Abstract
The Southern Ocean is experiencing complex climate change, and the Amundsen Sea is one of the regions that has responded most rapidly to climate change. Due to their role in ecosystems, environmental sensitivity and high endemism, Antarctic demersal fish are a favorable group that can act as an indicator of the response of Antarctic organisms to climate change. However, our knowledge of Antarctic fish fauna is insufficient, with knowledge gaps even in their taxonomy. This situation is greatly influenced by the limitations of traditional taxonomy and thus calls for alternative solutions such as DNA barcoding. In this study, DNA barcoding analyses of 69 fish samples obtained from the Amundsen Sea were conducted using the mitochondrial COI gene. Based on the molecular species delimitation results, 13 fish species were found to belong to two orders, six families, and 12 genera. Both the maximum likelihood and Bayesian inference methods showed that the phylogenetic relationships of Bathydraconidae were paraphyletic, which was consistent with previous phylogenetic research. Our research showed that the COI gene, as a DNA barcode, is not only suitable for the identification of Antarctic fish species but also reflects some phylogenetic characteristics that might provide important evidence and support for studies of Antarctic fish phylogenetic relationships. In summary, our study provides an important reference for fish diversity and taxonomy in the Amundsen Sea, which may further enhance our understanding of the biodiversity, taxonomy and biogeography of fish in this area.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Southern Ocean occupies almost 10% of the ocean area on Earth (Joyner 1998). It is the only ocean that surrounds Earth and is not divided by continents. This gives it a unique ocean current system. The Antarctic Circumpolar Current (ACC) travels around Antarctica in a clockwise direction, driven by sustained westerly winds (Allison et al. 2010). It prevents warm water from flowing from lower latitudes to higher latitudes, making the Southern Ocean one of the coldest oceans on Earth (Tynan 1998). Organisms in the Southern Ocean have adapted to the polar climate after millions of years of evolution (Clarke and Johnston 1996). However, complex climate change occurs in Antarctica, especially in West Antarctica, such as the Amundsen Sea (Jun et al. 2020). Changes in the marine environment, especially temperature (McGlone et al. 2010), salinity (Haumann et al. 2016), and dissolved oxygen (Keller et al. 2016), may have important effects on the marine ecosystem and biological community structure (La et al. 2019). As one of the most widely distributed and richest species groups in the ocean, fish are a key component in maintaining the balance of the marine ecosystem. They not only serve a basic ecological function but also play an important role in indicating the operating status of the ecosystem (Hunt et al. Jr 2002; Vander Zanden et al. 2011). Modern Antarctic fish fauna, whether in terms of biodiversity, abundance, or biomass, are mainly dominated by Notothenioidei, including Artedidraconidae, Bathydraconidae, Channichthyidae, Harpagiferidae, and Nototheniidae (Mintenbeck et al. 2012). These fish live in cold, oxygen-rich, and stable ocean environments and are highly endemic (Mintenbeck and Torres 2017). These characteristics, along with the roles the fish play in the ecosystem, make Antarctic fishes a favorable group that can act as an indicator of environmental change in the Southern Ocean.
Even in the vast ocean area of the Southern Ocean, there are only approximately 370 species of fish described that account for ~ 2% of all fish species worldwide, and this number is an underestimate (Eastman 2000). Ice cover, lack of deep-sea samples, low sampling frequency and insufficient traditional taxonomy may be the reasons for underestimation (Alt et al. 2021). Unfortunately, the situation of the fish fauna of the Amundsen Sea is even worse because the Amundsen Sea is located in a remote location relative to scientific research stations and routes (Griffiths et al. 2011). There have been only limited observation records and an underwater observation survey report (Eastman et al. 2012), while studies based on molecular taxonomy have not yet been reported. Currently, the Amundsen Sea is among the places where the sea temperature in the Southern Ocean rises most obviously (Kim et al. 2021). The rapid rise in sea temperature has led to a decrease in sea ice cover and a sustained decline in the ice shelf (Haumann et al. 2016). Meanwhile, the benthic ecosystem in Antarctica is vulnerable (Pineda-Metz et al. 2020), and glacier retreat (Sahade et al. 2015) and associated iceberg scouring (Gutt and Piepenburg 2003; Barnes and Souster 2011) have a huge impact on benthic communities, including Antarctic fish, which mostly belong to demersal fish (Mintenbeck et al. 2012). Moreover, the decline in salinity and dissolved oxygen (Yager et al. 2012; Randall-Goodwin et al. 2015) also brings challenges to fish survival that cannot be ignored. As one of the important indicator groups of climate change, the lack of information on the composition of fish communities in the Amundsen Sea will seriously affect the evaluation of the structure and function of its marine ecosystem. Therefore, a fish diversity baseline inventory is urgently needed, and clarifying the characteristics of Amundsen Sea fish diversity patterns can help us better understand the impacts of climate change on Amundsen Sea marine ecosystems.
Traditional fish classification is based on morphological identification, which is time consuming and depends on the experience of the taxonomist (Steinke et al. 2009). However, the morphologies of sibling species are similar, which can easily lead to misidentification. In particular, the amazing diversity of sizes, colors, and shapes in different life stages of fish is a challenge to taxonomists (Zhang and Hanner 2012). Moreover, the taxonomic division of some fish in the Southern Ocean is controversial (De Broyer et al. 2014). All these problems require new solutions. Species identification based on molecular biology has emerged to give taxonomists more choices and has the potential to become a universal method. This method is expected to become one of the most convincing types of classification evidence (Hebert et al. 2003a). DNA barcoding is increasingly advocated for in the identification of species. DNA barcoding based on the cytochrome c oxidase subunit Ι (COΙ) mitochondrial gene has been applied to the identification of species (Hebert et al. 2003b). A COΙ fragment of 650 bp has enough sequence diversity to reflect significant species-level differences and has demonstrated high efficiency and accuracy in species identification on a global scale, such as in Japanese marine fish (Zhang and Hanner 2011), Indian marine fish (Lakra et al. 2011), Cuban freshwater fish (Lara et al. 2010), Indo-Pacific coral reef fish (Hubert et al. 2012), and even birds (Hebert et al. 2004), mammals (Francis et al. 2010), and bivalves (Mikkelsen et al. 2007), among others. In this paper, the COΙ-based molecular identification method is applied to Antarctic fish of the Amundsen Sea. Our research aims to provide fundamental taxonomic information for fish species of the Amundsen Sea and thus provide a solid scientific basis for the ecological assessment and biological conservation of the Southern Ocean.
Materials and methods
Specimen collections
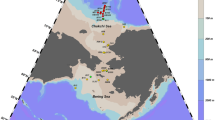
All specimens were collected at Xuelong icebreaker research vessels during the 36th Chinese National Antarctic Research Expedition (CHINARE) in 2020. Specimens were caught by a bottom trawling net (2.2 m wide, 0.65 m high, and 6.5 m long, 20 mm mesh diameter). Every net was employed for approximately 10 ~ 15 min at speeds of 2 ~ 3 kn. All samples were collected from 5 stations (Fig. 1) in the Amundsen Sea. All caught fish were sorted at -20 °C and provisionally identified. Muscle samples were stored in 95% ethanol for DNA extraction. Morphological identification followed Gon’s classification method (Graeme 1992). Finally, all fish were fixed in 10% formaldehyde and stored as voucher samples at the Third Institute of Oceanography, Ministry of Natural Resources.
DNA preparation, PCR and sequencing
DNA extraction was carried out with muscle tissue by using a DNeasy Blood and Tissue Kit [Qiagen, Hilden, Germany]. Some steps followed those of Hellberg et al. (2014). Microtubes of 1.5 mL [Axygen, New York, America] and ethanol (99.7%) [Xilong Scientific, Guangdong, China] were prepared in advance. Muscle samples (approximately 30 mg) were weighed into 1.5 mL microtubes, and then the steps in the manufacturer’s instructions were followed. Finally, DNA was stored at -20 ℃ until PCR amplification. The primers in this study were designed by Ward (2005) and were used for COΙ amplification.
All PCRs had a total volume of 25 µL and included 17.25 µL of ultrapure water, 2 µL of dNTPs (2.5 mM), 2.5 µL of 10 × PCR buffer (including Mg2+) (20 mM), 1 µL of each primer, 0.25 µL of Taq polymerase [TaKaRa, Kusatsu, Japan] (5 U/µL), and 1 µL of DNA template. Amplifications were performed using a SensoQuest LabCycler [SensoQuest, Germany] gradient thermal cycler. PCR cycling consisted of an initial step of 4 min at 95 ℃ and 35 cycles of 30 s at 94 ℃, 30 s at 50 ℃, and 30 s at 72 ℃, followed by a final extension at 72 ℃ for 10 min. PCR products were loaded onto 1% agarose gels and selected for sequencing, and all PCR products were purified and sequenced by Personal Biotechnology Co., Ltd.
DNA identification and phylogenetic analysis
All COI sequences were edited using DNASTAR Lasergene SeqMan Pro 7.1 and aligned manually using Sequencher 4.1 To facilitate the calculation of the genetic distance, two additional data points from the NCBI database were added for each species with fewer than three fish. We used two DNA identification methods to access taxonomic units: assembly of species by automatic partitioning (ASAP) (Puillandre et al. 2021) and Bayesian phylogenetics and phylogeography (BPP) (Yang et al. 2014) to infer putative species boundaries based on the COΙ gene. ASAP uses single locus sequence alignments to create species partitions; it is based on the implementation of a hierarchical clustering algorithm and compares only pairwise genetic distances. All aligned COΙ sequences were calculated by ASAP (https://bioinfo.mnhn.fr/abi/public/asap/asapweb.html) with the JC69 (Jukes-Cantor) model to compute the distance and default settings (split groups below probability 0.01, keep 10 best scores). BPP is a Bayesian Markov chain Monte Carlo (MCMC) tool for analyzing DNA sequences under the multispecies coalescent (MSC) model. The ultrametric tree with haplotypes was reconstructed using BEAST v1.10.4 (Drummond et al. 2012). The parameters in BEAUti use the GTR model and gamma shape site model. The number of gamma categories is 4, the relaxed clock is uncorrelated, and the chain length is 30,000,000 iterations for MCMC. The taxonomic units calculated by the ASAP and BPP were compared with the sequences of known species in the NCBI database to determine the taxonomic authenticity of the species. The taxonomic units with ≥ 98% similarity to the known sequences were the same species (Murphy et al. 2016), and those with < 98% and ≥ 95% similarity to the known sequences were the same genus (Ratnasingham & Hebert 2013).
The suitable genetic distance model was calculated by jModelTest v2.1.10 (Posada 2008). Genetic distances were calculated using the Kimura two-parameter (K2P) distance model (Kimura 1980) with 1000 bootstrap replicates and uniform rates using MEGA X (Kumar et al. 2018). Intra- and interspecies genetic distances and pairwise distance were considered. We used the online tool SMS to find suitable models of nucleotide substitution under the Akaike information criterion (AIC). A BI tree and ML tree were used to construct the phylogenetic relationships. The BI tree was constructed using MrBayes v3.1.2 (Huelsenbeck et al. 2001), and MCMC analysis was run with 10,000,000 generations, sampling every 1000 generations. We used PhyML3.0 (Guindon et al. 2010) to build an ML tree with GTR and 0.186 gamma shape parameters as substitution models, NII for tree improvement, and the aLRT SH-like fast likelihood method. Finally, the majority-rule consensus tree was reconstructed and displayed using Figtree v1.4.4.
Results
Morphological and DNA identification
A total of 69 fish samples were collected in this study. Most of them were adults and well preserved, but some individuals were small or damaged during preservation and thus difficult to identify. The identification was greatly limited by the poor Antarctic fish classification literature. In this study, 12 morphological species were identified by morphological characteristics and keys (Appendix 1).
All COΙ fragments were successfully amplified and sequenced. The sequences of the COΙ gene with high quality (no double peaks, short fragments or background noise) were aligned and contained no insertions, deletions, or stop codons. The length of the COΙ sequences was 652 bp after alignment, including 237 polymorphic sites (223 parsimony-informative sites, 14 singleton variable sites). The average base composition was A = 21.03%, C = 27.90%, G = 19.71%, and T = 31.36% on average, with a slight bias against G and C. The best classification result in ASAP (second-best model) supported 69 sequences representing 11 taxonomic units. Artedidraco lonnbergi and Dolloidraco longedorsalis were potentially one taxonomic unit. Lycenchelys sp. and Ophthalmolycus amberensis were also in the same situation. However, BPP showed a different result from ASAP (Fig. 2). BPP confirmed that 69 COI sequences belonged to 13 taxonomic units, and this result is basically consistent with the result of traditional morphological identification. Altogether, molecular methods proved that 69 sequences belonged to 13 species of fish, 12 genera, 6 families, and 2 orders (Table 1). The newly isolated nucleotide sequences were deposited in GenBank under accession numbers (Appendix 1).
Genetic distance and phylogeny analysis
The uncorrected K2P pairwise distance within species was below 1%, averaged 0.31%, and ranged from 0 to 1.01%. The genetic distance between species varied between 1.84% and 29.9% (Fig. 3). The best-fitting model was GTR + G, and the gamma distribution shape parameter was 0.186. Two phylogenetic trees, the BI tree and ML tree, showed similar topologies, and the majority-rule consensus tree was used to show the phylogenetic relationship of fish. The tree supported a branch of Bathydracinidae nested within Channichthyidae. Most individuals in the tree clustered together in groups of the same species.
Discussion
Effectiveness of COΙ barcoding and species identification
The accuracy of DNA barcoding is the key to species identification, which depends on the degree of intra- and interspecific variation of the selected gene fragments. The less intra- and interspecific overlap there is, the more effective the barcoding. Intraspecific variations are generally similar among species (Waugh 2007). However, the range of interspecific differences varies depending on the size of the selected group and geographic populations. The use of means for intraspecific and interspecific genetic distance comparisons does not allow for the detection of problematic cases. Therefore, we compared the minimum interspecific distance with the maximum intraspecific genetic distance (Meier et al. 2008). In this study, the minimum interspecific distance was 1.84%, the maximum intraspecific genetic distance was 1.01%, and the barcoding gap was between 1.01% and 1.84%.
We used two different methods to infer the putative species boundaries, namely ASAP and BPP. ASAP is based on single-marker pairwise genetic distance and avoids the heavy computational burden of phylogenetic reconstruction. It does not require any biological a priori insights and can quickly come up with relevant species hypotheses (Puillandre et al. 2021). BPP can accurately assign identity at the species level without knowing species boundaries in advance, even when analyzing rare taxa with only one locus available (Yang and Rannala 2017). The classification of most species is consistent. BPP and morphology have obtained similar results, while ASAP has some differences. As the BPP results were consistent with the BLAST results against the GenBank database, BPP was likely to show more accurate species identification results. However, it is worth noting that there are ten results displayed by ASAP. We consider the classification results of only the first- and second-best scores. If barcoding gaps or other prior conditions are considered, ASAP can achieve the same results as BPP. Overall, DNA identification can provide simple and reliable species classification results and shows the uniqueness of the method when morphology is difficult to perform.
In this study, 12 morphological species were identified, and 13 species were identified by DNA barcoding. Lycenchelys sp. was misidentified as O. amberensis. It is important to note that Lycenchelys sp. has been previously identified by Rock (2008), who supported that the individual was from a valid species without a morphological description. There are few data related to this species in the online database. The morphological characteristics of this sample in this study were impaired, and more specimens and more detailed descriptions of this species are still needed to determine its taxonomic status. Accurate taxonomic status and species identification require a combination of morphological and genetic findings. DNA barcoding shows the difference between the two species only at the genetic level but lacks support from morphological characteristics. The morphological characteristics of species are the scientific basis for their taxonomic status and biological studies, but traditional taxonomy relies on the experience of taxonomists. Therefore, a combination of molecular and traditional morphological methods for species identification is necessary.
Phylogenetic relationships
The COΙ gene is a short nucleotide fragment from mitochondria and is not the best choice for phylogenetic analysis; however, the topology of its phylogenetic tree might still have reference value (Steinke et al. 2009). The tree topology based on COΙ barcoding is usually related to the delineation of clusters. Although the ML tree was based on a priori inference and Bayesian inference was based on a posteriori inference, the topology supported by the results was basically the same (Fig. 4). In particular, they both supported that Bathydracinidae were paraphyletic. Previous studies reported similar results (Derome et al. 2002; Bargelloni et al. 2004). Multiple nuclear markers and multiple studies also confirmed that Bathydracinidae are paraphyletic (Near et al. 2004; Rock et al. 2008). In terms of the phylogenetic relationship, our COI-based phylogenetic signal further verifies the topological structure revealed by other studies.
The demersal fish fauna in the Amundsen Sea
In recent decades, with deepening research and the emergence of commercial fishing, increasing information about the community structure and classification of fish in the Southern Ocean has been discovered. In general, Notothenioidei, including Artedidraconidae, Bathydraconidae, Channichthyidae, Harpagiferidae, and Nototheniidae, has an absolute advantage in terms of number, accounting for most of the total species biodiversity (Eastman and McCune 2000; Eastman 2004). Additionally, there are some typical deep-sea fish groups, such as Liparidae and Zoarcidae. Some Antarctic fish diversity studies based on molecular taxonomy have been applied in the Ross Sea (Smith et al. 2012), Prydz Bay (Li et al. 2018), Scotia Sea (Rock et al. 2008), Dumont d’Urville Sea (Dettai et al. 2011), and Antarctic Peninsula (Mabragaña et al. 2016) and verified the aforementioned Antarctic fish diversity pattern.
In this study, 13 species of fish were identified in the surveyed seas, most of which belonged to Artedidraconidae, Bathydraconidae, Channichthyidae, and Nototheniidae in addition to Liparidae and Zoarcidae. Harpagiferiade did not appear in our study because these species are usually distributed in the sub-Antarctic region (Navarro et al. 2019), but the Amundsen Sea is located at high latitudes. Relatively speaking, there were only a few sampling stations with shallow sampling depths, which may be why we missed those typical deep-sea groups. At present, the fish fauna of the Amundsen Sea area have been studied by underwater observations. Our results supported that Notothenioidei dominates both in abundance and biomass. This is consistent with the aforementioned general pattern of the Southern Ocean fish fauna. The fish we caught were also roughly similar to the fauna observed by Eastman et al. (2012); however, our study provided more detailed assignment at the species level, with some additional exclusive species recorded. In particular, Ophthalmolycus amberensis, Chaenodraco wilsoni, Dacodraco hunteri, Akarotaxis nudiceps, Artedidraco lonnbergi and Vomeridens infuscipinnis might be recorded for the first time in the Amundsen Sea. It should also be noted that Eastman’s data came from underwater photography, and some species are difficult to identify by morphology; in contrast, our results are based on molecular taxonomy analysis of fish catches. From this perspective, our identification results are undoubtedly more credible.
To the best of our knowledge, our study is the first on the molecular taxonomy of fish in the Amundsen Sea. Our results provide important taxonomic information on the demersal fish fauna in the Amundsen Sea. This is of great significance for understanding the biodiversity, taxonomy and biogeography of fish in the Amundsen Sea. However, we believe that there remain many unknowns about the diversity of demersal fish in this area that should be explored. Broader sampling of latitudes, deeper sampling depths, and higher sampling densities are all necessary for future research. Finally, the integration of molecular identification and morphological identification is suggested to ensure precise taxonomy in future studies of Antarctic fishes.
Conclusions
This study illustrates the fauna and phylogenetic relationships of fish in the Amundsen Sea based on the 36th CHINARE. The results show that DNA barcoding is an effective method for identifying Antarctic fish, especially in the case of sample morphological damage. Thirteen species from six families of Antarctic fishes were identified, and six species were first recorded in the Amundsen Sea region. Our study provides reliable information on the distribution and classification of demersal fishes in the Amundsen Sea, which is highly similar to that in other parts of the Southern Ocean. The Amundsen Sea is geographically remote, but as one of the areas with the most rapid climate change, fish research in this area is an important part of the exploration of the Antarctic ecosystem affected by climate change. More surveys should be conducted to better understand fish in the Amundsen Sea and explore the profound impact of climate change on fish in polar regions.
References
Allison LC, Johnson HL, Marshall DP, Munday DR (2010) Where do winds drive the Antarctic Circumpolar Current? Geophys Res Lett 37. https://doi.org/10.1029/2010GL043355
Alt KG, Cunze S, Kochmann J, Klimpel SJAP (2021) Parasites of Three Closely Related Antarctic Fish Species (Teleostei: Nototheniinae) from Elephant Island. Acta Parasitol 67(1): 218-232.
Bargelloni L, Zane L, Derome N, Lecointre G, Patarnello T (2004) Molecular zoogeography of Antarctic euphausiids and notothenioids: from species phylogenies to intraspecific patterns of genetic variation. Antarct Sci 12:259–268
Barnes DK, Souster T (2011) Reduced survival of Antarctic benthos linked to climate-induced iceberg scouring. Nat Clim Change 1:365–368
Clarke A, Johnston IA (1996) Evolution and adaptive radiation of Antarctic fishes. Trends Ecol Evol 11:212–218
Broyer CD, Koubbi P, Griffiths H, Grant SA (2014) Biogeographic atlas of the Southern Ocean. Scientific Committee on Antarctic Research Cambridge. UK
Derome N, Chen WJ, Dettaı A, Bonillo C, Lecointre G (2002) Phylogeny of Antarctic dragonfishes (Bathydraconidae, Notothenioidei, Teleostei) and related families based on their anatomy and two mitochondrial genes. Mol Phylogenet Evol 24:139–152
Dettai A, Adamowizc SJ, Allcock L, Arango CP, Barnes DKA, Barratt I et al (2011) DNA barcoding and molecular systematics of the benthic and demersal organisms of the CEAMARC survey. Polar Sci 5:298–312
Drummond AJ, Suchard MA, Xie D, Rambaut A (2012) Bayesian Phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol, 29: 1969–1973
Eastman J, McCune A (2000) Fishes on the Antarctic continental shelf: evolution of amarine species flock? J Fish Biol 57:84–102
Eastman JT (2000) Antarctic notothenioid fishes as subjects for research in evolutionary biology. Antarct Sci 12:276–287
Eastman JT (2004) The nature of the diversity of Antarctic fishes. Polar Biol 28:93–107
Eastman JT, Amsler MO, Aronson RB, Thatje S, McClintock JB, Vos SC et al (2012) Photographic survey of benthos provides insights into the Antarctic fish fauna from the Marguerite Bay slope and the Amundsen Sea. Antarct Sci 25:31–43
Francis CM, Borisenko AV, Ivanova NV, Eger JL, Lim BK, Guillen-Servent A et al (2010) The role of DNA barcodes in understanding and conservation of mammal diversity in southeast Asia. PLoS One 5: e12575
Dewitt HH , Heemstra PC , Gon O (1992) Fishes of the Southern Ocean. Copeia 1:260.
Griffiths HJ, Danis B, Clarke A (2011) Quantifying Antarctic marine biodiversity: The SCAR-MarBIN data portal. Deep Sea Res Part II 58:18–29
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel OJSb (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Gutt J, Piepenburg D (2003) Scale-dependent impact on diversity of Antarctic benthos caused by grounding of icebergs. Mar Ecol Prog Ser 253:77–83
Haumann FA, Gruber N, Münnich M, Frenger I, Kern SJN (2016) Sea-ice transport driving Southern Ocean salinity and its recent trends. Nature 537:89–92
Hebert PDN, Cywinska A, Ball SL, deWaard JR (2003a) Biological identifications through DNA barcodes. Proc Biol Sci 270:313–321
Hebert PDN, Ratnasingham S, deWaard JR (2003b) Barcoding animal life: cytochrome c oxidase subunit 1 divergences among closely related species. Proc Biol Sci 270(Suppl 1):S96–99
Hebert PDN, Stoeckle MY, Zemlak TS, Francis CM (2004) Identification of Birds through DNA Barcodes. PLoS Biol 2:e312
Hellberg RS, Kawalek MD, Van KT, Shen Y, Williams-Hill DM (2014) Comparison of DNA Extraction and PCR Setup Methods for Use in High-Throughput DNA Barcoding of Fish Species. Food Anal Methods 7:1950–1959
Hubert N, Meyer CP, Bruggemann HJ, Guerin F, Komeno RJ, Espiau B et al (2012) Cryptic diversity in Indo-Pacific coral-reef fishes revealed by DNA-barcoding provides new support to the centre-of-overlap hypothesis. PLoS One 7: e28987
Huelsenbeck JP, Ronquist F, Nielsen R, Bollback, JPJS (2001) Bayesian inference of phylogeny and its impact on evolutionary biology. Science 294: 2310–2314
Hunt GL Jr, Stabeno P, Walters G, Sinclair E, Brodeur RD, Napp JM, et al (2002) Climate change and control of the southeastern Bering Sea pelagic ecosystem. Deep Sea Res Part II 49: 5821–5853
Joyner CC, Columbia, SC (1998) 369 Governing the frozen commons: The Antarctic regime and environmental protection. Polar Record 35(195):353.
Jun S-Y, Kim J-H, Choi J, Kim S-J, Kim B-M, An S-I (2020) The internal origin of the west-east asymmetry of Antarctic climate change. Sci Adv 6:eaaz1490
Keller DP, Kriest I, Koeve W, Oschlies AJ (2016) Southern Ocean biological impacts on global ocean oxygen. Geophys Res Lett 43:6469–6477
Kim S-Y, Lim D, Rebolledo L, Park T, Esper O, Muñoz P, et al (2021) A 350-year multiproxy record of climate-driven environmental shifts in the Amundsen Sea Polynya, Antarctica. Global and Planetary Change 205: 103589.
Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J Mol Evol 16:111–120
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547
La HS, Park K, Wahlin A, Arrigo KR, Kim DS, Yang EJ et al (2019) Zooplankton and micronekton respond to climate fluctuations in the Amundsen Sea polynya. Antarctica Sci Rep 9:10087
Lakra WS, Verma MS, Goswami M, Lal KK, Mohindra V, Punia P et al (2011) DNA barcoding Indian marine fishes. Mol Ecol Resour 11:60–71
Lara A, Ponce de Leon JL, Rodriguez R, Casane D, Cote G, Bernatchez L et al (2010) DNA barcoding of Cuban freshwater fishes: evidence for cryptic species and taxonomic conflicts. Mol Ecol Resour 10:421–430
Li Y, Zhang L, Song P, Zhang R, Wang L, Lin L (2018) Fish diversity and molecular taxonomy in the Prydz Bay during the 29th CHINARE. Acta Oceanol Sin 37:15–20
Mabragaña E, Delpiani S, Rosso J, González-Castro M, Antoni MD, Hanner R et al (2016) Barcoding Antarctic fishes: species discrimination and contribution to elucidate ontogenetic changes in Nototheniidae. DNA Barcoding in Marine Perspectives 213–242
McGlone MS, Turney CS, Wilmshurst JM, Renwick J, Pahnke KJNG (2010) Divergent trends in land and ocean temperature in the Southern Ocean over the past 18,000 years. Nature Geoscience 3:622–626
Meier R, Zhang G, Ali F (2008) The use of mean instead of smallest interspecific distances exaggerates the size of the “barcoding gap” and leads to misidentification. Syst Biol 57:809–813
Mikkelsen NT, Schander C, Willassen E (2007) Local scale DNA barcoding of bivalves (Mollusca): a case study. Zoolog Scr 36:455–463
Mintenbeck K, Barrera-Oro ER, Brey T, Jacob U, Knust R, Mark FC et al (2012) Impact of Climate Change on Fishes in Complex Antarctic Ecosystems. Academic Press 46: 351-426.
Mintenbeck K, Torres JJ (2017) Impact of Climate Change on the Antarctic Silverfish and Its Consequences for the Antarctic Ecosystem. The Antarctic Silverfish: a Keystone Species in a Changing Ecosystem. Springer, Cham 253-286.
Murphy KR, Kalmanek EA, Cheng CHC (2016) Diversity and biogeography of larval and juvenile notothenioid fishes in McMurdo Sound, Antarctica. Polar Biol 40:161–176
Navarro JM, Paschke K, Ortiz A, Vargas-Chacoff L, Pardo LM, Valdivia N (2019) The Antarctic fish Harpagifer antarcticus under current temperatures and salinities and future scenarios of climate change. Prog Oceanogr 174:37–43
Near TJ, Pesavento JJ, Cheng CH (2004) Phylogenetic investigations of Antarctic notothenioid fishes (Perciformes: Notothenioidei) using complete gene sequences of the mitochondrial encoded 16S rRNA. Mol Phylogenet Evol 32:881–891
Pineda-Metz SE, Gerdes D, Richter CJNc (2020) Benthic fauna declined on a whitening Antarctic continental shelf. Nature, 11:1–7
Posada D (2008) jModelTest: phylogenetic model averaging. Mol Biol Evol 25:1253–1256
Puillandre N, Brouillet S, Achaz G (2021) ASAP: assemble species by automatic partitioning. Mol Ecol Resour 21:609–620
Ratnasingham S, Hebert PD (2013) A DNA-based registry for all animal species: the Barcode Index Number (BIN) system.PloS one, 8(7), e66213
Randall-Goodwin E, Meredith M, Jenkins A, Yager P, Sherrell R, Abrahamsen EP, Guerrero R, Yuan X, Mortlock R, Gavahan KJESotA (2015) Freshwater distributions and water mass structure in the Amundsen Sea Polynya region, AntarcticaFreshwater distributions and water mass structure in the ASP region. Elementa: Science of the Anthropocene, https://doi.org/10.12952/journal.elementa.000065
Rock J, Costa FO, Walker DI, North AW, Hutchinson WF, Carvalho GR (2008) DNA barcodes of fish of the Scotia Sea, Antarctica indicate priority groups for taxonomic and systematics focus. Antarct Sci 20:253–262
Sahade R, Lagger C, Torre L, Momo F, Monien P, Schloss I, Barnes DK, Servetto N, Tarantelli S, Tatián MJSA (2015) Climate change and glacier retreat drive shifts in an Antarctic benthic ecosystem. Science Advances, 1, e1500050
Smith PJ, Steinke D, Dettai A, McMillan P, Welsford D, Stewart A, Ward RD (2012) DNA barcodes and species identifications in Ross Sea and Southern Ocean fishes. Polar Biol 35:1297–1310
Steinke D, Zemlak TS, Boutillier JA, Hebert PDN (2009) DNA barcoding of Pacific Canada’s fishes. Mar Biol 156:2641–2647
Tynan CT (1998) Ecological importance of the southern boundary of the Antarctic Circumpolar Current. Nature 392:708–710
Vander Zanden MJ, Vadeboncoeur Y, Chandra SJE (2011) Fish reliance on littoral–benthic resources and the distribution of primary production in lakes. Ecosystems 14:894–903
Waugh J (2007) DNA barcoding in animal species: progress, potential and pitfalls. BioEssays 29:188–197
Yager PL, Sherrell RM, Stammerjohn SE, Alderkamp A-C, Schofield O, Abrahamsen EP et al (2012) ASPIRE: the Amundsen Sea Polynya international research expedition. Oceanography 25: 40–53
Yang Z, Rannala BJMB (2014) Unguided species delimitation using DNA sequence data from multiple loci. Mol Biol Evol 31:3125–3135
Yang Z, Rannala BJME (2017) Bayesian species identification under the multispecies coalescent provides significant improvements to DNA barcoding analyses. Mol Ecol 26:3028–3036
Zhang J-B, Hanner R (2011) DNA barcoding is a useful tool for the identification of marine fishes from Japan. Biochem Syst Ecol 39:31–42
Zhang J, Hanner R (2012) Molecular approach to the identification of fish in the South China Sea. PLoS ONE 7:e30621
Acknowledgements
This work was supported by the “Impact and Response of Antarctic Seas to Climate Change” (IRASCC2020-2022-NO.01-02-02B & 02–03), Ministry of Natural Resources of the People’s Republic of China, Chinese Arctic and Antarctic Administration. We thank all the teammates and crew of the XUELONG R/V for their efforts in collecting specimens during the CHINARE-36 cruise.
Author information
Authors and Affiliations
Contributions
Shuai Cao: Conceptualization, Data curation, Writing-original draft preparation. Yuan Li: Data curation, Software, Writing- reviewing and editing. Ran Zhang: Investigation, Visualization. Xing Miao: Investigation, Writing-reviewing and editing. Longshan Lin: Supervision, Conceptualization, Writing - reviewing and editing, Resources. Hai Li: supervision, conceptualization, writing-original draft preparation, methodology, data curation, validation.
Corresponding authors
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Appendix 1 Information of samples and species identification
Appendix 1 Information of samples and species identification
Sample No. | Sample site | Longitude (°/W) | Latitude (°/S) | Sample Depth(m) | Molecular identification | Morphological identification | Genbank voucher No. | Similarity (%) | Genbank No. |
|---|---|---|---|---|---|---|---|---|---|
AN1 | A11-1 | 113.35 | 73.52 | 627 | Dacodraco hunteri | Dacodraco hunteri | HQ712963.1 | 99.85 | OK493632 |
AN2 | A11-4 | 117.32 | 72.25 | 523 | Lycenchelys sp. | Ophthalmolycus amberensis | EU326372.1 | 99.35 | OK493633 |
AN3 | A11-4 | 117.32 | 72.25 | 523 | Trematomus scotti | Trematomus scotti | KX676179.1 | 99.85 | OK493645 |
AN4 | A11-4 | 117.32 | 72.25 | 523 | Trematomus scotti | Trematomus scotti | HQ713279.1 | 99.69 | OK493646 |
AN5 | A11-4 | 117.32 | 72.25 | 523 | Trematomus scotti | Trematomus scotti | KX676176.1 | 99.54 | OK493647 |
AN6 | A11-4 | 117.32 | 72.25 | 523 | Trematomus scotti | Trematomus scotti | KX676176.1 | 99.69 | OK493648 |
AN7 | A11-4 | 117.32 | 72.25 | 523 | Trematomus scotti | Trematomus scotti | EU326433.1 | 100.00 | OK493649 |
AN8 | A11-4 | 117.32 | 72.25 | 523 | Trematomus scotti | Trematomus scotti | KX676176.1 | 99.85 | OK493650 |
AN9 | A11-4 | 117.32 | 72.25 | 523 | Trematomus scotti | Trematomus scotti | KX676176.1 | 100.00 | OK493651 |
AN10 | A11-4 | 117.32 | 72.25 | 523 | Trematomus scotti | Trematomus scotti | KX676176.1 | 100.00 | OK493652 |
AN11 | A11-4 | 117.32 | 72.25 | 523 | Trematomus scotti | Trematomus scotti | KX676181.1 | 99.85 | OK493653 |
AN12 | A11-4 | 117.32 | 72.25 | 523 | Trematomus scotti | Trematomus scotti | KX676176.1 | 100.00 | OK493654 |
AN13 | A11-4 | 117.32 | 72.25 | 523 | Trematomus scotti | Trematomus scotti | KX676179.1 | 99.54 | OK493655 |
AN14 | A11-1 | 113.35 | 73.52 | 627 | Vomeridens infuscipinnis | Vomeridens infuscipinnis | HQ713358.1 | 100.00 | OK493677 |
AN15 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676176.1 | 100.00 | OK493681 |
AN16 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | HQ713279.1 | 99.85 | OK493682 |
AN17 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676176.1 | 99.85 | OK493683 |
AN18 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676176.1 | 99.85 | OK493684 |
AN19 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676176.1 | 100.00 | OK493685 |
AN20 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676176.1 | 99.85 | OK493686 |
AN21 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676177.1 | 100.00 | OK493687 |
AN22 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676177.1 | 99.85 | OK493688 |
AN23 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676177.1 | 99.85 | OK493689 |
AN24 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676181.1 | 99.85 | OK493690 |
AN25 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676176.1 | 99.39 | OK493691 |
AN26 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | EU326433.1 | 99.85 | OK493692 |
AN27 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676171.1 | 99.69 | OK493693 |
AN28 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | HQ713279.1 | 99.54 | OK493694 |
AN29 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | EU326433.1 | 100.00 | OK493695 |
AN30 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676176.1 | 99.85 | OK493696 |
AN31 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676175.1 | 100.00 | OK493697 |
AN32 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676176.1 | 100.00 | OK493698 |
AN33 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676176.1 | 100.00 | OK493699 |
AN34 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | HQ713279.1 | 99.54 | OK493700 |
AN35 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676176.1 | 99.85 | OK493701 |
AN36 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676177.1 | 100.00 | OK493702 |
AN37 | A11-3 | 113.58 | 73.41 | 423 | Trematomus scotti | Trematomus scotti | KX676176.1 | 100.00 | OK493703 |
AN38 | A11-1 | 113.35 | 73.52 | 627 | Trematomus scotti | Trematomus scotti | KX676171.1 | 99.69 | OK493704 |
AN39 | A11-1 | 113.35 | 73.52 | 627 | Trematomus scotti | Trematomus scotti | KX676177.1 | 100.00 | OK493705 |
AN40 | A11-1 | 113.35 | 73.52 | 627 | Trematomus scotti | Trematomus scotti | KX676176.1 | 99.69 | OK493706 |
AN41 | A11-1 | 113.35 | 73.52 | 627 | Trematomus scotti | Trematomus scotti | KX676173.1 | 99.85 | OK493707 |
AN42 | A11-1 | 113.35 | 73.52 | 627 | Trematomus scotti | Trematomus scotti | KX676173.1 | 100.00 | OK493708 |
AN43 | A4-3 | 112.99 | 72.91 | 438 | Chaenodraco wilsoni | Chaenodraco wilsoni | NC_039158.1 | 99.09 | OK493709 |
AN44 | A11-1 | 113.35 | 73.52 | 627 | Chionodraco myersi | Chionodraco myersi | DQ526430.1 | 99.70 | OK493710 |
AN45 | A11-1 | 113.35 | 73.52 | 627 | Chionodraco myersi | Chionodraco myersi | DQ526430.1 | 99.56 | OK493711 |
AN46 | A11-1 | 113.35 | 73.52 | 627 | Macrourus whitsoni | Macrourus whitsoni | MT157320.1 | 97.99 | OK493712 |
AN47 | A11-1 | 113.35 | 73.52 | 627 | Macrourus whitsoni | Macrourus whitsoni | MT157320.1 | 100.00 | OK493713 |
AN48 | A4-3 | 112.99 | 72.91 | 438 | Chaenodraco wilsoni | Chaenodraco wilsoni | NC_039158.1 | 99.24 | OK493714 |
AN49 | A11-1 | 113.35 | 73.52 | 627 | Dolloidraco longedorsalis | Dolloidraco longedorsalis | NC_057667.1 | 99.56 | OK493715 |
AN50 | A11-1 | 113.35 | 73.52 | 627 | Dolloidraco longedorsalis | Dolloidraco longedorsalis | NC_057667.1 | 99.56 | OK493716 |
AN51 | A11-1 | 113.35 | 73.52 | 627 | Dolloidraco longedorsalis | Dolloidraco longedorsalis | NC_057667.1 | 99.56 | OK493717 |
AN52 | A11-2 | 115.10 | 73.02 | 693 | Dolloidraco longedorsalis | Dolloidraco longedorsalis | NC_057667.1 | 99.71 | OK493718 |
AN53 | A4-3 | 112.99 | 72.91 | 438 | Artedidraco lonnbergi | Artedidraco lonnbergi | HQ712823.1 | 100.00 | OK493719 |
AN54 | A11-1 | 113.35 | 73.52 | 627 | Trematomus loennbergii | Trematomus loennbergii | NC_048965.1 | 99.27 | OK493720 |
AN55 | A4-3 | 112.99 | 72.91 | 438 | Trematomus loennbergii | Trematomus loennbergii | NC_048965.1 | 99.56 | OK493721 |
AN56 | A11-1 | 113.35 | 73.52 | 627 | Akarotaxis nudiceps | Akarotaxis nudiceps | NC_057664.1 | 99.09 | OK493722 |
AN57 | A11-1 | 113.35 | 73.52 | 627 | Akarotaxis nudiceps | Akarotaxis nudiceps | NC_057664.1 | 99.41 | OK493723 |
AN58 | A4-3 | 112.99 | 72.91 | 438 | Ophthalmolycus amberensis | Ophthalmolycus amberensis | JN641043.1 | 100.00 | OK493724 |
AN59 | A11-4 | 117.32 | 72.25 | 523 | Gerlachea australis | Gerlachea australis | NC_057668.1 | 99.56 | OK493725 |
AN60 | A4-3 | 112.99 | 72.91 | 438 | Macrourus whitsoni | Macrourus whitsoni | MT157320.1 | 99.56 | OK493726 |
AN61 | A11-1 | 113.35 | 73.52 | 627 | Dacodraco hunteri | Dacodraco hunteri | HQ712963.1 | 99.85 | OK493727 |
AN62 | A4-3 | 112.99 | 72.91 | 438 | Chaenodraco wilsoni | Chaenodraco wilsoni | NC_039158.1 | 98.69 | OK493728 |
AN63 | A4-3 | 112.99 | 72.91 | 438 | Trematomus loennbergii | Trematomus loennbergii | NC_048965.1 | 99.41 | OK493730 |
AN64 | A4-3 | 112.99 | 72.91 | 438 | Trematomus loennbergii | Trematomus loennbergii | HQ713304.1 | 99.85 | OK493731 |
AN65 | A11-1 | 113.35 | 73.52 | 627 | Chionodraco myersi | Chionodraco myersi | DQ526430.1 | 99.56 | OK493732 |
AN66 | A11-1 | 113.35 | 73.52 | 627 | Vomeridens infuscipinnis | Vomeridens infuscipinnis | HQ713358.1 | 100.00 | OK493740 |
AN67 | A11-1 | 113.35 | 73.52 | 627 | Macrourus whitsoni | Macrourus whitsoni | MT157320.1 | 99.70 | OK493741 |
AN68 | A11-1 | 113.35 | 73.52 | 627 | Akarotaxis nudiceps | Akarotaxis nudiceps | NC_057664.1 | 99.70 | OK493743 |
AN69 | A4-3 | 112.99 | 72.91 | 438 | Gerlachea australis | Gerlachea australis | NC_057668.1 | 99.70 | OK493745 |
Rights and permissions
About this article
Cite this article
Cao, S., Li, Y., Miao, X. et al. DNA barcoding provides insights into Fish Diversity and Molecular Taxonomy of the Amundsen Sea. Conservation Genet Resour 14, 281–289 (2022). https://doi.org/10.1007/s12686-022-01273-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12686-022-01273-4