Abstract
The CRISPR/Cas9 genome editing system has been in the spotlight compared to programmable nucleases such as ZFNs and TALENs due to its simplicity, versatility, and high efficiency. CRISPR/Cas9 has revolutionized plant genetic engineering and is broadly used to edit various plants' genomes, including those transformation-recalcitrant species such as oil palm. This review will comprehensively present the CRISPR-Cas9 system's brief history and underlying mechanisms. We then highlighted the establishment of the CRISPR/Cas9 system in plants with an emphasis on the strategies of highly efficient guide RNA design, the establishment of various CRISPR/Cas9 vector systems, approaches of multiplex editing, methods of transformation for stable and transient techniques, available methods for detecting and analyzing mutations, which have been applied and could be adopted for CRISPR/Cas9 genome editing in oil palm. In addition, we also provide insight into the strategy of DNA-free genome editing and its potential application in oil palm.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Oil palm is the primary commodity crop in Malaysia, with oil palm planted area in 2022 reaching 5.67 million hectares (Parveez et al. 2023). This golden crop belongs to the family Arecaceae (formally known as Palmeae) of the monocot order Arecales and the genus Elaeis (Ong et al. 2020). There are two species of Elaeis, namely, E. guineensis Jacq. and E. oleifera, originated from West Africa and Latin America, respectively. Although both species can produce edible oil from fruit mesocarp and kernel, only the African oil palm E. guineensis is commercially planted due to its high oil yield. The oil yield per hectare of commercial oil palm is ten times higher than any other oilseed crop, making it the world's most efficient and valuable oil crop (Parveez et al. 2020). The demand for fats and oils derived from oil palm is increasing yearly due to the growing global population. Therefore, improving oil yield is necessary to meet food and industrial requirements. Oil palm improvement through breeding techniques is the key to increasing the oil yield without using more land (Parveez et al. 2015). However, oil palm breeding has a few limitations, such as a narrow genetic base and a highly heterozygous nature, because it is cross-pollinated, perennial, and requires heavy resources (land and other inputs). Therefore, the genetic engineering approach is one of the possible solutions to produce oil palm with high oil productivity, high oil quality, and other desired novel traits, thus ensuring the sustainability of the oil palm industry.
In oil palm genetic engineering, various works have been extensively conducted, such as the identification, characterization, and isolation of promoters and functional genes from oil palm, the construction of numerous transformation vectors, and the optimization of various oil palm transformation methods (Masani et al. 2018, 2022; Fizree et al. 2023). The oil palm genetic transformation was initially started using a biolistic method followed by Agrobacterium, polyethylene-glycol (PEG)-mediated transformation, and DNA microinjection (Parveez et al. 1998; Masli et al. 2009; Masani et al. 2014). Among the target goals of oil palm genetic engineering are producing high oleic acid, high palmitoleic acid, high lycopene, high ricinoleic acid, and biodegradable plastics (Parveez et al. 2015; Masani et al. 2018).
Numerous molecular approaches have been explored to induce mutations in plant species, such as gene silencing by RNA interference; however, this method does not allow for targeted genome editing. Novel genome-editing alternatives known as zinc finger nucleases (ZFNs) and TAL effector nucleases (TALENs) have been discovered (Iqbal et al. 2020). ZFNs and TALENs allow for precise nucleotide modifications of a gene of interest. However, because protein engineering is necessary for editing the gene of interest, implementing both technologies has been time-consuming and expensive (Gaj et al. 2013; Bortesi and Fischer 2015). In 2013, a simple yet versatile solution, the clustered regularly interspaced short palindromic repeat (CRISPR)-associated protein9 (Cas9) or CRISPR/Cas9, for targeted gene editing was discovered (Doudna and Charpentier 2014). Compared to ZFNs and TALENs, the CRISPR/Cas9 system is more practical and cost-effective.
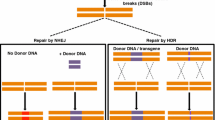
The CRISPR/Cas9 system is a versatile genome-editing technology that uses short RNAs as guides or guide RNA (gRNA) to target multiple genes (Doudna and Charpentier 2014). It was initially discovered as an essential element in Escherichia coli in the 1980s for archaeal and bacterial adaptive immunity (Ishino et al. 1987). However, until it was revealed that Streptococcus thermophilus could develop resistance to a bacteriophage by integrating a genetic segment of an infectious virus into its CRISPR locus, the system's extraordinary function was unknown (Barrangou et al. 2007). This approach relies on a double-stranded break (DSB) driven by a CRISPR RNA (crRNA) transcript to disrupt the target DNA (Bortesi and Fischer 2015). Two forms of DNA repair emerge after DSB of targeted DNA, known as non-homologous end-joining (NHEJ) and homology-directed repair (HDR) pathways. NHEJ is an error-prone mechanism that thus results in poor repair, causing gene function to be disrupted. On the other hand, HDR utilizes a template to repair DNA and makes new DNA complementary to the repair template (Belhaj et al. 2015). Thus far, numerous gene-editing types of research have been accomplished in over 70 different crop species (Timofejeva and Singh 2023), including monocotyledons such as sorghum (Jiang et al. 2013), rice (Shan et al. 2014), maize (Sant’Ana et al. 2020), soybean (Kim and Choi 2021), and dicotyledon plants, such as Populus (Fan et al. 2015), tomato (Nicolia et al. 2021), and pea (Li et al. 2023).
Enhancing plant qualities has always been the prime objective in agriculture genome editing efforts. Although there is a vast array of plant species, the breeding objectives remain comparable. For instance, amplifying the concentrations of select unique secondary metabolites, lengthening the preservation period of fruits, modifying the growth pattern of trees, optimizing yield potential, and fortifying resistance to plant pests and diseases. Through genome editing, these desired traits could be generated by modifying the related genes. For example, the use of CRISPR/Cas9 genome editing has led to the achievement of producing wheat plants with resistance to powdery mildew disease, a condition caused by the fungal pathogen Blumeria graminis (Wang et al. 2014). The utilization of CRISPR/Cas9 technology enables researchers to effectively enhance disease resistance in cucumber (Cucumis sativus) by precisely targeting the gene that encodes translational initiation factor eIF4E (Chandrasekaran et al. 2016). This precision editing results in the development of cucumber plants that possess robust and versatile immunity against various viruses.
Genome editing via CRISPR/Cas9 has also been used to increase crop productivity. For example, CRISPR/Cas9 has been used to simultaneously knock out three major genes that are responsible for regulating the size of the rice grains. The generated plants produced an increase in grain size and weight by up to 20%-30% when compared with the wild type (Xu et al. 2016). Similarly, the CRISPR/Cas9 system was also used to knock out the oleoyl-CoA desaturase (FAD2) gene to produce Camelina sativa plants with an increased oleic acid from 16 to 50%, and concurrently decreased linoleic and linolenic acids to least than 4% and 10%, respectively (Jiang et al. 2017). In addition, in weed control management, CRISPR/Cas9 has been utilized to develop plants that possess tolerance to herbicides. For example, CRISPR/Cas9 targeting the phosphoenolpyruvate-binding site in flax (Linum usitatissimum) produced edited flax lines that are tolerant to glyphosate (Sauer et al. 2016).
In 2013, the publication of the whole-genome sequence of African oil palm species (Singh et al. 2013) provided a wide-ranging chance to transition towards oil palm genome editing. The first report on successful genome editing in oil palm was published by Yeap et al. (2021), targeting the phytoene desaturase (EgPDS) and brassinosteroid-insensitive 1 (EgBRI1) genes. Based on the report, 62.5% to 83.33% mutation efficiencies were obtained using the biolistic transformation of oil palm immature embryos with CRISPR/Cas9 vectors. The report demonstrated that the CRISPR/Cas9 gene-editing system could induce site-targeted mutagenesis in the oil palm genome. The successful adaptation of the CRISPR/Cas9 system was later reported by Bahariah et al. (2023), targeting the genes responsible for increasing oleic acid content in oil palm. Later, the strategy in designing and selecting efficient gRNAs for oil palm study was reported by Jamaludin et al. (2023), marking a closer step to using CRISPR/Cas9 in oil palm genetic engineering.
Even though attempts are made towards an efficient CRISPR/Cas9 system, various plant species may differ in terms of their efficiencies and challenges (Cardi et al. 2023). The difficulties in oil palm transformation due to the size and complexity of genomic may raise the risk of off-target mutations and reduce gene-editing specificity. The delivery of CRISPR/Cas9 reagents into oil palm tissues could be further hampered by its poor transformation efficiency coupled with long plant generation time. Therefore, developing strategies that ensure efficient expression of large numbers of different gRNAs simultaneously from a single vector would allow more extensive use of the multiplex capability of the CRISPR/Cas9 system in oil palm. Implementation from available CRISPR/Cas9 protocols that have been developed in other evolutionary closely related species, such as rice, can also be adopted. Applying a DNA-free CRISPR/Cas9 system via ribonucleoprotein (RNP) in oil palm can avoid the red tape involving genetically modified organism (GMO) regulations. Therefore, this review mainly discusses the strategies and applications of CRISPR/Cas9 genome editing in plants to identify the workable CRISPR/Cas9 system for oil palm that could also lead to DNA-free gene-edited oil palm for a sustainable future.
Discovery of CRISPR/Cas9 genome editing system
Three decades ago, in 1987, a group of researchers in Japan discovered unknown DNA sequences made up of short repeating nucleotides flanked by a short segment (Ishino et al. 1987). In 2002, the strange bacterial sequence was identified as clustered regularly interspaced short palindromic repairs (CRISPR) and CRISPR-associated protein 9 (Cas9) or CRISPR/Cas9, a versatile, highly potential tool for genome editing (Jansen et al. 2002). Using whole-genome sequencing (WGS) technology to study CRISPR revealed that this system was commonly found in viruses and plasmids, thus possibly encoding for bacterial immunity system (Jansen et al. 2002). Consequent biochemical and genetic studies validated the previous speculation, demonstrating that the CRISPR/Cas9 system is involved in detecting and protecting mobile genetic elements (Barrangou et al. 2007; Charpentier and Doudna 2013).
The Cas9 gene was later discovered to work with the gRNA to locate and cut the invading DNA at the conserved region, also known as the proto-spacer adjacent motifs (PAM) sequence (Carroll 2012). Further research discovered that crRNA and trans-activating CRISPR RNA (tracrRNA) were required for Cas9 to form a functional complex for CRISPR/Cas9 activity (Deltcheva et al. 2011). Intriguingly, this dual CRISPR RNA and trans-activating CRISPR RNA can be recreated as gRNA that is enough to drive Cas9 to induce DSB in target DNA (Jinek et al. 2012). The true potential of genome editing applications using the CRISPR/Cas9 system fully emerged when research revealed that the Cas9 guided by the gRNA could work in various types of cells and organisms and cut the genome with high specificity (Charpentier and Doudna 2013). Subsequently, when DSB occurred due to the action of Cas9, two DNA repair mechanisms, NHEJ or HDR, were activated. Both pathways will result in DNA modification at the repair or target sites. Fast-forward to the discovery of the CRISPR/Cas9 system, this technology has been advanced vigorously, focussing on efficiency, enhancing specificity, and expanding its application in various organisms, including plants.
CRISPR/Cas9 versatility, architecture, and mechanisms
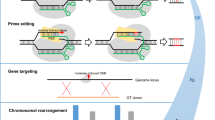
The Streptococcus pyogenes Cas9 or SpCas9 gene is a 4107 bp full-length coding region originating from the type II prokaryotic CRISPR adaptive immune system (Wu et al. 2014). It consists of two separate nuclease domains, namely RuvC and HNH domains. Both can cleave one of the target DNA strands and simultaneously generate a blunt-ended DSB. The gRNA is a fusion of crRNA and tracrRNA. It is crucial for CRISPR activity to recognize and cleave the target genomic sequence and maintain stability, thus improving the CRISPR/Cas9 system (Okada et al. 2022). The target recognition sequence, including PAM, will be bound with the Cas9 protein to form a Cas9/gRNA complex. This complex specificity towards its target site vitally depends on the hybridization of the target recognition sequence (without PAM) of gRNA to that specific target site. During the process, the PAM sequence acts as the binding signal for Cas9. The target sequence will initially recognize and move towards its NGG PAM sequence, allowing the Cas9/gRNA complex to locate its target DNA site (Barman et al. 2020). The complex will then separate the target DNA strands to facilitate the complementary base-pairing of gRNA with the target DNA strand. The RuvC and HNH domains will then cut off one of the two strands at three bases upstream of PAM, subsequently generating blunt-ended DSB. The DSB will be repaired either by an error-prone mechanism, so-called non-homologous end-joining (NHEJ), or an error-free mechanism, namely a homology-directed repair pathway (HDR) (Fig. 1). Both pathways can be used to induce gene modifications at the target loci. However, the choice of repair mechanism depends on various factors, including the stage of the cell cycle, the nature of DSB, and the availability of the repair templates (Ceccaldi et al. 2016).
Schematic illustration of CRISPR/Cas9 genome editing mechanisms. The Cas9–gRNA complex binds any genomic region with a PAM sequence (shown in pink). The 20 nucleotides of the gRNA sequence (without PAM) complement the genomic sequence immediately upstream of PAM (red). The Cas9 endonuclease will cleave the DNA genomic region at three nucleotides upstream of PAM (indicated by scissor), introducing the double-strand break (DSB). The DSB will be repaired by either an error-prone NHEJ pathway or HDR, an error-free mechanism. The target gene would be knockout if the indels were introduced at the exon or gene promoter. In contrast, if the HDR pathway repairs the DSB, gene replacement will be introduced to the genomic region adjacent to the DSB
Selection of the target gene for the establishment of the CRISPR/Cas9 system
Selection of the target gene is based on a case-by-case basis, mainly depending on the objectives of the research and the characteristics of the target gene, including the mutant’s phenotyping efficiency and the gene copy number (Shan et al. 2020). The target gene sequence and genomic structure identification, such as intron–exon structure in the target plants, can be obtained from a BLAST search using the available transcriptomic data (Shan et al. 2018). Further sequence confirmation and additional information on the corresponding genomic sequence can be obtained using PCR amplification and sequencing of the target amplicons. This information will facilitate the identification and selection of suitable gRNA. For an initial demonstration of CRISPR/Cas9 principles, loss-of-function gene activity is commonly being used by targeting the gene that produces a mutant with an obvious change in the phenotype, i.e., knockout of phytoene desaturase (PDS) that produces mutants with the albino phenotype (Charrier et al. 2019; Dai et al. 2021) (Table 1). This type of study has been applied as the first CRISPR/Cas9 application in various plant species, including oil palm (Yeap et al. 2021; Jamaludin et al. 2023), because it is more straightforward than targeting the gene controlling the complex traits.
Furthermore, the efficiency of CRISPR/Cas9 genome editing can also be evaluated by inducing mutation in the exogenous gene (DNA or gene originating from outside of the studied organism, also known as transgene) in the transgenic mutant (Kishi-Kaboshi et al. 2017). For example, Bhowmik et al. (2018) have designed gRNAs to target the red fluorescent protein gene from Discosoma coral (DsRED) in transgenic wheat microspores. With the availability of the DsRED sequences, the gRNA sequence can be efficiently designed. The absence of the red fluorescence signals indicated the successful knockout of the DsRED gene by CRISPR/Cas9 in transgenic wheat microspores. For oil palm, a series of DsRED gene constructs was developed and successfully tested in oil palm calli as a visual reporter marker (Fizree et al. 2019) and could be potentially used to evaluate CRISPR/Cas9 genome editing in oil palm.
Parameters for an efficient gRNA design
In CRISPR/Cas9 genome editing, designing a gRNA for gene targeting is the first step in CRISPR/Cas9 experiments (Doench et al. 2014). The sequence of gRNA can significantly affect the target DNA's cleavage efficiency, which also influences the probability of off-target binding and Cas9 cleavage (Zischewski et al. 2017). Therefore, designing the correct gRNA is critical for the success of CRISPR/Cas9 experiments. There are several important parameters to consider when designing a gRNA. The binding of each Cas nuclease to the target sequence happens only in the presence of a PAM sequence (Wu et al. 2014). For that reason, the target sites in the genome can be targeted by different Cas nuclease proteins that are restricted by the positions of particular PAM sequences (Molla and Yang 2020). This is because each Cas nuclease isolated from different bacterial species recognizes a different sequence of PAM (Kleinstiver et al. 2015). For example, SpCas9 nuclease cleaves the nucleotide upstream of the PAM sequence 5'-NGG-3' (where "N" can be any nucleotide base), while PAM sequence 5'-NNGRR(N)-3' is required by SaCas9 (Staphylococcus aureus) to target DNA regions for CRISPR editing (Ran et al. 2015). Although the PAM sequence itself plays an essential role in the cleavage of target sequences, it must not be included in the sequence of a gRNA.
Doench et al. (2014), Wu et al. (2014), Wong et al. (2015), Liang et al. (2016), and Bruegmann et al. (2019) reported that the presence of specific motifs in the gRNA sequence and gRNA secondary structure could enhance the activity of the gRNA in CRISPR/Cas9 studies (Fig. 2). For example, there should be favorable nucleotides at several positions. Guanine was preferred directly after the PAM motif (position 20) but unfavored at position 16 (Doench et al. 2014). In addition, cytosine was unfavorable at positions 3 and 20 of the gRNA sequence but favored at position 16 over guanine (Wu et al. 2014). Furthermore, adenine was also preferred in the middle of the gRNA sequence (Doench et al. 2014), and purine residue was preferred in the last gRNA nucleotide sequence (Bruegmann et al. 2019). On the other hand, all gRNAs having TTTT (poly-T) were also disfavoured, because a stretch of identical contiguous nucleotides will be recognized by U6 promoters and terminate the RNA polymerase III transcription (Wong et al. 2015). An early study on non-functional gRNA motifs also suggested that repetitive uracil (UUU) in the seed region of gRNA (10–12 nucleotides before the PAM sequence) or at the last six bases of the gRNA could also impair the CRISPR/Cas9 activity (Wu et al. 2014). Other than repetitive UUU, four repetitive G nucleotides were also disfavoured, because it was reported that the GGGG could significantly lead to low CRISPR/Cas9 efficiency. This GGGG was also prone to forming a guanine tetrad. This secondary structure causes the gRNA sequence to be less accessible for gRNA target recognition and binding (Wong et al. 2015).
The characteristics of efficient gRNAs based on the 20-bp guide sequence and gRNA secondary structure analysis. These characteristics were based on research of highly active gRNAs reported by Doench et al. (2014), Wu et al. (2014), Wong et al. (2015), Liang et al. (2016), and Bruegmann et al. (2019)
Another critical aspect of the high activity of gRNA is the secondary structure, which also affects the efficiency of the designed gRNA. Extensive studies reported that some structural motifs were identified to influence the effectiveness of gRNA positively (Doench et al. 2014; Wong et al. 2015). Although the current gRNA structure is based on bacterial crRNA and tracrRNA, which have been broadly used in eukaryotes, it has been shown that the gRNA secondary structure is prone to structural changes that may affect the CRISPR/Cas9 activity. Therefore, gRNA secondary structures should be analyzed to select the efficient gRNA that can produce high editing efficiency using the CRISPR/Cas9 system. The secondary structure analysis of designed gRNA can be done using the bioinformatics tool such as RNA fold Webserver by applying the Zuker–Stiegler algorithms and Andronescu model based on minimum free energy (MFE) representation (Bruegmann et al. 2019).
Typically, the secondary structure of a gRNA includes a crRNA sequence that encompasses 20 nucleotides of gRNA and a repeat region comprising 12 nucleotides, accompanied by a tracrRNA sequence obtained from a combination of an anti-repeat region and three stem-loop structures (Liang et al. 2016). By linking together, the artificial tetraloop structures form a single RNA transcript structure. The repeat and anti-repeat section of the tetraloop are commonly called the RAR stem-loop (GAAA), stem-loop 1 (CUAG), stem-loop 2 (GAAA), and stem-loop 3(AGU) (Jamaludin et al. 2023). Doench et al. (2014) and Wong et al. (2015) reported that the last three base pairs of the gRNA (seed region), 18–20 nucleotide position of the gRNA sequence, and 51–53 nucleotide position on the secondary structure should be unpaired and freely accessible to allow the binding of Cas9 to the target sequence.
The base-pairing of the guide sequence with its target DNA can be impacted by a stable complex between the gRNA sequence and other nucleotide bases (Liang et al. 2016). There are three types of base-pairing scores, namely, the total base-pairing scores (TBP), consecutive base-pairing scores (CBP), and internal base-pairing scores (IBP). The TBP score measures the total base pairs in the gRNA guide sequence combined with other sequences, whereas the CBP score represents the count of bases in the guide sequence combined with other sequences. Meanwhile, the internal base pair in the gRNA is indicated by the IBP score. To ensure efficiency in selecting gRNA, Liang et al. (2016) emphasized the importance of considering sequences with less than 12 TBP scores, a maximum of 7 CBP, and 6 IBP.
Based on these guidelines, Bahariah et al. (2023) have designed four gRNA for targeting palmitoyl-ACP-thioesterase (EgPAT) and oleoyl-CoA desaturase (EgFAD2) genes in an attempt to alter these genes for potential increase in oil palm oleic acid content. The study revealed that gRNA with high GC content of 60% and 65% could induce 100% indels rate. By considering these crucial motifs and preferable secondary gRNA structure, seven efficient gRNAs were selected from 167 gRNAs predicted by CRISPR-GE software to completely down-regulate the expression of oil palm phytoene desaturase (EgPDS) (Jamaludin et al. 2023). These seven gRNA sequences consisted of specific motifs such as adenine at position 9–11, no cytosine at position 3, and purine residue in the last 4 nucleotides, with the secondary structure having stem-loop 2 and stem-loop 3. The In vitro cleavage assay demonstrated that these seven sgRNA were efficient for cleavaging the corresponding target regions of the EgPDS gene.
Type of vectors for CRISPR/Cas9 system
Besides selecting the high-efficiency gRNA, the type of CRISPR/Cas9 vector system used can also affect the CRISPR/Cas9 editing efficiency. The efficiency of the CRISPR/Cas9 system varies amongst species, depending on the regulatory elements used, such as the species-based codon-optimized Cas9, promoters, and terminators for regulating the expression of Cas9 gene and gRNA sequence. An efficient vector system is generally required to produce a high expression level of gRNA and Cas9 genes, particularly for oil palm that is recalcitrant to genetic transformation. In the CRISPR/Cas9 vector system, two different RNA polymerase systems are employed; RNA polymerase II (Pol II) is responsible for driving the expression of the Cas9 gene, while RNA polymerase III (Pol III) controls the expression of gRNA. RNA polymerase III promoters, such as U3 or U6 small nuclear RNA (snRNA) gene promoters, are typically utilized to transcribe gRNAs in cells (Zhang et al. 2017) (Table 1). To initiate transcription, these Pol III promoters depend on a highly specific 5′ nucleotide, where the U6 promoter necessitates 5′-Guanine (G) and the U3 promoter demands 5′-Adenine (A) (Jiang et al. 2013). It is usually followed by a poly ‘T’ (five-to-eight Ts) that functions as a transcription termination signal (Kor et al. 2023). Thus, enhanced specificity can be achieved by including a specific nucleotide at the 5′ end of the gRNA sequences.
The OsU3 and OsU6 promoters from Oryza are commonly used for the monocots plants (Zafar et al. 2020; Bahariah et al. 2021, 2023; Tripathi et al. 2021; Yeap et al. 2021). Meanwhile, AtU6 and AtU3 promoters isolated from Arabidopsis thaliana are broadly used for the dicot plant (Jang et al. 2021; Pavese et al. 2021; Zhang et al. 2021a; Ly et al. 2023). The U6 promoters from Arabidopsis, Oryza, and Camelina were more efficient than their U3 counterparts (Zhou et al. 2023). In wheat (Triticum aestivum L.), the TaU3 promoter exhibited higher efficiency compared to the TaU6, OsU3, and OsU6 promoters (Zhang et al. 2021b; Kor et al. 2023). In barley (Hordeum vulgare), the HvU3 promoter showed higher activity than the OsU3 promoter (Lee et al. 2021). Therefore, it is recommended to isolate U3 and U6 from oil palm and experimentally evaluate the compatibility and efficiency of these promoters to regulate gRNA before further analysis is made.
Nevertheless, it is noteworthy that these U6 and U3 promoters may not always work for all targeted genes because of the absence of spatiotemporal specific control, which is ubiquitously present and expressed in all cells, tissues, and at all stages of plant growth and development (Xie and Yang 2013). For cloning multiple gRNA into the vector, different U3 or U6 promoters are recommended to regulate the gRNAs. This minimizes the risk of transgene silencing (Ma et al. 2015). Otherwise, the polycistronic tRNA–gRNA strategy can be utilized (Xie et al. 2015). In addition, for the gRNA scaffold, the long version of the gRNA scaffold (76 nucleotides) was shown to have higher efficiency compared to the short version (42 nucleotides) (Hsu et al. 2013). Both scaffolds are commonly used in most CRISPR/Cas9 studies in plants, because they are easier to design and show higher efficiency than the dual crRNA:tracrRNA system (Miao et al. 2013).
Aside from that, for Cas9, various plant constitutive promoters were utilized (Ma et al. 2016). This includes the CaMV35S promoter from the virus and Ubiquitin promoters from several plant species, such as maize, rice, and Arabidopsis (Zhang and Showalter 2020). It was reported that the maize ubiquitin promoter was more efficient for CRISPR/Cas9 editing in both monocot and dicot plants than the CaMV35S promoter from cauliflower mosaic virus (Feng et al. 2018). One factor contributing to this is the CaMV35S promoters reduced activity in embryogenic cells and vulnerability to transgene silencing (Jiang et al. 2014).
Nevertheless, the CaMV35S and maize Ubiquitin promoters have been used for regulating the Cas9 gene due to their ability to drive high transgene expression in oil palm cells efficiently (Yeap et al. 2021; Bahariah et al. 2023; Jamaludin et al. 2023). It also recommended employing oil palm endogenous constitutive promoters for driving the Cas9 gene, such as ubiquitin extension protein 2 (UEP2), which was demonstrated to regulate the transgene in oil palm tissues well (Fizree et al. 2019; Masura et al. 2019). Other promoters such as Arabidopsis egg-cell (E.C) specific promoters, including E.C 1.1 and E.C 1.2 (DD45), pollen-specific promoters such as LAT52, sporogenous cell-specific promoters such as SPL, and the Yao promoter were utilized to express the Cas9 gene in the early developmental stages, thus, increase the chance of obtaining homozygous and heritable mutations (Zhang and Showalter 2020).
To improve Cas9 expression in plants, the Cas9 gene was also modified with plant codons by attaching the nuclear localization signal (NLS) at both ends of Cas9 for gene editing in various crops, including oil palm (Yeap et al. 2021). It was reported that up to 83.33% mutation efficiency was achieved when oil palm codon-optimized Cas9 gene coupled with gRNA targeting EgPDS gene were transformed in oil palm immature embryos. However, no comparison was made to determine the editing efficiency between the SpCas9 gene and the oil palm codon-optimized Cas9 gene. The NLS is commonly used to transport the Cas9 gene into the nucleus of eukaryotes. According to reports, the codon-optimized Cas9 gene resulted in greater editing effectiveness in various plant species when compared to the original Cas9 gene (Jiang et al. 2013; Nekrasov et al. 2013; Ma et al. 2015). For instance, the codon-optimized Cas9 gene consisting of 50–400 bp GC-rich sequence, mimicking the Gramineae genes, was proven to be highly efficient in rice (Ma et al. 2015).
Multiplex CRISPR/Cas9 genome editing system
Previous CRISPR/Cas9 systems in plant protoplasts have shown the utilization of a single guide, with mutation frequencies of 0.1% in grape, 6.9% in apple, 19% in rice, and 44% in tobacco (Woo et al. 2015; Malnoy et al. 2016). This approach depends on a single gRNA to target a gene of interest by inducing a single DSB at the targeted locus and relies on the NHEJ to cause a frameshift mutation. However, this gene-knockout study is predicted to be less efficient in oil palm. Because oil palm is a diploid crop species with low transformation efficiency, multiple gRNAs may be required to edit the oil palm gene efficiently. Multiplex genome editing system consists of multiple gRNAs with distinct target sequences that bind to individual Cas9 to form the Cas9/gRNA complexes and subsequently bind and cleave the target genes in the same cell simultaneously (Cong et al. 2013).
In early works of CRISPR/Cas9 in plants, gRNA delivery was performed by co-transformation using several plasmids, such as in Arabidopsis, rice, and tomato (Timofejeva and Singh 2023). The method is tedious, even though multiple-site mutations were successfully achieved in these species. This type of strategy also can decrease the effectiveness of CRISPR/Cas9 for multiple gene editing in plant genomes. Therefore, various vector systems with multiplexing abilities were introduced. For example, the pYLCRISPRCas9 vector system designed by Ma et al. (2015) allows the assembly of multiple gRNA expression cassettes into a single vector system using golden gate cloning or the Gibson assembly method. This vector system is being used to develop a CRISPR/Cas9 system in oil palm (Bahariah et al. 2023; Jamaludin et al. 2023). In Bahariah et al.’s (2023) work, two CRISPR/Cas9 constructs carrying two gRNAs, targeting EgPAT or EgFAD2 gene, and one CRISPR/Cas9 construct consisting of all four gRNAs were successfully constructed. Meanwhile, depending on the position of seven gRNAs in the EgPDS genomic sequence, Jamaludin et al. (2023) successfully created two CRISPR/Cas9 constructs, assembling two gRNAs and one CRISPR/Cas9 construct with three gRNAs.
Alternatively, an endogenous tRNA-processing RNase, Csy4 endoribonucleases, and ribozymes were also engineered to control multiple gRNAs from a single transcript so-called polycistronic transcript system (Xie et al. 2015; Cermak et al. 2017). These multiple gRNAs were separated by the spacer sequences in the constructed plasmid. The functional gRNAs will be released due to the cleaving of the primary transcript by the respective enzymes (tRNA, Csy4, or ribozymes) at spacer sequences–gRNA junctions. Therefore, these polycistronic transcript systems can be utilized for multiplex genome editing in various living organisms, including plants (Ma et al. 2019; Huang et al. 2020; Li et al. 2021; Yang et al. 2022). Subsequently, it enables gene editing of multiple members of a gene family, genes with multiple functionalities involving a complex trait, or multiple sites of single genes. These polycistronic transcript systems are beneficial to editing oil palm genes involved in agronomical valuable traits usually regulated by multiple genes.
Transformation methods of CRISPR/Cas9 vector into plants
CRISPR/Cas9 genome editing in the plant required the delivery of DNA constructs consisting of Cas9 and gRNA into the cells. The delivery or transformation methods used in the recent applications of the CRISPR/Cas9 system in oil palm and other species are shown in Table 1. Yeap et al. (2021) reported the successful CRISPR/Cas9 gene editing in oil palm by targeting the EgBRI1 gene, which was determined using the biolistic transformation in immature embryos. Mutations were observed at the target sites of the EgBRI1 gene in oil palm shoots displaying dwarf phenotypes. Meanwhile, Bahariah et al. (2023) demonstrated that the biolistic transformation of embryogenic calli successfully produced oil palm plantlets with an edited EgPAT gene.
Other than that, Agrobacterium-mediated transformation can be another option for the delivery of CRISPR/Cas9 constructs into plant cells. This transformation method has been widely used for stable integration of CRISPR/Cas9 into plant genomes in many crop species, including apple (Charrier et al. 2019), banana (Zhang et al. 2022), potato (Ly et al. 2023), and plants listed in Table 1. Ali et al. (2015) and Yin et al. (2015) reported an improved Agrobacterium transformation utilizing the T-DNA embedded viral replicon, which was reported to enhance the NHEJ and HDR-mediated genome editing compared to the conventional T-DNA in tomato (Cermak et al. 2015). For oil palm, Bahariah et al. (2023) revealed a 28% mutation rate obtained through Agrobacterium-mediated transformation compared with a 6% mutation rate when embryogenic calli were transformed using biolistic. This finding suggested that Agrobacterium-mediated transformation is more efficient for altering the oil palm genome and, more importantly, for subsequently generating non-GMO gene-edited oil palms through crossing and segregation procedures.
Other than using protoplasts via polyethylene-glycol (PEG)-mediated transformation or callus via particle bombardment and Agrobacterium, immature embryos are currently being used for CRISPR/Cas9 studies. Additionally, immature embryos can be used for a DNA-free transformation system using RNP or mRNA (Svitashev et al. 2016; Liang et al. 2017). On the other hand, a new technology utilizing cell-penetrating peptides or CPPs has also been introduced. This strategy used infiltration of CPPs, positively charged short peptides that can move across cellular membranes of mature plant tissue. Interestingly, this technology has been shown to be capable of binding site-specific nucleases (Rádis-Baptista et al. 2017).
Recently, an efficient transformation method, namely a cut-dip-budding (CBD) delivery system, was developed to allow CRISPR/Cas9 genome editing in plants without going through the tissue culture process (Cao et al. 2022). This system delivered T-DNA of CRISPR/Cas9 gene construct from Agrobacterium strain into plant cells by infecting explants that can be generated to transformed shoots and subsequently to genome edited plants which was demonstrated in plants that recalcitrant to the genetic transformation such as dandelion Taraxacum kok-saghyz (Cao et al. 2023) and three succulent varieties Kalanchoe blossfeldiana, Crassula arborescens, and Sansevieria trifasciata (Lu et al. 2024) (Table 1). Hence, it would be intriguing to investigate a CBD delivery system for potentially editing the oil palm genome without requiring tissue culture.
Strategies of transient assay for developing CRISPR/Cas9 system
The use of transient assays to investigate the efficiency of the CRISPR/Cas9 system is highly recommended before utilizing the system in stable transformation (Shan et al. 2020). This is because stable transformation of recalcitrant plants such as oil palm is laborious and time-consuming compared to transient assays, which are more convenient and quicker. For now, two transient assay strategies are widely being used: protoplast transfection and leaf-cell agroinfiltration (Nekrasov et al. 2013; Xie and Yang 2013) (Fig. 3A). For example, Liang et al. (2017) reported using the transient system to test CRISPR/Cas9 efficiency in wheat protoplasts. Furthermore, for protoplast transfection, plant regeneration from the transfected protoplasts is also possible in some species, such as potatoes, tobacco, and lettuce (Andersson et al. 2018; Lin et al. 2018; Choi et al. 2022).
Strategies of transient assays and screening of mutation for CRISPR/Cas9 genome editing system. A Protoplast transient assay and agroinfiltration are commonly used for CRISPR/Cas9 proof-of-concept studies in new target species. B The CRISPR-induced mutation can be detected using SURVEYOR or T7EI assay, which uses enzymes sensitive to mismatched double-stranded DNA sequences. If indel is present, the heteroduplex containing the unpaired nucleotide (mismatch) will be cut by the SURVEYOR/T7EI enzyme, as shown in the gel image. C PCR/RE assay is based on the availability of the restriction enzyme in the CRISPR/Cas9 target site. The wild-type alleles will be digested in the presence of the restriction enzyme site. However, if the alleles are mutated, the restriction enzyme site is destroyed; therefore, the uncleaved DNA fragments will appear on the gel image, indicating the targeted mutagenesis by the CRISPR/Cas9 system
In the protoplast transient assay, the steps include the isolation of protoplasts, transfection with CRISPR/Cas9 constructs, and protoplast culture. Generally, enzymes such as macerozyme and cellulase are used to remove plant cell walls to isolate protoplasts. The CRISPR/Cas9 constructs are delivered into protoplasts via various techniques, including electroporation, polyethylene-glycol (PEG), or microinjection (Lin et al. 2018; Wu et al. 2020; Zhou et al. 2020). Because of its simplicity and speed of the transformation process, the protoplast transient assay is an appealing model for evaluating the mutagenesis efficiency of a CRISPR/Cas9 system, particularly in assessing efficient gRNA. The appropriate expression of the CRISPR/Cas9 in the transfected protoplasts will potentially produce targeted editing in the sample (Andersson et al. 2018; Fan et al. 2020; Yu et al. 2021). For oil palm, the functionality of oil palm codon-optimized Cas9, and the effectiveness of the designed gRNAs were confirmed by an electroporation-mediated protoplast transient system (Yeap et al. 2021). The study revealed that oil palm codon-optimized Cas9 successfully promoted cleavage frequency of up to 25.49% in protoplasts transformed with gRNA targeting the EgPDS gene. Meanwhile, a significant result of 21% editing efficiency generating large DNA fragment deletion of 304 bp was obtained when oil palm protoplasts were transformed with CRISPR/Cas9 construct targeting EgFAD2 gene via PEG-mediated transformation (Bahariah et al. 2023).
Besides, protoplasts can be transformed with RNP comprising Cas9 and gRNA only (Svitashev et al. 2016; Badhan et al. 2021) (Table 2), which can avoid the integration of foreign DNA, resulting in non-transgenic plants. For example, using PEG-mediated delivery, Cas9/gRNA ribonucleoprotein was used to generate transgene-free lettuce (Woo et al. 2015) and potato (Andersson et al. 2018). On the other hand, the agroinfiltration method required the suspension of Agrobacterium containing T-DNA plasmid expressing the gRNA and Cas9 genes. Vacuum infiltration or direct injection is used to introduce the Agrobacterium culture into the plant’s tissues (usually leaves). The CRISPR/Cas9 components can be transferred through transfer DNAs (T-DNAs) into the host plant genome. Genome editing processes might then occur in the transformed cells. Numerous plant species, including Nicotiana, Solanum, and Lactuca, have established protocols for efficient and routine agroinfiltration (Wroblewski et al. 2005). Transformed cells can easily be detected, because the T-DNA segment contains a visual reporter gene such as green fluorescent protein (GFP), as Jiang et al. (2013) reported. Additionally, various researches have been published utilizing the Agrobacterium rhizogenes-mediated hairy root transformation for rapid analysis of CRISPR/Cas9 efficiency in many legume species (Belhaj et al. 2015; Barman et al. 2020).
Screening of mutation and identification of mutant’s genotype
Several methods are used to detect the CRISPR/Cas9-induced mutation in various species, including oil palm (Table 1). These methods include polymerase chain reaction/restriction enzyme (PCR/RE) assays, T7 endonuclease I assay (T7EI), SURVEYOR nuclease assay, high-resolution melting (HRM) analysis-based assay, and PAGE-based genotyping assay (Shan et al. 2014; Thomas et al. 2014; Zhu et al. 2014). Although these assays are commonly used to screen for induced mutation, they have limitations, such as labour-intensive, time-consuming, sequence-limited, and expensive. The SURVEYOR assay and T7EI assay are commonly being used to detect mutation in CRISPR-treated samples, because these methods are more straightforward, cheaper, and applicable for any target sequence as both depend on the detection and digestion of mismatched heteroduplex DNA; however, their detection sensitivity is very low (Fig. 3B). In contrast, the PCR/RE assay is more reliable in mutation detection with a more straightforward methodology but requires restriction enzyme sites near the target sequences (Cong et al. 2013; Shan et al. 2014) (Fig. 3C). The method of PAGE-based genotyping assay is time-consuming and insensitive as it includes the denaturation and annealing of the PCR amplicons containing the induced mutations and detection using the native PAGE (Zhu et al. 2014). The HRM analysis-based technique is more efficient than PAGE-based assay as it depends on the melting temperature (Tm) distinction between the edited and non-edited PCR amplicons; however, it requires expensive equipment (Thomas et al. 2014).
Various software tools have been developed to detect and identify the CRISPR/Cas9 site-targeted mutation using Sanger sequencing data by analyzing the chromatogram files of Sanger sequenced-PCR products. The software tools are freely accessible and allow simultaneous analysis of multiple chromatograms in a short time. The most frequent software used are ICE (Interference of CRISPR Edits), TIDE (Tracking of indels by decomposition), TIDER (Tracking of Insertion, Deletions, and Recombination events), DsDecode (The Degenerate Sequence Decode), and CRISPR-Detector (Table 1). For example, ICE is the preferred software to quantify the presence of indels in the samples consisting of edited and unedited DNA mixtures, especially at low editing efficiency (Hasley et al. 2021). In addition, ICE analysis of Sanger sequencing data has shown a comparable or higher accuracy than next-generation sequencing (NGS). ICE used the untransformed or control sequences as the wild-type genomic reference to detect the presence of indels in the transformed samples. For oil palm, Bahariah et al. (2023) used ICE to evaluate the editing efficiency, indels type, and mutation genotype for samples derived from protoplasts transformed via PEG-mediated transformation, and embryogenic calli transformed via Agrobacterium- and biolistic-mediated transformations. The report emphasizes that ICE was a suitable software tool for the detection of mutations in gene-edited oil palm compared with other available software.
On the other hand, Yeap et al. (2021) employed TIDE for the detection of mutations in oil palm. TIDE is an assay that accurately determines the identity of indels and the frequency of induced mutagenesis in a cell pool by analyzing two resulting raw sequences obtained from one pair of standard PCR and Sanger sequencing reactions (Brinkman and Van Steensel 2019). TIDER is an upgraded version of TIDE that can predict the frequency of targeted mutagenesis, especially for small nucleotide changes induced by CRISPR homology-directed repair that uses a donor template (CRISPR-HDR) (Brinkman and Van Steensel 2019). TIDER software requires an additional sequencing trace, which can be prepared using a simple two-step PCR reaction. The DsDecode program can automatically decode the sequencing chromatograms with homozygous mutations, heterozygous or biallelic, into allelic sequences (Xie et al. 2019). This software can simultaneously read up to 30 chromatograms in a few minutes. For analyzing whole-genome sequence (WGS), which consists of both small and large datasets, CRISPR-Detector uses FASTQ sequencing files from control and CRISPR-treated samples (Huang et al. 2023). This software will process the data by aligning and sorting the reads, identifying indels, substitutions, and structural variants (SVs) while removing the genetic background and finalizing output to present the results.
The term mutation frequency is often used to describe the percentage of CRISPR/Cas9-induced mutation regenerated plants in which the targeted mutation can be detected at the locus or loci of interest (Shan et al. 2020). Previously, high mutation frequencies have been reported in rice and Arabidopsis (Ma et al. 2015). The genotype of the regenerated mutants depends on the time for the targeted CRISPR/Cas9-mutagenesis event to occur. Suppose a mutagenesis event happens before the first embryogenic cell divides. In that case, that is, in the early phase of plant regeneration and the locus on only one of the two sister chromatids was mutated, a diploid plant may be heterozygous or homozygous if both alleles were mutated with the same mutation or biallelic if both alleles were mutated. However, the breaks were repaired, resulting in different alleles (Shan et al. 2020).
However, in many cases, CRISPR/Cas9-targeted mutagenesis often occurs later in plant development and independently in different tissues (Jarvise et al. 2021). Thus, it produces chimeric plants with cells of different genotypes consisting of wild type, homozygous, heterozygous, or biallelic (Song et al. 2022). Early gene function studies can be conducted if the CRISPR/Cas9-induced homozygous and biallelic mutations occur in the first generation. This has been reported in tomato and rice (Brooks et al. 2014; Zhang et al. 2014). However, suppose the regenerated plants do not exhibit homozygous or biallelic mutations in the first generations or in the primary transformants. In that case, the phenotype analysis for loss-of-function activity can be analyzed in the later generation (Xu et al. 2015). This has been demonstrated in various species, including wheat (Wang et al. 2014) and Brassica napus (Yang et al. 2017), which obey the Mendelian heritability.
Proof of concept study of CRISPR/Cas9 system using PDS Gene
Establishing a CRISPR/Cas9 system in perennial plants such as oil palm can be more challenging than the annual plants. Many critical parameters, such as the ploidy level, genome heterozygosity, growth cycle, and physiological characteristics of the species, need to be carefully considered (De Bruyn et al. 2020). A high ploidy level will cause an increased workload for this gene-editing system to edit all copies of the target gene (Shan et al. 2020). Another challenge in introducing CRISPR/Cas9 study in oil palm is the long growth and complex physiology. These two factors are related, because long-growth cycle plants are usually perennials, primarily woody plants such as oil palms. Compared with small or annual plants, oil palms have a very long regeneration time; thus, the phenotyping of mutants can be complex. In addition, outcrossing and evaluating the inheritance of mutated alleles in the subsequent generations of most trees are tricky due to its dioecious nature (Bewg et al. 2018). Therefore, utilizing a target gene that can produce specific phenotypes at early plant development is advisable to identify the workable CRISPR/Cas9 system in oil palm (Bahariah et al. 2023; Jamaludin et al. 2023).
Phytoene desaturase (PDS) is the most utilized marker gene for CRISPR proof of principle studies (Shan et al. 2018). A marker gene is one in which mutants produce an obvious phenotype that allows easy visual screening. The PDS gene encodes an essential plant enzyme in the carotenoid biosynthetic pathway. It catalyzes the formation of one of the double bonds during the conversion of phytoene into lycopene. Therefore, additional copies of the PDS gene may increase the plant carotenoid content (Steinbrenner and Sandmann 2006). However, the most exciting criterion of the PDS gene is that it can be used as a photo-bleaching marker in plants, which can be easily exploited for proof-of-concept studies using CRISPR/Cas9. Silencing the PDS gene could disrupt the biosynthesis of carotenoid, chlorophyll, and gibberellins and produce albino plants (Jamaludin et al. 2023). Several studies have demonstrated the utilization of the PDS gene in various crops to test the efficiency of the CRISPR/Cas9 system (Fan et al. 2015; Yeap et al. 2021). In addition, PDS is typically a single copy gene in plant genomes, including oil palm, which is another advantage in its application.
Yeap et al. (2021) employed the EgPDS gene as a model gene to establish the CRISPR/Cas9 system in oil palm before applying the approach targeting the EgBRI1 gene. The group described the development of a transient protoplast assay to quickly define the efficiency of the CRISPR/Cas9 system in oil palm by targeting the EgPDS gene. From the five gRNAs tested, only two gRNAs, gPDS4 and gPDS5, successfully induced mutations in oil palm protoplasts with editing efficiencies of 83.33% and 62.50%, respectively. Using gPDS4 and gPDS5, they successfully generated oil palm shoots with chimeric albinism phenotype (Yeap et al. 2021). Unlike Yeap et al. (2021), who used single gRNA CRISPR/Cas9 constructs, Jamaludin et al. (2023) generated CRISPR/Cas9 constructs consisting of multiple gRNAs to modify the EgPDS gene in oil palm. However, no data from transformation works were reported in the study.
The model plant, N. tabacum, is an allotetraploid with a genome size of 4.5 G. It consists of high (> 70%) content of repetitive DNA (Sierro et al. 2014). It has become an essential model plant species for the study of CRISPR/Cas9 genome editing. Although N. tabacum contains four PDS genes (two each from the N. sylvestris and N. tomentosiformis progenitors), at which mutation of all four genes is necessary to obtain albino plants, various research has been successfully conducted utilizing tobacco as a model plant system for CRISPR/Cas9 study (Jiang et al. 2013; Nekrasov et al. 2013). The successful regeneration of the albino tobacco plantlet from the CRISPR/Cas9 transfected protoplast was also reported (Lin et al. 2018).
Oil Palm future prospect via CRISPR-RNP system
CRISPR/Cas9 is commonly implemented in plants to enhance agricultural qualities due to its ability to knock out genes. However, this procedure often involves transgenic intermediates that may raise legislation concerns and probably cause public rejection. Therefore, direct delivery of Cas9 protein and gRNA that can eliminate the integration of transgenes and reduce the risk of off-target activity, insertional mutagenesis, and immune response is more favorable (He and Zhao 2020). Previous research has shown successful delivery and CRISPR/Cas9 editing using laboratory-transcribed Cas9 and gRNAs into plant tissues (Woo et al. 2015; Klimek-Chodacka et al. 2021). This protein-based delivery system is also known as CRISPR–ribonucleoprotein (RNP) or CRISPR–RNP complex. This system requires two essential components in the formulation, the gRNA and the Cas9 protein, combined using a simple laboratory method to form a negatively charged recombinant Cas9 and gRNA or RNP complex. The RNP complex will then be delivered into the plant cell as one single reagent. In the cells, the RNP complex is susceptible to being degraded by both proteases and RNases, thus making it the most unstable format (Lin et al. 2022). However, for genome editing applications, the CRISPR/Cas9 reagents are only temporarily needed until the intended mutagenesis has happened; the presence of the CRISPR reagents is no longer favored as it can lead to off-target activity. Therefore, due to the rapid degradation of RNP in the cell, this format is more desirable to decrease the potential risk of off-target activity (Subburaj et al. 2022).
In contrast, when plasmid DNA is transformed into plant cells, the Cas9 gene may be randomly incorporated into the plant genome. The constitutive expression may increase off-target activity (Liang et al. 2018). These off-target effects can bring about the legislation concerns about GMO. The plasmid DNA or CRISPR/Cas9 expression vectors are usually delivered into the plant cells, where the Cas9 needs to be expressed first and then pre-assembled with the gRNA to form a complex, to finally able to cleave the target region, produce double-strand breaks and then, generate indels during the repair mechanisms. Unlike plasmid-based delivery system, RNP complexes only have to be pre-assembled by incubating the Cas9 protein with gRNA for 15 min at 37ºC in the laboratory (Klimek-Chodacka et al. 2021). The reaction produced a functional molecular complex that can be used directly for cellular genome editing. The RNP entered the nucleus using the NLS-fused Cas9, which is commonly used for RNP. Consequently, the action of RNP is more specific and quicker, since this complex can work immediately without needing intracellular transcription and translation in the genome (Liang et al. 2017). For these reasons, RNP is considered the most efficient CRISPR/Cas9 genome editing strategy for plants, including oil palm.
The delivery of RNP into plant tissues can be categorized into two approaches: physical approaches, such as microinjection, biolistic, and membrane deformation, or synthetic carrier approaches, such as lipid nanoparticles, inorganic nanoparticles, and nanogels (Zhang et al. 2021c). The PEG-mediated transformation has been the most commonly used technique to deliver RNP into the plant’s cells, including crop species such as apple and grapevine (Malnoy et al. 2016), rubber (Fan et al. 2020), banana (Wu et al. 2020), and recently in tomato (Lin et al. 2022), papaya (Elias et al. 2023), and oil palm (Norfaezah et al. 2024). PEG caused the protoplast to clump, thus promoting the proximal interaction between RNP and the cell surface (Masani et al. 2014). Previous research on protoplast PEG-mediated transformation has shown that an average of 10% editing rate was obtained using RNP (Svitashev et al. 2016; Andersson et al. 2018). Andersson et al. (2018) also reported that delivery of RNP using protoplast PEG-mediated transformation was efficient in editing all four copies of the PDS gene in 2%-3% of the regenerated shoots in potato species. Protoplast PEG-mediated transformation was also proven efficient in other crops, such as grapevine, apple, potatoes, lettuce, soybean, petunia, and wheat (Woo et al. 2015; Malnoy et al. 2016; Kim et al. 2018; Subburaj et al. 2022).
For oil palm, editing efficiencies of 63.6% up to 100% were obtained when protoplasts were transformed with multiple RNP, two combinations of RNP (RNP1/RNP2 or RNP3/RNP4), and three combinations of RNP (RNP5/RNP6/RNP7), targeting EgPDS gene (Norfaezah et al. 2024). The high editing efficiencies achieved in the study were due to the established efficient protocol for the preparation of RNP and protoplast transformation, including the two most influenced parameters, the amount of Cas9 protein and heat-treatment applied to transformed protoplasts. However, implementing protoplast PEG-mediated transformation in those plants is challenging due to the absence of regeneration protocols, and even if the protocols are available, the regeneration rate is meagre, such as in oil palm (Masani et al. 2013). Masani et al. (2014) also introduced the protoplast microinjection system for oil palm transformation work. However, no RNP delivery application through oil palm protoplast microinjection has yet been reported.
The Cas9 gene and gRNA can also be transformed directly into plant cells via particle bombardment. The RNP is coated with gold or tungsten microparticles before being bombarded into the target cell walls or membrane using helium shock (Stewart et al. 2018). This strategy produced high frequencies of mutated alleles in maize embryo cells, as Svitashev et al. (2016) reported. This strategy has also efficiently produced transgenic rice with inherited mutation (Shan et al. 2014). Optimization study of particle-bombardment delivery techniques into the plant cells using gold-activated silica nanoparticles has been described in detail by Martin-Ortigosa and Wang (2014). Hamada et al. (2017) also reported the utilization of in-planta particle bombardment (iPB) targeting mature plant tissue to avoid the regeneration of immature cells in wheat. This strategy is beneficial for those species that cannot be regenerated through the protoplast system but are amenable to callus regeneration. This approach was also successfully utilized in other species, such as wheat (Liang et al. 2017) and rice (Banakar et al. 2019).
Moreover, RNP can also be delivered via protoplast electroporation. This strategy exploited the disruption of cell membranes using an electrical pulse. Subsequently, producing temporary nanopores on the plant membranes will allow the transportation of RNP biomolecules (Boukany et al. 2011). Altogether, the DNA-free mutants produced using CRISPR-RNP technology are undifferentiated from the natural species. Therefore, they could be exempted from GMO legislation. Thus, CRISPR-RNP could be the new direction of futuristic breeding for sustainable oil palm agriculture. The summary of recent CRISPR-RNP applications in various plant species is shown in Table 2.
Conclusion and prospects
The advancement in CRISPR/Cas9 technology will enormously facilitate oil palm basic research, breeding, and functional studies of target genes. The development of CRISPR/Cas9 systems that are workable in oil palm would provide an easy-to-use platform to address diverse questions in oil palm functional genomics, such as the study of physiological traits, optimization of certain beneficial metabolic pathways, and genotype–phenotype relationship of oil palm genes. Site-specific mutagenesis by CRISPR/Cas9 system will enable deeper accessibility to gene expression study of various genes and targeting multiple loci despite the genomic complexity of the oil palm genome, which was previously difficult to access using the conventional molecular methods. Various platforms are being developed to increase the editing efficiency and specificity of the CRISPR/Cas9 system (Fig. 4), subsequently providing unprecedented potential for genetic studies of oil palm agronomical and economically beneficial traits. This includes higher proportions of desaturated fatty acids in palm oil, slow height increments, low lipase activity in mature fruits, improved nutritional values of palm oil, higher resistance to disease, and better environmental adaptability. The application of DNA-free genome editing, such as CRISPR-RNP, to produce DNA-free gene-edited oil palm may become an efficient strategy to address issues in current GMO regulations and public acceptance. This genome editing system will play a pivotal role in the sustainable agricultural development of oil palm.
Various approaches have been developed for establishing efficient and robust CRISPR/Cas9 gene editing system in oil palm (Masani et al. 2022; Fizree et al. 2023; Yeap et al. 2021; Bahariah et al. 2023; Jamaludin et al. 2023; Norfaezah et al. 2024). These approaches, (1) identification of suitable target genes and bioinformatics tools for designing efficient gRNA, (2) simple and efficient protocol to synthesise gRNA molecules and development of in-house robust CRISPR/Cas9 vector system, (3) development of multiplex genome editing system for editing valuable target traits such as oil palm palmitoyl-ACP thioesterase (EgPAT) and oleoyl-CoA desaturase (EgFAD2) genes for increasing oleic acid content, oil palm gibberellin acid 20 oxidase (EgGA20ox) genes for reducing tree height increment and oil palm virescens (EgVIR) genes for generating virescens fruits type, (4) efficient transformation, selection, and plant regeneration system for delivery plasmid-based CRISPR/Cas9 into embryogenic calli and protoplasts, (5) efficient RNP-based transformation and plant regeneration system utilizing oil palm embryogenic calli and protoplasts, and (6) evaluation of suitable screening method for detecting mutations in transformed plants, are in pipeline for developing a DNA-free genome editing protocol for oil palm improvements
Abbreviations
- Cas9:
-
CRISPR-associated protein9
- CBD:
-
Cut-dip-budding
- CBP:
-
Consecutive base-pairing
- CRISPR:
-
Clustered regularly interspaced short palindromic repeat
- crRNA:
-
CRISPR RNA
- DSB:
-
Double-stranded break
- GMO:
-
Genetically modified organism
- gRNA:
-
Guide RNA
- HDR:
-
Homology-directed repair
- IBP:
-
Internal base-pairing
- NHEJ:
-
Non-homologous end-joining
- NLS :
-
Nuclear localization signal
- PAM:
-
Proto-spacer adjacent motifs
- PDS:
-
phytoene desaturase
- PEG:
-
Polyethylene-glycol
- RNP:
-
Ribonucleoprotein
- TALENs:
-
TAL effector nucleases
- TBP:
-
Base-pairing
- tracrRNA:
-
Trans-activating CRISPR RNA
References
Ali Z, Abul-Faraj A, Li L, Ghosh N, Piatek M, Mahjoub A, Aouida M, Piatek A, Baltes NJ, Voytas DF, Dinesh-Kumar S, Mahfouz MM (2015) Efficient virus-mediated genome editing in plants using the CRISPR/Cas9 system. Mol Plant 8(8):1288–1291. https://doi.org/10.1016/j.molp.2015.02.011
Andersson M, Turesson H, Olsson N, Fält AS, Ohlsson P, Gonzalez MN, Samuelsson M, Hofvander P (2018) Genome editing in potato via CRISPR-Cas9 ribonucleoprotein delivery. Physiol Plant 164(4):378–384. https://doi.org/10.1111/ppl.12731
Badhan S, Ball AS, Mantri N (2021) First report of CRISPR/Cas9 mediated DNA-free editing of 4CL and RVE7 genes in chickpea protoplasts. Int J Mol Sci 22(1):396. https://doi.org/10.3390/ijms22010396
Bahariah B, Masani MYA, Rasid OA, Parveez GKA (2021) Multiplex CRISPR/Cas9-mediated genome editing of the FAD2 gene in rice: a model genome editing system for oil palm. J Gen Eng Biotechnol 19(1):1–13
Bahariah B, Masani MYA, Fizree MPMAA, Rasid OA, Parveez GKA (2023) Multiplex CRISPR/Cas9 gene-editing platform in oil palm targeting mutations in EgFAD2 and EgPAT genes. J Gen Eng Biotechnol 21(1):3. https://doi.org/10.1186/s43141-022-00459-5
Banakar R, Eggenberger AL, Lee K, Wright DA, Murugan K, Zarecor S, Dill CJL, Sashital DG, Wang K (2019) High-frequency random DNA insertions upon co-delivery of CRISPR-Cas9 ribonucleoprotein and selectable marker plasmid in rice. Sci Rep 9(1):1–13. https://doi.org/10.1038/s41598-019-55681-y
Barman A, Deb B, Chakraborty S (2020) A glance at genome editing with CRISPR–Cas9 technology. Curr Genet 66(3):447–462. https://doi.org/10.1007/s00294-019-01040-3
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315(5819):1709–1712. https://doi.org/10.1126/science.1138140
Belhaj K, Chaparro-Garcia A, Kamoun S, Patron NJ, Nekrasov V (2015) Editing plant genomes with CRISPR/Cas9. Curr Opin Biotechnol 32:76–84. https://doi.org/10.1016/j.copbio.2014.11.007
Bewg WP, Ci D, Tsai CJ (2018) Genome editing in trees: from multiple repair pathways to long-term stability. Front Plant Sci 9:1732. https://doi.org/10.3389/fpls.2018.01732
Bhowmik P, Ellison E, Polley B, Bollina V, Kulkarni M, Ghanbarnia K, Song H, Gao C, Voytas DF, Kagale S (2018) Targeted mutagenesis in wheat microspores using CRISPR/Cas9. Sci Rep 8(1):6502. https://doi.org/10.1038/s41598-018-24690-8
Bortesi L, Fischer R (2015) The CRISPR/Cas9 system for plant genome editing and beyond. Biotechnol Adv 33(1):41–52. https://doi.org/10.1016/j.biotechadv.2014.12.006
Boukany PE, Morss A, Liao WC, Henslee B, Jung H, Zhang X, Yu B, Wang X, Wu Y, Li L, Gao K, Hu X, Zhao X, Hemminger O, Lu W, Lafyatis GP, Lee LJ (2011) Nanochannel electroporation delivers precise amounts of biomolecules into living cells. Nat Nanotechnol 6(11):747–754. https://doi.org/10.1038/nnano.2011.164
Brinkman EK, Van Steensel B (2019) Rapid quantitative evaluation of CRISPR genome editing by TIDE and TIDER. CRISPR Gene Edit Methods Protocol. https://doi.org/10.1007/978-1-4939-9170-9_3
Brooks C, Nekrasov V, Lippman ZB, Van Eck J (2014) Efficient gene editing in tomato in the first generation using the clustered regularly interspaced short palindromic repeats/CRISPR-associated9 system. Plant Physiol 166(3):1292–1297. https://doi.org/10.1104/pp.114.247577
Bruegmann T, Deecke K, Fladung M (2019) Evaluating the efficiency of gRNAs in CRISPR/Cas9 mediated genome editing in poplars. Int J Mol Sci 20(15):3623. https://doi.org/10.3390/ijms20153623
Cao X, Xie H, Song M, Lu J, Ma P, Huang B, Wang M, Tian Y, Chen F, Peng J, Lang Z, Li G, Zhu JK (2022) Cut-dip-budding delivery system enables genetic modifications in plants without tissue culture. Innovation (camb) 4(1):100345. https://doi.org/10.1016/j.xinn.2022.100345
Cao X, Xie H, Song M, Zhao L, Deng S, Tian Y, Li G, Lang Z, Zhu JK (2023) Extremely simplified cut-dip-budding method for genetic transformation and gene editing in Taraxacum kok-saghyz. The Innovation Life 1(3):100040
Cardi T, Murovec J, Bakhsh A, Boniecka J, Bruegmann T, Bull SE (2023) CRISPR/Cas-mediated plant genome editing: outstanding challenges a decade after implementation. Trends Plant Sci 28(10):1144–1165. https://doi.org/10.1016/j.tplants.2023.05.012
Carroll D (2012) A CRISPR approach to gene targeting. Mol Ther 20(9):1658–1660. https://doi.org/10.1038/mt.2012.171
Ceccaldi R, Sarangi P, D’Andrea AD (2016) The fanconi anaemia pathway: new players and new functions. Nat Rev Mol Cell Biol 17(6):337–349. https://doi.org/10.1038/nrm.2016.48
Cermak T, Baltes NJ, Cegan R, Zhang Y, Voytas DF (2015) High-frequency, precise modification of the tomato genome. Genome Biol 16:1–15. https://doi.org/10.1186/s13059-015-0796-9
Cermak T, Curtin SJ, Gil-Humanes J, Cegan R, Kono TJY, Konecna E, Belanto JJ, Starker CG, Mathre JW, Greenstein RL, Voytas DF (2017) A multipurpose toolkit to enable advanced genome engineering in plants. Plant Cell 29(6):1196–1217. https://doi.org/10.1105/tpc.16.00922
Chandrasekaran J, Brumin M, Wolf D, Leibman D, Klap C, Pearlsman M, Sherman A, Arazi T, Gal-On A (2016) Development of broad virus resistance in non-transgenic cucumber using CRISPR/Cas9 technology. Mol Plant Pathol 17(7):1140–1153. https://doi.org/10.1111/mpp.12375
Charpentier E, Doudna JA (2013) Rewriting a genome. Nature 495(7439):50–51. https://doi.org/10.1038/495050a
Charrier A, Vergne E, Dousset N, Richer A, Petiteau A, Chevreau E (2019) Efficient targeted mutagenesis in apple and first-time edition of pear using the CRISPR-Cas9 system. Front Plant Sci 10:40. https://doi.org/10.3389/fpls.2019.00040
Choi SH, Ahn WS, Jie EY, Cho HS, Kim SW (2022) Development of late-bolting plants by CRISPR/Cas9-mediated genome editing from mesophyll protoplasts of lettuce. Plant Cell Rep 41(7):1627–1630. https://doi.org/10.1007/s00299-022-02875-w
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F (2013) Multiplex genome engineering using CRISPR/Cas systems. Science 339(6121):819–823. https://doi.org/10.1126/science.1231143
Dai X, Yang X, Wang C, Fan Y, Xin S, Hua Y, Wang K, Huang H (2021) CRISPR/Cas9-mediated genome editing in Hevea brasiliensis. Ind Crops Prod 164:113418. https://doi.org/10.1016/j.indcrop.2021.113418
De Bruyn C, Ruttink T, Eeckhaut T, Jacobs T, De Keyser E, Goossens A, Van Laere K (2020) Establishment of CRISPR/Cas9 genome editing in witloof (Cichorium intybus var. foliosum). Front Genome Ed. https://doi.org/10.3389/fgeed.2020.604876
Deltcheva E, Chylinski K, Sharma CM, Gonzales K, Chao Y, Pirzada ZA, Eckert MA, Vogel J, Charpentier E (2011) CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471(7340):602–607. https://doi.org/10.1038/nature09886
Doench JG, Hartenian E, Graham DB, Tothova Z, Hegde M, Smith I, Sullender M, Ebert BL, Xavier RJ, Root DE (2014) Rational design of highly active sgRNAs for CRISPR-Cas9–mediated gene inactivation. Nat Biotechnol 32(12):1262–1267. https://doi.org/10.1038/nbt.3026
Doudna JA, Charpentier E (2014) The new frontier of genome engineering with CRISPR-Cas9. Science 346(6213):1258096. https://doi.org/10.1126/science.1258096
Elias MJ, Hasley J, Tian M, Christopher DA (2023) Development of a mesophyll protoplast-based system for gene editing of papaya. In Vitro Cell Dev Biol Plant 59:517–535. https://doi.org/10.1007/s11627-023-10373-1
Fan D, Liu T, Li C, Jiao B, Li S, Hou Y, Luo K (2015) Efficient CRISPR/Cas9-mediated targeted mutagenesis in Populus in the first generation. Sci Rep 5(1):1–7. https://doi.org/10.1038/srep12217
Fan Y, Xin S, Dai X, Yang X, Huang H, Hua Y (2020) Efficient genome editing of rubber tree (Hevea brasiliensis) protoplasts using CRISPR/Cas9 ribonucleoproteins. Ind Crops Prod 146:112146. https://doi.org/10.1016/j.indcrop.2020.112146
Feng C, Su H, Bai H, Wang R, Liu Y, Guo X, Liu C, Zhang J, Yuan J, Birchler JA, Han F (2018) High-efficiency genome editing using a dmc1 promoter-controlled CRISPR/Cas9 system in maize. Plant Biotechnol J 16(11):1848–1857. https://doi.org/10.1111/pbi.12920
Fizree PMAA, Shaharuddin NA, Ho CL, Masura SS, Manaf MAA, Parveez GKA, Masani MYA (2019) Evaluation of transient DsRED gene expression in oil palm embryogenic calli. Sci Hortic 257:108679. https://doi.org/10.1016/j.scienta.2019.108679
Fizree MPMAA, Masani MYA, Shaharuddin NA, Chai-Ling H, Abd Manaf MA, Parveez GKA (2023) Efficient PEG-mediated transformation of oil palm mesophyll protoplasts and its application in functional analysis of oil palm promoters. S Afr J Bot 155:187–195. https://doi.org/10.1016/j.sajb.2023.02.025
Gaj T, Gersbach CA, Barbas CF (2013) ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 31(7):397–405. https://doi.org/10.1016/j.tibtech.2013.04.004
González MN, Massa GA, Andersson M, Decima Oneto CA, Turesson H, Storani L, Olsson N, Fält AS, Hofvander P, Feingold SE (2021) Comparative potato genome editing: Agrobacterium tumefaciens-mediated transformation and protoplasts transfection delivery of CRISPR/Cas9 components directed to StPPO2 gene. Plant Cell Tissue Organ Cult 145(2):291–305. https://doi.org/10.1007/s11240-020-02008-9
Hamada H, Linghu Q, Nagira Y, Miki R, Taoka N, Imai R (2017) An in planta biolistic method for stable wheat transformation. Sci Rep 7(1):1–8. https://doi.org/10.1038/s41598-017-11936-0
Hasley JAR, Navet N, Tian M (2021) CRISPR/Cas9-mediated mutagenesis of sweet basil candidate susceptibility gene ObDMR6 enhances downy mildew resistance. PLoS ONE 16(6):e0253245. https://doi.org/10.1371/journal.pone.0253245
He Y, Zhao Y (2020) Technological breakthroughs in generating transgene-free and genetically stable CRISPR-edited plants. aBIOTECH 1(1):88–96. https://doi.org/10.1007/s42994-019-00013-x
Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V, Li Y, Fine EJ, Wu X, Shalem O, Cradick TJ, Marraffini LA, Bao G, Zhang F (2013) DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 31(9):827–832. https://doi.org/10.1038/nbt.2647
Huang L, Wang D, Chen H, Hu J, Dai X, Liu C, Li A, Shen X, Qi C, Sun H, Zhang D, Chen T, Jiang Y (2023) CRISPR-detector: fast and accurate detection, visualization, and annotation of genome-wide mutations induced by genome editing events. J Genet Genom 5(8):563–572. https://doi.org/10.1016/j.jgg.2023.03.010
Huang X, Wang Y, Xu J, Wang N (2020) Development of multiplex genome editing toolkits for citrus with high efficacy in biallelic and homozygous mutations. Plant Mol Biol 104(3):297–307. https://doi.org/10.1007/s11103-020-01043-6
Iqbal Z, Iqbal MS, Ahmad A, Memon AG, Ansari MI (2020) New prospects on the horizon: genome editing to engineer plants for desirable traits. Curr Plant Biol 24:100171. https://doi.org/10.1016/j.cpb.2020.100171
Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A (1987) Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol 169(12):5429–5433. https://doi.org/10.1128/jb.169.12.5429-5433.1987
Jamaludin N, Bahariah B, Shaharuddin NA, Ho CL, Rasid OA, Parveez GKA, Masani MYA (2023) Designing gRNAs targeting oil palm phytoene desaturase (EgPDS) gene and construction of vectors for oil palm CRISPR/Cas9 study. J Oil Palm Res 35(1):133–146
Jang HA, Bae EK, Kim MH, Park SJ, Choi NY, Pyo SW, Lee C, Jeong HY, Lee H, Choi YI, Ko JH (2021) CRISPR-knockout of CSE gene improves saccharification efficiency by reducing lignin content in hybrid poplar. Int J Mol Sci 22(18):9750. https://doi.org/10.3390/ijms22189750
Jansen R, Embden JDV, Gaastra W, Schouls LM (2002) Identification of genes that are associated with DNA repeats in prokaryotes. Mol Microbiol 43(6):1565–1575. https://doi.org/10.1046/j.1365-2958.2002.02839.x
Jarvis BA, Romsdahl TB, McGinn MG, Nazarenus TJ, Cahoon EB, Chapman KD, Sedbrook JC (2021) CRISPR/Cas9-induced fad2 and rod1 mutations stacked with fae1 confer high oleic acid seed oil in pennycress (Thlaspi arvense L.). Front Plant Sci 12:652319. https://doi.org/10.3389/fpls.2021.652319
Jiang W, Zhou H, Bi H, Fromm M, Yang B, Weeks DP (2013) Demonstration of CRISPR/Cas9/sgRNA-mediated targeted gene modification in Arabidopsis, tobacco, sorghum and rice. Nucleic Acids Res 41(20):e188. https://doi.org/10.1093/nar/gkt780
Jiang W, Yang B, Weeks DP (2014) Efficient CRISPR/Cas9-mediated gene editing in Arabidopsis thaliana and inheritance of modified genes in the T2 and T3 generations. PLoS ONE 9(6):e99225. https://doi.org/10.1371/journal.pone.0099225
Jiang WZ, Henry IM, Lynagh PG, Comai L, Cahoon EB, Weeks DP (2017) Significant enhancement of fatty acid composition in seeds of the allohexaploid, Camelina sativa, using CRISPR/Cas9 gene editing. Plant Biotechnol J 15(5):648–657. https://doi.org/10.1111/pbi.12663
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 337(6096):816–821. https://doi.org/10.1126/science.1225829
Kim D, Alptekin B, Budak H (2018) CRISPR/Cas9 genome editing in wheat: enhancing quality and productivity for global food security-a review. Funct Integr Genom 23(3):265. https://doi.org/10.1007/s10142-023-01190-1
Kim H, Choi J, Won KH (2020) A stable DNA-free screening system for CRISPR/RNPs-mediated gene editing in hot and sweet cultivars of Capsicum annuum. BMC Plant Biol 20:449. https://doi.org/10.1186/s12870-020-02665-0
Kim H, Choi J (2021) A robust and practical CRISPR/crRNA screening system for soybean cultivar editing using LbCpf1 ribonucleoproteins. Plant Cell Rep 40:1059–1070. https://doi.org/10.1007/s00299-020-02597-x
Kishi-Kaboshi M, Aida R, Sasaki K (2017) Generation of gene-edited Chrysanthemum morifolium using multicopy transgenes as targets and markers. Plant Cell Physiol 58(2):216–226. https://doi.org/10.1093/pcp/pcw222
Kleinstiver BP, Prew MS, Tsai SQ, Nguyen NT, Topkar VV, Zheng Z, Joung JK (2015) Broadening the targeting range of Staphylococcus aureus CRISPR-Cas9 by modifying PAM recognition. Nat Biotechnol 33(12):1293–1298. https://doi.org/10.1038/nbt.3404
Klimek-Chodacka M, Gieniec M, Baranski R (2021) Multiplex site-directed gene editing using polyethylene glycol-mediated delivery of CRISPR gRNA: Cas9 ribonucleoprotein (RNP) complexes to carrot protoplasts. Int J Mol Sci 22(19):10740. https://doi.org/10.3390/ijms221910740
Kor SD, Chowdhury N, Keot AK, Yogendra K, Chikkaputtaiah C, Sudhakar RP (2023) RNA Pol III promoters-Key players in precisely targeted plant genome editing. Front Gen 13:989199. https://doi.org/10.3389/fgene.2022.989199
Lee JH, Won HJ, Hoang NTP, Lee SM, Kim HY, Jung JH (2021) Improving lignocellulosic biofuel production by CRISPR/Cas9-mediated lignin modification in barley. GCB Bioenergy 13(4):742–752. https://doi.org/10.1111/gcbb.12808
Lee MH, Lee J, Choi SA, Kim YS, Koo O, Choi SH, Ahn WS, Jie EY, Kim SW (2020) Efficient genome editing using CRISPR-Cas9 RNP delivery into cabbage protoplasts via electro-transfection. Plant Biotechnol Rep 14:695–702. https://doi.org/10.1007/s11816-020-00645-2
Li G, Liu R, Xu R, Varshney RK, Ding H, Li M, Yan X, Huang S, Li J, Wang D, Ji Y, Wang C, He J, Luo Y, Gao S, Wei P, Zong X, Yang T (2023) Development of an Agrobacterium-mediated CRISPR/Cas9 system in pea (Pisum sativum L.). The Crop J 11(1):132–139. https://doi.org/10.1016/j.cj.2022.04.011
Li J, Zhang S, Zhang R, Gao J, Qi Y, Song G, Li W, Li Y, Li G (2021) Efficient multiplex genome editing by CRISPR/Cas9 in common wheat. Plant Biotechnol J 19(3):427–429. https://doi.org/10.1111/pbi.13508
Liang G, Zhang H, Lou D, Yu D (2016) Selection of highly efficient sgRNAs for CRISPR/Cas9-based plant genome editing. Sci Rep 6(1):1–8. https://doi.org/10.1038/srep21451
Liang Z, Chen K, Li T, Zhang Y, Wang Y, Zhao Q, Liu J, Zhang H, Liu C, Ran Y, Gao C (2017) Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes. Nat Commun 8(1):1–5. https://doi.org/10.1038/ncomms14261
Liang Z, Chen K, Zhang Y, Liu J, Yin K, Qiu JL, Gao C (2018) Genome editing of bread wheat using biolistic delivery of CRISPR/Cas9 in vitro transcripts or ribonucleoproteins. Nat Protoc 13(3):413–430. https://doi.org/10.1038/nprot.2017.145
Lin CS, Hsu CT, Yang LH, Lee LY, Fu JY, Cheng QW, Wu FH, Hsiao HCW, Zhang Y, Zhang R, Chang WJ, Yu CT, Wang W, Liao LJ, Gelvin SB, Shih MC (2018) Application of protoplast technology to CRISPR/Cas9 mutagenesis: from single-cell mutation detection to mutant plant regeneration. Plant Biotechnol J 16(7):1295–1310. https://doi.org/10.1111/pbi.12870
Lin CS, Hsu CT, Yuan YH, Zheng PX, Wu FH, Cheng QW, Wu YL, Wu TL, Lin S, Yue JJ, Cheng YH, Lin SI, Shih MC, Sheen J, Lin YC (2022) DNA-free CRISPR-Cas9 gene editing of wild tetraploid tomato Solanum peruvianum using protoplast regeneration. Plant Physiol 188(4):1917–1930. https://doi.org/10.1093/plphys/kiac022
Liu W, Rudis MR, Cheplick MH, Millwood RJ, Yang JP, Ondzighi-Assoume CA, Montgomery GA, Burris KP, Mazarei M, Chesnut JD, Stewart CN (2020) Lipofection-mediated genome editing using DNA-free delivery of the Cas9/gRNA ribonucleoprotein into plant cells. Plant Cell Rep 39:245–257. https://doi.org/10.1007/s00299-019-02488-w
Lu J, Li S, Deng S, Wang M, Wu Y, Li M, Dong J, Lu S, Su C, Li G, Lang Z, Zhu JK (2024) A method of genetic transformation and gene editing of succulents without tissue culture. Plant Biotechnol J. https://doi.org/10.1111/pbi.14318
Ly DNP, Iqbal S, Fosu-Nyarko J, Milroy S, Jones MG (2023) Multiplex CRISPR-Cas9 gene-editing can deliver potato cultivars with reduced browning and acrylamide. Plants 12(2):379. https://doi.org/10.3390/plants12020379
Ma C, Zhu C, Zheng M, Liu M, Zhang D, Liu B, Li Q, Si J, Ren X, Song H (2019) CRISPR/Cas9-mediated multiple gene editing in Brassica oleracea var. capitata using the endogenous tRNA-processing system. Hortic Res 6:20. https://doi.org/10.1038/s41438-018-0107-1
Ma X, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R, Wang B, Yang Z, Li H, Lin Y, Xie Y, Shen R, Chen S, Wang Z, Chen Y, Guo J, Chen L, Zhao X, Liu YG (2015) A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8(8):1274–1284. https://doi.org/10.1016/j.molp.2015.04.007
Ma X, Zhu Q, Chen Y, Liu YG (2016) CRISPR/Cas9 platforms for genome editing in plants: developments and applications. Mol Plant 9(7):961–974. https://doi.org/10.1016/j.molp.2016.04.009
Malnoy M, Viola R, Jung MH, Koo OJ, Kim S, Kim JS, Velasco R, Kanchiswamy NC (2016) DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front Plant Sci 7:1904. https://doi.org/10.3389/fpls.2016.01904
Martin-Ortigosa S, Wang K (2014) Proteolistics: a biolistic method for intracellular delivery of proteins. Transgenic Res 23(5):743–756. https://doi.org/10.1007/s11248-014-9807-y
Masani MYA, Noll G, Parveez GKA, Sambanthamurthi R, Prüfer D (2013) Regeneration of viable oil palm plants from protoplasts by optimizing media components, growth regulators and cultivation procedures. Plant Sci 210:118–127. https://doi.org/10.1016/j.plantsci.2013.05.021
Masani MYA, Noll GA, Parveez GKA, Sambanthamurthi R, Prüfer D (2014) Efficient transformation of oil palm protoplasts by PEG-mediated transfection and DNA microinjection. PLoS ONE 9(5):e96831. https://doi.org/10.1371/journal.pone.0096831
Masani MYA, Izawati AMD, Rasid OA, Parveez GKA (2018) Biotechnology of oil palm: current status of oil palm genetic transformation. Biocatal Agric Biotechnol 15:335–347. https://doi.org/10.1016/j.bcab.2018.07.008
Masani MYA, Parveez GKA, Noll G, Fizree MPMA, Sambanthamurthi R, Pruefer D (2022) Protoplast isolation and transformation in oil palm. Methods Mol Biol 2464:187–202
Masli DIA, Kadir APG, Yunus AMM (2009) Transformation of oil palm using Agrobacterium tumefaciens. J Oil Palm Res 21:643–652
Masura SS, Parveez GKA, Abd RO (2019) Isolation of an oil palm constitutive promoter derived from ubiquitin extension protein (uep2) gene. J Oil Palm Res 31(1):28–41
Miao J, Guo D, Zhang J, Huang Q, Qin G, Zhang X, Wan J, Gu H, Qu LJ (2013) Targeted mutagenesis in rice using CRISPR-Cas system. Cell Res 23(10):1233–1236. https://doi.org/10.1038/cr.2013.123
Molla KA, Yang Y (2020) Predicting CRISPR/Cas9-induced mutations for precise genome editing. Trends Biotechnol 38(2):136–141. https://doi.org/10.1016/j.tibtech.2019.08.002
Nekrasov V, Staskawicz B, Weigel D, Jones JD, Kamoun S (2013) Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat Biotechnol 31(8):691–693. https://doi.org/10.1038/nbt.2655
Nicolia A, Andersson M, Hofvander P, Festa G, Cardi T (2021) Tomato protoplasts as cell target for ribonucleoprotein (RNP)-mediated multiplexed genome editing. Plant Cell Tissue Organ Cult 144(2):463–467. https://doi.org/10.1007/s11240-020-01954-8
Norfaezah J, Masani MYA, Fizree MPMA, Bahariah B, Shaharuddin NA, Ho CL, Rasid OA, Parveez GKA (2024) DNA-free CRISPR/Cas9 genome editing system for oil palm protoplasts using multiple ribonucleoproteins (RNPs) complexes. Ind Crops Prod 208:117795. https://doi.org/10.1016/j.indcrop.2023.117795
Okada K, Aoki K, Tabei T, Sugio K, Imai K, Bonkohara Y, Kamachi Y (2022) Key sequence features of CRISPR RNA for dual-guide CRISPR-Cas9 ribonucleoprotein complexes assembled with wild-type or HiFi Cas9. Nucleic Acids Res 50(5):2854–2871. https://doi.org/10.1093/nar/gkac100
Ong AL, Teh CK, Mayes S, Massawe F, Appleton DR, Kulaveerasingam H (2020) An improved oil palm genome assembly as a valuable resource for crop improvement and comparative genomics in the Arecoideae subfamily. Plants 9(11):1476. https://doi.org/10.3390/plants9111476
Parveez GKA, Christou P (1998) Biolistic-mediated DNA delivery and isolation of transgenic oil palm (Elaeis guineensis Jacq.) embryogenic callus cultures. J Oil Palm Res 10(2):29–38
Parveez GKA, Rasid OA, Masani MYA, Sambanthamurthi R (2015) Biotechnology of oil palm: strategies towards manipulation of lipid content and composition. Plant Cell Rep 34(4):533–543. https://doi.org/10.1007/s00299-014-1722-4
Parveez GKA, Hishamuddin E, Loh SK, Ong-Abdullah M, Salleh KM, Bidin MNIZ, Sundram S, Hassan ZAZ, Idris Z (2020) Oil palm economic performance in Malaysia and R&D progress in 2019. J Oil Palm Res 32(2):159–190
Parveez GKA, Rasid OA, Ahmad MN, Taib HM, Bakri MAM, Hafid SRA, Ismail TNMT, Loh SK, Ong-Abdullah M, Zakaria K, Idris Z (2023) Oil palm economic performance in Malaysia and R&D progress in 2022. J Oil Palm Res 35(2):193–216
Pavese V, Moglia A, Corredoira E, Martínez MT, Torello MD, Botta R (2021) First report of CRISPR/Cas9 gene editing in Castanea sativa Mill. Front Plant Sci 12:728516. https://doi.org/10.3389/fpls.2021.728516
Rádis-Baptista G, Campelo IS, Morlighem JÉR, Melo LM, Freitas VJ (2017) Cell-penetrating peptides (CPPs): From delivery of nucleic acids and antigens to transduction of engineered nucleases for application in transgenesis. J Biotechnol 252:15–26. https://doi.org/10.1016/j.jbiotec.2017.05.002
Ran FA, Cong L, Yan WX, Scott DA, Gootenberg JS, Kriz AJ, Zetsche B, Shalem O, Wu X, Makarova KS, Koonin EV, Sharp PA, Zhang F (2015) In vivo genome editing using Staphylococcus aureus Cas9. Nature 520(7546):186–191. https://doi.org/10.1038/nature14299
Sant’Ana RRA, Caprestano CA, Nodari RO, Agapito-Tenfen SZ, (2020) PEG-delivered CRISPR-Cas9 ribonucleoproteins system for gene-editing screening of maize protoplasts. Genes 11(9):1029. https://doi.org/10.3390/genes11091029
Sauer NJ, Narváez-Vásquez J, Mozoruk J, Miller RB, Warburg ZJ, Woodward MJ, Mihiret YA, Lincoln TA, Segami RE, Sanders SL, Walker KA, Beetham PR, Schöpke CR, Gocal GF (2016) Oligonucleotide-mediated genome editing provides precision and function to engineered nucleases and antibiotics in plants. Plant Physiol 170(4):1917–1928. https://doi.org/10.1104/pp.15.01696
Shan Q, Wang Y, Li J, Gao C (2014) Genome editing in rice and wheat using the CRISPR/Cas system. Nat Protoc 9(10):2395–2410. https://doi.org/10.1038/nprot.2014.157
Shan S, Mavrodiev EV, Li R, Zhang Z, Hauser BA, Soltis PS, Soltis DE, Yang B (2018) Application of CRISPR/Cas9 to Tragopogon (Asteraceae), an evolutionary model for the study of polyploidy. Mol Ecol Resour 18(6):1427–1443. https://doi.org/10.1111/1755-0998.12935
Shan S, Soltis PS, Soltis DE, Yang B (2020) Considerations in adapting CRISPR/Cas9 in nongenetic model plant systems. Appl Plant Sci 8(1):e11314. https://doi.org/10.1002/aps3.11314
Sierro N, Battey JN, Ouadi S, Bakaher N, Bovet L, Willig A, Goepfert S, Peitsch MC, Ivanov NV (2014) The tobacco genome sequence and its comparison with those of tomato and potato. Nat Commun 5(1):1–9. https://doi.org/10.1038/ncomms4833
Singh R, Ong-Abdullah M, Low ETL, Manaf MAA, Rosli R, Nookiah R, Ooi LCL, Ooi SE, Chan KL, Halim MA, Azizi N, Nagappan J, Bacher B, Lakey N, Smith SW, He D, Hogan M, Budiman MA, Lee EK, DeSalle R, Kudrna D, Goicoechea JL, Wing RA, Wilson RK, Fulton RS, Ordway JM, Martienssen RA, Sambanthamurthi R (2013) Oil palm genome sequence reveals divergence of interfertile species in old and new worlds. Nature 500(7462):335–339. https://doi.org/10.1038/nature12309
Song H, Ahn JY, Yan F, Ran Y, Koo O, Lee GJ (2022) Genetic dissection of CRISPR-Cas9 mediated inheritance of independently targeted alleles in tobacco α-1, 3-Fucosyltransferase 1 and β-1, 2-Xylosyltransferase 1 Loci. Int J Mol Sci 23(5):2450. https://doi.org/10.3390/ijms23052450
Steinbrenner J, Sandmann G (2006) Transformation of the green alga Haematococcus pluvialis with a phytoene desaturase for accelerated astaxanthin biosynthesis. Appl Environ Microbiol 72(12):7477–7484. https://doi.org/10.1128/aem.01461-06
Stewart MP, Langer R, Jensen KF (2018) Intracellular delivery by membrane disruption: mechanisms, strategies, and concepts. Chem Rev 118(16):7409–7531. https://doi.org/10.1021/acs.chemrev.7b00678
Subburaj S, Zanatta CB, Nunn JA, Hoepers AM, Nodari RO, Agapito-Tenfen SZ (2022) A DNA-free editing platform for genetic screens in soybean via CRISPR/Cas9 ribonucleoprotein delivery. Front Plant Sci 13:939997. https://doi.org/10.3389/fpls.2022.939997
Subburaj S, Zanon Agapito-Tenfen S (2023) Establishment of targeted mutagenesis in soybean protoplasts using CRISPR/Cas9 RNP delivery via electro-transfection. Front Plant Sci 14:1255819. https://doi.org/10.3389/fpls.2023.1255819
Svitashev S, Schwartz C, Lenderts B, Young JK, Mark Cigan A (2016) Genome editing in maize directed by CRISPR-Cas9 ribonucleoprotein complexes. Nat Commun 7(1):1–7. https://doi.org/10.1038/ncomms13274
Thomas HR, Percival SM, Yoder BK, Parant JM (2014) High-throughput genome editing and phenotyping facilitated by high resolution melting curve analysis. PLoS ONE 9(12):e114632. https://doi.org/10.1371/journal.pone.0114632
Timofejeva L, Singh D (2023) Methods and applications of CRISPR technology in plant sciences, 2022. Front Plant Sci 14:1207257. https://doi.org/10.3389/fpls.2023.1207257
Tripathi JN, Ntui VO, Shah T, Tripathi L (2021) CRISPR/Cas9-mediated editing of DMR6 orthologue in banana (Musa spp.) confers enhanced resistance to bacterial disease. Plant Biotechnol J 19(7):1291
Wang Y, Cheng X, Shan Q, Zhang Y, Liu J, Gao C, Qiu JL (2014) Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 32(9):947–951. https://doi.org/10.1038/nbt.2969
Wong N, Liu W, Wang X (2015) WU-CRISPR: characteristics of functional guide RNAs for the CRISPR/Cas9 system. Genome Biol 16(1):1–8. https://doi.org/10.1186/s13059-015-0784-0
Woo JW, Kim J, Kwon SI, Corvalán C, Cho SW, Kim H, Kim SG, Kim ST, Kim JS (2015) DNA-free genome editing in plants with pre-assembled CRISPR-Cas9 ribonucleoproteins. Nat Biotechnol 33(11):1162–1164. https://doi.org/10.1038/nbt.3389
Wroblewski T, Tomczak A, Michelmore R (2005) Optimization of Agrobacterium-mediated transient assays of gene expression in lettuce, tomato and Arabidopsis. Plant Biotechnol J 3(2):259–273. https://doi.org/10.1111/j.1467-7652.2005.00123.x
Wu S, Zhu H, Liu J, Yang Q, Shao X, Bi F, Hu C, Huo H, Chen K, Yi G (2020) Establishment of a PEG-mediated protoplast transformation system based on DNA and CRISPR/Cas9 ribonucleoprotein complexes for banana. BMC Plant Biol 20(1):1–10. https://doi.org/10.1186/s12870-020-02609-8
Wu X, Kriz AJ, Sharp PA (2014) Target specificity of the CRISPR-Cas9 system. Quant Biol 2(2):59–70. https://doi.org/10.1007/Fs40484-014-0030-x
Xie K, Minkenberg B, Yang Y (2015) Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc Natl Acad Sci USA 112(11):3570–3575. https://doi.org/10.1073/pnas.1420294112
Xie K, Yang Y (2013) RNA-guided genome editing in plants using a CRISPR-Cas system. Mol Plant 6(6):1975–1983. https://doi.org/10.1093/mp/sst119
Xie X, Ma X, Liu YG (2019) Decoding Sanger sequencing chromatograms from CRISPR-induced mutations. Plant Genom Edit CRISPR Syst. https://doi.org/10.1007/978-1-4939-8991-1_3
Xu R, Yang Y, Qin R, Li H, Qiu C, Li L, Wei P, Yang J (2016) Rapid improvement of grain weight via highly efficient CRISPR/Cas9-mediated multiplex genome editing in rice. J Genet Genomics 43(8):529–532. https://doi.org/10.1016/j.jgg.2016.07.003
Xu RF, Li H, Qin RY, Li J, Qiu CH, Yang YC, Ma H, Li L, Wei PC, Yang JB (2015) Generation of inheritable and “transgene clean” targeted genome-modified rice in later generations using the CRISPR/Cas9 system. Scientific Rep 5(1):11491. https://doi.org/10.1038/srep11491
Yang H, Wu JJ, Tang T, Liu KD, Dai C (2017) CRISPR/Cas9-mediated genome editing efficiently creates specific mutations at multiple loci using one sgRNA in Brassica napus. Sci Rep 7(1):7489. https://doi.org/10.1038/s41598-017-07871-9
Yang T, Ali M, Lin L, Li P, He H, Zhu Q, Sun C, Wu N, Zhang X, Huang T, Li CB, Li C, Deng L (2022) Recoloring tomato fruit by CRISPR/Cas9-mediated multiplex gene editing. Hortic Res 10(1):214. https://doi.org/10.1093/hr/uhac214
Yeap WC, Khan NCM, Jamalludin N, Muad MR, Appleton DR, Kulaveerasingam H (2021) An efficient clustered regularly interspaced short palindromic repeat (CRISPR)/CRISPR-associated protein 9 mutagenesis system for oil palm (Elaeis guineensis). Front Plant Sci 12:773656. https://doi.org/10.3389/fpls.2021.773656
Yin K, Han T, Liu G, Chen T, Wang Y, Yu AYL, Liu Y (2015) A geminivirus-based guide RNA delivery system for CRISPR/Cas9 mediated plant genome editing. Sci Rep 5(1):14926. https://doi.org/10.1038/srep14926
Yu J, Tu L, Subburaj S, Bae S, Lee GJ (2021) Simultaneous targeting of duplicated genes in Petunia protoplasts for flower color modification via CRISPR-Cas9 ribonucleoproteins. Plant Cell Rep 40:1037–1045. https://doi.org/10.1007/s00299-020-02593-1
Zafar K, Khan MZ, Amin I, Mukhtar Z, Yasmin S, Arif M, Ejaz K, Mansoor S (2020) Precise CRISPR-Cas9 mediated genome editing in super basmati rice for resistance against bacterial blight by targeting the major susceptibility gene. Front Plant Sci 11:575. https://doi.org/10.3389/fpls.2020.00575
Zhang H, Zhang J, Wei P, Zhang B, Gou F, Feng Z, Mao Y, Yang L, Zhang H, Xu N, Zhu JK (2014) The CRISPR/Cas9 system produces specific and homozygous targeted gene editing in rice in one generation. Plant Biotechnol J 12(6):797–807. https://doi.org/10.1111/pbi.12200
Zhang Y, Zhou P, Bozorov TA, Zhang D (2021a) Application of CRISPR/Cas9 technology in wild apple (Malus sieverii) for paired sites gene editing. Plant Methods 17(1):1–9. https://doi.org/10.1186/s13007-021-00769-8
Zhang J, Zhang H, Li S, Li J, Yan L, Xia L (2021b) Increasing yield potential through manipulating of an ARE1 ortholog related to nitrogen use efficiency in wheat by CRISPR/Cas9. J Integr Plant Biol 63(9):1649–1663. https://doi.org/10.1111/jipb.13151
Zhang S, Shen J, Li D, Cheng Y (2021c) Strategies in the delivery of Cas9 ribonucleoprotein for CRISPR/Cas9 genome editing. Theranostics 11(2):614. https://doi.org/10.7150/thno.47007
Zhang S, Wu S, Hu C, Yang Q, Dong T, Sheng O, Deng G, He W, Dou T, Li C, Sun C, Yi G, Bi F (2022) Increased mutation efficiency of CRISPR/Cas9 genome editing in banana by optimized construct. Peer J 10:e12664. https://doi.org/10.7717/peerj.12664
Zhang T, Gao Y, Wang R, Zhao Y (2017) Production of guide RNAs in vitro and in vivo for CRISPR using ribozymes and RNA polymerase II promoters. Bio-Protoc 7(4):2148–2148
Zhang Y, Showalter AM (2020) CRISPR/Cas9 genome editing technology: a valuable tool for understanding plant cell wall biosynthesis and function. Front Plant Sci 11:589517. https://doi.org/10.3389/fpls.2020.589517
Zhou J, Li D, Wang G, Wang F, Kunjal M, Joldersma D, Liu Z (2020) Application and future perspective of CRISPR/Cas9 genome editing in fruit crops. J Integr Plant Biol 62(3):269–286. https://doi.org/10.1111/jipb.12793
Zhou J, Luan X, Liu Y, Wang L, Wang J, Yang S, Liu S, Zhang J, Liu H, Yao D (2023) Strategies and methods for improving the efficiency of CRISPR/Cas9 gene editing in plant molecular breeding. Plants 12(7):1478. https://doi.org/10.3390/plants12071478
Zhu X, Xu Y, Yu S, Lu L, Ding M, Cheng J, Song G, Gao X, Yao L, Fan D, Meng S, Zhang X, Hu S, Tian Y (2014) An efficient genotyping method for genome-modified animals and human cells generated with CRISPR/Cas9 system. Sci Rep 4(1):6420. https://doi.org/10.1038/srep06420
Zischewski J, Fischer R, Bortesi L (2017) Detection of on-target and off-target mutations generated by CRISPR/Cas9 and other sequence-specific nucleases. Biotechnol Adv 35(1):95–104. https://doi.org/10.1016/j.biotechadv.2016.12.003
Acknowledgements
The authors would like to thank the MPOB management for permission to publish this review. The authors would also like to acknowledge all the Transgenic Technology Group (TTG) members of MPOB for their advice and kind assistance.
Author information
Authors and Affiliations
Contributions
Masani MYA conceived the idea, edited and final critically write up of the manuscript, and submitted the manuscript. Norfaezah J carried out literature survey, drafted the manuscript, and prepared the tables and figures. Bahariah B, Fizree MPMAA, Sulaiman WNSW, Shaharuddin NA, Rasid OA, and Parveez GKA reviewed and edited the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
The authors with this declare that the paper is based on our writing except for citations which have been duly acknowledged. We also state that it has not been previously or concurrently submitted for any other journals.
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this review paper.
Ethical approval
Not applicable.
Research involving human participants and/or animals
Not applicable.
Informed consent
All authors have read and approved this review paper.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Masani, M., Norfaezah, J., Bahariah, B. et al. Towards DNA-free CRISPR/Cas9 genome editing for sustainable oil palm improvement. 3 Biotech 14, 166 (2024). https://doi.org/10.1007/s13205-024-04010-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13205-024-04010-w