Abstract
The antimutagenic activity of probiotic strains has been reported over several decades of studying the effects of probiotics. However, this activity is rarely considered an important criterion when choosing strains to produce probiotic preparations and functional food. Meanwhile, the association of antimutagenic activity with the prevention of oncological diseases, as well as with a decrease in the spread of resistant forms in the microbiota, indicates its importance for the selection of probiotics. Besides, an antimutagenic activity can be associated with probiotics’ broader systemic effects, such as geroprotective activity. The main mechanisms of such effects are considered to be the binding of mutagens, the transformation of mutagens, and inhibition of the transformation of promutagens into antimutagens. Besides, we should consider the possibility of interaction of the microbiota with regulatory processes in eukaryotic cells, in particular, through the effect on mitochondria. This work aims to systematize data on the antimutagenic activity of probiotics and emphasize antimutagenic activity as a significant criterion for the selection of probiotic strains.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the modern anthropogenic environment, humans and animals are regularly exposed to various mutagens, in amounts that significantly exceed natural ones. Some of the widely used drugs can have a mutagenic effect [1], including those acting on the human microbiota, increasing the likelihood of antibiotic resistance. Anticancer drugs [2, 3] or antibiotics [4] are examples of such preparations. Moreover, mutagens are formed even during heat treatment of food [2]. Reactive oxygen species produced by both body cells and microbiota can also be considered promutagens, so probiotics’ antioxidant potential should also be considered [3].
In addition to exogenous mutagens that enter the human body from the environment, there are also endogenous mutagens. Reactive oxygen species (ROS), produced in mitochondria, are an integral part of cellular metabolism. ROS take part in signaling pathways in the cell, but they can lead to oxidative stress and damage cellular molecules: lipids, proteins, and DNA, thereby causing mutations [5, 6].
A high mutational burden leads to an increase in the level of several diseases and systemic effects. Also, the accumulation of mutations at critical points of the genome is considered one of the most likely mechanisms of aging. And even though now it is believed that the accumulation of mutations by cells to a greater extent affects the likelihood of malignization than the general fitness, over time there is more and more evidence that healthy cells accumulate many somatic mutations with age. That is, among others, associated with the risk of age-related diseases [7].
Antimutagens, or substances and structures capable of inactivating mutagens or reducing their effect on the body, have been found in various natural sources, including probiotic microorganisms.
However, in the search for new probiotic strains and screening and selection of components for complex probiotic preparations, little attention is paid to the criterion of antimutagenicity. The antimutagenic activity of probiotic strains is rarely considered an important criterion when choosing strains to produce probiotic preparations and functional food. Meanwhile, the association of antimutagenic activity with the prevention of oncological diseases, as well as with a decrease in the spread of resistant forms in the microbiota, indicates its importance for the selection of probiotics.
Besides, an antimutagenic activity can be associated with probiotics' broader systemic effects, such as geroprotective activity.
Mostly, the positive effect of probiotics on the host organism is explained by the following mechanisms:
-
1.
Probiotic microorganisms are in an antagonistic relationship with pathogenic fungi and bacteria.
-
2.
Probiotics release metabolites that have a positive effect on the host organism.
-
3.
Probiotics can metabolize hazardous and toxic compounds coming from the external environment into less toxic ones.
-
4.
Probiotics release substances that can interfere with the regulatory processes in the host’s body.
-
5.
There are also cases of specific interactions, such as some strains’ ability to suppress tumor cells’ growth.
However, a variety of systemic effects can also be observed. Some studies indicate that probiotics affect the mental state of the host [8], interfere with the regulation of metabolism, the work of hormonal systems [9], gene expression [10], and other regulatory mechanisms. Gradually, numerous facts have accumulated about the ability of probiotic microorganisms to effectively correct pathological manifestations of diseases not associated with infections, particularly allergies, toxicosis of various natures, etc. [11]. This nonspecific stimulating activity may be associated with the release of metabolites that protect host cells from the most destructive effects of stress — the generation of reactive oxygen species and DNA damage.
It is also known that probiotics play an immunomodulatory role, have anticancer effects, and help lower cholesterol levels. These functions are associated with the release of metabolites such as bacteriocins, biosurfactants, exopolysaccharides, and siderophores [12].
The antimutagenic activity of probiotics is generally viewed primarily in the context of the mechanism of anticancer action. A search for reviews focusing on the antimutagenic effects of probiotics reveals that no review has emerged that specifically addresses the antimutagenic effects of probiotics. The most recent review on a related topic, containing a mention of the antimutagenic activity of probiotics, also focuses on the anticancer effect of exopolysaccharides of lactic acid bacteria [13].
Indeed, it is the antimutagenic activity that allows some probiotic strains to reduce cancer incidence in hosts, which has been shown in various mammalian models. Thus, it has been shown that the administration of lactobacilli and Bifidobacterium effectively reduces DNA damage caused by chemical carcinogens in the gastric and colon mucosa in rats [14]. Lyophilized cultures of Bifidobacterium longum, introduced into the diet of rats, inhibited tumors of the liver, colon, and mammary gland caused by food mutagens [15]. Several spore-forming probiotics, such as Bacillus subtilis var. natto, also exhibit anticancer properties [16, 17].
Nevertheless, an antimutagenic activity can be the basis for other systemic effects, such as, for example, slowing down aging processes, including reproductive aging. It has been shown that the stabilization of mitochondrial DNA observed under the action of probiotic Bacillus preparations may be associated with the prolongation of reproductive age in chickens [18]. It should also be noted that metabolites of probiotic bacteria exhibit the ability to suppress the SOS response [19], which can also be attributed to antimutagenic effects.
This work aimed to systematize data on the antimutagenic activity of probiotics and emphasize antimutagenic activity as a significant criterion for selecting potential probiotic strains. The novelty of this work is that for the first time, antimutagenic activity is considered as an independent criterion for screening probiotics. We believe that putting together the available data on antimutagenic activity in a form of a critical review, with emphasis on the importance of this criterion for the future selection of probiotic candidates, may inspire researchers to use the criterion of antimutagenic activity in the selection of probiotics.
Probiotics with Antimutagenic Activity
Among all genera of probiotic bacteria, representatives of lactic acid bacteria (LAB) and Bifidobacterium are most often mentioned as sources of antimutagenic compounds. Several studies showed antimutagenic activity against heterocyclic amines, N-nitroso compounds, benzo (a) pyrene, and aflatoxin B [20,21,22]. Both live cultures of LAB and their fermentation products demonstrate antimutagenic and anticarcinogenic activity [23, 24].
Probiotics strains with antimutagenic activity are also found in other groups of microorganisms. For example, the Escherichia coli Nissle 1917 (EcN) strain is one of the oldest probiotics [25] that exhibits antimutagenic activity. Presumably, 4-nitroquinoline-1-oxide (4-NQO) is deactivated by the E. coli cell’s metabolic systems with the formation of decay products of 4-aminoquinoline. However, the exact mechanisms of deactivation of benzo(a)pyrene have not yet been established [26]. This kind of activity is quite typical for probiotics, for example, Lacticaseibacillus (formerly Lactobacillus) rhamnosus IMC501 can also convert 4-NQO into a non-genotoxic metabolite [27].
LAB with Antimutagenic Properties
It was stated that probiotic bacteria characteristic of the microbiota of goats, isolated from healthy goat feces and belonging to the genera Lactobacillus, Enterococcus, and Bifidobacterium, could reduce the mutagenicity of sodium azide and benzopyrene in the Ames test and reduce the risk of gastrointestinal cancer [28]. Among the substances whose mutagenic effects can be reduced by probiotic lactobacilli, heterocyclic aromatic amines should also be noted [29].
Lactobacillus and Bifidobacterium produce extracellular bioactive compounds with antimutagenic properties against benzo[a]pyrene (BaP) and sodium azide (SA). Interestingly, the common antimutagenic effects in exponential and stationary growth phases were different. Lactobacillus exhibit this activity mainly in the stationary growth phase [30].
Lactobacillus acidophilus (isolated from commercially available yogurt), Lactobacillus gasseri (P79), Weissela confusa (formerly Lactobacillus confusus) (DSM20196), Streptococcus thermophilus (NCIM 50,083), Bifidobacterium breve, and Bifidobacterium longum (isolated from child stools) reduced the DNA-damaging effect of methylnitronitrosoguanidine (MNNG) in rat intestinal cells. It is peptidoglycan fraction from lactobacilli that exhibit antimutagenic effects [14].
It has been shown that six strains of L. acidophilus and nine strains of Bifidobacteria show antimutagenic activity against the following mutagenic compounds: MNNG; 4-nitro-O-phenylenediamine; 4-nitroquinoline-N-oxide; Aflatoxin B; 2-amino-3-methyl-3H-imidazoquinoline; PhIP, и 2-Amino-3-methyl-9H-pyrido[2,3-b]indole. The effect strongly depends on strains [22]. L. acidophilus LA 106 fermented milk significantly decreased mutagenic effects by MNNG (by 77%) [31].
Lactiplantibacillus (formerly Lactobacillus) plantarum and Staphylococcus xylosus reduced the mutagenic activity of biogenic amines in the production of sausages [32].
Streptococcus thermophilus and Lactobacillus bulgaricus fermented milk reduced the effects of 4-nitroquinoline-N-oxide (a direct-acting mutagen) and 2-aminofluorene (a mutagen requiring S9 activation) [33].
There is evidence that palmitic acid produced by Lactobacillus delbrueckii ssp. bulgaricus and Streptococcus salivarius ssp. thermophilus in yogurt has antimutagenic effects on MNNG [34].
The possible mechanism of Lacticaseibacillus (formerly Lactobacillus) casei ATCC 393 antimutagenic effect can be connected with involvement and support in polyamines metabolism (putrescine, spermidine, and spermine) in host-organism cells [35].
It should be noted that not all LAB have antimutagenic activity. Moreover, Sharma M. et al. (2020) have shown that out of 60 LAB isolated from various sources, only 10 isolates showed antigenotoxicity of more than 30%, and four showed cytotoxicity of 70–80% [36]. In another research, only 4 strains from 25 isolates exhibited a pronounced antimutagenic activity [37].
Bifidobacterium with Antimutagenic Properties
Bifidobacterium bifidum, Bifidobacterium lactis, and Bifidobacterium longum showed significantly higher antimutagenic potential against benzo(a)pyrene than Bifidobacterium adolescentis, Bifidobacterium breve, and Bifidobacterium infantis. In particular, the activity of bifidobacteria on benzo [a] pyrene was noted by Lo et al. [21]. Bifidobacterium pseudocatenulatum G4 and B. longum are able to directly bind heterocyclic amines [38]. Bifidobacterium longum exhibited anti-mutagenic properties in fermented milk [20] and have shown the ability to bind dietary carcinogens [39].
Bacillus with Antimutagenic Properties
As for the representatives of the genus Bacillus, there is less research reported on their antioxidant and antimutagenic activity, although this is gradually changing. However, it should be noted that bacteria of this genus began to be considered as probiotic bacteria later than LAB.
Caldini et al. studied the effect of 16 Bacillus strains from pharmaceutical probiotic preparations and laboratory collections (B. subtilis, Bacillus firmus, Bacillus megaterium, Bacillus pumilus) on genotoxicity caused by the standard mutagen 4-nitroquinoline-1-oxide (4-NQO) using the SOS chromotest, with E. coli PQ37 as a test organism [40]. It was found that the activity of 0.1 mm 4-NQO decreased after co-incubation with Bacillus suspension with a titer of 108 CFU/mL. All isolates showed the ability to deactivate 4-NQO, with genotoxicity inhibition ranging from 92.9 to 100%. The authors associate the observed effect with the modification of the 4-NQO molecule.
In a later work [41], the inhibitory effect of 21 bacilli strains on four genotoxins was investigated in vitro using the same method. All strains exhibited high inhibitory activity against 4-nitroquinoline-1-oxide and N-methyl-N′-nitro-nitro-nitrosoguanidine (direct genotoxic agents), while against 2-amino-3,4-dimethylimidazo [4,5-f]-quinoline and aflatoxin B1 (indirect genotoxic agents), inhibitory activity was high or moderate. Antigenotoxicity was observed in vegetative cells but not in heat-treated cells or spore suspensions. The spectroscopy showed that the properties of genotoxin molecules were changed after incubation with cells, and all strains retained high viability after exposure to genotoxins.
It has been shown that the Bacillus coagulans strain GKN316 can efficiently metabolize furfural, 5-hydroxymethylfurfural (HMF), vanillin, syringaldehyde, and p-hydroxybenzaldehyde (pHBal), converting them into less toxic corresponding alcohols in situ [42].
Other Gut Bacteria with Antimutagenic Properties
Cell extracts and Streptococcus faecalis cells reduce the mutagenic effect of 2-nitrofluorene in the Salmonella Typhimurium TA1538 strain. This is manifested through several mechanisms involving extracellular and intracellular factors. Presumably, thiol compounds are extracellular factors. Desmutagens affecting the biotransformation of a mutagen within a cell include thermally stable compounds, possibly of proteinaceous nature, with a molecular weight of less than 12 kDa [43].
Another intestinal microorganism Enterococcus faecium M-74 had a more significant antimutagenic effect under similar conditions in a live state and when selenium was added to the medium [44].
Most of the above studies have some methodological drawbacks. The researchers choose xenobiotics as genotoxic substances (such as MNNG and NQO, rather exotic for living organisms) that must be inactivated by probiotics. Meanwhile, living organisms do not encounter these compounds that often. Even if considering the anthropogenic environment, among all the most frequently used experimental models of promutagens and mutagens, benz(a)pyrene is the only compound that humans and animals have to deal with. If we consider probiotics as a factor in preventive therapy against diseases caused by mutagenic factors, models based on more typical substances that threaten the human body in the modern world should be used for screening and selection of promising targeted probiotic strains. As stated earlier in the “Introduction” section, many drugs are mutagens and therefore can be used for similar models.
Probiotic strains with antimutagenic activity and their sources were summarized in Table 1.
Analyzing the data summarized in Table 1, we can conclude that among all references to probiotics with antimutagenic activity, representatives of lactobacilli are in the lead. According to our meta-analysis, reports of them represent about 43% of all mentions of antimutagenic probiotics. In 23% of cases, bifidobacteria are mentioned; 20% of papers mention representatives of the genus Bacillus; 9%, Streptococcus sp.; and in 5% of cases, other bacteria.
Antioxidant Activity of Probiotics
Antioxidant activity, although part of antimutagenic activity, requires separate consideration. The mechanisms responsible for it are usually more specific than those that provide antimutagenic activity.
Antioxidant activity of probiotics is shown for many strains, among which Lactobacillus species are most studied and used in medicine and the food industry.
Lactobacillus with Antioxidant Properties
Chooruk et al. showed that in a series of 201 strains of lactobacilli isolated from the human oral cavity, antioxidant activity is, to some extent, inherent in all of the isolated strains, and in a large number of strains, it was significant. The most prominent strains belonged to L. fermentum, L. paracasei, and L. rhamnosus [52]. It was shown that L. plantarum ATCC14917 enhanced the antioxidant activity of apple juice [53], and the Levilactobacillus (formerly Lactobacillus) brevis KCCM 12203P strain possessed both antioxidant and immunomodulatory activity [54].
Bifidobacterium with Antioxidant Properties
Bifidobacterium probiotic species are less studied due to the difficulty of their cultivation in the laboratory. However, there is evidence of the antioxidant activity of their representatives. It was shown that B. longum LTBL16 has high antioxidant activity [55]. B. lactis strain HN019 reduced the level of oxidative stress in patients with metabolic syndrome and reduced the level of inflammation [56]. The intake of Bifidobacterium bifidum ATCC 29,521 had a beneficial effect on the structure of the intestinal microbiota, in addition to antioxidant effects [57].
Other Gut Bacteria with Antioxidant Properties
Among genera that are less clearly (or, perhaps, more controversially) associated with probiotic activity, there are also probiotic species with antioxidant properties. For instance, Ent. faecium strains isolated from various fermented foods were reported as having antioxidant properties [58]. Streptococcus salivarius ssp. thermophillus strain exhibited high antioxidant activity and caused a significant decrease in the level of markers of oxidative stress in liver cells of mice [59]. Representatives of the genus Bacillus also demonstrated high antioxidant activity levels, both in vitro and in vivo [60, 61].
Bacterial Consortia with Antioxidant Properties
Antioxidant activity is observed not only in individual strains but also in microbial consortia. For instance, the kefir grains consortium’s bacteria exhibit antioxidant properties, and their beneficial effect on the condition of patients with Alzheimer’s disease (AD) has been shown to be connected with such activity. Moreover, in this study antioxidant defense mechanisms were involved in improving physiological and cognitive functions, and probiotics were able to reduce the ROS-mediated pro-inflammatory response, which is part of the pathogenesis of AD [62].
Antioxidant activity can overlap with antimutagenic activity since a decrease in ROS levels will reduce the total number of mutations in the genome. However, there are methods to consider them separately. Such an approach, in particular, can be the use of bacterial biosensors — a method that we used and was shown to be successful for the stage of primary screening of probiotics for veterinary medicine (see Table 3 and the corresponding links). In this case, the marker is the expression of bacterial genes responsible for responding to specific ROS (superoxide anion radical, hydrogen peroxide) or DNA damage (individual genes of the SOS response system).
Possible Mechanisms of Action
The mechanisms of antimutagenic action of probiotics are still the subject of discussion. It should be noted that the antimutagenic properties of probiotics can benefit the host not only through direct interaction of metabolites of probiotic bacteria with host cells but also indirectly by reducing the intensity of mutational processes in the microbiota. As shown earlier [3], this process slows down the emergence and spread of antibiotic resistance factors in the microbiota, reducing possible complications after antibiotic therapy.
At the molecular level, two groups of mechanisms can be distinguished:
-
1.
Direct binding to mutagens.
-
2.
Mutagens transformation [28]
On the cellular level, there are also two ways to fight mutagens:
-
3.
Production and/or excretion of antimutagenic metabolites.
-
4.
Production of antimutagenic substances via fermentative transformation of a substrate.
The latter two mechanisms are not necessarily ways of dealing with dangerous mutagens for themselves but could also arise as a side effect of other processes and gain a foothold as an advantage in symbiotic relationships.
On a systemic level:
-
5.
Indirect influence on the level of spontaneous mutagenesis and the expression of genes of the host defense systems.
Binding of Mutagens
The correlation between lactobacilli’s ability to bind mutagens and their antimutagenic activity has been persuasively shown by Stidl et al. [47].
The cell wall components can also play a significant role in binding and deactivating mutagens [29, 63]. Specifically, such a mechanism should be characteristic of the inactivation of amines [29]. Morotomi and Mutai found that the ability of probiotic Lactobacillus to bind to mutagenic products of tryptophan pyrolysis is pH-dependent and decreases with the addition of metal salts [64]. The authors concluded that the effect of amine binding by the bacterium L. casei appears to be related to cation-exchange mechanisms.
Studies comparing Gram-positive and Gram-negative bacteria’s ability to bind the pyrolysis products of tryptophan have shown that Gram-positive strains are consistently more effective [65]. This fact can be taken as an indication of the cell wall structure’s important role in the inactivation of mutagens. Subsequent studies have shown that the binding of amines with Gram-positive and Gram-negative bacteria occurs in the peptidoglycan layer and the outer membrane, respectively [48]. Sreekumar and Hosono suggested that the HA binding receptors are carbohydrate fragments of the cell wall and that glucose molecules play a crucial role in the binding reaction [20].
Transformation of Mutagens
The possibility of inhibiting certain stages of the transformation of promutagens into mutagens is also considered as a possible mechanism of probiotics’ action. It has been found, for example, that L. delbrueckii ssp. bulgaricus 191R releases hydrophobic metabolites with antimutagenic activity against MNNG and 3,2′-dimethyl-4- aminobiphenyl (DMAB). The exact mechanism or active substance has not been identified, but the authors consider the inhibition of cytochrome P450 1A2, inhibition of subsequent activating enzymes such as acetylase, reaction with N-hydroxy DMAB or other already activated forms of DMAB, or increased DNA repair as mechanisms of its activity [61].
In some studies, the effects of temperature-inactivated cells were compared with those of living cells, and it was found that the latter has a consistently higher antimutagenic activity [22, 66]. This observation suggests that living bacteria can produce metabolites or catalyze reactions that detoxify amines. Another study investigated the potential antimutagenic effects of various organic acids (lactic acid, butyric acid, and acetic acid) on IQ, PhIP, and Trp-P-1 and some non-amine carcinogens [20]. These acids are products of microbial fermentation of fibers and other polysaccharides [67]. Butyric acid inhibited the mutagenic effects of amines in the Salmonella enterica serovar Typhimurium TA98 test, while no such effects were observed with other acids.
It was shown that L. rhamnosus 231 has several mechanisms of antimutagenic action: adsorption (for example, acridine orange) and biotransformation with subsequent detoxification (for example, MNNG and 2-amino-3,8-dimethylimidazo[4,5-f]quinoxaline) [51].
Production and/or Excretion of Antimutagenic Metabolites
Considering the release of effector molecules, one can assume two possibilities: probiotics can release antimutagenic metabolites or transform the substrate so that antimutagen compounds are obtained. Methodologically, it can be quite difficult to differentiate these two mechanisms by studying the effects of a particular strain.
Thus, it has been shown that the main contribution to the antimutagenic activity of sour milk fermented by the probiotic strain L. plantarum is made by peptides less than 3 kDa and 3–10 kDa in size. However, it is unknown whether they are produced by the bacterium or obtained during the proteolysis of milk proteins [68].
The study of antimutagenic metabolites of the probiotic strain L. rhamnosus MD 14 showed that they belong to thermosensitive protein compounds and organic acids [35]. However, on the other hand, Bacillus metabolites, which have antimutagenic activity and can inhibit the SOS response in E. coli, exhibited thermal stability [19].
Fermentation of soy milk by lactic acid bacteria (Strep. thermophilus, L. acidophilus) and bifidobacteria (B. infantis, B. longum) significantly increased its antimutagenic properties against 4-nitroquinoline-N′-dimethyl -biphenyl (DMAB). The mutagenic effect of these compounds was also reduced by pretreating S. Typhimurium TA 100 cells with fermented soy milk [69, 70].
The degree of proteolysis of proteins in yogurt by Lact. acidophilus (ATCC® 4356 ™), L. casei (ATCC® 393 ™), and Lacticaseibacillus (formerly Lactobacillus) paracasei subsp. paracasei (ATCC® BAA52 ™) correlated with its antimutagenic activity. The released peptides showed high activity in trapping radicals with 1,1-diphenyl-2-picrylhydrazyl and 2,2′-azino-bis (3-ethylbenzothiazoline-6-sulfonic acid) [71]. It is known, L. plantarum KLAB21 produces glycoproteins with antimutagenic activity [72].
The differentiation of the mechanisms of cell-mediated antimutagenic activity from that of metabolite-mediated can be performed via testing cell-free preparations together with probiotic strain’s cells.
Systemic Effects
One more direction for implementing the antimutagenic effects of probiotics could be pointed out: these effects can be achieved by indirectly influencing the level of spontaneous mutagenesis. Recently, increasing information has appeared that probiotics can influence the expression of host genes, interfering with the work of its regulatory cascades, such as, the p38 MAP kinase pathway [73, 74]. It should be noted that these effects were observed both under the action of living cells and under the influence of cell-free preparations [74]. Such a change in the host’s cellular homeostasis may be associated, among other things, with the level of mutagenesis in the cells.
Maintaining the balance of prooxidants/antioxidants in the cell can also be one of the indirect pathways.
Mechanisms of Antioxidant Activity
As mentioned above, antioxidant activity can be considered a special case of antimutagenic activity, usually provided by different mechanisms that vary from strain to strain.
In general, we can describe antioxidant mechanisms by the binding or transformation of prooxidants/ROS, the release of antioxidants or the conversion of substrate molecules into antioxidants, as well as the regulation of the host defense systems.
Prooxidants/ROS Binding and Transformation
Another possible mechanism for the antioxidant action of probiotic bacteria is metal chelation. For example, Lactobacillus helveticus CD6 can produce substances that bind Fe2+ ions into chelates [75].
Bacteria have their own Fe-SOD and Mn-SOD enzyme systems to protect against free radicals [76], while Mn-SOD is similar to the Mn-SOD of eukaryotic cells mitochondria. These enzymes can reduce the number of prooxidant molecules in the environment. It has been shown that two strains of L. fermentum, which has a high production of glutathione peroxidase (GPx), also had significant antioxidant properties [77, 78]. There are genetically modified strains of lactobacilli that carry catalase genes, and their use has been shown to reduce the severity of Crohn’s disease in mice [79].
Producing of Antioxidant Metabolites or Substrate Transformation
A significant part of the antioxidant properties of food products, for example, milk and dairy products, is provided by protein substances: casein fraction and albumin [80] and short peptides [81]. Probiotic bacteria, which are used to produce fermented dairy products, can increase the number of antioxidant peptides in products due to their proteolytic activity. It has been shown for different genera of LAB, for example, for the symbiotic cultures of L. delbrueckii ssp. bulgaricus and Strep. thermophilus, as well as monocultures of L. acidophilus, L. casei, and B. bifidum [82]. At the same time, it is noted that the addition of probiotic strains increases the antioxidant properties of the fermented product compared to the unfermented one [83]. There is a correlation between the level of the strain’s proteolytic activity and the final product’s antioxidant properties, as was shown by the example of various Lactobacillus species [71, 83]. Solieri et al. compared the proteolytic and antioxidant activity of 39 non-starter lactobacilli from different cheeses [83]. Sah showed the same effect on L. acidophilus (ATCC4356), L. casei (ATCC 393), and L. paracasei ssp. paracasei (ATCC BAA52). Furthermore, even if there was no difference in antioxidant activity between the fermented and non-fermented products, there was a better bioavailability of antioxidants from the fermented product [47] as demonstrated with five strains of Bifidobacterium longum ssp. Longum [84]. We should note the synergistic effect of co-cultivation of different strains, as was shown for Lact. acidophilus (ATCC4356), L. casei (ATCC 393), and L. paracasei ssp. paracasei (ATCC BAA52) [71].
The other way is the production of low-molecular antioxidant molecules such as glutathione [85], butyrate [86, 87], and folate [75]. It has been shown that probiotic bacteria have enzyme complexes to produce antioxidant molecules, such as glutathione-producing L. fermentum ME-3 enzymatic system [85]. These molecules can be absorbed by the host and exhibit their properties in the host’s cells and tissues, for example, reducing the effects of oxidative stress in the liver [87,88,89].
Considering extracellular metabolites, some specific qualities should be mentioned, specifically: small size (associated with the ability to penetrate membranes); resistance to proteinases and other environmental factors; existence in a variety of isoforms; and the ability to reform the structure quickly. In particular, bacterial oligopeptides and lipopeptides synthesized both ribosomally and nonribosomally usually correspond to those criteria. It is known that the non-ribosomal synthesis of oligopeptides occupies a more significant share in the metabolism of Bacillus. Its products do not exceed several kDa in size, and a significant number of them are thermostable and are not hydrolyzed by proteinase K. Such resistance is provided by the atypical amino acids and stereoisomers in the structure of nonribosomally synthesized bacilli peptides [90]. These peptides are often considered to be antimicrobial and antifungal agents; however, recent data indicate their participation in regulatory processes [91]. For Lactobacillus, Enterococcus, or Bifidobacterium probiotic strains, the synthesis of ribosomal peptides with a broad spectrum of activities is more typical.
It is known that endogenous eukaryotic peptides that regulate the prooxidant/antioxidant balance are involved in the body’s response to oxidative stress that occurs during pathological processes and stressful conditions [92]. In addition to the signaling effect that allows peptides to normalize the cell’s oxidative status, they have antioxidant properties [93].
Besides, peptides of various origins (including synthetic peptides) can regulate the processes of cell proliferation and apoptosis [94], as well as penetrate the nucleus and nucleolus and bind there with DNA and histone proteins, affecting gene expression [91, 95]. Thus, peptides released by the microbiota should not be disregarded as they may have similar effects.
Systemic Effects
Presently, the influence of probiotic bacteria on the host organisms signaling pathways is an actively studied topic. For instance, Lactobacillus spp. influence the Nrf2-Keap1-AREA pathway. Nrf2 activates many genes, including those involved in the detoxification of xenobiotics and ROS [96]. An extracellular polysaccharide from the Bacillus sp. LBP32 inhibits NFκB production, preventing macrophage inflammatory responses, and ROS production [97]. L. rhamnosus GG improved the state of intestinal epithelial cells under severe oxidative stress through the production of soluble proteins p40 and p75, which acted through the mechanism of activation of mitogen-activated protein kinases (MAPKs), as well as inhibition of protein kinase C (PKC) [98]. However, specific mechanisms and specific signaling molecules that regulate these pathways are often not described in published reports.
It is proved that YD1 peptide, isolated from Bacillus amyloliquefaciens CBSYD1, has antioxidant activity and an effect on the host organism, similar to NF-E2-related factor-2 (Nrf-2) [46]. B. amyloliquefaciens SC06 strain reduced the level of damage to pig intestinal epithelial cells by modulating the Nrf2/Keap1 pathway and ROS production [60]. B. megaterium SF185 also protected CACO-2 intestinal epithelial cells from the effects of hydrogen peroxide [99]. However, in general, the antioxidant effect of bacillary probiotics is described much less than that of probiotics based on LAB and bifidobacteria.
The search for metabolites that provide these and other effects listed above seems to be a promising topic for further research.
However, the relationship between the antimutagenic activity of probiotic bacteria and the production of antioxidants is currently at the initial stage of the study. The ability of lacto- and bifidobacteria to produce substances that inactivate ROS has been reliably confirmed by experiments [100, 101], but for spore-forming probiotics, such activity has so far been described in very fragmentary terms.
Postbiotics as Antioxidants and Antimutagens
In recent years, researchers have been using the term “postbiotics” for the products of the probiotic microorganism’s activity that can positively affect the host organism, even in the absence of living cells. Among the representatives of this group, compounds with antioxidant immunomodulatory and anticancer properties have been identified.
Postbiotics are functional bioactive compounds generated in a matrix during fermentation. Postbiotics can include many different components, such as metabolites, short-chain fatty acids (SCFA), microbial cell fractions, functional proteins, extracellular polysaccharides (EPS), cell lysates, teichoic acid, peptidoglycan derived muropeptides, and pili-like structures [102].
As the variety of substances included in this group, the properties of postbiotics are diverse. They are able to exert immunomodulatory effects; for example, postbiotics obtained from Bifidobacterium breve C50 and Strep. thermophilus 065 induce high IL-10 production through TLR-2 and also stimulate Th1 responses [103, 104]. The use of L. paracasei CBA L74 postbiotics in infants led to a change in the levels of immune biomarkers in the blood and amelioration of the disease’s progression [105].
The cell-free supernatants of L. acidophilus, L. casei, Lactococcus lactis, Limosilactobacillus (formerly Lactobacillus) reuteri, and Saccharomyces boulardii demonstrate an antioxidant activity in addition to immunomodulatory effect [106]. Postbiotic exopolysaccharides from L. plantarum 70,810 have antitumor properties, inhibiting the proliferation of HepG-2, BGC-823, and HT-29 tumor cells [107]. Exopolysaccharides from L. helveticus MB2 showed the ability to bind ferrous ions, which provides one of the well-known mechanisms of antioxidant activity [108]. Exopolysaccharides of several wild lactobacilli strains also showed antioxidant properties [109].
As mentioned above, the production of folate, glutathione, and other antioxidant molecules is one of the mechanisms of probiotics’ antioxidant activity. Moreover, such products can be considered postbiotics [75, 85]. For example, folate-producing L. helveticus CD6 cell-free supernatants demonstrate antioxidant properties [75].
Some of the more specific properties of postbiotics are presented in Table 2.
Antimutagenic Effect of Probiotics in Mitochondria
In recent years, there was increasing evidence indicating that mitochondria can, in addition to a well-studied energy function, also serve as a kind of a “regulatory center” for eukaryotic cells [110,111,112,113]. Changes in mitochondrial function affect many processes, ranging from aging to diabetes [114, 115], and many of the proteins that play a crucial role in signaling cascades are found in mitochondria. It is especially relevant for systems regulating oxidative status, which provide an accurate balance of pro- and antioxidant activity [116]. For example, regulation of the nfe2l2/AP1 pathway that controls the antioxidant system can be directly initiated by changes in the mitochondrial membrane state. The same works for a number of other cascade processes and regulators: the antioxidant defense pathway regulated by MAPK10 kinase and the NFE2L2/AP1 pathway in general, the thioredoxin 2/peroxiredoxin 3 system, etc. [117,118,119].
This set of data could lead to an interesting hypothesis that the antimutagenic effect of probiotics can be realized indirectly. Namely, it can be carried out through the influence on redox homeostasis through interaction with mitochondria. Stefanaki et al. showed that the intestinal microbiota and its secreted metabolites could interact with mitochondria [120]. The prokaryotic origin of mitochondria is likely to contribute to such interactions [121].
The possibility of interaction of probiotics with host mitochondria is supported, for example, by the following studies. In Nakagawa et al. experiments, the L. gasseri SBT2055 probiotic increased the lifespan of Caenorhabditis elegans [73]. It was noted that the number of mitochondria significantly increased when the host was fed with the LG2055 strain, as compared to the control. The probiotic intake slowed down the age-related decline in mitochondrial function that is characteristic of aging. The transmembrane potential of the mitochondrial membrane was significantly higher in old worms fed with LG2055 than in their peers fed with the standard E. coli OP50. At the same time, life extension was observed both when feeding with live and dead LG2055 cells.
Emerging data indicate the role of ROS, nitric oxide, short-chain fatty acids, and hydrogen sulfide in cross-linking between microbiota-mitochondria and redox signaling [122]. Several studies show that the microbiota modulates mitochondrial activity and enhances the interaction between the host and the microbiota. Moreover, the effects can be both positive and negative, depending on which strain is involved — pathogenic or probiotic [123, 124]. Apparently, the microbiota can control mitochondrial activity and redox homeostasis [122].
We have previously shown the effect of probiotic Bacillus strains on mitochondrial DNA stability in birds [18]. It was shown that B. subtilis strains caused an increase in the expression of the genes associated with antioxidant activity in the liver and mitochondria compared as compared to the control group [62]. Biogenic selenium nanoparticles synthesized by L. casei ATCC 393 can protect the barrier function of the intestinal epithelium from oxidative damage by alleviating ROS-mediated mitochondrial dysfunction via the Nrf2 signaling pathway [125]; the positive effect of probiotics in Alzheimer’s disease also appears to be associated with effects on redox homeostasis, DNA damage levels, and mitochondrial activity [62].
There are three main mechanisms for the implementation of these effects that are being discussed in related publications:
-
1.
Microbiota can control mitochondrial activity and redox homeostasis.
-
2.
Microbiota can influence the expression of nuclear genes by stimulating the insertion of bacterial DNA.
-
3.
Mitochondrial DNA insertions occur in the host's somatic cells and may be triggered by microbiota activity.
Concerning point one, apparently, the mediator molecules secreted by the microbiota modulate mitochondrial activity and biogenesis. Depending on their concentration, these molecules affect mitochondrial homeostasis, which controls various cellular functions, in particular, ROS signaling, innate immune response, and energy metabolism [122]
The theory of endosymbiosis, according to which mitochondria originate from bacterial endosymbionts, also suggests that mitochondria may have signaling pathways that respond to bacterial signals [126, 127].
Thus, the antioxidant/antimutagenic effect on mitochondria should be considered one of the criteria for bacterial strains’ probiotic potential. Such a test might be difficult to perform at a stage of initial screening, but at the stage of animal tests, it is possible to measure the level of mutagenesis in mitochondria using the Comet Assay, PCR and any other available method.
The Effect of Complex Preparations
It is interesting to note that antimutagenic activity increases with the use of a complex of strains compared with a monoculture. Several studies report that probiotic bacteria work better in combination than individually. For example, a complex of four strains isolated from goats showed better activity than the same strains separately [28]. Given this, another way of realizing the antimutagenic properties of probiotic Bacillus strains seems to be possible: since probiotic Bacillus, in particular B. subtilis, improves the viability of normal intestinal microbiota, such as representatives of Lactobacillus and Bifidobacterium; this may, in turn, lead to active production of antimutagenic metabolites by the latter and, thus, improve the antimutagenic potential of microbiota in general. It was found that the viability of lactobacilli when combined with bacilli is increased significantly. The authors speculate that this effect may be due to the release of catalase and subtilisin from B. subtilis [128, 129].
The fact that often complex probiotic preparations can be more effective also makes sense in the context of the diversity of the spectrum of metabolites secreted by different groups of probiotics, since different groups of microorganisms secrete different antimutagenic metabolites, apparently, complementing each other’s activity (see, for example, [111, 112]).
Antimutagenic Action as a Criterion for Screening
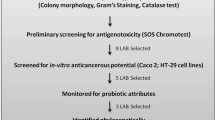
The current approach to selecting potential probiotics can be summarized in the following scheme (Fig. 1).
After isolation of strains from natural sources and their preliminary identification, they are checked for several criteria, such as safety (hemolytic activity, ability to adhere to mammalian cells, production of lytic enzymes, toxins, biogenic amines), biological activity, and ability to colonize the host and survive in the internal environment (such parameters as hydrophobicity of the cell surface, ability to adhere to mucin, to the intestinal epithelium, and autoaggregation screening are considered) [130]. These studies can be performed in a different order or simultaneously. However, they always precede the study of the effects of a probiotic directly on the host organism.
All screening procedures preceding animal and/or human testing are naturally aimed at predicting the strain’s probiotic properties before it is introduced into the host. There are various model systems based on cell cultures, single-cell biosensors, and in vitro tests. Biological activity, which is most often evaluated at stage 4, is understood very broadly in different studies. The main effects include antimicrobial, immunomodulatory (for example, in models of co-cultivation of bacteria with epithelial cells and immune cells that mimic in vivo interactions), anti-inflammatory, antitumor properties (this section sometimes includes antimutagenic), and the ability to interact with certain specific metabolites or produce them. All these tests are conducted quite randomly, making it difficult to compare the results of different studies [130].
We consider it essential to distinguish antimutagenic and antioxidant activity as a mandatory criterion for screening probiotic strains in vivo (Point 4A of Fig. 1) since it can be associated with many systemic effects.
The most common methods used to assess antimutagenic activity are summarized in Table 3.
Conclusion
Thus, we can conclude that antimutagenic activity is an important property of probiotic strains. Many data obtained based on in vitro experiments using model mutagens shows that many probiotic strains can inactivate these substances or reduce their effect, exhibiting, in particular, anticarcinogenic properties. The mechanisms of antimutagenic activity of probiotics can be associated with (a) binding of mutagens, (b) transformation of mutagens, and (c) inhibition of the transformation of promutagens into antimutagens. Effector molecules that carry out these processes can be part of cell structures, be secreted extracellularly, or be obtained due to the transformation of the substrate by bacteria. The possibility of an indirect decrease in the level of mutagenesis in cells due to the interaction of metabolites of probiotics with the host’s regulatory cascades requires separate consideration. Antimutagenic activity may be associated with the broader systemic effects of probiotics, such as geroprotective activity, and should be considered an essential criterion in selecting probiotic strains.
Also, it is interesting to study the effect of probiotics on mitochondria. The evolutionary relationship between bacteria and mitochondria suggests that mitochondria may have previously unknown signaling pathways that respond to bacterial signals. Besides, mitochondria play a significant role in the production and control of ROS in the cell. It means the pathways of probiotics’ systemic antioxidant effects can be implemented through them.
Probiotics with antimutagenic properties can be used as adjunctive therapy in the treatment of genotoxic drugs, as well as to prevent the mutagenic effect of environmental pollutants on humans and animals. However, the use of these strains should not be limited only to this area, since antimutagenic properties can lead to wider systemic effects that still require further study.
Availability of Data and Material
All data generated or analyzed during this study are included in this published article.
References
Mazanko MS, Chistyakov VA, Prazdnova EV et al (2016) Dioxidine induces bacterial resistance to antibiotics. Mol Gen Microbiol Virol 31:227–232. https://doi.org/10.3103/S0891416816040066
Silva MJ, Costa P, Dias A, Valente M, Louro H, Boavida MG (2005) Comparative analysis of the mutagenic activity of oxaliplatin and cisplatin in the Hprt gene of CHO cells. Env Mol Mutagenesis 46(2):104–115. https://doi.org/10.1002/em.20138
Chistyakov VA, Prazdnova EV, Mazanko MS, Churilov MN, Chmyhalo VK (2018) Increase in bacterial resistance to antibiotics after cancer therapy with platinum-based drugs. Mol Biol (Mosk) 52(2):232–236. https://doi.org/10.1134/S0026893317050077
Bhattacharya P, Mukherjee S, Mandal SM (2020) Fluoroquinolone antibiotics show genotoxic effect through DNA-binding and oxidative damage. Spectrochim Acta A Mol Biomol Spectrosc 227:117634. https://doi.org/10.1016/j.saa.2019.117634
DeBalsi KL, Hoff KE, Copeland WC (2017) Role of the mitochondrial DNA replication machinery in mitochondrial DNA mutagenesis, aging and age-related diseases. Ageing Res Rev 33:89–104. https://doi.org/10.1016/j.arr.2016.04.006
Giorgi C, Marchi S, Simoes ICM et al (2018) Mitochondria and reactive oxygen species in aging and age-related diseases. Int Rev Cell Mol Biol 340:209–344. https://doi.org/10.1016/bs.ircmb.2018.05.006
Zhang L, Vijg J (2018) Somatic mutagenesis in mammals and its implications for human disease and aging. Annu Rev Genet 52:397–419. https://doi.org/10.1146/annurev-genet-120417-031501
Petra AI, Panagiotidou S, Hatziagelaki E et al (2015) Gut-microbiota-brain axis and its effect on neuropsychiatric disorders with suspected immune dysregulation. Clin Ther 37:984–995. https://doi.org/10.1016/j.clinthera.2015.04.002
Zhou Y, Li S, Pang Q, Miao Z (2020) Bacillus amyloliquefaciens BLCC1-0238 can effectively improve laying performance and egg quality via enhancing immunity and regulating reproductive hormones of laying hens. Probiotics Antimicrob Proteins 12:246–252. https://doi.org/10.1007/s12602-019-9524-1
Li B, Etareri Evivie S, Lu J et al (2018) Lactobacillus helveticus KLDS1.8701 alleviates d -galactose-induced aging by regulating Nrf-2 and gut microbiota in mice. Food Funct 9:6586–6598. https://doi.org/10.1039/C8FO01768A
Vanderhoof J, Mitmesser S (2010) Probiotics in the management of children with allergy and other disorders of intestinal inflammation. Benef Microbes 1:351–356. https://doi.org/10.3920/BM2010.0034
Kanmani P, Satish Kumar R, Yuvaraj N et al (2013) Probiotics and its functionally valuable products-a review. Crit Rev Food Sci Nutr 53:641–658. https://doi.org/10.1080/10408398.2011.553752
Wu J, Zhang Y, Ye L, Wang C (2020) The anti-cancer effects and mechanisms of lactic acid bacteria exopolysaccharides in vitro: a review. Carbohydr Polym. 117308. https://doi.org/10.1016/j.carbpol.2020.117308
Pool-Zobel BL, Neudecker C, Domizlaff I et al (1996) Lactobacillus- and Bifidobacterium-mediated antigenotoxicity in the colon of rats. Nutr Cancer 26:365–380. https://doi.org/10.1080/01635589609514492
Reddy BS, Rivenson A (1993) Inhibitory effect of Bifidobacterium longum on colon, mammary, and liver carcinogenesis induced by 2-amino-3-methylimidazo[4,5-f]quinoline, a food mutagen. Cancer Res 53:3914–3918
Yun S-I (2004) The binding of Bacillus natto isolated from natto to heterocyclic pyrolysates. World J Microbiol Biotechnol 20:469–474. https://doi.org/10.1023/B:WIBI.0000040396.11236.e0
Hong HA, Duc LH, Cutting SM (2005) The use of bacterial spore formers as probiotics. FEMS Microbiol Rev 29:813–835. https://doi.org/10.1016/j.femsre.2004.12.001
Makarenko MS, Chistyakov VA, Usatov AV et al (2019) The impact of Bacillus subtilis KATMIRA1933 supplementation on telomere length and mitochondrial DNA damage of laying hens. Probiotics Antimicrob Proteins 11:588–593. https://doi.org/10.1007/s12602-018-9440-9
Prazdnova EV, Mazanko MS, Bren AB et al (2019) SOS response inhibitory properties by potential probiotic formulations of Bacillus amyloliquefaciens B-1895 and Bacillus subtilis KATMIRA1933 obtained by solid-state fermentation. Curr Microbiol 76:312–319. https://doi.org/10.1007/s00284-018-01623-2
Sreekumar O, Hosono A (2011) The antimutagenic properties of a polysaccharide produced by Bifidobacterium longum and its cultured milk against some heterocyclic amines. Can J Microbiol 44(11):1029–1036. https://doi.org/10.1139/w98-103
Lo P-R, Yu R-C, Chou C-C, Huang E-C (2004) Determinations of the antimutagenic activities of several probiotic bifidobacteria under acidic and bile conditions against benzo[a]pyrene by a modified Ames test. Int J Food Microbiol 93:249–257. https://doi.org/10.1016/j.ijfoodmicro.2003.11.008
Lankaputhra WEV, Shah NP (1998) Antimutagenic properties of probiotic bacteria and of organic acids. Mutat Res 397:169–182. https://doi.org/10.1016/S0027-5107(97)00208-X
Commane D, Hughes R, Shortt C, Rowland I (2005) The potential mechanisms involved in the anti-carcinogenic action of probiotics. Mutat Res 591:276–289. https://doi.org/10.1016/j.mrfmmm.2005.02.027
Ljungh A, Wadström T (2006) Lactic acid bacteria as probiotics. Curr Issues Intest Microbiol 7:73–89
Nissle A (1918) Die antagonistische behandlung chronischer darmstörungen mit colibakterien. Med Klin 2:29–30
Janosch D, Dubbert S, Eiteljörge K et al (2019) Anti-genotoxic and anti-mutagenic activity of Escherichia coli Nissle 1917 as assessed by in vitro tests. Benef Microbes 10:449–461. https://doi.org/10.3920/BM2018.0113
Bocci A, Sebastiani B, Trotta F, Federici E, Cenci G (2015) In vitro inhibition of 4-nitroquinoline-1-oxide genotoxicity by probiotic Lactobacillus rhamnosus IMC501. J Microbiol Biotechnol 25:1680–1686. https://doi.org/10.4014/jmb.1501.01086
Apás AL, González SN, Arena ME (2014) Potential of goat probiotic to bind mutagens. Anaerobe 28:8–12. https://doi.org/10.1016/j.anaerobe.2014.04.004
Knasmüller S, Steinkellner H, Hirschl AM et al (2001) Impact of bacteria in dairy products and of the intestinal microflora on the genotoxic and carcinogenic effects of heterocyclic aromatic amines. Mutat Res 480–481:129–138. https://doi.org/10.1016/S0027-5107(01)00176-2
Chalova VI, Lingbeck JM, Kwon YM, Ricke SC (2008) Extracellular antimutagenic activities of selected probiotic Bifidobacterium and Lactobacillus spp. as a function of growth phase. J Environ Sci Health B 43:193–198. https://doi.org/10.1080/03601230701795262
Hosoda M, Hashimoto H, Morita H et al (1992) Antimutagenicity of milk cultured with lactic acid bacteria against N-methyl-N’-nitro-N-nitrosoguanidine. J Dairy Sci 75:976–981. https://doi.org/10.3168/jds.S0022-0302(92)77839-4
Kim HS, Lee SY, Kang HJ et al (2019) Effects of six different starter cultures on mutagenicity and biogenic amine concentrations in fermented sausages treated with vitamins C and E. Food Sci Anim Resour 39:877–887. https://doi.org/10.5851/kosfa.2019.e66
Bodana AR, Rao DR (1990) Antimutagenic activity of milk fermented by Streptococcus thermophilus and Lactobacillus bulgaricus. J Dairy Sci 73:3379–3384. https://doi.org/10.3168/jds.S0022-0302(90)79033-9
Bakalinsky AT, Nadathur SR, Carney JR, Gould SJ (1996) Antimutagenicity of yogurt. Mutat Res 350:199–200. https://doi.org/10.1016/0027-5107(95)00113-1
Irecta-Nájera CA, Del Rosario H-López M, Casas-Solís J et al (2017) Protective effect of Lactobacillus casei on DMH-induced colon carcinogenesis in mice. Probiotics Antimicrob Proteins 9:163–171. https://doi.org/10.1007/s12602-017-9253-2
Sharma M, Chandel D, Shukla G (2020) Antigenotoxicity and cytotoxic potentials of metabiotics extracted from isolated probiotic, Lactobacillus rhamnosus MD 14 on Caco-2 and HT-29 human colon cancer cells. Nutr Cancer 72:110–119. https://doi.org/10.1080/01635581.2019.1615514
Ahmadi MA, Ebrahimi MT, Mehrabian S et al (2014) Antimutagenic and anticancer effects of lactic acid bacteria isolated from Tarhana through Ames test and phylogenetic analysis by 16S rDNA. Nutr Cancer 66:1406–1413. https://doi.org/10.1080/01635581.2014.956254
Faridnia F, Hussin A, Saari N et al (2010) In vitro binding of mutagenic heterocyclic aromatic amines by Bifidobacterium pseudocatenulatum G4. Benef Microbes 1:149–154. https://doi.org/10.3920/BM2009.0035
Bolognani F, Rumney CJ, Rowland IR (1997) Influence of carcinogen binding by lactic acid-producing bacteria on tissue distribution and in vivo mutagenicity of dietary carcinogens. Food Chem Toxicol 35:535–545. https://doi.org/10.1016/S0278-6915(97)00029-X
Caldini G, Trotta F, Cenci G (2002) Inhibition of 4-nitroquinoline-1-oxide genotoxicity by Bacillus strains. Res Microbiol 153:165–171. https://doi.org/10.1016/S0923-2508(02)01302-5
Cenci G, Caldini G, Trotta F, Bosi P (2008) In vitro inhibitory activity of probiotic spore-forming bacilli against genotoxins. Lett Appl Microbiol l46:331–337. https://doi.org/10.1111/j.1472-765X.2007.02314.x
Jiang T, Qiao H, Zheng Z et al (2016) Lactic acid production from pretreated hydrolysates of corn stover by a newly developed Bacillus coagulans strain. PLoS ONE 11:e0149101. https://doi.org/10.1371/journal.pone.0149101
Vorob’eva LI, Cherdyntseva TA, Abilev SK, (1995) Antimutagenic action of bacterial culture liquid on mutagenesis induced by 2-nitrofluorene in Salmonella typhimurium strains. Genetika 31:901–907
Belicová A, Križková L, Dobias J et al (2004) Synergic activity of selenium and probiotic bacterium Enterococcus faecium M-74 against selected mutagens in Salmonella assay. Folia Microbiol 49:301–305. https://doi.org/10.1007/BF02931047
Chistyakov VA, Prazdnova EV, Mazanko MS, Bren AB (2018) The use of biosensors to explore the potential of probiotic strains to reduce the SOS response and mutagenesis in bacteria. Biosensors 8:25. https://doi.org/10.3390/bios8010025
Verdenelli MC et al (2010) Investigation of the antigenotoxic properties of the probiotic Lactobacillus rhamnosus IMC 501® by gas chromatography-mass spectrometry. Ital J Food Sci 22(4):473–478
Stidl R, Sontag G, Koller V, Knasmüller S (2008) Binding of heterocyclic aromatic amines by lactic acid bacteria: Results of a comprehensive screening trial. Mol Nutr Food Res 52:322–329. https://doi.org/10.1002/mnfr.200700034
Nadathur SR, Gould SJ, Bakalinsky AT (1994) Antimutagenicity of fermented milk. J Dairy Sci 77:3287–3295. https://doi.org/10.3168/jds.S0022-0302(94)77269-6
Chang JH, Shim YY, Cha SK, Chee KM (2010) Probiotic characteristics of lactic acid bacteria isolated from kimchi. J Appl Microbiol 109(1):220–230. https://doi.org/10.1111/j.1365-2672.2009.04648.x
Kumar A, Singh NK, Sinha PR (2010) Inhibition of 1,2-dimethylhydrazine induced colon genotoxicity in rats by the administration of probiotic curd. Mol Biol Rep 37(3):1373–1376. https://doi.org/10.1007/s11033-009-9519-1
Ambalam P, Dave JM, Nair BM, Vyas BR (2011) In vitro mutagen binding and antimutagenic activity of human Lactobacillus rhamnosus 231. Anaerobe 17(5):217–222. https://doi.org/10.1016/j.anaerobe.2011.07.001
Chooruk A, Piwat S, Teanpaisan R (2017) Antioxidant activity of various oral Lactobacillus strains. J Appl Microbiol 123:271–279. https://doi.org/10.1111/jam.13482
Li Z, Teng J, Lyu Y et al (2019) Enhanced antioxidant activity for apple juice fermented with Lactobacillus plantarum ATCC14917. Molecules 24:51. https://doi.org/10.3390/molecules24010051
Song MW, Jang HJ, Paik K-TK, H-D, (2019) Probiotic and antioxidant properties of novel Lactobacillus brevis KCCM 12203P isolated from kimchi and evaluation of immune-stimulating activities of its heat-killed cells in RAW 264.7 cells. J Microbiol Biotechnol 29:1894–1903. https://doi.org/10.4014/jmb.1907.07081
Huang G, Pan H, Zhu Z, Li Q (2020) The complete genome sequence of Bifidobacterium longum LTBL16, a potential probiotic strain from healthy centenarians with strong antioxidant activity. Genomics 112:769–773. https://doi.org/10.1016/j.ygeno.2019.05.015
Bernini LJ, Simão ANC, de Souza CHB et al (2018) Effect of Bifidobacterium lactis HN019 on inflammatory markers and oxidative stress in subjects with and without the metabolic syndrome. Br J Nutr 120:645–652. https://doi.org/10.1017/S0007114518001861
Wang B, Xu H, Xu F et al (2015) Efficacy of oral Bifidobacterium bifidum ATCC 29521 on microflora and antioxidant in mice. Can J Microbiol 62(3):249–262. https://doi.org/10.1139/cjm-2015-0685
Pieniz S, Andreazza R, Okeke BC et al (2015) Antimicrobial and antioxidant activities of Enterococcus species isolated from meat and dairy products. Braz J Biol 75:923–931. https://doi.org/10.1590/1519-6984.02814
Riane K, Ouled-Haddar H, Alyane M et al (2019) Assessment of Streptococcus salivarius sp thermophilus antioxidant efficiency and its role in reducing paracetamol hepatotoxicity. Iran J Biotechnol 17:58–66. https://doi.org/10.30498/ijb.2019.91761
Rahman MdS, Hee Choi Y, Seok Choi Y et al (2018) A novel antioxidant peptide, purified from Bacillus amyloliquefaciens, showed strong antioxidant potential via Nrf-2 mediated heme oxygenase-1 expression. Food Chem 239:502–510. https://doi.org/10.1016/j.foodchem.2017.06.106
Wang Y, Wu Y, Wang Y et al (2017) Bacillus amyloliquefaciens SC06 alleviates the oxidative stress of IPEC-1 via modulating Nrf2/Keap1 signaling pathway and decreasing ROS production. Appl Microbiol Biotechnol 101:3015–3026. https://doi.org/10.1007/s00253-016-8032-4
Ton AMM, Campagnaro BP, Alves GA et al (2020) Oxidative stress and dementia in Alzheimer’s patients: effects of synbiotic supplementation. Oxid Med Cell Longev 2020:2638703. https://doi.org/10.1155/2020/2638703
Morotomi M, Mutal M (1986) In vitro binding of potent mutagenic pyrolyzates to intestinal bacteria. J Natl Cancer Inst 77:195–201. https://doi.org/10.1093/jnci/77.1.195
Zhang XB, Ohta Y (1991) Binding of mutagens by fractions of the cell wall skeleton of lactic acid bacteria on mutagens. J Dairy Sci 74:1477–1481. https://doi.org/10.3168/jds.S0022-0302(91)78306-9
Zhang XB, Ohta Y (1993) Antimutagenicity of cell fractions of microorganisms on potent mutagenic pyrolysates. Mutat Res Genet Toxicol Environ Mutagen 298:247–253. https://doi.org/10.1016/0165-1218(93)90003-V
Zhang XB, Ohta Y, Hosono A (1990) Antimutagenicity and binding of lactic acid bacteria from a Chinese cheese to mutagenic pyrolyzates. J Dairy Sci 73:2702–2710. https://doi.org/10.3168/jds.S0022-0302(90)78955-2
Cummings J (1983) Fermentation in the human large intestine: evidence and implications for health. Lancet 321:1206–1209. https://doi.org/10.1016/S0140-6736(83)92478-9
Aguilar-Toalá JE, Santiago-López L, Peres CM et al (2017) Assessment of multifunctional activity of bioactive peptides derived from fermented milk by specific Lactobacillus plantarum strains. J Dairy Sci 100:65–75. https://doi.org/10.3168/jds.2016-11846
Hsieh M-L, Chou C-C (2006) Mutagenicity and antimutagenic effect of soymilk fermented with lactic acid bacteria and bifidobacteria. Int J Food Microbiol 111:43–47. https://doi.org/10.1016/j.ijfoodmicro.2006.04.034
Hsieh M-L, Fang SW, Yu R-C, Chou C-C (2007) Possible mechanisms of antimutagenicity in fermented soymilk prepared with a coculture of Streptococcus infantis and Bifidobacterium infantis. J Food Prot 70:1025–1028. https://doi.org/10.4315/0362-028X-70.4.1025
Sah BNP, Vasiljevic T, McKechnie S, Donkor ON (2014) Effect of probiotics on antioxidant and antimutagenic activities of crude peptide extract from yogurt. Food Chem 156:264–270. https://doi.org/10.1016/j.foodchem.2014.01.105
Rhee C-H, Park H-D (2001) Three glycoproteins with antimutagenic activity identified in Lactobacillus plantarum KLAB21. Appl Environ Microbiol 67:3445–3449. https://doi.org/10.1128/AEM.67.8.3445-3449.2001
Nakagawa H, Shiozaki T, Kobatake E et al (2016) Effects and mechanisms of prolongevity induced by Lactobacillus gasseri SBT2055 in Caenorhabditis elegans. Aging Cell 15:227–236. https://doi.org/10.1111/acel.12431
Savustyanenko AV (2016) Mechanisms of action of probiotics based on Bacillus subtilis. Actual Infectology [RU] 2(11):35–44. https://doi.org/10.22141/2312-413x.2.11.2016.77529
Ahire JJ, Mokashe NU, Patil HJ, Chaudhari BL (2013) Antioxidative potential of folate producing probiotic Lactobacillus helveticus CD6. J Food Sci Technol 50:26–34. https://doi.org/10.1007/s13197-011-0244-0
Landis GN, Tower J (2005) Superoxide dismutase evolution and life span regulation. Mech Ageing Dev 126:365–379. https://doi.org/10.1016/j.mad.2004.08.012
Kullisaar T, Zilmer M, Mikelsaar M et al (2002) Two antioxidative lactobacilli strains as promising probiotics. Int J Food Microbiol 72:215–224. https://doi.org/10.1016/S0168-1605(01)00674-2
Kim HS, Chae HS, Jeong SG et al (2005) In vitro antioxidative properties of lactobacilli. Asian-australas J Anim Sci 19:262–265.
LeBlanc JG, del Carmen S, Miyoshi A et al (2011) Use of superoxide dismutase and catalase producing lactic acid bacteria in TNBS induced Crohn’s disease in mice. J Biotechnol 151:287–293. https://doi.org/10.1016/j.jbiotec.2010.11.008
Zulueta A, Maurizi A, Frígola A et al (2009) Antioxidant capacity of cow milk, whey and deproteinized milk. Int Dairy J 19:380–385. https://doi.org/10.1016/j.idairyj.2009.02.003
Power O, Jakeman P, FitzGerald RJ (2013) Antioxidative peptides: enzymatic production, in vitro and in vivo antioxidant activity and potential applications of milk-derived antioxidative peptides. Amino Acids 44:797–820. https://doi.org/10.1007/s00726-012-1393-9
Gjorgievski N, Tomovska J, Dimitrovska G et al (2014) Determination of the antioxidant activity in yogurt. J HygEng Des 8:88–92
Solieri L, Rutella GS, Tagliazucchi D (2015) Impact of non-starter lactobacilli on release of peptides with angiotensin-converting enzyme inhibitory and antioxidant activities during bovine milk fermentation. Food Microbiol 51:108–116. https://doi.org/10.1016/j.fm.2015.05.012
Gagnon M, Savard P, Rivière A et al (2015) Bioaccessible antioxidants in milk fermented by Bifidobacterium longum subsp. longum strains. Bio Med Res Int 2015:16938. https://doi.org/10.1155/2015/169381
Kullisaar T, Songisepp E, Aunapuu M et al (2010) Complete glutathione system in probiotic Lactobacillus fermentum ME-3. Appl Biochem Microbiol 46:481–486. https://doi.org/10.1134/S0003683810050030
BerniCanani R, Sangwan N, Stefka AT et al (2016) Lactobacillus rhamnosus GG-supplemented formula expands butyrate-producing bacterial strains in food allergic infants. ISME J10:742–750. https://doi.org/10.1038/ismej.2015.151
Canani RB, Filippis FD, Nocerino R et al (2017) Specific signatures of the gut microbiota and increased levels of butyrate in children treated with fermented cow’s milk containing heat-killed Lactobacillus paracasei CBA L74. Appl Environ Microbiol 83(19):e01206-e1217. https://doi.org/10.1128/AEM.01206-17
Endo H, Niioka M, Kobayashi N et al (2013) Butyrate-producing probiotics reduce nonalcoholic fatty liver disease progression in rats: new insight into the probiotics for the gut-liver axis. PLoS One 8:e63388. https://doi.org/10.1371/journal.pone.0063388
Lutgendorff F, Nijmeijer RM, Sandström PA et al (2009) Probiotics prevent intestinal barrier dysfunction in acute pancreatitis in rats via induction of ileal mucosal glutathione biosynthesis. PLoS One 4:e4512. https://doi.org/10.1371/journal.pone.0004512
Yang H, Li X, Li X et al (2015) Identification of lipopeptide isoforms by MALDI-TOF-MS/MS based on the simultaneous purification of iturin, fengycin, and surfactin by RP-HPLC. Anal Bioanal Chem 407:2529–2542. https://doi.org/10.1007/s00216-015-8486-8
Vasilchenko AS, Rogozhin EA (2019) Sub-inhibitory effects of antimicrobial peptides. Front Microbiol 10:1160. https://doi.org/10.3389/fmicb.2019.01160
Kozina LS (2007) Effects of bioactive tetrapeptides on free-radical processes. Bull Exp Biol Med 143:744–746. https://doi.org/10.1007/s10517-007-0230-8
Christen S, Peterhans E, Stocker R (1990) Antioxidant activities of some tryptophan metabolites: possible implication for inflammatory diseases. Proc Natl Acad Sci U S A 87:2506–2510. https://doi.org/10.1073/pnas.87.7.2506
Chalisova NI, Lopatina NG, Kamishev NG et al (2012) Effect of tripeptide lys-glu-asp on physiological activity of neuroimmunoendocrine system cells. Bull Exp Biol Med 153:569–572. https://doi.org/10.1007/s10517-012-1768-7
Ashapkin V, Khavinson V, Shilovsky G et al (2020) Gene expression in human mesenchymal stem cell aging cultures: modulation by short peptides. Mol Biol Rep 47:4323–4329. https://doi.org/10.1007/s11033-020-05506-3
Jones RM, Desai C, Darby TM et al (2015) Lactobacilli modulate epithelial cytoprotection through the Nrf2 pathway. Cell Rep 12:1217–1225. https://doi.org/10.1016/j.celrep.2015.07.042
Diao Y, Xin Y, Zhou Y et al (2014) Extracellular polysaccharide from Bacillus sp. strain LBP32 prevents LPS-induced inflammation in RAW 264.7 macrophages by inhibiting NF-κB and MAPKs activation and ROS production. Int Immunopharmacol 18:12–19. https://doi.org/10.1016/j.intimp.2013.10.021
Seth A, Yan F, Polk DB, Rao RK (2008) Probiotics ameliorate the hydrogen peroxide-induced epithelial barrier disruption by a PKC- and MAP kinase-dependent mechanism. Am J Physiol Gastrointest Liver Physiol 294:G1060–G1069. https://doi.org/10.1152/ajpgi.00202.2007
Mazzoli A, Donadio G, Lanzilli M et al (2019) Bacillus megaterium SF185 spores exert protective effects against oxidative stress in vivo and in vitro. Sci Rep 9:12082. https://doi.org/10.1038/s41598-019-48531-4
Shen Q, Shang N, Li P (2011) In vitro and in vivo antioxidant activity of Bifidobacterium animalis 01 isolated from centenarians. Curr Microbiol 62:1097–1103. https://doi.org/10.1007/s00284-010-9827-7
Achuthan AA, Duary RK, Madathil A et al (2012) Antioxidative potential of lactobacilli isolated from the gut of Indian people. Mol Biol Rep 39:7887–7897. https://doi.org/10.1007/s11033-012-1633-9
Wegh CAM, Geerlings SY, Knol J et al (2019) Postbiotics and their potential applications in early life nutrition and beyond. Int J Mol Sci 20:4673. https://doi.org/10.3390/ijms20194673
Hoarau C, Lagaraine C, Martin L et al (2006) Supernatant of Bifidobacterium breve induces dendritic cell maturation, activation, and survival through a toll-like receptor 2 pathway. J Allergy Clin Immunol 117:696–702. https://doi.org/10.1016/j.jaci.2005.10.043
Ménard S, Laharie D, Asensio C et al (2005) Bifidobacterium breve and Streptococcus thermophilus secretion products enhance T helper 1 immune response and intestinal barrier in mice. Exp Biol Med (Maywood) 230:749–756. https://doi.org/10.1177/153537020523001008
Corsello G, Carta M, Marinello R et al (2017) Preventive effect of cow’s milk fermented with Lactobacillus paracasei CBA L74 on common infectious diseases in children: a multicenter randomized controlled trial. Nutrients 9:669. https://doi.org/10.3390/nu9070669
De Marco S, Sichetti M, Muradyan D et al (2018) Probiotic cell-free supernatants exhibited anti-inflammatory and antioxidant activity on human gut epithelial cells and macrophages stimulated with LPS. Evid Based Complement Alternat Med 2018:1756308. https://doi.org/10.1155/2018/1756308
Wang K, Li W, Rui X et al (2014) Characterization of a novel exopolysaccharide with antitumor activity from Lactobacillus plantarum 70810. Int J Biol Macromol 63:133–139. https://doi.org/10.1016/j.ijbiomac.2013.10.036
Li W, Ji J, Chen X et al (2014) Structural elucidation and antioxidant activities of exopolysaccharides from Lactobacillus helveticus MB2-1. CarbohydrPolym 102:351–359. https://doi.org/10.1016/j.carbpol.2013.11.053
Khalil ES, Abd Manap MY, Mustafa S et al (2018) Probiotic properties of exopolysaccharide-producing Lactobacillus strains isolated from tempoyak. Molecules 23:398. https://doi.org/10.3390/molecules23020398
Skulachev VP (2010) How to cancel the program of body aging? Russ J Gen Chem 80:1523–1541. https://doi.org/10.1134/S1070363210070492
Skulachev VP (2011) Aging as a particular case of phenoptosis, the programmed death of an organism (A response to Kirkwood and Melov “On the programmed/non-programmed nature of ageing within the life history”). Aging 3:1120–1123. https://doi.org/10.18632/aging.100403
Anisimov VN, Egorov MV, Krasilshchikova MS et al (2011) Effects of the mitochondria-targeted antioxidant SkQ1 on lifespan of rodents. Aging 3:1110–1119. https://doi.org/10.18632/aging.100404
Chae S, Ahn BY, Byun K et al (2013) A systems approach for decoding mitochondrial retrograde signaling pathways. Sci Signal 6(264):rs4. https://doi.org/10.1126/scisignal.2003266
Shinjo S, Jiang S, Nameta M et al (2017) Disruption of the mitochondria-associated ER membrane (MAM) plays a central role in palmitic acid–induced insulin resistance. Exp Cell Res 359:86–93. https://doi.org/10.1016/j.yexcr.2017.08.006
Bajpai P, Darra A, Agrawal A (2018) Microbe-mitochondrion crosstalk and health: an emerging paradigm. Mitochondrion 39:20–25. https://doi.org/10.1016/j.mito.2017.08.008
Zolotukhin P, Kozlova Y, Dovzhik A et al (2013) Oxidative status interactome map: towards novel approaches in experiment planning, data analysis, diagnostics and therapy. Mol Biosyst 9:2085–2096. https://doi.org/10.1039/C3MB70096H
Lo S-C, Hannink M (2008) PGAM5 tethers a ternary complex containing Keap1 and Nrf2 to mitochondria. Exp Cell Res 314:1789–1803. https://doi.org/10.1016/j.yexcr.2008.02.014
Niture SK, Jain AK, Shelton PM, Jaiswal AK (2011) Src subfamily kinases regulate nuclear export and degradation of transcription factor Nrf2 to switch off Nrf2-mediated antioxidant activation of cytoprotective gene expression. J Biol Chem 286:28821–28832. https://doi.org/10.1074/jbc.M111.255042
Zolotukhin PV, Belanova AA, Prazdnova EV et al (2016) Mitochondria as a signaling Hub and target for phenoptosis shutdown. Biochemistry 81:329–337. https://doi.org/10.1134/S0006297916040039
Stefanaki C, Bacopoulou F, Michos A (2018) The impact of probiotics’ administration on glycemic control, body composition, gut microbiome, mitochondria, and other hormonal signals in adolescents with prediabetes – a randomized, controlled trial study protocol. Contemp Clin Trials Commun 11:55–62. https://doi.org/10.1016/j.conctc.2018.06.002
Zorov DB, Plotnikov EY, Silachev DN et al (2014) Microbiota and mitobiota. Putting an equal sign between mitochondria and bacteria. Biochemistry 79:1017–1031. https://doi.org/10.1134/S0006297914100046
Saint-Georges-Chaumet Y, Attaf D, Pelletier E, Edeas M (2015) Targeting microbiota-mitochondria inter-talk: microbiota control mitochondria metabolism. Cell Mol Biol (Noisy-le-grand) 61:121–124
Edeas M, Weissig V (2013) Targeting mitochondria: strategies, innovations and challenges: the future of medicine will come through mitochondria. Mitochondrion 13:389–390. https://doi.org/10.1016/j.mito.2013.03.009
Prasun P (2019) Mitochondrial Medicine. In: Prasun P (ed) Chapter 1 - functions of mitochondria. Academic Press, pp 1–3
Xu C, Qiao L, Ma L et al (2019) Biogenic selenium nanoparticles synthesized by Lactobacillus casei ATCC 393 alleviate intestinal epithelial barrier dysfunction caused by oxidative stress via Nrf2 signaling-mediated mitochondrial pathway. Iran J Neonatol 14:4491–4502. https://doi.org/10.2147/IJN.S199193
Pallen MJ (2011) Time to recognise that mitochondria are bacteria? Trends Microbiol 19:58–64. https://doi.org/10.1016/j.tim.2010.11.001
Lobet E, Letesson J-J, Arnould T (2015) Mitochondria: a target for bacteria. Biochem Pharmacol 94:173–185. https://doi.org/10.1016/j.bcp.2015.02.007
Hosoi T, Ametani A, Kiuchi K, Kaminogawa S (2011) Improved growth and viability of lactobacilli in the presence of Bacillus subtilis (natto), catalase, or subtilisin. Can J Microbiol 46(10):892–897. https://doi.org/10.1139/w00-070
Zhang Y-R, Xiong H-R, Guo X-H (2014) Enhanced viability of Lactobacillus reuteri for probiotics production in mixed solid-state fermentation in the presence of Bacillus subtilis. Folia Microbiol 59:31–36. https://doi.org/10.1007/s12223-013-0264-4
Papadimitriou K, Zoumpopoulou G, Foligné B et al (2015) Discovering probiotic microorganisms: in vitro, in vivo, genetic and omics approaches. Front Microbiol 6:58. https://doi.org/10.3389/fmicb.2015.00058
Zeiger E (2019) The test that changed the world: the Ames test and the regulation of chemicals. Mutat Res 841:43–48. https://doi.org/10.1016/j.mrgentox.2019.05.007
Ragavan ML, Das N (2020) In vitro studies on therapeutic potential of probiotic yeasts isolated from various sources. Curr Microbiol 77(10):2821–2830. https://doi.org/10.1007/s00284-020-02100-5
Lee SB, Cosmas B, Park HD (2020) the antimutagenic and antioxidant activity of fermented milk supplemented with Cudrania tricuspidata powder. Foods 9(12):1762. https://doi.org/10.3390/foods9121762
Funding
EVP, MSM, VAC, AAB, AGR, and EYK were financially supported by the Ministry of Science and Higher Education of the Russian Federation within the framework of the state task in the field of scientific activity (Southern Federal University, no. 0852–2020-0029). VAC and MLC acknowledge the support of the Government of the Russian Federation (contract No. 075–15-2019–1880).
Author information
Authors and Affiliations
Contributions
Evgeniya V. Prazdnova conceived the original idea and performed the overall analysis. Maria S. Mazanko performed the analysis on antioxidant activity of probiotics. Vladimir A. Chistyakov supervised the project. Anna A. Bogdanova prepared figures and schemes and systematized the list of references. Aleksandr G.Refeld performed the analyses of recent reviews. Evgeniya Y.Kharchenko performed meta-analysis. Michael L. Chikindas was in charge of the overall direction and planning.
Corresponding author
Ethics declarations
Ethics Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Prazdnova, E.V., Mazanko, M.S., Chistyakov, V.A. et al. Antimutagenic Activity as a Criterion of Potential Probiotic Properties. Probiotics & Antimicro. Prot. 14, 1094–1109 (2022). https://doi.org/10.1007/s12602-021-09870-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12602-021-09870-9