Abstract
Knowledge of genetic diversity and relatedness of cultivars; is essential for a successful breeding scheme. The present study was conducted to assess the genetic diversity of 30 Iranian Chrysanthemum morifolium cultivars using sequence-related amplified polymorphism (SRAP) molecular markers and morphological traits. For this purpose, 30 cultivars were planted in a completely randomized block design with three replications at the Research Farm of Isfahan University of Technology in 2015. The evaluation of morphological traits during the two consecutive years showed a wide range of variations among different cultivars. The results obtained by the analysis of variance also revealed the significant effects of cultivar, environment and cultivar × environment interaction on all studied traits. Correlation analysis of variables further showed that flowering durability was affected by plant morphology. Results of cluster analysis based on the phenotypic data divided the cultivars into six groups. Molecular analysis using SRAP markers produced a total of 213 identifiable fragments with an average of 21.3 polymorphic bands per combination. The polymorphic information content (PIC) value was 0.46. Along with the Bayesian structure analysis which showed that the highest value of the maximum delta log likelihood (ΔK) was K = 3, cluster analysis also grouped the cultivars into three main clusters. The high value of admixture revealed ancestral relationships between the studied cultivars. Overall, our findings revealed the potential of SRAP markers and morphological traits for the analysis of genetic diversity in Chrysanthemum. This can be, in turn, used in selecting the desired parents in the hybrid breeding programs as well as developing superior cultivars for ornamental applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Genetic diversity is crucial for the successful exploitation of novel traits in commercial ornamental plants. In traditional breeding, the recognition of ornamental plant cultivars has been based on morphological parameters as a common index (Fu et al. 2008). DNA markers have widely been employed in characterizing genetic resources; generally considered as fingerprinting, these markers contribute to assessing the genetic diversity and supplying the knowledge necessary for ecology, evolution, and population genetics (Arzani and Ashraf 2016). The use of molecular markers along with morphological traits can provide new insights useful for further breeding programs in ornamental plants (Martí et al. 2002; Da Silva 2004; Hassanein and Al-Soqeer 2017). Among molecular markers, sequence-related amplified polymorphism (SRAP) is a novel genetic marker system recognized as an effective marker in cultivars identification and analysis. This marker, which serves as a PCR-based technique, amplifies open reading frames (ORF) (Mokhtari et al. 2013).
Chrysanthemum (Chrysanthemum morifolium L.), as a short-day and perennial herbaceous member from the Asteraceae family (Roein et al. 2015), is one of the most popular and economically important ornamental and medicinal species in Asian and European countries (Zhang et al. 2011a). The breeding of perennial and heterozygous crops depends mostly on individual selection and hybridization (Yao et al. 2008). Today, different cultivars of Chrysanthemum have been obtained in different countries through different methods, especially by hybridization. Theses cultivars exhibit a wide range of diversity in their morphological traits. Hence, evaluation and identification of valuable cultivars and their cultivation can be of great importance for the ornamental purposes. Interestingly, most Chrysanthemum cultivars look like each other in terms of the morphological traits. So, more accurate evaluation of the cultivars based on DNA molecular markers seems to be necessary. There are some reports on the genetic variation of C. morifolium based on different molecular tools and different countries; these include ISSR, SRAP, RAPD and AFLP markers from India, China, Pakistan and Iran, respectively (Palai and Rout 2011; Shao et al. 2010; Naz et al. 2015; Roein et al. 2014).
In Iran, many C. morifolium cultivars with different flower colors have been introduced; however, not much information is available on their genetic diversity. SRAP markers have been used to characterize Chinese medicinal Chrysanthemum (Shao et al. 2010), but no similar studies have been reported on the genetic variation of the Iranian Chrysanthemum cultivars. In addition, there is still comparatively little work on using molecular markers to discriminate desirable plant types to improve plants. Furthermore, most of the previous studies on the variations of C. morifolium cultivars have been carried out regardless of the environmental effects (E) and their interactions with the genotype (G), G × E (Shao et al. 2010; Banerji et al. 2012; Roein et al. 2014). Therefore, the present study was aimed to investigate the morphological variation in C. morifolium during two constitutive years and its relationship with the molecular markers data.
In the light of the mentioned information, the present study was carried out (1) to evaluate the morphological characteristics of different Chrysanthemum cultivars including vegetative and reproductive characters in 2 years, (2) to assess the extent of genetic diversity and the population structure of Chrysanthemum cultivars by using SRAP markers, and (3) to explore the suitable parents for targeted hybridization in Chrysanthemum.
2 Materials and methods
2.1 Plant materials
Thirty cultivars of Chrysanthemum (Chrysanthemum morifolium Ramat.), which had been provided by the Iranian Research Center for Ornamental Plants, Mahallat, Iran, were used in this study (Table 1). Stock plants were grown at the research farm of Isfahan University of Technology during two constitutive years, including 2015 and 2016, using a completely randomized block design with three replications.
2.2 Phenotypic evaluation
The agro-phenological and morphological characteristics were recorded for the germplasm. The following phenotypic traits were evaluated: plant height, flower head diameter, flower dry weight, flower fresh weight, number of flowers per plant, yield of flowers per plant, diameter of the plant canopy, days for the color appearance of the flower bud, and flowering durability (days to 15% flower senescence).
2.3 DNA extraction and PCR amplification
Genomic DNA was extracted using the young leaves of Chrysanthemum cultivars according to the modified Murray and Thompson’s (1980) method. The quality of the extracted DNA was estimated using agarose gel electrophoresis and staining with ethidium bromide.
A set of ten different SRAP primer (Table 2) combinations were employed for the analysis of Chrysanthemum samples (Ghafouri and Rahimmalek 2018; Talebi et al. 2015). DNA amplification was performed in a 10 μl reaction mixture. PCR mixture consisted of 1 μl genomic DNA, 5 μl of the Master Mix Red (Ampliqon, Finland), 1 μl of each primer (0.5 μM), and distilled water to the final volume. PCR amplification program included 5 min of the initial denaturation at 94 °C, 5 cycles during 1 min at 94 °C, 1 min annealing at 35 °C, and 1 min elongation at 72 °C; these were followed by 35 cycles during 1 min at 94 °C, 1 min at 50 °C, 1 min at 72 °C, and a final stage of 5 min at 72 °C. Separation of DNA amplified fragments was performed on 12% acrylamide gels [acrylamide: bisacrylamide (29:1), 1 × TBE] at 230 V for 2.5 h. Then, the gels were stained with silver nitrate.
2.4 Data analysis
The morphological data obtained during 2 years were subjected to the analysis of variance (ANOVA) using a split-plot completely randomized block design with the SAS v9 software (SAS institute, 2002). Mean comparisons were performed using the Fisher’s (protected) least significant difference (LSD, p ≤ 0.05) test. The genotypic and phenotypic variances as well as heritability were calculated for the phenotypic traits by the following formula:
MSG refers to genetic mean squares, MES is the environmental mean squares, \(\sigma_{g}^{2}\) is the genetic variance, \(\sigma_{p}^{2}\) is the phenotypic variance, GCV refers to the genotypic coefficients of variation, and PCV represents the phenotypic coefficients of variation.
The cultivars were clustered for all morphological traits by the STATGRAPHICS, ver.6, statistical package using the Ward method. Pearson’s correlation coefficient was calculated using SPSS (version16; SPSS Inc. Chicago, IL, USA).
For each SRAP marker, the presence (1) or absence (0) of the bands was scored. To estimate the number of genetically distinct clusters (K), the STRUCTURE 2.3.4. program was used. Analysis was conducted using Markov chain Monte Carlo from 5000 to 50,000 repeats (MCMC), which were correlated with the allele frequencies (K = 2–10). Cluster analysis was conducted based on the Neighbor-Joining method using MEGA, version 4.0. Polymorphic information content (PIC) was calculated according to the formula: PIC = 1−∑P2ij, where Pij is the frequency of the jth allele for the ith marker. The correlation between the morphological traits and the molecular markers was estimated using the Mantel’s test. The cophenetic correlation coefficient was also obtained using the COPH routine to check the goodness of fit between the clustering and the similarity coefficient matrix.
3 Results
3.1 Morphological evaluation
The results of the 2-year analysis of variance (ANOVA) revealed the significant effect of cultivar, year and cultivar × year interactions on all studied morphological traits (Table 3). The highest phenotypic (PCV) and genotypic (GCV) coefficients of variation were attributed to the number of flowers per plant in both years (i.e., > 57%). Among the studied traits, the highest heritabilities were obtained for the flower fresh weight and the days to color appearance of the flower bud, while flower yield recorded the lowest heritability (Table 4).
During the first year, cultivars Mehrnoosh2 (100 cm), Afsaneh (28 g), Romina (184 days) and Taraneh (38.66 days) exhibited the highest values for the canopy diameter, flower dry weight, days to color appearance of the flower bud and flowering durability, respectively; on the other hand, in the second year, the highest values of these traits were recorded for the cultivars Tarannom2 (83.33 cm), Armita2 (24 g), Poya3 (189.33 days), and Afsaneh (37.66 days). In 2015, the plant height varied from 22 cm (Poya3) to 89.66 cm (Tarannom2) with an average value of 46.82, while in 2016, it ranged from 17.66 cm (Taraneh) to 112.66 cm (Tarannom2), with the average value being 49.57 cm. Flower head diameter varied from 2.9 in the cultivar Romina to 5.3 in the cultivar Ashna. The highest flower yield (245.19 g) and number of flowers (521.67) in both years belonged to the cultivar Shokoh. Also, the highest fresh weight was recorded for the cultivar Taraneh with an average value of 148.90 g (Table 5).
3.2 Correlation analysis of the morphological traits
Correlation analysis was conducted for all morphological traits obtained in 2 years (Table 6). The highest positive and significant correlation was observed between the fresh and dry weight of flower (r = + 0.924), as well as between flower head diameter and flower dry weight (r = + 0.753); these were followed by canopy diameter and plant height (r = + 0.665). Moreover, a negative correlation was found between the duration of flowering and days to flowering (r = − 0.753). In addition, flowering durability established significant correlations with flower dry weight (r = + 0.52**), flower head diameter (r = + 0.38*), days to color appearance of the flower bud (r = − 0.75**), and plant height (r = − 0.43*).
3.3 Hierarchical cluster analysis of morphological traits
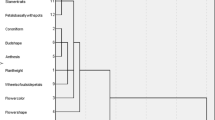
Cluster analysis was performed based on the morphological traits obtained in 2 years; all the 30 studied cultivars were divided into six groups using the Ward method (Fig. 1). Cluster 1 was characterized mostly by small flowers, the least vase life and flower weight, but plant height, flower yield, and flower number were high. Cluster 2 consisted of seven cultivars with the lowest plant height, canopy diameter, and flower yield. The cultivars in the third group with the medium flower durability, late flowering time, and low flower yield were classified in two separate subgroups based on their flower diameter. Having the large flowers was characteristic of the first subgroup of the third main group. Cluster 4 was characterized by the lower number of flowers, as well as the large flower head diameter and long flowering durability. The cultivar Taraneh in this group with the lowest plant height was placed in a separate subgroup. Cluster 5 comprised the cultivars Hour and Fariba2 with the earliest flowering times. Finally, Cluster 6 was characterized by the early flowering time and appropriate flowering durability. Whenever the purpose of breeding is the production of cultivars for cut flower applications, the cultivars with the higher height, attractive colors, large flowers and high vase life are suitable for selection. Therefore, the crossing of the first group with the fourth and fifth groups could be effective. If the purpose is to produce pot or garden cultivars, having features such as lower height and the high number of flowers per plant can be effective. The probability of reaching this goal with the crossing of the cultivar Taraneh, which had small shrubs and large flowers, was like that with the cultivars for the first group, such as Shokoh, which had many flowers.
3.4 SRAP amplification and levels of polymorphism
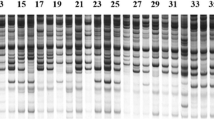
Ten primer combinations were selected based on clarity, sharpness, and number of bands in a prescreening assay performed by 20 primer combinations on 30 cultivars. A total of 222 identifiable fragments were obtained; out of these, 213 bands (95.94%) were polymorphic, ranging from 11 to 33 with an average of 21.3 polymorphic bands per combination. The Shannon’s information index (I) and the Nei’s gene diversity index (H) were calculated from this analysis to be 0.53 and 0.35, respectively. Shao et al. (2010) reported lower values (I: 0.34, H: 0.21) for the Chinese Chrysanthemum cultivars in comparison with the Iranian ones. The average polymorphism information content (PIC) value was 0.46 (Table 7). The highest (0.496) and lowest values (0.433) of the PIC were attributed to Me2-Em3 and Me5-Em2 primer combinations. Figure 2 shows the pattern of Me4-Em3 SRAP primer combination in different cultivars.
3.5 Phylogenetic relationships and principal coordinate analysis (PCoA) of the molecular data
Cluster analysis of the molecular data based on the NJ method divided the cultivars into three major groups (Fig. 3). The first group included 11 individuals, the cluster II had eight cultivars, and eleven individuals were assigned to the cluster III.
Principal coordinate analysis (PCoA) was conducted to ensure a better view of the genetic relationships in Chrysanthemum cultivars. According to the PCoA analysis, the first three principal coordinates accounted for 17.78% of the total variation, such that the contribution of the first component was 8.89% and those of the second and third components were 4.63% and 4.24% of the total variation respectively (Fig. 4). According to Mohammadi and Prasanna (2003), cluster analysis could be more sensitive for the detection of germplasm relationships than PCoA, as the first two or three principles explained less than 25% of the total variation.
The Bayesian approach for clustering information from SRAP primer combinations revealed the highest value of ΔK as K = 3. Genetic admixture was observed among the three clusters, which could be related to the amount of cross pollination and the level of heterozygosity of Chrysanthemum. According to a membership value of 0.75, the STRUCTURE analysis divided 15 cultivars into three clusters (Fig. 5). Cultivars with membership probabilities less than 0.75 were attributed to an admixture group. Cluster a included four cultivars (20, 21, 18, 11), cluster b consisted of six cultivars (26, 31, 14, 30, 13, 25), and cluster c comprised five cultivars (5, 10, 1, 6, 7). Clustering according to structure analysis was in line with the clustering-based NJ method. The diversity parameters in each cluster were calculated (Table 8). Among the three groups, the admixed cluster showed the highest diversity parameters (H and PIC), and the cluster a exhibited the lowest ones. Also, based on the genetic similarity index, genetic distances ranged from 0.202 in Ashna and Hour to 0.545 in Fariba2 and Sahel with a mean of 0.370.
Genetic structure of 30 Chrysanthemum cultivars as inferred by using structure software with ten SRAP marker data sets and K = 3 clusters. a Red cluster; b green cluster; and c blue cluster. The X and Y axis shows the cultivars and the proportion membership of each individual into various clusters, respectively
4 Discussion
Given that biodiversity is fundamental to the breeders’ programs and there is a wealth of empirical evidence regarding the benefits of plant genetic resources in developing improved plant cultivars, biotechnological approaches could substantively enrich this use (Arzani and Ashraf 2016). The results obtained from the phenotypic data revealed a wide range of variation among all studied cultivars. In agreement with our findings, previous studies reported a high phenotypic variation among the Indian and Chinese Chrysanthemum cultivars (Kulkarni and Reddy 2004; Shao et al. 2010).
According to the analysis of variance in two constitutive years, the significant interaction effect between the genotypic and environmental factors as well as higher PCV values against the GCV values reflected the important effect of environmental conditions on the morphological traits among the Chrysanthemum plants. Different patterns of temperature and rainfall during the 2 years of field experiments led to a high level of phenotypic variation for all traits. The lower amount of rainfall in the second year rather than the first one (the monthly rainfall average in 2016 ~ 29.1 < 49.1 ~ in 2015 cm) might have reduced the average of flower head and canopy diameter, flower number and yield, and flower durability in the second year. Flowering time and vegetative period are important factors in selecting flowers and introducing new cultivars. The cultivars Fariba2 and Hour were early flowering, while Romina and Poya3 were recorded as the latest flowering cultivars.
Furthermore, in the ornamental plants, freshness and flower durability are critical traits for the selection of the superior cultivars. In the present study, the cultivars Taraneh, Atashgoon and Fariba2 had the highest duration of flowering (> 33d). The tallest cultivars were Tarannome2 and khalijfars2, while the shortest one was Taraneh during both years. The effect of genotypic variation on the plant height has also been reported by Ahmad et al. (2012). The dwarf and compact Chrysanthemum cultivars are suitable for pot cultures, while the erect and tall growing ones are suitable for use as cut flowers. Therefore, among the studied cultivars, “Taraneh” was found to be the best one for pot culture due to its maximum vase life and minimum plant height; also, the cultivars “Tarannome2”, “Khalijfars2”, “Ashna”, “Shokoh”, “Azin2”, “Sahar” and “Marmar” could be ideal for applying as cut flowers. Gaikwad et al. (2002) also recommended nine elite Indian Chrysanthemum cultivars as cut flowers based on some morphological traits including flowering time, yield and vase life.
Correlation analyses also showed that by increasing the vegetative stage, the flowering period of the plant, which was responsible for its ornamental value, was decreased. The results of the correlation analysis revealed that the cultivars with larger flower diameter had a higher fresh and dry weight, less flower numbers, and a higher flowering durability. Also, it could be deduced that the cultivars with the higher plant height, in addition to canopy diameter, had more flowers and less flowering durability. Heritability, as an index estimating the reliability of phenotypic values, is necessary for effective breeding programs (Falconer et al. 1996). The results showed the high percentage of heritability for the days to color appearance of the flower bud, flower fresh weight, flower dry weight and plant height, which revealed the potential of these traits for the genotypic selection of Chrysanthemum germplasm. Similar observations have been made in the case of blooming time in Chrysanthemum (Roein et al. 2014; Zhang et al. 2011b). The low heritability was recorded for flowering durability, thereby suggesting the indirect selection based on the flower dry weight and days to color appearance of the flower bud, which had high heritabilities and high correlation with flowering durability, would be more effective. Selection of parents could be useful in wide crossing for developing different hybrid combinations and new Chrysanthemum cultivars. According to the morphological data, the highest Euclidean distance was observed between the cultivars “Taraneh” and “Shokoh” (9.15), as well as between the cultivars Tarannom2 and Forogh (7.95), while the lowest one was observed between the cultivars Mitra and Ziba2 (0.76). So, the more diverse cultivars could be introduced as the parental materials.
Previous studies have reported the potential of SRAP markers in cultivar identification (Shao et al. 2010). In the present research, a large number of polymorphic bands assessed by applying SRAP markers confirmed that SRAP analysis could be useful for identifying various Chrysanthemum cultivars. In the present research, the average percentage of polymorphic bands (96%) was higher than that reported in the Chinese Chrysanthemum cultivars using SRAP (81.87%) and AFLP (93%) markers (Shao et al. 2010; Han et al. 2007). Accordingly, 213 polymorphic bands were produced as it was in range of many previously published reports for the SRAP markers (Talebi et al. 2015; Aneja et al. 2013; Shao et al. 2010). The PIC value is often used to determine the degree of gene diversity among different cultivars (Feng et al. 2016). According to Xie et al. (2010), the applied SRAP primers revealed moderate polymorphism among the studied cultivars (0.25 < PIC < 0.5), which was suitable for the analysis of genetic variation in the studied cultivars.
In the present study, less than 25% of the total variation was explained with the first three principal components, which showed that there was a low correlation between the primer combinations and markers distribution across the genome. So, the results of PCoA did not confirm the clustering analysis.
Subgroups showed a membership value of the cultivars that was higher than 0.75, suggesting the presence of admixture. According to STRUCTURE software analysis, the mixed population ancestry was observed among the separate groups. It could be the result of the incorporation of more than a single gene pool as well as gene flow between different cultivars during hybridization programs (Soriano et al. 2016). The admixture models for the individual cultivars could also provide useful information regarding the hybrid origins (Chen et al. 2008). SRAP phylogenies showed agreement with the previous molecular phylogenetic data explaining the hybrid lineage among Vitis varieties and the related species (Robarts and Wolfe 2014). Admixtures have also been reported in the Iranian and Indian Chrysanthemum cultivars based on AFLP and RAPD markers, respectively (Roein et al. 2014). In the present study, twenty-three out of 30 cultivars showed some degree of admixture. Fifteen cultivars with membership probabilities > 0.75 were assigned to three groups (Fig. 5a–c). The remaining cultivars, viz. Atashgoon, Azin2, Sahar, Marmar, Sahand2, Khalijfars2, Hour, Erica, Ashna, Bolor, Atash2, Romina, Dorna2, Aydin and Taraneh, were highly variable in respect to the molecular level (membership value < 0.75), consequently forming the admixed group. These cultivars could be suggested for the improvement of mapping population and tagging QTLs of important ornamental characteristics (Kumar et al. 2017). Also, evaluation of genetic diversity values (H) for each group showed that the admixed group had a higher genetic diversity index (0.33), revealing that this group had more allelic richness and genetic diversity. The highest genetic distance was obtained between the clusters a and b (0.287); this was followed by the distance between the clusters b and c (0.280). These differences might be attributed to the phenotypic variation of the cultivars in the groups. It indicated that the cultivars in these clusters were more diverse and could be introduced as the parental candidates to improve the new Chrysanthemum cultivars.
The clusters inferred by structure analysis could also be used for distinguishing the flower colors. White flowers were only found in the admixed group, while purple and red ones were the most abundant in the group b. The yellow color was also dominant in the clusters a and c. Similarly, recent studies have reported the application of SRAP markers to differentiate among flower color, which can lead to association with the behavior of pollinators or seed-dispersing organisms (Robarts and Wolfe 2014). Also, according to Li et al. (2013), the Chrysanthemum cultivars were separated in terms of their flower size by using the UPGMA clustering results, which had been obtained from conserved DNA-derived polymorphism (CDDP). Furthermore, based on the genetic similarity index, the greatest genetic distances were observed between the cultivars Fariba2 (which belonged to the cluster c) and Sahel (which belonged to the cluster b), while the most similar cultivars were Ashna and Hour (both belonged to the admixed group). Ashna and Hour possessed a semi-double flower head type. The cultivar Sahel had purple flowers, needed medium days for the flowering stage, with single flower head type and the medium plant height, while the cultivar Fariba2 had a yellow flower color, early flowering time, and a semi-double flower head type with the low plant height. The high genetic distance between some cultivars including Sahel and Fariba2, as well as Sahel and Forough, could provide valuable genetic information to introduce the parental materials for the purpose of breeding programs.
The Mantel test did not show any significant correlation between the results of morphological and molecular data (r = − 0.16, p > 0.05). The negative correlation showed that the two methods (morphological and molecular approach) were distinct and highly variable (Kumar et al. 2013). Choudhary et al. (2017) also suggested that molecular markers should not be seen as a substitute of morphological traits as each marker system evaluates different aspects of genetic variation. Moreover, Solouki et al. (2008) suggested the role of environmental factors in determining the low correlation between phenotypic and molecular data, which could be due to the differentiation of traits-related genes from those associated with the adaptive response of the cultivars (Soriano et al. 2016).
5 Conclusion
Chrysanthemum morifolium is commercially an important plant with various applications. For the first time, the genetic diversity and population structure of Iranian Chrysanthemum cultivars were analyzed using SRAP markers and morphological traits in two constitutive years. Remarkable phenotypic and molecular variations were observed among the studied cultivars. Overall, the results of the present study showed that the morphological traits as well as SRAP molecular markers could be valuable resources to facilitate germplasm identification. Selection of divergent parents could be useful in wide crossing for developing different hybrid combinations and new Chrysanthemum cultivars. Finally, according to the results of the two morphological and molecular parts for Sahel and Forough cultivars as well as those for Taraneh and Shokoh, a relatively high genetic distance was observed; therefore, these cultivars could be recommended for use as the parental crosses.
References
Ahmad HM, Ahsan M, Ali Q, Javed I (2012) Genetic variability, heritability and correlation studies of various quantitative traits of mungbean (Vigna radiate L.) at different radiation levels. Int Res J Microbiol 3:352–362
Aneja B, Yadav NR, Yadav RC, Kumar R (2013) Sequence related amplified polymorphism (SRAP) analysis for genetic diversity and micronutrient content among gene pools in mungbean [Vigna radiata (L.) Wilczek]. Physiol Mol Biol Plants 19:399–407
Arzani A, Ashraf M (2016) Smart engineering of genetic resources for enhanced salinity tolerance in crop plants. Crit Rev Plant Sci 35:146–189
Banerji BK, Batra A, Dwivedi AK (2012) Morphological and biochemical characterization of Chrysanthemum. J Hortic Sci 7:51–55
Chen H, Morrell PL, Ashworth VE, De La Cruz M, Clegg MT (2008) Tracing the geographic origins of major avocado cultivars. J Hered 100:56–65
Choudhary SB, Sharma HK, Kumar AA, Maruthi RT, Mitra J, Chowdhury I, Singh BK, Karmakar PG (2017) SSR and morphological trait based population structure analysis of 130 diverse flax (Linum usitatissimum L.) accessions. C R Biol 340:65–75
Da Silva JAT (2004) Ornamental Chrysanthemums: improvement by biotechnology. Plant Cell Tissue Org Cult 79:1–18
Falconer DS, Mackay TF, Frankham R (1996) Introduction to quantitative genetics. Trends Genet 12:280
Feng SG, He RF, Jiang MY, Lu JJ, Shen XX, Liu JJ, Wang ZA, Wang HZ (2016) Genetic diversity and relationships of medicinal Chrysanthemum morifolium revealed by start codon targeted (SCoT) markers. Sci Hortic 201:118–123
Fu X, Ning G, Gao L, Bao M (2008) Genetic diversity of Dianthus accessions as assessed using two molecular marker systems (SRAPs and ISSRs) and morphological traits. Sci Hortic 117:263–270
Gaikwad AM, Katwate SM, Nimbalkar CA (2002) Evaluation of Chrysanthemum cultivars for growth and yield. South Indian Hortic 50:624–628
Ghafouri F, Rahimmalek M (2018) Genetic structure and variation in different Iranian myrtle (Myrtus communis L.) populations based on morphological, phytochemical and molecular markers. Ind Crops Prod 123:489–499
Han J, Hu N, Li YG, Shang FD (2007) Genetic diversity of Chrysanthemum cultivars revealed by AFLP analysis. Acta Hortic Sin 34:1041
Hassanein AMA, Al-Soqeer AA (2017) Morphological and genetic diversity of Moringa oleifera and Moringa peregrina genotypes. Hortic Environ Biotechnol 59:251–261
Kulkarni BS, Reddy BS (2004) Vegetative growth, flower yield and quality of different Chrysanthemum cultivars. J Ornam Hortic 7:32–36
Kumar S, Sharma R, Kumar V, Vyas GK, Rathore A (2013) Combining molecular-marker and chemical analysis of Capparis deciduas (Capparaceae) in the Thar desert of Western Rajasthan (India). Rev Biol Trop 61:311–320
Kumar S, Kumar M, Yadav HK, Sharma S (2017) Genetic diversity and population structure analysis of Chrysanthemum (Dendranthema grandiflora Tzvelev) germplasm based on RAPD markers. J Environ Biol 38:457
Li T, Li Y, Ning H, Sun X, Zheng C (2013) Genetic diversity assessment of Chrysanthemum germplasm using conserved DNA-derived polymorphism markers. Sci Hortic 162:271–277
Martí C, Uberhuaga E, Pérez C (2002) Application of RAPD markers in the characterisation of Chrysanthemum varieties and the assessment of somaclonal variation. Euphytica 127:247–253
Mohammadi SA, Prasanna BM (2003) Analysis of genetic diversity in crop plants-salient statistical tools and considerations. Crop Sci 43:1235–1248
Mokhtari N, Rahimmalek M, Talebi M, Khorrami M (2013) Assessment of genetic diversity among and within Carthamus species using sequence-related amplified polymorphism (SRAP) markers. Plant Syst Evol 299:1285–1294
Murray MG, Thompson WF (1980) Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8:4321–4326
Naz S, Shehzadi I, Shahzadi K, Ilyas S (2015) Assessment of genetic variability in different Chrysanthemum varieties of Pakistan using RAPD markers. J Anim Plant Sci 25:554–560
Palai SK, Rout GR (2011) Characterization of new variety of Chrysanthemum by using ISSR markers. Hortic Bras 29:613–617
Robarts DW, Wolfe AD (2014) Sequence-related amplified polymorphism (SRAP) markers: a potential resource for studies in plant molecular biology. Appl Plant Sci 2:1400017
Roein Z, Asil MH, Sabouri A, Dadras AR (2014) Genetic structure of Chrysanthemum genotypes from Iran assessed by AFLP markers and phenotypic traits. Plant Syst Evol 300:493–503
Roein Z, Hassanpour Asil M, Sabouri A (2015) Identification of AFLP markers associated with flowering time and ornamental traits in Chrysanthemum. Iran J Genet Plant Breed 4:37–47
Shao QS, Guo QS, Deng YM, Guo HP (2010) A comparative analysis of genetic diversity in medicinal Chrysanthemum morifolium based on morphology, ISSR and SRAP markers. Biochem Syst Ecol 38:1160–1169
Solouki M, Mehdikhani H, Zeinali H, Emamjomeh AA (2008) Study of genetic diversity in chamomile (Matricaria chamomilla) based on morphological traits and molecular markers. Sci Hortic 117:281–287
Soriano JM, Villegas D, Aranzana MJ, Moral LFG, Royo C (2016) Genetic structure of modern durum wheat cultivars and mediterranean landraces matches with their agronomic performance. PLoS One 11:e0160983
Talebi M, Rahimmalek M, Norouzi M (2015) Genetic diversity of Thymus daenensis subsp. daenensis using SRAP markers. Biologia 70:453–459
Xie WG, Zhang XQ, Cai HW, Liu W, Peng Y (2010) Genetic diversity analysis and transferability of cereal EST-SSR markers to orchardgrass (Dactylis glomerata L.). Biochem Syst Ecol 38:740–749
Yao MZ, Chen L, Liang YR (2008) Genetic diversity among tea cultivars from China, Japan and Kenya revealed by ISSR markers and its implication for parental selection in tea breeding programmes. Plant Breed 127:166–172
Zhang F, Chen S, Chen F, Fang W, Chen Y, Li F (2011a) SRAP-based mapping and QTL detection for inflorescence-related traits in Chrysanthemum (Dendranthema morifolium). Mol Breed 27:11–23
Zhang F, Chen S, Chen F, Fang W, Deng Y, Chang Q, Liu P (2011b) Genetic analysis and associated SRAP markers for flowering traits of Chrysanthemum (Chrysanthemum morifolium). Euphytica 177:15–24
Acknowledgements
The authors would like to thank Mr. Mohammad Reza Shafie at the National Research Center of Ornamental Plants, Mahallat, Iran, for providing the genetic materials used in this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interests.
Additional information
Communicated by Inhwa Yeam.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hodaei, M., Rahimmalek, M. & Arzani, A. Genetic diversity of Iranian Chrysanthemum morifolium cultivars using morphological traits and sequence-related amplified polymorphism (SRAP) markers. Hortic. Environ. Biotechnol. 60, 753–765 (2019). https://doi.org/10.1007/s13580-019-00137-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13580-019-00137-5