Abstract
Metabolic Dysfunction-Associated Steatotic Liver Disease (MASLD) is a worldwide leading cause of liver-related associated morbidities and mortality. Currently, there is a lack of reliable non-invasive biomarkers for an accurate of MASLD. Hence, this study aimed to evidence the functional role of miRNAs as potential biomarkers for MASLD assessment. Data from 55 participants with steatosis (MASLD group) and 45 without steatosis (control group) from the Fatty Liver in Obesity (FLiO) Study (NCT03183193) were analyzed. Anthropometrics and body composition, biochemical and inflammatory markers, lifestyle factors and liver status were evaluated. Circulating miRNA levels were measured by RT-PCR. Circulating levels of miR-122-5p, miR-151a-3p, miR-126-5p and miR-21-5p were significantly increased in the MASLD group. These miRNAs were significantly associated with steatosis, liver stiffness and hepatic fat content. Logistic regression analyses revealed that miR-151a-3p or miR-21-5p in combination with leptin showed a significant diagnostic accuracy for liver stiffness obtaining an area under the curve (AUC) of 0.76 as well as miR-151a-3p in combination with glucose for hepatic fat content an AUC of 0.81. The best predictor value for steatosis was obtained by combining miR-126-5p with leptin, presenting an AUC of 0.95. Circulating miRNAs could be used as a non-invasive biomarkers for evaluating steatosis, liver stiffness and hepatic fat content, which are crucial in determining MASLD.
Clinical trial registration
• Trial registration number: NCT03183193 (www.clinicaltrials.gov).
• Date of registration: 12/06/2017.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
One of the leading causes of liver-related associated morbidities and mortality is non-alcoholic fatty liver disease (NAFLD), which is characterized by an excess storage of triglycerides (TG) and fatty acids in the hepatocytes [55]. In recent years, the incidence of this disease has increasing, reaching a global prevalence about 30% [63]. This disease encompasses a wide range of histopathology that goes from simple hepatic steatosis to non-alcoholic steatohepatitis (NASH), and can reach into fibrosis, cirrhosis and cellular hepatocarcinoma (HCC) [55].
In June 2023, a multi-society Delphi consensus statement introduced a new terminology, replacing NAFLD and Metabolic Dysfunction Associated Fatty Liver Disease (MAFLD) with Metabolic Dysfunction-Associated Steatotic Liver Disease (MASLD). Similarly, NASH was replaced with Metabolic Dysfunction-Associated Steatohepatitis (MASH). These new terms encompass the presence of at least one to five cardiometabolic risk factors in their diagnostic criteria [51]. Even with the revised definition, about 99% of patients previously diagnosed with NAFLD meet the diagnostic criteria for MASLD [28].
Due to interactions among genetic, metabolic, and environmental factors, there is an interindividual variation in the MASLD phenotype and its progression, which currently remain unclear [66]. MASLD is strongly associated with chronic liver disease, cardiovascular disease (CVD), type 2 diabetes mellitus (T2DM), obesity, insulin resistance (IR), hypertension, hyperlipidemia, and metabolic syndrome [55, 62].
Liver biopsy continues to be the reference technique for the diagnosis and prognosis of the disease. However, it carries a risk for the patient because it is an invasive procedure, and it can cause complications, as well as errors in the sample [30]. Imaging modalities such as ultrasonography (US), computed tomography (CT), and magnetic resonance imaging (MRI) provide non-invasive options for detecting MASLD. However, these techniques are frequently associated with significant time consumption and high costs [57]. There is a need for improved non-invasive tools that facilitate the diagnosis and staging of MASLD, along with the development of methods to identify those at risk of disease progression [30].
MASLD involves many signaling molecules contributing to hepatic metabolism, inflammatory, oxidative, and fibrotic processes, making it quite complex [45]. These include microRNAs (miRNAs), which are small non-coding RNAs that play important roles in the regulatory processes of gene expression and target numerous genes involved in the glucose and lipid metabolism, inflammation, cell proliferation, apoptosis and necrosis, in turn involved in the pathogenesis of MASLD [10, 11].
Although it is not fully understanding the exact molecular drivers and biological pathways responsible for disease progression, genetic variations likely play a role in the complexity and differences observed in the disease phenotype [48]. Alteration in circulating miRNAs play an important role in hepatocyte function, liver injury, viral hepatitis, ALD, MASLD, liver fibrosis progression, and HCC [43]. Moreover, miRNAs regulate multiple signaling pathways involved in lipid accumulation, IR, oxidative stress, and inflammatory responses, thereby contributing to the development and advancement of MASLD [37]. MiR-122, a key liver-specific miRNA, plays a role regulating liver metabolism and maintaining fatty acid balance, previously correlated with steatosis severity in MASLD [5], playing an important role in MASLD pathogenesis as well as miR-21 and miR-192 [60]. In addition, miR-126-5p could impact on pathogenesis of liver fibrosis and it is associated with T2DM, obesity, metabolic syndrome [4], adiposity, lipid and glucose metabolism and inflammation as well as miR-222 [8]. Additionally, miR-151 which is linked to TNF-α [40], and miR-15b-3p and miR-29b-3p, which are especially related with high expression in hepatic stellate cells, have been associated with liver tumor development, suggesting their involvement in hepatic lesions [31, 60].
Despite some associations between those miRNAs and MASLD, a comprehensive understanding of the complex involvement of miRNAs in the disease is needed. In this context, the aim of the present study was examining the functional role of miRNAs as hepatic status biomarkers using simple samples such as serum for the diagnosis and management of MASLD.
Material and methods
Study participants
This research is a cross-sectional analysis that include the evaluation of baseline measurements of 55 subjects with overweight or obesity (BMI ≥ 27.5 kg/m2 < 40 kg/m2) with MASLD (diagnosed by ultrasonography) obtained from a randomized controlled trial, the FLiO (Fatty liver in obesity) study (www.clinicaltrials.gov; NCT03183193), and 45 subjects with normal-weight (BMI < 25 kg/m2) without MASLD (confirmed by ultrasonography) obtained from the EHGNA study (a FLiO study continuation). From them, miRNAs circulating levels have been obtained. The exclusion criteria have already been described previously [42].
The study protocol was approved by the Research Ethics Committee of the University of Navarra (ref. 54/2015). Everyone gave written informed consent before enrolling in the study. All procedures were performed in compliance with significant national regulations, institutional policies, and in accordance with the Declaration of Helsinki and following the CONSORT 2010 guidelines.
Anthropometric and body composition evaluation
Anthropometric measurements (BW and waist circumference) and body composition (Lunar iDXA, enCORE 14.5, Madison, WI) were determined under fasting conditions in the Nutritional Intervention Unit of the University of Navarra, according to standardized procedures [14]. Body Mass Index (BMI) was calculated as BW divided by height squared (Kg/m2).
Biochemical determinations
Blood samples were collected, processed and stored at -80 degrees, until further analysis [18]. Biochemical determinations such as blood glucose, glycosylated hemoglobin (HbA1c), triglycerides (TG), total cholesterol (TC), high-density lipoprotein cholesterol (HDL-c), alanine aminotransferase (ALT), aspartate aminotransferase (AST) and gamma glutamyl transferase (GGT) were measured in a Pentra C-200 autoanalyzer (HORIBA ABX, Madrid, Spain) Spain) with specific commercial kits and following the manufacturer’s instructions (Cobas 8000, Roche Diagnostics, Switzerland). Low-density lipoprotein (LDL-c) levels were calculated using the Friedewald formula [25]: LDL-c = TC—HDL-c—TG/5. The Castelli’s Risk Index (CRI) was calculated with the formula: Total cholesterol/HDL-c, as previously described [23]. The fatty liver index (FLI) was calculated using serum TG, BMI, waist circumference, and GGT concentrations, as mentioned [6]. FLI values < 30 rule out hepatic steatosis, and values ≥ 60 indicate hepatic steatosis. Concentrations of insulin, leptin, chemerin, retinol-binding protein (RBP4) and adiponectin were measured using ELISA kits (Demeditec; Kiel-Wellsee, Germany) in a Triturus autoanalyzer (Grifols, Barcelona, Spain). Leukocyte cell-derived chemotaxin-2 (LECT2) was analyzed using the Triturus Autoanalyzer (Grifols, Barcelona, Spain) with specific kits (Biovendor LLC, North Carolina, United States). Insulin resistance was estimated through the Homeostasis Model Assessment Index (HOMA-IR), which was calculated using a previously described formula [14].
Imaging techniques in the evaluation of liver status
The liver status assessment was carried out under fasting conditions at the University of Navarra Clinic by highly qualified personnel. To determine the presence of hepatic steatosis, ultrasonography was performed (Siemens ACUSON S2000 and S3000), according to the methodology already described [9]. To determine liver fat content and hepatic volume, magnetic resonance was used (Siemens Aera 1.5 T, Erlangen Germany), applying the DIXON technique [9]. Multi-echo T2 corrected single breath-hold spectroscopy (Histo) of a single voxel and multi-echo 3D gradient echo imaging with Dixon reconstruction and T2 correction were included by quantitative sequences [18]. ARFI elastography was performed to determine liver stiffness, using the value obtained from 10 valid ARFI measurements of each subject as the mean value [9].
Dietary intervention and lifestyle
Dietary intake was assessed with a semiquantitative food frequency questionnaire (FFQ) with 173 items, validated in Spain for daily energy and nutrient intake [22]. Daily food consumption was estimated by multiplying the portion size by the consumption frequency and dividing as described elsewhere [46]. The nutrient composition of the specified serving size for each food was estimated using Spanish food composition tables [46]. Adherence to the Mediterranean diet (MedDiet) was assessed with a 17-item screening questionnaire, with a final score ranging from 0 to 17, with a higher score indicating better MedDiet adherence [26]. Physical activity was estimated using a validated Spanish version of Minnesota leisure-Time Physical Activity Questionnaire [42]. The physical activity was classified into four different categories (sedentary, light, moderate or vigorous) based on the International Physical Activity Questionnaire (IPAQ) [41].
RNA isolation, reverse transcription and Real-Time PCR (RT-PCR)
Serum was isolated from whole blood by centrifugation at 1100 g at 4 °C for 15 min (Modelo5415R, Eppendorf AG, Hamburgo, Germany) and then, samples were frozen (-80 ºC) until RNA reverse transcription. After RNA extraction, total RNA of the serum sample was isolated with RNeasy Serum/Plasma Advanced Kit (Qiagen, Hilden, Germany) following manufacturer’s instructions. For the procedure, absolute ethanol (1L) and Isopropanol (2-Propanol, gradient HPLC grade) were used. cDNA was synthesized using 4 μl of miRNA sample, miRCURY LNA RT Kit (Qiagen, Hilden, Germany), which allowed the detection of the miRNAs of interest. Once the reverse transcription was realized, the protocol for the expression of miRNA isolated from serum was carried out. Quantitative PCR (qPCR) was performed with the CFX384 Touch Real-Time PCR system (Bio-Rad, Hercules, CA, USA) using the miRCURY SYBR® Green PCR Kit (Qiagen, Hilden, Germany). Finally, miRNAs Relative Quantities (RQs) were calculated with the formula 2−ΔCt. Normalization factor (NF) was calculated using the geometric mean of RQs of all expressed miRNAs per sample and the normalized relative quantities were obtained dividing RQs by the sample specific NF as previously described [44]. The values were expressed as fold change (FC) of each miRNA with respect to the exogenous reference gene Unisp6 [44], the spike-in used to assess the quality of the cDNA synthesis and qPCR process.
In Silico evaluation
Based on scientific literature, we selected the miRNAs that have been previously associated with MASLD and its comorbidities [12, 35, 66]. The association between miRNAs, validated target genes and diseases focusing on those miRNAs showing significant differences between control and intervention groups were analyzed. Once the relevant miRNAs were identified, we linked them to previously validated genes in humans by utilizing all publicly available database through miRWalk (http://mirwalk.umm.uni-heidelberg.de/; accessed on 11/06/2023). Among the genes associated with our miRNAs, we selected those with significant results. Subsequently, we specifically chose genes associated with MASLD or fatty liver disease, relying on the data provided by the DisGeNet (https://www.disgenet.org; accessed on 26/06/2023) database. To investigate interactions between the selected miRNAs and their target genes a network was created using miRNet (https://www.mirnet.ca/accessed on 28/06/2024). We amplified and highlighted in black those target genes that are shared between our specific miRNAs, which were marked in blue. The genes were obtained through miRTaBase v8.0 and TarBase v8.0 (ID type miRbase). Furthermore, to discover metabolic pathways and biological process from validated target genes associated with each miRNA, we used GeneCodis (https://genecodis.genyo.es/; accessed on 28/06/2024), choosing homo sapiens as a main organism.
Statistical analyses
The normality of the variables was evaluated using Shapiro–wilk test. Data presentation depended on distribution (mean ± standard deviation (SD) or median ± interquartile range (IQR)). Differences between groups were assessed using the Student’s t test or the Mann–Whitney U test and categorical variables with X2 test. Differences between miRNAs were presented as median ± (SE). The association between variables was evaluated using the Pearson correlation coefficient or Spearman's rho (p), as appropriate.
Univariate and multivariate logistic regressions analyses were performed with liver steatosis, liver stiffness or hepatic fat content variables as dependent variables and miRNAs and other covariables as independent variables. Receiver Operating Characteristic Curve Analyses (ROC) and the areas under the ROC curve (AUROC) were calculated to assess the predictive power of circulating miRNAs and their combinations with hepatic steatosis degree, liver stiffness and hepatic fat content. Regression models were adjusted by potential cofounders including sex, age, physical activity and inflammatory and biochemical markers related to MASLD (adiponectin, RBP4, TG, glucose, LECT2, leptin and chemerin). For the regressions analyses, steatosis degree was categorized into two groups (without steatosis vs mild, moderate and severe steatosis) and liver stiffness (m/s) (≤ 1.31 vs > 1.31) and hepatic fat content (%) (≤ 4.4 vs > 4.4) according to the median. Optimism-corrected value was used to validate the results obtained using the Tibshirani´s enhanced bootstrap method described previously [29].
Statistical analyses were carried out with the statistical program Stata version 15.0 (StataCorp 2011, College Station, TX, USA). All p values presented were two-tailed. Differences were considered statistically significant when p < 0.05.
Generative IA and IA-assisted technologies in the writing process
During the preparation of this work the author used OpenAI in order to enhance readability and language of the manuscript. After using this tool, the author reviewed and edited the content as needed and take full responsibility for the content of the publication.
Results
Overview of baseline characteristics
Characteristics of MASLD (n = 55) and control (n = 45) groups are shown in Table 1. Significant differences were found between both groups in body composition, anthropometric measures and biochemical determinations, clearly proving worse health status in MASLD group (Supplementary Fig. 1). Hepatic transaminases such as ALT and GGT, hepatic fat content, hepatic volume and steatosis degree were significantly increased in MASLD (p < 0.001). LECT2, chemerin, RBP4 and leptin were significantly increased in MASLD group, nevertheless adiponectin was significantly lower (p < 0.001). Lipid profile including total cholesterol, HDL and LDL was significantly lower in MASLD group than controls, although CRI was significantly higher in MASLD group as well as TG. MedDiet adherence score was significantly lower in MASLD group compared to controls (p < 0.001). No significant differences were found in physical activity between both groups.
Analyses of circulating miRNAs levels and their associations with MASLD and metabolic factors
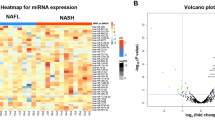
Eight miRNAs related with MASLD were selected and its circulating levels were analyzed (miR-21-5p, miR-151a-3p, miR-192-5p, miR-15b-3p, miR-29b-3p, miR-126-5p, miR-222-3p and miR-122-5p). Only circulating miR-122-5p, miR-151a-3p, miR-126-5p and miR-21-5p levels were significantly higher in MASLD group in comparison with control group (Fig. 1).
Although they were not strong, positive associations were found between variables such as hepatic fat content, steatosis degree and liver stiffness and circulating miRNAs such as miR-122-5p, miR-151a-3p, miR-126-5p and miR-21-5p (Fig. 2).
Correlation plots between liver stiffness, hepatic fat content, steatosis degree and miRNAs. Liver stiffness and (A) miR-122-5p, (B) miR-151a-3p, (C) miR-126-5p and (D) miR-21-5p. Hepatic fat content and (E) miR-122-5p, (F) miR-151a-3p, (G) miR-126-5p and (H) miR-21-5p. Steatosis degree and (I) miR-122-5p, (J) miR-151a-3p, (K) miR-126-5p and (L) miR-21-5p
Moreover, transaminases which are involved in MASLD, such as ALT (r = 0.46), AST (r = 0.38) and GGT (r = 0.47) were very positively correlated with miR-122-5p levels (p < 0.001) (Supplementary Tables 1, 2). Similarly, body composition and anthropometric measures were significantly associated with all miRNAs previously mentioned as well as HOMA index. Insulin was also positively associated with miR-122-5p, miR-151a-3p and miR-126-5p, and LECT2 was significantly correlated with miR-122-5p and miR-151a-3p. MiR-21-5p was also negatively associated with the MedDiet adherence score and adiponectin and was positively associated with chemerin.
Logistic regression analyses for MASLD and related variables
Univariate analysis between hepatic fat content, steatosis degree and liver stiffness as dependent variables and miRNAs as independent variables (Tables 2, 3 and 4), demonstrated a moderate capacity to predict steatosis degree and liver stiffness (AUROCs 0.62–0.68). Furthermore, miR-21-5p was able to predict hepatic fat content with a moderate capacity (AUROC 0.62). MiR-126-5p was the only that did not predict liver stiffness (Tables 2, 3 and 4).
Contributing factors such as adiponectin, RBP4, TG, glucose, LECT2, leptin and chemerin were analyzed to evaluate its ability to predict MASLD-associated factors (Supplementary Table 3). The optimism-corrected values were used to validate AUROC results.
Multivariable regressions were performed including the contributing factors selected previously. The first model (model 1) was adjusted by sex, age and physical activity. The rest of the models were created from model 1 and adding different MASLD-related variables such as adiponectin (model 2), RBP4 (model 3), TG (model 4) and glucose (model 5), LECT (model 6), leptin (model 7) and chemerin (model 8).
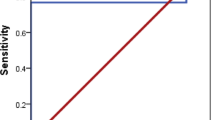
The most predictive capacity for liver stiffness with a moderate capacity was for miR-21-5p or miR-151a-3p both with leptin (model 7, AUC 0.76 in both, p < 0.05) (Table 2, Fig. 3). Other models with similar predictive capacity were miR-21-5p with chemerin, with an AUC of 0.70 and miR-151a-3p with LECT2 with an AUC of 0.72. Mir-122-5p with adiponectin had also predictive capacity but less that the other models with an AUC of 0.66. However, miR-126-5p was the only miRNA, that had not statistical significance in the predictive capacity for liver stiffness. For hepatic fat, miR-151a-3p with glucose (model 5) had the most ability to predict it with a high prediction capacity (model 5, AUC 0.81, p < 0.05, Fig. 3). In addition, miR-21-5p had also a high ability to predict hepatic fat with the third model that include RBP4 (AUC 0.80, p < 0.05,) (Table 3). Finally, the major value for predict the presence of steatosis was obtained combining miR-126-5p with leptin (model 7) obtaining an AUC of 0.95 (Fig. 3). A good capacity of prediction was achieved also using miR-122-5p, miR-151a-3p or miR-21-5p with adiponectin (model 2) with an AUC of 0.89 in all of them and miR-151-a3p or miR-21-5p with LECT2 (model 6) with an AUCs of 0.90, and 0.91, respectively (Table 4). We performed combinations of miRNAs with factors associated for MASLD (Supplementary Tables 4–6), but no combination was able to surpass the predictive capacity of each individual miRNAs.
In silico evaluation
The in silico analysis to further explore the potential role of miRNA in MASLD revealed that several genes associated with MASLD are regulated by specific microRNAs (Supplementary Table 7). Genes such as CREB1, MAPK1 or PTPN2 are extensively linked to MASLD and regulated by miR-122-5p. Additionally, miR-126-5p regulates important genes like SORT1 and PRKAA2, while miR-151a-3p regulates HADH and MCL1, as well as others. Furthermore, miR-21-5p regulates genes such as CCR7 and DNM1L, including additional ones. All these genes are regulated through different signaling pathways including JAK-STAT, PI3K-Akt, TNFα, chemerin, NOD-like receptor, mTOR, among others (Supplementary Tables 8–11). Additionally, the top 10 metabolic pathways according to p-adjusted value were depicted for the target genes associated with each miRNA in the Supplementary Fig. 2. Furthermore, a network of interactions between our specific miRNAs and their target genes were generated. The network shows a high density of interactions, with 4042 target genes identified. Some genes, such as CALD1, CCNG1, TNPO1, PURB, MYCBP2, MTPN, SNTB2, CBX5, CREBRF, BRWD1 and RBM12 were shared by miR-122-5p, miR-151-3p, miR-21-5p and miR-126-5p (Supplementary Fig. 3).
Discussion
MASLD is becoming a global health challenge, and there is a need for newer disease prediction and prognosis biomarker. Liver biopsy is the reference technique for diagnosis and prognosis of MASLD, however it has limitations and carries a risk for the patient because it is an invasive and complex method [30], so in this sense, the need arises to continue investigating new methods allowing to know the stage of the disease in a simple and more informative way.
Non-invasive imaging techniques like ultrasound, CT, MRI, and proton magnetic spectroscopy aim to replace biopsies but depend on operator skills and costly equipment. Ultrasound lacks objective analysis, while quantitative methods like proton density fat fraction (PDFF) require specialized equipment. Transient elastography (TE), the most common non-invasive MASLD diagnostic tool, has significant sampling variability, with probe positioning affecting results in over 30% of patients [58]. ARFI can have diagnostic accuracy comparable to that of TE, indicating its potential as a valuable tool in liver assessment. However, ARFI has limitations, including the requirement for the operator to define the region of interest and obtain a series of liver stiffness measurements (LSM) [33].
Despite the intricate biological complexity of miRNAs, they serve as reliable circulating biomarkers for MASLD diagnosis across various disease stages [39]. These small non-coding RNA molecules, typically ≤ 25 nucleotides long, regulate gene expression post-transcriptionally by binding to the 3' untranslated region of their target mRNAs [43]. MASLD is associated with alterations in hepatic miRNA expression patterns across early, intermediate, and advanced stages. Specific miRNA species are implicated in steatosis development and the progression of MASL to MASH and cirrhosis [30]. MiRNAs are excellent biomarkers due to their high stability, protected by vesicles and proteins, enabling resistance to external insults. Detection techniques based on PCR are extremely sensitive, capable of identifying even a single molecule [17]. Additionally, miRNAs can be detected in various bodily fluids including serum, plasma, whole blood, urine, and saliva. These miRNAs remain stable under different conditions such as temperature, pH variations, and over time, making them easy to measure, making them useful as diagnostic and prognostic indicator for diseases [11].
In the present study, we were able to observe that those participants with MASLD, whom had hepatic steatosis proven by ultrasonography, clearly had higher fat content and liver stiffness than in the control group, whom had lack of the disease. Elevated values of liver transaminases such as ALT and GGT, but not AST, were observed in MASLD group in comparison with control group. Moreover, it had been shown that ALT and GGT are biomarkers of liver disease, however AST could be more related with alcoholic liver or autoimmune diseases [53].
MiRNAs are interrelated to both inflammation and metabolic control [38]. HOMA-IR, HbA1c, insulin and blood glucose were increased in MASLD group in comparison with control group. HOMA-IR had been showed to be a good predictor for hepatic steatosis and fat content and insulin as a risk factor for MASLD [18]. As well, in hepatic IR, insulin, continues stimulating lipogenesis, which ends up producing hyperglycemia, hyperlipidemia, hepatic steatosis and T2DM [10]. MASLD can be associated with disturbances in glucose metabolism, and subsequently, elevated blood glucose levels may support the fat accumulation. In hepatic steatosis, the hepatic glucose utilization rate is increased, possibly as a result of IR and elevated insulin levels in the bloodstream [15, 34]. Despite healthier lifestyles, the control group showed higher LDL-c and TC levels, not yet reaching pathological thresholds [27]. Elevated HDL-c in controls might drive the TC increase, while low HDL-c links to MASLD [16]. MASLD group exhibited significantly increased TG levels, crucial in MASLD pathogenesis by promoting fatty acid deposition, aligning with previous studies considering TG as a risk factor for MASLD [65]. Additionally, higher atherogenic indices in MASLD suggest elevated cardiovascular risk and less favorable lipid profiles compared to controls [23].
A decreasing in adiponectin levels was observed in the MASLD group. Adiponectin acts as a protective agent against hepatic steatosis, inflammation, and fibrosis [24]. Similarly, we found adiponectin to be a protective factor against MASLD, predicting hepatic fat content, steatosis degree, and liver stiffness. Additionally, LECT2, leptin, and chemerin levels were increased in subjects with MASLD. Previous observations indicate a general increase in chemerin and LECT2 levels in obesity and IR states [36, 49]. Leptin had been observed to increase with the severity of MASLD, suggesting a possible compensatory mechanism against fat accumulation [49]. Recent studies showed that RBP4 levels in adipose and circulating tissue are associated with IR, dyslipidemia and T2DM and therefore linked to MASLD [47]. Our results showed that RBP4, in addition to being elevated in subjects with MASLD compared to the control group, could predict hepatic steatosis and hepatic fat content with a moderate capacity.
Furthermore, our miRNAs analyses revealed that only miR-21-5p, miR-151a-3p, miR-126-5p and miR-122-5p had significantly higher circulating levels in patients from the MASLD group compared with controls. Previous studies demonstrated that miR-122-5p was increased and positively correlated with markers of MASLD severity [66] and with body weight, TG, and body insulin insensitivity [2] and distinguished MASLD from healthy controls [39]. Similar, higher levels of miR-21 are effective biomarkers for MASLD diagnosis and play a key role in the development of the disease [54]. Other study showed that circulating miR-15b-3p, miR-21-5p, miR-29b-3p, miR-126-5p, miR-151a-3p and miR-192-5p were increased more than twice in a NASH group compared with the MASLD group [35], indicating a relation between miRNAs and the severity of the disease.
The analysis focused on miRNAs significantly differing between groups, finding positive associations between body composition, anthropometric measures, and all miRNAs. The MASLD group had lower adherence to the MedDiet and higher energy intake, aligning with obesity being a key factor in MASLD development, consistent with the negative correlation between miRNAs and MedDiet observed in our study. It had been shown that a balanced nutrition and moderate weight loss were the best therapeutic approach for MASLD [3]. Additionally, significant negative correlations were observed between our four miRNAs and adiponectin, particularly evident in miR-21-5p, supporting the documented role of miR21 in upregulating adiponectin mRNA expression [32]. On the other hand, chemerin, which participates in regulating angiogenesis, inflammation and cell proliferation [49], was positively correlated with miR-21-5p, which is positively correlated with steatosis, lobular inflammation, serum ALT and hepatic activity [66]. LECT2 positively correlated with miR-122-5p and miR-151a-3p, paralleling its significant levels in MASLD individuals with metabolic syndrome components and associations with obesity and anthropometric measures [36]. Likewise, liver stiffness, steatosis, and hepatic fat content correlated with miR-21-5p, miR-122-5p, miR-151a-3p, and miR-126-5p, consistent with findings in other studies [7, 20, 50, 64].
Our logistic regression analyses demonstrated the predictive capability of miRNAs for MASLD, even after adjusting for potential variables related to the disease such as leptin or glucose, among others. We obtained that the combination of miR-21-5p or miR-151a-3p with leptin for predict liver stiffness with an AUC of 0.76 for both options, miR-151a-3p with glucose was also the best combination to predict the hepatic fat content with an AUC of 0.81, and miR-126-5p with leptin to predict the presence or absence of hepatic steatosis and therefore, the disease, with an AUC of 0.95. Similar results were obtained in other study in which miR-122 discriminate subjects with MASLD from controls with an AUC of 0.85 [52]. In addition, it had been shown that miR-21-5p, miR151a-3p and miR-126,5p individually showed a diagnostic accuracy for MASH with an AUC of 0.73, 0.75 and 0.69, respectively for MASH as well as the combination of miR-21-5p, miR-151a-3p, miR-192-5p and miR-4449 with an AUC of 0.87 [35]. Furthermore, other study included miR-122-5p, miR-1290, miR-27b-3p and miR-192-5p with An AUC of 0.85 for MASLD [56]. In our study the combination of various miRNAs together does not provide better results than the miRNA individually for the diagnosis of our variables associated with MASLD suggesting that each miRNA could had been involved differently depending on the stage of the disease and could impact other miRNAs by predicting MASLD. Other non-invasive scores such as FLI index, Hepatic Steatosis Index (HIS), Steatotest, and NAFLD Liver fat Score showed a moderate diagnostic performance for MASLD (AUC from 0.68- 0.87), but it is important to highlight that the integration of these scores into everyday clinical practice is often limited by concerns about their diagnostic effectiveness [1]. Our study provides evidence of the predictive capacity of miR-122-5p, miR-151a-3p, miR-126-5p and miR-21-5p for hepatic steatosis, hepatic stiffness, and fat content, offering a non-invasive and promising method compared to liver biopsy and others non-invasive tools with a higher diagnostic performance. Furthermore, it integrates multiple diagnostic modalities and clinical variables for a comprehensive disease evaluation, promoting a personalized approach in medicine.
When we performed the in silico analysis, we found different target genes involved in MASLD, which are being regulated by the four miRNAs and which participate and contribute to the development and progression of the disease. In general, these genes are implicated in the regulation of lipid metabolism, inflammatory response, energy homeostasis and apoptosis in the liver through numerous pathways such as PI3K-AKT, JAK-STAT, mTOR, AMPK, cAMP, NF-kappa B, IL-6 and TNFα signaling pathways, among others [13, 19]. In addition, identifying shared target genes is also important as it underscores the complexity of gene regulation by multiple miRNAs. Such interactions may have significant implications for biological processes and disease mechanisms, underscoring the importance of considering miRNA co-regulation in molecular studies. Their dysfunctions can contribute to the accumulation of hepatic fat, chronic inflammation and metabolic imbalance observed in the development and progression of MASLD. Advancing our knowledge about this molecular mechanism will provide a better understanding of the diseases and enable the development of more effective therapeutic approaches. Studying the expression of miRNAs of interest in MASLD is crucial to understand its pathogenesis, diagnose the condition, and identify potential therapeutic targets. Additionally, their expression in various tissues can help in the development of miRNA-based therapies and serve as biomarkers for disease assessment.
However, the study has some limitations that should be acknowledged. First, miRNA level expression was normalized with the exogenous Unisp6, suggested by Vigneron et al. [59]. Not all studies had been controlled with this reference gene. Further evaluations are required using other different standardized controls in the applications of miRNAs as potential biomarkers for MASLD for reducing the technical variability among experimental replicates. Secondly, for the evaluation of MASLD non-invasive imaging techniques for the assessment of hepatic steatosis were used instead of a liver biopsy, which is the most reliable method in the detection of MASH or fibrosis in MASLD patients. However, this method is an invasive procedure, very expensive, with possible error in samples and complications that occurs during it procedure [66]. Thirdly, in our study, MASLD group consisted of individuals with MASLD and obesity, while our controls were subjects with normal-weight without MASLD. Although it would have been more interesting to have controls without MASLD and with obesity, it was challenging because most of the patients with obesity that we recruited, had MASLD. Ensuring uniform sample characteristics in a scientific study is crucial for maintaining research quality, result validity, and facilitating other researchers; ability to comprehend, replicate, and build upon your work, thus advancing scientific knowledge effectively and reliably. Fifthly, we only evaluated a few specific miRNAs. A comprehensive analysis of the full miRNA profiles in the blood could have identified other miRNAs that, alone or in combination, might serve as better predictors. Lastly, it is worth noting that this study evaluated a sample of patients at baseline. MicroRNAs (miRNAs), as epigenetic modifiers, can determine not only early disease evaluation but also the risk of progression and prognosis. Therefore, it would be highly interesting to observe changes in miRNA expression over time and assess differential aspects in miRNA expression patterns when different diets are applied.
Conclusion
In summary, findings of this study demonstrated that the combination of miRNAs including miR-151a-3p or miR-21-5p or miR-126-5p along leptin and miR-151a-3p with glucose can be used as non-invasive biomarker for the comprehensive assessment of steatosis, liver stiffness and hepatic fat content, which are critical factors in determining the presence of MASLD. These findings do not only highlight the promise of miRNAs as epigenetic regulators but also accentuate their application in the early assessment of the disease with potential in precision medicine.
Data availability
Data available on request due to privacy/ethical restrictions by sending an email to the corresponding author.
References
Abdelhameed F, Kite C, Lagojda L, Dallaway A, Chatha KK, Chaggar SS, Dalamaga M, Kassi E, Kyrou I, Randeva HS (2024) Non-invasive Scores and Serum Biomarkers for Fatty Liver in the Era of Metabolic Dysfunction-associated Steatotic Liver Disease (MASLD): A Comprehensive Review From NAFLD to MAFLD and MASLD. Curr Obes Rep. https://doi.org/10.1007/s13679-024-00574-z
Alrob OA, Khatib S, Naser SA (2017) MicroRNAs 33, 122, and 208: a potential novel targets in the treatment of obesity, diabetes, and heart-related diseases. J Physiol Biochem 73(2):307–314. https://doi.org/10.1007/s13105-016-0543-z
Anania C, Perla FM, Olivero F, Pacifico L, Chiesa C (2018) Mediterranean diet and nonalcoholic fatty liver disease. World J Gastroenterol 24(19):2083–2094. https://doi.org/10.3748/wjg.v24.i19.2083
Ando Y, Yamazaki M, Yamada H, Munetsuna E, Fujii R, Mizuno G, Ichino N, Osakabe K, Sugimoto K, Ishikawa H et al (2019) Association of circulating miR-20a, miR-27a, and miR-126 with non-alcoholic fatty liver disease in general population. Sci Rep 9(1):18856. https://doi.org/10.1038/s41598-019-55076-z
Bandiera S, Pfeffer S, Baumert TF, Zeisel MB (2015) miR-122–a key factor and therapeutic target in liver disease. J Hepatol 62(2):448–457. https://doi.org/10.1016/j.jhep.2014.10.004
Bedogni G, Bellentani S, Miglioli L, Masutti F, Passalacqua M, Castiglione A, Tiribelli C (2006) The Fatty Liver Index: a simple and accurate predictor of hepatic steatosis in the general population. BMC Gastroenterol 6:33. https://doi.org/10.1186/1471-230X-6-33
Boštjančič E, Bandelj E, Luzar B, Poljak M, Glavač D (2015) Hepatic expression of miR-122, miR-126, miR-136 and miR-181a and their correlation to histopathological and clinical characteristics of patients with hepatitis C. J Viral Hepat 22(2):146–157. https://doi.org/10.1111/jvh.12266
Brandão-Lima PN, Carvalho GB, Payolla TB, Sarti FM, Rogero MM (2022) Circulating microRNA Related to Cardiometabolic Risk Factors for Metabolic Syndrome: A Systematic Review. Metabolites 12(11):1044. https://doi.org/10.3390/metabo12111044
Cantero I, Elorz M, Abete I, Marin BA, Herrero JI, Monreal JI, Benito A, Quiroga J, Martínez A, Huarte MP et al (2019) Ultrasound/Elastography techniques, lipidomic and blood markers compared to Magnetic Resonance Imaging in non-alcoholic fatty liver disease adults. Int J Med Sci 16(1):75–83. https://doi.org/10.7150/ijms.28044
Chao HW, Chao SW, Lin H, Ku HC, Cheng CF (2019) Homeostasis of glucose and lipid in non-alcoholic fatty liver disease. Int J Mol Sci 20(2):298. https://doi.org/10.3390/ijms20020298
Chiam K, Mayne GC, Wang T, Watson DI, Irvine TS, Bright T, Smith LT, Ball IA, Bowen JM, Keefe DM et al (2020) Serum outperforms plasma in small extracellular vesicle microRNA biomarker studies of adenocarcinoma of the esophagus. World J Gastroenterol 26(20):2570–2583. https://doi.org/10.3748/wjg.v26.i20.2570
de Conti A, Ortega JF, Tryndyak V, Dreval K, Moreno FS, Rusyn I, Beland FA, Pogribny IP (2017) MicroRNA deregu-lation in nonalcoholic steatohepatitis-associated liver carcinogenesis. Oncotarget 8(51):88517–88528. https://doi.org/10.18632/oncotarget.19774
De Gregorio E, Colell A, Morales A, Marí M (2020) Relevance of SIRT1-NF-κB Axis as Therapeutic Target to Ameliorate Inflammation in Liver Disease. Int J Mol Sci 21(11):3858. https://doi.org/10.3390/ijms21113858
De la Iglesia R, Lopez-Legarrea P, Abete I, Bondia-Pons I, Navas-Carretero S, Forga L, Martinez JA, Zulet MA (2014) A new dietary strategy for long-term treatment of the metabolic syndrome is compared with the American Heart Association (AHA) guidelines: the MEtabolic Syndrome REduction in NAvarra (RESMENA) project. Br J Nutr 111(4):643–652. https://doi.org/10.1017/S0007114513002778
Della Pepa G, Lupoli R, Masulli M, Boccia R, De Angelis R, Gianfrancesco S, Piccolo R, Rainone C, Rivellese AA, Annuzzi G, Bozzetto L (2024) Blood glucose control and metabolic dysfunction-associated steatotic liver disease in people with type 1 diabetes. J Endocrinol Invest. https://doi.org/10.1007/s40618-024-02333-2
Deprince A, Haas JT, Staels B (2020) Dysregulated lipid metabolism links NAFLD to cardiovascular disease. Mol Metab 42:101092. https://doi.org/10.1016/j.molmet.2020.101092
Di Mauro S, Scamporrino A, Filippello A, Di Pino A, Scicali R, Malaguarnera R, Purrello F, Piro S (2021) Clinical and molecular biomarkers for diagnosis and staging of NAFLD. Int J Mol Sci 22(21):11905. https://doi.org/10.3390/ijms222111905
Elorz M, Benito-Boilos A, Marin BA, Pérez Díaz Del Campo N, Herrero JI, Monreal JI, Tur JA, Martínez JA, Zulet MA, Abete I (2022) Neck Circumference for NAFLD Assessment during a 2-Year Nutritional Intervention: The FLiO Study. Nutrients 14(23):5160. https://doi.org/10.3390/nu14235160
Ersahin T, Tuncbag N, Cetin-Atalay R (2015) The PI3K/AKT/mTOR interactive pathway. Mol BioSyst 11(7):1946–1954. https://doi.org/10.1039/c5mb00101c
Ezaz G, Trivedi HD, Connelly MA, Filozof C, Howard K, Parrish ML, Kim M, Herman MA, Nasser I, Afdhal NH et al (2020) Differential Associations of Circulating MicroRNAs With Pathogenic Factors in NAFLD. Hepatol Commun 4(5):670–680. https://doi.org/10.1002/hep4.1501
Fang Z, Dou G, Wang L (2021) MicroRNAs in the Pathogenesis of Nonalcoholic Fatty Liver Disease. Int J Biol Sci 17(7):1851–1863. https://doi.org/10.7150/ijbs.59588
Fernández-Ballart JD, Piñol JL, Zazpe I, Corella D, Carrasco P, Toledo E, Perez-Bauer M, Martínez-González MA, Salas-Salvadó J, Martín-Moreno JM (2010) Relative validity of a semi-quantitative food-frequency questionnaire in an elderly Mediterranean population of Spain. Br J Nutr 103(12):1808–1816. https://doi.org/10.1017/S0007114509993837
Fernández-Macías JC, Ochoa-Martínez AC, Varela-Silva JA, Pérez-Maldonado IN (2019) Atherogenic index of plasma: novel predictive biomarker for cardiovascular illnesses. Arch Med Res 50(5):285–294. https://doi.org/10.1016/j.arcmed.2019.08.009
Francisco V, Sanz MJ, Real JT, Marques P, Capuozzo M, AitEldjoudi D, Gualillo O (2022) Adipokines in non-alcoholic fatty liver disease: Are we on the road toward new biomarkers and therapeutic targets? Biology 11(8):1237. https://doi.org/10.3390/biology11081237
Friedewald WT, Levy RI, Fredrickson DS (1972) Estimation of the concentration of low-density lipoprotein cholesterol in plasma, without use of the preparative ultracentrifuge. Clin Chem 18:499–502. https://doi.org/10.1093/clinchem/18.6.499
Galmes-Panades AM, Konieczna J, Abete I, Colom A, Rosique-Esteban N, Zulet MA, Vázquez Z, Estruch R, Vidal J, Toledo E et al (2019) Lifestyle factors and visceral adipose tissue: Results from the PREDIMED-PLUS study. PLoS ONE 14(1):e0210726. https://doi.org/10.1371/journal.pone.0210726
Grundy SM, Stone NJ, Bailey AL, Beam C, Birtcher KK, Blumenthal RS, Braun LT, de Ferranti S, Faiella-Tommasino J, Forman DE et al (2019) 2018 AHA/ACC/AACVPR/AAPA/ABC/ACPM/ADA/AGS/APhA/ASPC/NLA/PCNA Guideline on the Management of Blood Cholesterol: Executive Summary: A Report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. J Am Coll Cardiol 73(24):3168–3209. https://doi.org/10.1016/j.jacc.2018.11.002
Hagström H, Vessby J, Ekstedt M, Shang Y (2024) 99% of patients with NAFLD meet MASLD criteria and natural history is therefore identical. J Hepatol 80(2):e76–e77. https://doi.org/10.1016/j.jhep.2023.08.026
Harrell F (2015) Applications to Linear Models, Logistic and Ordinal Regression, and Survival Analysis. In Regression Modeling Strategies. New York, NY: Springer
Hochreuter MY, Dall M, Treebak JT, Barrès R (2022) MicroRNAs in non-alcoholic fatty liver disease: Progress and perspectives. Mol Metab 65:101581. https://doi.org/10.1016/j.molmet.2022.101581
Huang YH, Lin KH, Chen HC, Chang ML, Hsu CW, Lai MW, Chen TC, Lee WC, Tseng YH, Yeh CT (2012) Identification of postoperative prognostic microRNA predictors in hepatocellular carcinoma. PLoS ONE 7(5):e37188. https://doi.org/10.1371/journal.pone.0037188
Kang M, Yan LM, Zhang WY, Li YM, Tang AZ, Ou HS (2013) Role of microRNA-21 in regulating 3T3-L1 adipocyte differentiation and adiponectin expression. Mol Biol Rep 40(8):5027–5034. https://doi.org/10.1007/s11033-013-2603-6
Kang YW, Baek YH, Lee JH, Roh YH, Kwon HJ, Moon SY, Son MK, Jeong JS (2024) Assessing the utility of acoustic radiation force impulse in the evaluation of non-alcoholic fatty liver disease with severe obesity or steatosis. Diagnostics (Basel) 14(11):1083. https://doi.org/10.3390/diagnostics14111083
Keramida G, Hunter J, Peters AM (2016) Hepatic glucose utilization in hepatic steatosis and obesity. Biosci Rep 36:e00402. https://doi.org/10.1042/BSR20160381
Kim TH, Lee Y, Lee YS, Gim JA, Ko E, Yim SY, Jung YK, Kang S, Kim MY, Kim H et al (2021) Circulating miRNA is a useful diagnostic biomarker for nonalcoholic steatohepatitis in nonalcoholic fatty liver disease. Sci Rep 11(1):14639. https://doi.org/10.1038/s41598-021-94115-6
Lan F, Misu H, Chikamoto K, Takayama H, Kikuchi A, Mohri K, Takata N, Hayashi H, Matsuzawa-Nagata N, Takeshita Y et al (2014) LECT2 functions as a hepatokine that links obesity to skeletal muscle insulin resistance. Diabetes 63(5):1649–1664. https://doi.org/10.2337/db13-0728
Li H, Liang J, Han M, Gao Z (2024) Polyphenols synergistic drugs to ameliorate non-alcoholic fatty liver disease via signal pathway and gut microbiota: A review. J Adv Res S2090–1232(24):00091–00092. https://doi.org/10.1016/j.jare.2024.03.004
Lischka J, Schanzer A, Hojreh A, Ba-Ssalamah A, de Gier C, Valent I, Item CB, Greber-Platzer S, Zeyda M (2021) Circulating microRNAs 34a, 122, and 192 are linked to obesity-associated inflammation and metabolic disease in pediatric patients. Int J Obes (Lond) 45(8):1763–1772. https://doi.org/10.1038/s41366-021-00842-1
Liu CH, Ampuero J, Gil-Gómez A, Montero-Vallejo R, Rojas Á, Muñoz-Hernández R, Gallego-Durán R, Romero-Gómez M (2018) miRNAs in patients with non-alcoholic fatty liver disease: A systematic review and meta-analysis. J Hepatol 69(6):1335–1348. https://doi.org/10.1016/j.jhep.2018.08.008
Mahmoudi A, Butler AE, Jamialahmadi T, Sahebkar A (2022) The role of exosomal miRNA in nonalcoholic fatty liver disease. J Cell Physiol 237(4):2078–2094. https://doi.org/10.1002/jcp.30699
Mantilla Toloza SC, Gómez-Conesa A (2007) El Cuestionario Internacional de Actividad Física. Un instrumento adecuado en el seguimiento de la actividad física poblacional. Rev Iberoam Fisioter Kinesiol 10(1):48–52. https://doi.org/10.1016/s1138-6045(07)73665-1
Marin-Alejandre BA, Cantero I, Perez-Diaz-Del-Campo N, Monreal JI, Elorz M, Herrero JI, Benito-Boillos A, Quiroga J, Martinez-Echeverria A, Uriz-Otano JI (2021) Effects of two personalized dietary strategies during a 2-year intervention in subjects with nonalcoholic fatty liver disease: A randomized trial. Liver Int 41(7):1532–1544. https://doi.org/10.1111/liv.14818
Martinez MA, Tural C, Franco S (2022) Circulating MicroRNAs as a Tool for Diagnosis of Liver Disease Progression in People Living with HIV-1. Viruses 14(6):1118. https://doi.org/10.3390/v14061118
Martínez-Fleta P, Vera-Tomé P, Jiménez-Fernández M, Requena S, Roy-Vallejo E, Sanz-García A, Lozano-Prieto M, López-Sanz C, Vara A, Lancho-Sánchez Á, Martín-Gayo E et al (2022) A Differential Signature of Circulating miRNAs and Cytokines Between COVID-19 and Community-Acquired Pneumonia Uncovers Novel Physiopathological Mechanisms of COVID-19. Front Immunol 12:815651. https://doi.org/10.3389/fimmu.2021.815651
Moore JB (2019) From sugar to liver fat and public health: Systems biology driven studies in understanding non-alcoholic fatty liver disease pathogenesis. Proc Nutr Soc 78(3):290–304. https://doi.org/10.1017/S0029665119000570
Moreiras O, Carbajal Á, Cabrera L, Cuadrado C (2003) Tablas de composición de alimentos. Ediciones Pirámide, Madrid, Spain
Nono Nankam PA, Blüher M (2021) Retinol-binding protein 4 in obesity and metabolic dysfunctions. Mol Cell Endocrinol 531:111312. https://doi.org/10.1016/j.mce.2021.111312
Pirola CJ, Sookoian S (2022) Metabolic dysfunction-associated fatty liver disease: advances in genetic and epigenetic implications. Curr Opin Lipidol 33(2):95–102. https://doi.org/10.1097/MOL.0000000000000814
Polyzos SA, Kountouras J, Mantzoros CS (2016) Adipokines in nonalcoholic fatty liver disease. Metabolism 65(8):1062–1079. https://doi.org/10.1016/j.metabol.2015.11.006
Pratama MY, Pascut D, Tamini S, Minocci A, Tiribelli C, Grugni G, Sartorio A (2020) Circulating microRNA associated to different stages of liver steatosis in prader-willi syndrome and non-syndromic obesity. J Clin Med 9(4):1123. https://doi.org/10.3390/jcm9041123
Rinella ME, Lazarus JV, Ratziu V, Francque SM, Sanyal AJ, Kanwal F, Romero D, Abdelmalek MF, Anstee QM, Arab JP et al (2023) A multisociety Delphi consensus statement on new fatty liver disease nomenclature. Hepatology 78(6):1966–1986. https://doi.org/10.1097/HEP.0000000000000520
Salvoza NC, Klinzing DC, Gopez-Cervantes J, Baclig MO (2016) Association of circulating serum miR-34a and miR-122 with dyslipidemia among patients with non-alcoholic fatty liver disease. PLoS ONE 11(4):e0153497. https://doi.org/10.1371/journal.pone.0153497
Sharma P (2022) Value of liver function tests in cirrhosis. J Clin Exp Hepatol 12(3):948–964. https://doi.org/10.1016/j.jceh.2021.11.004
Shatoor AS, Al Humayed S, Almohiy HM (2022) Astaxanthin attenuates hepatic steatosis in high-fat diet-fed rats by suppressing microRNA-21 via transactivation of nuclear factor erythroid 2-related factor 2. J Physiol Biochem 78(1):151–168. https://doi.org/10.1007/s13105-021-00850-9
Sodum N, Kumar G, Bojja SL, Kumar N, Rao CM (2021) Epigenetics in NAFLD/NASH: Targets and therapy. Pharmacol 167:105484. https://doi.org/10.1016/j.phrs.2021.105484
Tan Y, Ge G, Pan T, Wen D, Gan J (2014) A pilot study of serum microRNAs panel as potential biomarkers for diagnosis of nonalcoholic fatty liver disease. PLoS One 9(8):e105192. https://doi.org/10.1371/journal.pone.0105192
Tobaruela-Resola AL, Riezu-Boj JI, Milagro FI, Mogna-Pelaez P, Herrero JI, Elorz M, Benito-Boillos A, Tur JA, Martínez JA, Abete I et al (2024) Multipanel approach including mirnas, inflammatory markers, and depressive symptoms for metabolic dysfunction-associated steatotic liver disease diagnosis during 2-year nutritional intervention. Nutrients 16(11):1547. https://doi.org/10.3390/nu16111547
Turchinovich A, Baranova A, Drapkina O, Tonevitsky A (2018) Cell-free circulating nucleic acids as early biomarkers for NAFLD and NAFLD-associated disorders. Front Physiol 9:1256. https://doi.org/10.3389/fphys.2018.01256
Vigneron N, Meryet-Figuière M, Guttin A, Issartel JP, Lambert B, Briand M, Louis MH, Vernon M, Lebailly P, Lecluse Y et al (2016) Towards a new standardized method for circulating miRNAs profiling in clinical studies: Interest of the exogenous normalization to improve miRNA signature accuracy. Mol Oncol 10(7):981–992. https://doi.org/10.1016/j.molonc.2016.03.005
Wang X, He Y, Mackowiak B, Gao B (2021) MicroRNAs as regulators, biomarkers and therapeutic targets in liver diseases. Gut 70(4):784–795. https://doi.org/10.1136/gutjnl-2020-322526
Xin S, Zhan Q, Chen X, Xu J, Yu Y (2020) Efficacy of serum miRNA test as a non-invasive method to diagnose nonalcoholic steatohepatitis: A systematic review and meta-analysis. BMC Gastroenterol 20(1):186. https://doi.org/10.1186/s12876-020-01334-8
Younossi Z, Tacke F, Arrese M, Chander Sharma B, Mostafa I, Bugianesi E, Wai-Sun Wong V, Yilmaz Y, George J et al (2019) Global Perspectives on Nonalcoholic Fatty Liver Disease and Nonalcoholic Steatohepatitis. Hepatology 69(6):2672–2682. https://doi.org/10.1002/hep.30251
Younossi ZM, Golabi P, Paik JM, Henry A, Van Dongen C, Henry L (2023) The global epidemiology of nonalcoholic fatty liver disease (NAFLD) and nonalcoholic steatohepatitis (NASH): A systematic review. Hepatology 77(4):1335–1347. https://doi.org/10.1097/HEP.0000000000000004
Yu F, Jiang Z, Chen B, Dong P, Zheng J (2017) NEAT1 accelerates the progression of liver fibrosis via regulation of microRNA-122 and Kruppel-like factor 6. J Mol Med (Berl) 95(11):1191–1202. https://doi.org/10.1007/s00109-017-1586-5
Yuan S, Chen J, Li X, Fan R, Arsenault B, Gill D, Giovannucci EL, Zheng JS, Larsson SC (2022) Lifestyle and metabolic factors for nonalcoholic fatty liver disease: Mendelian randomization study. Eur J Epidemiol 37(7):723–733. https://doi.org/10.1007/s10654-022-00868-3
Zhang Z, Moon R, Thorne JL, Moore JB (2023) NAFLD and vitamin D: Evidence for intersection of microRNA-regulated pathways. Nutr Res Rev 36(1):120–139. https://doi.org/10.1017/S095442242100038X
Acknowledgements
The authors are very grateful to all the participants of the study as well as to Bertha Araceli Marin-Alejandre, Nuria Pérez-Díaz-Del-Campo, Irene Cantero, Veronica Ciaurriz, Vanessa Bullon- Vela and Cristina Galarregui, and to the professional staff of the FLiO study (CUN, CIN and CHN). The authors also thank LABORATORIOS CINFA, S.A. and VISCOFAN, S.A. for financial support of the Centre for Nutrition Research. Finally, the authors wish to express their gratitude to the Government of Navarra, CIBERobn, and Fundació La Marató de TV3 for the financial support of the project.
Funding
Open Access funding provided thanks to the CRUE-CSIC agreement with Springer Nature. This research was funded by the Health Department of the Government of Navarra, with the FLiO (61/2015) and EHGNA (PC082-083–084) projects, CIBERobn (Physiology of Obesity and Nutrition, CB12/03/30002 and CB12/03/30038), Fundació La Marató de TV3 (201630.10) and the Centre for Nutrition Research (CIN) of the University of Navarra.
Author information
Authors and Affiliations
Contributions
Conceptualization, A.L.T-R., F.I.M., I.A. and M.A.Z.; methodology, A.L.T-R., F.M.I., M.E., A.B-B., J.I.H., P. M-P., I.A. and M.A.Z.; validation, A.L.T-R., F.I.M., I.A. and M.A.Z.; formal analysis, A.L.T-R., F.I.M., I.A. and M.A.Z; investigation, A.L.T-R, F.I.M., I.A. and M.A.Z; resources, A.L.T-R., F.I.M., I.A. and M.A.Z.; data curation, A.L.T-R., F.I.M., I.A. and M.A.Z.; writing- original draft preparation, A.L.T-R., F.I.M., I.A. and M.A.Z.; writing -review and editing, A.L.T-R., F.I.M., I.A. and M.A.Z; visualization, A.L.T-R., F.I.M., I.A. and M.A.Z.; supervision, A.L.T-R., I.A. and M.A.Z.; project administration, I.A. and M.A.Z.; funding acquisition, J.A.T., J.A.M., I.A. and M.A.Z. All the authors reviewed and approved the final manuscript of this publication. Itziar Abete and M. Angeles Zulet contributed equally to this work.
Corresponding author
Ethics declarations
Ethics approval
This research was performed under the principles of the Declaration of Helsinki, and it was approved by the Ethics Committee of the University of Navarra (ref. 54/2015).
Conflict of interest
The authors declare no competing interests.
Informed consent
Informed consent was obtained from all the participants involved in this study.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Key Points
• Circulating miRNAs were significantly increased in the MASLD group.
• Steatosis degree, liver stiffness and hepatic fat content are key MASLD determinants.
• Combining miRNAs with leptin or glucose provides diagnostic values for liver variables.
• Circulating miRNAs can be used as non-invasive biomarkers for MASLD assessment.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tobaruela-Resola, A.L., Milagro, F.I., Elorz, M. et al. Circulating miR-122-5p, miR-151a-3p, miR-126-5p and miR-21-5p as potential predictive biomarkers for Metabolic Dysfunction-Associated Steatotic Liver Disease assessment. J Physiol Biochem (2024). https://doi.org/10.1007/s13105-024-01037-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s13105-024-01037-8