Abstract
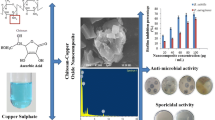
The growing resistance to existing antimicrobial formulations poses a great concern for global public health and demands the development of alternatives. Chitosan was synthesized from Archachatina marginata shell and characterized using Fourier transform infrared (FTIR), X-ray diffractometry (XRD), scanning electron microscopy (SEM), transmission electron microscopy (TEM), energy-dispersive X-ray fluorescence (EDXRF), and energy-dispersive X-ray spectroscopy (EDS) and its antibacterial activity assessed against Staphylococcus aureus (hemolytic and clinical strain), Staphylococcus saprophyticus DSM 18669, Escherichia coli 0157, Klebsiella pneumonia EO1 16S ribosomal RNA gene, and Salmonella typhi ATCC 13311. FTIR revealed hydroxyl, carbonyl, and amide as the main functional groups. XRD identified crystalline features, while SEM images indicated a grainy surface with clustering of uneven particles with TEM images showed polygonal particles, rod-like structures, and spherical voids on the surface with diameters ranging from 5.80 to 31.05 nm. EDXRF and EDS indicated calcium, carbon, and oxygen as the main elements. Chitosan produced the highest inhibition zone of 39 mm against Staphylococcus saprophyticus DSM, while there was no effect on Salmonella typhi ATCC 1331, and the minimum inhibitory concentration was between 0.0781 and 0.3125 mg/mL for tested strains. Chitosan’s antibacterial activity compared favorably to levofloxacin, a synthetic drug as a benchmark. This suggests that chitosan from Archachatina marginata shell is a potential antibacterial agent for multiple applications.
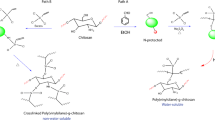
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Antibacterial resistance has grown significantly around the globe in recent years and is now acknowledged as a significant medical concern in the majority of healthcare settings [1, 2]. The burden of infectious illnesses is increased by the significant mortality attributable to resistance [3, 4]. Resistance is not a recent phenomenon; resistance genes are widely distributed in nature and engage in intricate ecological interactions [5, 6]. Antibacterial resistance was barely noticed in the past as newer antibiotics gradually became accessible and were easily changed and enhanced for therapeutic use, despite the fact that heavy usage of antimicrobial medications started to impose new survival pressure on important microorganisms. Researches have provided a fair description of the resistance situation, coordinated action plans have been put in place, and the urgent need for new antibiotics remains evident [7,8,9,10]. Annually, more than 2.8 million antibiotic-resistant illnesses occur in the United States (US) leading to more than 35,000 fatalities. Additionally, in 2017, at least 12,800 Americans died, and almost 223,900 needed hospital care for Clostridium difficile, a Gram-positive bacterium [11]. According to a World Health Organization (WHO) report in 2022, high levels of resistance in bacteria leading to life-threatening bloodstream infections as well as increasing resistance to treatment in several bacteria causing common infections is mind-boggling. This was contained in the Global Antimicrobial Resistance and Use Surveillance System (GLASS) based on data collected in 87 countries [12].
Nanoparticles (NPs) are defined as materials or particles having a size between 1 and 100 nm. Even though some metals such as silver or copper exhibit inhibitory action against bacteria in their bulk forms, other metals only have it as NPs. Induction of oxidative stress, non-oxidative mechanisms, and, in a minor way, the interaction of released metal ions with functional groups of proteins and nucleic acids are three processes that are hypothesized to occur concurrently, but those mechanisms of action have not yet been fully explained. Metal NP antibacterial action is specifically influenced by variables such as size, zeta potential (electrokinetic potential), charge, surface shape, and crystal structure. NPs have the ability to damage bacterial membranes and prevent the development of biofilms. Also, greater biofilm inhibition is provided by smaller NPs (e.g., Ag, ZnO, Mg, or NO NPs) [13,14,15].
The residues of the seafood industry, mainly crab shells, shrimp shells, krill shells, and fish scales, form important resources for the extraction of chitin and chitosan [16]. Snail shell waste is a bountiful source for chitosan production, and literature is still scanty on its use for the production, characterization, and antimicrobial activity of chitosan [17, 18]. In Nigeria, snails are mostly found in the southern part of the country with distinct species such as the African giant land snail (Archachatina marginata), tiger snail (Achatina achatina), land snail (Achatina fulica), Limcolana aurora sp., and other garden snails [19, 20]. Snails are available in large quantities during the rainy season, and after consumption, their shells are discarded indiscriminately leading to environmental menace. The presence of carbohydrate (86.83/100–92.76/100 g) in snail shell makes it suitable for the production of chitosan in substantial quantity when compared to seafood waste [21, 22].
Chitosan is produced either by enzymatic hydrolysis in the presence of a chitin deacetylase or (partial) deacetylation of chitin involving the removal of acetyl group, proteins, and minerals in the solid form using concentrated sodium hydroxide [23,24,25]. The degree of deacetylation (DD) in the range of 50–95%, the purity of the product, and the sequence of the amino and acetamido groups are factors that affect the physical characteristics of chitosan [26, 27]. Chitosan is an excellent antimicrobial substance against various microbial pathogens [28, 29]. The antimicrobial activity of chitosan is affected by its solubility, which in turn is affected by the elevated quantities of free amino groups in the chain. The fundamental change in its chemical structure, low molecular weight, water solubility, and the degree of deacetylation (DDA) complements the antimicrobial behavior of chitosan [30]. Further, chitosan is also applied as an activator of plant defenses [31], a supplement for food preservation and food additives [32], and a corrosion inhibitor [33].
To address the epidemic of antimicrobial resistance, it is vital to develop new classes of antibacterial drugs with nanoparticle properties from renewable sources such as shells of mollusks. Therefore, the purpose of this work was to synthesize chitosan from Archachatina marginata shell, an agricultural waste; characterize, and examine its antibacterial activity against clinical strains of Staphylococcus aureus (β-hemolytic), Staphylococcus saprophyticus DSM 18669, Escherichia coli 0157, Klebsiella pneumonia EO1 16S ribosomal RNA gene, and Salmonella typhi ATCC 13311.
2 Experimental
2.1 Materials
Sodium hydroxide, BDH (Poole, England); hydrochloric acid, Analar (Essex England); sodium hypochlorite, Shandong Retouch (China); acetic acid, Analar (Essex England); media and discs, Hi-Media (India); autoclave, Express Laboratories (USA); microtitre plates, Labsystems Oy (Helsink, Finland); McFarland standard, Sigma-Aldrich (UK); saline solution, Biomed Diagnostics (USA); Archachatina marginata shell obtained from Mushin Market Lagos, Nigeria. The chemicals were used as received.
2.2 Preparation of Chitosan
Modifications were made to the method outlined [34]. Using a magnetic stirrer and heating it at 80 °C with constant stirring for 6 h, we deproteinized 50 g powdered of Archachatina marginata shell. After this process, the residue was neutralized with deionized water and dried at 100 °C for 1 h. 1 M HCl solution was used to demineralize the dried residue while it was stirred at 30 °C for 3 h. The residue was then washed to a neutral state with deionized water and dried at 90 °C for 1 h. After 4 h of deacetylation in 12.5 M NaOH solution at 30 °C with continual stirring, the resulting chitin was washed and dried in the manner outlined above. The powder was neutralized by rinsing in double-distilled water until the pH was at 7. The powder was dried at 90 °C for 1 h after being decoloured by soaking in a 10% sodium hypochlorite solution at 25 °C for 4 h with constant stirring. A medium-density polythene packaging material was used to store the chitosan powder that was the product of the process. The process is illustrated in Scheme 1.
2.3 Characterization of Chitosan
2.3.1 Fourier Transform Infrared Spectroscopy
Functional groups were determined by Fourier transform infrared spectroscopy (FTIR) using a Perkin Elmer Spectrum 100 FT-IR spectrophotometer. KBr was used as the diluent and mixed with 1% chitosan powder by using an agate mortar. The mixture was compressed to obtain tablets, and scanning was carried out over the frequency of 4000 to 400 cm−1 at resolution of 4 cm−1.
2.3.2 X-Ray Diffractometry
Crystal definition and size were examined by X-ray diffractometer (ARL’XTRA Thermo Fisher Scientific Company Switzerland: serial number 197492086) operated at 30 kV and 100 mA. The pattern was recorded by Cu Kα radiation with λ = 1.5406 Å and nickel monochromator. Scanning was done at a scan rate of 2°/min with the scan angle from 10 to 90°. The size of chitosan crystallite was calculated using Scherer’s equation [35].
where D is the average crystallite size, K is the Scherer coefficient (0.89), λ is the X-Ray wavelength (λ – 1.5406 Å), Ɵ is Bragg’s angle (Ɵ), and β is the full width at maximum (FWHM) in radians.
2.3.3 Scanning Electron Microscopy and Transmission Electron Microscopy
Scanning electron microscopy (SEM) and transmission electron microscopy (TEM) were used to analyze cellular morphology, size, and shape (TEM). The Phenom-World PRO:X 800–07334 transmission microscope from Thermo Fisher Scientific Company in Switzerland, serial number MVE01570775, was utilized for the SEM examination. The sputter coater was used to provide a 25-nm gold coating to an aluminum stub that held the sample at 25 mm in diameter. The sample then adhered to the carbon disc. The samples were then scanned using a 5-kV accelerating voltage and a backscattered electron detector with a focused fine stream of secondary electrons.
Transmission electron microscopy examination was performed with a Tecnai G2 Spirit TWIN FEI, Netherlands, LaB6 microscope. A carbon/formvar support film-covered copper grid was used to hold the liquid sample. Extra material was wiped off a filter paper after 15 s. After 15 s, a drop of the negative stain (1% uranyl acetate) was applied and blotted. The specimen container containing the grid was then put into a Phillips/FEI CM 120 BioTwin TEM operating at 200 kV for imaging.
2.3.4 Energy-Dispersive X-Ray Fluorescence and Energy-Dispersive X-Ray Spectroscopy
Energy-dispersive X-ray fluorescence (EDXRF) (Thermo Scientific X-ray Fluorescence (XRF) Epsilon Spectrometer) was used to identify the elements present. After removing oxygen and moisture from the sample holders for 10 min using a vacuum pump, the samples were ready to be analyzed using the XRF spectrometer. At 200 kV, an X-ray micro-analyzer (Oxford 6587 INCA, Oxford Instruments, Abingdon, UK) was coupled to a transmission electron microscope (TEM) detector (Phillips/FEI CM 120 BioTwin) to perform energy-dispersive X-ray (EDX) analysis in scanning mode.
2.3.5 Degree of Deacetylation (DDA)
The degree of deacetylation (DDA) was calculated from FTIR data as reported [36].
A1629.85 and A3450.65 cm−1 indicate absolute heights of absorption bands of amide and hydroxyl groups, respectively, while 1.33 denotes the ratio of A1629.85 and A3450.65 cm−1 for fully N-acetylated chitosan.
2.4 Chemical Composition
Moisture content, ash, and protein content were determined using the procedure described [37].
2.5 Antimicrobial Assay
The antibacterial activity of chitosan was evaluated against Staphylococcus aureus (hemolytic and clinical strain), Staphylococcus saprophyticus DSM 18669, Escherichia coli 0157, Klebsiella pneumonia EO1 16S ribosomal RNA gene, and Salmonella typhi ATCC 13311. The strains were clinical isolates obtained from stock cultures at the Pharmaceutical Technology Department, Faculty of Pharmacy, University of Lagos, Idi-Araba, Lagos, Nigeria.
2.6 Preparation of Culture Media
Mueller–Hinton agar (Oxoid, England) was used as the growth medium for the microorganisms. Thirty-seven grams of the dehydrated bacteriological culture media was weighed and dissolved in distilled water (100 mL) according to the manufacturer’s specification. The resultant suspension was dispensed into a clean conical flask and autoclaved at 121 °C for 15 min and then poured into previously sterilized Petri dishes. The plates were allowed to cool and solidify. The molten agar gels formed were dispensed in 15 mL portions into Petri dishes for potency test. The cultures were all sub-cultured on nutrient agar and incubated at 37 °C for 24 h prior to use.
2.7 Antimicrobial Profile (Zone of Inhibition)
The agar diffusion technique was used for the antimicrobial activity [38]. Antimicrobial discs containing chitosan at a concentration of 40 mg/mL (0.2 g chitosan dissolved in 5 mL (1%) acetic acid) were dispensed with the aid of sterile forceps onto the potato dextrose agar (PDA) plates in such a way that the minimum center-to-center distance was 24 mm and the discs were no closer than 10 to 15 mm from the edge of the Petri dish. The following method was used to make standard solutions of 0.05, 0.025, 0.0125, and 0.0063 mg/mL of pure levofloxacin (Sigma-Aldrich, UK). Levofloxacin standard was prepared by dissolving 0.05 g in 10 mL of sterile distilled water (diluent), yielding a stock concentration of 5000 mg/mL solution from which a twofold dilution was performed. The bacterial strains were tested using levofloxacin as a positive control. After 24 h in the 37 °C incubator, the plates were examined. The zone reader was used to obtain measurements in triplicate, and the average zone value was then calculated in millimeters.
2.8 Determination of Minimum Inhibitory Concentration (MIC)
A broth microdilution assay was used for determining chitosan’s minimum inhibitory concentration (MIC) [39]. Standard Bioscreen C 100 well microtitre plates were used to dilute chitosan in a Giolitti and Cantoni broth (GCB) medium with a pH of 6. McFarland 0.5 turbidity standard was used to calibrate bacterial cell suspensions (2 × 108 cfu/mL). The test isolates were inoculated into microtitre plates containing 10 mL of prepared normal saline, and the plates were incubated at 37 °C for 24 h. After incubation, the suspension was further diluted in normal saline to generate a final inoculum 108 cfu/mL dilution of the microorganism; at this time, the concentration of the test microorganisms is approximately 1.5 × 108 cfu/mL. From a concentration of 40 mg/mL, chitosan was diluted twofold in sterile broth (20, 10, 5, 2.5, 1.25, 0.625, 0.3125, 0.1563, 0.0781, 0.0391, 0.0195, 0.0978 mg/mL) respectively.
2.9 Statistical Analysis
Data obtained from antimicrobial tests were analyzed using means and mean deviation (means of triplicate determinations), and a test of significance was carried out using ANOVA (GraphPad Software Inc., San Diego, USA).
3 Results and Discussion
3.1 Fourier Transform Infrared Spectroscopy (FTIR)
The FTIR spectrum (Fig. 1) shows absorption bands at 3418 cm−1 (OH stretching), 2967 cm−1 (CH3 stretching), 2517 cm−1 (CH2 stretch), 1721 cm−1 (carbonyl stretch), 1637 cm−1 (NH primary amine), 1346 cm−1 (C-O starch in the primary alcoholic group), 1198 cm−1 (free amino group (NH2) at glucosamine C2 position), 1123 cm−1 (C–O–C stretching), and 1066 cm−1. These obtained vibration patterns served as an indicator of the presence of elements such as carbon, nitrogen, and oxygen which are the primary constituents of chitosan. This is similar to the result obtained from rose shrimp (Parapenaeus longirostris), crab (Callinectes amnicola), and shrimp (Penaeus notialis) shell wastes [40,41,42,43]. Deacetylation can be identified by the appearance of broad and sharp wavenumbers at 1743, 1633, and 1598 cm−1 [44].
3.2 X-Ray Diffractometry (XRD)
The X-ray diffraction patterns of chitosan showed a total of 29 peaks out of which 11 are prominent (Fig. 2). The prominent peaks are 23.04°, 26.21°, 29.41°, 33.13°, 36.13°, 37.95°, 41.18°, 43.13°, 45.18°, 49.89°, 50.23°, and 52.44° assigned to 111, 111, 111, 200, 200, 200, 220, 220, 220, 311, 311, and 311 respectively. This is based on the Bragg reflection reference of the International Centre Diffraction Data (ICDD) database on crystalline and amorphous materials. The XRD referencing of chitosan in this study was also based on the findings presented in a previous study [45,46,47]. The two sharp peaks at 29.41° and 36.13° denote the presence of calcite and aragonite mineral substance [48]. Calcium is the predominant element in both minerals hence the characteristic peak which is sharp and broad [49, 50]. The presence of the sharp peaks is an evidence of the crystalline structure of chitosan [51]. Also, the average crystallite size of 22.47 nm calculated using Scherer’s equation suggests the formation of a nanoparticle. In comparison with commercial chitosan obtained from crustacean shells, findings from this study differ; 11.7°, 20.02°, and 20.04° were reported [16, 52]. This might be attributed to the difference in mineral composition of chitosan (Table 1). The XRD patterns of chitosan obtained in this study match closely with those of chitosan extracted from crab shell [53], chitosan extracted from Pinna bicolor pen shell [54], chitosan obtained from Rhizopus oryzae NRRL [55], chitosan extracted from green mussel shells [56], chitosan synthesized from crab shell (Scylla sp.) marine shrimps (Penaeus semisulcatus), and horn snail shell (Telescopium telescopium) [57, 58].
3.3 Energy-Dispersive X-Ray Fluorescence (EDXRF)
A significant signal was observed at 3.60 keV, which is a typical binding energy of calcium [43, 59]. Strontium, silver, nickel, copper, tin, and zinc also showed less prominent peaks (Fig. 3). The elemental composition of commercial chitosan obtained from the shells of crab is consistent with the findings from this study, which include the detection of Ca, Y, Ag, Nb, K, Na, Mg, and Al [60]. The detection of alkaline metals, alkaline earth metals, rare earth metals, and transition metals may be due to the mineral composition of gastropod and snail shells [61, 62]. However, the lack of K and L lines in the equipment prevented the detection of additional components such as carbon, hydrogen, and oxygen in chitosan.
3.4 Energy-Dispersive X-Ray Spectroscopy (EDS)
Carbon, oxygen, calcium, copper, silicon, and magnesium were all detected as shown in Fig. 4 with calcium being the element with the strongest peak thus confirming the composition of chitosan [18]. The lack of sulfur and phosphorus was indicative of the efficacy of the chemical treatment involved in the synthesis of chitosan from A. marginata shell. This is consistent with results obtained from chitosan synthesized from ruhi (Labeo rohita) scales Callinectes amnicola, Penaeus notialis, and shrimp (Penaeus monodon) [43, 53, 63].
3.5 Scanning Electron Microscopy (SEM)
Images of chitosan at three different magnifications in order to understand the surface shape and particle dispersion are shown in Fig. 5. × 1500 (a) and × 1000 (b) show rough, grainy surface with clustering of uneven particles [25, 47, 58], while × 300 (c) reveals particles that were polygonal, rod-like, and circular without agglomeration with crystalline feature [49]. Comparable morphological details were similar to the work by [18]; however, it was at a magnification level 100 times higher than that employed in the present work. Image processing application, ImageJ, was used to quantify chitosan particle size. The measured sizes of the particles collected were between 11.06 and 34.37 nm. This confirms the nanometric nature of chitosan by showing that its constituent particles interacted strongly due to their high surface energy. Nanoparticles with high dispersion offer a myriad of advantages across various sectors. In healthcare, nanotechnology plays a pivotal role in advancing diagnostic techniques, enabling precise pharmaceutical administration, and facilitating the development of biomedical implants [64]. This unique characteristic of high dispersion sets nanoparticles apart from other substances, making them valuable resources in industries ranging from healthcare to technology [65].
3.6 Transmission Electron Microscopy (TEM)
Images of chitosan at two different scales to understand shape, particle size, and distribution are shown in Fig. 6. Five hundred nanometers (a) shows chitosan surface as dense, well organized with few gaps in between consisting of pseudo-spherical particles and randomly spread as flakes [43]. Two hundred nanometers (b) indicates polygonal particles, rod-like structures, and spherical voids on the surface [49]. These micrographs are similar to results obtained from Callinectes amnicola, Penaeus notialis, and Penaeus semisulcatus shell wastes [43, 57]. The abundance of pores (shown as dark spots in both images) and their close spatial proximity along the microstructure are evidence of nanometric particles exhibited by chitosan. The chitosan particle sizes were measured using the ImageJ software from the images. The sizes varied from 5.80 to 31.05 nm. SEM image results are verified by the TEM images.
3.7 Degree of Deacetylation (DDA)
The FTIR data indicated that 75.39% of the chitosan was deacetylated, which was lower than the value reported [18]; however, it was closer to 76.70% [66] and 73.6% [67]. The values obtained in this study are comparable to chitosan synthesized from shrimp (Macrobrachium jelskii) shell, fungal biomass, Litopenaeus vannamei, and pink shrimp (Penaeus brasiliensis) [68,69,70]. In practice, DDA values between 30 and 95% are suitable for various applications [71]. The degree of deacetylation of chitosan is often determined by FTIR spectroscopy. Deacetylation degree (DDA) is the mole fraction of deacetylated units in the polymer chain and has a significant impact on chitosan characteristics such as chemical reactivity, biodegradability, and biocompatibility. Deacetylation is the process of removing acetyl groups from a chitin molecular chain, leaving behind a full amino group (-NH2) with acetyl residue in smaller proportion; the specifics of this process rely on the sample preparation and analytical techniques utilized [72, 73]. Overall, the physical, chemical, and biological characteristics of chitosan are influenced by the source of the chitin derivative, the time, temperature, and alkaline concentration utilized during the extraction process [53].
3.8 Chemical Composition
The chemical components of chitosan are shown in Table 2. The percentages of moisture, ash, and protein are as follows: 2.21% [74], 6.05% [75], and [67]. In respect to moisture content, a lower value indicates higher self-stability and quality [76]. Generally, the allowable range of moisture content in chitosan for multiple uses is 5–15% [77].
This research found a greater ash content than reported works [18, 66]. This is affected by the concentration of acid, reaction duration, and operating temperature as they relate to the demineralization process’ overall scope. Ash levels below 1% in chitosan are recommended for use in biomedical applications because of their beneficial effects on cell viability and other biological processes [78]. Yet, chitosan with an ash concentration of more than 1% has been employed in a number of biological applications [79, 80].
The high protein content of chitosan in this study is connected with the level of deacetylation, which converted acetyl groups into amino groups [18, 66,67,68,69,70].
In this study, the chemical composition has indicated that chitosan was successfully synthesized from Archachatina marginata shell and comparable in values with chitosan from crab, shrimp, and mussel shells.
3.9 Antimicrobial Evaluation
3.9.1 Zone of Inhibition
Chitosan exhibited different degrees of activities against both Gram-positive and Gram-negative species based on the observed zones of inhibition which ranged from 23.67 to 39.00 mm comparable with levofloxacin (Table 3). Figure 7 shows the response of S. aureus (hemolytic), K. pneumonia strain E01 16S ribosomal RNA gene, and E. coli strain 0157 to chitosan except S. typhi ATCC 13311, while Fig. 8 shows the response of levofloxacin to these isolates. In this study, chitosan activity against these isolates is comparable with commercial chitosan, chitosan extracted from shrimp (Penaeus semisulcatus) shell, horse mussel (Modiolus modiolus) shell, and prawn (Macrobrachium lamarrei) shell [81,82,83,84].
Chitosan’s efficacy against some of these isolates was higher than that of the levofloxacin, a standard drug. The zone of inhibition of chitosan against S. saprophyticus DSM 18669 was higher (39.00 mm) than levofloxacin (31.33 mm at the highest concentration of 0.05 and 18.33 mm at the lowest concentration of 0.0063 mg/mL). Also, levofloxacin at a concentration of 0.0063 mg/mL did not show inhibitory activity against S. aureus (hemolytic) and E. coli strain 0157. The zone of inhibition of chitosan was statistically different from levofloxacin at 0.05, 0.025, and 0.0125 mg/mL against S. aureus, S. saprophyticus DSM 18669, E. coli, and K. pneumonia. This implies that levofloxacin would require concentrations greater than 0.05 mg/mL to have the same effect as chitosan against S. aureus (hemolytic) and S. saprophyticus DSM 18669, while chitosan would require concentrations above 40 mg/mL to have the same effect as levofloxacin at concentrations of 0.05 and 0.025 mg/mL against E. coli strain 0157 and K. pneumonia E01 16S ribosomal RNA gene. The non-inhibitory effect of chitosan on S. typhi ATCC 13311 implies that a concentration above 40 mg/mL would be required possibly due to plasmic composition of the isolate among other factors.
Chitosan was not assessed at various concentrations owing to its status as a novel product, allowing for the arbitrary selection of concentration levels as long as they demonstrate biological efficacy against the targeted organisms. An inhibitory effect and noticeable impact on the strains were observed at a concentration of 40 mg/mL. There exists no definitive guideline concerning the utilization of a singular concentration within antimicrobial testing protocols. Studies conducted by authors provided empirical evidence supporting the efficacy of a singular concentration when combating various strains or isolates of microorganisms [85,86,87,88].
Regarding the mechanism of chitosan antibacterial activity, two mechanisms are proposed, viz direct interaction of chitosan nanoparticles which increases bacteria membrane permeability and inhibits their growth or interference with anionic channels due to negatively charged thinner layer of peptidoglycan cell wall and lipopolysaccharides [89, 90]. The second mechanism occurs due to the binding of chitosan occasioned by its high surface area to volume ratio, binding to the DNA causing disruption of mRNA generation in bacterial cells. This causes membrane disruption and ultimately death of the bacteria cells by allowing internal components to seep out [91,92,93].
Results in this study showed that chitosan nanoparticle was more effective against Gram-positive bacteria than Gram-negative bacteria [91]. In spite of this, some research suggests that chitosan is more effective against Gram-negative bacteria than it is against Gram-positive bacteria [94]. There is no indication that Gram-positive bacteria are more resistant than Gram-negative bacteria. Particle size, culture population, pH, natural organic matter, and ionic strength are all factors that may alter chitosan’s efficacy as an antibacterial agent [89].
Largely, the activity of chitosan in this study compares favorably with functionalized grapheme by azo‑pyridinium compounds [95], triazole-tethered derivatives of curcumin [96], glycoside‑ and acyclic nucleoside–based 6‑cyclohexyl‑4‑aryl‑2‑oxonicotinonitrile [4], thiochromene compounds [97], and pyrazole bearing biologically active imidazolyl, pyridine, and quinoxaline derivatives [98].
3.9.2 Minimum Inhibitory Concentration
Bacterial isolates were subjected to MIC testing with chitosan concentrations ranging from 20 to 0.0098 mg/mL. Table 4 displays the minimum inhibitory concentrations (MICs) of chitosan against the isolates. The MIC of chitosan against the bacterial isolates was between 0.3125 and 0.0.0781 mg/mL. MIC of an antibacterial agent is the lowest concentration shown to inhibit the growth of germs when tested by visual inspection, a reader, or absorbance measurement at 660 nm [99]. Clear microtitre plates show no growth, whereas muddy wells suggest resistance by the isolates; hence in this study, the MIC was determined by visual examination (macroscopic assessment). Even at the lowest doses of 0.0391, 0.0195, and 0.0098 mg/mL, the plates were turbid, showing that the isolates were resistant to the chitosan. Nonetheless, the wells became clearer at concentrations of 20, 10, 5, 2.5, 1.25, 0.625, 0.3125, 0.1563, and 0.0781 mg/mL, all of which corresponded to the MIC values.
The variation in the antimicrobial efficacy of chitosan nanoparticles against Gram-positive and Gram-negative bacteria, as observed in agar diffusion experiments compared to MIC evaluations, can be attributed to multiple factors related to the composition of bacterial cell walls, nanoparticle properties, and assay methodologies [100, 101].
Gram-positive bacteria are characterized by the presence of a substantial peptidoglycan layer within their cellular envelopes, rendering them susceptible to interference from nanoparticles. The existence of this layer offers a larger surface area for nanoparticles to engage with, resulting in a more notable inhibitory impact in agar diffusion assays. On the other hand, Gram-negative bacteria have a peptidoglycan layer that is thinner and is surrounded by an outer membrane made up of lipopolysaccharides (LPS) and proteins. This external membrane functions as a barricade, constraining the entry and effectiveness of nanoparticles, consequently diminishing the inhibitory effects observed in agar diffusion tests [102].
The dimensions, morphology, electrostatic potential, and constitution of nanoparticles can collectively affect their interactions with bacterial cells [88]. Certain nanoparticles might exhibit particular preferences for distinct cell wall constituents, chemical binding with functional membrane proteins, and adsorption of the NP responsible for mechanical deformation, leading to cell rupture and death or demonstrate enhanced permeation across specific bacterial lipid bilayers [103, 104]. During trials involving agar diffusion, chitosan nanoparticles were placed on the surface of the agar medium, where their interaction with bacterial cells primarily took place at the interfacial boundary. The magnitude and electrostatic nature significantly influence the diffusion kinetics and efficacy of nanoparticles. During MIC evaluations, nanoparticles are frequently dispersed in aqueous nutrient broths to improve their proximity and interaction with bacterial cells. Therefore, a decrease in the required nanoparticle concentration to hinder bacterial proliferation might be accomplished, particularly concerning Gram-negative strains, which may demonstrate heightened susceptibility to distinct properties of chitosan nanoparticles in solution [105, 106].
Agar diffusion tests and MIC assays are two methodologies employed for assessing the antibacterial efficacy of nanoparticles. The agar diffusion tests offer a qualitative evaluation through the measurement of the zone of inhibition surrounding nanoparticles contained in a disc or well. This technique primarily demonstrates the capability of nanoparticles to engage with bacteria at the surface. In contrast, MIC assays yield a quantitative assessment by establishing the minimum nanoparticle concentration needed to completely hinder bacterial proliferation. This approach considers several variables including nanoparticle concentration, exposure duration, and bacterial growth dynamics to furnish a more precise and reliable evaluation of the effectiveness of antibacterial agents [107].
The increased suppression of Gram-positive microorganisms noted in agar diffusion experiments could be attributed to the increased thickness of the peptidoglycan layer found in their cellular membranes, thus providing a more extensive area for interaction with nanoparticles. On the contrary, in MIC assays, a decreased amount of nanoparticles may be adequate to impede the growth of Gram-negative bacteria, influenced by various factors like nanoparticle characteristics and testing techniques that facilitate enhanced interaction and penetration into bacterial cells.
4 Conclusion
Chitosan from A. marginata shell possesses similar properties with chitosan obtained from other sources such as shrimp, crayfish, and crab shells. The nanoparticle nature of the chitosan was confirmed by FTIR, SEM, TEM, XRD, EDXRF, and EDS. Evaluation of potency against Staphylococcus aureus (β-hemolytic), Staphylococcus saprophyticus DSM 18669, Escherichia coli strain 0157, and Klebsiella pneumonia strain EO1 16S ribosomal RNA gene was remarkable except Salmonella typhi ATCC 13311. The antibacterial activity of chitosan compared favorably with levofloxacin appearing to have a broad-spectrum activity. Also, Gram-positive bacteria were more susceptible than Gram-negative bacteria. The antibacterial activity of chitosan from A. marginata makes it a potential green material against microorganisms that are developing resistance against antibiotic drugs.
Data Availability
Data used for this study and other supplementary information are available on reasonable request.
References
Livermore, D. M. (2009). Has the era of untreatable infections arrived? Journal of Antimicrobial Chemotherapy, 64(Suppl. 1), i29-36.
Garrido-Maestu, A., Ma, Z., Chen, N., Ko, S., Tong, Z., & Jeong, K. C. (2018). Engineering of chitosan-derived nanoparticles to enhance antimicrobial activity against foodborne pathogen Escherichia coli O157: H7. Carbohydrate Polymers, 197, 623–630.
De Kracker, M. E. A., Davey, P. G., Grundmann, H., BURDEN Study Group. (2011). Mortality and hospital stay associated with resistant Staphylococcus aureus and Escherichia coli bacteremia: Estimating the burden of antibiotic resistance in Europe. PLoS Medicine, 8, 1001104.
Hassan, A. E., Abou-elkhair, R. A. I., Mohamed, S. M., Moustafa, A. H., Fikry, R. M., & Yassen, F. A. (2019). Glycoside- and acyclic nucleoside-based 6-cyclohexyl-4-aryl-2-oxonicotinonitrile: Synthesis and antimicrobial evaluation. Journal of the Iranian Chemical Society, 16, 1891–1902. https://doi.org/10.1007/s13738-019-01662-x
Davies, J., & Davies, D. D. (2010). Origins and evolution of antibiotic resistance. Microbiology and Molecular Biology Reviews, 74, 417–33.
Rolain, J. M., Canton, R., & Cornaglia, G. (2012). Emergence of antibiotic resistance: Need for a new paradigm. Clinical Microbiology and Infection, 18, 615–616.
Cars, O., Hedin, A., & Heddini, A. (2011). The global need for effective antibiotics—Moving towards concerted action. Dru Res Up, 14, 68–69.
Theuretzbacher, U. (2012). Accelerating resistance, inadequate antibacterial drug pipelines and international responses. International Journal of Antimicrobial, 39, 295–9.
Carlet, J., Jarlier, V., Harbarth, S., Voss, A., Goossens, H., & Pittet, D. (2012). Ready for a world without antibiotics? The pensieres antibiotic resistance call to action. Antimicrobial Resistance and Infection Control, 1, 11. https://doi.org/10.1186/2047-2994-1-11
IDA (Infectious Diseases Society of America). (2010). The 10 _ ‘20 initiative: Pursuing a global commitment to develop 10 new antibacterial drugs by 2020. Clinical Infectious Diseases, 50, 1081–1083.
Antibiotic Resistance Threats in the United States CDCs (Centre for Disease Control) AR Threats Report (2019). https://www.cdc.gov/drugresistance/pdf/ar-threats-2013-508. Accessed 26 March 2022.
The Global Antimicrobial Resistance and Use Surveillance System (GLASS) Report World Health Organization (WHO) (2022). https://www.who.int/news/item/09-12-2022-report-signals-increasing-resistance-to-antibiotics-in-bacterial-infections-in-humans-and-need-for-better-datatibiotics-i. Accessed 7 July 2023.
Zamborini, F. P., Bao, L., & Dasari, R. (2012). Nanoparticles in measurement science. Analytical Chemistry, 84, 541–576.
Webster TJ, Seil I (2012) Antimicrobial applications of nanotechnology: methods and literature. International Journal of Nanomedicine, 7:2767. https://doi.org/10.2147/IJN.S24805
Wang, L., Hu, C., & Shao, L. (2017). The antimicrobial activity of nanoparticles: Present situation and prospects for the future. International Journal of Nanomedicine, 12, 1227–1249.
Varma, R. (2020). Vasudevan S (2020) Extraction, characterization, and antimicrobial activity of chitosan from horse mussel Modiolus modiolus. ACS Omega, 5, 20224–20230.
Johney, J., & Eagappan, K. (2016). Ragunathan RR (2016) Microbial extraction of chitin and chitosan from Pleurotus spp, its characterization and antimicrobial activity. International Journal of Current Pharmaceutical Research, 9, 88–93.
Bello, V. E., & Olafadehan, O. A. (2021). Comparative investigation of RSM and ANN for multi-response modelling and optimization studies of derived chitosan from Archachatina marginata shell. Alexandria Engineering Journal, 60, 3869–3899.
Sogbesan, O. A., & Ugwumba, A. A. A. (2012). Culture and nutrient values of Limicolaria aurora Jay Mollusca Achatinidae raised in two different substrates. Revista Cientı´fica UDO Agrı´cola, 12, 479–485.
Igbinosa, I. B., Isaac, C., Adamu, H. O., & Adeleke, G. (2016). Parasites of edible land snails in Edo State, Nigeria. Helminthologia, 53, 331–335. https://doi.org/10.1515/helmin-2016-003
Agoha, E. E. C., Mazi, E. A. (2009). Biopolymers from African giant snail shells waste: Isolation and characterization. In: Do¨ ssel, O., Schlegel, W.C. (eds) World Congress on Medical Physics and Biomedical Engineering, September 7–12, Munich, Germany. IFMBE Proceedings 25:249–251. https://doi.org/10.1007/978-3-642-03900-3_71
Ademolu, K. O., Akintola, M. Y., Olalonye, A. O., & Adelabu, B. A. (2018). Traditional utilization and biochemical composition of six mollusc shells in Nigeria. Revista de Biologia Tropical, 63, 459–464.
Houndonougbo, M. F., Chrysostome, C. A. A. M., Odoulami, R. C., & Codjia, J. T. C. (2012). Snail shell as an efficient mineral feedstuff for layer hens: Effects and Optimum Rate. Livestock Research for Rural Development, 24, 162.
Zargar, V., Asghari, M., & Dashti, A. (2015). A review on chitin and chitosan polymers: Structure, chemistry, solubility, derivatives and applications. ChemBioEng, 2(00), 1–24.
Oyekunle, D. T., & Omoleye, J. A. (2019). New process for synthesizing chitosan from snail shells. Journal of Physics: Conference Series, 1299, 012089.
Muzzarelli, R. A. A. (1990). Modified chitosans and their chemical behavior Polym. Prepr. Am. Chem. Soc., Div. Polym. Chem. 31(1):626. https://doi.org/10.1002/chin.199111346
Campana Filho, S., De Britto, D., Curti, E., Abreu, F., Cardoso, M., & Battisti, M. (2007). Extraction, structures and properties of α- and β-chitin. Quimica Nova, 300, 644–650.
Friedman, M., & Juneja, V. K. (2010). Review of antimicrobial and antioxidative activities of chitosans in food. Journal of Food Protection, 73, 1737–1761.
Kong, M., Chen, X. G., Xing, K., & Park, H. J. (2010). Antimicrobial properties of chitosan and mode of action: A state-of-the-art review. International Journal of Food Microbiology, 144(1), 51–63. https://doi.org/10.1016/jijfoodmicro.2010.09.012
Qin, C., Du, Y., Xiao, L., Li, Z., & Gao, X. (2002). Enzymic preparation of water-soluble chitosan and their antitumor activity. International Journal of Biological Macromolecules, 31, 111–117.
Cota-Arriola, O., Cortez-Rocha, M. O., Burgos-Hernandez, A., Ezquerra-Brauer, J. M., & Plascencia-Jatomea, M. (2013). Controlled release matrices and micro/nanoparticles of chitosan with antimicrobial potential: Development of new strategies for microbial control in agriculture. Journal of the Science of Food and Agriculture, 93, 1525–1536.
Morin-Crini, N., Lichtfouse, E., Torri, G., & Crini, G. (2019). Applications of chitosan in food, pharmaceuticals, medicine, cosmetics, agriculture, textiles, pulp and paper, Biotechnology and Environmental Chemistry. Environmental Chemistry Letters, 17, 1667–1692. https://doi.org/10.1007/s10311-019-00904-x
Okoronkwo, A. E., Olusegun, S. J., & Oluwasina, O. O. (2015). The inhibitive action of chitosan extracted from Archachatina marginata shells on the corrosion of plain carbon steel in acid media. Anti-Corrosion Methods and Materials, 62(1), 13–18. https://doi.org/10.1108/ACMM-10-2013-1307
Kewboonruang, S., Phatrabuddha, N., Sawangwong, P., & Pitaksanurat, S. (2016). Comparative studies on the extraction of chitin – Chitosan from golden apple snail shells at the control field. IOSR Journal of Polymer and Textile Engineering, 3, 34–41. https://doi.org/10.9790/019X-03013441
Kaviya, S., Santhanalakshmi, J., Viswanathan, B., Muthumary, J., & Srinivasan, K. (2011). Biosynthesis of silver nanoparticles using Citrus sinensis peel extract and its antibacterial activity.Spectrochimica Acta Part A. Molecular and Biomolecular Spectroscopy, 79, 594–8. https://doi.org/10.1016/j.saa.2011.03.040
Mohanasrinivasan, V., Mishra, M., Paliwal, J. S., Singh, S. K., Selvarajan, E., Suganthi, V., & Subathra, D. C. (2014). Studies on heavy metal removal efficiency and antibacterial activity of chitosan prepared from shrimp shell waste. Biotech, 3(4), 167–75. https://doi.org/10.1007/s13205-013-0140-6
AOAC. (2019). Association of Official Analytical Chemists. Official Methods of Analysis 21st Edition International, Gaithersburg, Maryland USA
Holubnycha, V., Kalinkevich, O., Ivashchenko, O., & Pogorielov, M. (2018). Antibacterial activity of in situ prepared chitosan/silver nanoparticles solution against methicillin-resistant strains of Staphylococcus aureus. Nanoscale Research Letters, 13, 71.
Limam, Z., Selmi, S., Sadok, S., & El Abed, A. (2011). Extraction and characterization of chitin and chitosan from crustacean by-products: Biological and physicochemical properties. African Journal of Biotechnology, 10(4), 640–647. https://doi.org/10.5897/AJB10.209
Dahmane, E. M., Taourirte, M., Eladlani, N., & Rhazi, M. (2014). Extraction and characterization of chitin and chitosan from Parapenaeus longirostris from Moroccan local sources. International Journal of Polymer Analysis and Characterization, 19, 342–351. https://doi.org/10.1080/1023666X.2014.902577
Kaya, M., Baran, T., & Karaarslan, M. A. (2015). New method for fast chitin extraction from shells of crab, crayfish and shrimp. Natural Product Research, 29, 1477–1480.
Antonino, R. S. C. M. Q., Fook, B. R. P. L., Lima, V. A. O. L., Rached, R. F., Lima, E. P. N., Lima, R. J. S., Covas, C. A. P., & Fook, M. V. L. (2017). Preparation and characterization of chitosan obtained from shells of shrimp (Litopenaeus vannamei Boone). Marine Drugs, 15, 141.
Olafadehan, O. A., Amoo, K. O., Ajayi, T. O., & Bello, V. E. (2021). Extraction and characterization of chitin and chitosan from Callinectes amnicola and Penaeus notialis shell wastes. Journal of Chemical Engineering and Materials Science, 12(1), 1–30. https://doi.org/10.5897/JCEMS2020.0353
Majekodunmi, S. O., Olorunsola, E. O., & Uzoaganobi, C. C. (2017). Comparative physicochemical characterization of chitosan from shells of two bivalve mollusks from two different continents. American Journal of Polymer Science, 7, 15–22. https://doi.org/10.5923/j.ajps.20170701.03
Ogawa, K., Oka, K., Miyanishi, T., Hirano, S. (1984). X-ray diffraction study on chitosan-metal complexes. https://doi.org/10.1016/B978-0-12-780950-2.50026-5
Arukalam, I. O., Timothy, U. J., Madu, I. O., & Achor, J. O. (2021). Improving the water barrier and anticorrosion performances of epoxy-chitosan coatings via silane modification. Journal of Bio- and Tribo-Corrosion, 7, 85. https://doi.org/10.1007/s40735-021-00512-9
Kahdestani, S. A., Mehrnoosh Hasan Shahriari, M. H., & Abdouss, M. (2020). Synthesis and characterization of chitosan nanoparticles containing teicoplanin using sol–gel. Polymer Bulletin. https://doi.org/10.1007/s00289-020-03134-2
Kumari, S., Rath, P., & Kumar, A. S. H. (2016). Chitosan from shrimp shell (Crangon crangon) and fish scales (Labeorohita): Extraction and characterization. African Journal of Biotechnology, 15, 1258–1268.
Bamiro, O. A., Salisu, A. O., Iyere, E. M., Olatundun, A., Adeleye, O. A., & Bakre, L. G. (2021). Characterization of chitosan obtained from snail shell (Achatina fulica) and its excipient potentials in metronidazole tablet formulation. Dhaka University Journal of Pharmaceutical Sciences, 20(1), 31–39. https://doi.org/10.3329/dujps.v20i1.50855
Ahmed, H. Y., Safwat, N., Shehata, R., Althubaiti, E. H., Kareem, S., Atef, A., Qari, S. H., Aljahani, A. H., Al-Meshal, A. S., Youssef, M., & Rokayya, S. R. (2022). Synthesis of natural nano-hydroxyapatite from snail shells and its biological activity: Antimicrobial, antibiofilm and biocompatibility. Membranes, 12, 408. https://doi.org/10.3390/membranes12040408
Rasti, H., Parivar, K., Baharara, J., Iranshahi, M., & Namvar, F. (2017). Chitin from the mollusc chiton: Extraction, characterization and chitosan preparation. Iranian Journal of Pharmaceutical Research: IJPR, 16, 366–379.
Govindan, S., Nivethaa, E. A. K., Saravanan, R., Narayanan, V., & Stephen, A. (2012). Synthesis and characterization of chitosan–silver nanocomposite. Applied Nanoscience, 2, 299–303. https://doi.org/10.1007/s13204-012-0109-5
Kumari, S., Kumar, A. S. H., Abanti, S., & Kumar, R. P. (2017). Physicochemical properties and characterization of chitosan synthesized from fish scales, crab and shrimp shells. International Journal of Biological Macromolecules, 104, 1697–1705. https://doi.org/10.1016/j.ijbiomac.2017.04.119[CrossRef]
Sudatta, B. P., Sugumar, V., Varma, R., & Nigariga, P. (2020). Extraction, characterization and antimicrobial activity of chitosan from pen shell, Pinna bicolor. International Journal of Biological Macromolecules, 163, 423–430.
Sebastian, J., Rouissi, T., Brar, S. K., Hegde, K., & Verma, M. (2019). Microwave-assisted extraction of chitosan from Rhizopus oryzae NRRL 1526 biomass. Carbohydrate Polymers, 219, 431–440.
Kaewprachu, P., & Jaisan, C. (2023). Physicochemical properties of chitosan from green mussel shells (Perna viridis): A comparative study. Polymers, 15, 2816. https://doi.org/10.3390/polym15132816
Thamilarasan, V., Sethuraman, V., Gopinath, K., Balalakshmi, C., Govindarajan, M., Mothana, R. A., Siddiqui, N. A., Khaled, M., & Benelli, G. (2018). Single step fabrication of chitosan nanocrystals using Penaeus semisulcatus: Potential as new insecticides, antimicrobials and plant growth promoters. Journal of Cluster Science, 29, 375–384. https://doi.org/10.1007/s10876-018-1342-1
Rajathy, T. J., Srinivasan, M., & Mohanraj, T. (2021). Physicochemical and functional characterization of chitosan from horn snail gastropod Telescopium telescopium. Journal of Applied Pharmaceutical Science, 11(2), 52–58. https://doi.org/10.7324/JAPS.2021.110207
Sundalian, M., Husein, S. G., Karlina, K., & Putri, D. (2022). Analysis and benefit of shells content of freshwater and land snails from gastropods class. Biointerface Research in Applied Chemistry, 12(1), 508–517. https://doi.org/10.33263/BRIAC121.508517
Sumaila, A., Ndamitso, M. M., Iyaka, Y. A., Abdulkareem, A. S., Tijani, J. O., & Idris, M. O. (2020). Extraction and characterization of chitosan from crab shells: Kinetic and thermodynamic studies of arsenic and copper adsorption from electroplating wastewater. Iraqi Journal of Science, 61(9), 2156–2171. https://doi.org/10.24996/ijs.2020.61.9.2
Angeli, J. L. F., Sartoretto, J. R., Kim, B. S. M., de Lima Ferreira, P. A., de Mahiques, M. M., & Figueira, R. C. L. (2021). Trace element fluxes during the “anthropocene” in a large South American industrial and port area (Santos and Sao Vicente Estuarine System, SE, Brazil). Environmental Monitoring and Assessment, 193, 594. https://doi.org/10.1007/s10661-021-09378-3
Ochi, D. O., & Babayemi, A. K. (2023). The physico-chemical properties and sorption potentials of snail shell particulates, chitin, chitosan, and oxalic acid modified chitosan from Achatina fulica shell. European Journal of Sustainable Development Research, 7(4), em0232. https://doi.org/10.29333/ejosdr/13476
Rahman, M. M., Maniruzzaman, M. (2023). Extraction, characterization and microstructure analysis of chitin and chitosan from shrimp (Penaeus monodon) waste shell. https://doi.org/10.2139/ssrn.4441334
Haruna, M. A., Magami, S. M. (2022). Nanoparticle dispersions for engineering application. 369–405. https://doi.org/10.1201/9781003280293-11
Blinova, A. V., & Rumyantsev, V. A. (2020). Nanotechnologies as the reality of modern dentistry (literature review). Endodontics Today, 18(2), 56–61. https://doi.org/10.36377/1683-2981-2020-18-2-56-61
Oyedeko, K. F. K., Akinyanju, A. S. A., Akinbomi, J., Lasisi, M. K., & Adesina, A. A. (2019). Preparation and characterization of synthetic chitosan using bio-waste (Snail Shell). Engineering & Technology Research Journal, 4(2), 38–46.
Muñoz, G., Valencia, C., Valderruten, N., & Ruiz-Durántez, E. (2015). Zuluaga F (2015) Extraction of chitosan from Aspergillus niger mycelium and synthesis of hydrogels for controlled release of betahistine. Reactive & Functional Polymers, 91, 1–10. https://doi.org/10.1016/j.reactfunctpolym.2015.03.008
Hennig EL (2009) Use of chitosan obtained from shrimp waste to evaluate the adsorption capacity of Fe3+ ions. M.Sc. Thesis, chemical technology and environmental, federal university of rio grande (FURG) Rio Grande, Brazil
Santos, M. C., Cirilo Oliveira, A. N., & Nunes, M. L. (2011). Determination of degree of deacetylation of chitosan obtained from shrimp saburica (Macrobrachium jelskii). Scientia Plena, 1877, 7–9.
Vilar Junior, J. C., Ribeaux, D. R., Alves da Silva, C. A., & De Campos-Takaki, G. M. (2016). Physicochemical and antibacterial properties of chitosan extracted from waste shrimp shells. International Journal of Microbiology, 2016, 5127515. https://doi.org/10.1155/2016/5127515
Sarbon, N. M., Sandanamsamy, S., Kamaruzaman, S. F. S., & Ahmad, F. (2014). Chitosan extracted from mud crab (Scylla olivicea) shells: Physicochemical and antioxidant properties. Journal of Food Science and Technology, 52(7), 4266–4275.
Khan, W. M., Prithiviraj, B., & Smith, D. L. (2002). Effect of foliar application of chitin and chitosan oligosaccharides on photosynthesis of maize and soybean. Photosynthetica, 40, 621–624. https://doi.org/10.1023/A:1024320606812
Boukhlifi, F. (2020). Modern spectroscopic techniques and applications quantitative analysis by IR: Determination of chitin/chitosan. InTechOpen, pp 1–24. https://doi.org/10.5772/intechopen.89708
Unuabonah, E. I., Adewuyi, A., Kolawole, M. O., Omorogie, M. O., Olatunde, O. C., Fayemi, S. O., Günter, C., Okoli, C. P., Agunbiade, F. O., & Taubert, A. (2017). Disinfection of water with new chitosan-modified hybrid clay composite adsorbent. Heliyon, 3(8), 2–30. https://doi.org/10.1016/j.heliyon.2017.e00379
Akinlabi, A. K., Mosaku, A. M., Adeogun, A. I., Falomo, A. A., Oladipo, G., Oni, S., Ndiayi, V. N., & Aminu, A. O. (2018). Synthesis and characterization of epoxidized natural rubber-chitosan (ENR-C 2h) for oil spillage sorption. Journal of Chemical Society of Nigeria, 43(1), 187–203.
Ocloo, F. C. K., Quayson, E. T., Adu-Gyamfi, A., Quarcoo, E. A., Asare, D., Serfor-Armah, Y., & Woode, B. K. (2011). Physicochemical and functional characteristics of radiation-processed shrimp chitosan. Radiation Physics and Chemistry, 80, 837–841.
Strusczyk, M. H. (2006). Global requirements for medical applications of chitin and its derivatives; Polish Chitin Society: Łódź, Poland Monograph XI, 95–101.
Devlieghere, F., Vermeulen, A., & Debevere, J. (2004). Chitosan: Antimicrobial activity, interactions with food components and applicability as a coating on fruit and vegetables. Food Microbiology, 21, 703–714.
Chung, Y. C., Su, Y. P., Chen, C. C., Jia, G., Wang, H. L., Wu, J. C., & Lin, J. G. (2004). Relationship between antibacterial activity of chitosan and surface characteristics of cell wall. Acta Pharmacologica Sinica, 25(7), 932–936.
Fernandez-Saiz, P., Lagaron, J. M., & Ocio, M. J. (2009). Optimization of the film forming and storage conditions of chitosan as an antimicrobial agent. Journal of Agriculture and Food Chemistry, 57, 3298–3307.
Li, Z., Yang, F., & Yang, R. (2015). Synthesis and characterization of chitosan derivatives with dual-antibacterial functional groups. Int J of Biol Macro, 75(378), 387. https://doi.org/10.1016/j.ijbiomac.2015.01.056
Li, J., Wu, Y., & Zhao, L. (2016). Antibacterial activity and mechanism of chitosan with ultra high molecular weight. Carbohydrate Polymers, 148, 200–205. https://doi.org/10.1016/j.carbpol.2016.04.025
Oliveira, M., Peixoto, C., & Amorim, R. (2019). Ultrastructural analysis of chitosan antibacterial activity against clinical isolates of Staphylococcus aureus and Escherichia coli. Advances in Microbiology, 9, 893–903. https://doi.org/10.4236/aim.2019.910055
Pokhrel, S., Lach, R., Grellmann, W., Wutzler, A., Lebek, W., Godehardt, R., Yadav, P. N., & Adhikari, R. (2016). Synthesis of chitosan from prawn shells and characterization of its structural and antimicrobial properties. Nepal Journal of Science and Technology, 17(1), 5–9.
Zhuravliova, O. A., Voeikova, T. A., Kuligin, V. S., & Debabov, V. G. (2021). Microbial synthesis and evaluation of bactericidal properties of cadmium sulfide nanoparticles. Journal of Microbiology, Epidemiology and Immunobiology. https://doi.org/10.36233/0372-9311-89
Zherebyateva, O. O., Aznabaeva, L., Ukubaeva, D. G., Mikhailova, E. A., Pankov, A. S., Bolshakova, O. I., Sarantseva, S. V., Lebedev, V. T., & Mironov, A. Y. (2023). Evaluation of nanoparticles as promising alternative antimicrobials. Disinfection Affairs. https://doi.org/10.35411/2076-457x-2023-2-5-14
Holt-Torres, P. S., Chen, Y., & Liu, H. H. (2023). Evaluation of antimicrobial activities of nanoparticles and nanostructured surfaces in vitro. JoVE (Journal of Visualized Experiments), 21(194), e64712. https://doi.org/10.3791/64712
Niu, B., & Zhang, G. (2023). Effects of different nanoparticles on microbes. Microorganisms, 11(3), 542. https://doi.org/10.3390/microorganisms11030542
Shukla, B. S., Gauthier, T. P., Correa, R., Smith, L., & Abbo, L. (2013). Treatment considerations in vancomycin-resistant Enterococcal bacteremia: Daptomycin or linezolid? A review. International Journal of Clinical Pharmacy, 35(5), 697–703.
Pasquina-Lemonche, L., Burns, J., Turner, R. D., Kumar, S., Tank, R., Mullin, N., Wilson, J. S., Chakrabarti, B., Bullough, P. A., Foster, S. J., & Hobbs, J. K. (2020). The architecture of the gram-positive bacterial cell wall. Nature, 582, 294–297.
Qian, J., Pan, C., & Liang, C. (2017). Antimicrobial activity of Fe-loaded chitosan nanoparticles Eng. Life Sciences, 17, 629–634. https://doi.org/10.1002/elsc.201600172
Gomes, L. P., Paschoalin, V. M. F., & Del Aguila, E. M. (2017). Chitosan nanoparticles: production, physicochemical characteristics and nutraceutical applications. Revista Virtual de Química, 9(1), 387–409.
Kefayat, A., Farahani, R. H., Rafienia, M., Hazrati, E., & Nekta, N. H. (2022). Synthesis and characterization of cellulose nanofibers/chitosan/cinnamon extract wound dressing with significant antibacterial and wound healing properties. Journal of the Iranian Chemical Society, 19, 1191–1202. https://doi.org/10.1007/s13738-021-02374-x
No, H. (2002). Antibacterial activity of chitosan and chitosan oligomers with different molecular weights. International Journal of Food Microbiology, 74(1–2), 65–72.
Omidi, S., Kakanejadifard, A., & Azarbani, F. (2018). Enhanced antibacterial activity of functionalized graphene by azo-pyridinium compounds. Journal of the Iranian Chemical Society, 15, 1467–1475. https://doi.org/10.1007/s13738-018-1344-9
Esmaeelzadeh, M., Salehi, P., Bararjanian, M., & Gharaghani, S. (2019). Synthesis of new triazole tethered derivatives of curcumin and their antibacterial and antifungal properties. Journal of the Iranian Chemical Society, 16, 465–477. https://doi.org/10.1007/s13738-018-1524-7
Bakr, R. B., & Azab, I. H. E. (2022). Elkanzi NAA (2022) Thiochromene candidates: design, synthesis, antimicrobial potential and in silico docking study. Journal of the Iranian Chemical Society, 19, 1413–1423. https://doi.org/10.1007/s13738-021-02391-w
Ibrahim, S. A., Rizk, H. F., El-Borai, M. A., & Sadek, M. E. (2021). Green routes for the synthesis of new pyrazole bearing biologically active imidiazolyl, pyridine and quinoxaline derivatives as promising antimicrobial and antioxidant agents. Journal of the Iranian Chemical Society, 18, 1391–1404. https://doi.org/10.1007/s13738-020-02119-2
Akinduti, P. A., Motayo, B., Idowu, O. M., Isibor, P. O., Olasehinde, G. I., Obafemi, Y. D., Ugboko, H. U., Oyewale, J. O., Oluwadun, A., Adeyemi, G. A. (2019).Suitability of spectrophotometric assay for determination of honey microbial inhibition. Journal of Physics: Conference Series, 1299, 012131. IOP Publishing. https://doi.org/10.1088/1742-6596/1299/1/012131
Mohammadi, A., Hashemi, M., & Hosseini, S. (2016). Effect of chitosan molecular weight as micro and nanoparticles on antibacterial activity against some soft rot pathogenic bacteria. Lwt - Food Science and Technology, 71, 347–355. https://doi.org/10.1016/J.LWT.2016.04.010
Ismatova, N. R. (2023). Antibacterial response of nanostructured chitosan hybrid materials. Biological and Medical Physics Series, 161-179. https://doi.org/10.1007/978-981-19-9646-7_7
Nam-Cha, S. H., Ocaña, A. V., Pérez-Tanoira, R., Aguilera-Correa, J. J., Domb, A. J., Ruiz-Grao, M. C., Cebada-Sánchez, S., López-Gónzalez, Á., Molina-Alarcón, M., Pérez-Martínez, J., & Pérez-Martínez, F. C. (2023). Methacrylate cationic nanoparticles activity against different gram-positive bacteria. Antibiotics (Basel), 12(3), 533. https://doi.org/10.3390/antibiotics12030533
Linklater, D. P., Baulin, V. A., Guével, X. L., Jleury, J. B., Hanssen, E., Juodkazis, S., Bryant, G., Crawford, R. J., Stoodley, P., & Ivanova, E. P. (2020). Antibacterial action of nanoparticles by lethal stretching of bacterial cell membranes. Advanced Materials, 32(52), 2005679. https://doi.org/10.1002/ADMA.202005679
Phakatkar, A. H., Ghildiyal, P., Wang, Y., Zachariah, M. R., Shokuhfar, T., & Shahbazian-Yassar, R. (2022). In-situ TEM studies on nanoparticle interactions with bacterial cells. Microscopy and Microanalysis, 28, 1104–1106. https://doi.org/10.1017/S1431927622004676
Helander, I., Nurmiaho-Lassila, E., Ahvenainen, R., Rhoades, J., & Roller, S. (2001). Chitosan disrupts the barrier properties of the outer membrane of gram-negative bacteria. International journal of food microbiology, 71(2–3), 235–244. https://doi.org/10.1016/S0168-1605(01)00609-2
Jung, E., Youn, D., Lee, S., No, H., Ha, J., & Prinyawiwatkul, W. (2010). Antibacterial activity of chitosans with different degrees of deacetylation and viscosities. International Journal of Food Science and Technology, 45, 676–682. https://doi.org/10.1111/J.1365-2621.2010.02186.X
Sarian, F. D., Ando, K., Tsurumi, S., Miyashita, R., Ute, K., & Ohama, T. (2022). Evaluation of the growth-inhibitory spectrum of three types of cyanoacrylate nanoparticles on gram-positive and gram-negative bacteria. Membranes (Basel)., 12(8), 782. https://doi.org/10.3390/membranes12080782.PMID:36005697;PMCID:PMC9414559
Acknowledgements
Appreciation to Mr. Usman Abdulrahman Department of Microbiology and Biotechnology, Faculty of Pharmacy, University of Lagos, Idi-Araba, Lagos State, Nigeria (microbial analysis); Engr. Abdulhamid Abdurahman and Mallam Musa Yusuf Central Laboratory, Umar Musa Yar’dua University Kastina, Kastina State, Nigeria (SEM, EDXRF and XRD analyses), and Dr Jane Reuben-Kalu Department of Plant Biotechnology, Tamil Nadu Agricultural University, Coimbatore, 641003, India (SEM and EDX analyses).
Author information
Authors and Affiliations
Contributions
RO: conceptualization and research design, data interpretation, supervision, and manuscript preparation; SA: conceptualization and research design, execution of research design, and manuscript preparation; FA: conceptualization and research design, data interpretation, supervision, and manuscript preparation; OA: provision of research resource (FTIR instrumentation) and data interpretation.
Corresponding author
Ethics declarations
Ethical Approval
This study did not involve use of human and animal specimens.
Consent for Publication
All authors gave their consent for publication.
Competing Interests
The authors declare no competing interests.
Informed Consent
All authors gave their approval for publication.
Funding
The study was self funded by the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Atanda, S.A., Shaibu, O.R., Agunbiade, F.O. et al. Facile Synthesis and Characterization of Chitosan Nanoparticles from Archachatina marginata Shell as Potential Solution to Antimicrobial Resistance. BioNanoSci. (2024). https://doi.org/10.1007/s12668-024-01399-9
Accepted:
Published:
DOI: https://doi.org/10.1007/s12668-024-01399-9