constant support and help. Financial supports from the Council of Scientific and Industrial Research New Delhi, the Science and Engineering Research Board DST Govt. of India, New Delhi, and the Uttarakhand Council of Science and Technology Dehradun are gratefully acknowledged.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Rawat, S., Jugran, A.K., Bhatt, I.D. et al. Genetic diversity analysis in natural populations of Roscoea procera Wall. from West Himalaya, India. Braz. J. Bot 39, 621–630 (2016). https://doi.org/10.1007/s40415-016-0260-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40415-016-0260-4