Abstract
Bacteriophages (or phages), the most abundant viral entity of the planet, are omni-present in all the ecosystems. On the basis of their unique characteristics and anti-bacterial property, phages are being freshly evaluated taxonomically. Phages replicate inside the host either by lytic or lysogenic mode after infecting and using the cellular machinery of a bacterium. Since their discovery by Twort and d’Herelle in the early 1900s, phage became an important agent for combating pathogenic bacteria in clinical treatments and its related research gained momentum. However, due to recent emergence of bacterial resistance on antibiotics, applications of phage (phage therapy) become an inevitable option of research. Phage particles become popular as a biotechnological tool and treatment of pathogenic bacteria in a range of applied areas. However, there are few concerns over the application of phage-based solutions. This review deals with the updated phage taxonomy (ICTV 2015 Release and subsequent revision) and phage biology and the recent development of its application in the areas of biotechnology, biosensor, therapeutic medicine, food preservation, aquaculture diseases, pollution remediation, and wastewater treatment and issues related with limitations of phage-based remedy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There are billions of viruses on the Earth. An estimation of their number reaches an astronomical figure of about 1031 viruses, which is much more than the number of stars in the universe (Weitz and Wilhelm 2013). Most of the viruses play a significant role in the global biogeochemical cycles and thrive by infecting microbes like bacteria, archaea, and microeukaryotes (Suttle 2007). Because of their astounding numbers, and intimate relationship with different microbes, they control both host populations and ecosystem functions (Weitz and Wilhelm 2012). Among the total viral population, bacteriophages, popularly known as phages (Greek “phagein” meaning “to eat”), are the group of viruses that infect and devour bacteria. This potential antibacterial property of phage is unique (Jamalludeen et al. 2009).
Phages are considered to be the most diverse and abundant entity on earth and are thought to exist in every ecosystem (Maranger and Bird 1995; Hendrix 2002; Hanlon 2007; Le Romancer et al. 2007) ranging from extremely hot environments like hot springs, the Sahara, to extremely cold environments like polar inland waters (Breitbart et al. 2004; Glud and Middleboe 2004; Prigent et al. 2005; Suttle 2005; Säwström et al. 2008; Lin et al. 2010). Sea water is among the hugely diverse and most dense natural environments for phages and other viruses; for example, the surface seawater has a concentration of approximately 10 million viruses per milliliter (Breitbart 2012). Similarly, forest floors and agricultural soils usually harbor a phage count of approximately108–109 per gram of soil (dry mass; Williamson et al. 2003; Williamson et al. 2005). It is also reported that to carry out important ecological functions, phages express a variety of auxiliary metabolic genes (Breitbart 2012; Weitz and Wilhelm 2013).
A single phage particle can hunt for a specific bacterium species or a subset of the same species. After infecting a bacterium, phages replicate inside it. Once a bacterial cell is infected, phages multiply exponentially using the cellular machinery of the bacteria including protein-synthesizing and energy-generating systems. However, the method of propagation may either be lytic or lysogenic in nature. The lytic phages cause lysis of the host bacterial cells to release progeny viruses (Young et al. 2000). On the other hand, lysogenic or the temperate phages integrate their nucleic acid (genome) within the host bacterial cell and replicate along with the host, conferring new properties to the host bacteria (Brock et al. 1988). Phages are classified on the basis of certain criteria including its host specificity, morphology, nucleic acid type, mode of infection, morphogenesis, phylogeny, serology, sensitivity to physical and chemical agents, and its environment (Abeles et al. 1984). All bacterial genera, including the cyanobacteria, archaebacterial, and mycoplasmas, are known to be vulnerable to a plethora of phages.
Brief history of bacteriophage
British bacteriologist Ernest Hanbury Hankin, in 1896, first reported antibacterial activity in the waters of the river Ganga and the river Yamuna in India (Hankin 1896: http://icmr.nic.in/buapril02.pdf). The then unidentified antibacterial agent was also noted as filterable and heat labile; it was found to limit the spread of cholera epidemic (Hankin 1896; Van Helvoort 1992; Sulakvelidze et al. 2001). However, how the implications of Hankin’s findings related to the existence of bacteriophages remain dubious in the scientific society.
In 1915, Frederick William Twort FRS (October 22, 1877–March 20, 1950), an English bacteriologist, superintendent of the Brown Institute for Animals and professor of bacteriology at the University of London, while growing viruses in a laboratory condition, found zones of clearance in the form of “transmissible glassy transformation” with micrococcus bacteria. He also concluded that this agent multiplied itself in the process of killing the bacteria (Twort 1915, 1922, 1949).
On the other hand, Félix d’Herelle (April 25, 1873–February 22, 1949), a French-Canadian microbiologist working at the Pasteur Institute in Paris, observed the bacteriophage phenomenon (in the year 1917) on severe hemorrhagic dysentery among French troops stationed at Maisons-Laffitte. He had observed the same phenomenon in 1910 when he was studying microbiology due to an epizootic locust infection in Mexico. d’Herelle prepared bacteria-free filtrates from fecal samples of patients and incubated the filtrate with the bacterium Shigella, isolated from those patients. His primary goal was to develop a vaccine against bacterial dysentery. For observing the growth of the bacteria, an aliquot of the filtrate-bacteria mixture was spread on agar medium and incubated. d’Herelle found the appearance of small, clear zones, which he primarily termed taches, then taches vierges, and later, plaques. This finding was presented in September 1917 and subsequently published (d’Herelle 1917, 1930; Summers 1999; Sulakvelidze et al. 2001). d’Herelle also proposed that the phenomenon was caused by a bacteria parasitizing virus and named “bacteriophage”—phages implying to “eat” or “devour” bacteria. As per the recollection of d’Herelle, the name “bacteriophage” was decided together with his wife Marie, and the word came into existence on 18 October 1916—the day before their youngest daughter’s birthday (d’Herelle 1930; Summers 1999; Sulakvelidze et al. 2001). Thus, the credit of discovery of bacteriophages goes independently to two scientists: F.W. Twort and Félix d’Herelle. Primarily, the scientific society first called it “Twort-d’Herelle phenomenon,” and later, the “bacteriophage phenomenon” (d’Herelle 1949; Duckworth 1976; Sulakvelidze et al. 2001).
Structure of bacteriophage
Length of phages varies widely and usually ranges from 24 to 200 nm (Mayer 2016). T4 phages are among the largest phages that are approximately 200 nm in length and 80–100 nm wide. Phages having an icosahedral shape or a shape with 20 sides and filaments are also common (http://www.microrao.com/micronotes/bacteriophage.pdf). After the first commercial production of transmission electron microscope (called “Hypermicroscope”) by Siemens and Halske Company (Germany) in 1939, photomicrography of bacteriophages came into the existence. Ruska (1940) and Pfankuch and Kausche (1940), both working in the same company, reported separately in the same journal and on same date showing the first pictures of bacteriophages to the scientific world (Ruska 1940; Pfankuch and Kausche 1940; Ackermann, 2011a, b). On the other hand, in North America, Prebus and Hillier of University of Toronto constructed separately the first electron microscope in 1938, and subsequently, commercialized the same after years of research by Hiller at Radio Corporation of America (RCA, Princeton, NJ; Prebus and Hillier 1939; Haguenau et al. 2003; Ackermann 2011a, b). Luria and Anderson, in 1942, reported different unstained coli and staphylococcal phage particles, which were renamed later as T2 (T-even type; Luria and Anderson 1942; Ackermann 2011a, b). Despite continental difference, huge political unrest (because of World War II) and extreme limitations on scientific exchange, interestingly, both the group of scientists were aware of other progress in the development of this extremely important scientific instrument (Pfankuch and Kausche 1940; Ackermann 2011a, b). Later on, using the “negative contrast” technique in an electron microscope, Brenner and his team first reported a clearer image of bacteriophages in 1959 (Brenner and Horne 1959; Brenner et al. 1959). In subsequent years, Bradley and Kay (1960) and Bradley (1967) elaborated fine structures of phages. The basic features of a typical bacteriophage, in this case the T4 bacteriophage, include a “head” or capsid and a “tail” (Fig. 1).
The head or capsid structure, regardless of its size or shape, is a congregation of one or more protein subunits called protomers. The morphological subunit of a capsid is a capsomere which protects the viral nucleic acid (genome). The tail is a hollow tube through which the nucleic acid passes when the bacteriophage infects a bacterial host cell. Some phages do not possess a tail. The T4 phage has additional structures, namely the base plate and tail fibers attached to the tail that aid the phage in attaching itself to a bacterial cell (Leiman et al. 2003).
Taxonomy of phages
As mentioned earlier, there are billions and billions of phages. Deciphering taxonomic characterization is a challenging task, especially for such nano-sized phage particles. In 1967, phages were first classified by Bradley and were subsequently approved by the International Committee on Taxonomy of Viruses (ICTV) and total 111 phages were listed in classification (Wildy 1971). Bradley’s classification projected six basic morphological types of tailed phages that further, categorized on the basis of morphotypes (contractile tails, long and noncontractile tails, and short tails), small isometric ssDNA viruses, filamentous phages, and small ssRNA phages. The regulating body for the viral taxonomic system (ICTV) characterizes phage particles taking into consideration numerous parameters like host range, physical characteristics (such as structure, capsid size, and shape), type of genomic material (single or double-stranded DNA or RNA), genome size, and resistance to organic solvents (Murphy et al. 1995). More than 96 % of phages are tailed and carries dsDNA as genetic material; however, they may vary in shape like cubic, filamentous, and pleomorphic (Ackermann 2011a, b). Polyphasic taxonomy was revised and emphasis was given on genomic relationship (Thompson et al. 2015). Earlier in 2008, only 18 genera and 36 species were listed among three caudoviral families, Myoviridae, Podoviridae, and Siphoviridae. Subsequently, the order caudovirales expanded and an updated and detailed bacteriophage classification was presented in ICTV Release 2015 (http://www.ictvonline.org/virusTaxonomy.asp; Krupovic et al. 2016). Some alterations have also been suggested by committee, like replacement of word “phage” with “virus” in prokaryotic virus taxon names, omission of infix “like” from prokaryotic virus genus names, discouragement of use of “Phi” and other Greek letters in prokaryotic virus genera, exclusion of hyphens, and encouragement to use of host genus name in replacement of taxon names is being encouraged to avoid confusion related to phage action on specific bacterial host (Krupovic et al. 2016; Pietilä et al. 2016). The overview of the various bacteriophage families has been included in Table 1 compiled from (ICTV 2015 Release; EC 47, London, UK, July 2015 Email ratification 2016 (MSL #30) for ready reference to the readers.
Mechanism of proliferation
Specific receptors (like lipopolysaccharides, teichoic acids, proteins, and flagella) on the surface of the host bacteria are required for the phage to infect the bacteria. Due to this specificity of the receptor present on the bacterial cell surface, phages can infect specific hosts only. However, in solution, this interaction with the host is a random phenomenon for the phages. Bacterial type (Gram-negative and Gram-positive), growth conditions, and virulence also influence the phage to attach on the host’s surface (Rakhuba et al. 2010). The outer membrane of Gram-negative bacteria has an external lipopolysaccharide (LPS) layer and embedded outer membrane proteins (OMPs) for transport and diffusion of nutrients. These act as phage receptors, and in some infection strategies, they are essential for adsorption of phage particles as well (Bugla-Ploskonska et al. 2007; Rakhuba et al. 2010). In comparison, teichoic acids (peptidoglycan interspersed with acid polysaccharides) present in the Gram-positive bacteria cell wall act as receptors for their corresponding phages (Brown et al. 2013; León and Bastías 2015).
The penetration processes in bacterial cell also vary in different phage groups. In general, myoviridae phage inserts its genetic material into the bacterial cell by using a syringe-like movement of its tail (Fig. 2). After receptor recognition, in a reversible binding mode, the phage particle attaches its base plate with the bacterial surface utilizing the flexing activity of tail fibers. The phage takes sufficient time to make its surface binding irreversible. Thereafter, with the help of ATP, the contraction of its tail takes place, along with insertion of its genetic material. On the contrary, podoviridae phage, which is devoid of the tail part of the myoviridae phage, inserts its genetic material after enzymatically degrading a portion of the bacterial cell membrane using its small, tooth-like tail fibers (Rakhuba et al. 2010; Brown et al. 2013; León and Bastías 2015).
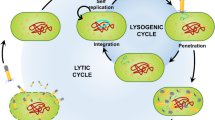
Phages undergo two possible life cycles: lytic and lysogenic cycles.
Lytic cycle
Phages, which are in their lytic phase, are called virulent phages. During the lytic cycle, lysis (death) of host bacterium occurs as the phage multiplies in great numbers within the host, following which the phage disintegrates and ruptures the host cell for release of new phage particles (Fig. 2).
The lytic cycle initiates with the attachment of the phage on the bacteria with the aid of a complex of proteins (Karlsson et al. 2003). Once the attachment of the viral particle is complete, the genetic material of the bacteriophage is inserted into the bacterial host cell. Upon penetration, the bacterial metabolic machinery is utilized by the phage to create multiple copies of its own genetic material (DNA or RNA). DNA viruses directly transcribe themselves into mRNA (messenger RNA) molecules that are then used to direct the host cell’s ribosomes. In case of RNA viruses (retroviruses), a unique enzyme—reverse transcriptase—transcribes the viral RNA into DNA, and thereafter, follows the path of DNA virus for transcription of the viral RNA. Towards the later stages of this translation, the newly translated proteins are assembled to form the capsid and the tail of the phages that break out of the host cell, bursting its cellular membrane. Each newly formed particle continues to infect new host cells and subsequently proliferates. In few cases, instead of the phage genome, host chromosome gets packed in to the capsid during phage replication and represents an example of horizontal gene transfer within the bacterial population via transduction (Madigan and Martinko 2006). A direct application of phages exhibiting only the lytic cycle is that they can be conveniently employed for tackling the problem of antibiotic-resistant pathogenic bacteria.
Lysogenic cycle
In comparison to the lytic cycle, the lysogenic phase is exhibited particularly by temperate phages (Campbell and Reece 2005), and results in the integration of the viral genetic material with the bacterial genome (called prophage), ensuring continued replication of the viral genetic material without any fatal consequences to the infected host (Inal 2003). However, due to incorporation of viral genetic material into the host, change in the phenotype of the infected bacteria is a common phenomenon. This conversion may induce the pathogenicity of bacteria, which is evident for many common bacterial strains (Brussow et al. 2004; Keen 2012). Prevention of such lysogenic conversion may also be done using hydrogen peroxide by production of a reactive oxygen species, glutathione, and overexpression of transcriptional repressors (Wagner et al. 2001; Liu et al. 2005; Ptashne 2006; Keen 2012).
Application of phages
Biotechnological relevance
Recently, phages have become an important instrument to biotechnologists. It is being studied for various purposes like drug designing, synthesis of novel protein, delivery of protein and DNA vaccines, detection of pathogenic bacteria, and screening of protein libraries, peptides, or antibodies (Hart et al. 1994; Sperinde et al. 2001; Clark and March 2004; Donnelly et al. 2015; Gao et al. 2015). Further, phages are being used as a substitute to antibiotics for many antibiotic-resistant bacterial strains, biocontrol agents in agriculture, aquaculture, and oil and petroleum industry (Haq et al. 2012). Researchers are also working upon synthesizing a novel peptide by exploiting phage display techniques, gene inclusion, and replication using host machinery (Smith 1985; Sidhu 2000; Haq et al. 2012; Wang et al. 2016). Phage display was first described by Smith (1985) to identify polypeptides with precise bait-binding activity; the technique has evolved with several versatile applications (Paschke 2006; Li and Caberoy 2010). The displayed molecule encoded by the phage genetic information will be expressed on its surface along with the coat protein. Researchers reported using phages like M13, Lambda, and T7 for phage display techniques (Benhar 2001; Willats 2002; Wang et al. 2016). A range of ligands can be prepared by fusing the coat protein gene with different gene-encoding peptides. Such ligands must have capabilities like detecting viable bacteria like Escherichia coli (Wang et al. 2016) and targeting specific pathogen proteins, recognizing specific receptors, and blocking interaction between ligand and receptor (Watson and Eveland 1965; Kodikara et al. 1991; Funatsu et al. 2002; Haq et al. 2012). Furthermore, bacteriophages are being modified as tunable nano-containers for the packaging and delivering of peptides (Kelly et al. 2015). In an interesting study, Dickerson et al. (2005) demonstrated the use of engineered filamentous bacteriophage in cocaine sequestration for blocking the behavioral effects of cocaine in rodent model. Furthermore, phage technology for detecting affinity of antibody to a particular antigen (pathogenic) or phage amplification for detecting pathogenic bacteria, like Mycobacterium tuberculosis, E. coli, Pseudomonas, etc., improvised research in this field (Stewart et al. 1989; Barry et al. 1996; Donnelly et al. 2015). Exploring phage sensitivity and specificity to design phage-based biosensors can be an improved alternative to antibody-based immunoassay techniques (Peltomaa et al. 2015).
Medicine and clinical application
About a century ago, in 1917, the bacteriophage era was started when Félix d’Herelle published a paper demonstrating “un bactériophage obligatoire” (d’Herelle 1917, 1949; Abedon et al. 2011). Bruynoghe and Maisin (1921) initiated bacteriophage therapy by treating patients having staphylococcal infections. “Phage therapy” has become a potential therapeutic option (Duckworth and Gulig 2002; Wright et al. 2009a, b; Sarker et al. 2012). Immense research has been initiated to employ phage therapy as an alternative (Matsuzaki et al. 2003; Quintin et al. 2005; Chatterjee et al. 2015; Cui 2015; Zschach et al. 2015; Abedon 2016; Maszewska et al. 2016). Phage based products were first developed in the commercial laboratory of d’Herelle in Paris. Treatment of various pathogens and antibiotic-resistance bacteria (like Salmonella spp. Clostridium difficile, and diarrheagenic E. coli) has been tried using phage (Mai et al. 2010; Frampton et al. 2012). Phage-based products were first developed in the commercial laboratory of d’Herelle in Paris. The company (later became the successful French company L’Oréal) produced five phage preparations (Bacté-coli-phage, Bacté-rhinophage, Bacté-intesti-phage, Bacté-pyo-phage, and Bacté-staphyphage) (Summers 1999; Sulakvelidze et al. 2001). In 1940s, Eli Lilly and Company (Indianapolis, IN, USA) developed seven phage products for human uses. These products were either in the form of a lysate in broth cultures (e.g., Colo-lysate, Ento-lysate, Neiso-lysate, and Staphylo-lysate) or in the form of a water soluble jelly (e.g., Colo-jel, Ento-jel, and Staphylo-jel) against targeted bacteria like, staphylococci, streptococci, and Escherichia coli. They were used for treating abscesses, septic wounds and vaginitis, mastoid infections, and respiratory tract infections (Sulakvelidze et al. 2001).
However, due to exponential growth of antibiotic-based drug companies and pharmaceutical giant companies, phage products became less popular (Alisky et al. 1998; Sulakvelidze and Kutter 2005). In favor of therapeutic uses of phages, recent reports on the application of phage cocktails for open septic wounds and burn injuries have contributed encouraging results. Phage cocktail containing 82 phages against Pseudomonas aeruginosa and 8 phages against Staphylococcus aureus was successfully applied on eight patients (Merabishvili et al. 2009; Wright et al., 2009a, b; Nilsson 2014). Recently, Pherecydes Pharma, a biotechnology company in France, have announced the phase I/II single-blind multicenter clinical trials of its bacteriophage based product ‘Phagoburn’ (http://www.pherecydes-pharma.com/phagoburn-clinical-study.html; Reardon 2014; Ravat et al. 2015; Servick 2016). Phagoburn contains bacteriophage cocktail targeting pathogenic strain of Escherichia coli and Pseudomonas aeruginosa and its infection in serious burn patients. The Phagoburn is a collaborative European project (www.phagoburn.eu) coordinated by French Ministry of Defense and being conducted in eleven major burns units in France, Switzerland and Belgium along with eight civilian hospitals. Phage therapy is important alternative to overcome critical limitations of antibiotic therapy due to emergence of bacterial resistance, like, in the case of Clostridium difficile infection (CDI). CDI is responsible for inducing the dysbiosis, which has extremely high recurrence rates. As per reports, treatment of CDI using current antibiotics has become more and more ineffective (Sangster et al. 2014). Further, prolonged use of antibiotics can also harm beneficial gut flora causing discomfort to the patients. Phage therapy against C. difficile involves specifically targeting the causative agent, sparing the other bacterial organisms of the human gut (Sangster et al. 2014, 2015). Similarly, treatment of nosocomial infections (common hospital-acquired infections) caused by Pseudomonas aeruginosa is a huge therapeutic challenge currently. High rate of morbidity and mortality is connected with the infection, along with a greater possibility of drug resistance in the bacteria during the course of therapy. With such limited scope for developing new drugs, scientists have also found considerable success in alternative treatment options including phage-based approaches (Viertel et al. 2014; Chatterjee et al. 2015). Babalova et al. (1968) used Shigella phages to treat bacterial dysentery and successfully controlled dysentery in patients. Till date, inadequate references are available on human clinical trials conducted and published; the reported cases are phase I studies and mostly from European Medicines Agency (EMA) or United States Food and Drug Administration (FDA) jurisdictions (Bruttin and Brüssow 2005; Merabishvili et al. 2009; Rhoads et al. 2009). In 2009, phase I clinical trial of bacteriophage cocktail (Biophage-PA) targeting three pathogenic strains of S. aureus, P. aeruginosa, and E. coli targeting venous ulcers was approved US FDA (Rhoads et al. 2009). Similarly, a randomized, double-blind, placebo-controlled phase I/II clinical trial approved by UK Medicines and Healthcare products Regulatory Agency (MHRA) and the Central Office for Research Ethics Committees (COREC) ethical review process was performed to treat chronic otitis against multidrug resistant bacteria P. aeruginosa and found lower bacterial count and significant improvement with a single input dose of 600,000 bacteriophages (Wright et al. 2009a, b). Bacteriophage application trials have shown that PFU count of 102–103 plaque forming unit (PFU) is adequate to counter 106–109 CFU per milliliter proliferation threshold of bacteria in vivo (Marza et al. 2006; Wright et al. 2009a, b; Parracho et al. 2012). However, the reported trials with bacteriophages have been performed with 105 and 109 PFU of individual bacteriophages (Bruttin and Brüssow 2005; Merabishvili et al. 2009; Rhoads et al. 2009; Wright et al. 2009a, b). Development of phage bioderm is another important area of application in clinical sector. It is a therapeutic autodegrading, non-toxic, biopolymer complex containing phages that help in healing of wounds and burns, osteomyelitis, and periodontal diseases (Mathur et al. 2003). Studies reported 20–95 % lysis of clinical isolates of Serratia marcescens using specific phage strains and phage type (Alisky et al. 1998). However, there are certain limitations on the application of phage bioderm, like reduction in phage activity due to development of antibodies against phages, induction of toxin genes, and fast discharge of bacterial endotoxins due to the lytic effect of phage (Mathur et al. 2003).
Biosensor development
IUPAC defines a biosensor as “a device that uses specific biochemical reactions mediated by isolated enzymes, immune-systems, tissues, organelles or whole cells to detect chemical compounds usually by electrical, thermal or optical signals” (Nagel et al. 1992; Hwang 2014). A biosensor is typically composed of bio-based recognition and transducer components, and electronic systems (signal amplification, processing, and display). Potential applications of biosensors are diverse and numerous, from defense security to pharmaceutical science to environmental monitoring and assessment (Kirsch et al. 2013). Biosensors have a number of advantages like sensitivity, specificity, speed and accuracy of detection, easy sample preparation, cost-effectiveness, etc. (Singh et al. 2013; Hwang 2014; Rackus et al. 2015). Like that of many other biomaterials, unique biological, geometrical, and mechanical characters of bacteriophages can be exploited for bacterial identification, pathogen detection, and biocontrol (Hwang 2014).
It is simpler to develop biosensor surfaces by surface adsorption of phages, though it may give inconsistent results due to unstable immobilization densities (Balasubramanian et al. 2007; Lakshmanan et al. 2007). However, chemically anchoring (cysteamine-modified and glutaraldehyde-activated gold substrate) phages on a detection platform display a consistent improvement in the phage density and detection (Cademartiri et al. 2010; Singh et al. 2013). For chemically functionalizing phage-based biosensors, selection (biopanning) purity of the phage suspension is an important criterion which should be devoid of various other bio-contaminating agents like other carbohydrates, proteins, and lipids (Naidoo et al. 2012). However, genetically modified or engineered phages are more appropriate to develop bio-probes than intact wild-type phages. Being biologically active, wild-type phages upon infection lyse the host bacterium that may lead to reduction of signal on a biosensor platform (Singh et al. 2010, 2013).
Recently, utilizing unique ability of phages to display peptides or proteins on their surface, called “phage display,” is becoming a powerful tool to screen diversity of targets like proteins, carbohydrates, small molecules, or an entire cell. For the phage display technology, lambda, f1, M13, fd, T4, and T7 phages have most widely been used (Smith 1985; Atias et al. 2008; Singh et al. 2013). This emerging technology can revolutionize diagnostics by creating molecules that are otherwise unavailable via conventional approaches. Researchers have successfully expressed cellular proteins and peptides, antibody (or its fragments), and antigen molecule on the surface of phages for developing pathogen detection biosensors, molecular imaging, and gene delivery (Petrenko 2008; Pande et al. 2010; Singh et al. 2013; Hwang 2014; Chuang et al. 2015).
Agriculture and food safety
Phage application in agriculture and food material is an upcoming area of development. Salmonella, Campylobacter, Listeria, and E. coli are common bacterial contaminants in food borne-related infections (DuPont 2007). Bacterial infection in crops is a grave problem that reduces the yields (Allen et al. 2011). High resistivity to multiple antibiotics was reported by a number of researchers in the agricultural field (Price et al. 2012; Zhu et al. 2013; Micallef et al. 2013; Popowska et al. 2012). To evade infection, a number of reports of phage application on various crops (like tomato, citrus, and onion) are available (Huff et al. 2005; Lang et al. 2007; Balogh et al. 2008; Jones et al. 2014). Phage products like AgriPhage™, Omnilytics are being used at a large scale for protecting crops from a number of bacterial diseases. Similarly, other phage products like LISTEX (for Listeria sp.; http://www.listex.eu/product/), Listshield™, and Intralytix were approved by the FDA (USA), for treating food products before they are being marketed (Meaden and Koskella 2013). Phages have been exploited to treat colibacillosis and prevent food-borne pathogens (Huff et al. 2005; Sharma et al. 2009). Phage therapy is applicable in restricting bacterial contamination on a variety of food materials like chicken flesh, meat, fruits, and vegetables (Flaherty et al. 2000; Goode et al. 2003; Allwood et al. 2004; Toro et al. 2005; Higgins et al. 2005; Fiorentin et al. 2005; Ravensdale et al. 2007; Sharma et al. 2009; Guenther et al. 2009; Marcó et al. 2012). Further, antimicrobial packaging against the activity of Listeria monocytogenes for cantaloupes and ready to eat meat and E. coli for alfalfa seeds and sprouts are available in the market (Lone et al. 2016).
Aquaculture industries
Aquaculture industries often undergo economic losses chiefly due to uncontrolled microbial diseases that threaten their development and sustainability (Almeida et al. 2009, Silva et al. 2014). Fish-based products are becoming more popular throughout the world as a cheap source of protein. For the last three decades, considerable growth of aquaculture and related industries has taken place. On the other hand, pathogenic bacteria like Flavobacterium psychrophilum, Photobacterium damselae, Vibrio anguillarum, Vibrio vulnificus, Aeromonas hydrophila, and Aeromonas salmonicida have become more prominent, causing heavy loss and/or diseases in humans due to consumption of contaminated aquaculture products (Subasinghe et al. 2001; Nakai and Park 2002; Flegel 2006; Saksida et al. 2006; FAO Report 2015). Mostly in marine and estuarine fisheries and occasionally in freshwater fisheries, vibriosis is a common disease that causes significant mortality in fish (up to 100 % mortality in larvae). This is also responsible for disease outbreaks in fisheries units. Bacterial genera Vibrio (with its various species like V. vulnificus, V. anguillarum, V. parahaemolyticus, V. alginolyticus, and V. salmonicida) and Photobacterium damselae are the causative agents of vibriosis (Noorlis et al. 2011; Higuera et al. 2013; Martínez-Díaz and Hipólito-Morales 2013; Silva et al. 2014). As antibiotic treatments have exhibited reduced effectiveness against multidrug-resistant bacteria, treatments with phages have proven efficient in case of V. anguillarum infection in fish larvae (Silva et al. 2014). However, the success of aquaculture phage therapy depends mainly on two factors: number of phages produced by each host cell after host lysis and on latent period for fresh infection to a new host by new phage particles. Ideally, for phage therapy in aquatic condition, maintaining abundant phage number and short period for fresh infection is required, which is a challenge for the days to come (Abedon et al. 2001; Crothers-Stomps et al. 2010; Silva et al. 2014). Table 2 has summarized the phage applicability and trials performed in recent era of antibiotic resistance bacteria.
Wastewater phages and their application in effluent water decontamination
Common wastewater treatment involves technology based on chemical precipitation, ion exchange, sedimentation, or coagulation-flocculation (Otte and Jacob 2006). These techniques are energy and maintenance intensive and require high-infrastructure facilities. Researchers across the world are trying to harness phage-based techniques in wastewater treatments to improve quality of effluent and sludge either released into the environment or reused (Fu and Wang 2011). This treatment has the potential to manage the problems of environmental wastewater processes like: reduction in pathogenic bacteria, foaming in activated sludge plants, digestibility, and water ability of sludge, and reduction in competition between functionally important bacterial populations and nuisance bacteria (Withey et al. 2005; Mulani et al. 2015). This is a highly potential area of research and development especially in a country like India; there is potential here for application of appropriate, affordable, sustainable, and eco-friendly approaches. Phage-mediated bactericidal effects in any wastewater treatment process have many controlling factors that lead to treatment performance. Wastewater is rich in microbial diversity that contains a wide variety of microbes and bacteriophages, among which, somatic coliphages are the most abundant. These coliphages are virulent, in nature, generally following lytic cycle for their reproduction. These phages belong to the families Myoviridae, Siphoviridae, Podoviridae, and Microviridae that mainly target the member of Enterobacteriaceae family especially E. coli (Hayes 1968). Coliphages find favorable environment in wastewater. However, their abundance depends upon factors like optimal condition and density of the host bacteria, pH, temperature, cations like calcium and magnesium, organic matter concentration, etc. (Grabow 1990; Grabow et al. 1980, 1998). For instance, fecal coliform bacteria closely related to E. coli, especially Klebsiella species, are heterotrophic and favor certain water environments for its multiplication and in turn support the multiplication of somatic coliphages (Grabow et al. 2000). It has been reported that phage titer against Salmonella typhi in both laboratory media and sewage was found to be 109–1010 per milliliter (Goyal et al. 1980). Successful phage replication requires a minimum of 104 host bacteria per milliliter. According to Goyal et al. (1987), phages under nutrient-limited conditions were found concentrated on the solid surface instead of flowing water. Coliphages also inhabit inside the gut of humans and other warm blooded animals, multiply within it, and are excreted along with fecal matter. Different pieces of information are available on phages in sewage. A number of studies have reported density of somatic coliphages to be 106–108 per milliliter (Bell 1976; Ignazitto et al., 1980; Havelaar and Hogeboom 1984; Havelaar et al. 1984; Tartera et al. 1989; Grabow et al. 1993; Grabow et al., 2000).
Interestingly, some members of coliphages showed similarity with human viruses like enteroviruses such as polio viruses (Family Picornaviridae) and F-RNA coliphages (Family Leviviridae) (Kamiko and Ohgaki 1993). Both viral particles consist of an icosahedral capsids (about 25 nm diameter) and a single strand (ss)-RNA genome. Due to these similarities, coliphages are considered to be valuable models or surrogates for human enteric viruses (Grabow 2001), which are also excreted along with fecal matter (Grabow et al. 1995). Although coliphages are shed regularly along with fecal matter, human enteric viruses are excreted generally during infection. Greater number of adenoviruses than enteroviruses have been constantly present in raw sewage around the world (Irving and Smith 1981; Hurst et al. 1988; Krikelis et al. 1985a, b), and according to Hurst et al. 1988, 80 % of infectious adenoviruses shed in the feces and present in raw sewage may be enteric adenoviruses. Recently, human fecal contamination was indicated using potential source indicators such as bacteriophages against Bacteroides (GA-17, GB-124, and ARABA 84) as well as sorbitol-fermenting Bifidobacteria (Wicki et al. 2015). Furthermore, according to Haramoto et al. (2015), coliphages belonging to the genogroup, GI F-RNA, may be used as a suitable indicator of virus reduction during wastewater treatment.
Impediments of phage application: interplay between phage and bacterial resistance
There is a tug of war between phage and bacterial infection and immunity. The selective pressure among these two entities is unique. Bacteria develop resistance to phage; subsequently, phages improve their antiviral mechanism and vice versa. This interplay between phage and bacteria is an interesting phenomenon; elevated mutation rates is a normal event for bacteria like Pseudomonas fluorescens when they are subjected to coevolve in the invitro systems (Buckling and Rainey 2002). Again, Pal et al. (2007) showed that in laboratory conditions, coevolution of P. fluorescens with its bacteriophage help in faster bacterial mutation rates. They have also found that 25 % of the P. fluorescens populations (approx. 200 bacterial generations) coevolving with the lytic phages had evolved 10 to 100-fold increase in mutation rates (Pal et al. 2007). For lytic bacteriophages, reciprocal selection for bacterial resistance and phage infectivity is essential. And this bacteria-phage interface generates more beneficial resistance mutations to the bacteria (Gomez and Buckling 2013). However, natural populations of the bacteria-phage coevolving in their natural environment (as for example, soil) may not follow the same mutation pattern as, in nature, several other factors may influence the selection rates for mutation. Innumerable microbial species including natural microbial and viral community (NMC) as a whole and other physico-chemical factors possibly will influence the rate of mutation (Gomez and Buckling 2013).
Bacterial resistance towards phage, therefore, may be due to several mechanisms. These include modification of phage attachment or adsorption sites (receptors) and CRISPR sequence-based adaptive immunity (Hyman and Abedon 2010). Alteration of phage adsorption sites/receptors is the most common phenomenon by which bacteria evade phage infection and become resistant to phage. However, phages can also adapt themselves to recognize these new modified receptors. Synthesis of exopolysaccharide (EPS) or masking proteins (like protein A of S. aureus) to mask the phage receptor is another strategy of bacteria. Again, phages conquer the barrier by cleaving the EPS layer using polysaccharide lyase or a polysaccharide hydrolase (Labrie et al. 2010, Örmälä and Jalasvuori 2013).
Further, self/non-self discrimination in prokaryotes is ubiquitous and provided by restriction-modification (RM) systems that defend hosts from exogenous DNA (Pleška et al. 2016). Bacterial system has the ability to recognize and modify the phage DNA. In many instances, the phage DNA is cleaved by the restriction endonucleases on entering into the bacterial cell through RM system and protects bacterial cell from foreign phage DNA attack (Pleška et al. 2016).
It has also been reported that phages adopt anti-restriction strategy to avoid recognition by endonuclease enzyme. For example, T4 phage escapes of restriction endonuclease attack as it contains hydroxymethylcytosine (HMC) instead of cytosine. However, some bacteria, again, modified their system to specially recognize hydroxymethylcytosine (HMC) and shatter phage DNA (Bickle and Kruger 1993; Borgaro and Zhu 2013). Yet again, anti-restriction strategy in Staphylococcus phage K is interesting where no 5′ GATC-3′ cleavage site is present; hence, DNA is protected from restriction (Kruger et al. 1988; Tock and Dryden 2005, Bryson et al. 2015).
CRISPER-Cas mechanism or Clustered regularly interspaced short palindromic repeats (CRISPRs) and the CRISPR-associated (cas) genes present a novel example of acquired resistance against viruses in prokaryotes. This mechanism is widely present in genome of bacteria and archaeal population and CRISPER-Cas loci were first described in 1987 in E. coli (Ishino et al. 1987). CRISPER is composed of 21–48 bp direct repeats separated by (26–72 bp) non-repetitive spacers, and at 5′ edge, this loci is flanked by a number of cas genes (4–20 in number) in bacterial strains. The variation in cas gene and spacer make this system more unique and robust (Marraffini 2015). Main function of CRISPER-Cas is to provide immunity against foreign DNA including phage genomic DNA or plasmid DNA (Makarova et al. 2006; Bolotin et al. 2005; Haft et al. 2005; Mojica et al. 2005; Pourcel et al. 2005; Marraffini and Sontheimer 2008). Further, as for example of antiphage activity of CRISPER-Cas mechanism in Streptococcus thermophilus, exposure to virulent phage gave rise to the phage-resistant mutants due to insertion of additional 30 bp spacer similar to protospacer of infecting phage (Barrangou et al. 2007). The event of acquiring immunity against phage can be explained in following steps like adaptation or spacer attainment, transcription of acquired spacer (small CRISPER RNAs (crRNAs), on recurrent phage attack this crRNAs form a complex with Cas protein), and immunity against phage (crRNAs-Cas complex direct nuclease to trace and chop the invading phage DNA; Marraffini 2015).
Conclusive remarks
Phages are omni-present, ubiquitous, and important part of nature. Being antibacterial and natural predators of bacteria, the role of phage can be extensive as nano-cleaner of ecosystem. Detailed classification of bacteriophages may be beneficial to mankind in the screening of therapeutic phages against a number of diseases and economically important phages help in industries and its preservation. High-throughput sequencing like metagenomics study of environmental samples can provide information of novel bacteriophages or new bacteriophage species. Various biological, environmental, medical, and pharmaceutical applications related to the use of phage are becoming more and more attractive. Further, employability of phage for sustainable wastewater treatment is a challenging approach. Labeled phages are used in situ bacterial detection using phage typing of clinical bacterial strains. Although discovered a century ago, their use in various fields are still restrictive. Comprehensive studies on basic molecular biology, host range increment, and genetics processes of these tiny antibacterial agents are required which will help them consider novel beneficial agent for bactericidal activities, even against multidrug-resistant bacteria.
References
Abedon ST (2016) Bacteriophage exploitation of bacterial biofilms: phage preference for less mature targets? FEMS Microbiol Lett. doi:10.1093/femsle/fnv246
Abedon ST, Herschler TD, Stopar D (2001) Bacteriophage latent-period evolution as a response to resource availability. Appl Environ Microbiol 13:4233–4241
Abedon ST, Sarah JK, Bob GB, Kutter EM (2011) Phage treatment of human infections. Bacteriophage. 1(2):66–85
Abeles AL, Snyder KM, Chattoraj DK (1984) P1 plasmid replication: replicon structure. J Mol Biol 17:307–324
Ackermann HW (2011a) Bacteriophage taxonomy Microbiology Australia, 90–94. Downloaded from http://journals.cambridgemedia.com.au/UserDir/CambridgeJournal/Articles/11%20ackermann244.pdf
Adriaenssens EM, Vaerenbergh JV, Vandenheuvel D (2012) T4-related bacteriophage LIMEstone isolates for the control of soft rot on potato caused by Dickeya solani. PLoS One 7:e33227
Alisky J, Iczkowski K, Rapoport A, Troitsky N (1998) Bacteriophages show promise as antimicrobial agents. J Infect 36:5–15
Allen HK, Looft T, Bayles DO, Humphrey S, Levine UY, Alt D, Stanton TB (2011) Antibiotics in feed induce prophages in swine fecal microbiomes. MBio 2:00260–00211
Allwood PB, Malik YS, Maherchandani S, Vought K, Johnson LA, Braymen C, Hedberg CW, Goyal SM (2004) Occurrence of Escherichia coli, noroviruses, and f-specific coliphages in fresh market-ready produce. J Food Protect 67:2387–2390
Almeida A, Cunha A, Gomes NCM, Alves E, Costa L et al (2009) Phage therapy and photodynamic therapy: low environmental impact approaches to inactivate microorganisms in fish farming plants. Mar Drugs 7:268–313
Atias D, Lobel L, Virta M, Marks RS (2008) Phage-displayed epitopes as bioreceptors for biosensors part two. Biological and molecular recognition systems. Wiley, Handbook of Biosensors and Biochips. doi:10.1002/9780470061565.hbb012
Babalova EG, Katsitadze KT, Sakvarelidze LA, Imnaishvili NS, Sharashidze TG, Badashvili VA, Kiknadze GP, Meĭpariani AN, Gendzekhadze ND, Machavariani EV, Gogoberidze KL, Gozalov EI, Dekanosidze NG (1968) Preventive value of dried dysentery bacteriophages. Zh Mikrobiol Epidemiol Immunobiol 45:143–145
Balasubramanian S, Sorokulova IB, Vodyanoy VJ, Simonian AL (2007) Lytic phage as a specific and selective probe for detection of Staphylococcus aureus—a surface plasmon resonance spectroscopic study. Biosens. Bioelectron. 22:948–955
Balogh B, Canteros BI, Stall RE, Jones JB (2008) Control of citrus canker and citrus bacterial spot with bacteriophages. Plant Dise 92:1048–1052
Balogh B, Jones JB, Momolet MT (2003) Improved efficacy of newly formulated bacteriophages for management of bacterial spot on tomato. Plant Dis 87:949–954
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P (2007) CRISPR provides acquired resistance against viruses in prokaryotes. Science 315:1709–1712
Barry MA, Dower WJ, Johnston SA (1996) Toward cell-targeting gene therapy vectors: selection of cell-binding peptides from random peptide presenting phage libraries. Nat Med 2:299–305
Bell RG (1976) The limitation of the ratio of fecal coliform to total coliphage as a water pollution index. Water Res 10:745–748
Benhar I (2001) Biotechnological applications of phage and cell display. Biotechnol Adv 19:1–33
Bickle TA, Kruger DH (1993) Biology of DNA restriction. Microbiol Rev 57:434–450
Biswas B, Adhya S, Washart P, Paul B, Trostel AN, Powell B, Carlton R, Merril CR (2002) Bacteriophage therapy rescues mice bacteremic from a clinical isolate of vancomycin-resistant Enterococcus faecium. Infect Immun 70:204–210
Bolotin A, Ouinquis B, Sorokin A, Ehrlich SD (2005) Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 151:2551–2561
Borgaro JG, Zhu Z (2013) Characterization of the 5-hydroxymethylcytosine-specific DNA restriction endonucleases. Nucleic Acids Res 41:4198–4206
Bradley DE (1967) Ultrastructure of bacteriophages and bacteriocins. Bacteriol Rev 31:230–314
Bradley DE, Kay D (1960) The fine structure of Bacteriophages. J Gen Microbiol 23:553–563
Breitbart M (2012) Marine viruses: truth or dare. Annu Rev Mar Sci 4:425–448
Breitbart M, Wegley L, Leeds S, Schoenfeld T, Rohwer F (2004) Phage community dynamics in hot springs. Appl Environ Microbiol 70:1633–1640
Brenner S, Horne RW (1959) A negative staining method for high-resolution electron microscopy of viruses. Biochim Biophys Acta 34:103–110. doi:10.1016/0006-3002(59)90237-9
Brenner S, Streisinger G, Horne RW, Champe SP, Barnett L, Benzer S, Rees MW (1959) Structural components of bacteriophage. J Mol Biol 1:281
Brock TD, Madigan, Michael T (1988) Biology of microorganisms, 5th edn. Prentice-Hall, USA, pp. 1988–1835
Brown S, Maria JPS, Walker S (2013) Wall teichoic acids of grampositive bacteria. In: S. Gottesman (ed) Annual Review of Microbiology, 67 (Palo Alto: Annual Reviews), pp 313–336
Brussow H, Canchaya C, Hardt W (2004) Phages and the evolution of bacterial pathogens: from genomic rearrangements to lysogenic conversion. Microbiol Mol Biol Rev 68:560–602
Bruttin A, Brüssow H (2005) Human volunteers receiving Escherichia coli phage T4 orally: a safety test of phage therapy. Antimicrob Agents Chemother 49:2874–2878
Bruynoghe R, Maisin J (1921) Essais de theârapeutique au moyen du bacteriophage. J Soc Biol 85:1120–1121
Bryson AL, Hwang Y, Sherrill-Mix S, Wu GD, Lewis JD, Black L, Clark TA, Bushman FD (2015) Covalent modification of bacteriophage T4 DNA inhibits CRISPR-Cas9. MBio 6:e00648. doi:10.1128/mBio.00648-15
Buckling A, Rainey PB (2002) Antagonistic coevolution between a bacterium and a bacteriophage. P Roy Soc B Biol Sci 269:931–936
Bugla-Ploskonska G, Futoma-Koloch B, Doroszkiewicz W (2007) Role of outer membrane proteins of gram-negative bacteria in interaction with human organism. Postepy Mikrobiol 46:139–152
Cademartiri R, Anany H, Gross I, Bhayani R, Griffiths M, Brook MA (2010) Immobilization of bacteriophages on modified silica particles. Biomaterials 31:1904–1910
Campbell NA, Reece JB (2005) Biology. Pearson, Benjamin Cummings, San Francisco, pp. 338–339
Chatterjee M, Anju CP, Biswas L, Anil Kumar V, Gopi Mohan C, Biswas R (2015) Antibiotic resistance in Pseudomonas aeruginosa and alternative therapeutic options. Int J Med Microbiol doi. doi:10.1016/j.ijmm.2015.11.004
Chuang CH, Wu TF, Chen CH, Chang KC, Ju JW, Huang YW, Van Nhan V (2015) Lab on a chip for multiplexed immunoassays to detect bladder cancer using multifunctional dielectrophoretic manipulations. Lab Chip 15:3056–3064
Clark JR, March JB (2004) Bacterial viruses as human vaccines? Expert Rev Vaccines 3:463–476
Crothers-Stomps C, Høj L, Bourne DG, Hall MR, Owens L (2010) Isolation of lytic bacteriophages against Vibrio harveyi. J Appl Microbiol 108:1744–1750
Cui Z (2015) Advances in the treatment of wound bacterial infection with phage. Zhonghua Shao Shang ZaZhi 31:389–391
D’Herelle F (1917a) Sur un microbe invisible antagoniste des bacillesdysente ´riques. C R Academy of Sciences (Paris) 165:373–375
D’Herelle F (1930) The bacteriophage and its clinical applications. Charles C Thomas, Springfield, Ill
Dalmasso M, Strain R, Neve H, Franz CMAP, Cousin FJ, Ross RP (2016) Three new Escherichia coli phages from the human gut show promising potential for phage therapy. PLoS One 11(6):e0156773. doi:10.1371/journal.pone.0156773
d’Herelle F (1917b) On an invisible microbe antagonistic to dysentery bacilli. Comptes Rendus Academie des Sciences 165:373–375
d’Herelle F (1949) The bacteriophage. Science News 14:44–59
Dickerson TJ, Kaufmann GF, Janda KD (2005) Bacteriophage-mediated protein delivery into the central nervous system and its application in immune pharmacotherapy. Expert Opin Biol Ther 5:773–781
Donnelly A, Yata T, Bentayebi K, Suwan K, Hajitou A (2015) Bacteriophage mediates efficient Gene transfer in combination with conventional Transfection reagents. Viruses 7:6476–6489
Duckworth DH (1976) Who discovered Bacteriophage? Bacteriol Rev 40:793–802
Duckworth DH, Gulig PA (2002) Bacteriophages: potential treatment for bacterial infections. BioDrugs 16:57–62
DuPont HL (2007) The growing threat of foodborne bacterial enteropathogens of animal origin. Clin Infect Dis 45:1353–1361
FAO Report (2015) Downloaded from: http://www.fao.org/3/a-i4910e.pdf. Accessed 09 Jan 2016
Fiorentin L, Vieira ND, Bavioni JW, Aves ES (2005) Use of lytic bacteriophages to reduce salmonella enteritidis in experimentally contaminated chicken cuts. Braz J Poult Sci 7:255–260
Flaherty JE, Jones JB, Harbaugh BK, Somodi GC, Jackson LE (2000) Control of bacterial spot on tomato in the greenhouse and field with h-mutant bacteriophages. Hort Sci 35:882–884
Flegel TW (2006) Detection of major penaeid shrimp viruses in Asia, a historical perspective with emphasis on Thailand. Aquaculture 258:1–33
Frampton RA, Pitman AR, Fineran PC (2012) Advances in Bacteriophage-mediated control of plant pathogens. Int J Microbiol Res . doi:10.1155/2012/326452Article ID 326452, 11 pages
Fu F, Wang Q (2011) Removal of heavy metal ions from wastewaters: a review. J. Environ Manag 92:407–418. doi:10.1016/j.jenvman.2010.11.011
Fujiwara A, Fujisawa M, Hamasaki R, Kawasaki T, Fujie M, Yamada T (2011) Biocontrol of Ralstonia solanacearum by treatment with lytic bacteriophages. Appl Env Microbiol 77:4155–4162
Funatsu T, Taniyama T, Tajima T, Tadakuma H, Namiki H (2002) Rapid and sensitive detection method of a bacterium by using a GFP reporter phage. Microbiol Immun 46:365–369
Gao C, Hong M, Geng J, Zhou H, Dong J (2015) Characterization of PI (breast cancer cell special peptide) in MDA-MB-231 breast cancer cells and its potential therapeutic applications. Int J Oncol 47:1371–1378
Glud RN, Middleboe M (2004) Virus and bacteria dynamics of coastal sediment: implication for benthic carbon cycling. Limnol Oceanogr 49:2073–2081
Golkar Z, Bagasra O, Jamil N (2013) Experimental phage therapy on multiple drug resistant Pseudomonas Aeruginosa infection in mice. J Antivir Antiretrovir:S10–005. doi:10.4172/jaa.S10-005
Gomez P, Buckling A (2013) Coevolution with phages does not influence the evolution of bacterial mutation rates in soil. ISME J 7:2242–2244
Goode D, Allen VM, Barrow PA (2003) Reduction of experimental salmonella and campylobacter contamination of chicken skin by application of lytic bacteriophages. Appl Environ Microbiol 69:5032–5036
Goyal SM, Gerba CP, Bitton G (1987) Phage ecology. Wiley, New York, p. 321
Goyal SM, Zerda KS, Gerba CP (1980) Concentration of coliphages from large volumes of water and wastewater. Appl Environ Microbiol 39:85–91
Grabow WOK (2001) Bacteriophages: update on application as models for viruses in water. Water SA 27:251–268
Grabow WOK (1990) Microbiology of drinking water treatment: reclaimed wastewater. In: McFeters GA (ed) Drinking water microbiology-progress and recent developments. Springer Verlag, New York, pp. 185–203
Grabow WOK, Burger JS, Nupen EM (1980) Evaluation of acid-fast bacteria, Candida albicans, enteric viruses and conventional indicators for monitoring wastewater reclamation systems. Prog Water Technol 12:803–817
Grabow WOK, Holtzhausen CS and De Villiers JC (1993) Research on Bacteriophages as Indicators of Water Quality. WRC Report No 321/1/93 Water Res Comm Pretoria 147
Grabow WOK, Neubrech TE, Holtzhausen CS, Jofre J (1995) Bacteroides fragilis And Escherichia coli bacteriophages: excretion by humans and animals. Water Sci Technol 31:223–230
Grabow WOK, Taylor MB, Clay CG and De Villiers JC (2000) Molecular detection of viruses in drinking water: Implications for safety and disinfection. Proc. 2nd Conf. of the Int. Life Sciences Inst.: The Safety of Water Disinfection: Balancing Chem. and Microb. Risks. Radisson Deauville Resort, Miami Beach, Florida, USA, pp 15–17
Grabow WOK, Vrey A, Uys M and De Villiers JC (1998) Evaluation of the application of bacteriophages as indicators of water quality. WRC Report No 540/1/98. Water Res. Commission, Pretoria, p 55
Guenther S, Huwyler D, Richard S, Loessner MJ (2009) Virulent bacteriophage for efficient biocontrol of listeria Listeria monocytogenes in ready-to-eat foods. Appl Environ Microbiol 75:93–100
Haft DH, Selengut J, Mongodin EF, Nelson KE (2005) A guild of 45 CRISPR-associated (Cas) protein families and multiple CRISPR/Cas subtypes exist in prokaryotic genomes. PLoS Comput Biol 1:474–483
Hagens S, Habe A, Ahsen UV, Gabain A, Blasi U (2004) Therapy of experimental pseudomonas infections with a non replicating genetically modified phage Antimicrob. Agents Chemother 48:3817–3822 . doi:10.1128/AAC0066-4804/04/$08.000
Haguenau F, Hawkes PW, Hutchison JL, Satiat-Jeunemaître B, Simon GT, Williams DB (2003) Key events in the history of the electron microscope. Microsc Microanal 9:96–138. doi:10.1017/S1431927603030113
Hankin EHL (1896) Action bactericide des Eaux de la Jumna et du Gange sur le vibrion du cholera. Annales de l’Institut Pasteur 10:511
Hanlon GW (2007) Bacteriophages: an appraisal of their role in the treatment of bacterial infections. Int J Antimicrob Agents 30:118–128
Ackermann HW (2011b) The first phage electron micrographs. Bacteriophage. 1(4):225–227
Haq IU, Chaudhry WN, Akhtar MN, Andleeb S, Qadri I (2012) Bacteriophages and their implications on future biotechnology: a review. Virol J 9:1–8
Haramoto E, Fujino S, Otagiri M (2015) Distinct behaviors of infectious F-specific RNA coliphage genogroups at a wastewater treatment plant. Sci Total Environ 1:32–38
Hart SL, Knight AM, Harbottle RP, Mistry A, Hunger HD, Cutler DF, Williamson R, Coutelle C (1994) Cell binding and internalization by filamentous phage; displaying a cyclic Arg-Gly-asp-containing peptide. The J Biol Chem 269:12468–12474
Havelaar AH, Hogeboom WM (1984) A method for the enumeration of male-specific bacteriophages in sewage. J Appl Bacteriol 56:439–447
Havelaar AH, Hogeboom WM, Pot R (1984) F-specific RNA bacteriophages in sewage; methodology and occurence. Water Sci Technol 17:645–655
Hayes W (1968) The genetics of bacteria and their viruses, 2nd edn. Blackwell Scientific Publications, Oxford
Hendrix RW (2002) Bacteriophages: evolution of the majority. Theor Popul Biol 61:471–480
Higgins J, Higgins S, Guenther KL, Huff W, Donoghue MA, Donoghue DJ, Hargis BM (2005) Use of specific bacteriophage treatment to reduce salmonella in poultry products. Poult Sci 84:1141–1145
Higuera G, Bastías R, Tsertsvadze G, Romero J, Espejo RT (2013) Recently discovered Vibrio anguillarum phages can protect against experimentally induced vibriosis in Atlantic salmon, Salmo salar. Aquaculture 392–395:128–133
Huff WE, Huff GR, Rath NC, Balog JM, Donoghue AM (2005) Alternatives to antibiotics: utilization of bacteriophage to treat colibacillosis and prevent foodborne pathogens. Poult Sci 84:655–659
Hurst CJ, McClellan KA, Benton WH (1988) Comparison of cytopathogenicity, immunofluorescence and in situ DNA hybridization as methods for the detection of adenoviruses. Water Res 22:1547–1552
Hwang I (2014) Virus outbreaks in chemical and biological sensors. Sensors 14(8):13592–13612. doi:10.3390/s140813592
Hyman P, Abedon ST (2010) Bacteriophage host range and bacterial resistance. Adv Appl Microbiol 70:217–248
Ignazitto G, Volterra L, Aulicino FA, D’angelo AM (1980) Coliphages as indicators in a treatment plant. Water Air Soil Pollut 13:391–398
Imbeault S, Parent S, Lagacé M, Uhland CF, Blais JF (2006) Using bacteriophages to prevent furunculosis caused by Aeromonas salmonicida in farmed brook trout. J Aquat Anim Health 18:203–214
Inal JM (2003) Phage therapy: a reappraisal of Bacteriophages as antibiotics. Archivum Immunologiae et Therapie Experimentalis 53:237–244
Irving LG, Smith FA (1981) One-year survey of enteroviruses, adenoviruses, and reoviruses isolated from effluent at an activated-sludge purification plant. Appl Environ Microbiol 41:51–59
Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A (1987) Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol 169:5429–5433
Jado I, Lopez R, Garcia E, Fenoll A, Casal J, Garcia P (2003) Phage lytic enzymes as therapy for antibiotic-resistant Streptococcus pneumoniae infection in a murine sepsis model. J Antimicrob Chemother 52:967–973
Jamalludeen N, She YM, Lingohr EJ, Griffiths M (2009) Isolation and characterization of virulent bacteriophages against Escherichia coli serogroups O1, O2, and O78. Poult Sci 88:1694–1702
Jones JB, Vallad GE, Iriarte FB, Obradović A, Wernsing MH, Jackson LE, Balogh B, Hong JC, Momo MT (2014) Considerations for using bacteriophages for plant disease control. Bacteriophage 2:208–214
Kamiko N, Ohgaki S (1993) Multiplication characteristics of FRNA phage and its utility as an indicator for pathogenic viruses. Water Sci Technol 27:133–136
Karlsson F, Borrebaeck C, Nilsson N, Malmborg-Hager AC (2003) The mechanism of bacterial infection by filamentous phages involves molecular interactions between TolA and phage protein 3 domains. J Bacteriol 8:2628–2634
Karunasagar I, Shivu MM, Girisha SK, Krohne G, Karunasagar I (2007) Biocontrol of pathogens in shrimp hatcheries using bacteriophages. Aquaculture 268:288–292
Keen EC (2012) Paradigms of pathogenesis: targeting the mobile genetic elements of disease. Front Cell Infect Microbiol 2:1–3
Kelly P, Anand P, Uvaydov A, Chakravartula S, Sherpa C, Pires E, O’Neil A, Douglas T, Holford M (2015) Developing a dissociative Nanocontainer for peptide drug delivery. Int J Environ Res Public Health 12:12543–12555
Kirsch J, Siltanen C, Zhou Q, Revzin A, Simonian A (2013) Biosensor technology: recent advances in threat agent detection and medicine. Chem Soc Reviews 42:8733–8768
Kodikara CP, Crew HH, Stewart GS (1991) Near on-line detection of enteric bacteria using lux recombinant bacteriophage. FEMS Microbiol Lett 67:261–265
Krikelis V, Markoulatos P, Spyrou N, Serie C (1985a) Detection of indigenous enteric viruses in raw sewage effluents of the city of Athens, Greece, during a two-year survey. Water Sci Technol 17:159–164
Krikelis V, Spyrou N, Markoulatos P, Serie C (1985b) Seasonal distribution of enteroviruses in domestic sewage. Can J Microbiol 31:24–25
Kruger DH, Barcak GJ, Smith HO (1988) Abolition of DNA recognition site resistance to the restriction endonuclease EcoRII. Biomed Biochim Acta 47:K1–K5
Krupovic M, Dutilh BE, Adriaenssens EM, Wittmann J, Vogensen FK, Sullivan MB, Rumnieks J, Prangishvili D, Lavigne R, Kropinski AM, Klumpp J, Gillis A, Enault F, Edwards RA, Duffy S, Clokie MR, Barylski J, Ackermann HW, Kuhn JH (2016) Taxonomy of prokaryotic viruses: update from the ICTV bacterial and archaeal viruses subcommittee. Arch Virol. doi:10.1007/s00705-015-2728-0
Kumari S, Harjai K, Chhibber S (2011) Bacteriophage versus antimicrobial agents for the treatment of murine burn wound infection caused by Klebsiella pneumoniae B5055. J Med Microbiol 60:205–210
Laanto E, Sundberg LR, Bamford JKH (2011) Phage specificity of the freshwater fish pathogen Flavobacterium columnare. Appl Environ Microbiol 77:7868–7872
Labrie SJ, Samson JE, Moineau S (2010) Bacteriophage resistance mechanisms. Nat Rev Microbiol 8:317–327
Lakshmanan RS, Guntupalli R, Hu J, Kim DJ, Petrenko VA, Barbaree JM, Chin BA (2007) Phage immobilized magnetoelastic sensor for the detection of salmonella typhimurium. J Microbiol Meth 71:55–60
Lang JM, Gent DH, Schwartz HF (2007) Management of Xanthomonas leaf blight of onion with bacteriophages and a plant activator. Plant Dis 91:871–878
Le Romancer M, Gaillard M, Geslin C, Prieur D (2007) Viruses in extreme environments. Reviews in Environ Sci Biotech 6:17–31
Lee CY, Kim SJ, Park BC, Han JH (2016) Effects of dietary supplementation of bacteriophages against enterotoxigenic Escherichia coli (ETEC) K88 on clinical symptoms of post-weaning pigs challenged with the ETEC pathogen. J Anim Physiol Anim Nutr (Berl). doi:10.1111/jpn.12513
Leiman PG, Kanamaru S, Mesyanzhinov VV, Arisaka F, Rossmann MG (2003) Structure and morphogenesis of bacteriophage T4. Cell Mole Life Sci 60:2356–2370
León M, Bastías R (2015) Virulence reduction in bacteriophages resistant bacteria. Front Microbiol 6:343. doi:10.3389/fmicb.2015.00343
Leverentz B, Conway WS, Alavidze Z, Janisiewicz WJ, Fuchs Y, Camp MJ, Chighladze E, Sulakvelidze A (2001) Examination of bacteriophage as a biocontrol method for salmonella on fresh-cut fruit: a model study. J Food Protect 64:1116–1121
Li W, Caberoy NB (2010) New perspective for phage display as an efficient and versatile technology of functional proteomics. Appl Microbiol Biotechnol 85:909–919
Lin L, Honh W, Ji X, Han J, Huang L, Wei Y (2010) Isolation and characterization of an extremely long tail Thermus bacteriophage from Tegchong hot springs in China. J Basic Micro 50:452–456
Liu Y, Zhang Q, Fang C, Zhu S, Tang Y, Huang S (2005) Effect of glutathione on UV induction in prophage lambda. Arch Microbiol 183:444–449
Lone A, Anany H, Hakeem M, Aguis L, Avdjian AC, Bouget M, Atashi A, Brovko L, Rochefort D, Griffiths MW (2016) Development of prototypes of bioactive packaging materials based on immobilized bacteriophages for control of growth of bacterial pathogens in foods. Int J Food Microbiol 217:49–58
Luria SE, Anderson TF (1942) The identification and characterization of bacteriophages with the electron microscope. Proc Natl Acad Sci U S A 28:127–130. doi:10.1073/pnas.28.4.127
Madigan M, Martinko J (2006) Brock biology of microorganisms, 11th edn. Prentice Hall, USA
Mai V, Ukhanova M, Visone L, Abuladze T, Sulakvelidze A (2010) Bacteriophage Administration Reduces the Concentration of Listeria monocytogenes in the Gastrointestinal Tract and Its Translocation to Spleen and Liver in Experimentally Infected Mice. Int J Microbiol 2010:624234. doi:10.1155/2010/624234
Makarova KS, Grishin NV, Shabalina SA, Wolf YI, Koonin EV (2006) A putative RNA-interference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action. Biol Direct 1:7
Maranger R, Bird DF (1995) Viral abundance in aquatic systems: a comparison between marine and fresh waters. Mar Ecol Prog Ser 121:217–226
Marcó MB, Moineau S, Quiberoni A (2012) Bacteriophages and dairy fermentations. Bacteriophage. 2:149–158
Markoishvili K, Tsitlanadze G, Katsarava R, Morris G, Sulakvelidze A (2002) A novel sustained-release matrix based on biodegradable poly (esteramide)s and impregnated with bacteriophages and an antibiotic shows promise in management of infected venous stasis ulcers and other poorly healing wounds. Int J Dermatol 41:453–458
Marraffini LA (2015) CRISPR-cas immunity in prokaryotes. Nature 526:55–61
Marraffini LA, Sontheimer EJ (2008) CRISPR interference limits horizontal gene transfer in staphylococci by targeting DNA. Science 322:1843–1845
Martínez-Díaz SF, Hipólito-Morales A (2013) Efficacy of phage therapy to prevent mortality during the vibriosis of brine shrimp. Aquaculture 400:120–124
Marza JA, Soothill JS, Boydell P, Collyns TA (2006) Multiplication of therapeutically administered bacteriophages in pseudomonas infected patients. Burns 32:644–666
Maszewska A, Wójcik E, Ciurzyńska A, Wojtasik A, Piątkowska I, Dastych J, Różalski A (2016) Differentiation of polyvalent bacteriophages specific to uropathogenic Proteus Mirabilis strains based on the host range pattern and RFLP. Acta Biochim Pol. doi:10.18388/abp.2015_1114
Mathur MD, Vidhani S, Mehndiratta PL (2003) Bacteriophage therapy: an alternative to conventional antibiotics. J Assoc Physicians India 51:593–596
Matsuzaki S, Yasuda M, Nishikawa H, Kuroda M, Ujihara T, Shuin T, Shen Y, Jin Z, Fujimoto S, Nasimuzzaman MÂD, Wakiguchi H, Sugihara S, Sugiura T, Koda S, Muraoka A, Imai S (2003) Experimental protection of mice against lethal Staphylococcus aureus infection by novel bacteriophage phi MR11. J Infect Dis 187:613–624
Mayer G (2016) Downloaded from: http://www.microbiologybook.org/mayer/phage.htm
McKenna F, Tarabily KAE, Hardy GESTJ, Dell B (2001) Novel in vivo use of a polyvalent Streptomyces phage to disinfest Streptomyces scabies-infected seed potatoes. Plant Pathol 50:666–675
Meaden S, Koskella B (2013) Exploring the risks of phage application in the environment. Front Microbiol 4:8
Merabishvili M, Pirnay JP, Verbeken G, Chanishvili N, Tediashvili M, Lashkhi N, Glonti T, Krylov V, Mast J, Parys LV, Lavigne R, Volckaert G, Mattheus W, Verween G, Corte PD, Rose T, Jennes S, Zizi M, Vos DD, Vaneechoutte M (2009) Quality-controlled small-scale production of a well-defined Bacteriophage cocktail for use in human clinical trials. PLoS One 4:1–104 . doi:10.1371/journal.pone.0004944e4944
Micallef SA, Goldstein RE, George A, Ewing L, Tall BD, Boyer MS, Joseph SW, Sapkota AR (2013) Diversity, distribution and antibiotic resistance of Enterococcus spp. recovered from tomatoes, leaves, water and soil on U.S. mid-Atlantic farms. Food Microbiol 36:465–474
Mojica FJM, Diez-Villasenor C, Garcia-Martinez J, Soria E (2005) Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol 60:174–182
Mulani MS, Azhar S, Azharuddin S, Tambe S (2015) Harnessing the power of bacteriophage for pathogenreduction in wastewater. Int J Curr Microbiol Appl Sci 2:152–161
Murphy FA, Fauquet CM, Bishop DHL, Ghabrial SA, Jarvis AW, Martelli GP, Mayo MA, Summers MD (eds) (1995) Virus taxonomy. Sixth report of the International Committee on Taxonomy of Viruses. Springer-Verlag, New York, N.Y
Nagel B, Dellweg H, Gierasch LM (1992) Glossary for chemists of terms used in biotechnology—(IUPAC recommendations 1992. Pure Appl Chem 64:143–168
Naidoo R, Singh A, Arya SK, Beadle B, Glass N, Tanha J, Szymanski CM, Evoy S (2012) Surface-immobilization of chromatographically purified bacteriophages for the optimized capture of bacteria. Bacteriophage 2:15–24
Nakai T, Park SC (2002) Bacteriophage therapy of infectious diseases in aquaculture. Res Microbiol 153:13–18
Nilsson AS (2014) Phage therapy--constraints and possibilities. J Med Sci 119:192–198
Noorlis A, Ghazali FM, Cheah YK, Tuan Zainazor TC, Ponniah J et al (2011) Prevalence and quantification of Vibrio species and Vibrio parahaemolyticus in freshwater fish at hypermarket level. Int Food Res J 18:689–695
Örmälä AM, Jalasvuori M (2013) Phage therapy: should bacterial resistance to phages be a concern, even in the long run? Bacteriophage 3:e24219. doi:10.4161/bact.24219
Otte ML, Jacob DL (2006) Constructed wetlands for phytoremediation: rhizofiltration, phytostabilisation and phytoextraction. In: Mackova M, Dowling D, Macek T (eds) Phytoremediation Rhizoremediation. Springer, Dordrecht, pp. 57–67
Pal C, Macia MD, Oliver A, Schachar I, Buckling A (2007) Coevolution with viruses drives the evolution of bacterial mutation rates. Nature 450:1079–1081
Pande J, Szewczyk MM, Grover AK (2010) Phage display: concept, innovations, applications and future. Biotechnol Adv 28:849–858
Park S, Nakai T (2003) Bacteriophage control of pseudomonas plecogl ossicida infection in ayu Plecoglossus altivelis. Dis Aquat Org 53:33–39
Park SC, Shimamura I, Fukunaga M, Mori KI, Nakai T (2000) Isolation of bacteriophages specific to a fish pathogen pseudomonas plecoglossicida as a candidate for disease control. Appl Environ Microbiol 66:1416–1422
Parracho HMRT, Burrowes BH, Enright MC, McConville ML, Harper DR (2012) The role of regulated clinical trials in the development of bacteriophage therapeutics. Journal of Molecular and Genetic Medicine 6:279–286
Paschke M (2006) Phage display systems and their applications. Appl Microbiol Biotechnol 70:2–11
Peltomaa R, Perolio IL, Benito-Peña E, Barderas R, Moreno-Bondi MC (2015) Application of bacteriophages in sensor development. Anal Bioanal Chem 1–24
Petrenko V (2008) Evolution of phage display: from bioactive peptides to bioselective nanomaterials. Expert Opin Drug Deliv 5:825–836
Pfankuch E, Kausche GA (1940) Isolierung und übermikroskopische Abbildung eines Bakteriophagen. Naturwissenschaften 28:46. doi:10.1007/BF01486932
Pietilä MK, Roine E, Sencilo A, Bamford DH, Oksanen HM (2016) Pleolipoviridae, a newly proposed family comprising archaeal pleomorphic viruses with single-stranded or double-stranded DNA genomes. Arch Virol 161(1):249–256. doi:10.1007/s00705-015-2613-x
Pourcel C, Salvignol G, Vergnaud G (2005) CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additional tools for evolutionary studies. Microbiology 151:653–663
Borah PK, Jindal JK, Verma JP (2000) Integrated management of bacterial leaf spot of mungbean with bacteriophages of Xav and chemicals. J Mycol Plant Pathol 30:19–21
Pleška M, Qian L, Okura R, Bergmiller T, Wakamoto Y, Kussell E, Guet CC (2016) Bacterial autoimmunity due to a restriction-modification system. Curr Biol 26:404–409
Popowska M, Rzeczycka M, Miernik A, Krawczyk-Balska A, Walsh F, Duffy B (2012) Influence of soil use on prevalence of tetracycline, streptomycin, and erythromycin resistance and associated resistance genes. Antimicrob Agents Chemother 56:1434–1443
Prebus A, Hillier J (1939) The construction of a magnetic electron microscope of high resolving power. Can J Res 17:49–65. doi:10.1139/cjr39a-004
Price LB, Stegger M, Hasman H, Aziz M, Larsen J, Andersen PS et al (2012) Staphylococcus aureus CC398: host adaptation and emergence of Methicillin resistance in livestock. MBio 3:00305–00311
Prigent M, Leroy M, Confalonieri F, Dutertre M, DuBow MS (2005) A diversity of bacteriophages forms and genomes can be isolated from the surface sands of Sahara Desert. Extremophiles 9:289–296
Ptashne M (2006) Lambda’s switch: lessons from a modules wap. Curr Biol 16:459–462
Quintin F, Kerrigan WC, Soothill JS (2005) Experimental bacteriophage protection against Staphylococcus aureus abscesses in a rabbit model. Antimicrob Agents Chemother 1220–1221
Rackus DG, Shamsi MH, Wheeler AR (2015) Electrochemistry, biosensors and microfluidics: a convergence of fields. Chem Soc Rev 44:5320–5340
Rahmani R, Zarrini G, Sheikhzadeh F, Zadeh NAM (2015) Effective phages as green antimicrobial agents against antibiotic-resistant hospital Escherichia coli jundishapur. J Microbiol 8(2):e17744
Rakhuba DV, Kolomiets EI, Dey ES, Novik GI (2010) Bacteriophage receptors, mechanisms of phage adsorption and penetrationinto host cell. Pol J Microbiol 59:145–155
Ravat F, Jault P, Gabard J (2015) Bactériophages et phagothérapie: utilisation de virus Naturels pour traiter les infections bactériennes. Ann Burns Fire Disasters 28:13–20
Ravensdale M, Blom TJ, Garza JAG, Svircev AM, Smith RJ (2007) Bacteriophages and the control of Erwinia carotovora subsp. Carotovora Can J Plant Pathol 29:121–130
Reardon S (2014) Phage therapy gets revitalized. Nature 510:15–16
Rhoads DD, Wolcott RD, Kuskowski MA, Wolcott BM, Ward LS, Sulakvelidze A (2009) Bacteriophage therapy of venous leg ulcers in humans: results of a phase I safety trial. J Wound Care 18:237–243
Rivas L, Coffey B, McAuliffe O, Coffey A, Ross RP, McDonnell MJ, Duffy G, Burgess CM (2010) In vivo and ex vivo evaluations of Bacteriophages e11/2 and e4/1c for use in the control of Escherichia coli O157:H7. Appl Environ Microbiol 76:7210–7216
Ruska H (1940) Über die Sichtbarmachung der bakteriophagen Lyse im Übermikroskop. Naturwissenschaften 28:45–46. doi:10.1007/BF01486931
Saksida S, Constantine J, Karreman GA, Neville C, Sweeting R, Beamish R (2006) Evaluation of sea lice, Lepeophtheirus salmonis, abundance levels on farmed salmon in British Columbia, Canada. In The Proceedings from the International Symposium 6–11 august on Veterinary Epidemiology and Economics XI, Cairns, Australia
Sangster W, Hegarty JP, Stewart DB (2014) Phage therapy for Clostridium difficile infection: an alternative to antibiotics? Seminars in Colon and Rectal Surgery 25(3):167–170
Sangster W, Hegarty JP, Stewart DB Sr (2015) Phage tail-like particles kill Clostridium Difficile and represent an alternative to conventional antibiotics. Surgery 157:96–103
Sarker SA, McCallin S, Barretto C, Berger B, Pittet AC, Sultana S, Krause L, Huq S, Bibiloni R, Bruttin A, Reuteler G, Brussow H (2012) Oral T4-like phage cocktail application to healthy adult volunteers from Bangladesh. Virology 434:222–232
Säwström CH, Lisle J, Anesio AM, Priscu JC, Laybourn-Parry J (2008) Bacteriophage in polar inland waters. Extremophiles 12:167–175
Servick K (2016) Beleaguered phage therapy trial presses on. Science 352:1506
Sharma M, Patel JR, Conway WS, Ferquson S, Sulakvelidze A (2009) Effectiveness of bacteriophage in reducing Escherichia coli o157:H7 on fresh cut cantaloupes and lettuce. J Food Protect 72:1481–1485
Sidhu SS (2000) Phage display in pharmaceutical biotechnology. Curr Opin Chem Biol 11:610–616
Silva YJ, Costa L, Pereira C, Mateus C, Cunha Â, Calado R, Gomes NCM, Pardo MA, Hernandez I, Almeida A (2014) Phage therapy as an approach to prevent Vibrio anguillarum infections in fish larvae production. PLoS One 9:e114197. doi:10.1371/journal.pone.0114197
Singh A, Arya SK, Glass N, Hanifi-Moghaddam P, Naidoo R, Szymanski CM, Tanha J, Evoy S (2010) Bacteriophage tail spike proteins as molecular probes for sensitive and selective bacterial detection. Biosens Bioelectron 26:131–138
Singh A, Poshtiban S, Evoy S (2013) Recent advances in Bacteriophage based biosensors for food-borne pathogen detection. Sensors 13:1763–1786. doi:10.3390/s130201763
Smith GP (1985) Filamentous fusion phage: novel expression vectors that display cloned antigens on the virion surface. Science 228:1315–1317
Sperinde JJ, Choi SJ, Szoka FC Jr (2001) Phage display selection of a peptide DNaseII inhibitor that enhances gene delivery. The Journal of Gene Medicine 3:101–108
Stewart GSAB, Smith T, Denyer S (1989) Genetic engineering for bioluminescent bacteria. Food Sci and Tech Today 3:19–22
Subasinghe RP, Bondad-Reantaso MG, McGladdery SE (2001) Aquaculture development, health and wealth. In Aquaculture in the Millennium. Technica l Proceedings of the Conference on Aquaculture in the Third Millennium; Suba singhe RP, Bueno P, Phillips MJ, Hough C, McGladdery SE, Arthur JR, Eds Bangkok Thailand
Sulakvelidze A, Alavidze Z, Morris JG Jr (2001) Bacteriophage therapy. Antimicrob Agents Chemother 45:649–659
Sulakvelidze A, Kutter E (2005) Bacteriophage therapy in humans. In: Kutter E, Sulakvelidze A (eds) Bacteriophages: biology and applications. CRC Press, USA, pp. 381–436
Summers WC (1999) Felix d’Herelle and the origins of molecular biology. Yale University Press, New Haven, Conn
Suttle CA (2005) Viruses in the sea. Nature 437:356–361
Suttle CA (2007) Marine viruses - major players in the global ecosystem. Nat Rev Microbiol 5:801–812
Sybesma W, Zbinden R, Chanishvili N, Kutateladze M, Chkhotua A, Ujmajuridze A, Mehnert U, Kessler TM (2016) Bacteriophages as potential treatment for urinary tract infections. Front Microbiol 7:465. doi:10.3389/fmicb.2016.00465
Tan D, Gram L, Middelboe M (2014) Vibriophages and their interactions with the fish pathogen Vibrio anguillarum. Appl Environ Microbiol 80:3128–3140
Tartera C, Lucena F, Jofre J (1989) Human origin of Bacteroides fragilis bacteriophages present in the environment. Appl Environ Microbiol 55:2696–2701
Thompson CC, Amaral GR, Campeão M, Edwards RA, Polz MF, Dutilh BE, Ussery DW, Sawabe T, Swings J (2015) Microbial taxonomy in the post-genomic era: rebuilding from scratch? Archiv Microbiol 197:359–370
Tock MR, Dryden DT (2005) The biology of restriction and anti-restriction. Curr Opin Microbiol 8:466–472
Toro H, Price SB, McKee AS, Hoerr FJ, Krehling J, Perdue M, Bauermeister L (2005) Use of bacteriophages in combination with competitive exclusion to reduce salmonella from infected chickens. Avian Dis 9:118–124
Twort FW (1915) An investigation on the nature of ultra-microscopic viruses. Lancet 2:1241–1243
Twort FW (1922) The bacteriophage: the breaking down of bacteria by associated filter passing lysins. Br Med J 2:293–296
Twort FW (1949) The discovery of the bacteriophage. Science News 14:33–34
Van Helvoort T (1992) Bacteriological and physiological research styles in the early controversy on the nature of the bacteriophage phenomenon. Med Hist 3:243–270
Verner-Jeffreys DW, Algoet M, Pond MJ, Virdee HK, Bagwell NJ, Roberts EG (2007) Furunculosis in Atlantic salmon (Salmo salar L.) is not readily controllable by bacteriophage therapy. Aquaculture 270:475–484
Viertel TM, Ritter K, Horz HP (2014) Viruses versus bacteria-novel approaches to phage therapy as a tool against multidrug-resistant pathogens. J Antimicrob Chemother 69:2326–2336
Vinod MG, Shivu MM, Umesha KR, Rajeeva BC, Krohne G, Indrani K, Iddya K (2006) Isolation of Vibrio harveyi bacteriophage with a potential for biocontrol of luminous vibriosis in hatchery environments. Aquaculture 255:117–124
Wagner P, Acheson D, Waldor M (2001) Human neutrophils and their products induce Shiga toxin production by enterohemorrhagic Escherichiacoli. Infect Immun 69:1934–1937
Wang Z, Wang D, Chen J, Sela DA, Nugen SR (2016) Development of a novel bacteriophage based biomagnetic separation method as an aid for sensitive detection of viable Escherichia coli. Analyst 141:1009–1016
Watanabe R, Matsumoto T, Sano G, Ishii Y, Tateda K, Sumiyama Y, Uchiyama J, Sakurai S, Matsuzaki S, Imai S, Yamaguchi K (2007) Efficacy of bacteriophage therapy against gut-derived sepsis caused by Pseudomonas aeruginosa in mice. Antimicrob Agents Che mother 51:446–452
Watson BB, Eveland WC (1965) The application of the phage fluorescent antiphage staining system in the specific identification of Listeria monocytogenes. I. Species specificity and immunofluorescent sensitivity of Listeria monocytogenes phage observed in smear preparations. Journal of Infectious Disease 115:363–369
Weitz JS, Wilhelm SW (2012) Ocean viruses and their effects on microbial communities and biogeochemical cycles. F1000 Biology Reports 4:17. doi:10.3410/B4-17
Weitz JS and Wilhelm SW (2013) An ocean of viruses. The Scientist (July 1st issue). http://www.the-scientist.com/?articles.view/articleNo/36120/title/An-Ocean-of-Viruses/. Downloaded 3 Jan 2016
Wicki M, Auckenthaler A, Felleisen R, Karabulut F, Niederhauser I, Tanner M, Baumgartner A (2015) Assessment of source tracking methods for application in spring water. J Water Health 13:473–488
Wildy P (1971) Classification and nomenclature of viruses. First report of the International Committee on Nomenclature of Viruses. S. Karger, Basel
Willats WG (2002) Phage display: practicalities and prospects. Plant Mol Biol 50:837–854
Williamson KE, Radosevich M, Wommack KE (2005) Abundance and diversity of viruses in six Delaware soils. Appl Environ Microbiol 71:3119–3125
Williamson KE, Wommack KE, Radosevich M (2003) Sampling natural viral communities from soil for culture-independent analyses. Appl Environ Microbiol 69:6628–6633
Wills QF, Kerrigan C, Soothill JS (2005) Experimental bacteriophage protection against Staphylococcus aureus abscesses in a rabbit model. Antimicrob Agents Chemother 49:1220–1221
Withey S, Cartmell E, Avery LM, Stephenson T (2005) Bacteriophages—potential for application in wastewater treatment processes. Sci Total Environ 339:1–18
Wlodarczyk KD, Vandenheuve D, Jang HB, Olszak T, Arabski M, Wasik S, Drabik M, Briers Y, Higgins G, Tyrrell J, Harvey BJ, Noben JP, Lavigne R, Kawa ZD (2016) A proposed integrated approach for the preclinical evaluation of phage therapy in pseudomonas infections (2016). Scientific RepoRts 6:28115. doi:10.1038/srep28115
Wright A, Hawkins CH, Anggård EE, Harper DR (2009a) A controlled clinical trial of a therapeutic bacteriophage preparation in chronic otitis due to antibiotic-resistant Pseudomonas aeruginosa; a preliminary report of efficacy. Clin Otolaryngol 34:349–357
Young R, Wang IN, Roof WD (2000) Phages will out: strategies of host cell lysis. Trends Microbiol 8:120–128
Zhu YG, Johnson TA, Su JQ, QiaoM GGX, Stedtfeld RD et al (2013) Diverse and abundant antibiotic resistance genes in Chineses wine farms. Proceedings of the National Academy of Sciences USA 110:3435–3440
Zschach H, Joensen KG, Lindhard B, Lund O, Goderdzishvili M, Chkonia I, Jgenti G, Kvatadze N, Alavidze Z, Kutter EM, Hasman H, Larsen MV (2015) What can we learn from a metagenomic analysis of a Georgian Bacteriophage cocktail? Viruses 7:6570–6589
Acknowledgments
The authors sincerely acknowledge Professor Andrew M. Kropinski, Departments of Food Science, Molecular and Cellular Biology and Pathobiology, University of Guelph, Canada and the chair of the Bacterial and Archaeal Viruses Subcommittee, for his kind suggestion and help related to bacteriophage taxonomy. Authors also acknowledge Director, Defense Research Laboratory for his kind support. Sincere thanks are also due to Mrs. Swagata Chatterjee for her contribution in making handmade figures. Further, sincere apology is being conveyed from the authors for the many colleagues whose works could not be referred to in this manuscript due to space limitations.
Author information
Authors and Affiliations
Corresponding author
Additional information
Sonika Sharma and Soumya Chatterjee contributed equally.
Rights and permissions
About this article
Cite this article
Sharma, S., Chatterjee, S., Datta, S. et al. Bacteriophages and its applications: an overview. Folia Microbiol 62, 17–55 (2017). https://doi.org/10.1007/s12223-016-0471-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12223-016-0471-x