Abstract
Carica papaya L. is an economically significant crop in tropical and subtropical regions, with a gross production value of $6.2 × 109 in 2020. However, various biotic and abiotic stresses threaten crop productivity. To enhance stress resistance, genetic engineering and traditional breeding have been employed. Unfortunately, these methods are limited by the scarcity of innate disease resistance genes in the genome and the poor fertility of interspecific hybrids. Therefore, to circumvent these limitations, we developed a papaya protoplast-based gene editing system. By optimizing protoplast isolation, 28% higher yields were achieved from older (≥75 d) plants at 1.11 × 108 ± 0.069 protoplasts per gram-fresh-weight. Protoplast viability was 89.87 ± 2.02%,. We established an efficient genetic transfection method and verified proper expression, cellular function and localization of GFP and PDI-mCherry fusions in the protoplasts. Using preassembled CRISPR-Cas9 ribonucleoprotein complexes, we successfully edited a mutant GFP transgene, resulting in a frame-shift restoration efficiency of 27.88 ± 1.65%. Next, the CpPDS and CpMLO6 genes were targeted, creating knockouts in three different papaya cultivars. The average CpPDS mutant frequency obtained was 42.31 ± 1.90%, of which 31.25 ± 1.46% were frame-shift knockouts, while 11.05 ± 1.37% were in-frame protein variants. The average CpMLO6 mutant frequency was 16.20 ± 1.53%, of which 13.71 ± 1.67% were frame-shift knockouts and 2.50 ± 0.26% were in-frame variants. Taken together, a DNA-free CRISPR-Cas9 gene editing system was successfully demonstrated in papaya protoplasts on multiple target genes for use in papaya crop improvement.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Papaya (Carica papaya L., family Caricaceae) is an economically important tropical and subtropical crop worldwide, with a global gross agricultural production value of $US 6.2 × 109 in 2020 (FAOSTAT 2020; Evans et al. 2021). The unripe fruit and tree trunk are rich sources of the proteolytic enzyme papain, which aids digestion and is a meat tenderizer (Brocklehurst et al. 1981; Krishna et al. 2008). Fresh ripe papaya ranks first among fresh fruit for vitamins C and A, calcium, iron, potassium, folate, riboflavin, niacin, thiamin, and fiber (Krishna et al. 2008; Hewajulige and Dhekney 2016). Besides its dietary health benefits, the cosmetic, pharmaceutical, textile, and food processing industries use papaya and its by-products (Carlos-Hilario and Christopher 2015; Hewajulige and Dhekney 2016; Evans et al. 2021). The agricultural importance of papaya necessitates efforts to genetically improve its resistance to biotic and abiotic threats that undermine its productivity.
The elucidation of the papaya genome sequence facilitated studies on gene identification, genetic engineering, and comparative genomics that are useful for crop improvement (Ming et al. 2008; Ming and Moore 2014). The relatively small papaya genome of 372 megabases is approximately 2.75 times larger than the Arabidopsis thaliana genome (Arumuganathan and Earle 1991). Papaya and Arabidopsis share a common ancestor, with both species belonging to the order Brassicales (Ming and Moore 2014). Although the Arabidopsis genome has helped identify papaya genes, papaya contains fewer homologous disease resistance genes than Arabidopsis (Ming et al. 2008).
The limited genetic resistance to diseases is of concern because disease-causing pathogens such as the pathogenic fungus, Oidium caricae-papayae, and the oomycete, Phytophthora palmivora (Nelson 2008; Cunningham and Nelson 2012) threaten the viability of the papaya industry. Oidium caricae-papayae causes powdery mildew on papaya, resulting in agricultural losses as infected fruit are unsuitable for the market (Cunningham and Nelson 2012). Phytophthora is a hemibiotrophic pathogenic oomycete that affects agronomically important food crops around the world (Lamour 2013). The Phytophthora palmivora species causes Phytophthora blight of papaya (Nelson 2008). Symptoms include fruit and root rot, stem canker, and severe structural damage, leading to plant death (Nelson 2008). Chemical and biological control agents have been used to control Phytophthora infection, with chemical fungicides proving the most effective (Agrios 2005; Nelson 2008); the latter have been associated with environmental health concerns (Zubrod et al. 2019). Therefore, developing eco-friendlier alternatives will be useful and welcomed for papaya. Genetic engineering and traditional breeding methods were used to moderately improve resistance to P. palmivora (Zhu et al. 2007a, b). However, the few innate disease resistance genes in its genome is compounded by F1 sterility of hybrids made with closely related species (Manshardt and Wenslaff 1989). In addition, genetic engineering with an interspecific resistance transgene led to progeny with low viability and vigor (Porter et al. 2014). Clearly, new approaches are needed for this crop. The evolving ability of pathogens to infect plants by overcoming resistance (R) gene-based immunity necessitates a gene editing system capable of developing new resistant varieties via the mutation of the susceptibility (S) loci (van Schie and Takken 2014).
A critical family of S-genes, which are involved in plant pathogen pathosystems, is the MLO gene family. The MLO gene is involved in host compatibility, playing a role in facilitating pathogen entry into the host. It also increases innate immunity upon disrupting its function (van Schie and Takken 2014). The MLO locus encodes a seven-transmembrane domain protein anchored in the plasma membrane whose exact biochemical function is unknown (Büschges et al. 1997; Devoto et al. 1999). Naturally occurring loss-of-function MLO mutants provided broad-spectrum resistance to powdery mildew (Büschges et al. 1997; Bai et al. 2008). Actin remodeling is critical for determining whether pathogenic fungi successfully invade host cells, and MLO may influence this process (Opalski et al. 2004). O. caricae and P. palmivora enter the host cell by similar mechanisms during the early biotrophic infection stage, with the leaves of young barley MLO5 mutant plants demonstrating improved resistance to P. palmivora (Le Fevre et al. 2016). The development of a gene editing method for papaya to generate CpMLO6 mutant lines will help facilitate resistance studies in the future.
Clustered regularly interspaced short palindromic repeats (CRISPR) and CRISPR-associated protein 9 (Cas9) are part of an innovative gene editing technology first identified as an adaptive immune response in bacteria against bacteriophage infection (Jinek et al. 2012; Sorek et al. 2013; Hsu et al. 2014). The type II CRISPR-Cas9 system comprises a Cas9 endonuclease bound with a transactivating CRISPR RNA (tracrRNA) fused to a CRISPR RNA (crRNA), to comprise a chimeric single-guide RNA (sgRNA) with a 20 nucleotide target sequence (Anzalone et al. 2020). The complex can target, bind, and cleave complementary DNA sequences, introducing double-strand breaks (DSB) into host DNA. Due to the cell’s inaccurate non-homologous end joining (NHEJ) DNA repair mechanism, erroneous repairs are often made, introducing a diverse range of insertion and deletion (indel) mutations that can dramatically alter or terminate targeted protein function (Chen et al. 2019; Zhang et al. 2021a).
Agrobacterium-mediated CRISPR-Cas9 gene editing has been used in various crop improvement studies (Tian et al. 2017; Kaur et al. 2018; Zhang et al. 2019a; Zhang et al. 2019b; Navet and Tian 2020). Recently, this traditional DNA-based method (Brewer and Chambers 2022) was used to edit the phytoene desaturase (CpPDS) gene in papaya. However, extended time constraints are introduced while regenerating calli tissue suitable for transformation, segregating transgenic from non-transgenic material via Mendelian segregation (Zhang et al. 2019a), and navigating regulatory processes for commercialization (El-Mounadi et al. 2020). Developing a DNA-free method for gene editing in papaya will provide the tools necessary to bypass these constraints. Specifically, using papaya protoplasts instead of embryogenic calli reduces the time and costs required to generate and screen transformants, because protoplast isolation takes hours rather than weeks or months to produce large quantities of suitable target protoplasts. They are homogenous with similar physiological and physical characteristics. Finally, protoplasts provide numerous independent events in a single experiment, as each represents a single in vivo system (Nadakuduti et al. 2019).
A DNA-free method for performing CRISPR-Cas9 gene editing in plant cells is the use of CRISPR-Cas9 ribonucleoprotein (RNP) complexes. These RNP complexes can enter plant protoplasts, perform stable gene edits, and then are degraded through normal cell turnover activities. It is considered a transgene-free gene editing method because no transgenic material remains in the host genome (He et al. 2022). As such, regulatory concerns associated with traditional gene editing methods are diminished (Metje-Sprink et al. 2019). The CRISPR-Cas9 RNP complex is formed using all the same components necessary for traditional CRISPR-Ca9 gene editing, although the components are complexed in vitro rather than in vivo, then transfected into the plant protoplast using methods such as PEG-meditated transfection (Metje-Sprink et al. 2019).
Delivery of the CRISPR-Cas9 RNP complexes into plant protoplasts using the PEG-mediated transfection method aids in transfection efficiency, especially when combined with Ca2+ and Mg2+, to increase phospholipid membrane permeability and improve transfection rates (Boss and Mott 1980; Maas and Werr 1989; Parray et al. 2020). This method has been used in several plant species, including A. thaliana (Woo et al. 2015), tobacco (Woo et al. 2015), rice (Woo et al. 2015), potato (Andersson et al. 2018; González et al. 2020), tomato (Lin et al. 2022), grapevine (Malnoy et al. 2016), apple (Malnoy et al. 2016), cabbage (Park et al. 2019), canola (Sidorov et al. 2021), and banana (Wu et al. 2020). Furthermore, complexing Cas9 endonuclease with a single-guide RNA (sgRNA), having a 2- to 10-fold molar excess of sgRNA, was effective for RNP-based gene editing in several plant species (Woo et al. 2015; Malnoy et al. 2016; Subburaj et al. 2016; Liang et al. 2017). The RNP delivery method has numerous advantages over traditional CRISPR-Cas9 delivery methods. Firstly, off-target effects are decreased, as there is no longer a transgenic CRISPR-Cas9 cassette continuously expressed in the host genome (Liang et al. 2017; He et al. 2022). Secondly, the time required for Cas9 transcription and translation prior to gene editing is eliminated because preassembled gene editors are delivered as an activated complex. Lastly, RNP delivery eliminates the concern of introducing insertion mutations that may occur using traditional stable transformation methods (Zhang et al. 2021b).
Considering these aforementioned advantages, we undertook a study to apply this DNA-free method to papaya protoplasts with the long-term goal of crop improvement. We improved the method to isolate intact viable protoplasts from papaya of various aged leaves and transfected them with reporter gene fusions for transient expression. Finally, DNA-free editing was conducted and confirmed by correcting a frame-shift mutant GFP transgene and deep amplicon sequence verification of targeted editing of endogenous papaya genes, CpPDS, and CpMLO6.
Materials and Methods
Growing Papaya Plants
Papaya Solo Sunrise and Sunset seeds were purchased from the University of Hawai’i Agricultural Diagnostic Service Center – Seed Program and Paramount Seeds Inc. (paramountseeds.com/collections/papaya). Papaya Solo Kapoho seeds were obtained by cleaning and drying the seeds from the fruit. Seeds were stored at 4°C. To germinate, the seeds were soaked for 24 h in 1 M KNO3 before transferring them into Sunshine Mix No. 4, mixed with 1/5 vol of perlite. Plants were grown under broad-spectrum light at 75–150 μmol m−2 s−2 for a 16-h daily photoperiod. Germination occurred approximately 10 d after planting. The age of a plant was calculated from the day of seedling emergence from the soil after germination. Plants were well watered with care taken to ensure that they were free from drought or overwatering stress, as both conditions could affect the quality of isolated protoplasts. The leaves were visually inspected to ensure that the selected tissue was free of abnormalities and signs of biotic or abiotic stress. Leaves were harvested when plants were 50–71 d old for the young age category and 79–136 d for the old age category. New, fully expanded leaves were selected with petioles no longer than 1 inch and no reddening.
Protoplast Isolation and Quantification
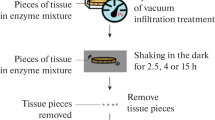
Leaves were collected and prepared according to methods previously described (Zhang et al. 2011), with a few modifications. Approximately 0.5 g of the newest fully expanded leaves from young or old plants was surface sterilized in a 50% EtOH solution for 30 s, and then transferred to a 20% Clorox solution for 7 min. Sterilized leaves were rinsed three times in ddH2O and then pat dried. A sharp scalpel was used to cut the leaves transversely into strips 1–2 mm wide. A sheet of white paper was used as a cutting surface to identify chlorophyll staining, indicating when the scalpel blades needed replacement. The protoplast isolation medium (1.3% Cellulase R-10, 0.3% Macerozyme R-10, 0.44 M D-mannitol, 20 mM MES at pH 5.8, 20 mM KCl, 10 mM CaCl2, 0.5% w/v PVP) was heated in a 55°C liquid bath for 10 min to deactivate proteases and ensure the enzymes were fully dissolved. The isolation medium was cooled to RT, and then, polyvinylpyrrolidone (PVP) was added to 0.5% w/v, vortexed to dissolve the PVP thoroughly, and then sterilized using a 0.20-μm filter. 0.5 g of leaf strips was added to 10 ml of isolation medium in a 50-ml conical tube, and a vacuum (10–12 mbar, 1.1–1.3 kPa) was applied for 30 min under low light. The vacuum was released every 5 min to resubmerge the leaf material in the solution and reapplied cyclically over the 30-min infiltration period. Vacuum-infiltrated leaves were poured onto a 100-mm Petri dish, spread evenly, sealed with micropore tape, placed in an orbital shaker, and incubated at 26°C with 60 rpm for 13–15 h in the dark. After incubation, protoplasts were washed in an ice-cold modified W5 buffer solution (154 mM NaCl, 125 mM CaCl2, 5 mM KCl, 2 mM MES, pH 5.8). Ten milliliters of W5 buffer was added to the Petri dish and mixed gently. Protoplasts were filtered through a 30–70-μm nylon mesh in a 50-ml conical tube. The protoplast solution was centrifuged for 5 min at 100 × g, at 4°C. The supernatant was carefully discarded, and the protoplast pellet was resuspended in 10 ml W5 buffer. The washes were repeated 2× until the supernatant was free of debris. Finally, the protoplast pellet was resuspended in 10 ml of W5 buffer and incubated on ice for 30 min. Protoplasts were quantified using a hemocytometer. The total number of protoplasts was divided by leaf GFW to determine the total number of papaya protoplasts isolated per GFW leaves.
Protoplast Viability Assay
The FDA hydrolysis assay (Larkin 1976) was used to determine the viability of freshly isolated papaya protoplasts. A 500-μl aliquot of protoplasts in W5 buffer was added to a 2-ml round-bottom tube. Ten microliters μl of 0.5% FDA (5 mg/ml in acetone stock solution) was added to the protoplasts to obtain a final concentration of 0.01% FDA. After 5 min, the protoplasts were analyzed and imaged using brightfield and epifluorescence microscopy with an Olympus BX51 upright compound epifluorescence microscope equipped with a Leica DFC7000T 2.8-megapixel cooled CCD camera located in the Biological Electron Microscope Facility at the University of Hawai’i at Mānoa. A Chroma bandpass eGFP filter with excitation (EX 480/20 nm, BA 495LP) and emission (EM 510/20 nm) was used for epifluorescence imaging. The fluorescing to the non-fluorescing protoplasts were quantified using a hemocytometer.
Transient Transfection of Papaya Protoplasts
Protoplasts were genetically transfected using the PEG-mediated transfection protocol (Yoo et al. 2007) with a few modifications. Freshly isolated protoplasts were centrifuged at 100 × g for 5 min. The protoplasts were resuspended in a modified MMg buffer solution (0.6 M Mannitol, 15 mM MgCl2, 4 mM MES at pH 5.8) to a concentration of 5 × 105 protoplasts ml−1. A 200-μl aliquot of protoplast in MMg was added to a 2-ml round bottom tube; then, 30–40 μg of the plasmid construct enhanced GFP(S65T), also termed eGFP, was added and mixed gently and thoroughly. The GFP(S65T) construct is based on the vector pBluescript KS(+), with eGFP driven by a CaMV 35S promoter and flanked by a NOS terminator (35S-eGFP). The eGFP emits an increased signal intensity with reduced photobleaching by replacing serine at site 65 with a threonine (Chiu et al. 1996). The total vol of plasmid DNA did not exceed 10% of the total vol of added protoplasts. An equal vol of PEG-CaCl2 (0.2 M D-mannitol, 0.1 M CaCl2, and 40% (w/v) polyethylene glycol 4000) was added, and then mixed by inversion. The transfection mixture was incubated for 30 min at RT in the dark. Transfection was stopped by adding 800 μl of W5 buffer at RT. The tube was gently inverted and then centrifuged for 3 min at 300 × g at RT. The supernatant was discarded, and the W5 buffer wash step was repeated twice. The protoplasts were then resuspended in 1 ml of WI buffer (0.6 M D-mannitol, 20 mM KCl, and 4 mM MES at pH 5.8) and incubated at RT in the dark for 16–18 h.
Brightfield microscopy, imaging, and quantification were performed as described above (“Growing Papaya Plants” section). Then, confocal images were obtained using a TCS Leica SP8 X confocal laser scanning microscope with a white light laser mounted on a Leica DF6 CFS upright microscope integrated with a high-speed DFC9000 GTC digital camera (Biological Electron Microscope Facility at the University of Hawai’i at Mānoa). Observations were made with a HC PL APO CS2 63× 1.4 NA oil immersion lens. Protoplasts were visualized using the eGFP excitation/emission spectrum of 488/505–525 nm with a time-gating of 0.3–6.0 ns to reduce chlorophyll autofluorescence. All images were acquired using Leica LAS X software.
Transient Cotransfection and Colocalization Analysis
Papaya leaf mesophyll protoplasts were cotransfected as described in the “An Improved Method for Isolating High-Yield, Highly Viable Papaya Protoplasts” section with a few modifications. For cotransfection, 15 μg of the construct PDI9:eGFP and 15 μg of the construct ER:mCherry were added. PDI9:eGFP is in pBluescript KS(+) driven by the CaMV 35S promoter ending with a NOS terminator. PDI9 is fused at the C-terminus to GFP(S65T), with a KDEL ER retention motif of PDI9 added to the C-terminus of GFP(S65T) (Yuen et al. 2013). The construct ER:mCherry in pBluescript KS(+) is driven by the CaMV 35S promoter ending with a NOS terminator (Cho et al. 2011). Images were obtained as described above (“An Improved Method for Isolating High-Yield, Highly Viable Papaya Protoplasts” section) with a few modifications. Observations were made with a HC PL APO CS2 100× 1.4 NA oil immersion lens. PDI9-GFP and ER-mCherry were visualized using the excitation/emission spectra of 488/505–525 nm and 543/585–615 nm, respectively. A time-gating of 0.3–6.0 ns was used to reduce chlorophyll autofluorescence.
Frame-Shift Restoration of the GFP Mutant (GFPm) Using CRISPR-Cas9 RNP
A recombinant Alt-R® SpCas9 nuclease containing a nuclear localization signal (NLS) and a C-terminal 6-His tag was purchased from Integrated DNA Technologies, Inc. (IDT; Coralville, IA). The custom design tool for IDT CRISPR-Cas9 gRNA (http://www.idtdna.com/site/order/oligoentry/index/crispr) was used to purchase a predesigned IDT Alt-R CRISPR-Cas9 sgRNA oligo integrated with a custom 20 nt (5′-ACCATGGTGAGCAAGGGGCG-3′) sequence targeting the GFP+1 frame-shift mutation site in GFPm (Liu et al. 2018). The GFPm gene is inserted in pUC19, driven by a CaMV 35S promoter and ended by a NOS terminator. The GFPm frame-shift mutant construct and the 20 nt sgRNA target sequence used in this study were provided by (Liu et al. 2018). Papaya leaf mesophyll protoplasts were cotransfected as described in the “An Improved Method for Isolating High-Yield, Highly Viable Papaya Protoplasts” section with several modifications. Ten microliters of RNP complex buffer (20 mM HEPES, 150 mM KCl, pH 7.5), 20 μg of SpCas9 enzyme, and 10 μg sgRNA were added to a 2-ml tube with sgRNA present in a molar excess of approximately 2.5× of SpCas9. The solution was mixed and then incubated at RT for 5 min to allow the RNP complexes to assemble. Forty micrograms of the GFPm construct and 200 μl of a 5 × 105 protoplasts ml−1 in MMg solution were added, and the solution was mixed gently by inversion. The transfection procedure then followed the “An Improved Method for Isolating High-Yield, Highly Viable Papaya Protoplasts” section, with an extended incubation time of 48 h. Protoplasts with GFP frame restored were quantified using a Leica SP8 X confocal laser scanning microscope and a hemocytometer using the excitation/emission spectrum of 488/505–525 nm.
Western Blot Analysis
Immunoblot analysis was performed on protoplast samples transfected with the GFPm plasmid in the presence or absence of RNP complex, as described in Fig. 4D, scaled up to 5-fold. A sample transfected with the plasmid pUC19 was included as a negative control. Samples were centrifuged at 300 × g for 3 min, and then, the supernatant was discarded. The protoplast pellet was resuspended by adding 80 μl protoplast protein extraction buffer (PEB) (10 mM HEPES pH 7.5, 100 mM NaCl, 1 mM EDTA pH 8.0, 10% glycerol, 0.5% Triton X-100, 625 μM PMSF) to the 50-ml conical tube containing the pelleted protoplasts. The resuspended protoplasts in PEB were then transferred to a 2-ml round bottom tube, and samples were vortexed vigorously for 30 s. Lysate protein levels were quantified using the Bio-Rad DC™ Protein Assay Kit II (Bio-Rad;5000112). For immunoblot detection, protein samples were mixed with Non-Reducing Lane Marker 5× sample buffer from Thermo Scientific and 200 mM DTT, and then heated for 4 min at 90°C. The samples were quenched on ice, separated by SDS-PAGE (5% polyacrylamide stacking gel at pH 6.8, 10% polyacrylamide resolving gel at pH 8.8), and transferred to a nitrocellulose membrane. Each well was loaded with 10 μg of total protein while the positive control contained 1 μg. Immunoblot analysis was performed using Invitrogen A6455 rabbit anti-GFP primary antibody at 1:2000 dilution and Advansta R-05072 goat anti-rabbit IgG (H+L) HRP conjugate secondary antibody at 1:20,000 dilution, and then visualized by chemiluminescence using the Advansta WesternBright™ ECL Western blotting detection kit K-12045-D20.
Editing of CpMLO6 and CpPDS in papaya protoplasts using CRISPR-Cas9 RNP
A phylogenetic tree was constructed to determine which MLO homolog in papaya would most likely contribute to the susceptibility to powdery mildew. Sixteen annotated non-low-quality papaya MLO-like protein sequences were aligned with MLO homologs from seven plant species belonging to clade IV of monocots and clade V of dicots (Supplemental file 1), which were functionally characterized to be susceptibility (S) genes to powdery mildew (Pessina et al. 2014). The resulting alignment was used to construct a phylogenetic tree using the Unweighted Pair Group Method with Arithmetic Mean (UPGMA) algorithm with bootstrap resampling over 1000 replicates. Papaya MLO-like protein 6 (CpMLO6, XP_021909409.1) was determined to be closely related to MLO homologs in dicot clade V (Supplemental file 1).
Candidate 20 nt sgRNA target sequences were identified using the EuPaGDT web server (http://grna.ctegd.uga.edu) with the mRNA sequences of CpPDS (NCBI Reference Sequence: XM_022033216.1) and CpMLO6 (NCBI Reference Sequence: XM_022053717.1) as the queries and the SunUp Papaya genome (Ming et al. 2008) assembly 1.0 (NCBI GenBank Assembly Accession: GCA_000150535.1) uploaded as a custom genome, for on- and off-target analyses, editing efficiency scores, and GC content. Candidate target sequences located in exons close to 5′ end, with no off-targets, editing efficiency scores higher than 0.4, and GC content between 40 and 65%, were selected for prediction of RNA secondary structures using the RNAstructure web server (http://rna.urmc.rochester.edu/RNAstructureWeb/). The candidate target sequences coupled with a gRNA scaffold were used for RNA structure prediction. The ones predicted to have a secondary structure similar to other functionally active sgRNAs were selected and subjected to in vitro DNA cleavage activity assay (Gumtow et al. 2018; Navet and Tian 2020; Hasley et al. 2021).
CpPDS-sgRNA16 (20nt target sequence: 5′-AGTGTTTCTGCGGCGAGCTT-3′) and CpMLO6-sgRNA254 (20nt target sequence: 5′-CGAAGTCAATTGGAGCCACG-3′) targeting CpPDS and CpMLO6, respectively, were selected for in vitro cleavage assays. sgRNAs containing 20 nt target sequence followed by 80 nt sgRNA scaffold sequence were synthesized by IDT. The primers CpMLO6_InVitro_F1 (5′-CCTTCATATGTCCGTATCACTG-3′), CpMLO6_InVitro_R1 (5′-ACATCCATACGCTACGTACTTC-3′), CpPDS_InVitro_F1 (5′-CCAGATAGACATTACCCAGAATC-3′), and CpPDS_InVitro_R1 (5′-CTATGTCCTGGAATGAACTTCAC-3′) were used to produce 1250 bp amplicons as the target DNA for in vitro cleavage assays and to obtain sequence information for CpMLO6 and CpPDS papaya cultivars Sunset, Sunrise, and Kapoho. Sanger sequence data provided by Genewiz, Inc (South Plainfield, NJ) for CpMLO6 and CpPDS were identical in all three cultivars. In vitro cleavage assays were performed using the GenCrispr sgRNA Screening Kit (GenScript, Piscataway, NJ) following the manufacturer’s instructions.
Editing of CpMLO6 and CpPDS in papaya protoplasts was performed with several modifications. Sixty micrograms of SpCas9 and 30 μg of sgRNA were added to a 2-ml round bottom tube. RNP complexing buffer (20 mM HEPES, 150 mM KCl, pH 7.5) was added to reach a final volume of 40 μl. Transfection protocols were performed as described in the “An Improved Method for Isolating High-Yield, Highly Viable Papaya Protoplasts” section with an extended incubation period of 48 h. DNA was extracted from gene edited protoplasts using the Macherey-Nagel, Inc (Allentown, PA) NucleoSpin Plant II Mini kit (Cat. No. 740770.50). PCR amplification using Phusion® High-Fidelity DNA Polymerase produced amplicons for Sanger and Illumina® next generation deep amplicon sequencing from Genewiz, Inc. Primers CpMLO6_sg254_F1 (5′-GAGAGCTCTGTACGAATCACTTG-3′) and CpMLO6_sg254_R1 (5′- GCACATTTATCAGTAGAGGCA-3′) amplified a 386 bp region spanning the CpMLO6 sgRNA254 target site. Primers CpPDS_sg16_F1 (5′-CCAGATAGACATTACCCAGAATC-3′) and CpPDS_sg16_R1 (5′-GGATTCACTAACCCTAAATGC-3′) amplified a 406 bp region spanning the CpPDS sgRNA16 target site. Amplicons were prepared according to Genewiz, Inc. sample submission guidelines for Amplicon-EZ® services. Amplicon-EZ results were processed using Geneious Prime software from Biomatters, Inc (Boston, MA). Illumina paired-end reads were paired and merged, and then processed using the Analyze CRISPR Editing Results tool. A minimum read frequency of 0.10% was applied, filtering mutant detection below the threshold. Mutants were screened against the WT amplicon to ensure they were exogenous to the WT genome.
Results
An Improved Method for Isolating High-Yield, Highly Viable Papaya Protoplasts
We initially focused on the protoplast isolation procedure to maximize the yield of viable mesophyll protoplasts. We examined the influence of leaf and plant age, the components of the isolation medium, enzyme concentration, incubation period, plant and leaf health, and vacuum infiltration during preliminary trials. The data shown here were obtained using the optimized conditions as described in the Methods. Plants were divided into two age categories: old (≥75 d) and young (<75 d) (Fig. 1). The leaf tissue of old plants (Fig. 1A, C) provided an average yield of 1.11 × 108 ± 0.069 protoplasts per GFW (Fig. 1E), whereas the leaf tissue of young plants (Fig. 1B, D) yielded 0.87 × 108 ± 0.079 protoplasts per GFW (Fig. 1E). The yield differences between old and young plants were statistically significant, with t(37) = 2.28, p = .028. Generally, both ages produced large numbers of intact protoplasts with minimal extraneous debris. However, protoplasts from older plants were more durable; they tolerated repeated centrifugation and washing steps without lysing during downstream procedures. Therefore, older plants were used for all subsequent protoplast isolations reported here. Interestingly, the leaf mesophyll protoplasts of papaya were relatively small at 15–25 μm, compared to protoplasts from other plant species, such as Arabidopsis, which ranged from 30 to 50 μm (Yoo et al. 2007).
Enzymatic isolation of papaya leaf mesophyll protoplasts from old (≥75 d) and young (<75 d) plants. (A) Representative leaf sample from an old plant and (B) a young plant. (C, D) Freshly isolated mesophyll protoplasts from old and young plants, respectively, viewed at 20× magnification. (E) A two-sample t-test was used to evaluate mean protoplast yield per gm FW of leaf tissue. The boxplot represents quartiles of data points over 23 total replicates. The horizontal lines represent median values. Equal variances were not assumed.
Freshly Isolated Papaya Protoplasts Maintain High Viability, as Assessed by the Fluorescein Diacetate Hydrolysis Assay
The viability of protoplasts was determined using fluorescein diacetate (FDA) staining (Larkin 1976). Immediately after protoplast isolation, protoplasts were treated with FDA and observed using bright field and epifluorescence microscopy (Fig. 2A, B). The viability of protoplasts from new leaf tissue of older plants was 89.87 ± 2.02% (Fig. 2C). The consistently high percentage of viable protoplasts obtained here was used for downstream transfection and gene editing.
Fluorescein diacetate (FDA) viability assay of freshly isolated papaya mesophyll protoplasts. (A) Representative image of papaya protoplasts counted with a hemocytometer utilizing epifluorescence and brightfield microscopy merged into a single composite image. (B) Representative image of papaya protoplasts viewed on a single-well concavity slide using epifluorescence and brightfield microscopy. Both images are merged into a single composite image and viewed at 10× magnification. (C) An interval plot of average protoplast viability as determined by FDA fluorescence is shown. Papaya plants were ≥75 d old.
An Optimized Method for Transient Transfection of Papaya Protoplasts Developed Using a Fluorescent Marker
We next focused on ensuring that isolated protoplasts could be genetically transfected to transiently express a reporter gene as a prerequisite for subsequent gene editing applications. The freshly isolated papaya protoplasts were transfected with plasmid DNA containing the 35S-eGFP construct. As expected, the successfully transfected protoplasts emitted green fluorescence throughout the cell (Cho et al. 2011) (Fig. 3A). No signal was detected from the untransfected protoplasts. Both transfected and non-trasnfected protoplasts were observed. The efficiency of protoplast transfection of the single construct was 43.79 ± 1.65% (Fig. 3D). PEG concentrations ranging from 25 to 40% were used, with 40% providing optimal results. When using fragile protoplasts from young plants, 25% PEG minimized cell damage while maintaining transfection with reduced efficiency.
Confocal laser scanning microscopy analysis of transfected papaya mesophyll protoplasts. (A) Transient transfection of the protoplasts with the fluorescent marker GFP(S65T). The fluorescent signal is detected throughout the cell as expected. The protoplasts were imaged using excitation/emission spectra of 488/505–525 nm. (B) Transient transfection of the protoplasts with the fluorescent protein fusion PDI9-GFP, observed localized to the ER as imaged using the excitation/emission spectra, 488/505–525 nm. (C) Cotransfection of the protoplasts with fluorescent protein fusion constructs PDI9-GFP and ER-mCherry (the ER marker Er-rk) as imaged using the excitation/emission spectra of 488/505–525 nm and 543/585–615 nm, respectively. (D) Interval plot of mean transfection efficiency using GFP(S65T). Data represents a triplicate of experiments. Interval bars represent one standard error from the mean.
Colocalization of the Fluorescent Fusion Proteins ER-mCherry and Arabidopsis PDI9-GFP Confirms Normal Cell Function in the Papaya Protoplasts
The ability to localize the resulting fusion proteins to the appropriate intracellular compartments was evaluated to verify normal cellular function and activity post-transfection. A fluorescent protein fusion, PDI9:eGFP plasmid, which contained the Arabidopsis protein disulfide isomerase 9 gene (AtPDI9) (Yuen et al. 2013), was delivered to protoplasts simultaneously with the plasmid containing the endoplasmic reticulum (ER) marker, ER-rk (Cho et al. 2011; Nelson et al. 2007). As in previous studies in Arabidopsis (Cho et al. 2011; Yuen et al. 2013), we observed that the PDI9-GFP fusion was localized to the ER in papaya using confocal laser scanning microscopy (Fig. 3B). The analysis further showed that PDI9-GFP strongly colocalized with the ER-mCherry marker, indicating that PDI9-GFP is expressed and localized to the ER lumen in papaya (Fig. 3C). The overall percent efficiencies of transfection for individual PDI9-GFP and ER-mCherry alone were 46.21 ± 1.22%, which is similar to the efficiency of 35S:eGFP above. However, for double transfections with two different constructs, the efficiency decreased to 27.33 ± 1.83%. In summary, plasmid DNA entered the nucleus of the host cell using the optimized transfection procedure, the reporter genes were properly expressed, and the resulting proteins were trafficked to the correct subcellular compartments. These results demonstrate proper cellular function post-transfection.
CRISPR-Cas9 Targeted Restoration of the GFP+1 Frame-Shift Mutation Using Preassembled Ribonucleoprotein Complexes
We then used the abovementioned parameters for protoplast transfection to test whether the preassembled CRISPR-Cas9 RNP gene editing complex could perform targeted editing in vivo. The assay utilized a GFP mutant (GFPm), which contained a single nt frame-shift mutation that disrupted the reading frame for GFP mRNA, thereby preventing the formation of GFP protein and fluorescence, as previously described (Liu et al. 2018). Upon CRISPR-Cas9 editing of the mutation site, the error-prone NHEJ DNA repair mechanism corrects the reading frame, restoring GFP protein function and fluorescence, which is readily observable.
Papaya protoplasts were cotransfected with GFPm and Cas9 complexed with a sgRNA targeting the GFPm mutation site (Liu et al. 2018). The protoplasts containing the CRISPR-edited frame-shift of GFP that restored the correct reading frame emitted green fluorescence when analyzed under confocal laser scanning microscopy (Fig. 4A). Unedited protoplasts, or those that were edited but did not restore the correct reading frame, did not emit fluorescence (Fig. 4B). Three in vivo protoplast gene editing experiments were completed, resulting in a frame-shift restoration efficiency of 27.88 ± 1.65% (Fig. 4C). The papaya-optimized PEG-mediated transient transfection method successfully introduced a CRISPR-Cas9 RNP-based gene editor that corrected the GFPm gene in vivo.
CRISPR-Cas9 RNP targeted gene editing and restoration of the GFP+1 (GFPm) frame-shift mutation in vivo in papaya mesophyll protoplasts. (A, B) Representative images of the reading frame corrected GFPm visualized using confocal laser scanning microscopy with the excitation/emission spectra, 488/505–525 nm. The untransfected, unedited, and non-frame-shift restored protoplasts do not emit a fluorescent signal. (C) An interval plot of mutant editing efficiency used protoplasts cotransfected with GFPm and a RNP complex targeting the GFP mutation site (+RNP) and without (-RNP). The data represents triplicate experiments. (D) Immunoblot detection assay of the frame-shift restored GFP protein using a GFP-specific antiserum on protoplast protein lysates. The arrowhead denotes a GFP band in the GFPm+RNP lane only. Protoplasts transfected with the pUC19 empty vector (no RNP) and plasmid with GFPm with (+) and without (-) RNP. (E) A corresponding Coomassie-stained protein gel is shown as loading control of protoplast protein lysates. Protein markers are shown in kDa.
Immunoblot analysis was performed using a GFP-specific antibody to confirm and quantify the presence of the functional GFP in the protein extracts from transfected protoplasts. The frame-shift restored GFP protein was detected in the sample cotransfected with the GFPm plasmid and the Cas9-sgRNA RNP complex targeting the GFP+1 frame-shift mutation site (Fig. 4D). No observable bands were detected in the sample transfected with the empty vector control pUC19, or GFPm plasmid in the absence of Cas9-sgRNA RNP complex despite the equal loading of protein in each treatment (Fig. 4D, E). The results further confirmed that the proper editing of GFPm occurred, detecting functionally restored GFP protein in protoplasts.
DNA-Free CRISPR-Cas9 Targeted Mutagenesis of Papaya Mildew Resistance Locus O-Like 6 and Phytoene Desaturase Gene
A critical next step in developing a DNA-free gene editing system was to target the papaya endogenous genes and edit them in vivo, producing knockout mutants. CpPDS (NCBI accession: XP_021888908.1) and CpMLO6 (NCBI accession: XP_021909409.1) were selected to develop the DNA-free gene editing system. CpPDS was chosen because its mutation was expected to produce a readily observable albino phenotype (Arias et al. 2006), as described in banana (Kaur et al. 2018), tobacco (Lin et al. 2018), and rice (Wang et al. 2017). The MLO6 locus was chosen to disrupt because the induced and naturally occurring MLO mutants display increased resistance to infection by powdery mildew-causing fungal pathogens (Opalski et al. 2004). In the papaya genome (Ming et al. 2008), 16 high-quality protein sequences were annotated as MLO-like proteins . We conducted a phylogenetic analysis of these sequences with representative MLO genes of Clade IV and Clade V from plants (Supplemental file 1), which contain known genes involved in susceptibility to powdery mildew in monocots and dicots, respectively (Pessina et al. 2014). We found that CpMLO6 was clustered with Clade V MLOs (Supplemental file 1), suggesting that CpMLO6 is a candidate S-gene MLO homolog in papaya and an ideal target for genome editing to disrupt susceptibilty. For each gene, a functional sgRNA was selected based on the analyses of on- and off-targets in the genome, gene editing efficiency score, GC content, and predicted secondary structure (Supplemental file 2), followed by in vitro DNA cleavage assays (Supplemental file 3). CpPDS-sgRNA16 (20nt target sequence: 5′-AGTGTTTCTGCGGCGAGCTT-3′) and CpMLO6-sgRNA254 (20nt target sequence: 5′-CGAAGTCAATTGGAGCCACG-3′) targeting CpPDS and CpMLO6, respectively, were able to successfully guide the cleavage of the target DNA.
To specifically edit the papaya CpPDS gene using an RNP containing CpPDS-sgRNA16 and Cas9, two replicates were performed in papaya cv. Solo Sunrise, which produced an average mutant frequency of 45.09% (Fig. 5A). 32.08% of the generated mutants resulted in frame-shift knockouts, while in-frame protein variants represented 13.01% (Fig. 5A). Of the 32.08% knockout mutants, 19.80% consisted of premature stop codons (Fig. 5A). Indel types ranged from +5 to −13 bp (Fig. 5B). The deletion of −3 bp occurred with the highest frequency at 10.44%, followed by the addition of +1 bp at 9.51%, deletion of −1 bp at 8.66%, and deletions of −4 bp at 6.08% and −6 bp at 3.45% (Fig. 5B). The specific types of mutations detected from next generation sequencing (NGS) data targeting CpPDS in cv. Solo Sunrise are provided (Tables 1, 2, Supplemental file 4). This experiment was repeated on two other papaya cultivars, Sunset and Kapoho. In cv. Solo Sunset, a mutant frequency of 37.22% was produced, with 28.24% being frame-shift knockout mutants and 8.73% being in-frame protein variants. Of the total frame-shift mutants detected, 17.81% produced premature stop codons (Fig. 5C). Indel types ranged from +1 to −64 bp, with the deletion of −1 bp occurring with the highest frequency at 11.35% followed by the addition of +1 bp at 7.95% and deletions of −3, −4, and −2 bp at 6.89%, 5.34%, and 1.74%, respectively (Fig. 5D). Targeting CpPDS in papaya cv. Solo Kapoho produced a mutant frequency of 41.81%. Frame-shift mutants accounted for 32.36% of the total mutants detected, while 9.46% were in-frame protein variants (Fig. 5E). 20.81% of frame-shift knockout mutants were resultant of premature stop codons (Fig. 5E). Indel types ranged from +5 to −53 bp (Fig. 5F). The indel type produced with the highest frequency was the deletion of −1 bp at 12.99% followed by the addition of +1 bp at 7.98% and the deletions of −4, −3, and −6 bp at 6.09%, 5.98%, and 2.54%, respectively (Fig. 5F). More detailed results obtained targeting CpPDS in cv. Solo Sunset and Kapoho are provided (Tables 3, 4, Supplemental file 4). The average mutant frequency obtained targeting CpPDS in all three cultivars was 42.31 ± 1.90% (Supplemental file 5). 31.25 ± 1.46% of the generated mutants were frame-shift knockouts, while in-frame protein variants accounted for 11.05 ± 1.37% (Supplemental file 5). Of the 31.25 ± 1.46% frame-shift knockout mutants detected, 19.56 ± 1.64% generated premature stop codons.
The measurement of DNA-free CRISPR-Cas9 gene editing efficiency in papaya protoplasts targeting the CpPDS gene as confirmed by NGS. The bar chart representations of total CpPDS CRISPR-Cas9 gene editing efficiency (indels—black) and proportion of editing events leading to frame-shift knockouts (gray) and in-frame protein variants (light gray). Premature stop codons are presented as a proportion of total frame-shift knockouts (striped). The summary of mutants (A) and indels (B) generated for the CpPDS gene from the cv. Solo Sunrise. The summary of mutants (C) and indels (D) generated for the CpPDS gene from the cv. Solo Sunset. The summary of mutants (E) and indels (F) generated for the CpPDS gene from the cv. Solo Kapoho.
A CRISPR-Cas9 RNP targeting the papaya locus, CpMLO6, using CpMLO6-sgRNA254, performed gene editing at the specified site in papaya cv. Solo Sunrise producing an average mutant frequency of 18.84% (Fig. 6A). 16.54% of the generated mutants resulted in frame-shift knockouts, while 2.30% were in-frame protein variants (Fig. 6A). The indel types ranged from +2 to −15 bp (Fig. 6B). The deletion of −1 bp occurred with the highest frequency at 6.96%, followed by the addition of +1 bp at 4.40%, deletion of −2 bp at 1.91%, deletion of −3 bp at 1.70%, and deletion of −4 bp at 1.29% (Fig. 6B). Total mutants detected from the NGS data obtained targeting CpMLO6 in cv. Solo Sunrise is provided (Tables 5, 6, Supplemental file 6). This process was repeated, targeting CpMLO6 in cv. Solo Sunset and Kapoho. In cv. Solo Sunset, a mutant frequency of 13.60% was observed, with frame-shift mutants representing 10.37% of the total mutations detected, while 3.23% were in-frame protein variants (Fig. 6C). Detected indels ranged from +1 to −10 bp with the deletion of −1 bp observed at the highest frequency, followed closely by the addition of +1 bp at 3.69% and the deletion of −3 bp at 2.64% (Fig. 6D). While targeting CpMLO6 in Solo Kapoho, a mutant frequency of 13.52% was produced (Fig. 6E). As a proportion of total mutants, 11.36% were frame-shift knockouts, while 2.16% were in-frame protein variants (Fig. 6E). Indels ranged from +1 to −10 bp with the deletion of −1 bp occurring with the highest frequency, followed by the addition of +1 bp at 2.47%, deletion of −3 bp at 1.19%, and deletion of −4 bp at 1.03% (Fig. 6F). Detailed results obtained targeting CpMLO6 in cv. Solo Sunset and Kapoho are provided (Tables 7, 8, Supplemental file 6). The average mutant frequency obtained targeting CpMLO6 in all three cultivars was 16.20 ± 1.53% (Supplemental file 5). 13.71 ± 1.67% of the generated mutants were frame-shift knockouts, while 2.50 ± 0.26% were in-frame protein variants (Supplemental file 5).
The measurement of DNA-free CRISPR-Cas9 gene editing efficiency in papaya protoplasts targeting the CpMLO6 gene as confirmed by NGS. Bar chart representations of total CpMLO6 CRISPR-Cas9 gene editing efficiencies (indels—black) and proportions of editing events leading to frame-shift knockouts (gray) and in-frame protein variants (light gray). The summary of mutants (A) and indels (B) generated for the CpMLO6 gene from cv. Solo Sunrise. The summary of mutants (C) and indels (D) generated for the CpMLO6 gene from the cv. Solo Sunset. The summary of mutants (E) and indels (F) generated for the CpMLO6 gene from the cv. Solo Kapoho.
Discussion
The Yield for Isolating Durable, Viable Papaya Leaf Mesophyll Protoplasts Has Substantially Improved
Protoplasts play an important role in plant research (Cocking 1972) with studies of protoplast fusion (Keller and Melchers 1973; Kao and Michayluk 1974; Zimmermann and Scheurich 1981) and hybridization (Kao et al. 1974; ; Melchers et al. 1978; Evans 1983; Dudits et al. 1987; Sihachakr et al. 1988), cell wall regeneration and cell division (Nagata and Takebe 1970; Kao et al. 1970; Vasil and Vasil 1972), nucleic acid delivery (Takebe and Otsuki 1969), cell signaling (Jang and Sheen 1994; Sheen 1996), stable transformation (Krens et al. 1983; Paszkowski et al. 1984), and transient transfection (Negrutiu et al. 1990; Abel and Theologis 1994). These early reports provide the basis for developing new uses of protoplasts, such as defining subcellular localizations of proteins (Yuen et al. 2013) and protein-protein interactions (Carrillo and Christopher 2022). Continuing along this line, we theorized that protoplasts offer an effective experimental system for gene editing of papaya. We demonstrated an improved method for isolating papaya mesophyll protoplasts for use in a DNA-free targeted gene editing system.
The results collected in 23 isolations reflect a 7-fold increase in the average protoplast yield of 1.11 × 108 ± 0.069 (Fig. 1E) compared to the previously published optimal protoplast yield of 1.56 × 107 ± 0.100 (Zhang et al. 2011). This increase in yield may be due to morphological or physiological differences between cultivars, or the extensive effort placed here in optimizing the protoplast isolation media. These optimizations include adjustments to enzyme concentration, incubation duration, temperature, pH, and osmolarity (Wu et al. 2017; Sangra et al. 2019; Adedeji et al. 2020; Xu et al. 2021). Plant tissues have been processed uniquely to remove the sometimes difficult to penetrate epidermal layer (Murata et al. 1994; Locatelli et al. 2003; Wu et al. 2009; Yuen et al. 2016). Vacuum infiltration is also widely used to enhance the removal of the formidable waxy cuticle on leaves (Newell and Luu 1985; Yoo et al. 2007; Yuen et al. 2013; Nanjareddy et al. 2016). The use of a cyclical vacuum infiltration and release method is believed to have improved protoplast yields. When the vacuum infiltration step was removed, the protoplast isolation failed. Additionally, when protoplasts were isolated under constant vacuum, protoplast yields were reduced. The cyclical vacuum infiltration method was critical for obtaining optimal results. Our papaya-optimized protoplast isolation method produces a significantly improved yield of highly viable and durable protoplasts suitable for research regardless of leaf age. However, protoplasts isolated from older plants were more durable in downstream steps and are thus recommended. The resulting viability of the protoplasts was 89.87 ± 2.02% (Fig. 2C), which is similar to the previously reported viability at 89.96 ± 2.89% (Zhang et al. 2011). In summary, the optimal conditions for obtaining viable papaya leaf mesophyll protoplast yields are 1.3% cellulase, 0.3% macerozyme, 0.44M mannitol, pH 5.8, 30-min vacuum infiltration with vacuum release every 5 min, and 13–15-h incubation period at 26°C and 60 rpm in the dark.
PEG-Mediated Transfection and Subcellular Localization of Fluorescent Fusion Proteins Were Optimized in Papaya Mesophyll Protoplasts
We used the eGFP reporter construct GFP(S65T) (Chiu et al. 1996) by itself and as the PDI9-GFP fusion protein for the following purposes: (a) To develop an efficient method for transient transfection of the protoplasts; (b) To verify proper GFP gene expression in vivo as an additional criterion for protoplast viability; (c) To confirm that transiently expressed proteins are properly targeted to subcellular compartments as indicators of preserved normal cellular functions. These results provided the foundation for subsequent gene editing experiments.
First, an optimized PEG-mediated method for transfection of protoplasts with reporter gene-plasmids was established for papaya. The fluorescent signal observed in transfected protoplasts confirmed plasmid uptake and viable expression of the construct (Fig. 3A). The efficiency of the PEG-mediated transient transfection method of 44%, reported here (Fig. 3D), is similar to or above the transfection efficiencies reported for some plant species, including potato at 49% (Konovalova et al. 2021), orchid hybrid ‘Ruili Beauty’ at 42% (Li et al. 2018), sugarcane at 41% (Wu et al. 2021), and cannabis at 23% (Matchett-Oates et al. 2021). Although efficiency results are below those reported for other plant species, including cassava at 71% (Wu et al. 2017), cabbage at 68% (Sivanandhan et al. 2021), cucumber at 57% (Huang et al. 2013), and the well-established Arabidopsis protoplast transfection method at 60–90% (Yoo et al. 2007). Interestingly, increased PEG concentrations and incubation times in the referenced plant species negatively affected transfection efficiency in some species, while positively affecting others, indicating that PEG-mediated protoplast transfection optimization is highly species-dependent. Although the efficiency of our papaya-optimized transfection method is less than 50%, as required for some applications (Yoo et al. 2007), it falls within the range of results reported for a diverse range of plant species.
Second, the Arabidopsis PDI9 gene encodes a protein folding chaperone of the ER secretory pathway (Yuen et al. 2013). We used cotransfection of PDI9:eGFP with the ER marker, ER:mCherry, to demonstrate that PDI9-GFP was clearly and correctly localized to the ER in papaya protoplasts as part of the secretory pathway (Fig. 3B). These results confirmed normal cell function and the ability of transfected protoplasts to viably express constructs delivered using our papaya-optimized PEG-mediated transfection method (Fig. 3C).
The CRISPR-Cas9 RNP-Based Gene Editing System in Papaya Corrected the GFP+1 Frame-Shift Mutation
The transient reporter assay used here containing the mutant frame-shift GFP (Liu et al. 2018) proved to be very effective in testing the competence of gene editing of papaya protoplasts. A CRISPR-Cas9 gene editing RNP complexed with a sgRNA (designed to target the GFPm mutation site) was able to edit the GFPm gene in the plasmid construct. A diverse range of indel mutations was introduced into the repaired GFPm sequence, some of which corrected the frame-shift and restored the wild-type (WT) GFP phenotype (Fig. 4A, B). The editing was confirmed using immunoblot analysis with a GFP-specific antibody to detect the fully translated GFP polypeptide.
The resulting GFPm editing efficiency of 27.88 ± 1.65% (Fig. 4C) was higher than the 3.3% reported by Liu et al. (2018). Significant differences existed between the methods used to introduce the CRISPR-Cas9 complexes into protoplasts. In switchgrass, protoplasts were cotransfected with a construct carrying a gene editing cassette and a separate construct carrying GFPm. Both constructs must be expressed and available in sufficient quantities to form complexes in the cell to edit the gene (Dewitt et al. 2017). In contrast, our system used the CRISPR-Cas9 complexed with the guide RNA and no plasmid expression of the editing components was needed.
From days 1 to 3, protoplast viability was reduced from 90 to ~40% when suspended in a mannitol-based solution, as reported in other plants (Poddar et al. 2020). The window of viable expression is not a significant concern using an RNP-based gene editing system, such as used here, which can perform targeted edits almost immediately upon entry into the cell (Kim et al. 2014). We believe this RNP-based system positively influenced the increased GFPm editing efficiency observed in papaya. Developing these parameters by using the in vitro complexed CRISPR-Cas9 RNP editor acting on a GFPm plasmid was a prerequisite to targeting genes of interest in the papaya genome.
The DNA-Free CRISPR-Cas9 RNP-Based System Successfully Edited Two Endogenous Genes, CpPDS and CpMLO6, of the Papaya Genome
The highest gene editing efficiencies for CpPDS and CpMLO6 reported here (46% and 19.15%, respectively (Figs. 5C and 6C), were similar to or above those reported for other plants when evaluated under similar conditions. For example, the PDS editing efficiencies were 0%, 1.33%, and 24.51% in oilseed rape, cabbage, and Chinese cabbage, respectively (Murovec et al. 2018), 1.8% in cabbage (Lee et al. 2020), and 0.92% in Cavendish banana (Wu et al. 2020). Results targeting MLO-like homologs were 11.3% in sweet pepper (Kim et al. 2020) and 0.1% in grapevine (Malnoy et al. 2016). The efficiency of RNP-based editing of various genes in plant protoplasts is highly variable between plant species and between gene targets (Zhang, et al. 2021). Such efficiencies include 60% in tomato (Lin et al. 2022), 2% in wild cabbage (Park et al. 2019), 9% in potato (Andersson et al. 2018), 45% in bread wheat (Liang et al. 2017), 7% in apple (Malnoy et al. 2016), and 44%, 19%, and 16% in tobacco, rice, and Arabidopsis, respectively (Woo et al. 2015). As described here, both mutant CpPDS and CpMLO6 produced indels in protoplasts at rates similar to or above those of other plant species. Results obtained while targeting CpPDS and CpMLO6 in the papaya cultivars Solo Sunrise, Sunset, and Kapoho were similar and consistent for their respective genes, with Sunrise consistently producing slightly higher editing efficiencies, while Sunset and Kapoho produced slightly reduced efficiencies. Therefore, an effective DNA-free gene editing system was established here to target the genome of papaya protoplasts, and the next step is the regeneration of these edited cells into whole plants.
Conclusions
This study used DNA-free CRISPR-Cas9 ribonucleoprotein complexes for gene editing in papaya protoplasts. The long-term goal is to enhance disease resistance for crop improvement. We first optimized the isolation of intact viable protoplasts from papaya leaves of various ages, providing a consistent and reliable source of plant material for gene editing experiments. Next, we tested the utility of the DNA-free CRISPR-Cas9 RNP gene editing system by correcting a frame-shift mutant GFP transgene, providing evidence of the method’s effectiveness. This approach involved preassembling Cas9 endonuclease with single-guide RNA complexes in vivo, then transfecting the protoplasts using PEG-mediated transfection. We then successfully employed this gene editing method to target the endogenous papaya genes, CpPDS and CpMLO6. We confirmed targeted editing of the endogenous papaya genes CpPDS and CpMLO6 through deep amplicon sequence verification. This transgene-free method eliminates the introduction of transgenic material into the host genome, reducing regulatory concerns and providing a more rapid and efficient alternative for papaya improvement. Furthermore, several advantages over traditional gene editing methods are realized, such as reduced off-target effects, decreased Cas9 transcription and translation time, and elimination of insertion mutations associated with stable transformation. The next step is to develop a regeneration protocol for the protoplasts, from different tissue sources, containing the edited CpPDS and CpMLO6 genes. The successful application of this method in papaya paves the way for future research and development to improve crop resistance to pathogens and other environmental threats.
References
Abel S, Theologis A (1994) Transient transformation of Arabidopsis leaf protoplasts: a versatile experimental system to study gene expression. Plant J 5:421–427
Adedeji OS, Naing AH, Kim CK (2020) Protoplast isolation and shoot regeneration from protoplast-derived calli of Chrysanthemum cv. White ND. Plant Cell Tissue Organ Cult 141:571–581
Agrios GN (2005) Plant Pathology, 5th edn. Elsevier Academic Press, Amsterdam
Andersson M, Turesson H, Olsson N, Fält A-S, Ohlsson P, Gonzalez MN, Samuelsson M, Hofvander P (2018) Genome editing in potato via CRISPR-Cas9 ribonucleoprotein delivery. Physiol Plant 164:378–384
Anzalone AV, Koblan LW, Liu DR (2020) Genome editing with CRISPR–Cas nucleases, base editors, transposases and prime editors. Nat Biotech 38:824–844
Arias RS, Dayan FE, Michel A, Howell JL, Scheffler BE (2006) Characterization of a higher plant herbicide-resistant phytoene desaturase and its use as a selectable marker. Plant Biotechnol J 4:263–273
Arumuganathan K, Earle ED (1991) Nuclear DNA content of some important plant species. Plant Mol Biol Report 9:208–218
Bai Y, Pavan S, Zheng Z, Zappel NF, Reinstädler A, Lotti C, De Giovanni C, Ricciardi L, Lindhout P, Visser R, Theres K, Panstruga R (2008) Naturally Occurring Broad-Spectrum Powdery Mildew Resistance in a Central American Tomato Accession Is Caused by Loss of Mlo Function. Mol Plant-Microbe Interact 21:30–39
Boss WF, Mott RL (1980) Effects of Divalent Cations and Polyethylene Glycol on the Membrane Fluidity of Protoplast. Plant Physiol 66:835–837
Brewer SE, Chambers AH (2022) CRISPR/Cas9-mediated genome editing of phytoene desaturase in Carica papaya L. J Hortic Sci Biotechnol 97:580–592
Brocklehurst K, Baines B, Kierstan M (1981) Papain and other constituents of Carica papaya L. Top Enzym Ferment Biotechnol 5:262–335
Büschges R, Hollricher K, Panstruga R, Simons G, Wolter M, Frijters A, Van Daelen R, Van Der Lee T, Diergaarde P, Groenendijk J, Töpsch S, Vos P, Salamini F, Schulze-Lefert P (1997) The Barley Mlo Gene: A Novel Control Element of Plant Pathogen Resistance. Cell 88:695–705
Carlos-Hilario LR, Christopher DA (2015) Improved Agrobacterium-mediated transformation of Carica papaya cultivar ‘Kapoho’ from embryogenic cell suspension cultures. In Vitro Cellular Dev Biol Plant 51:580–587
Carrillo R, Christopher DA (2022) Development of a GFP biosensor reporter for the unfolded protein response-signaling pathway in plants: incorporation of the bZIP60 intron into the GFP gene. Plant Signal Behav 17(1):2098645
Chen K, Wang Y, Zhang R, Zhang H, Gao C (2019) CRISPR/Cas Genome Editing and Precision Plant Breeding in Agriculture. Annu Rev Plant Biol 70:667–697
Chiu W-l, Niwa Y, Zeng W, Hirano T, Kobayashi H, Sheen J (1996) Engineered GFP as a vital reporter in plants. Curr Biol 6:325–330
Cho EJ, Yuen CYL, Kang B-H, Ondzighi CA, Staehelin LA, Christopher DA (2011) Protein disulfide isomerase-2 of Arabidopsis mediates protein folding and localizes to both the secretory pathway and nucleus, where it interacts with maternal effect embryo arrest factor. Mol Cells 32:459–475
Cocking EC (1972) Plant Cell Protoplasts-Isolation and Development. Annu Rev Plant Physiol 23:29–50
Cunningham B, Nelson S (2012) Powdery Mildew of Papaya in Hawaii. http://hdl.handle.net/10125/33214
Devoto A, Piffanelli P, Nilsson I, Wallin E, Panstruga R, von Heijne G, Schulze-Lefert P (1999) Topology, subcellular localization, and sequence diversity of the Mlo family in plants. J Biol Chem 274:34993–35004
Dewitt MA, Corn JE, Carroll D (2017) Genome editing via delivery of Cas9 ribonucleoprotein. Methods 121-122:9–15
Dudits D, Maroy E, Praznovszky T, Olah Z, Gyorgyey J, Cella R (1987) Transfer of resistance traits from carrot into tobacco by asymmetric somatic hybridization: Regeneration of fertile plants. Proc Natl Acad Sci U S A 84:8434–8438
El-Mounadi K, Morales-Floriano ML, Garcia-Ruiz H (2020) Principles, applications, and biosafety of plant genome editing using CRISPR-Cas9. Front Plant Sci 11. https://doi.org/10.3389/fpls.2020.00056
Evans DA (1983) Agricultural Applications of Plant Protoplast Fusion. Bio-Technology 1:253–261
Evans EA, Balen FH, Crane JH (2021) An overview of us papaya production, trade, and consumption. University of Florida IFAS Extension Pub. #FE914, pp 1–8
FAOSTAT (2020) Food and Agriculture Organization of the United Nations. https://www.fao.org/faostat/en/#data/QV
González MN, Massa GA, Andersson M, Turesson H, Olsson N, Fält A-S, Storani L, Décima Oneto CA, Hofvander P, Feingold SE (2020) Reduced enzymatic browning in potato tubers by specific editing of a polyphenol oxidase gene via ribonucleoprotein complexes delivery of the CRISPR/Cas9 System. Front Plant Sci 10. https://doi.org/10.3389/fpls.2019.01649
Gumtow R, Wu D, Uchida J, Tian M (2018) A Phytophthora palmivora Extracellular Cystatin-Like Protease Inhibitor Targets Papain to Contribute to Virulence on Papaya. Mol Plant-Microbe Interact 31:363–373
Hasley JAR, Navet N, Tian M (2021) CRISPR/Cas9-mediated mutagenesis of sweet basil candidate susceptibility gene ObDMR6 enhances downy mildew resistance. PLoS One 16:e0253245
He Y, Mudgett M, Zhao Y (2022) Advances in gene editing without residual transgenes in plants. Plant Physiol 188:1757–1768
Hewajulige IGN, Dhekney SA (2016) Papayas. In: Caballero B, Finglas PM, Toldrá F (eds) Encyclopedia of Food and Health. Academic Press, Oxford, pp 209–212
Hsu PD, Lander ES, Zhang F (2014) Development and Applications of CRISPR-Cas9 for Genome Engineering. Cell 157:1262–1278
Huang H, Wang Z, Cheng J, Zhao W, Li X, Wang H, Zhang Z, Sui X (2013) An efficient cucumber (Cucumis sativus L.) protoplast isolation and transient expression system. Sci Hortic 150:206–212
Jang JC, Sheen J (1994) Sugar sensing in higher plants. Plant Cell 6:1665–1679
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna Jennifer A, Charpentier E (2012) A Programmable Dual-RNA–Guided DNA Endonuclease in Adaptive Bacterial Immunity. Science 337:816–821
Kao KN, Constabel F, Michayluk MR, Gamborg OL (1974) Plant Protoplast Fusion and Growth of Intergeneric Hybrid Cells. Planta 120:215–227
Kao KN, Keller WA, Miller RA (1970) Cell division in newly formed cells from protoplasts of soybean. Exp Cell Res 62:338–340
Kao KN, Michayluk MR (1974) A Method for High-frequency Intergeneric Fusion of Plant Protoplasts. Planta 115:355–367
Kaur N, Alok A, Shivani KN, Pandey P, Awasthi P, Tiwari S (2018) CRISPR/Cas9-mediated efficient editing in phytoene desaturase (PDS) demonstrates precise manipulation in banana cv. Rasthali Gen Funct Integr Gen 18:89–99
Keller WA, Melchers G (1973) The Effect of High pH and Calcium on Tobacco Leaf Protoplast Fusion. Zeitschrift für Naturforschung C A J Biosci 28:737–741
Kim H, Choi J, Won K-H (2020) A stable DNA-free screening system for CRISPR/RNPs-mediated gene editing in hot and sweet cultivars of Capsicum annuum. BMC Plant Biol 20:449. https://doi.org/10.1186/s12870-020-02665-0
Kim S, Kim D, Cho SW, Kim J, Kim J-S (2014) Highly efficient RNA-guided genome editing in human cells via delivery of purified Cas9 ribonucleoproteins. Genome Res 24:1012–1019
Konovalova LN, Strelnikova SR, Zlobin NE, Kharchenko PN, Komakhin RA (2021) Efficiency of Transient Expression in Protoplasts of Various Potato Cultivars. Appl Biochem Microbiol 57:800–807
Krens FA, Wullems GJ, Schilperoort RA (1983) Transformation of Plant Protoplasts in Vitro. Springer, New York, Boston, MA, pp 387–408
Krishna L, Paridhavi M, Patel JA (2008) Review on nutritional, medicinal and pharmacological properties of papaya (Carica papaya L.). Indian J Nat Prod Resour 7:364–373
Lamour K (2013) Phytophthora a global perspective. CAB International; Cambridge, Mass
Larkin PJ (1976) Purification and viability determinations of plant protoplasts. Planta 128:213–216
Le Fevre R, O’Boyle B, Moscou MJ, Schornack S (2016) Colonization of Barley by the Broad-Host Hemibiotrophic Pathogen Phytophthora palmivora Uncovers a Leaf Development–Dependent Involvement of Mlo. Mol Plant-Microbe Interact 29:385–395
Lee MH, Lee J, Choi SA, Kim Y-S, Koo O, Choi SH, Ahn WS, Jie EY, Kim SW (2020) Efficient genome editing using CRISPR–Cas9 RNP delivery into cabbage protoplasts via electro-transfection. Plant Biotechnol Rep 14:695–702
Li J, Liao X, Zhou S, Liu S, Jiang L, Wang G (2018) Efficient protoplast isolation and transient gene expression system for Phalaenopsis hybrid cultivar ‘Ruili Beauty’. In Vitro Cell Dev Biol Plant 54:87–93
Liang Z, Chen K, Li T, Zhang Y, Wang Y, Zhao Q, Liu J, Zhang H, Liu C, Ran Y, Gao C (2017) Efficient DNA-free genome editing of bread wheat using CRISPR/Cas9 ribonucleoprotein complexes. Nat Commun 8:14261
Lin C-S, Hsu C-T, Yang L-H, Lee L-Y, Fu J-Y, Cheng Q-W, Wu F-H, Hsiao HCW, Zhang Y, Zhang R, Chang W-J, Yu C-T, Wang W, Liao L-J, Gelvin SB, Shih M-C (2018) Application of protoplast technology to CRISPR/Cas9 mutagenesis: from single-cell mutation detection to mutant plant regeneration. Plant Biotechnol J 16:1295–1310
Lin CS, Hsu CT, Yuan YH, Zheng PX, Wu FH, Cheng QW, Wu YL, Wu TL, Lin S, Yue JJ, Cheng YH, Lin SI, Shih MC, Sheen J, Lin YC (2022) DNA-free CRISPR-Cas9 gene editing of wild tetraploid tomato Solanum peruvianum using protoplast regeneration. Plant Physiol 188:1917–1930
Liu Y, Merrick P, Zhang Z, Ji C, Yang B, Fei S-Z (2018) Targeted mutagenesis in tetraploid switchgrass (Panicum virgatum, L.) using CRISPR/Cas9. Plant Biotechnol J 16:381–393
Locatelli F, Vannini C, Magnani E, Coraggio I, Bracale M (2003) Efficiency of transient transformation in tobacco protoplasts is independent of plasmid amount. Plant Cell Rep 21:865–871
Maas C, Werr W (1989) Mechanism and optimized conditions for PEG mediated DNA transfection into plant protoplasts. Plant Cell Rep 8:148–151
Malnoy M, Viola R, Jung M-H, Koo O-J, Kim S, Kim J-S, Velasco R, Nagamangala Kanchiswamy C (2016) DNA-Free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Front Plant Sci 7. https://doi.org/10.3389/fpls.2016.01904
Manshardt RM, Wenslaff TF (1989) Interspecific Hybridization of Papaya with Other Carica Species. J Am Soc Hortic Sci 114:689–694
Matchett-Oates L, Mohamaden E, Spangenberg GC, Cogan NOI (2021) Development of a robust transient expression screening system in protoplasts of Cannabis. In Vitro Cellular & Developmental Biology - Plant. https://doi.org/10.1007/s11627-021-10178-0
Melchers G, Sacristán MD, Holder AA (1978) Somatic hybrid plants of potato and tomato regenerated from fused protoplasts. Carlsb Res Commun 43:203–218
Metje-Sprink J, Menz J, Modrzejewski D, Sprink T (2019) DNA-Free Genome Editing: Past, Present and Future. Front Plant Sci 9:1957
Ming R, Hou S, Feng Y, Yu Q, Dionne-Laporte A, Saw JH, Senin P, Wang W, Ly BV, Lewis KLT, Salzberg SL, Feng L, Jones MR, Skelton RL, Murray JE, Chen C, Qian W, Shen J, Du P et al (2008) The draft genome of the transgenic tropical fruit tree papaya (Carica papaya Linnaeus). Nature 452:991–996
Ming R, Moore PH (2014) Genetics and Genomics of Papaya. Springer New York, New York, NY, USA
Murata T, Fukuoka H, Kishimoto M (1994) Plant Regeneration from Mesophyll and Cell Suspension Protoplasts of Sweet Potato, Ipomoea batatas (L.) Lam. Japan J Breeding 44:35–40
Murovec J, Guček K, Bohanec B, Avbelj M, Jerala R (2018) DNA-Free Genome Editing of Brassica oleracea and B. rapa Protoplasts Using CRISPR-Cas9 Ribonucleoprotein Complexes. Front Plant Sci 9:1594
Nadakuduti SS, Starker CG, Ko DK, Jayakody TB, Buell CR, Voytas DF, Douches DS (2019) Evaluation of methods to assess in vivo activity of engineered genome-editing nucleases in protoplasts. Front Plant Sci 10. https://doi.org/10.3389/fpls.2019.00110
Nagata T, Takebe I (1970) Cell Wall Regeneration and Cell Division in Isolated Tobacco Mesophyll Protoplasts. Planta 92:301–308
Nanjareddy K, Arthikala M-K, Blanco L, Arellano ES, Lara M (2016) Protoplast isolation, transient transformation of leaf mesophyll protoplasts and improved Agrobacterium-mediated leaf disc infiltration of Phaseolus vulgaris: tools for rapid gene expression analysis. BMC Biotechnol 16:53. https://doi.org/10.1186/s12896-016-0283-8
Navet N, Tian M (2020) Efficient targeted mutagenesis in allotetraploid sweet basil by CRISPR/Cas9. Plant Direct 4(6):e00233
Negrutiu I, Dewulf J, Pietrzak M, Botterman J, Rietveld E, Wurzer-Figurelli EM, Ye D, Jacobs M (1990) Hybrid genes in the analysis of transformation conditions: II. Transient expression vs stable transformation—analysis of parameters influencing gene expression levels and transformation efficiency. Physiol Plant 79:197–205
Nelson BK, Cai X, Nebenführ A (2007) A multicolored set of in vivo organelle markers for co-localization studies in Arabidopsis and other plants. Plant J 51:1126–1136
Nelson S (2008) Phytophthora Blight of Papaya. http://hdl.handle.net/10125/12413
Newell CA, Luu HT (1985) Protoplast culture and plant regeneration in Glycine canescens F.J. Herm. Plant Cell Tissue Organ Cult 4:145–149
Opalski KS, Schultheiss H, Kogel K-H, Hückelhoven R (2004) The receptor-like MLO protein and the RAC/ROP family G-protein RACB modulate actin reorganization in barley attacked by the biotrophic powdery mildew fungus Blumeria graminis f.sp. hordei. Plant J 41:291–303
Park S-C, Park S, Jeong YJ, Lee SB, Pyun JW, Kim S, Kim TH, Kim SW, Jeong JC, Kim CY (2019) DNA-free mutagenesis of GIGANTEA in Brassica oleracea var. capitata using CRISPR/Cas9 ribonucleoprotein complexes. Plant Biotechnol Rep 13:483–489
Parray ZA, Hassan MI, Ahmad F, Islam A (2020) Amphiphilic nature of polyethylene glycols and their role in medical research. Polym Test 82:106316
Paszkowski J, Shillito RD, Saul M, Mandák V, Hohn T, Hohn B, Potrykus I (1984) Direct gene transfer to plants. EMBO J 3:2717–2722
Pessina S, Pavan S, Catalano D, Gallotta A, Visser RG, Bai Y, Malnoy M, Schouten HJ (2014) Characterization of the MLO gene family in Rosaceae and gene expression analysis in Malus domestica. BMC Genomics 15:618
Poddar S, Tanaka J, Cate JHD, Staskawicz B, Cho M-J (2020) Efficient isolation of protoplasts from rice calli with pause points and its application in transient gene expression and genome editing assays. Plant Methods 16:151. https://doi.org/10.1186/s13007-020-00692-4
Porter BW, Christopher DA, Zhu YJ (2014) Genomics of Papaya Disease Resistance. In: Ming R, Moore PH (eds) Genetics and Genomics of Papaya. Springer New York, New York, NY, pp 277–307
Sangra A, Shahin L, Dhir SK (2019) Optimization of Isolation and Culture of Protoplasts in Alfalfa ( Medicago sativa ) Cultivar Regen-SY. Am J Plant Sci 10:1206–1219
Sheen J (1996) Ca2+-Dependent Protein Kinases and Stress Signal Transduction in Plants. Science 274:1900–1902
Sidorov V, Wang D, Nagy ED, Armstrong C, Beach S, Zhang Y, Groat J, Yang S, Yang P, Gilbertson L (2021) Heritable DNA-free genome editing of canola (Brassica napus L.) using PEG-mediated transfection of isolated protoplasts. In Vitro Cell Dev Biol Plant. https://doi.org/10.1007/s11627-021-10236-7
Sihachakr D, Haicour R, Serraf I, Barrientos E, Herbreteau C, Ducreux G, Rossignol L, Souvannavong V (1988) Electrofusion for the production of somatic hybrid plants of Solanum melongena L. and Solanum khasianum C.B. Clark Plant Sci 57:215–223
Sivanandhan G, Bae S, Sung C, Choi S-R, Lee G-J, Lim Y-P (2021) Optimization of Protoplast Isolation from Leaf Mesophylls of Chinese Cabbage (Brassica rapa ssp. pekinensis) and Subsequent Transfection with a Binary Vector. Plants 10:2636
Sorek R, Lawrence CM, Wiedenheft B (2013) CRISPR-Mediated Adaptive Immune Systems in Bacteria and Archaea. Annu Rev Biochem 82:237–266
Subburaj S, Chung SJ, Lee C, Ryu S-M, Kim DH, Kim J-S, Bae S, Lee G-J (2016) Site-directed mutagenesis in Petunia × hybrida protoplast system using direct delivery of purified recombinant Cas9 ribonucleoproteins. Plant Cell Rep 35:1535–1544
Takebe I, Otsuki Y (1969) Infection of Tobacco Mesophyll Protoplasts by Tobacco Mosaic Virus. Proc Natl Acad Sci U S A 64:843–848
Tian S, Jiang L, Gao Q, Zhang J, Zong M, Zhang H, Ren Y, Guo S, Gong G, Liu F, Xu Y (2017) Efficient CRISPR/Cas9-based gene knockout in watermelon. Plant Cell Rep 36:399–406
van Schie CCN, Takken FLW (2014) Susceptibility Genes 101: How to Be a Good Host. Annu Rev Phytopathol 52:551–581
Vasil IK, Vasil V (1972) Totipotency and Embryogenesis in Plant Cell and Tissue Cultures [with Discussion]. In Vitro 8:117–127
Wang M, Mao Y, Lu Y, Tao X, Zhu J-K (2017) Multiplex Gene Editing in Rice Using the CRISPR-Cpf1 System. Mol Plant 10:1011–1013
Woo JW, Kim J, Kwon SI, Corvalán C, Cho SW, Kim H, Kim S-G, Kim S-T, Choe S, Kim J-S (2015) DNA-free genome editing in plants with preassembled CRISPR-Cas9 ribonucleoproteins. Nat Biotechnol 33:1162–1164
Wu F-H, Shen S-C, Lee L-Y, Lee S-H, Chan M-T, Lin C-S (2009) Tape-Arabidopsis Sandwich - a simpler Arabidopsis protoplast isolation method. Plant Methods 5:16
Wu J-Z, Liu Q, Geng X-S, Li K-M, Luo L-J, Liu J-P (2017) Highly efficient mesophyll protoplast isolation and PEG-mediated transient gene expression for rapid and large-scale gene characterization in cassava (Manihot esculenta Crantz). BMC Biotechnol 17:29–29
Wu S, Zhu H, Liu J, Yang Q, Shao X, Bi F, Hu C, Huo H, Chen K, Yi G (2020) Establishment of a PEG-mediated protoplast transformation system based on DNA and CRISPR/Cas9 ribonucleoprotein complexes for banana. BMC Plant Biol 20:1
Wu Z-D, Hu X, Zan F-G, Pan Y-B, Burner DM, Luo Z-Y, Liu J-Y, Zhao L-P, Yao L, Zhao Y, Liu X-L, Xia H-M, Yang K, Zhao J, Zhao P-F, Qin W, Chen X-K, Wu C-W (2021) Subcellular Localization of the D27 Protein in Sugarcane (Saccharum spp. Hybrids) Using an Optimized Protoplast Isolation, Purification, and Transient Gene Expression Protocol. Sugar Tech 23:316–325
Xu X-f, Zhu H-y, Ren Y-f, Feng C, Ye Z-h, Cai H-m, Wan X-c, Peng C-y (2021) Efficient isolation and purification of tissue-specific protoplasts from tea plants (Camellia sinensis (L.) O. Kuntze). Plant Methods 17:84–84
Yoo S-D, Cho Y-H, Sheen J (2007) Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis. Nat Protoc 2:1565–1572
Yuen C, Matsumoto K, Christopher D (2013) Variation in the Subcellular Localization and Protein Folding Activity among Arabidopsis thaliana Homologs of Protein Disulfide Isomerase. Biomolecules 3:848–869
Yuen CYL, Shek R, Kang B-H, Matsumoto K, Cho EJ, Christopher DA (2016) Arabidopsis protein disulfide isomerase-8 is a type I endoplasmic reticulum transmembrane protein with thiol-disulfide oxidase activity. BMC Plant Biol 16:1–15
Zhang A, Liu Y, Wang F, Li T, Chen Z, Kong D, Bi J, Zhang F, Luo X, Wang J, Tang J, Yu X, Liu G, Luo L (2019a) Enhanced rice salinity tolerance via CRISPR/Cas9-targeted mutagenesis of the OsRR22 gene. Molec Breed 39:1
Zhang J, Shen W, Yan P, Li X, Zhou P (2011) Factors that influence the yield and viability of protoplasts from Carica papaya L. African J Biotech 10:5137–5142
Zhang S, Shen J, Li D, Cheng Y (2021a) Strategies in the delivery of Cas9 ribonucleoprotein for CRISPR/Cas9 genome editing. Theranostics 11:614–648
Zhang Y, Iaffaldano B, Qi Y (2021b) CRISPR ribonucleoprotein-mediated genetic engineering in plants. Plant Commun 2:100168–100168
Zhang Z, Hua L, Gupta A, Tricoli D, Edwards KJ, Yang B, Li W (2019b) Development of an Agrobacterium-delivered CRISPR/Cas9 system for wheat genome editing. Plant Biotechnol J 17:1623–1635
Zhu YJ, Agbayani R, Moore PH (2007a) Ectopic expression of Dahlia merckii defensin DmAMP1 improves papaya resistance to Phytophthora palmivora by reducing pathogen vigor. Planta 226:87–97
Zhu YJ, Agbayani R, Nishijima W, Moore P (2007b) Characterization of disease resistance of Carica papaya to Phytophthora. Acta horticulturae, 10.17660/ActaHortic.2007.740.32:265-269
Zimmermann U, Scheurich P (1981) High Frequency Fusion of Plant Protoplasts by Electric Fields. Planta 151:26–32
Zubrod JP, Bundschuh M, Arts G, Brühl CA, Imfeld G, Knäbel A, Payraudeau S, Rasmussen JJ, Rohr J, Scharmüller A, Smalling K, Stehle S, Schulz R, Schäfer RB (2019) Fungicides: An Overlooked Pesticide Class? Environ Sci Technol 53:3347–3365
Acknowledgements
We thank Dr. Bing Yang for the GFPm construct and advice on its use, and Mrs. Tina Weatherby for expert advice on microscopy.
Dr. Bing Yang
Bond Life Sciences Center
Division of Plant Sciences
University of Missouri
1201 Rollins Street, Columbia, MO 65211
Tina Weatherby
Biological Electron Microscope Facility
Life Sciences Building
University of Hawai’i at Mānoa
1800 East-West Road, Honolulu, HI 96822
Funding
This work was supported by the University of Hawai’i at Mānoa and the USDA-NIFA-AFRI award 2020-67013-31549.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible Editor: Yong Eui Choi
Supplementary information
ESM 1
Supplemental file 1: Phylogenetic analysis of the candidate S-gene MLO-like protein homologs in papaya. (PDF 392 kb)
ESM 2
Supplemental file 2: The secondary structure of selected sgRNA candidates targeting Carica papaya endogenous genes, CpPDS and CpMLO6. (PDF 379 kb)
ESM 3
Supplemental file 3: In vitro DNA cleavage activity of CpPDS-sgRNA16 and CpMLO6-sgRNA254. (PDF 247 kb)
ESM 4
Supplemental file 4: Table 1: Indels generated during initial DNA-free CRISPR-Cas9 targeted gene editing of CpPDS in papaya cv. Solo Sunrise. Table 2: Indels generated during replicate DNA-free CRISPR-Cas9 targeted gene editing of CpPDS in papaya cv. Solo Sunrise. Table 3: Indels generated during DNA-free CRISPR-Cas9 targeted gene editing of CpPDS in papaya cv. Solo Sunset. Table 4: Indels generated during DNA-free CRISPR-Cas9 targeted gene editing of CpPDS in papaya cv. Solo Kapoho. (PDF 2013 kb)
ESM 5
Supplemental file 5: Summary of DNA-free CRISPR-Cas9 targeted gene editing of CpPDS and CpMLO6 in papaya cultivars Solo Sunrise, Sunset, and Kapoho. (PDF 206 kb)
ESM 6
Supplemental file 6: Table 5: Indels generated during initial DNA-free CRISPR-Cas9 targeted gene editing of CpMLO6 in papaya cv. Solo Sunrise. Table 6: Indels generated during replicate DNA-free CRISPR-Cas9 targeted gene editing of CpMLO6 in papaya cv. Solo Sunrise. Table 7: Indels generated during DNA-free CRISPR-Cas9 targeted gene editing of CpMLO6 in papaya cv. Solo Sunset. Table 8: Indels generated during DNA-free CRISPR-Cas9 targeted gene editing of CpMLO6 in papaya cv. Solo Kapoho. (PDF 1700 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Elias, M.J., Hasley, J., Tian, M. et al. Development of a Mesophyll Protoplast-Based System for Gene Editing of Papaya. In Vitro Cell.Dev.Biol.-Plant 59, 517–535 (2023). https://doi.org/10.1007/s11627-023-10373-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11627-023-10373-1