Abstract
The green phyllodocids Eulalia clavigera and E. viridis are a known European pseudo-cryptic complex, but questions about its distribution and evidence of additional lineages in previous studies call for an investigation of the real diversity within the complex.
We analyze DNA sequences (mtCOI-5P, ITS, and 28S rRNA) of different populations of E. clavigera from intertidal and subtidal marine waters along the North East Atlantic, Mediterranean Sea, the Azores and Webbnesia (Madeira, Savage islands and Canaries), and populations of E. viridis from the Scandinavia. This provided compelling evidence for the existence of six additional divergent evolutionary lineages, three of the most abundant being described here as new species: Eulalia feliciae sp. nov., intertidal and unique to the Western Mediterranean, Eulalia madeirensis sp. nov., subtidal and unique to the Madeira Island (Portugal), and Eulalia xanthomucosa sp. nov., mostly subtidal and occurring in the British Isles and southern France. Complementary morphometric analyses showed that E. feliciae sp. nov. and E. madeirensis sp. nov. formed two independent morphometric clusters, while E. xanthomucosa sp. nov. often overlapped with E. clavigera sensu stricto (s. s.), although being unique in showing a yellow coloration and parapodial cirri on median segments larger in relation to its body size.
Recent biotechnological findings based on “E. clavigera” specimens highlight the importance of formally describing cryptic complexes, since each lineage chemistry might be unique and may have a range of distinct effects and applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Biodiversity may vary at genetic, species, and ecosystem levels, with the true animal diversity being appraised in-deep by using molecular tools to detect cryptic or pseudo-cryptic species. Pseudo-cryptic complexes constitute a substantial fraction of biodiversity and appear to be frequent among marine benthic invertebrates (Miglietta et al., 2011; Nygren, 2014) in well-known taxa and areas (e.g., Bleidorn et al., 2006; Carr et al., 2011; Grosse et al., 2021; Jolly et al., 2006; Leite et al., 2020; Lobo et al., 2017). Despite the increasing evidence of cryptic species occurrence, the lack of formal taxonomic description (Fernandez-Triana, 2022) hinders accurate estimates of their contribution to biodiversity (Delić et al., 2017; Fišer et al., 2018; Hutchings & Kupriyanova, 2018); therefore, limiting our understanding of their evolutionary and ecological significance, as well as their recognition in large scale biomonitoring programs using high throughput sequence technologies.
The homogeneously green phyllodocid, Eulalia viridis (Linnaeus, 1767), has been reported from throughout the northern hemisphere (Eibye-Jacobsen, 1991, 1993). It is common in intertidal and subtidal coastal areas and marinas, down to 50 m depth (Ushakov, 1972), usually living on rocky reefs in crevices, among algae, mussel beds, Balanus spp. blocks, Dendropoma reefs, Posidonia oceanica (Linnaeus) Delile, 1813 meadows, and coralligenous formations (Bonse et al., 1996; Viéitez et al., 2004; Çinar, 2005). However, in Mediterranean Sabellaria alveolata (Linnaeus, 1767) reefs, it is replaced by Eulalia ornata Saint-Joseph 1888, faint yellow but sometimes greenish species morphologically similar to E. viridis, except for the pigmentation pattern (Schimmenti et al., 2016).
Based on isoelectric focusing and morphological data, a correlation between exclusive isoenzymes and protein patterns, the morphology and size of the mid-body dorsal cirri, and the size of the proboscideal papillae allowed to discriminate between two distinct groups of Eulalia populations (Bonse et al., 1996). The morphotype from the North Sea and Scandinavian coasts, showing smaller papillae and slender dorsal cirri, corresponded to E. viridis, while that from France and England showing larger papillae and significantly thicker dorsal cirri was attributed to Eulalia clavigera (Audouin & Milne-Edwards, 1833), hitherto synonymized with E. viridis. The reproductive biology of phyllodocids is poorly known, and these two species are not an exception. They have planktonic larval stages and reproduce once a year (Meyer, 1938), but Northern Europe populations also differ locally in reproduction time, with reproductive cycle starting four to six weeks earlier in Sweden than in England and France (Olive, 1975; Pleijel, 1993). The green Eulalia sp. is a predator feeding mostly on mussels and barnacles (Rodrigo et al., 2015), with occasional scavenger habits also being observed (Morton, 2011). Molecular studies based on the mitochondrial cytochrome c oxidase subunit I (COI) also allowed distinguishing two populations of Eulalia viridis from Kandalaksha Bay (Russia) and Portugal, which showed more than 20% Kimura’s two parameter (K2P) genetic divergence (Hardy et al., 2011; Lobo et al., 2016). Their highly similar morphology and the large number of discriminating genetic markers certainly imply the existence of a pseudo-cryptic species complex. A South-Western population of E. clavigera from Patagonia did not show any genetic differentiation toward the North-Eastern Atlantic specimens, supporting a non-native origin of the Patagonian population (Langeneck et al., 2019). However, Mediterranean specimens examined by Langeneck et al. (2019) represented a distinct lineage compared to the Patagonia/North East Atlantic clade.
Given the high number of species complexes among phyllodocid genera, such as Notophyllum (Nygren et al., 2010) or Eumida (Nygren & Pleijel, 2011; Teixeira et al., 2022), and among other polychaete families (Grosse et al., 2021; Lobo et al., 2016; Sampieri et al., 2021), the actual diversity and distribution of European species of the Eulalia viridis/clavigera species group is questioned. In this study, we use a combined multi-locus and morphometric approach to compare the European populations of E. clavigera, from the United Kingdom to Portugal, the Azores and Webbnesia (Madeira, Savage islands and Canaries), and the Western and Eastern Mediterranean Sea, as well as molecular data for populations of E. viridis from the Scandinavia to investigate the possible occurrence of additional diagnosable species within the Eulalia viridis/clavigera complex.
Methods
Taxon sampling and molecular data retrieval
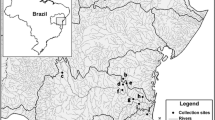
We sampled a total of 131 specimens of Eulalia, belonging to the Eulalia clavigera/viridis complex, three of Eulalia aurea Gravier, 1896 and one of Phyllodoce Lamarck, and 1818 distributed along the European coasts, including the Azores and Webbnesia (Fig. 1). Worms were collected at low tide in rocky beaches among algae and mussels in marinas or subtidal areas up to 34 meters in depth and fixed in 96% ethanol. Samples were harvested in continental Portugal (Canto Marinho, Leixoes, Aveiro, and Nazaré), Santa Maria (Azores) and Madeira islands, Spain (La Coruña and the islands of Tenerife, Gran Canaria, and La Palma), France (Roscoff, Morgat, Banyuls, and Corsica), Great Britain (Plymouth and Cornwall), Norway (Espevær , Grimstad, Bergen, Trondheim, and Finmark), Sweden (Koster), Italy (Livorno, Ischia island and Taranto), and Croatia (Istria) (Table 1, Table S1).
Sampling sites. Abbreviations as in Table 1
We sequenced a partial segment of COI 5’ end (mtCOI-5P) from 119 specimens and a representative number per location for the ITS-regions (i.e., ITS1, 5.8S rRNA, and ITS2) and 28S rRNA (Table 2). MtCOI-5P sequences belonging to Eulalia cf. clavigera from Capraia Island and port of Stintino, Italy (Langeneck et al., 2019), were mined from GenBank for comparison purposes. Eulalia aurea and Phyllodoce sp. were used as outgroups. DNA was extracted, amplified, sequenced, and assembled as described in Nygren et al. (2010) or Lobo et al. (2016), depending of the lab and materials available, from a few parapodia or a small portion of the posterior end of the specimens. Morphological and molecular vouchers are deposited in the Research Collection of Marine Invertebrates of the Department of Biology of the University of Aveiro (CoBI-DBUA), in the French Muséum national d'Histoire naturelle (MNHN; specimens MNHN-IA-2021-654 and MNHN-IA-2021-655), Scripps Oceanographic Institution (SIO; French Mediterranean specimen BI-2014/15-077), National Museum of Science and Natural History (MUHNAC, Portugal; specimen MB29-000385 from the Savage islands) and in Arne Nygren’s private collection (ANPC; specimens MTE040-20, MTE042-20, MTE052-20, MTE053-20, MTE054-20, MTE055-20, MTE057-20, MTE079-20, MTE080-20, MTE081-20, and MTE088-20) using the Barcode Of Life Data (BOLD) Process ID ("MTE", http://v4.boldsystems.org/) for Arne Nygren’s specimens.
The full dataset and its metadata can be accessed at BOLD Systems under the project “Eulalia Species Complex (DS-MTE)” (DOI: https://doi.org/10.5883/DS-MTEC), except for the four COI sequences from Langeneck et al. (2019). GenBank accession numbers: OP898309-OP898427 (mtCOI-5P), OP897856-OP897897 (ITS region), and OP897898-OP897939 (28S) (Table S1).
Phylogeny and genetic distances
Maximum likelihood (ML) and Bayesian inference (BI) were used for the phylogenetic analyses. The nuclear markers (ITS-regions and 28S) and mtCOI-5P locus were aligned with MAFFT online (https://mafft.cbrc.jp/alignment/server/, Katoh & Standley, 2013) and concatenated with MEGA 10.0.05 (Kumar et al., 2018). Highly variable regions, extensive gaps, and poorly aligned positions (mainly present in the ITS-regions) were eliminated using Gblocks 0.91b (http://phylogeny.lirmm.fr/phylo_cgi/one_task.cgi?task_type=gblocks; Castresana, 2000), allowing all options for a less stringent selection and not allowing many contiguous non-conserved positions, so that it becomes more suitable for a phylogenetic analyses. We used MrBayes 3.1.2 (Ronquist & Huelsenbeck, 2003) for Bayesian analysis. Best-fit models were selected using the Akaike Information Criterion (AIC) in the JModeltest software (Darriba et al., 2012; Guindon & Gascuel, 2003). For mtCOI-5P, we applied the Hasegawa-Kishino-Yano gamma distributed rates across sites (HKY + G) for the first two positions and the HKY model with equal rates across sites for the third position. Regarding the concatenated ITS with 28S, we applied the General Time Reversible (GTR) model with equal rates across sites. The number of generations was set to 107, and the sample frequency every 500 generations. Twenty-five percent of the samples were discarded as burn-in (burninfrac = 0.25). The resulting tree files were checked for convergence in the effective sampling sizes (ESSs > 200) with Tracer 1.6 software (Rambaut et al., 2018) and then analyzed in Figtree 1.4.3 (https://tree.bio.ed.ac.uk/software/figtree/). The final concatenated tree was edited with the software Inkscape 0.92.3 (https://www.inkscape.org). ML phylogenies were performed in MEGA 10.0.05 with 1000 bootstrap runs with the GTR model and equal rates across sites for the concatenated dataset. Only the BI tree is shown in Results, including the ML support if topologies are similar. The alignments (FASTA and NEXUS format) for each individual marker and the concatenated ones are all publicly available at Figshare (DOI: https://doi.org/10.6084/m9.figshare.21657416).
The mean genetic distances (K2P) within and between molecular operational taxonomic units (MOTUs) were calculated in MEGA 10.0.05 using the same GBlock alignment as indicated for nuclear loci.
MOTU clustering
To depict MOTUs, we delineated by applying three delineation methods to both the concatenated mitochondrial and nuclear alignments, except for mtCOI-5P, where we also applied the Barcode Index Number (BIN) implemented in BOLD (Ratnasingham & Hebert, 2013). The Automatic Barcode Gap Discovery (ABGD, Puillandre et al., 2012) was implemented online with default settings using the K2P distance matrix (https://bioinfo.mnhn.fr/abi/public/abgd/abgdweb.html). The Generalized Mixed Yule Coalescent (GYMC) single threshold model (Fujisawa & Barraclough, 2013) and the Poisson Tree Processes (bPTP, Zhang et al., 2013) were implemented online (https://species.h-its.org/). The Bayesian ultrametric tree for the GYMC was generated with BEAST 2.4.6 (Bouckaert et al., 2014) with the appropriate best model (based on AIC criteria; GTR equal rates), and four independent runs for 5—107 Markov Chain Monte Carlo (MCMC) generations, sampled every 5,000 generations. The Tracer 1.6 software was used to estimate ESS convergence > 200 for all parameters. The consensus tree was obtained using TreeAnnotator 2.4.6 (Bouckaert et al., 2014) and loaded into Figtree. ML phylogenies contributed for the bPTP results. A final consensus MOTUs was chosen using the majority rule (i.e., most common number of MOTUs).
Genetic diversity and structure
Haplotype networks were made through the PopART software (Leigh & Bryant, 2015). The Templeton, Crandall, and Sing method (TCS, Clement et al., 2002) was used to evaluate the relationship between haplotypes and their geographical distribution. GBlocks was not applied in this analysis to avoid underestimating the number of nuclear haplotypes. Indices of genetic diversity (number of haplotypes, h; haplotype diversity, hd; polymorphic sites, S; nucleotide diversity, Pi; Fu & Li D and Tajima D statistical tests) were estimated based on mtCOI-5P for each MOTU using DNASP 5.10 (Librado & Rozas, 2009).
Morphometric analysis
Specimens from four lineages of Eulalia were morphometrically compared against each other to complement the molecular data. The remaining lineages had less than three available specimens with a very small size, being therefore unsuitable for morphometric studies and thus have not been named. A minimum of five specimens per population with optimal conditions having all the selected morphological characters and whenever possible, similar in size, were chosen.
The following characters were selected and measured (Fig. 2, supplementary Table S2): the number of segments; the length (mm) of the worm from the prostomium to the pygidium, chaetigerous lobes, terminal antennae, palps, middle antenna, dorsal and ventral tentacular cirrus from the second segment, dorsal and ventral cirri from median segments, and head; the width (mm) of the worm with and without parapodia (to the tip of the chaetigerous lobes, excluding chaetae) from median segments, head and dorsal and ventral cirri from median segments; and distance between eyes and height (mm) of the chaetigerous lobes. The distance between eyes was measured from the center of the eyespots to avoid possible fixation biases, following Martin et al. (2017). Measurements were done with a LEICA MC170 HD stereo microscope, with an incorporated measurement software. Photographs of live specimens were taken with a Canon EOS1100D camera.
Scheme of the morphotype of Eulalia clavigera, showing the morphometric measurements used for analysis. a Anterior end. b Parapodia. Abbreviations: CLL, length of chaetigerous lobes; CLH, height of chaetigerous lobes; AL, length of antennae; PL, length of palps; MAL, length of middle antenna; DTL, length of dorsal tentacular cirri on segment 2; VTL, length of ventral tentacular cirri on segment 2; DCL, length of dorsal cirri; VCL, length of ventral cirri; HL, length of head; WWP, width of worm with parapodia; WW, width of worm without parapodia; HW, width of head; DCW, width of dorsal cirri; VCW, width of ventral cirri; DE, distance between the eyes
To minimize bias based on size variability, measurements taken to analyze the inter-lineage differences were compared as proportions to create scatter plots between relevant morphological characters and evaluate relationships between the measurements, similar to previous studies (Ravara et al., 2010; Teixeira et al., 2020, 2022). The significance of the inter-population differences using ratios of proportion data (e.g., dividing the length of dorsal cirri against the length of ventral cirri) was also explored by one-way analysis of similarity (ANOSIM), with normalized data based on Euclidean distance resemblance matrices, using PRIMER (2008, version 6.1.11, Plymouth, UK). All remaining analyses were conducted using Microsoft Excel (Office 365 ProPlus).
Results
Phylogenetic reconstruction
Without any variation in the different delineation methods, nine MOTUs are retrieved from the concatenated Bayesian phylogenetic tree (Fig. 3a), belonging to monophyletic clades with low intraspecific variation (<3%): the previously described E. clavigera s.s. (MOTU 4) and E. viridis (MOTU 7), six undescribed lineages of Eulalia, and the MOTU GB1 from Langeneck et al. (2019). Besides MOTU 4, major clade A englobes four additional MOTUs genetically close to E. clavigera s.s. with high bootstrap support: MOTU 1 (unique to the western Mediterranean), MOTU 2 (unique to the subtidal habitats from Madeira), MOTU 3 (the unnamed Eulalia KRO53, from the Eastern Mediterranean Sea), and lastly, MOTU GB1 (the Mediterranean Eulalia cf. clavigera).
Phylogenetic tree and respective COI haplotypes and MOTU locations. a Phylogenetic tree reconstructed using Bayesian inference based on concatenated COI, ITS regions and 28S sequences, with information regarding the different MOTU delineation methods. BINs were used only for COI. MOTU GB1 only have COI sequences and was not present in BOLD systems preventing BIN analysis. Only the bootstrap support over 0.85 BI and 85 ML is shown. Each different consensus MOTU is represented by the respective number, with the different colors corresponding to the respective geographic distribution. Live photo belong to the specimen DBUA0002474.02.v01, measuring around 45 mm in length (WL) and exhibiting greenish color. b Haplotype network based on COI for all the analyzed MOTUs and outgroups (OUTG). Each haplotype is represented by a circle and number of haplotypes are according to the displayed scale. Numbers correspond to the number of mutational steps between haplotypes. Lines without numbers means only one mutation between haplotypes
MOTUs 5 and 6 (major clade B) are sister to each other and fall outside the main “clavigera” clade. MOTU 5 is present in the British Isles and the Western Mediterranean. The subtidal MOTU 6, together with MOTUs 3 and 8, also subtidal, includes few and very small specimens in relatively poor conditions or exhausted in the DNA analysis and has not been named or used in the morphometric analysis. Eulalia aurea was revealed as an actual ingroup for the E. clavigera/viridis species complex.
Genetic distances
The global mean genetic distances (K2P) for the Eulalia complex and outgroups is detailed in Table 3. Apart from the outgroups and Eulalia IT2-1 (MOTU 8), the mean intraspecific distances are 0.93% (0.0–3.3%), with an average congeneric distance of 17.9% (7.1–25.5%) for mtCOI-5P; 1.4% (0.0–3.9%) and 17.2% (4.4–32.6%), respectively, for the ITS region intra- and inter-specific divergence; and 0.04% (0–0.4%) and 2.7% (0–5.9%) for the 28S intra- and inter-specific divergence, respectively. The populations of E. clavigera s.s. from continental Europe, and Azores and the Canary islands (MOTU 4) show mtCOI-5P maximum distances up to 3.3% and no considerable divergence (<1%) in the nuclear markers. Eulalia IT2-1 has a particularly high inter-specific distance in the nuclear markers, reaching more than 60% for the ITS region and 12% for the 28S locus, similar to those in the Phyllodoce sp. (outgroup).
Haplotype networks
Only the 28S network (Fig. 4b) fail to discriminate all the identified MOTUs from the concatenated dataset. It is characterized by a star-shape phylogeny, with most of the unique haplotypes closely related to the common central haplotype, which is composed by MOTUs 1, 3, and 4. However, MOTU 1 also has a distinct haplotype from the common central one, with a similar number of mutations apart, as the outgroup E. aurea.
Haplotypes networks based on ITS (a) and 28S (b) for all MOTUs and outgroups, except MOTU GB1. Each haplotype is represented by a circle and number of haplotypes are according to the displayed scale. Colors indicate the geographic location of the haplotype. Numbers correspond to the number of mutational steps between haplotypes. Lines without numbers means only one mutation between haplotypes
The mtCOI-5P (Fig. 3b) and ITS (Fig. 4a) networks reveal geographically structured populations within MOTU 4, between continental Europe, and Azores and the Canary islands, corresponding to two distinct clades in the BI tree. However, they do not have diverged enough to be divided into two separate MOTUs. Correlations of certain COI haplotypes or parts of the haplotype network with a specific biogeographic region can be found in Madeira (MOTU 2), Scandinavia (MOTU 7), Eastern Mediterranean (MOTU 3), and South of France (MOTU 1).
The mtCOI-5P haplotype diversity is relatively high (Hd > 0.89 to 0.985, Table 4) or even extreme, with almost all specimens having an unique haplotype as in MOTUs 1 (8 haplotypes in 10 specimens) and 2 (11 in 12). None of the MOTUs have a significant Tajima D and Fu and Li’s D tests, with the neutral model of nucleotide substitutions being accepted for all the lineages.
Morphometric measurements
The analyses of the most significative morphometric proportions show the formation of independent clusters among the analyzed species (Figs. 5a-h), except for the often-overlapping MOTU 5 and MOTU 4 (E. clavigera s. s.). However, despite E. clavigera s. s. being considerably larger (both in number of segments and body length and width), MOTU 5 shows similar measurements for dorsal and ventral cirri, head, dorsal tentacular cirri on segment 2, and antennae (Fig. 5a, g). This was further highlighted in Table 5, where very significative proportion ratios (ANOSIM R values close to 1 with p value less than 5%) between these two species were found, in particular, for the length of the dorsal cirri of median segments vs. body width and length (Fig. 5e, f). Global ANOSIM tests for the proportion ratios used in Fig. 5 were also considerably high and the different ratio values are summarized in Table 6 for all the analyzed lineages. The continental Europe and Canarian populations of E. clavigera s.s. (MOTU 4) do not differ considerably, except for the larger worm length and number of segments for the latter (Fig. 5f and Table 6). MOTUs 1 and 2 can be mainly distinguished from E. clavigera s. s. based on the comparisons between antennae length vs. palp length, length of dorsal tentacular cirri on segment 2 vs. worm width, and distance between eyes vs. head length (Table 6). MOTU 2 shows smaller proportions than the remaining MOTUs, while MOTU 1 mainly appears in an intermediate position. Moreover, besides geographic distribution and live coloration, the length between the chaetigerous lobe vs. ventral cirri seem to be the most significative ratio distinguishing MOTUs 1 and 2 (Table 5), with MOTU 2 having longer chaetigerous lobes and smaller ventral cirri.
Scatter plots with the most considerable proportions in distinguishing E. clavigera s.s. (populations from mainland Europe and Canary Islands), E. feliciae sp. nov., E. madeirensis sp. nov., and E. xanthomucosa sp. nov. Significance of the proportion ratios by analysis of similarity (ANOSIM) between all the lineages (Global R) is also indicated. a Morphometric proportions between the length of the ventral cirri (VCL) and the length of the dorsal cirri (DCL). b Between the length of the head (HL) and the length of the antennae (AL). c Between the length of the chaetigerous lobe (CLL) and the length of the ventral cirri (VCL). d Between the length of the head (HL) and the width of the head (HW). e Between the width of the worm from median segment (WW) and length of the dorsal cirri (DCL). f Between the length of the worm (WL) and the length of the dorsal cirri (DCL). g Between the width of the worm (WW) and length of the dorsal tentacular cirrus on segment 2 (DTL). h Between the lenght of the head (HL) and the distance between the eyes (DE)
Live, relaxed specimens of Eulalia (WL: worm length). a Eulalia feliciae sp. nov., specimen DBUA0002478.01.v07, dorsal view, greenish coloration. b Eulalia madeirensis sp. nov., specimen DBUA0002479.01.v03, dorsal view, faint yellowish/light green coloration. c Eulalia xanthomucosa sp. nov., specimen from the Natural History Museum, London (left) and specimen BI-2014/15–077 (right), dorsal view, bright yellow coloration
Taxonomic account
Eulalia clavigera/viridis species complex.
Diagnosis
Body anteriorly stout and posteriorly tapered, variable in size (length, width, and number of segments). Living specimens may be yellow, but usually green including the pharynx, varying in intensity; once preserved, the pigment fades off into a greenish hue and turning to brownish when aging. Darker, glandular spots usually present laterally from prostomium, in dorsal cirri, along posterior segmental margins but missing in mid-dorsal regions and basally in parapodia. Prostomium rounded triangular, wider than long. Eyes medium-large, rounded, occasionally partly covered by segment 1. Distance between eyes (from center of eyespots) either shorter or as long as head length. Median antenna well in front of eyes, as long as apical ones. Palps as long or slightly longer than antennae. Proboscis widest distally, densely covered with rounded to conical papillae. Terminal ring with varying number of papillae. Longest tentacular cirri may be shorter or slightly longer than body width from median segments. Tentacular cirri of segment 1 reaching about segments 3–5, half as long as dorsal tentacular cirri of segment 2. Dorsal tentacular cirri of segment 2 reaching about segments 6-9, almost two times as long as ventral tentacular cirri (reaching about segments 3-6, often thick and slightly flattened). Dorsal tentacular cirri of segment 3 reaching about segments 6-9, as long as dorsal tentacular cirri of segment 2. Dorsal cirri of median segments asymmetrically lanceolate, about twice as long than wide. Dorsal cirri 2-3 times longer than ventral cirri in median segments. Ventral cirri rounded longer than wide, either smaller or slightly longer than chaetigerous lobes in median segments. Chaetae usually from segment 3, occasionally 1-2 chaetae arising from anterior side of ventral cirrophores of segment 2. Rostrum of chaetal shaft with one large tooth and numerous smaller ones either side, decreasing in size proximally. Blades short. Two pygidial cirri about three times as long as wide, with pointed tips. Median pygidial papilla absent.
Remarks
The description of four, from the eight European lineages belonging to the E. clavigera/viridis complex, is provided below, namely, for E. clavigera s.s., E. feliciae sp. nov., E. madeirensis sp. nov., and E. xanthomucosa sp. nov. Eulalia viridis, found both in intertidal and subtidal habitats is also part of the complex, as described in Pleijel (1993). The undescribed intertidal Eulalia cf. clavigera (MOTU GB1), first detected in Langeneck et al. (2019), belongs to the complex, as well as the two remaining rarer and unnamed subtidal lineages from the current study (Eulalia KRO53 and Eulalia IS-BA).
Besides E. clavigera s.s. which is widespread in the North East Atlantic, Azores, Webbnesia, and Western Mediterranean, the remaining lineages seem to be locally restricted. Intertidal specimens from the Azores and Canary archipelagos and North East Atlantic are somewhat similar regarding size, color pattern, and morphological features. Specimens belonging to the “clavigera” clade (clade A, Fig. 3a) from the Mediterranean or the subtidal habitats of Madeira (Portugal) are often smaller, may present a yellowish-green (apart from the usual bright or emerald green) live color, more pointed dorsal cirri, and slightly more elongate prostomium. Kato et al. (2001) compiled several Eulalia species having green pigmentation, but the only one having uniform pigmentation was E. viridis. The latter study apparently overlooked the re-description of E. viridis from Bonse et al. (1996), which reinstated E. clavigera. Schimmenti et al. (2016) added additional data by summarizing the main morphological differences, including coloration, between E. ornata, E. viridis, and E. clavigera s. s.
The original description highlighted changes in coloration after fixation, where the overall color of Phyllodoce clavigera, now Eulalia clavigera, is reported as bright green but, through the action of ethanol, changes to metallic brown (Audouin and Milne-Edwards, 1833). Langeneck et al. (2019) indicated a homogeneous coloration in their specimens, but living animals are paler ventrolaterally. Moreover, the pharynx distal papillae change their shape depending on the sample treatment and non-relaxed specimens have globose, low papillae whereas osmotic shocked specimens have them thin, better defined.
Key to the five European Eulalia species analyzed in this study:
-
1. Live specimens with bright yellow coloration; length of the dorsal cirri more than three times longer than ventral cirri of median segments ... E. xanthomucosa sp. nov.
-
— Live specimens with greenish coloration; length of the dorsal cirri less than three times longer than ventral cirri of median segments ..................................................... 2
-
2(1). Ventral cirri of median segments slightly longer or as long as chaetigerous lobe; relatively large eyes; unique to the Scandinavian region .............................. E. viridis
-
— Ventral cirri of median segments clearly shorter than chaetigerous lobe; medium-sized eyes; absent in Scandinavian region ............................................................... 3
-
3(2). Palps as long as antennae; dorsal tentacular cirri on segment 2 clearly shorter than body width of median segments ............................................... E. clavigera s. s.
-
— Palps slightly longer than antennae; dorsal tentacular cirri on segment 2 as long or slightly longer than body width of median segments ........................................... 4
-
4(3). Small-sized worm; clearly longer chaetigerous lobes than ventral cirri (1.5x); subtidal and unique to Madeira Island (Portugal) .......................... E. madeirensis sp. nov.
-
— Small to medium-sized worm; chaetigerous lobes as long or slightly longer than ventral cirri (1.2x); usually intertidal and unique to the Western Mediterranean ... E. feliciae sp. nov.
Eulalia clavigera (Audouin & Milne Edwards, 1833) s.s.
Phyllodoce clavigera Audouin and Milne-Edwards 1833: 226–228, PL. XVI, Fig. 9–13.
Eulalia clavigera: Bonse et al., 1996: 40–45, Fig. 14 (redescr., syn.); Alós, 2004: 193–196, Fig. 69 (SEM photographs).
? Eulalia viridis: Morgado and Amaral, 1983: 51 (non Linnaeus, 1767).
Material examined
Portugal: Aveiro, 6 spms, DBUA0002468.01.v01-v06, 40° 33′ 32.4″ N–8° 46′ 19.2″ W, low tide, among rocks with algae and mussels, collected by Marcos A. L. Teixeira and Ascensão Ravara, 27/07/2018; Canto Marinho, 7 spms, DBUA0002469.02.v01-v07, 41° 44′ 13.2″ N–8° 52′ 33.6″ W, low tide, among rocks with algae and mussels, collected by Marcos A. L. Teixeira, 20/05-/2019; Areosa, 3 spms, DBUA0002469.01.v01-v03, 41° 42′ 36.0″ N–8° 52′ 12.0″ W, low tide, among rocks with algae and mussels, collected by Marcos A. L. Teixeira, 20/03/2018; Leixões, 1 spm, DBUA0002470.01.v01, 41° 10′ 58.8″ N–8° 42′ 18.0″ W, marina in pontoon scrapings, collected by Sofia Duarte, 23/06/2020; Nazaré, 3 spms, DBUA0002493.01.v01-v03, 39° 36′ 13.0″ N–9° 04′ 44.0″ W, low tide, among rocks with algae, mussels, barnacles, and Sabellaria tubes, collected by Ascensão Ravara, 26/07/2021; Santa Maria (Azores), 2 spm, DBUA0002477.01.v01-v02, 36° 57′ 03.6″ N–25° 01′ 04.8″ W, low tide, among rocks with algae and mussels, collected by Ana Costa, 07/05/2019; Savage islands, 1 spm, MB29-000,385, 30° 08′ 23.9″ N–15° 51′ 57.6″ W, low tide, among rocks with algae, kindly provided by the National Museum of Science and Natural History (MUHNAC, Portugal), collected in 22/06/2010. Spain: Ferrol lagoon (La Coruña), 5 spms, DBUA0002473.01.v01-v05, 43° 30′ 07.2″ N–8° 09′ 32.4″ W, low tide, among rocks with algae and mussels, collected by Julio Parapar, 03/02/2015; Tenerife (Canary islands), 11 spms, DBUA0002476.01.v01-v11, 28° 34′ 15.6″ N–16° 20′ 02.4″ W, low tide, among rocks with algae, collected by Marcos A. L. Teixeira, Pedro E. Vieira, José Carlos Hernández, and Beatriz Alfonso, 05/04/2019; La Palma (Canary islands), 10 spms, DBUA0002476.02.v01-v10, 28° 48′ 18.0″ N–17° 45′ 43.2″ W, low tide, among rocks with algae, collected by Marcos A. L. Teixeira and Pedro E. Vieira, 09/04/2019; Gran Canaria (Canary islands), 5 spms, DBUA0002476.03.v01-v05, 27° 59′ 06.0″ N–15° 22′ 33.6″ W, low tide, among rocks with algae, collected by Marcos A. L. Teixeira and Pedro E. Vieira, 06/04/2019. France: Roscoff, 8 spm, DBUA0002471.01.v01-v08, 48° 43′ 33.6″ N–3° 58′ 40.8″ W, low tide, among rocks with algae and mussels, collected by Arne Nygren, 20/03/2018; Morgat, 2 spms, DBUA0002472.01.v01-v02, low tide, among rocks with algae, 48° 13′ 20.3″ N–4° 29′ 42.5″ W, collected by Nicolas Lavesque, 16/06/2018. Great Britain: Plymouth, 12 spms, DBUA0002474.01.v01-v05, DBUA0002474.02.v01-v05 and DBUA0002474.03. v01-v02, 50° 21′ 25.2″ N–4° 07′ 40.8″ W, low tide, among rocks with algae and mussels, collected by Arne Nygren and Fredrik Pleijel, 18/03/2006. Italy: Livorno, 3 spms, DBUA0002475.01.v01-v03, 43° 32′ 24.0″ N–10° 18′ 00.0″ E, marina in pontoon scrapings, collected by Joachim Langeneck, 20/09/2019.
Measurements
Medium to large-sized worms. Complete specimens up to 275 segments, 68 mm total length and 2 mm maximum width if parapodia included (smallest specimens: 30 mm long, 1.6 mm wide, and 155 segments).
Diagnosis
Living specimens deep green, fading to a greenish hue when preserved and turning to brownish when aging. Prostomium rounded triangular, 1.7 times wider than long. Eyes medium-sized, rounded, occasionally partly covered by segment 1. Distance between eyes (from center of eyespots) either shorter than or as long as head length. Palps as long as antennae. Proboscis widest distally, densely covered with rounded to conical papillae. Terminal ring with varying number of papillae. Longest tentacular cirri clearly shorter than body width from median segments. Tentacular cirri of segment 1 reaching about segment 3, half as long as dorsal tentacular cirri of segment 2. Dorsal tentacular cirri of segment 2 reaching about segment 7, usually 1.9 times as long as ventral tentacular cirri (reaching about segment 3-4, often thick and slightly flattened). Dorsal tentacular cirri of segment 3 reaching about segment 7, as long as dorsal cirri of segment 2. Dorsal cirri of median segments asymmetrically lanceolate, about twice as long than wider. Dorsal cirri 2.5 times longer than ventral cirri in median segments. Ventral cirri rounded 1.5 times longer than wide; twice as short as chaetigorous lobes.
Molecular data
ITS, 28S, and mtCOI-5P sequences as in specimens DBUA0002468.01.v01-v05, DBUA0002469.01.v01-v03, DBUA0002469.02.v01-v05, DBUA0002470.01.v01, DBUA0002471.01. v01-v05, DBUA0002472.01.v01-v02, DBUA0002473.01.v01-v05, DBUA0002474.01.v01-v05, DBUA0002474.02.v01-v05, DBUA0002474.03.v01-v02, DBUA0002475.01.v01-v03, DBUA000 2476.01.v01-v05, DBUA0002476.02.v01-v07, DBUA0002476.03.v01-v05, DBUA0002477.01.v01-v02, and MB29-000385 (Table S1). Eulalia clavigera s. s. clearly differs from the remaining species of the complex, grouping in MOTU 4 (Fig. 3a). Interspecific mtCOI-5P mean distances to the closest and distant neighbor are 7.5% (K2P, Eulalia KRO53) and 23.3% (K2P, E. viridis), respectively. DOI for the species’ Barcode Index Number (BIN): https://doi.org/10.5883/BOLD:AAY5110.
Distribution and habitat
North East Atlantic Ocean (United Kingdom, France, Iberian Peninsula) to the Western Mediterranean (Western Italy) and the Canaries, Azores, and Savage archipelagos; introduced in the South-western Atlantic Ocean (Langeneck et al., 2019). Type locality: Brittany, France.
Usually present in intertidal rocky areas with algae, mussels, and barnacles, in association with reefs of Sabellaria and in marinas among the algae attached to pontoons.
Reproduction
The available data most likely belong to different lineages (i.e., Scandinavian E. viridis, and English and French E. clavigera). It has planktonic larvae and reproduces once a year (Meyer, 1938), but the Northern European populations differ in reproduction time, with reproductive cycle starting 4-6 weeks earlier in Swedish than in English and French specimens (Olive, 1975; Pleijel, 1993).
Remarks
Eulalia clavigera was synonymized with E. viridis by McIntosh (1908) and later reinstalled by Bonse et al. (1996). These two species slightly differ in prostomial, parapodial, and pharynx papillation features, although the length-to-width ratio of dorsal cirri is the most distinguishing character (Bonse et al., 1996). Eulalia viridis has smaller papillae and slender dorsal cirri than E. clavigera, the former being restricted to Scandinavian and Northern Sea coasts and seeming to be an intertidal or subtidal northern boreal and sub-arctic species. Conversely, E. clavigera is a temperate species, mostly found in intertidal rocky beaches, occurring from Great Britain to the Western Mediterranean Sea, but also in the Azores, Savage, and Canary Islands. Besides the feeding on mussels and barnacles reported in previous studies (Rodrigo et al., 2015), the intertidal Canarian specimens were observed preying on other smaller polychaetes as well.
Phyllodoce gervillei from Granville (France) was erected by Audouin and Milne Edwards (1833), stating that it was identical to P. clavigera, except for lacking median antenna and having smaller tentacular cirri. It was also synonymized under E. viridis by McIntosh (1908), who considered the absence of antennae as accidental. However, the type locality of P. gervillei led us to assume it is most probably a synonym of E. clavigera.
Specimens from the type locality of E. clavigera (Brittany, France) were collected for this study and grouped in MOTU 4 (Fig. 3a). Only the number of segments and worm length allowed partial separation of the continental European from Canarian populations (Fig. 5f, Table 6). Similarly, despite grouping in two distinct clades and having unique mtCOI-5P and ITS haplotypes, they only diverge up to 3.3% (mtCOI-5P, K2P) and cluster in the same MOTU. Eulalia clavigera s. s. usually shows larger proportions in most diagnostic characters than those of the other three species of the complex here described (Fig. 5a-h), especially the longer chaetigerous lobe vs. smaller ventral cirri, the head length/width ratio, and much larger body width vs. smaller dorsal tentacular cirri of segment 2, opposed to the inverse ratio seen in the remaining species of the complex (Table 6). Only E. xanthomucosa sp. nov. can often share the same morphometric clusters, especially regarding the length of the dorsal cirri, although it tends to show considerably less segments and shorter body length and width than E. clavigera s.s.
Eulalia feliciae sp. nov.
(Fig. 6a).
Urn:lsid:zoobank.org:act: 9BE4ED06-563E-43C0-9873-D105D88032E4.
Material examined
Type material. France: Banyuls, 1 spm, holotype and hologenophore, DBUA0002478.01.v05, 42° 28′ 48.0″ N–3° 08′ 06.0″ E, near shore at 0.5–1 m depth, rocky beach, collected by Arne Nygren and Fredrik Pleijel, 22/04/2001; 5 spms, paratype and paragenophores, DBUA0002478.01.v01-v04 and DBUA0002478.01.v06, 42° 28′ 48.0″ N–3° 08′ 06.0″ E, near shore at 0.5–1 m depth, rocky beach, collected by Arne Nygren and Fredrik Pleijel, 22/04/2001.
Other material. France: Banyuls, 2 spms, DBUA0002478.01.v07and MTE040-20, 42° 28′ 48.0″ N–3° 08′ 06.0″ E, near shore at 0.5–1 m depth and subtidal at 10 m depth respectively, among algae, rocks, and mussels, collected by Arne Nygren and Fredrik Pleijel, 02/04/2009; 2 spms, DBUA0002478.01.v08 and MTE042-20, 42° 28′ 48.0″ N–3° 08′ 06.0″ E, subtidal at 10 m depth and near shore at 0.5–1 m depth respectively, among rocks with hydroids, collected by Arne Nygren and Fredrik Pleijel, 05/04/2009.
Measurements
Small to medium-sized worms. Complete specimens up to 135 segments, 14 mm total length and 0.6 mm maximum width if parapodia included (smallest: 9 mm long, 0.5 mm wide, and 93 segments). Holotype lacking posterior end, 14 mm long, 0.6 mm wide, and 135 segments.
Diagnosis
Living specimens deep emerald green (Fig. 6a) fading to a greenish hue once preserved, also when aging. Prostomium rounded triangular, 1.3 times wider than long. Eyes medium-sized, rounded, occasionally partly covered by segment 1. Distance between eyes (from center of eyespots) shorter than head length. Palps slightly longer than antennae. Proboscis widest distally, densely covered with rounded to conical papillae. Longest tentacular cirri slightly longer than body width from median segments. Tentacular cirri of segment 1 reaching segment 3-4, half as long as dorsal tentacular cirri of segment 2. Dorsal tentacular cirri of segment 2 reaching about segment 6-7, usually 1.8 times as long as ventral tentacular cirri (reaching segment 4, often thick and slightly flattened). Dorsal tentacular cirri of segment 3 reaching about segment 6-7, as long as dorsal tentacular cirri of segment 2. Dorsal cirri of median segments asymmetrically lanceolate, about 2.4 times longer than wide. Dorsal cirri 2.2 times longer than ventral cirri in median segments. Ventral cirri of median segments twice longer than wide; slightly shorter than chaetigerous lobes.
Molecular data
ITS, 28S, and mtCOI-5P sequences as in specimens DBUA0002478.01.v01-v08, MTE040-20, and MTE042-20 (Table S1). Eulalia feliciae sp. nov. clearly differs from the remaining species of the complex, grouping in MOTU 1 (Fig. 3a). Inter-specific mtCOI-5P mean distances to the closest and distant neighbor are 13.9% (K2P, E. clavigera s.s.) and 22% (K2P, E. viridis), respectively. DOI for the species’ Barcode Index Number (BIN): https://doi.org/10.5883/DS-MTEFBIN (BIN: BOLD:AEC0502).
Etymology
The new species is named after Felicia Ulltin, a former master student under the supervision of the last author, whose enthusiasm and love for polychaetes are unmatched and an inspiration for future marine researchers.
Distribution
Mediterranean Sea: South of France. Usually present in intertidal or subtidal rocky areas among algae, hydroids and mussels.
Remarks
Besides the molecular data and geographical distribution, E. feliciae sp. nov. can be distinguished from E. clavigera s.s. and the remaining species from the complex mostly by the deep emerald green coloration in vivo. Moreover, it shows morphometric proportions in most diagnostic characters larger when compared to E. madeirensis sp. nov. but smaller against E. clavigera s. s. and E. xanthomucosa sp. nov. (Fig. 5a-h). The most significative proportions are the ratio between the length of the dorsal vs. ventral cirri, the length of chaetigorous lobe vs. ventral cirri, the length of antennae vs. head, the head length/width, or the distance between eyes vs. head length (Table 6). Moreover, the dorsal tentacular cirri on segment 2 is longer than body width, opposed to the inverse ratio seen in E. clavigera s.s.; chaetigerous lobes are only as long or slightly longer than ventral cirri (1.2x), compared to the much larger ratio seen in the other described species; and similar to E. madeirensis sp. nov., it shows slightly longer palps than antennae (Table 6). Furthermore, the dorsal and ventral cirri length ratio is considerably smaller (2.2 times), when compared to E. xanthomucosa sp. nov. (3.2 times), E. madeirensis sp. nov. (2.6 times), and E. clavigera s.s (2.5 times).
Eulalia madeirensis sp. nov.
(Fig. 6b).
Urn:lsid:zoobank.org:act: A107B872-462C-4E36-B15C-5CE02AA71FB7.
Material examined
Type material. Portugal: Madeira (Funchal), 1 spm, holotype and hologenophore, DBUA0002479.01.v02, 32° 38′ 09.6″ N–16° 55′ 51.6″ W, subtidal, 11 m depth, collected by Arne Nygren, 21/09/2009; 4 spms, paratypes and paragenophores, DBUA0002479.01.v01, DBUA0002479.01.v03, DBUA0002479.01.v04-v06, 32° 38′ 09.6″ N–16° 55′ 51.6″ W, subtidal, 11 m depth, collected by Arne Nygren, 21/09/2009.
Other material. Portugal: Madeira (Funchal), 1 spm, MTE052-20, 32° 38′ 09.6″ N–16° 55′ 51.6″ W, subtidal, 11 m depth, collected by Arne Nygren, 21/09/2009; Madeira (Porto Moniz), 4 spms, DBUA0002479.02.v01, MTE053-20, MTE055-20, and MTE057-20, 32° 51′ 38.6″ N–17° 09′ 06.3″ W, subtidal, 11 m depth, collected by Arne Nygren, 30/09/2009.
Measurements
Small-sized worms. Complete specimens up to 115 segments, 10 mm total length, and 0.4 mm maximum width if parapodia included (smallest specimen: 4 mm long, 0.3 mm wide, and 52 segments). Holotype lacking posterior end, 10 mm long, 0.4 mm wide, and 115 segments.
Diagnosis
Living specimens yellowish to light green (Fig. 6b), fading to greenish brown once preserved. Prostomium rounded triangular, 1.4 times wider than long. Eyes medium-sized, rounded, occasionally partly covered by segment 1. Distance between eyes (from center of eyespots) slightly shorter than head length. Palps slightly longer than antennae. Proboscis widest distally, densely covered with rounded to conical papillae. Longest tentacular cirri slightly longer than body width from median segments. Tentacular cirri of segment 1 reaching segment 3-4, as less than half long as dorsal tentacular cirri of segment 2. Dorsal tentacular cirri of segment 2, reaching segment 8, usually 1.7 times longer than ventral tentacular cirri (reaching segment 4-5, being often thick and slightly flattened). Dorsal tentacular cirri of segment 3 reaching about segment 8, as long as dorsal cirri from segment 2. Dorsal cirri of median segments asymmetrically lanceolate, about twice longer than wide. Dorsal cirri 2.6 times longer than ventral cirri in median segments. Ventral cirri of median segments rounded 1.6 times longer than wide; 1.5 times shorter than chaetigorous lobes, much shorter in posterior body half.
Molecular data
ITS, 28S, and mtCOI-5P sequences as in specimens DBUA0002479.01.v01-v06, DBUA0002479.02.v01, MTE052-20- MTE055-20, and MTE057-20 (Table S1). Eulalia madeirensis sp. nov. clearly differs from the remaining species of the complex, grouping in MOTU 2 (Fig. 3a). Inter-specific mtCOI-5P mean distances to the closest and distant neighbor are 11.4% (K2P, E. clavigera s.s.) and 23.3% (K2P, E. viridis), respectively. DOI for the species’ Barcode Index Number (BIN): https://doi.org/10.5883/BOLD:AEC0503.
Etymology
The new species is named after Madeira, the main island of the archipelago and the unique remote location where this species can be found so far.
Distribution and habitat
Atlantic Ocean: Exclusive to Madeira Island (Portugal), in subtidal environments up to 11 m depth.
Remarks
Besides the molecular data, bathymetric ranges and geographical distribution, E. madeirensis sp. nov. can be distinguished from E. clavigera s.s. and the remaining species of the complex mainly by the yellowish light green coloration in vivo, its smaller size and smaller morphometric proportions in most diagnostic characters (Fig. 5h). The most significative ones being the ratio between the length of the dorsal vs. ventral cirri, the length of chaetigorous lobe vs. ventral cirri, the length of antennae vs. head, the head length/width ratio, or the distance between eyes vs. head length (Table 6). Moreover, the dorsal tentacular cirri on segment 2 is longer than body width, opposed to the inverse ratio seen in E. clavigera s.s.; chaetigerous lobes are longer than ventral cirri (1.5x); and similar to E. feliciae sp. nov., it shows slightly longer palps than antennae (Table 6). Furthermore, the dorsal and ventral cirri length ratio is considerably smaller (2.6 times), when compared to E. xanthomucosa sp. nov. (3.2 times).
Further subtidal sampling in other islands from the Azores and Webbnesia is required to see if this species is endemic to Madeira or widespread throughout the different archipelagos.
Eulalia xanthomucosa sp. nov.
(Fig. 6c).
urn:lsid:zoobank.org:act: 70600CFB-9A4D-43D7-9AC5-97BF79FFA046.
Material examined
Type material. United Kingdom: Cornwall (Newlyn Marina), 1 spm, holotype and hologenophore, DBUA0002480.01.v07, 50° 06′ 10.8″ N–5° 32′ 49.2″ W, subtidal at 25 m depth, among coralligenous samples, collected by David Fenwick, 02/06/2016; 3 spms, paratypes and paragenophores, DBUA0002480.01.v01-v03, 50° 06′ 10.8″ N–5° 32′ 49.2″ W, lower shore in a rock crevice, collected by David Fenwick, 02/07/2016; 3 spms, paratypes and paragenophores, DBUA0002480.01.v04-v06, 50° 06′ 10.8″ N–5° 32′ 49.2″ W, subtidal at 25 m depth, in rock crevices at Laminaria zones and among coralligenous, collected by David Fenwick, 22–08-2017.
Other material. France: Banyuls, 1 spm, BI-2014/15–077, 42° 28′ 48.0″ N–3° 08′ 06.0″ E, subtidal at 25 m depth, among algae and boulders, collected by Fredrik Pleijel, 07/04/2009; 1 spm, DBUA0002481.01.v01, 42° 50′ 37.0″ N–3° 14′ 12.0″ E, subtidal at 25 m depth, among coralligenous, collected by Felicia Ulltin, 15/09/2020. France: Corsica Island, 2 spms, MNHN-IA-2021–654 and MNHN-IA-2021–655, 41°26'49.2"N–8°54'00.0"E26.8′ N, subtidal at 34 m depth, collected by the CORSICABENTHOS expeditions, 23/10/2020.
Measurements
Medium to large-sized worms. Complete specimens up to 230 segments, 104 mm total length, and 2.378 mm maximum width if parapodia included (smallest specimen: 12 mm long, 0.397 mm wide, and 89 segments). Holotype lacking posterior end, 26 mm long, 1.2 mm wide, and 128 segments.
Diagnosis
Living specimens bright yellow, enhanced by the prevalent yellowish mucus (Figs. 6c and 7a, b), fading to brownish once preserved. Prostomium rounded triangular, 1.1 times wider than long. Eyes medium to largemedium-sized, rounded, occasionally partly covered by segment 1. Distance between eyes (from center of eyespots) clearly shorter than head length. Palps as long as antennae. Proboscis not seen. Longest tentacular cirri slightly longer than body width from median segments. Tentacular cirri of segment 1 reaching segment 4-5, half as long as dorsal tentacular cirri of segment 2. Dorsal tentacular cirri of segment 2 reaching about segment 8-9, usually 1.8 times as long as ventral tentacular cirri (reaching segment 5-6, often thick and slightly flattened). Dorsal tentacular cirri of segment 3 reaching about segment 8-9, as long as dorsal cirri from segment 2. Dorsal cirri of median segments asymmetrically lanceolate, about 2.3 times longer than wide. Dorsal cirri 3.2 times longer than ventral cirri in median segments. Ventral cirri of median segments 1.5 times longer than wide; 1.7 times shorter than chaetigerous lobes.
Live, relaxed specimens from E. xanthomucosa sp. nov. exhibiting yellow coloration and high prevalence of yellowish mucus. a Mucus present in the posterior end of the body (specimen from David Fenwick’s private collection). b Mucus present in the median part of the body (specimen from David Fenwick’s private collection)
Molecular data
ITS, 28S, and mtCOI-5P sequences as in specimens DBUA0002480.01.v01-v07, DBUA0002481.01.v01, BI-2014/15–077, MNHN-IA-2021–654, and MNHN-IA-2021–655 (Table S1). Eulalia xanthomucosa sp. nov. clearly differs from the remaining species of Eulalia, grouping in MOTU 5 (Fig. 3a). Inter-specific mtCOI-5P mean distances to the closest and distant neighbor are 12.1% (K2P, Eulalia IS-BA) and 20.4% (K2P, E. feliciae sp. nov.), respectively. DOI for the species’ Barcode Index Number (BIN): https://doi.org/10.5883/BOLD:AEC0501.
Etymology
The new species is named based on its unique bright yellow (“xantho” from ancient Greek) coloration, which is further enhanced by the prevalent and equally yellowish mucus.
Distribution and habitat
Atlantic Ocean: United Kingdom, Cornwall; Mediterranean Sea: France, Banyuls and Corsica Island. Occasional lower intertidal but typically shallow sublittoral in rock crevices at Laminaria zones, among coralligenous material in marinas.
Remarks
This species was registered at the Natural History Museum as Eulalia sp. “Emits Yellow Mucus A” (tvk NHMSYS0021180023, https://www.aphotomarine.com/worm_eulalia_species_28-09-11.html). Eulalia xanthomucosa sp. nov. differs from E. clavigera s.s. and the remaining species from the complex in live coloration (bright yellow instead of green, with a prevalent presence of yellow mucus; Fig. 7a, b), but may be confused with the also yellowish E. aurea. However, it also differs from both E. aurea and E. clavigera s. s. in having dorsal cirri of median segments much longer in relation to body size (Fig. 5e, f). This feature is more noticeable the larger the specimen is (Fig. 6c). Eulalia aurea further differs from E. xanthomucosa sp. nov. by the presence of two mid-dorsal red lines (in live specimens only), and two lateral darker lines (Pleijel, 1993). Moreover, the dorsal tentacular cirri on segment 2 is usually longer than body width, opposed to the inverse ratio seen in E. clavigera s.s. The latter species has been found living together with E. xanthomucosa sp. nov. in Newlyn Marina (Cornwall, United Kingdom), but it usually occurs higher on the shore than E. xanthomucosa sp. nov. Marinas are a nurse area for numerous species of epibionts that occur as part of the biofouling community (Hadfield, 2011; Nedved & Hadfield, 2008). Based on our observations, it seems E. xanthomucosa sp. nov., a predatory phyllodocid, is recruited to the Newlyn Marina due to the plenty availability of food.
Eulalia xanthomucosa sp. nov. presents larger morphometric proportions in most of the diagnostic characters when compared with E. feliciae sp. nov. and E. madeirensis sp. nov., (Fig. 5a-h) especially between the length of the dorsal vs. ventral cirri, the length of chaetigerous lobe vs. ventral cirri, the length of antennae vs. head, the head length/width ratio, or the distance between eyes vs. head length (Table 6). However, E. clavigera s.s. may have a similar antennae vs. palps ratio and it can often share the same morphometric cluster (Fig. 5a, b, h), but with the analyzed specimens being considerably larger (i.e., longer, wider and with more segments) than the ones from E. xanthomucosa sp. nov. Additionally, the eyes are clearly larger for the new species (https://www.aphotomarine.com/images/marine_worms2/worm_eulalia_sp_nov_29-04-22_9.jpg).
Some specimens from E. xanthomucosa sp. nov. can also reach similar size as the larger ones belonging to E. clavigera s.s. (e.g., DBUA0002481.01.v01: up to 230 segments and 104 mm total length and 2.378 mm maximum width, including parapodia).
Discussion
Recently, a hidden biotechnological potential was uncovered in marine invertebrates, which might offer a wide array of natural products, showing properties compatible with anesthetics, fluorescent probes, and even antibiotics and pesticides (Rodrigo & Costa, 2019). By analyzing the phylogeny of toxin mixtures, Rodrigo et al. (2021a) show that annelids are uniquely positioned in the evolution of animal venoms. In particular, using the toxin-containing mucus present in the green Eulalia sp., which based on collection site (mainland Portugal) corresponds to E. clavigera s.s. in our study, revealed possible applications in anti-cancer therapeutics (Rodrigo et al., 2021b) and fluorescent probes for biotechnological applications using a protein mixture from the mucus (Rodrigo, 2020). This highlights the importance of formally describing cryptic complexes, since biochemical features might be unique to each lineage and can have a range of distinct effects and applications.
Molecular tools allowed us to unravel the hidden diversity within Eulalia, revealing compelling evidence on the existence of eight European MOTUs within the E. clavigera and E. viridis pseudo-cryptic complex. Combining molecular, morphometric, coloration, and geographical approaches, we concluded that three of these lineages merit being described as new species. Mean mtCOI-5P distances (17.9%) between lineages fit the annelid species distinction range (Nygren et al., 2018; Ravara et al., 2017; Sampieri et al., 2021), including phyllodocids (e.g. between 22 lineages belonging to the Eumida sanguinea (Örsted, 1843) species complex, Teixeira et al., 2022), and the MOTU delineation was congruent among all the methods employed.
We also found a clear geographic structure for most of these European MOTUs. Eulalia viridis (MOTU 7) is unique to the Scandinavia and Northern Sea and seems to be a northern boreal and sub-arctic species, occurring both inter-tidally and sub-tidally waters, in agreement with previous works (Bonse et al., 1996; Kato et al., 2001). Eulalia clavigera s. s. (MOTU 4) is a temperate species mostly found in intertidal rocky shores, occurring from Great Britain to the Western Mediterranean, as well as in the Azores, Savage, and Canary archipelagos. Its presence was also confirmed in Argentina (Langeneck et al., 2019). This species seems to be one of the most dominant taxa in Tenerife rocky beaches, including heavily human populated artificial pools in tourist zones, while morphologically similar individuals (lacking molecular data) have been reported from Brazil (Langeneck et al., 2019), with the specimens of E. viridis from southern Brazil (Morgado & Amaral, 1983) might actually belong to E. clavigera as well.
New continental species
Eulalia clavigera s. s. was considered a Mediterranean relict (Langeneck et al., 2019), while most Mediterranean shallow-water green Eulalia probably belong to either one or several different species. At least four different MOTUs seem to be exclusive to the Mediterranean Sea (Fig. 3a, b), a known biodiversity hotspot (Bianchi & Morri, 2000) for cryptic (Calvo et al., 2009; Langeneck et al., 2020; Taboada et al., 2017) and exotic (Galil, 2009; Zenetos et al., 2008) species. The alternating glacial and inter-glacial periods has been often suggested as one of the reasons explaining the high number of Mediterranean species. Under interglacial, the Mediterranean had a warm and arid climate leading to a deficient water balance, with the entrance of Atlantic surface waters through the Strait of Gibraltar playing a key role. This possibly allowed the introduction and maintenance of (sub)tropical littoral biota (Bianchi et al., 2012), while the North East Atlantic boreal species found refugia in the Mediterranean during glacial periods (Gómez & Lunt., 2007; Maggs et al., 2008; Schmitt et al., 2021). The survival of part of this fauna despite the different environmental (including water temperature) and depth conditions over time sustains the Mediterranean “biodiversity pump” hypothesis, as a possible outcome of the climatic events of the Quaternary (Bianchi & Morri, 2000).
Eulalia feliciae sp. nov. (MOTU 1) seems to be sympatric with E. clavigera s. s. (MOTU 4) and E. xanthomucosa sp. nov. (MOTU 5) in the Western Mediterranean. Together with a specimen of E. clavigera s.s. reported in Langeneck et al. (2019), these three species were collected inter-tidally in Banyuls-sur-Mer. However, as far as we know, E. xanthomucosa sp. nov. seems to be more abundant sub-tidally (mainly in recreational marinas), also occurs in Great Britain, and is characteristically yellowish instead of greenish, being thus similar to E. aurea coloration-wise. Live coloration is one of the most important taxonomic features in this genus, as most species are similarly brownish when preserved and almost impossible to distinguish based on morphological features (Schimmenti et al., 2016). Eulalia xanthomucosa sp. nov. was indeed the most divergent species of the complex and, besides coloration, it differs from E. clavigera s.s. morphologically in having longer dorsal and ventral cirri relative to body length and width. These morphological differences appear to parallel the molecular divergence data, e.g., the interspecific nuclear genetic distances tripled (Table 3) compared to those distances found between MOTUs within the major “clavigera” clade (clade A, Fig. 3a). The specimens from this clade (with the exception of the population from Madeira) also shared the 28S haplotypes. However, this seems to be common in other closely related marine species (Borges et al., 2012; Vieira et al., 2019). The ancestral central haplotype in the 28S network (Fig. 4b) might suggest the possibility of vicariance-driven speciation through a single colonization event and subsequent diversification (Meißner et al., 2014). The subtidal Eulalia IT2-1 (MOTU 8) also has a particularly high inter-specific distance in the nuclear markers, compared to the remaining complex, mirroring the values found for the outgroup (Phyllodoce sp.). MOTU 8 belongs to a very small specimen apparently fitting with the morphotype of E. viridis (i.e., pointed mid-body dorsal cirri, bright large red eyes). However, it is molecularly highly divergent, showing evidence of an entirely new, yet undescribed species of Eulalia outside the E. clavigera/viridis species complex.
The unnamed MOTU 3 from Croatia is genetically close to E. clavigera s.s. (mtCOI-5P, 7.5%; ITS, 4.8%; no 28S variation), suggesting a recent speciation, unlikely driven by the Messinian salinity crisis (from 6 to 5.33 MY, e.g., Hupało et al., 2019). Instead, this might be explained by environmental driven selection promoting local adaptation (Peijnenburg et al., 2004). The small-size and type locality of MOTU 3 agree with Eulalia virens Ehlers, 1864, a junior synonym of E. viridis originally described for the Adriatic Sea (Read and Fauchald, 2022). Eulalia virens is mainly characterized by its small size (54 segments, 7 mm in length, and 0.5 mm in width), but further sampling is required to allow elucidating if both designations belong or not to the same morphotype/species.
Additional un-sampled European MOTUs of Eulalia might still be uncovered. For example, the species Eulalia (Eumida) microceros Claparède, 1868, from the Gulf of Naples (synonymized to E. viridis) is characterized by its large size (5 cm long, 3 mm wide, and 300 segments). This far surpasses any of the analyzed green specimens of Eulalia from continental Europe in this study (Table 6, Table S2), suggesting it might be a large E. clavigera s.s. (based on the type locality and original description, PL. XVI, Fig. 4), or another large similar morphotype, different from those we have analyzed. Besides molecular data, morphometry, coloration and geographic restricted locations, reproductive features and gametogenesis could also contribute in future studies to discriminate closely related species, as seen in Sampieri et al. (2020), in which two cryptic Laeonereis Hartman, 1945 (family Nereididae) lineages from the West Atlantic coast were distinguished using both mtCOI-5P and histological data.
The Azores and Webbnesia archipelagos
Despite the high incidence of marine invertebrate endemisms in the Azorean ecoregion and Webbnesia (Desiderato et al., 2019; Vieira et al., 2019), no additional intertidal MOTUs were recorded in the Azores and Canary archipelagos. These volcanic islands were never in contact with the continent, were formed at different times, are hundreds of kilometers apart, possess a range of unique geological and climatic conditions, and their biota are the result of dispersal from distant geographical sources and in situ evolution and diversification (Fernández-Palacios et al., 2011). However, their intertidal populations of Eulalia do not differ substantially from the continental ones, with the exception of two partial morphometric markers and completely sorted mtCOI-5P and ITS haplotypes (Figs. 3b, 4a, and 5f). Conversely, a new species (E. madeirensis sp. nov., MOTU 2) was found in the subtidal populations from Madeira, which agrees with previous findings of cryptic lineages at different depths, e.g. three MOTUs in Phyllodoce madeirensis Langerhans, 1880 (Martin et al., 2021). Additional sampling efforts in Canarian and Azorean subtidal habitats may reveal new species of Eulalia. Intertidal populations of Eulalia from South Eastern Atlantic (Patagonia, Argentina) also failed to display any molecular or morphological divergence from the European E. clavigera s. s. (Langeneck et al., 2019). This may suggest a recent anthropogenically-mediated colonization for both the Canary and the South American populations. Indeed, neither E. clavigera nor E. viridis were recorded during the intensive surveys done in the 70’s, unlike the abundant populations observed recently in Puerto Madryn, Argentina (Orensanz J. M., personal communication in Langeneck et al., 2019). Furthermore, the first records of the E. clavigera in Canary date at least from 1976 (Sosa et al., 1976; Núñez et al., 2005; BDBC, https://www.biodiversidadcanarias.es/biota/?lang=en). However, the specimens from the Azores and Canary archipelagos do not share mtCOI-5P or ITS haplotypes with mainland Europe (Figs. 3b and 4a, respectively), suggesting an older non-anthropogenically driven colonization, instead. A recent unintentional introduction of E. clavigera s. s. by shipping activities, either with ballast waters or in fouling communities, has been suggested for the Patagonian populations by Schwindt et al. (2014), in a similar way as proved for small benthic marine fishes, chordates, invertebrates, and plankton, introduced either as eggs, larvae, or juveniles, and being first recorded from regions with major commercial ports and heavy international shipping (Cuesta et al., 2016; Lockett & Gomon, 2001; Wonham et al., 2000).
Conclusions
Besides E. viridis (restricted to Northern Europe, both in intertidal and subtidal areas) and E. clavigera s.s., we have found six rarer, locally restricted MOTUs within the Eulalia clavigera/viridis species complex, with an additional one from the Western Mediterranean reported in a previous study (MOTU GB1). Eulalia clavigera s. s. is instead widespread in Europe, being particularly abundant in temperate intertidal areas from the North East Atlantic (ranging from Portugal to the Great Britain), including the Azores and Webbnesia (i.e., Savage and Canary islands), but also with a small presence in the Western Mediterranean, usually in Marina environments. The close genetic proximity but lack of shared haplotypes between the continental and islandic populations of the latter species allows discarding a recent anthropogenic introduction through shipping. Moreover, three new species were described: Eulalia feliciae sp. nov., intertidal and unique to the Western Mediterranean, Eulalia madeirensis sp. nov., subtidal and unique to the Madeira Island (Portugal), and Eulalia xanthomucosa sp. nov., mostly subtidal, with an unique bright yellow coloration within the complex and occurring in the British Isles and southern France. Three unnamed lineages (MOTU GB1, and both the subtidal Eulalia KRO53 and Eulalia IS-BA) need additional specimens with good structural integrity to attempt a formal description, with the molecular and biogeographical data gathered in this work being a great starting point for future studies.
Availability of data and materials
New sequence data and specimen metadata were uploaded in the project “Eulalia species complex” (DS-MTE) within BOLD (https://v4.boldsystems.org/) and in the following link: https://doi.org/10.5883/DS-MTEC. The alignments (FASTA and NEXUS formats) for each marker (mtCOI-5P, ITS and 28S) and the concatenated one (mtCOI-5P + ITS + 28S) are all publicly available online at Figshare (DOI: https://doi.org/10.6084/m9.figshare.2165741). GenBank accession numbers: OP898309-OP898427 (mtCOI-5P), OP897856-OP897897 (ITS), and OP897898-OP897939 (28S)). See online supplemental Table S1 for more details. The new biological material is deposited at the Biological Research Collection (Marine Invertebrates) of the Department of Biology of the University of Aveiro (CoBI at DBUA), Portugal. The specimen from Banyuls (France) belonging to E. xanthomucosa sp. nov was donated to SCRIPPS Oceanography Institution (SIO), one specimen from the Savage islands was kindly loaned by the National Museum of Science and Natural History (MUHNAC, Portugal), while the two specimens from Corsica are deposited at the Muséum national d’Histoire naturelle (MNHN). All specimens available upon request, including the ones from Arne Nygren’s personal collection (ANPC, tagged with the BOLD Process ID: "MTE").
References
Alós, C. (2004). Familia Phyllodocidae Örsted, 1843. In Vieitez, J. M., Alós, C., Parapar, J., Besteiro, C., Moreira, J., Nüñez, J., Laborda, J., & San Martín, G. (Eds.), Fauna Iberica 25 (pp. 105–209). Madrid: Annelida Polychaeta I. Museo Nacional de Ciencias Naturales. CSIC.
Audouin, J. V., & Milne-Edwards, H. (1833). Classification des Annélides et description de celles qui ornalt les côtes de la France. Annales des sciences naturelles, Paris. (series 1), 29, 195–269.
Barfuss, M. H. (2012). Molecular studies in Bromeliaceae: implications of plastid and nuclear DNA markers for phylogeny, biogeography, and character evolution with emphasis on a new classification of Tillandsioideae. Vienna: University of Vienna, 244. Available at: http://othes.univie.ac.at/24037
Bely, A. E., & Wray, G. A. (2004). Molecular phylogeny of naidid worms (Annelida: Clitellata) based on cytochrome oxidase I. Molecular Phylogenetic and Evolution, 30(1), 50–63. https://doi.org/10.1016/S1055-7903(03)00180-5
Bianchi, C. N., & Morri, C. (2000). Marine biodiversity of the Mediterranean Sea: Situation, problems and prospects for future research. Marine Pollution Bulletin, 40(5), 367–376. https://doi.org/10.1016/S0025-326X(00)00027-8
Bianchi, C. N., Morri, C., Chiantore, M., Montefalcone, M., Parravicini, V., & Rovere, A. (2012). Mediterranean Sea biodiversity between the legacy from the past and a future of change. In N. Stambler (Ed.), Life in the Mediterranean Sea: A look at habitat changes (pp. 1–55). Nova Science Publishers Inc.
Bleidorn, C., Kruse, I., Albrecht, S., & Bartolomaeus, T. (2006). Mitochondrial sequence data expose the putative cosmopolitan polychaete Scoloplos armiger (Annelida, Orbiniidae) as a species complex. BMC Evolutionary Biology, 6(1), 47. https://doi.org/10.1186/1471-2148-6-47
Bonse, S., Schmidt, H., Eibye-Jacobsen, D., & Westheide, W. (1996). Eulalia viridis (Polychaeta: Phyllodocidae) is a complex of two species in northern Europe: Results from biochemical and morphological analysis. Cahiers De Biologie Marine, 37, 33–48.
Borges, L. M. S., Sivrikaya, H., le Roux, A., Shipway, J. R., Cragg, S. M., & Costa, F. O. (2012). Investigating the taxonomy and systematics of marine wood borers (Bivalvia : Teredinidae) combining evidence from morphology, DNA barcodes and nuclear locus sequences. Invertebrate Systematics, 26(6), 572–582. https://doi.org/10.1071/IS12028
Bouckaert, R., Heled, J., Kühnert, D., Vaughan, T., Wu, C. H., Xie, D., Suchard, M. A., Rambaut, A., & Drummond, A. J. (2014). BEAST 2: A software platform for Bayesian evolutionary analysis. PLOS Computational Biology, 10(4), e1003537. https://doi.org/10.1371/journal.pcbi.1003537
Calvo, M., Templado, J., Oliverio, M., & Machardom, A. (2009). Hidden Mediterranean biodiversity: Molecular evidence for a cryptic species complex within the reef building vermetid gastropod Dendropoma petraeum (Mollusca: Caenogastropoda). Biological Journal of the Linnean Society, 96(4), 898–912. https://doi.org/10.1111/j.1095-8312.2008.01167.x
Carr, C. M., Hardy, S. M., Brown, T. M., Macdonald, T. A., & Hebert, P. D. N. (2011). A tri-oceanic perspective: DNA barcoding reveals geographic structure and cryptic diversity in Canadian polychaetes. PLoS ONE, 6(7), e22232. https://doi.org/10.1371/ornal.pone.0022232
Castresana, J. (2000). Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Molecular Biology and Evolution, 17(4), 540–552. https://doi.org/10.1093/oxfordjournals.molbev.a026334
Çinar, M. E., & Gӧnlügür-Demirci, G. (2005). Polychaete assemblages on shallow-water benthic habitats along the Sinop Peninsula (Black Sea, Turkey). Cahiers De Biologie Marine, 46, 253–263.
Claparède, É. (1868). Les annélides chétopodes du Golfe de Naples. Mémoires De La Société De Physique Et D’histoire Naturelle De Genève, 19(2), 313–584.
Clement, M., Snell, Q., Walke, P., Posada, D., & Crandall, K. (2002). TCS: estimating gene genealogies. In Proceedings 16th International Parallel and Distributed Processing Symposium (pp. 7). Presented at the Proceedings 16th International Parallel and Distributed Processing Symposium. IPDPS 2002, Ft. Lauderdale, FL: IEEE. https://doi.org/10.1109/IPDPS.2002.1016585
Cuesta, J. A., Almón, B., Pérez-Dieste, J., Trigo, J. E., & Bañón, R. (2016). Role of ships’ hull fouling and tropicalization process on European carcinofauna: New records in Galician waters (NW Spain). Biological Invasions, 18(3), 619–630. https://doi.org/10.1007/s10530-015-1034-9
Darriba, D., Taboada, G. L., Doallo, R., & Posada, D. (2012). jModelTest 2: More models, new heuristics and parallel computing. Nature Methods, 9(8), 772–772. https://doi.org/10.1038/nmeth.2109
Delić, T., Trontelj, P., Rendoš, M., & Fišer, C. (2017). The importance of naming cryptic species and the conservation of endemic subterranean amphipods. Scientific Reports, 7(1), 3391. https://doi.org/10.1038/s41598-017-02938-z
Desiderato, A., Costa, F. O., Serejo, C. S., Abbiati, M., Queiroga, H., & Vieira, P. E. (2019). Macaronesian islands as promoters of diversification in amphipods: The remarkable case of the family Hyalidae (Crustacea, Amphipoda). Zoologica Scripta, 48(3), 359–375. https://doi.org/10.1111/zsc.12339
Ehlers, E. H. (1864). Die Borstenwürmer (Annelida Chaetopoda) nach systematischen und anatomischen Untersuchungen dargestellt. Retrieved February 13, 2022, from https://www.biodiversitylibrary.org/page/1985759
Eibye-Jacobsen, D. (1991). A revision of Eumida malmgren 1865 polychaeta phyllodocidae. Steenstrupia. Retrieved October 25, 2021, from https://eurekamag.com/research/006/964/006964164.php
Eibye-Jacobsen, D. (1993). On the phylogeny of the Phyllodocidae (Polychaeta Annelida): An alternative. Journal of Zoological Systematics and Evolutionary Research, 31(3), 174–197. https://doi.org/10.1111/j.1439-0469.1993.tb00188.x
Fernández-Palacios, J. M., de Nascimento, L., Otto, R., Delgado, J. D., García-del-Rey, E., Arévalo, J. R., & Whittaker, R. J. (2011). A reconstruction of Palaeo-Macaronesia, with particular reference to the long-term biogeography of the Atlantic island laurel forests. Journal of Biogeography, 38(2), 226–246. https://doi.org/10.1111/j.1365-2699.2010.02427.x
Fernandez-Triana, J. L. (2022). Turbo taxonomy approaches: Lessons from the past and recommendations for the future based on the experience with Braconidae (Hymenoptera) parasitoid wasps. ZooKeys, 1087, 199–220. https://doi.org/10.3897/zookeys.1087.76720
Fišer, C., Robinson, C. T., & Malard, F. (2018). Cryptic species as a window into the paradigm shift of the species concept. Molecular Ecology, 27(3), 613–635. https://doi.org/10.1111/mec.14486
Folmer, O., Black, M., Hoeh, W., Lutz, R., & Vrijenhoek, R. (1994). DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from metazoan invertebrates. Molecular Marine Biology and Biotechnology, 3, 294–299.
Fujisawa, T., & Barraclough, T. G. (2013). Delimiting species using single-locus data and the Generalized Mixed Yule Coalescent approach: A revised method and evaluation on simulated data sets. Systematic Biology, 62(5), 707–724. https://doi.org/10.1093/sysbio/syt033
Galil, B. S. (2009). Taking stock: Inventory of alien species in the Mediterranean Sea. Biological Invasions, 11(2), 359–372. https://doi.org/10.1007/s10530-008-9253-y
Grosse, M., Capa, M., & Bakken, T. (2021). Describing the hidden species diversity of Chaetozone (Annelida, Cirratulidae) in the Norwegian Sea using morphological and molecular diagnostics. ZooKeys, 1039, 139–176. https://doi.org/10.3897/zookeys.1039.61098
Gómez, A., & Lunt, D. H. (2007). Refugia within refugia: Patterns of phylogeographic concordance in the Iberian Peninsula. In S. Weiss & N. Ferrand (Eds.), Phylogeography of Southern European Refugia (pp. 155–188). Springer.
Guindon, S., & Gascuel, O. (2003). A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Systematic Biology, 52(5), 696–704. https://doi.org/10.1080/10635150390235520
Hadfield, M. G. (2011). Biofilms and marine invertebrate larvae: What bacteria produce that larvae use to choose settlement sites. Annual Review of Marine Science, 3, 453–470. https://doi.org/10.1146/annurev-marine-120709-142753
Hardy, S. M., Carr, C. M., Hardman, M., Steinke, D., Corstorphine, E., & Mah, C. (2011). Biodiversity and phylogeography of Arctic marine fauna: Insights from molecular tools. Marine Biodiversity, 41(1), 195–210. https://doi.org/10.1007/s12526-010-0056-x
Hassouna, N., Mithot, B., & Bachellerie, J. P. (1984). The complete nucleotide sequence of mouse 28S rRNA gene. Implications for the process of size increase of the large subunit rRNA in higher eukaryotes. Nucleic Acids Research, 12(8), 3563–3583. https://doi.org/10.1093/nar/12.8.3563
Hupało, K., Teixeira, M. A. L., Rewicz, T., Sezgin, M., Iannilli, V., Karaman, G. S., Grabowski, M., & Costa, F. O. (2019). Persistence of phylogeographic footprints helps to understand cryptic diversity detected in two marine amphipods widespread in the Mediterranean basin. Molecular Phylogenetics and Evolution, 132, 53–66. https://doi.org/10.1016/j.ympev.2018.11.013
Hutchings, P., & Kupriyanova, E. (2018). Cosmopolitan polychaetes – fact or fiction? Personal and Historical Perspectives. Invertebrate Systematics, 32(1), 1–9. https://doi.org/10.1071/IS17035
Jolly, M. T., Viard, F., Gentil, F., Thiébaut, E., & Jollivet, D. (2006). Comparative phylogeography of two coastal polychaete tubeworms in the Northeast Atlantic supports shared history and vicariant events. Molecular Ecology, 15(7), 1841–1855. https://doi.org/10.1111/j.1365-294X.2006.02910.x
Kato, T., Pleijel, F., & Mawatari, S. F. (2001). Eulalia gemina (Phyllodocidae: Polychaeta), a new species form Shirahama, Japan. Proceedings of the Biological Society of Washington, 114, 381–388.
Katoh, K., & Standley, D. M. (2013). MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Molecular Biology and Evolution, 30(4), 772–780. https://doi.org/10.1093/molbev/mst010
Kumar, S., Stecher, G., Li, M., Knyaz, C., & Tamura, K. (2018). MEGA X: Molecular evolutionary genetics analysis across computing platforms. Molecular Biology and Evolution, 35(6), 1547–1549. https://doi.org/10.1093/molbev/msy096
Langeneck, J., Diez, M. E., Nygren, A., Salazar-Vallejo, S., Carrera-Parra, L. F., Vega Fernández, T., Badalamenti, F., Castelli, A., & Musco, L. (2019). Worming its way into Patagonia: An integrative approach reveals the cryptic invasion by Eulalia clavigera (Annelida: Phyllodocidae). Marine Biodiversity, 49(2), 851–861. https://doi.org/10.1007/s12526-018-0864-y
Langeneck, J., Scarpa, F., Maltagliati, F., Sanna, D., Barbieri, M., Cossu, P., Mikac, B., Galletti, M. C., Castelli, A., & Casu, M. (2020). A complex species complex: The controversial role of ecology and biogeography in the evolutionary history of Syllis gracilis Grube, 1840 (Annelida, Syllidae). Journal of Zoological Systematics and Evolutionary Research, 58(1), 66–78. https://doi.org/10.1111/jzs.12336
Leigh, J. W., & Bryant, D. (2015). Popart: Full-feature software for haplotype network construction. Methods in Ecology and Evolution, 6(9), 1110–1116. https://doi.org/10.1111/2041-210X.12410
Leite, B. R., Vieira, P. E., Teixeira, M. A. L., Lobo-Arteaga, J., Hollatz, C., Borges, L. M. S., Duarte, S., Troncoso, J. S., & Costa, F. O. (2020). Gap-analysis and annotated reference library for supporting macroinvertebrate metabarcoding in Atlantic Iberia. Regional Studies in Marine Science, 36, 101307. https://doi.org/10.1016/j.rsma.2020.101307
Librado, P., & Rozas, J. (2009). DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics, 25(11), 1451–1452. https://doi.org/10.1093/bioinformatics/btp187
Linnaeus, C. V. (1767). Sistema naturae, 12th Edition. Stockholm. pp 1866.
Lobo, L., Ferreira, M. S., Antunes, I. C., Teixeira, M. A. L., Borges, L. M. S., Sousa, R., Gomes, P. A., Costa, M. H., Cunha, M. R., & Costa, F. O. (2017). Contrasting morphological and DNA barcode-suggested species boundaries among shallow-water amphipod fauna from the southern European Atlantic coast. Genome, 60(2), 147–157. https://doi.org/10.1139/gen-2016-0009
Lobo, J., Teixeira, M. A. L., Borges, L. M. S., Ferreira, M. S. G., Hollatz, C., Gomes, P. T., Sousa, R., Ravara, A., Costa, M. H., & Costa, F. O. (2016). Starting a DNA barcode reference library for shallow water polychaetes from the southern European Atlantic coast. Molecular Ecology Resources, 16(1), 298–313. https://doi.org/10.1111/1755-0998.12441
Lockett, M. M., & Gomon, M. F. (2001). Ship mediated fish invasions in Australia: Two new introductions and a consideration of two previous invasions. Biological Invasions, 3(2), 187–192. https://doi.org/10.1023/A:1014584201815
McIntosh, W. C. (1908). A monograph of British Annelids Part I. Polychaeta. Nephthydidae to Syllidae. Ray Society of London, II., 2, 1–232.
Maggs, C. A., Castilho, R., Foltz, D., Henzler, C., Jolly, M. T., Kelly, J., Olsen, J., Perez, K. E., Stam, W., Väinölä, R., Viard, F., & Wares, J. (2008). Evaluating signatures of glacial refugia for North Atlantic benthic marine taxa. Ecology, 89(11), S108–S122. https://doi.org/10.1890/08-0257.1
Martin, D., Aguado, M. T., Fernández Álamo, M. A., Britayev, T. A., Böggemann, M., Capa, M., Faulwetter, S., Fukuda, M. V., Helm, C., Petti, M. A. V., Ravara, A., & Teixeira, M. A. L. (2021). On the diversity of Phyllodocida (Annelida: Errantia), with a focus on Glyceridae, Goniadidae, Nephtyidae, Polynoidae, Sphaerodoridae, Syllidae, and the Holoplanktonic Families. Diversity, 13(3), 131. https://doi.org/10.3390/d13030131
Martin, D., Meca, M. A., Gil, J., Drake, P., & Nygren, A. (2017). Another brick in the wall: Population dynamics of a symbiotic species of Oxydromus (Annelida, Hesionidae), described as new based on morphometry. Contributions to Zoology, 86(3), 181–211. https://doi.org/10.1163/18759866-08603001
Meißner, K., Bick, A., Guggolz, T., & Gõtting, M. (2014). Spionidae (Polychaeta: Canalipalpata: Spionida) from seamounts in the NE Atlantic. Zootaxa, 3786(3), 201–245. https://doi.org/10.11646/zootaxa.3786.3.1
Meyer, A. (1938). Der Rogen und die Entwicklung der trochophora von Eulalia viridis. Biologia Generalis, 14, 334–389.
Miglietta, M. P., Faucci, A., & Santini, F. (2011). Speciation in the sea: Overview of the symposium and discussion of future directions. Integrative and Comparative Biology, 51(3), 449–455. https://doi.org/10.1093/icb/icr024
Morgado, E. H., & Amaral, A. C. Z. (1983). Anelídeos poliquetos associados ao briozoário Schizoporella unicornis (Johnston): IV. Phyllodocidae e Hesionidae. Revista Brasileira De Zoologia, 2, 49–54. https://doi.org/10.1590/S0101-81751983000200002
Morton, B. (2011). Predator–prey-scavenging interactions between Nucella lapillus, Carcinus maenas and Eulalia viridis all exploiting Mytilus galloprovincialis on a rocky shore recovering from tributyl-tin (TBT) pollution. Journal of Natural History, 45(39–40), 2397–2417. https://doi.org/10.1080/00222933.2011.596637
Nedved, B. T., & Hadfield, M. G. (2008). Hydroides elegans (Annelida: Polychaeta): A model for biofouling research. In H. C. Flemming, R. Venkatesan, S. P. Murthy, & K. E. Cooksey (Eds.), Marine and Industrial Biofouling (pp. 203–218). Springer.
Núñez, J., Brito, M. C., & Docoito, J. R. (2005). Annelid polychaetes from Canaries: Catalogue of species, distribution and habitats. Vieraea, 33, 297–332.
Nygren, A., Eklöf, J., & Pleijel, F. (2009). Arctic-boreal sibling species of Paranaitis (Polychaeta, Phyllodocidae). Marine Biology Research, 5(4), 315–327. https://doi.org/10.1080/17451000802441301
Nygren, A. (2014). Cryptic polychaete diversity: A review. Zoologica Scripta, 43(2), 172–183. https://doi.org/10.1111/zsc.12044
Nygren, A., Eklöf, J., & Pleijel, F. (2010). Cryptic species of Notophyllum (Polychaeta: Phyllodocidae) in Scandinavian waters. Organisms Diversity & Evolution, 10(3), 193–204. https://doi.org/10.1007/s13127-010-0014-2
Nygren, A., Parapar, J., Pons, J., Meißner, K., Bakken, T., Kongsrud, J. A., Oug, E., Gaeva, D., Sikorski, A., Johansen, R. A., Hutchings, P. A., Lavesque, N., & Capa, M. (2018). A mega-cryptic species complex hidden among one of the most common annelids in the North East Atlantic. PLoS ONE, 13(6), e0198356. https://doi.org/10.1371/journal.pone.0198356
Nygren, A., & Pleijel, F. (2011). From one to ten in a single stroke – resolving the European Eumida sanguinea (Phyllodocidae, Annelida) species complex. Molecular Phylogenetics and Evolution, 58(1), 132–141. https://doi.org/10.1016/j.ympev.2010.10.010
Olive, P. J. W. (1975). A vitellogenesis promoting influence of the prostomium in the polychaete Eulalia viridis (müller) (phyllodocidae). General and Comparative Endocrinology, 26(2), 266–273. https://doi.org/10.1016/0016-6480(75)90145-8
Peijnenburg, K. T. C. A., Breeuwer, J. A. J., Pierrot-Bults, A. C., & Menken, S. B. J. (2004). Phylogeography of the planktonic Chaetognath Sagitta setosa reveals isolation in European Seas. Evolution, 58(7), 1472–1487. https://doi.org/10.1111/j.0014-3820.2004.tb01728.x
Pleijel, F. (1993). Polychaeta Phyllodocidae. Marine Invertebrates of the Scandinavia, 8, 1–159.
Puillandre, N., Lambert, A., Brouillet, S., & Achaz, G. (2012). ABGD, Automatic Barcode Gap Discovery for primary species delimitation. Molecular Ecology, 21(8), 1864–1877. https://doi.org/10.1111/j.1365-294X.2011.05239.x
Rambaut, A., Drummond, A. J., Xie, D., Baele, G., & Suchard, M. A. (2018). Posterior summarization in Bayesian phylogenetics using tracer 1.7. Systematic Biology, 67(5), 901–904. https://doi.org/10.1093/sysbio/syy032
Ratnasingham, S., & Hebert, P. D. N. (2013). A DNA-based registry for all animal species: The Barcode Index Number (BIN) system. PLoS ONE, 8(7), e66213. https://doi.org/10.1371/journal.pone.0066213
Ravara, A., Cunha, M. R., Pleijel, F. (2010). Nephtyidae (Annelida, Polychaeta) from southern Europe. Zootaxa, 2682, 1–68. https://doi.org/10.11646/zootaxa.2682.1.1
Ravara, A., Ramos, D., Teixeira, M. A. L., Costa, F. O., & Cunha, M. R. (2017). Taxonomy, distribution and ecology of the order Phyllodocida (Annelida, Polychaeta) in deep-sea habitats around the Iberian margin. Deep Sea Research Part II: Topical Studies in Oceanography, 137, 207–231. https://doi.org/10.1016/j.dsr2.2016.08.008
Read, G., & Fauchald, K. (Ed) (2022). World Polychaeta Database. Eulalia virens Ehlers, 1864. Retrieved January 10, 2022, from World Register of Marine Species at: https://www.marinespecies.org/aphia.php?p=taxdetails&id=339528
Rodrigo, A. P., Costa, M. H., de Matos, A. P. A., Carrapiço, F., & Costa, P. M. (2015). A Study on the digestive physiology of a marine polychaete (Eulalia viridis) through microanatomical changes of epithelia during the digestive cycle. Microscopy and Microanalysis, 21(1), 91–101. https://doi.org/10.1017/S143192761401352X
Rodrigo, A. P., & Costa, P. M. (2019). The hidden biotechnological potential of marine invertebrates: The polychaeta case study. Environmental Research, 173, 270–280. https://doi.org/10.1016/j.envres.2019.03.048
Rodrigo A. P. (2020). The biotechnological value of a novel potent marine biotoxin from the polychaete worm Eulalia viridis: chemical and toxicological evaluation. Universidade Nova Lisboa. PhD thesis. Retrieved January 10, 2022, from https://hdl.handle.net/10362/116178
Rodrigo, A. P., Grosso, A. R., Baptista, P. V., Fernandes, A. R., & Costa, P. M. (2021a). A transcriptomic approach to the recruitment of venom proteins in a marine annelid. Toxins, 13(2), 97. https://doi.org/10.3390/toxins13020097
Rodrigo, A. P., Mendes, V. M., Manadas, B., Grosso, A. R., Alves de Matos, A. P., Baptista, P. V., Costa, P. M., & Fernandes, A. R. (2021b). Specific antiproliferative properties of proteinaceous toxin secretions from the marine annelid Eulalia sp. onto ovarian cancer cells. Marine Drugs, 19(1), 31. https://doi.org/10.3390/md19010031
Ronquist, F., & Huelsenbeck, J. P. (2003). MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics, 19(12), 1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Sampieri, B. R., Steiner, T. M., Baroni, P. C., da Silva, C. F., Teixeira, M. A. L., Vieira, P. E., Costa, F. O., & Amaral, A. C. Z. (2020). How oogenesis analysis combined with DNA barcode can help to elucidate taxonomic ambiguities: a polychaete study-based approach. Biota Neotropica, 20. https://doi.org/10.1590/1676-0611-BN-2020-0959
Sampieri, B. R., Vieira, P. E., Teixeira, M. A. L., Seixas, V. C., Pagliosa, P. R., Amaral, A. C. Z., & Costa, F. O. (2021). Molecular diversity within the genus Laeonereis (Annelida, Nereididae) along the west Atlantic coast: Paving the way for integrative taxonomy. PeerJ, 9, e11364. https://doi.org/10.7717/peerj.11364
Schimmenti, E., Musco, L., Brutto, S. L., Mikac, B., Nygren, A., & Badalamenti, F. (2016). A Mediterranean record of Eulalia ornata (Annelida: Phyllodocidae) corroborating its fidelity link with the Sabellaria alveolata-reef habitat. Mediterranean Marine Science, 17(2), 359–370. https://doi.org/10.12681/mms.1485
Schmitt, T., Fritz, U., Delfino, M., Ulrich, W., & Habel, J. C. (2021). Biogeography of Italy revisited: Genetic lineages confirm major phylogeographic patterns and a pre-Pleistocene origin of its biota. Frontiers in Zoology, 18(1), 34. https://doi.org/10.1186/s12983-021-00418-9
Schwindt, E., Gappa, J. L., Raffo, M. P., Tatián, M., Bortolus, A., Orensanz, J. M., Alonso, G., Diez, M. E., Doti, B., Genzano, G., Lagger, C., Lovrich, G., Piriz, M. L., Mendeza, M. M., Savoya, V., & Sueiro, M. C. (2014). Marine fouling invasions in ports of Patagonia (Argentina) with implications for legislation and monitoring programs. Marine Environmental Research, 99, 60–68. https://doi.org/10.1016/j.marenvres.2014.06.006
Sosa, A., Núñez, J., & Bacallado, J.J. (1976). Contribución al estudio de los poliquetosen Canarias. I: Aphroditidae, Amphinomidae, Phyllocidae y Eunicidae. Vieraea, 6(2), 231–252
Taboada, S., Leiva, C., Bas, M., Schult, N., & McHugh, D. (2017). Cryptic species and colonization processes in Ophryotrocha (Annelida, Dorvilleidae) inhabiting vertebrate remains in the shallow-water Mediterranean. Zoologica Scripta, 46(5), 611–624. https://doi.org/10.1111/zsc.12239
Teixeira, M. A. L., Vieira, P. E., Pleijel, F., Sampieri, B. R., Ravara, A., Costa, F. O., & Nygren, A. (2020). Molecular and morphometric analyses identify new lineages within a large Eumida (Annelida) species complex. Zoologica Scripta, 49(2), 222–235. https://doi.org/10.1111/zsc.12397
Teixeira, M. A. L., Vieira, P. E., Ravara, A., Costa, F. O., & Nygren, A. (2022). From 13 to 22 in a second stroke: Revisiting the European Eumida sanguinea (Phyllodocidae: Annelida) species complex. Zoological Journal of the Linnean Society, 196(1), 169–197.
Ushakov, P. L. (1972). Polychaetes of the sub-order Phyllodociforma of the Polar Basin and the North Western part of the Pacific. In Russian. Translated by the Israel Program for Scientific Translations: Jerusalem, 1974. Fauna SSSR, 102, 1–271.
Vieira, P. E., Desiderato, A., Holdich, D. M., Soares, P., Creer, S., Carvalho, G. R., Costa, F. O., & Queiroga, H. (2019). Deep segregation in the open ocean: Macaronesia as an evolutionary hotspot for low dispersal marine invertebrates. Molecular Ecology, 28(7), 1784–1800. https://doi.org/10.1111/mec.15052
Viéitez, J. M., Alós, C., Parapar, J., Besteiro, C., Moreira, J., Núñez, J., Laborda, A. J. & San Martín, G. (2004). Annelida Polychaeta I. In Ramos, M. A. et al. (Eds). Fauna Ibérica, Vol. 25 (530 pp). Museo Nacional de Ciencias Naturales CSIC, Madrid
Wonham, M. J., Carlton, J. T., Ruiz, G. M., & Smith, L. D. (2000). Fish and ships: Relating dispersal frequency to success in biological invasions. Marine Biology, 136(6), 1111–1121. https://doi.org/10.1007/s002270000303
Zenetos, A., Meric, E., Verlaque, M., Galli, P., Boudouresque, C. F., Giangrande, A., Cinar, M., & Bilecenoglu, M. (2008). Additions to the annotated list of marine alien biota in the Mediterranean with special emphasis on Foraminifera and Parasites. Mediterranean Marine Science, 9(1), 119–166. https://doi.org/10.12681/mms.146
Zhang, J., Kapli, P., Pavlidis, P., & Stamatakis, A. (2013). A general species delimitation method with applications to phylogenetic placements. Bioinformatics, 29(22), 2869–2876. https://doi.org/10.1093/bioinformatics/btt499
Acknowledgements
The authors wish to thank Sofia Duarte for the Portuguese Eulalia specimens, Ana Costa for the specimens from the Azores, Nicolas Lavesque and Felicia Ulltin for the northern and southern French samples, respectively and Julio Parapar for the northern Spanish specimens, Additionally, to Jorge Núñez and Beatriz Alfonso for all the assistance and knowledge provided during the Canary islands sampling campaign. Moreover, we would like to thank the reviewer for taking the time reviewing and improving upon the original version of our manuscript.
Thanks are due to the Corsica program. The CORSICABENTHOS expeditions (PI: Line Le Gall), with a focus on the small benthic biota, are the marine component of the “Our Planet Reviewed” program. The Corsica program is run by Muséum National d’Histoire Naturelle in partnership with Université de Corse Pasquale Paoli and Office de l’Environnement de la Corse (OEC), with the support of Office Français de la Biodiversité (OFB) and Collectivité Territoriale de Corse (CTC). CORSICABENTHOS 2 took place in October 2020 in collaboration with Réserve Naturelle des Bouches de Bonifacio. The organizers are grateful to Medeleine Cancemi, Jean-François Cubells, Jean-Michel Culioli, and Jean-Michel Palazzi for their support.
Funding
This study was supported by the project ATLANTIDA–Platform for the monitoring of the North Atlantic Ocean and tools for the sustainable exploitation of the marine resources, with the reference NORTE-01–0145-FEDER-000040, co-financed by the European Regional Development Fund (ERDF), through Programa Operacional Regional do Norte (NORTE 2020). Thanks are due, for the financial support to CESAM (UIDB/50017/2020 + UIDP/50017/2020), to Portuguese Foundation for Science and Technology and Ministry of Education and Science (FCT/MEC) through national funds, and the co-funding by the FEDER, within the PT2020 Partnership Agreement and Compete 2020. Marcos A. L. Teixeira was supported by a PhD grant from FCT co-financed by ESF (SFRH/BD/131527/2017) and from the DNAqua-Net STSM grant “Rich and hidden biodiversity not yet barcoded in the Canary archipelago (Spain) as an opportunity to enrich the DNA barcode reference library for European polychaetes,” under the EU Cost action CA15219–Developing new genetic tools for bio-assessment of aquatic ecosystems in Europe. Pedro E. Vieira was supported by national funds through the Portuguese Foundation for Science and Technology (FCT, I.P.) in the scope of the project (early detection and monitoring of non-indigenous species in coastal ecosystems based on high-throughput sequencing tools, PTDC/BIA-BMA/29754/2017). Ascensão Ravara was supported by national funds, through FCT, I.P., in the scope of the framework contract foreseen in the numbers 4, 5, and 6 of the article 23, of the Decree-Law 57/2016, of August 29, changed by Law 57/2017, of July 19. Arne Nygren was supported by the Norwegian Taxonomy Initiative [https://www.biodiversity.no/Pages/135523] (Cryptic polychaete species in Norwegian waters, knr 49–13, pnr 70184228), the Swedish Taxonomy Initiative [https://www.artdatabanken.se/en/the-swedish-taxonomy-initiative/] (Polychaete species complexes in Swedish waters, dnr 140/07 1.4 and 166/08 1.4), and Kungliga Fysiografiska sällskapet Nilsson-Ehle donationerna [https://www.fysiografen.se/sv/].
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation and data collection were performed by Marcos A. L. Teixeira, Pedro E. Vieira, Joachim Langeneck, José Carlos Hernández, David Fenwick, Fredrik Pleijel, Ascensão Ravara, and Arne Nygren. Analyses were performed by Marcos A. L. Teixeira. The first draft of the manuscript was written by Marcos A. L. Teixeira, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no completing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
13127_2022_597_MOESM1_ESM.xlsx
Supplementary file1: Voucher data, origin of the specimens and GenBank accession numbers for each of the analyzed genetic markers original to this study and molecular metadata used for comparison purposes or as outgroups. (XLSX 21 KB)
13127_2022_597_MOESM2_ESM.xlsx
Supplementary file2: Measurements for all the specimens used in morphometry belonging to Eulalia clavigera with populations from the Macaronesia islands and mainland Europe, Eulalia feliciae sp. nov., Eulalia madeirensis sp. nov. and Eulalia xanthomucosa sp. nov. (XLSX 50 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Teixeira, M.A.L., Vieira, P.E., Fenwick, D. et al. Revealing the diversity of the green Eulalia (Annelida, Phyllodocidae) species complex along the European coast, with description of three new species. Org Divers Evol 23, 477–503 (2023). https://doi.org/10.1007/s13127-022-00597-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s13127-022-00597-1