Abstract
Genetic erosion in cultivated wheat provides a good reason for studying genetic diversity in crop wild relatives. In the present study, genetic diversity of 32 accessions belonging to T. boeoticum species collected from different parts of Iran were evaluated using 13 morphological traits as well as ten inter-simple sequence repeat primers. Statistical analysis for morphological traits showed significant differences among accessions (except number of fertile tillers and total tillers per plant). In principal component analysis, the first three PCA showed 82.65 % of the total morphological variation. Based on the morphological traits, accessions were separated into two main groups by cluster analysis. In molecular analysis, polymerase chain reactions amplified 105 DNA fragments, out of which, 95 (90.47 %) were polymorphic. From geographic perspective, the accessions sampled from western and southwestern of Iran showed the highest and lowest polymorphism, respectively. However, the maximum values of effective number of alleles (Ne), Nei’s gene diversity (He) and Shannon’s information index (I) was related to accessions collected from NW regions. Also, according to cluster analysis and PCoA plot genetic diversity was not related to geographical distribution. Overall, our results revealed a remarkable level of genetic diversity among studied Iranian T. boeoticum accessions; especially accessions collected from Kermanshah and Lorestan provinces, which can be of interest for future breeding programs. So, conservation of germplasm of these areas is recommended.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genetic erosion in cultivated wheat suggests research for assessing genetic diversity in its wild relatives. The genus Triticum L., at diploid level includes three species Triticum monococcum L., T. urartu Gandilyan and T. boeoticum Boiss. (Morrison and Damania 1993). There are many studies showing that T. monococcum, as one of the first crops in Fertile Crescent, was domesticated through T. boeoticum (Dvorak et al. 1998). T. urartu, instead was never domesticated, though it played an important role in wheat evolution, since it donated the A genome to all tetraploid and hexaploid wheat (Kilian et al. 2007). The main geographical origins of T. boeoticum are in the central and eastern parts of the Fertile Crescent valley, that is part of Armenia, Azerbaijan, Iran, Iraq, Lebanon, Syria and Turkey (Miller 1987; Zohary and Hopf 1973). Iran is a main center of distribution of wild wheat’s with associated compositions of Triticum and Aegilops as the richest wheat gene pool has been found in this region (Tabatabaei and Maassoumi 2001). The origins of wild wheat’s in the different parts of Iran especially northwest and west regions (east of the Fertile Crescent), suggest this area as an ideal one for discovering suitable genes to be transferred into cultivated wheat. In particular, T. boeoticum in these area has high levels of genetic diversity and may provide significant information regarding its potential for breeding purposes (Naghavi et al. 2010).
It is assumed that evaluation of genetic diversity is very important to maximize the efficiency of breeding programs. Genetic diversity of many plant species has so far been investigated using morphological characters and molecular markers. Morphological markers are usually used to estimate genetic diversity, but are not entirely successful due to environmental influences. By contrast, molecular markers provide valuable information, especially in studies on genetic diversity and genetic relationships between different plant species, because they show genetic differences in greater detail without interference from environmental factors (Ghahremani-Majd and Dashti 2014). Genetic diversity among Iranian Einkorn populations has been studied using a large number of marker systems, including morpho-physiological characters (Salimi et al. 2005) floristic index (Naghavi et al. 2009) seed storage protein (Sofalian and Valizadeh 2009) isozymes (Cheniany et al. 2007) and molecular markers (Chhuneja et al. 2008; Kojima et al. 1998; Naghavi et al. 2010; Nasernakhaei et al. 2013). Inter-simple sequence repeats (ISSRs) are the most popular markers based on polymerase chain reaction. These markers have successfully been used to genetic identification (Fahima et al. 1998) estimate genetic relationship among different accessions (Roder et al. 1998) genetic diversity analysis (Mousavifard et al. 2015) and genomic fingerprinting (Fernandez et al. 2002). Furthermore, these markers are known to be repeatable, highly polymorphic as well as very reproducible and highly informative (Bornet and Branchard 2001).
In the present study, we aimed to evaluate genetic variability among populations of T. boeoticum sampled from the northwest, west and southwest of Iran using agro-morphological characters and molecular data obtained from ISSR markers.
Materials and methods
Plant material and field experiment
Thirty-two accessions of T. boeoticum Boiss. collected from the northwest to southwest of Iran were analyzed. Accessions are preserved in the Gene Bank of the University of Ilam. Detailed information for these accessions is listed in Table 1. Seeds of all populations were put in double layered rolled filter papers. Then the rolled papers were transferred into plastic bags to avoid moisture loss. Seeds were allowed to germinate in a germinator at 20 °C for 4 days. After germination seedlings were cultured in plastic pots filled with soil. In order to vernalization the pots transferred into a growth chamber (2–4 °C) for 4 weeks. After this period, the plant materials were planted at a randomized complete block design with three replications in Research Station of the Faculty of Agriculture, University of Tabriz, Iran (Latitude 38°05′N, Longitude 46°17′E, and Altitude 1360 m above sea level) in 2010–2011 cropping seasons, and 13 characters were recorded following IPGRI handbook. Each experimental plot consisted of 1-m rows and plants in the row spaced 20 cm apart. The climate is characterized by mean annual precipitation of 245.75 mm per year, mean annual temperature of 10 °C, and annual maximum temperature of 16.6 °C.
Agro-morphological traits
The agro-morphological characters were: heading data, plant height, leaf length (second leaf from top), leaf width (second leaf from top), main spike length, number of total tillers, number of fertile tillers, number of spikelet per spike, number of seeds per spike, weight of 100-seeds, seed yield per plant, biomass and harvest index. Heading date was recorded as number of days from planting until 50 % of the heads in a plot. Plant height was measured from ground to the tip of the main spike at maturity. Length and width of leaves were measured from leaf sheath to the tip of leaf. Also main spike length was measured excluding awns. To avoid the loss of seed and estimate grain yield per plant, all spikes were protected with trap. Finally, total above ground dry weight was measured as biomass. Then data analyzed using SAS v9.1 software (SAS 2004). Statistical analysis included: Analysis of variance (ANOVA), simple statistics (i.e.; mean, variance and C.V.), phenotypic correlation, principal components analysis (PCA) and cluster analysis.
DNA extraction and molecular analysis
From each accession, after the seed germination and growth, DNA was isolated from the leaves of greenhouse-grown plants according to the Piccolo et al. (2012) protocol. DNA quality was also analyzed by 1 % agarose. After optimizing polymerase chain reaction (PCR), ten ISSR primers were selected from the set of Biotechnology Laboratory, University of British Columbia, Canada. The PCRs were performed in the reaction mixture 25 µL volume, with 1.5 µL of isolated DNA from each sample, 1 µL of each primer, 12.5 µL Master kit (Thermo12.6), 10 µL ddH2O. Amplification was run at 94 °C for 7 min, followed by 40 cycles of denaturation at 94 °C for 30 s, primer annealing at 34–45 °C (varied for each primer) for 45 s and primer elongation at 72 °C for 2 min. The final extension was 7 min at 72 °C. The amplification reaction products were detected by 1.5 % denaturing agarose gels stained with Ethidium Bromide under UV light.
To characterize genetic variation, some of parameters namely; Shannon’s information index (Is), Nei’s gene diversity (He) and the observed number of alleles (Ne) were calculated for accessions. All of the above calculations were carried out using PopGen 1.31 software (Yeh et al. 1997). Cluster analysis based on Jaccard’s similarity coefficients matrix (Jaccard 1908) was constructed using NTSYS-pc software (Rholf 2000). Principal coordinated analysis (PCoA) was also carried out to show accessions distribution in a scatter-plot by GenAlEx 6.41 software (Peakall and Smouse 2006). Also, correlation between ISSR data and agro-morphological traits was measured using Mantel test (Mantel 1967) statistics by NTSYS-pc software (Rholf 2000).
Results
Agro-morphological analysis
Analysis of variance showed significant differences among accessions for all characters except number of fertile tillers and total tillers per plant. Mean values, ranges of variability and coefficients of variance for the different characters are presented in Table 2. The CV values for six of the agro-morphological characters indicated a high level of variation, namely seed yield per plant, number of total and fertile tillers per plant, weight of 100-seeds, harvest index, and biomass. By contrast, low levels of variation among the accessions were found for heading data, leaf width and plant height. Simple correlation coefficient analysis also revealed the existence of significant positive or negative correlations among different characters (data not presented). Taking into account of agro-morphological means comparison we identified the best accessions for each character (Table 3). For instance, the greatest amount of plant height, leaf width and grain yield per plant belonged to the Songhor (a region in Kermanshah province) accession (IUGB-0320). Also, the highest number of fertile tillers and total tillers per plant recorded for accessions IUGB-0052, IUGB-0120 and IUGB-0316 (collected from Sefid-Dasht; a region in Lorestan province).
The PCA showed that three components explained 82.65 % of the total variation contributed by agro-morphological traits (Table 4). The first component explained 52.89 % of the total variation including plant height, leaf width, spike length, number of total tillers per plant, number of spikelet and seeds per spike, weight of 100-seeds, grain yield and biomass per plant. The second factor indicated 17.82 % of the total variation and presented heading data, leaf length, weight of 100-seeds, and harvest index. The third factor showed 11.94 % of the total variation and containing number of fertile tillers. The agro-morphological cluster analysis divided the accessions into two main groups at distance level of 15 (Fig. 1). The first main group (A) was divided into two sub-clusters at distance level of 10. In the sub-cluster Ia, the accessions in view of leaf length, weight of 100- seeds, seed yield per plant, and harvest index were higher than the others. In the sub-cluster Ib, the accessions higher in view of heading data, plant height, leaf width, spike length, number of total tillers and fertile tillers per plant, spike length, number of seed per spike, seed yield per plant, and biomass than other groups. The second main group (B) comprised other accessions of T. boeoticum, which had lower values for all characters.
Molecular analysis
In PCRs, the 10 ISSR primers generated 105 bands across 40 Iranian T. boeoticum accessions studied, out of which, 95 (90.47 %) were polymorphic (Table 5). The number of bands per primer ranged from 7 (UBC825) to (UBC826, UBC849, UBC856 and UBC887) with an average of 10.5 bands per primer. The PIC values for the primers in the present study varied from 0.39 (UBC841)–0.50 (UBC825) with an average 0.47. The primer UBC887 showed the highest value of marker index (5.8) while the primer UBC855 revealed the highest value of resolving power (10.9). Genetic similarity was estimated from the Jaccard’s similarity coefficient value for all accessions of T. boeoticum considering ISSR marker individually (coefficient matrix not shown). Pairwise genetic similarity of 32 accessions ranged from 0.11 to 0.86. The accessions IUGB-0113 and IUGB-316 from Qazvin and Lorestan had the highest genetic similarity (0.86) whereas the lowest similarity was detected in IUGB-114 from Qazvin and IUGB-407 from Kermanshah (0.11). These results recommend that, ISSR markers are suitable for the determination of genetic diversity of T. boeoticum.
From a geographical viewpoint, we subdivided the distribution range of the accessions studied into three geographical regions, northwest (NW), west (W) and southwest (SW) with 6, 19 and 6 samples, respectively (see Table 1). The maximum value of Na and PPL was found in the W region, whereas, the maximum Ne, Ho and I related to NW region. The genetic diversity parameters estimated for these regions shown in Table 6. The Na and PPL indexes indicated that the accessions collected from W region had the greatest value (0.27, 0.42 for He and I, respectively), while the maximum Ne, He and I related to accessions sampled from NW region. The results showed that the genetic diversity within the accessions collected from W and NW was greater than was found within the SW accessions.
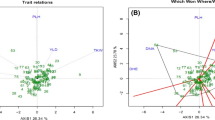
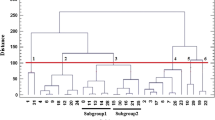
In order to study genetic relationships among accessions cluster analysis was performed based on Jaccard’s similarity coefficients matrix. The dendrogram constructed using UPGMA cluster analysis grouped all accessions into 5 major clusters (Fig. 2). Cluster A consisted of three accessions; cluster B consisted of 19 accessions; cluster C and D consisted of three accessions each and cluster E included four accessions. Also principal coordinate analysis (PCoA) showed that, the variance explained by the first and second component was 24.97 and 22.50 % respectively. Therefore, the first two components explained 47.47 % of the total variation. Results of scatter diagram (Fig. 3) using the first two components showed that genetic variation did not match with the geographical distribution, which was confirmed by cluster analysis. The Mantel test results showed a weak negative correlation (r = −0.013, p = 0.430) between morphological traits and ISSR markers. Despite of low correlation between agro-morphological and ISSR matrices, there were similar grouping of accessions in the respective dendrograms (sub-cluster Ia in Fig. 1 and cluster B in Fig. 2). However, there was some difference between the two dendrograms. For instance, the accessions IUGB-0004 (Kurdistan), IUGB-0012 (Kermanshah), IUGB-0019 (Kermanshah), and IUGB-0120 (Kermanshah), which has morphologically similarities in sub-cluster Ib (Fig. 1) were grouped in separate clusters (Group D and E) in ISSR analysis.
Discussion
In the present study there was a significant variation for all measured morphological characters except number of fertile tillers and total tillers per plant among T. boeoticum accessions. This variation illustrates a high level of genetic diversity and will help the future breeding programs to produce cultivars that are potentially suitable for improvement wheat (Sheikhehpour et al. 2014). Therefore, agronomic and morphological characteristics still have the advantage of providing a direct tool in the field for assessing plant performance, select and conserve germplasms. Similarly, many studies (Naghavi et al. 2009; Salimi et al. 2005; van Slageren 1994; Zhang et al. 2015) indicated that morphological characters are very helpful in identification and evaluation of genetic diversity in wild wheat germplasms. Most characters were highly variable between accessions, except for plant height, spike length and heading data, for which the CV was, <20 % (Table 2). The PCA showed that, the three first principle components accounted for 84.65 % of total genetic variation among accessions (Table 4). The first three principal components mainly explained by morphological traits. Bandou et al. (2009) using PCA analysis of morphological indices for Algeria Aegilops germplasms, found that morphological variability can be used to identify subspecies. Also, the agro-morphological cluster analysis clustered all accessions into two main groups (Fig. 1). Taking into account of agro-morphological analysis we identified the best accessions for each character (Table 3). For instance, one accession collected from Kermanshah (IUGB-0320) had the greatest amount of plant height as well as leaf width and seed yield per plant. Also, accessions collected from Lorestan (IUGB-0052, IUGB-0120 and IUGB-0360) recorded the highest number of fertile tillers and total tillers per plant. Our finding showed that these areas and their related accessions could be considered as an ideal source of morphological variation, thus detection of genetic diversity of them is very useful for breeding programs.
Furthermore, there was a significant variation among T. boeoticum accessions based on ISSR markers. The markers detected 10.5 polymorphic bands per primer with an average polymorphism of 90.47 %. This result detected a high level of polymorphism among accessions using ISSR, indicating the high efficiency of this DNA-marker to reveal genetic diversity of T. boeoticum accessions. Previously, several researchers screened the genetic diversity of Iranian T. boeoticum. Naghavi et al. (2010) using RAPD, AFLP and SSR markers showed that there is a vast variation in the T. boeoticum from west of Iran. Mousavifard et al. (2015) using ISSR marker also demonstrated high variability among the diploid wheats collected from northwest to southwest regions of Iran. From the geographic point of view, the genetic diversity among the accessions collected from west (W) and northwest (NW) was greater than was found among the SW accessions (Table 6). In contrary, the highest genetic polymorphism was detected among samples collected from the western region (W). Although the most accessions belonged to the west region, the polymorphism level may be affected by sample size, though the previous study (Naghavi et al. 2010) also reported high genetic diversity in Iranian populations of T. boeoticum collected in west Iran. Five accessions collected around Lorestan province (in the west Iran) showed close relationship, characterizing a distinct population. However, accessions collected from Kermanshah are genetically very divers. Accessions collected from West Azerbaijan, East Azerbaijan (northwest), and Qazvin (IUGB-0003, IUGB-0216, IUGB-0113, respectively), despite being geographical distant, were clustered with accessions collected from Kermanshah province (west).
According to the clustering pattern obtained by agro-morphological traits (Fig. 1) and ISSR data (Fig. 2) as well as PCoA plot (Fig. 3), the genetic variation did not agree with the geographical distribution, as it can be seen that accessions sampled from the same geographical area occur in different clusters, and on the contrary some of accessions from different areas clustered together. Similarly, in the studies conducted by Naghavi et al. (2009), Ebrahimi et al. (2009), Ghahremani-Majd and Dashti (2014) and Morshedloo et al. (2015) there was no relationship between geo-graphical origin and genetic diversity. Based on abovementioned results, it can be suggested that there are different sub-gene pools within the T. boeoticum in Iran. Similarly, Malaki et al. (2006) using AFLP markers found a high level of genetic diversity in the T. boeoticum accessions collected from Kermanshah and Lorestan provinces. In our study, there was a weak negative correlation between the clustering of the accessions based on the SSR markers and agro-morphological traits (r = −0.013). A poor correlation was often observed between morphological traits and molecular markers variations in many crop species (Zhang et al. 2015). Wilson et al. (1977) reported that, the morphological change and molecular divergence might be quite independent, responding to different evolutionary pressures and following different rules. There are several reasons for the disagreement between the results based on morphological and molecular variations; (1) morphological variation is strongly correlated with environmental conditions (Johns et al. 1997) (2) a change in a nucleotide repetition can results in a change in molecular pattern. Thus, microsatellite markers are not subjected to either natural or artificial selection compared to morphology characters (Semagn 2002) and (3) the lack of correlation between ISSR data and morphological markers may be affected by the fact that both types of markers follow different evolutionary paths (Wendel and Doyle 1998).
Conclusion
Crop wild relatives are increasingly used in wheat breeding programs as donors of genes conferring resistance to biotic and abiotic stresses, as well as of DNA that contribute to the improvement of grain quality. T. boeoticum has been reported as a valuable source of desirable genes conferring higher protein quality and amino-acid content (Ciaffi et al. 1998) bread making quality (Waines 1994) and resistance to biotic stresses (Hovhannisyan et al. 2011). Skovmand et al. (2002) also reported that in the Iranian mountains, especially Zagros Mountains (West of Iran), there are recourses of drought-tolerance and salt-tolerance. In conclusion, morphological genetic diversity observed in 32 accessions of T. boeoticum, which originated from different eco-geographical regions of Iran, revealed that they are rich in unexploited potentially useful agronomical as well as morpho-physiological traits. Therefore, sampling natural populations of T. boeoticum and evaluation of sampled material could bring to light more desirable DNA. Moreover, in this study molecular markers showed a high level of genetic diversity within T. boeoticum germplasms in Iran. It can be suggested that exploring this highly diverse gene pool may results in discovery valuable alleles for studies in wheat evolution, adaptation and improvement. Finely, these kinds of studies need to be further strengthened through more comprehensive sampling methods and advanced molecular techniques.
References
Bandou H, Rodriguez-Quijano M, Carrillo J, Branlard G, Zaharieva M, Monneveux P (2009) Morphological and genetic variation in Aegilops geniculata from Algeria. Plant Syst Evol 277:85–97. doi:10.1007/s00606-008-0106-z
Bornet B, Branchard M (2001) Nonanchored inter simple sequence repeat (ISSR) markers: reproducible and specific tools for genome fingerprinting. Plant Mol Biol Report 19:209–215. doi:10.1007/BF02772892
Cheniany M, Ebrahimzadeh H, Salimi A, Niknam V (2007) Isozyme variation in some populations of wild diploid wheats in Iran. Biochem Syst Ecol 35:363–371. doi:10.1016/j.bse.2006.12.006
Chhuneja P, Kaur S, Garg T, Ghai M, Kaur S, Prashar M, Bains N, Goel R, Keller B, Dhaliwal H (2008) Mapping of adult plant stripe rust resistance genes in diploid A genome wheat species and their transfer to bread wheat. Theor Appl Genet 116:313–324. doi:10.1007/s00122-007-0668-0
Ciaffi M, Dominici L, Lafiandra D (1998) High molecular weight glutenin subunit variation in wild and cultivated einkorn wheats (Triticum spp., Poaceae). Plant Syst Evol 209:123–137. doi:10.1007/BF00991528
Dvorak J, Luo M, Yang Z (1998) Genetic evidence on the origin of Triticum aestivum L., The origins of agriculture and crop domestication. In: Proceedings of the Harlan symposium. ICARDA, Aleppo, pp 235–251
Ebrahimi R, Zamani Z, Kashi A (2009) Genetic diversity evaluation of wild Persian shallot (Allium hirtifolium Boiss.) using morphological and RAPD markers. Sci Hortic 119:345–351. doi:10.1016/j.scienta.2008.08.032
Fahima T, Roder M, Grama A, Nevo E (1998) Microsatellite DNA polymorphism divergence in Triticum dicoccoides accessions highly resistant to yellow rust. Theor Appl Genet 96:187–195. doi:10.1007/s001220050726
Fernandez M, Figueiras A, Benito C (2002) The use of ISSR and RAPD markers for detecting DNA polymorphism, genotype identification and genetic diversity among barley cultivars with known origin. Theor Appl Genet 104:845–851. doi:10.1007/s00122-001-0848-2
Ghahremani-Majd H, Dashti F (2014) Genetic diversity of Persian shallot (Allium hirtifolium Boiss.) populations based on morphological traits and RAPD markers. Plant Syst Evol 300:1021–1030. doi:10.1007/s00606-013-0940-5
Hovhannisyan NA, Dulloo ME, Yesayan AH, Knüpffer H, Amri A (2011) Tracking of powdery mildew and leaf rust resistance genes in Triticum boeoticum and T. urartu, wild relatives of common wheat. Czech J Genet Plant 47:45–57
Jaccard P (1908) Nouvelles recherches sur la distribution florale. Bulletin de la Societe Vaudoise Sciences Naturelles 44:223–270
Johns MA, Skroch PW, Nienhuis J, Hinrichsen P, Bascur G, Muñoz-Schick C (1997) Gene pool classification of common bean landraces from Chile based on RAPD and morphological data. Crop Sci 37:605–613. doi:10.2135/cropsci1997.0011183X003700020049x
Kilian B, Ozkan H, Deusch O, Effgen S, Brandolini A, Kohl J, Martin W, Salamini F (2007) Independent wheat B and G genome origins in outcrossing Aegilops progenitor haplotypes. Mol Biol Evol 24:217–227. doi:10.1093/molbev/msl151
Kojima T, Nagaoka T, Noda K, Ogihara Y (1998) Genetic linkage map of ISSR and RAPD markers in Einkorn wheat in relation to that of RFLP markers. Theor Appl Genet 96:37–45. doi:10.1007/s001220050706
Malaki M, Naghavi MR, Alizadeh H, Potki P, Kazemi M, Pirseyedi SM, Mardi M, Tabatabaei F (2006) Study of genetic variation in wild diploid wheat (Triticum boeoticum) from Iran using AFLP markers. IJB 4:269–274
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Cancer Res 27:209–220
Miller T (1987) Systematics and evolution wheat breeding. Springer, Germany
Morrison L, Damania A (1993) Triticum–Aegilops systematics: taking an integrative approach biodiversity and wheat improvement. Wiley, New York
Morshedloo MR, Moghadam MRF, Ebadi A, Yazdani D (2015) Genetic relationships of Iranian Hypericum perforatum L. wild populations as evaluated by ISSR markers. Plant Syst Evol 301:657–665. doi:10.1007/s00606-014-1103-z
Mousavifard SS, Saeidi H, Rahiminejad MR, Shamsadini M (2015) Molecular analysis of diversity of diploid Triticum species in Iran using ISSR markers. Genet Resour Crop Evol 62:387–394. doi:10.1007/s10722-014-0160-z
Naghavi M, Maleki M, Tabatabaei S (2009) Efficiency of floristic and molecular markers to determine diversity in Iranian populations of T. boeoticum. World Acad Sci Eng Technol 3:42–44
Naghavi M, Malaki M, Alizadeh H, Pirseiedi M, Mardi M (2010) An assessment of genetic diversity in wild diploid wheat Triticum boeoticum from west of Iran using RAPD, AFLP and SSR markers. J Agric Sci Technol 11:585–598
Nasernakhaei F, Rahiminejad MR, Saeidi H, Tavasoli M (2013) Taxonomic identity of the Iranian diploid Triticum as evidenced by nrDNA ITS analysis. Phytotaxa 143:43–53. doi:10.11646/phytotaxa.143.1.2
Nei M (1973) Analysis of gene diversity in subdivided populations. Proc Natl Acad Sci 70:3321–3323
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295. doi:10.1111/j.1471-8286.2005.01155.x
Piccolo SL, Alfonzo A, Conigliaro G, Moschetti G, Burruano S, Barone A (2012) A simple and rapid DNA extraction method from leaves of grapevine suitable for polymerase chain reaction analysis. Afr J Biotechnol 11:10305–10309. doi:10.5897/AJB11.3023
Rholf F (2000) Numerical taxonomy and multivariate analysis system: version 2.10. Exeter Software, Setauket, New York
Roder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Salimi A, Ebrahimzadeh H, Taeb M (2005) Description of Iranian diploid wheat resources. Genet Resour Crop Evol 52:351–361. doi:10.1007/s10722-005-2256-y
SAS (2004) STAT User’s guide. v. 9.1. SAS Inst Inc., Cary
Semagn K (2002) Genetic relationships among ten endod types as revealed by a combination of morphological, RAPD and AFLP markers. Hereditas 137:149–156. doi:10.1034/j.1601-5223.2002.01600.x
Shannon CE, Weaver W (1949) Mathematical theory of communication. University of Illinois Press, Urbana
Sheikhehpour S, Bahraminejad S, Cheghamirza K (2014) Morphological and molecular genetic variations of oat genotypes grown in Kermanshah, Iran. Mol Biol Rep 41:4023–4030. doi:10.1007/s11033-014-3271-x
Skovmand B, Rajaram S, Ribaut J, Hede A (2002) Wheat genetic resources. FAO Plant Production and Protection Series (FAO), Rom, Italy
van Slageren MW (1994) Wild wheats: a monograph of Aegilops L. and Amblyopyrum (Jaub. & Spach) Eig (Poaceae). Wageningen Agricultural University Papers
Sofalian O, Valizadeh M (2009) Investigation of seed storage proteins in some wild wheat progenitors using SDS-PAGE and Acid-PAGE. Not Bot Horti Agrobot 37:179–182
Tabatabaei SF, Maassoumi TR (2001) Triticum boeoticum ssp. thaoudar “exists” in Iran. Cereal Res Commun 29:121–126
Waines JG (1994) High temperature stress in wild wheats and spring wheats. Funct Plant Biol 21:705–715. doi:10.1071/PP9940705
Wendel JF, Doyle JJ (1998) Phylogenetic incongruence: window into genome history and molecular evolution molecular systematics of plants II. Springer, Berlin
Wilson AC, Carlson SS, White TJ (1977) Biochemical evolution. Annu Rev Biochem 46:573–639. doi:10.1146/annurev.bi.46.070177.003041
Yeh F, Yang R, Boyle T (1997) POPGENE, the user friendly shareware for population genetic analysis. Molecular Biology and Biotechnology Center. University of Alberta, Edmonton
Zhang Z, Gao J, Kong D, Wang A, Tang S, Li Y, Pang X (2015) Assessing genetic diversity in Ziziphus jujuba ‘Jinsixiaozao’ using morphological and microsatellite (SSR) markers. Biochem Syst Ecol 61:196–202. doi:10.1016/j.bse.2015.06.021
Zohary D, Hopf M (1973) Domestication of pulses in the old world legumes were companions of wheat and barley when agriculture began in the Near East. Science 182:887–894. doi:10.1126/science.182.4115.887
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Pour-Aboughadareh, A., Mahmoudi, M., Moghaddam, M. et al. Agro-morphological and molecular variability in Triticum boeoticum accessions from Zagros Mountains, Iran. Genet Resour Crop Evol 64, 545–556 (2017). https://doi.org/10.1007/s10722-016-0381-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-016-0381-4