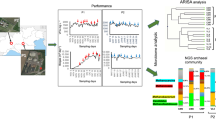
Abstract
Vinasse is a by-product from the ethanol industry and can be used for methane production through anaerobic digestion process driven by microbial consortia. The microorganisms involved must be known to obtain an ideal methane production. The present work evaluated the production of methane and byproducts from different ratios substrate/biomass (S0/X0), using sludge from an effluent treatment of the vegetable oil industry as inoculum in media containing vinasse. Also, the microbial community of the best methane production bioassay was characterized by high-throughput DNA sequencing. The following chemical parameters were evaluated: methanogenic activity, chemical oxygen demand, carbohydrate consumption, and production of volatile fatty acids. The highest methane production occurred at S0/X0 ratio of 1.5, which produced 59.78 mmol CH4 L−1. A great variety of microorganism genera was identified by high-throughput DNA sequencing, showing differences in the microbial consortia of the initial and final sampling times. At the final sampling point, the classes Bacteroidia (Porphyromonadaceae—OTU genera unknown 42.26% and Bacteroides genus 10.58%) and the class Betaproteobacteria (Proteobacteria-Comamonadaceae OTU) were identified as the dominant bacteria. The most abundant archaeal genera in the bioassay were Methanosaeta, Methanomassiliicoccaceae OTU vadinCA11, and Methanobacterium. The identification of the microorganisms of consortia involved in anaerobic digestion can collaborate on technologies to increase methane production through microbial isolation, bioaugmentation, and co-cultures.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Ethanol represents one of the major alternatives for reducing dependence on fossil fuels. Brazil is the world’s largest producer of ethanol from sugarcane, producing about 28 billion ton per year [1]. The bioethanol production is expected to keep growing for the next decade to reach 134 billion liters in 2024 with biggest producers expected to be USA, Brazil, European Union, and China [2]. However, for each liter of ethanol produced, 12–15 l of vinasse is generated. Vinasse is a liquid by-product of high organic load, dark brown color, low pH, high salt content, and highly polluting [3, 4]. It contains a chemical oxygen demand (COD) of 20 to 100 g COD.L−1, some organic solids in suspension as minerals, residual sugar, and some volatile compounds. Currently, vinasse is discharged directly onto farmland by fertigation, but its prolonged use as fertilizer leads to the salinization of the soil and desertification [5, 6]. Also, sprinkling vinasse on sugarcane crops delays their maturation and leads to a decreased sugar content [7]. The high production of vinasse demands a search for better environmental management of this by-product.
Methane is a sustainable alternative for energy production, being produced from agricultural byproducts as a substrate [8, 9]. The production of methane through vinasse biodigestion helps to reduce the polluting potential of this residue, producing value-added byproducts as well as a high-quality biofertilizer for agriculture [3].
The process of vinasse bioconversion can be performed using inocula of natural microbial communities, such as sludge from effluent treatment plants [8, 10]. The microorganism consortia involved in this process shows a series of complex metabolic reactions involving several species of bacteria and archaea [11]. They act symbiotically at intermediary stages, such as hydrolysis, acidogenesis, acetogenesis, and methanogenesis [12,13,14]. The knowledge of the microbial communities is essential for understanding and developing the process of methanogenesis [15,16,17]. In this context, the objective of this work was to evaluate the production of methane and by-product of vinasse in bioassays with different S0/X0 ratios and to use high-throughput sequencing to identify the microbial community associated with the best methane bioassay.
2 Materials and methods
2.1 Operational conditions
Sugarcane vinasse provided by the company Guarani SP, Brazil, was used as a culture medium for the bioassays (Table 1). It was centrifuged at 10,000 rpm for 5 min to remove coarse solids. The biomass used was granular sludge from UASB reactor, from vegetable oil treatment station (S 29° 51′ 41.04″; W 51° 10′ 45.12″). The inoculum that was lightly macerated to reduce granulometry and was directly inoculated in the culture medium with vinasse.
The experiments were performed in triplicate, using four different substrate/biomass ratios (S0/X0 = 0.5, 1.0, 1.5, and 1.7) (Table 2). Bioreactors with a total volume of 610 mL (366 mL of medium and 244 mL of headspace) run for 14 days under mesophilic conditions. The pH was adjusted periodically to 6.8–7.2 using KOH 3M to maintain optimum conditions for archaea development. The experiments were kept under stirring in an orbital shaker at 140 rpm. Nitrogen sprinkled for 10 min to maintain in anaerobiosis of the medium. During the fermentation, initial (0 h), intermediary (168 h), and final (336 h) samples were collected, and sent to chemical and biological analyses. Gaseous samples were collected from the headspace every 24 h.
2.2 Chemical analyses
The analysis of methane production was performed using gas chromatography (DaniMaster AS), with CarboxenTM 1006 PLOT capillary column (30 m × 0.53 mm), with thermal conductivity detector (TCD), ultrapure nitrogen gas as the carrier gas. The amount of methane produced was analyzed every 24 h for 14 days. A calibration curve was performed using standard methane gas. Analyses were also made from liquid samples of volatile fatty acids (acetic, propionic, butyric, isobutyric, valeric, and isovaleric), where samples were centrifuged at 1000 rpm for 10 min, filtered at 0.2 membrane (Millipore), and diluted in 5 × methanol. They were analyzed using gas chromatography (GC/MS, Shimadzu-QP 2010 Ultra) equipped with a DN-FFAP (30 mx 0.32 mm × 0.25 μm) column with flame ionization detector (FID), with helium as carrier gas and synthetic, and air and nitrogen as auxiliary gases. The column temperature was 100 °C for 5 min, increasing 7 °C per min to 200 °C. The temperatures of the injector and detector were 200 °C and 250 °C, respectively. The volatile acids were identified by the respective retention times, compared with those obtained in the standard curves. For the development of the standard curves, synthetic solutions of the studied compounds were used. The analyses of the composition of the vinasse and the COD were determined following the standard methods [18]. The total carbohydrate concentration of the samples was determined by a colorimetric method using sucrose as standard [19].
2.3 Experimental data fitting
The experimental data in triplicate were adjusted using the Statistica software (version 10). The modified Gompertz model [20] was used to estimate the kinetic parameters (Rm, Hmax and λ) for the methane production in the different substrate/biomass ratios (Eq. 1).
where M is the cumulative methane production (mL); λ is the lag phase time (h); P is the ultimate methane production (mL); Rm is methane production rate (mL.h−1); and e is the constant (2.72).
2.4 High throughput DNA sequencing
To identify which microorganisms are parts of the methane production, the initial sample (IS, 0 h) and final sample (FS, 336 h) from the bioassay, S0/X0 = 1.5 ratio, were sequenced. The genomic DNA from the microbial consortia showing the best methane production was extracted from 250 μL of the sample with the DNeasy PowerSoil Kit (Qiagen), according to the manufacturer’s protocol. The extracted DNA was amplified by PCR with primers 515F (5′-GTGCCAGCMGCCGCGGTAA-3′) and 806R (5′-GGACTACVSGGGTATCTAAT-3′) for the 16S rRNA gene. The amplification conditions followed 35 cycles (95 °C, 30 s; 50 °C, 1 min; 72 °C, 1 min) after an initial denaturation at 95 °C for 3 min [21] in an Applied Biosystems Model 2720 thermal cycler. The samples were analyzed on 1% agarose gel. PCR products from the initial samples and final fermentation sample after the highest methane production were submitted to high throughput DNA sequencing on the Ion PGM System (Thermo Fisher) following the manufacturer’s instructions (Thermo Fisher). The libraries construction was performed with the Ion Plus Fragment Library kit for short amplicons (≤ 350 pb), from 100 ng of the amplification product. A barcode adapter of the IonXpress Barcode adapters was added to each sample. All procedures were performed according to the manufacturer’s protocol. For quantification and equalization of the libraries, the Ion Library Equalizer kit was used, according to the manufacturer’s instructions. For emulsion PCR and enrichment steps, the Ion PGM Template OT2 200 kit was used with the Ion One Touch 2 system. For the sequencing, an Ion 316 chip was used with the Ion PGM Sequencing 200 v2 kit, according to the manufacturer’s instructions. Fragments of the 16S rRNA gene generated by the sequencing were submitted to quality control using the PRINSEQ program [22]. The replicate sequences were identified, sorted, and filtered to exclude unique sequences using the USEARCH v7.0.1090 program [23]. The clusters were assembled using a 99% minimum identity and chimeras were removed using the RDP reference database [24]. The taxonomic attribution was obtained using QIIME v1.7 based on 97% sequence similarities with the GreenGenes 13.8 database [25, 26]. Sequences were deposited in the National Center for Biotechnology Information (NCBI) under BioProject PRJNA471743.
3 Results and discussion
3.1 Methane production
All bioassays produced methane independent of the S0/X0 ratio used (Table 3). The best final production of methane P = 613.7 mL was found in the bioassay with a ratio of S0/X0 = 1.5, with a maximum production rate of 1.9 mL CH4 h−1 and lag phase (λ) at 59 h.
Comparing these results with other studies, it can be seen that Kiyuna et al. [27] found their best methane production of 541.4 ml CH4 in a vinasse study, and Shin et al. [28] in a work using another substrate (swine manure and food debris) presented 601 mL of CH4. The best methane yield (59.78 mmol CH4 L−1) was observed in 264 h (Fig. 1).
3.2 Carbohydrate consumption and removal COD
In the bioassays, the highest percentages of carbohydrate intake were observed in the S0/X0 = 1.7 (89.3%), 1.5 (78.6%), and 1.0 (72.7%) ratios. The removal of COD was less effective in the S0/X0 = 1.7 (53.7%) ratio and the higher removal efficiency occurred in S0/X0 = 0.5 with 89.4% (Table 3). In general, it is observed that the higher the availability of organic load in the medium, the higher the difficulty in the removal of COD. The ability of microbial consortia to reduce the organic load of vinasse is already well reported in the literature, reducing the impact in the environment [29, 30].
When analyzing the methane volume per gram of COD removed, it is observed that the S0/X0 value 0.5 to 1 is more advantageous than the higher ratios. In the same way, the conversion of substrate to methane is higher in S0/X0 0.5 and 1 (14.7%) and lower in S0/X0 (8.4 to 9.3%). The ratio S0/X0 = 1.5 was the dilution chosen because it presented the best-accumulated methane production, although COD removal and the lag phase time were not more efficient. These results can be explained by the influence of the fermentation parameters in the microbial communities. As can be observed, the higher substrate concentration (S0/X0 = 1.7) obtained higher intakes of carbohydrates, indicating a possible change in the metabolic pathways or predominance of microorganisms. These results, however, require further investigation. On the other hand, it is a fact that the microbial communities strongly alter the substrate composition, as can be observed in the fermentation of S0/X0 = 1.5 (Table 1). After 336 h of bioassay, it was noted the reduction of the concentration of potassium, iron, calcium, and the increase of phosphorus and total manganese.
3.3 Volatile fatty acids present in the fermentation
The acids analyzed in the fermentation process production of methane from sugarcane vinasse were acetic, propionic, butyric, isobutyric, valeric, and isovaleric (Fig. 2).
Chromatographic data show that in the initial vinasse samples volatile fatty acids were detected and that these acids were consumed during the anaerobic decomposition which can be seen in Fig. 2b–d, with exception of the sample S0/X0 = 0.5.
The volatile fatty acids have been reported as the primary precursors of methane production. Syntrophobacter groups, as sulfate reducers, are capable of degrading propionate in syntrophic association with methanogens, are the main propionate oxidizers [31, 32]. The Syntrophobacter group was found to be the most abundant among propionate degraders present in anaerobic digesters and the species described in the literature are S. fumaroxidans, S. wolinii, S. pfennigii, S. sulfatireducens, and HP1.1 strain of the Syntrophobacter genus [33,34,35,36,37]. On the other hand, a massive accumulation of the acids can also act as inhibitors of methanogenic microorganisms due to a decrease in the pH of the medium, to thermodynamic reasons, and the acids toxicity, destabilizing the cellular biochemical reactions of the consortia [38]. Acid production has become a useful process for the chemistry of renewable energies. These acid derivatives can be purified and used in the food, textile, pharmaceutical, leather, and plastic industries. The values obtained in the market are higher for the acids than for the methane, not to mention the ease of storage, transport, and greater safety [39, 40].
3.4 Microbial community identification
Samples from the S0/X0 = 1.5 ratio assay were submitted to high throughput DNA sequencing. A total of 78,892 and 86,051 DNA sequences were obtained from samples IS and FS, respectively.
Thirty-six phyla were identified in the two samples, and a larger number of taxa were observed in the SI than in the FS. A higher number of phyla in the SI compared with FS was also observed in other studies of anaerobic digestion of residues [11, 29, 41]. The environment to which the microorganisms are submitted in the fermentation tests to produce methane leads to a selection of microorganisms that act in the process and the number of taxa tends to decrease. Species in syntrophy can occur by the combined metabolic activity of microorganisms, allowing the microbial community to survive with minimal energy resources [8, 42]. Preliminary studies on biogas plants and anaerobic reactors presented similar results about the identified microorganisms [43,44,45].
Proteobacteria (25.6%), Bacteroidetes (18.7%), and Euryarchaeota (17.6%) were the most representative phyla in IS. In the FS, the most prevalent phyla were Bacteroidetes (58.5%), Firmicutes (14.1%), and Proteobacteria (13.1%) (Fig. 3). These phyla were also the most prevalent microorganisms in anaerobic fermentation of vinasse in the study of methanogenic analysis of desulphurization system used to treat biogas from vinasse methanization [29]. Bacteroidetes, the most abundant phylum in our study, is composed of strict anaerobes capable of degrading protein-rich substrates involved in the hydrolysis and acidogenesis of anaerobic digestion [46, 47].
Phylum Firmicutes contain individuals capable of fermenting various organic substrates and producing spores. The genus Clostridia, for example, can catabolize proteins, lipids, and complex carbohydrates [17, 43, 44, 48]. Proteobacteria, Deltaproteobacteria, and Gammaproteobacteria consist of microorganisms involved in acidogenesis, nitrification, and other metabolic processes [49, 50].
An OTU belonging to the order Bacteroidales was the most prevalent microorganism in the IS, corresponding to 10.96% of the total sequences (Table 4). Bacteria of the genera Syntrophomonas, Syntrophus, Flavobacterium, Candidatus Cloacomonas, Anaerolinaceae, Sulfuricurvum, Sulfurimonas, Thermomonas, and Kosmotoga also occurred in abundance higher in IS than in FS, and all were observed in abundance higher than 1% of the total sequences.
This result is in accordance with Krause et al. [51] that performed a study on a total community DNA sample from an agricultural biogas reactor continuously fed with maize silage, green rye, and small proportions of chicken manure. Microorganisms from the class Bacteroidia were identified in other studies using a methanogenic reactor to treat livestock waste, and they might be related to the fermentation of glucose, producing acetate and propionate [52,53,54]. Firmicutes is an acetate-oxidizing bacteria, in syntrophic cooperation with hydrogenotrophic methanogens [55].
Syntrophomonas is a bacterial genus capable of fermenting fatty acids in syntrophy with microorganisms that use hydrogen and formate [56]. Since vinasse is rich in organic acids—previous studies have reported concentrations of volatile fatty acids in the order of 19 g L−1 in sugarcane molasses [49].
When analyzing the most prevalent microorganisms in FS, we can see the unknown genus of the family Porphyromonadaceae with the highest number of copies of RNA16S genes with 42.26% followed by Bacteroides with 10.58% and OTU Comamonadaceae with 7.94% of abundance (Table 4). These microorganisms have been related to some stage of anaerobic digestion, such as hydrolysis, acidogenesis, or acetogenesis [49, 50, 57].
A broad community of microorganisms is involved, including methanogens. These Archaea are the biologic key to the process because they accomplish the methane-forming reaction [58]. In this study, eight archaeal genera (or respective OTU) were observed in IS and FS in abundance higher than 1% of the total sequences. The most abundant microorganism of the phylum Crenarchaeota was an OTU belonging to the class MCG (pGrfC26) (2.39% in IS and 0.1% in FS). The most prevalent genera or respective OTU from the phylum Euryarchaeota in FS were Methanosaeta 1.97%, VadinCA11 (OTUs belonging to the family Methanomassiliicoccaceae) (1.15%), and Methanobacterium 0.7% (Table 5). These genera were probably the methane producers observed in the biogas [15].
The genus Methanosaeta is composed by acetoclastic species, capable of forming long and thin filaments that provide granules and syntrophy. Methanobacterium also produce filaments, but short, and are the most closely related hydrogenotrophic species to the final anaerobic digestion phase [59]. The genus Methanosarcina is well known to be together with Methanosaeta, the largest producer of methane, but it appears only in 0.43% in the FS sample. This reduced participation of Methanosarcina can be explained because this microorganism is sensible to propionic acid [60] and this acid was found in SF from 206.2 to 69.8 mg L−1 in this study.
The genus Methanosaeta was related to anaerobic digestion in the analysis of the microbial community of 22 anaerobic mesophilic digesters treating urban sewage [61]. In a study with anaerobic reactors for large-scale methane production, using 16S rRNA gene sequencing, [62, 63] found archaeal OTUs affiliated to the orders Methanosarcinales, Methanomicrobiales, phylum Crenarchaeota, and the candidate group Arc I. In summary, these groups of prokaryotes found in our anaerobic digestion ensure the production of methane, including hydrolysis, acidogenesis, acetogenesis, and vinasse methanogenesis. Also at low abundance (lower than 0.02% of the total sequences), the genus Methanospirillum was reported by Tsushima et al. [64], a hydrogenotrophic archaea at low temperatures.
With regard to the abundance of each taxon, there was an increase of some microorganisms in IS when compared with FS. The genera Chryseobacterium, Flavobacterium, Sphingobacterium, Pseudomonas, Syntrophus, Sulfurimonas, and the OTUs SHA-114, FCPU426, Hyd24-12, and ABY1 were at least 100 times more abundant in IS than in FS. The genus Ruminococcus and the OTUs BSV43 and R4-41B were at least 100 times more abundant in FS than in IS (Fig. 4).
4 Conclusions
The evaluation of the methanogenic activities showed that the bioassay with higher efficiency in producing methane after 236 h was in the S0/X0 = 1.5 ratio, with a value of 59.78 mmol CH4 L−1. In the microbial consortia, not all the microorganisms present could be identified at the genus level. Although not all the microorganisms present in the consortia could be identified at the genus level, a great variety was identified by high-throughput sequencing. This technique was able to distinguish the communities from the initial and final sampling times.
At the final sampling point from the highest methane production, the most prevalent phyla were Bacteroidetes, Firmicutes, and Proteobacteria. It can be observed the consumption of the acids during the production of methane was observed, especially the propionic and the presence of syntrophic groups like Syntrophobacter. A genus unknown from the order Bacteroidales was the most prevalent in the highest production of methane, followed by Bacteroides and an OTU belonging to the family Comamonadaceae. As regards Archaea, the most prevalent were the genera Methanosaeta, Methanobacterium, Methanosarcina (Euryarchaeota), and OTU VadinCA1.
The identification of the most prevalent genera and the relation of compounds generated in the anaerobic digestion serve to understand the participation of the microorganisms in the process and develop strategies in the bioprospection of products of biotechnological interest. The identification of the microorganisms of consortia involved in anaerobic digestion can collaborate on technologies to increase methane production through microbial isolation, bioaugmentation, and co-cultures.
References
_Agência Nacional de Petróleo, Gás natural e biocombustíveis. Produção de etanol anidro e hidratado, segundo grandes regiões e unidades da Federação – 2008-2017. http://www.anp.gov.br/publicacoes/anuario-estatistico/anuario-estatistico-2018. Accessed 20 Oct 2018
Hoarau J, Caro Y, Grondin I, Petit T (2018) Sugarcane vinasse processing: toward a status shift from waste to valuable resource. A review. J Water Process Eng 24:11–25. https://doi.org/10.1016/j.jwpe.2018.05.003
Sindhu R, Gnansounou E, Binod P, Pandey A (2016) Bioconversion of sugarcane crop residue for value added products–an overview. Rene Ener 98:203–215. https://doi.org/10.1016/j.renene.2016.02.057
Satyawali Y, Balakrishanan M (2008) Wastewater treatment in molasses-based alcohol distilleries for COD and color removal: a review. Environ Manag 86:481–497. https://doi.org/10.1016/j.jenvman.2006.12.024
Sydney EB, Novak AC, Rosa D, Medeiros ABP, Brar SK, Larroche C, Soccol CR (2018) Screening and bioprospecting of anaerobic consortia for biohydrogen and volatile fatty acid production in a vinasse based medium through dark fermentation. Process Biochem 67(1–7):1–7. https://doi.org/10.1016/j.procbio.2018.01.012
Mariano AP, Maciel Filho R (2012) Improvements in biobutanol fermentation and their impacts on distillation energy consumption and wastewater generation. Bioenerg Res 5(2):504–514. https://doi.org/10.1007/s12155-011-9172-0
Sydney EB, Larroche C, Novak AC, Nouaille R, Sarma SJ, Brar SK, Letti AJ, Soccol VT, Soccol CR (2014) Economic process to produce biohydrogen and volatile fatty acids by a mixed culture using vinasse from sugarcane ethanol industry as nutrient source. Bioresour Technol 159:380–386. https://doi.org/10.1016/j.biortech.2014.02.042
Matsakas L, Gao Q, Jansson S, Rova U, Christakopoulos P (2017) Green conversion of municipal solid wastes into fuels and chemicals. Electron J Biotechnol 26:69–83. https://doi.org/10.1016/j.ejbt.2017.01.004
Kamdem I, Hiligsmann S, Vanderghem C, Jacquet N, Tiappi FM, Richel A, Jackes P, Thonart P (2018) Enhanced biogas production during anaerobic digestion of steam-pretreated lignocellulosic biomass from Williams Cavendish Banana plants. Waste Biomass Valor 9(2):175–185. https://doi.org/10.1007/s12649-016-9788-6
Wu YR, He J (2013) Characterization of anaerobic consortia coupled lignin depolymerization with biomethane generation. Bioresour Technol 139:5–12. https://doi.org/10.1016/j.biortech.2013.03.103
Cisneros-Pérez C, Etchebehere C, Celis LB, Carrillo-Reyes J, Alatriste-Mondragón F, Razo-Flores E (2017) Effect of inoculum pretreatment on the microbial community structure and its performance during dark fermentation using anaerobic fluidized-bed reactors. Int J Hydrog Energy 42(15):9589–9599. https://doi.org/10.1016/j.ijhydene.2017.03.157
Siegert I, Banks C (2005) The effect of volatile fatty acid additions on the anaerobic digestion of cellulose and glucose in batch reactors. Process Biochem 40(11):3412–3418. https://doi.org/10.1016/j.procbio.2005.01.025
Krieg NR, Staley JT, Brown DR, Hedlund BP, Paster BJ, Ward NL, Ludwig W, Whitman WB (2010) The Bacteroidetes, Spirochaetes, Tenericutes (Mollicutes), Acidobacteria, Fibrobacteres, Fusobacteria, Dictyoglomi, Gemmatimonadetes, Lentisphaerae, Verrucomicrobia, Chlamydiae, and Planctomycetes, 2nd edn. Springer, New York, pp 25–469
Lee SH, Park JH, Kim SH, Yu BJ, Yoon JJ, Park HD (2015) Evidence of syntrophic acetate oxidation by Spirochaetes during anaerobic methane production. Bioresour Technol 190:543–549. https://doi.org/10.1016/j.biortech.2015.02.066
Kundu K, Sharma S, Sreekrishnan TR (2017) Influence of process parameters on anaerobic digestion microbiome in bioenergy production: towards an improved understanding. Bioenerg Res 10(1):288–303. https://doi.org/10.1007/s12155-016-9789-0
Wongwilaiwalin S, Mhuantong W, Tangphatsornruang S, Panichnumsin P, Champreda V, Tachaapaikoon C (2016) Isolation of cellulolytic microcosms from bagasse compost in co-digested fibrous substrates. Biomass Convers Biorefin 6(4):421–426. https://doi.org/10.1007/s13399-016-0199-5
Xia C, Kumar A, Chen X, Tucker M, Liang Y (2018) Conversion of corn stover hydrolysates to acids: comparison between Clostridium carboxidivorans P7 and microbial communities developed from lake sediment and an anaerobic digester. Biomass Convers Biorefin 8(1):169–178. https://doi.org/10.1007/s13399-017-0239-9
APHA (2012) Standard methods for water and wastewater examination, 19th edn. Am Public Health Assoc, Washington DC
Dubois M, Gilles A, Hamilton JK, Rebers PA, Smith F (1956) Colorimetric method of determination of sugars and related substances. Anal Chem 28:350–355. https://doi.org/10.1021/ac60111a017
Lay JJ, Li Y, Noike T (1997) Influences of pH and moisture content on the bmethane production in high-solids sludge digestion. Water Res 31(6):1518–1524. https://doi.org/10.1016/S0043-1354(96)00413-7
Bates ST, Berg-Lyons D, Caporaso JG, Walters WA, Knight R, Fierer N (2011) Examining the global distribution of dominant archaeal populations in soil. The ISME Journal 5(5):908–917. https://doi.org/10.1038/ismej.2010.171
Schmieder R, Edwards R (2011) Quality control and preprocessing of metagenomic datasets. Bioinformatics. 27(6):863–864. https://doi.org/10.1093/bioinformatics/btr026
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461. https://doi.org/10.1093/bioinformatics/btq461
Cole JR, Wang Q, Fish JA, Chai B, Mcgarrell DM, Sun Y, Brown CT, Porras-Alfaro A, Kuske CR, Tiedje J (2013) Ribosomal database project: data and tools for high throughput rRNA analysis. Nucleic Acids Res 42(D1):D633–D642. https://doi.org/10.1093/nar/gkt1244
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
De Santis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72(7):5069–5072. https://doi.org/10.1128/AEM.03006-05
Kiyuna LSM, Fuess LT, Zaiat M (2017) Unraveling the influence of the COD/sulfate ratio on organic matter removal and methane production from the biodigestion of sugarcane vinasse. Bioresour Technol 232:103–112. https://doi.org/10.1016/j.biortech.2017.02.028
Shin JD, Han SS, Eom KC, Sung SH, Park SW, Kim HO (2008) Predicting methane production potential of anaerobic co-digestion of swine manure and food waste. Environ Engineer Res 13(2):93–97. https://doi.org/10.4491/eer.2008.13.2.093
Dias MF, Colturato LF, Oliveira JP, Leite LR, Oliveira G, Chernicharo CA, Araújo JC (2016) Metagenomic analysis of a desulphurisation system used to treat biogas from vinasse methanisation. Bioresour Technol 205:58–66. https://doi.org/10.1016/j.biortech.2016.01.007
Cruz-Salomón A, Meza-Gordillo R, Rosales-Quintero A, Ventura-Canseco C, Lagunas-Rivera S, Carrasco-Cervantes J (2017) Biogas production from a native beverage vinasse using a modified UASB bioreactor. Fuel. 198:170–174. https://doi.org/10.1016/j.fuel.2016.11.046
Khan MA, Ngo HH, Guo WS, Liu Y, Nghiem LD, Hai FI, Deng LJ, Wang J, Wu Y (2016) Optimization of process parameters for production of volatile fatty acid, biohydrogen and biomethane from anaerobic digestion. Bioresour Technol 219:738–748. https://doi.org/10.1016/j.biortech.2016.08.073
Li H, Chang J, Liu P, Fu L, Ding D, Lu Y (2015) Direct interspecies electron transfer accelerates syntrophic oxidation of butyrate in paddy soil enrichments. Environ Microbiol 17(5):1533–1547. https://doi.org/10.1111/1462-2920.12576
Harmsen HJM, Van Kuijk BLM, Plugge CM, Akkermans ADL, de Vos WM, Stams AJM (1998) Syntrophobacter fumaroxidans sp. nov., a syntrophic propionate-degrading sulfate-reducing bacterium. Int J Syst Bacteriol 48:1383–1387. https://doi.org/10.1099/00207713-48-4-1383
Liu Y, Balkwill DL, Aldrich HC, Drake GR, Boone DR (1999) Characterization of the anaerobic propionate-degrading syntrophs Smithella propionica gen. nov. sp. nov. and Syntrophobacter wolinii. Int J Syst Bacteriol 49:545–556. https://doi.org/10.1099/00207713-49-2-545
Wallrabenstein C, Hauschild E, Schink B (1994) Pure culture and cytological properties of Syntrophobacter wolinii. FEMS Microbiol Lett 123:249–254. https://doi.org/10.1111/j.1574-6968.1994.tb07232.x
Chen S, Liu X, Dong X (2005) Syntrophobacter sulfatireducens sp. nov., a novel syntrophic, propionate-oxidizing bacterium isolated from UASB reactors. Int J Syst Evol Microbiol 55:1319–1324. https://doi.org/10.1099/ijs.0.63565-0
Zellneri G, Busmann A, Rainey FA, Diekmann H (1996) A syntrophic propionate-oxidizing, sulfate-reducing bacterium from a fluidized bed reactor. Syst Appl Microbiol 19(3):414–420. https://doi.org/10.1016/S0723-2020(96)80071-3
Hajarnis SR, Ranade DR (1994) Effect of propionate toxicity on some methanogens at different pH values and in combination with butyrate. Proceeding of 7th International Symposium on Aerobic Digestion. Cape Town, South Africa
Den Boer E, Łukaszewska A, Kluczkiewicz W, Lewandowska D, King K, Reijonen T, Kuhmonen T, Jääskeläinen A, Heitto A, Laatikainen R, Hakalehto E (2016) Volatile fatty acids as an added value from biowaste. Waste Manag 58:62–69. https://doi.org/10.1016/j.wasman.2016.08.006
Fei X, Zekkos D, Raskin L (2015) Archaeal community structure in leachate and solid waste is correlated to methane generation and volume reduction during biodegradation of municipal solid waste. Waste Manag 36:184–190. https://doi.org/10.1016/j.wasman.2014.10.027
Gomaa MA, Abed RMM (2017) Potential of fecal waste for the production of biomethane, bioethanol and biodiesel. J Biotechnol 253:14–22. https://doi.org/10.1016/j.jbiotec.2017.05.013
Morris BE, Henneberger R, Huber H, Moissl-Eichinger C (2013) Microbial syntrophy: interaction for the common good. FEMS Microbiol Rev 37(3):384–406. https://doi.org/10.1111/1574-6976.12019
Luo G, Angelidaki I (2014) Analysis of bacterial communities and bacterial pathogens in a biogas plant by the combination of ethidium monoazide, PCR and ion torrent sequencing. Water Res 60:156–163. https://doi.org/10.1016/j.watres.2014.04.047
Sundberg C, Al-Soud WA, Larsson M, Alm E, Yekta SS, Svensson BH, Sorensen SJ, Karlsson A (2013) 454 pyrosequencing analyses of bacterial and archaeal richness in 21 full-scale biogas digesters. FEMS Microbiol Ecol 85(3):612–626. https://doi.org/10.1111/1574-6941.12148
Nelson MC, Morrison M, Yu Z (2011) A meta-analysis of the microbial diversity observed in anaerobic digesters. Bioresour Technol 102(4):3730–3739. https://doi.org/10.1016/j.biortech.2010.11.119
Vartoukian SR, Palmer RM, Wade WG (2007) The division ‘Synergistes. Anaerobe. 13(3–4):99–106. https://doi.org/10.1016/j.anaerobe.2007.05.004
Jumas-Bilak E, Roudiere L, Marchandin H (2009) Description of ‘Synergistetes’ phyl. nov. and emended description of the phylum ‘Deferribacteres’ and of the family Syntrophomonadaceae, phylum ‘Firmicutes. Int J Syst Evol Microbiol 59(5):1028–1035. https://doi.org/10.1099/ijs.0.006718-0
Kröber M, Bekel T, Diaz NN, Goesmann A, Jaenicke S, Krause L, Miller D, Viehover P, Puhler A, Runte KJ, Schlüter A (2009) Phylogenetic characterization of a biogas plant microbial community integrating clone library 16S-rDNA sequences and metagenome sequence data obtained by 454-pyrosequencing. J Biotechnol 142:38–49. https://doi.org/10.1016/j.jbiotec.2009.02.010
Juste-Poinapen NM, Turner MS, Rabaey K, Virdis B, Batstone DJ (2015) Evaluating the potential impact of proton carriers on syntrophic propionate oxidation. Sci Rep 5:18364. https://doi.org/10.1038/srep18364
Yang L, Huang Y, Zhao M, Huang Z, Miao H, Xu Z, Ruan W (2015) Enhancing biogas generation performance from food wastes by high-solids thermophilic anaerobic digestion: effect of pH adjustment. Int Biodeterior Biodegradation 105:153–159. https://doi.org/10.1016/j.ibiod.2015.09.005
Krause L, Diaz NN, Edwards RA, Gartemann KH, Krömeke H, Neuweger H, Puhler A, Runte KJ, Schluter A, Stoye J, Tauch A, Goesmann A, Szczepanowski R (2008) Taxonomic composition and gene content of a methane-producing microbial community isolated from a biogas reactor. J Biotechnol 136(1–2):91–101. https://doi.org/10.1016/j.jbiotec.2008.06.003
Nishiyama T, Ueki A, Kaku N, Watanabe K, Ueki K (2009) Bacteroides graminisolvens sp. nov., a xylanolytic anaerobe isolated from a methanogenic reactor treating cattlewaste. Int J Syst Evol Microbiol 59(8):1901–1907. https://doi.org/10.1016/j.watres.2018.04.043
Ueki A, Abe K, Kaku N, Watanabe K, Ueki K (2008) Bacteroides propionicifaciens sp. nov., isolated from rice-straw residue in a methanogenic reactor treating waste from cattle farms. Int J Syst Evol Microbiol 58(2):346–352. https://doi.org/10.1099/ijs.0.65486-0
Kampmann K, Ratering S, Kramer I, Schmidt M, Zerr W, Schnell S (2012) Unexpected stability of Bacteroidetes and Firmicutes communities in laboratory biogas reactors fed with different defined substrates. Appl Environ Microbiol, AEM-06394 78:2106–2119. https://doi.org/10.1128/AEM.06394-11
Campanaro S, Treu L, Kougias PG, Luo G, Angelidaki I (2018) Metagenomic binning reveals the functional roles of core abundant microorganisms in twelve full-scale biogas plants. Water Res 140:123–134. https://doi.org/10.1016/j.watres.2018.04.043
Sieber JR, Sims DR, Han C, Kim E, Lykidis A, Lapidus AL, McInerney MJ (2010) The genome of Syntrophomonas wolfei: new insights into syntrophic metabolism and biohydrogen production. Environ Microb 12(8):2289–2301. https://doi.org/10.1111/j.1462-2920.2010.02237.x
Fuess LT, Garcia ML (2014) Implications of stillage land disposal: a critical review on the impacts of fertigation. J Environ Manag 145:210–229. https://doi.org/10.1016/j.jenvman.2014.07.003
Garrity GM, Holt JG (2001) The road map to the manual. In: Bergey’s manual of systematic bacteriology. Springer New York, New York, pp 119–166
Traversi D, Villa S, Acri M, Pietrangeli B, Degan R, Gilli G (2011) The role of different methanogen groups evaluated by real-time qPCR as high-efficiency bioindicators of wet anaerobic co-digestion of organic waste. AMB Express 1(1):28
Wang P, Wang H, Qiu Y, Ren L, Jiang B (2017) Microbial characteristics in anaerobic digestion process of food waste for methane production–a review. Bioresour Technol 248:29–36. https://doi.org/10.1016/j.biortech.2017.06.152
De Vrieze J, Hennebel T, Boon N, Verstraete W (2012) Methanosarcina: the rediscovered methanogen for heavy duty biomethanation. Bioresour Technol 112:1–9. https://doi.org/10.1016/j.biortech.2012.02.079
Raskin L, Zheng DD, Griffin ME, Stroot PG, Misra P (1995) Characterization of microbial communities in anaerobic bioreactors using molecular probes. Anton Leeuw 68:297–308
Riviere D, Desvignes V, Pelletier E, Chaussonnerie S, Guermazi S, Weissenbach J, Sghir A (2009) Towards the definition of a core of microorganisms involved in anaerobic digestion of sludge. ISME J 3(6):700–714. https://doi.org/10.1038/ismej.2009.2
Tsushima I, Yoochatchaval W, Yoshida H, Araki N, Syutsubo K (2010) Microbial community structure and population dynamics of granules developed in expanded granular sludge bed (EGSB) reactors for the anaerobic treatment of low-strength wastewater at low temperature. J Environ Sci Heal A 45(6):754–766. https://doi.org/10.1080/10934521003651531
Acknowledgments
Luiz Gustavo dos Anjos Borges thanks PEGA/PUCRS. We also thank High-Performance Computing Lab - LAD/PUCRS for allowing access to run the high-throughput computational analyses.
Funding
This study was financially supported by the PETROBRAS.
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Iltchenco, J., Almeida, L.G., Beal, L.L. et al. Microbial consortia composition on the production of methane from sugarcane vinasse. Biomass Conv. Bioref. 10, 299–309 (2020). https://doi.org/10.1007/s13399-019-00426-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13399-019-00426-0