Abstract
This research was conducted to study the genetic diversity in safflower (Carthamus tinctorius L.) using agro-morphological traits and RAPD markers. Sixteen selected lines derived from landraces growing in various agro-climatic regions of Iran along with four exotic genotypes were evaluated in a randomized complete block design with three replications under field conditions. Days to emergence, days to initial flowering, days to flowering, days to maturity, plant height, branches per plant, capitula per plant, seeds per capitulum, 1,000-seed weight, seed yield per plant, seed yield, and reaction to powdery mildew (Leveillula taurica Arnaud) were evaluated in this study. Genetic diversity of the genotypes was assessed by RAPD markers. The results indicated significant differences among genotypes for the agro-morphological traits and clustering based on these traits classified the genotypes into five groups. Analysis of the RAPD markers revealed 15 polymorphic primers out of 50 used primers. Based on RAPD data, the highest genetic similarity was observed between the cultivars of “AC Sunset,” “AC Sterling” from Canada and the lowest relatedness observed between a local breeding line “E2428” and genotype “GE62923” from Germany. Cluster analysis based on RAPD markers and 54% coefficient of similarity divided the genotypes into five distinct groups. Comparing the clusters based on agro-morphological traits with those from molecular markers showed slight similarities. The finding of high genetic variation for agro-morphological traits and polymorphism at DNA level reveal that agronomic traits can be improved by selection programs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Safflower (Carthamus tinctorius L.) is one of the oldest oilseed crops and is widely grown under the hot, dry climate of the Middle East, the center of its origin and diversity (Knowles and Ashri 1995). This crop was first domesticated and grown to use its flowers for fabric dyes, food coloring, and for medicinal purposes (Harlan 1992; Weiss 2000); however, it is currently grown as an oilseed crop worldwide. Safflower has some agronomic advantages such as drought resistance and adaptation to arid and semiarid climatic conditions (Weiss 2000).
The assessment of the genetic diversity in crop species is of interest for the conservation of genetic resources, broadening of the genetic base, and practical applications in breeding programs. Safflower possesses considerable diversity across different regions of the world (Knowles 1989). In order to design an appropriate breeding program, it is important to know how much the phenotypic variation of a trait is heritable (Kearsey and Pooni 1996), since the efficiency of a selection program is mainly dependent on the magnitude of genetic variation and heritability of a trait (Falconer and Mackay 1996).
Genetic diversity of a crop species has been investigated using either agro-morphological traits or molecular markers. Random amplified polymorphic DNA (RAPD) markers (Williams et al. 1990) has provided a powerful tool for the investigation of genetic diversity. The RAPD procedure works with anonymous genomic markers, requires only small amount of DNA, and is simple and less labor than the other DNA markers (Caetano-Anolles et al. 1991).
Genetic diversity of some safflower germplasm has been previously investigated based on the agro-morphological traits (Ashri 1975; Jaradat and Shahid 2006), biochemical traits (Han and Li 1992; Fernandez-Martinez et al. 1993) and molecular markers including isozymes (Han and Li 1992), RAPDs (Maali Amiri et al. 2001; Sehgal and Raina 2005; Vilatersana et al. 2005), ISSRs (Ash et al. 2003; Yu-Xia et al. 2007) and AFLPs (Sehgal and Raina 2005).
The present study was conducted to assess the genetic diversity of different local and exotic genotypes of safflower using both agro-morphological traits and RAPD markers.
Materials and methods
Field experiment
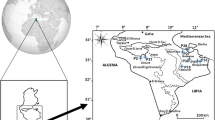
Sixteen breeding lines selected from various local populations of safflower growing in different climatic conditions of Iran along with four genotypes obtained from Canada and Germany were used in this study (Table 1). This experiment was carried out at the research farm of Isfahan University of Technology, Isfahan (32°32′ N and 51°32′ E, 1,630 m asl) on a Typic Haplargid soil with a texture of clay loam, pH 7.5 and organic matter content of 1%, in 2005. A randomized complete block design with three replications was used. Each plot consisted of four rows 50 cm apart and 4 m in length with inter-row plant distance of 10 cm. Plot fertilized with 200 kg N/ha and 200 kg P/ha prior to sowing and 100 kg N/ha was top dressed 30 days after planting.
Days to 50% emergence, initial flowering, 50% flowering and maturity, and also reaction to powdery mildew (Leveillula taurica Arnaud) were visually recorded for each plot. The disease severity was scored based on a 0–9 scale in which 0 = no symptoms, 1 = trace to 2%, 2 = 2–4%, 3 = 4–7%, 4 = 7–15%, 5 = 15–35%, 6 = 35–65%, 7 = 65–85%, 8 = 85–95%, and 9 = >95% leaf area infected, respectively. The disease developed naturally in field plots, starting in mid-season (mid-June) and progressively worsened toward maturity. In addition, plant height and yield components including branches per plant, capitula per plant, seeds per capitula, 1,000-seed weight, and seed yield per plant were recorded on ten plants per plot and their average were used. Two middle rows of each plot was harvested to determine the seed yield.
RAPD analysis
A bulked genomic DNA was extracted from the young leaves (Murray and Thampson 1998) of ten plants per genotypes and quantified electrophoretically using lambda standard DNA (GeneCraft, Munster, Germany) on 0.8% agarose gels. Fifty ten-mer arbitrary primers from A, B, C, H, K, and I series of Operan Technologies were used for PCR reaction.
The PCR reaction was performed in a 25 μl volume using a Techgen PCR thermocycler (Biometra, Germany). The reaction mixture contained: 2.5 μl 10× buffer, 0.2 mM dNTPs, 2 mM MgCl2, 20 ng template DNA, 0.8 μM primer and 1 U Taq DNA polymerase (Biotherm, Germany). DNA thermal cycler programmed to 1 cycle of 5 min at 94°C (denaturation), 1 min at 36°C (annealing), and 1 min at 72°C (extension); followed by 40 cycles of 1 min at 94°C, 1 min at 36°C, and 1 min at 72°C ending with 1 cycle of 5 min at 72°C (final extension).
PCR amplified products were separated by electrophoresis in 1.2% agaros gels using Biometra Model S2 gel electrophoresis equipped with Biometra Model PS9009TC power supply at 50 W for 2 h in 1× TBE buffer. Gels were stained with ethidium bromide (0.5 μg/ml). DNA banding patterns were visualized using Biometra Gel Documentation Model P755874X02.
Data analysis
Agro-morphological traits
Analysis of variance (ANOVA) of data for agro-morphological traits was performed using General Linear Model of SAS program (SAS Institute 1997). The data of powdery mildew disease was square root transformed before subjected to ANOVA. Agro-morphological data was used to generate a genetic similarity matrix with SIMQUAL routine Dice from the NTSYS-pc statistical package (Rohlf 1992). The genetic similarity matrix was transferred into distance matrix using TRANSFER subroutine from NTSYS-pc, resulting in distance matrix equivalent to that of Nei and Li (1979). Clustering of genotypes based on the agro-morphological traits was performed by the SAHN in NTSYS-pc and the genetic distance matrix using unweighted pair group method with arithmetic mean (UPGMA) (Rohlf 1992).
RAPD markers
Amplification products were scored for the presence (1) or absence (0) of bands and binary matrix was assembled for RAPD markers. The binary matrix was subjected to statistical analyses using NTSYS-pc (Rohlf 1992). Dice’s similarity coefficient was employed to compute pairwise genetic similarities. The corresponding dendrogram was constructed by applying UPGMA clustering algorithm (Rohlf 1992). To check the goodness of fit of a cluster analysis to the associated similarity matrix, co-phenetic correlation was computed (Rohlf 1992).
Number of polymorphic bands, percentage of polymorphism and polymorphic information content (PIC) were calculated for each primer (Anderson et al. 1993). Three dimension plots showing the relationships among the genotypes were generated by principal coordinate analysis (PCoA) (Rohlf 1972).
Analysis of molecular variance (AMOVA) was performed to estimate variance components for RAPD data, partitioning the variation into within and among geographical areas including local regions of Isfahan, Hamedan, Kordestan, Markazi, and Khorasan, and countries of Canada and Germany, using Arliquin2.0 software (Excoffier et al. 1992).
Combination of agro-morphological and RAPD data
The agro-morphological and RAPD data were transformed to a single matrix of similarity coefficient using the Dice method (Nei and Li 1979). The resulting similarities between lines were visualized in dendrogram using the UPGMA clustering algorithm.
To investigate the association between genetic and geographical distances, interpoint distances between genotypes were calculated in kilometers and then the relationships between two matrices were assessed using the Mantel test (Mantel 1967).
Results
Agro-morphological traits
The results of ANOVA showed that genotypes significantly differed for agro-morphological traits (Table 2). Based on genetic coefficient of variations, relatively higher range of genetic variation was observed for days to 50% emergence, seeds per capitula, seed yield, and seed yield per plant (Table 3). The highest and the lowest genetic variation were observed for seeds per capitulum and days to maturity, respectively. Seeds per capitulum varied from 12.3 to 27.8 and days to maturity ranged from 100.7 to 127.4 days. Plant height, seed yield per plant, and seed yield ranged from 57.1 to 135.8 cm, 7.2 to 16.4 g, and 1,088 to 2,168 kg/ha, respectively. Small difference between phenotypic and genetic coefficient of variations and high broad-sense heritabilities for most of the traits indicated that most of the observed variation was due to genetic factors (Table 3). The lowest and highest broad-sense heritabilities were estimated for days to 50% emergence (69.3%) and days to 50% flowering (96%), respectively (Table 3). The heritability estimates for economical traits of seed yield (83%) and seed yield per plant (82.9%) were high. The reaction of safflower genotypes to powdery mildew was narrowly varied ranging from 1.6 to 2.6.
Clustering based on agro-morphological traits clustered the genotypes into five main groups including 4, 4, 8, 3, 1 genotypes, respectively (Fig. 1). Cluster 1 included GE62923, GE62916 genotypes from Germany and AC Sunset and AC Sterling cultivars from Canada. Breeding lines C128, C121, C4110, C111 derived from Kouseh landrace were located in cluster 2, and breeding lines of A3 derived from Azarbayjan landrace, M420 from Markazi landrace, S411 and S3110 from Khorasan landrace, H21 from Hamedan, and E2428, E2417, E1431 from Isfahan landrace were grouped in cluster 3. Cluster 4 possessed breeding lines H27 from Hamedan landrace and K21, K15 from Kordestan landrace. Cluster 5 included breeding line M112 selected from Markazi landrace.
RAPD markers
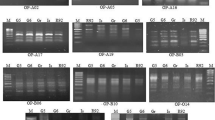
Out of 50 used primers, 15 showed polymorphism and generated a total of 132 polymorphic bands (Table 4). The number of polymorphic amplified products ranged from 3 for primers OPH07, OPH10, and OPI04 to 16 for primer OPC08. PIC ranged from 0.26 to 0.71, indicating that the least informative primer was OPI04 and the most was OPC08 (Table 4). An example of an RAPD pattern produced using the OPC05 primer is presented in Fig. 2.
RAPD agarose gel profile of 20 safflower genotypes using primer OPC05. M: 100 bp molecular size ladder; numbers correspond to the genotypic number listed in Table 1
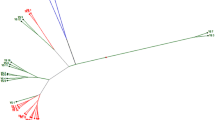
Coefficient of similarities based on RAPD data ranged from 0.3 to 0.89. The lowest genetic similarity was observed between local breeding line E2428 (from Isfahan landrace) and GE62923 genotype from Germany, and the highest one belongs to AC Sunset and AC Sterling cultivars from Canada (Table 5). Grouping the genotypes based on RAPD data revealed five main clusters (Fig. 3). Cluster 1 comprised E2428 and E1431 breeding lines (from Isfahan landrace), AC Sunset, and AC Sterling cultivars (from Canada), cluster 2 possessed breeding lines C121, C128, and C111 (from Kouseh landrace), K15 breeding line (from Kordestan landrace), E2417 breeding line (from Isfahan landrace), M112 and M420 breeding lines (from Markazi landrace), S3110 and S411 breeding lines (from Kordestan landrace), A3 breeding lines (from Azarbayjan landrace), and GE62916 genotype (from Germany). Cluster 3 included breeding lines H21, H27 (from Hamedan landrace), K21 (from Kordestan landrace); clusters 4 and 5 contained C4110 breeding line (from Kouseh landrace) and GE62923 genotype (from Germany), respectively.
To display the relationships among the safflower genotypes in term of their position relative to three coordinate axes, PCoA was performed. The first three components derived from this analysis explained 78% of the variation (Table 6). The three-dimensional plot also showed that the Iranian and exotic genotypes were clustered in the same groups (Fig. 4).
AMOVA for RAPD data indicated that there were no significant differences among the geographical regions in which the used genotypes were derived (data not shown).
The relationship between the genetic distances estimated based on agro-morphological traits and molecular markers was not statistically significant (r = 0.47). Comparing the clusters based on agro-morphological traits with those from molecular markers indicated that there was a slight similarity between these clustering.
Clustering based on combination of agro-morphological traits and RAPDs
The results of clustering based on combination agro-morphological and RAPD data grouped the genotypes into four distinct clusters (Fig. 5) which was mostly similar to that of the RAPDs, except that breeding lines K15 and C4110 were grouped in different clusters. The Mantel test suggested that genetic distances between genotypes were not correlated with geographic distances (r = 0.34, P = 0.06).
Discussion
There was a wide range of variation among genotypes for plant height and days to maturity. Broad-sense heritability estimates for these traits were also relatively high which in turn indicate that they can be easily improved in breeding programs. Safflower cultivars with lower plant height facilitate mechanical harvesting (Weiss 2000). The early maturing cultivars have certain advantages that foster their use, such as early harvesting before damage of biotic and abiotic stresses, to cultivate the crop in a short growing season, or to facilitate planting of the succeeding crop in a rotation system.
Number of branches and capitula per plants, seeds per capitula, and 1,000-seed weight showed high levels of variability and relatively high broad-sense heritabilities. High broad-sense heritability for number of branches per plant in this study was in agreement with that of Kavani et al. (2000). These results indicate that selection can be effectively employed for improvement these yield components. Increasing in branches and capitula per plant as the most important yield components in safflower can be used as the selection criteria to enhance the seed yield (Ashri et al. 1974; Able and Driscoll 1976; Lahane et al. 1999). However, based on the results of present study, direct selection for seed yield can be also effective, since a wide range of genetic variation and high broad-sense heritability was observed for this trait. Improvement of seed yield is the main objective of breeding programs in safflower.
Powdery mildew caused by the fungus L. taurica Arnaud (Braun 1987) is one of the major diseases of safflower in the world and can be severely damage the crop in dry regions (Holiday 1980). Most of the safflower genotypes had low reaction to powdery mildew implying that they can be used in breeding programs (Jan 1988).
Dendrogram based on the combination of agro-morphological and RAPDs was more similar to that obtained by RAPD data. The high degree of polymorphism detected by RAPD marker and the possibility of screening a higher number of anonymous loci imply that this marker can efficiently classify safflower germplasm.
Despite of differences between two dendrograms obtained by either agro-morphological or RAPD data, A3, S3110, S411, and M420 were classified into same clusters in both dendrograms, indicating of their close relationship. The genetic relatedness between S3110 and S411 was expected, since they were derived from the same population. Observation of no significant difference among the geographical regions based on AMOVA for RAPD data and the clustering pattern of genotypes revealed that genetic and geographic distances were not related. These results were confirmed by the Mantel test showing a low coefficient of correlation between genetic and geographic distances. This can be due to the exchange of plant materials across the regions during the history of safflower cultivation.
References
Able GH, Driscoll MF (1976) Sequential trait development and breeding for high yield in safflower. Crop Sci 16:213–216
Anderson JA, Churchill GA, Autroque JE, Tanksley SD, Swells ME (1993) Optimising selection for plant linkage map. Genome 36:181–186
Ash GJ, Raman R, Crump NS (2003) An investigation of genetic variation in Carthamus lanatus in New South Wales, Australia, using intersimple sequence repeats (ISSR) analysis. Weed Res 43:208–213
Ashri A (1975) Evaluation of the germplasm collection of safflower Carthamus tinctorius L. V: Distribution and regional divergence for morphological characters. Euphytica 24:651–659
Ashri A, Zimmer DE, Urie AL, Cahaner A, Marani A (1974) Evaluation of the world collection of safflower (Carthamus tinctorius L). IV. Yield and yield components and their relationship. Crop Sci 14:799–802
Braun U (1987) A monograph of the Erysiphales (Powdery mildews). Beihefte zur Nova Hedwigia 89. J. Cramer, Berlin
Caetano-Anolles G, Bassam BJ, Gresshoff PM (1991) DNA amplification fingerprinting using very short arbitrary oligonucleotide primers. Biotech 9:553–556
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction sites. Genetics 131:479–491
Falconer DS, Mackay TFC (1996) Introduction to quantitative genetics. Longman, Harlow, UK
Fernandez-Martinez J, Rio M, Haro A (1993) Survey of safflower (Carthamus tinctorius L) germplasm for variants in fatty acid composition and other seed characters. Euphytica 19:115–122
Han Y, Li D (1992) Evaluation of safflower (Carthamus tinctorius L.) germplasm-analysis in fatty acid composition of seeds of domestic and exotic safflower varieties. Bot Res 6:28–35
Harlan J (1992) Crops and man, 2nd edn. ASA, CSSA, Inc., Madison, WI, USA
Holiday P (1980) Fungus diseases of tropical crops. Cambridge Univ. Press, Cambridge, UK
Jan CC (1988) Registration of a powdery mildew resistant sunflower germ plasm pool, PM1. Crop Sci 28:1040
Jaradat AA, Shahid M (2006) Patterns of phenotypic variation in a germplasm collection of Carthamus tinctorius L. from the Middle East. Euphytica 53:225–244
Kavani RH, Shukla PT, Madariya RB (2000) Analysis of variability for seed yield and related characters in safflower (Carthamus tinctorius L.). Madras Agric J 87:449–452
Kearsey MJ, Pooni HS (1996) The genetical analysis of quantitative traits. Chapman and Hall, London
Knowles PF (1989) Centers of plant diversity and conservation of crop germplasm: safflower. Econ Bot 23:324–329
Knowles PF, Ashri A (1995) Safflower Carthamus tinctorius (Compositae). In: Smartt J, Simmonds NW (eds) Evolution of crop plants, 2nd edn. Longman Scientific and Technical, London, pp 47–50
Lahane PS, Mukewar AM, Zope JS, Kalpande HV, Kalpande VV (1999) Genetic variability for different traits in safflower (Carthamus tinctorius L.). J Soil Crops 9:130–132
Maali Amiri RM, Yazdi-Samadi B, Ghannadha MR, Abd-Mishani C (2001) Detection of DNA polymorphism in landrace populations of safflower in Iran using RAPD-PCR technique. Iran J Agri Sci 32:737–745
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Murray MG, Thampson WF (1998) Rapid isolation of high molecular weight plant DNA. Nucleic Acid Res 8:4321–4325
Nei M, Li WH (1979) Mathematical model for studying in genetic variation. Proc Natl Acad Sci USA 76:5269–5273
Rohlf FJ (1972) An empirical comparison of three ordination techniques in numerical taxonomy. Syst Zool 21:271–280
Rohlf FJ (1992) NTSYS-PC: numerical taxonomy and multivariate analysis system version 2.02 k. State University of New York, Stony Brook, NY
SAS Institute (1997) SAS/STAT Software: changes and enhancements through Release 6.12. SAS Institute Inc., Cary, NC
Sehgal D, Raina SN (2005) Genotyping safflower (Carthamus tinctorius) cultivars by DNA fingerprints. Euphytica 146:67–76
Vilatersana R, Garnatje T, Susanna A, Garcia-Jacas N (2005) Taxanomic problems in Carthamus (Asteraceae): RAPD markers and sectional classification. J Bot Linn Soc 147:375–385
Weiss EA (2000) Oil Seed Crops. Blackwell Science Ltd, Oxford, UK
Williams JGK, Kubelik AR, Livak KJ, Rafaleski JA, Tingey SV (1990) DNA Polymorphism amplified by arbitrary primers are useful as genetic markers. Nucleic Acids Res 18:6531–6535
Yu-Xia Y, Wei W, You-Liang Z, Li C, Ren-Jian L, Chun-Yan H (2007) Genetic diversity and relationships among safflower (Carthamus tinctorius L.) analyzed by inter-simple sequence repeats (ISSRs). Genet Res Crop Evol 54:1043–1051
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Amini, F., Saeidi, G. & Arzani, A. Study of genetic diversity in safflower genotypes using agro-morphological traits and RAPD markers. Euphytica 163, 21–30 (2008). https://doi.org/10.1007/s10681-007-9556-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10681-007-9556-6