Abstract
Bamboo, a gramineous plant belonging to the family Poaceae, comprises of 1575 species from 116 genera across the globe. It has the ability to grow and evolve on degraded land and hence, can be utilized in the various applications as an alternative for plastic and wood. DNA barcoding, a long genomic sequence, identifies barcode region which shows species-specific nucleotide differences. This technology is considered as advanced molecular technique utilized for characterization and classification of the various species by applying distinctive molecular markers. Recent investigations revealed the potential application of various barcode regions such as matK, rbcL, rpoB, rpoC1, psbA-trnH, and ITS2, in identification of many bamboo species from different genus. In this review we comprehensively discussed the relevance of DNA barcoding as a tool in classification/identification of various bamboo species. We highlighted the methodology, how this advance technology overcomes the challenges associated with traditional methods along with prospects for future research.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
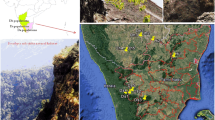
Bamboo placed to monocotyledonous plant under Poaceae family and sub family Bambusoideae [1]. There have been diversities of bamboo species found across the globe including India (Fig. 1). Bamboo has three interesting characteristics: peak development pace of 1 m per day (unique to only a few species), blossoming cycle (3 to 120 years) and widespread rhizome (sympodial and monopodial) [2,3,4,5,6]. Bamboo from base to top, as its root (rhizome), stem (culm), and leaf is used in various applications like food, fire wood, hand crafts and therapeutic [7, 8]. Bamboo is widely utilized in pulp and paper industries because of its fibre content [9, 10]. Its natural physical structure of culms (pole) is an appealing component for construction and building businesses, such as material for fortified cement beams [11, 12]. For the past two-decade bamboo has been utilized as fuel, electricity, biochar and charcoal which is an efficient way of controlling environmental contamination [13,14,15,16,17]. Previous studies on physico-chemical properties and mechanical strength of bamboo (rhizome, culm, and leaves) confirms that bamboo species are being utilized in numerous different ventures to deliver various items like furnishings, woven items, trimming/painstaking work, flooring, fibreboard, boards, mash, paper, development/building material, rustic uses, fuel, non-private/horticulture, pressing, and transport [2, 18,19,20]. Bamboo plantations and its biomass has been practiced in improvement of ecological restoration of degraded/contaminated lands and conversion of mine degraded sites into carbon sinks [11, 21]. Bamboo farming is done using rhizome, tissue culture, or small branch cutting directly without any further verification leads to mis identification and mis classification of bamboo species.
Several traditional identification and classification systems were deployed in the field of bamboo research previously. Researchers/investigators classified bamboo in to 67 genera in nine sub‐tribes, on the basis of floral characters such as flower [22], inflorescence [23], ovary and seed [24]. Bamboo species have an unusual life cycle, which diverges from one species to other; vast difference in vegetative growth phase with minimum from one year to maximum up to 120 years. Moreover, some of the species have not been reported flowering in their life cycle [25]. Further, floristic identification of these sterile bamboos is a serious problem and it does not have any solution in systematic classification, where, taxonomic studies are depended completely on floristic characters; moreover, vegetative characters could rapidly change or influenced by various environmental factors, such as soil, climatic factors etc., [26, 27]. Therefore, classification or identification of these bamboos using molecular approach is the best option to overcome problems in systematic classification [28]. Furthermore, utilizing DNA markers could provide solutions to systematic classification as an alternative approach in the studies of bamboo phylogeny and identification (Table 1). In 1997, Molecular identification studies by Clark and Kobayashi on woody bamboos have confirmed a linkage between temperate and tropical woody bamboos (New World and Old World) [29, 30].
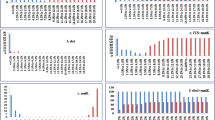
Currently, the application of advance molecular-based technology has become more prominent for characterization, taxonomic and systemic studies of organisms [46,47,48]. Also, several published researches emerge out in last decade related to application of barcode in identification of plants species (Fig. 2). These methodologies consist of RAPD (random amplified polymorphic DNA) [49], AFLP (amplified fragment length polymorphism), RFLP (restriction fragment length polymorphism), ISSR (inter simple sequence repeat), iPBS (inter primer binding site), and SCoT (start codon targeted). These molecular markers give accurate and detailed information regarding the genetic diversity [41, 50, 51], phylogenetic interactions [52, 53] and species-specific marker [54]. DNA barcoding is only one of its kind methods for characterization and classification of several organisms at species level as well genus level. The last decade witnessed the potentialities of DNA barcoding as a promising tool to supplement the species recognition in plants as well as in animals. RFLP, RADP and AFLP markers used to investigate species identification and creating phylogenetic relationships between various species of Bambusa, Dendrocalamus and Phyllostachys species [55,56,57,58]. Some other researchers used ISSR markers and expressed sequence tag (EST) based primers to find molecular relationship among the bamboo species [59, 60].
The present review is designed to consider above facts and focus to prepare a framework with respect to the application and mechanism of DNA barcoding in identification and characterization of bamboo species. A detailed discussion on, how this method overrides the traditional identification and classification methods, along with opportunities, challenges and prospects for future research.
Traditional System of Classification Versus DNA Barcoding
Traditional system of plant identification i.e., floral taxonomy has several problems that hinder it progress. Few plants species are only known from their original descriptions and several others are yet to be identified [61]. Morphological categorization of plant species is generally reliant on vegetative features that are simply governed by environmental variables [26]. Morphological systematic is based on morphological features, comparative anatomy and other vegetative characteristics [62]. Systematic identification would be beneficial in visual determination of species at various levels of classification by using their macromorphological characters, however, there are many drawbacks in the systematic [59]. Morphological characteristics, the properties of culm/culm sheaths and leaves are highly crucial to distinguish or identify bamboo species only in its matured state or in the farming field but this method is not always reliable. Table 2 describes 7 key attributes of bamboo which are used for species identification in the traditional method by most of the people who work in the bamboo sector from farm to industry. Bamboo culms height, diameter, and internode length are the countable quality yet can't be used in species identification purpose (Fig. 3). These three attributes (culms height, width, and internode length) are not valuable to distinguish among same genus species like Dendrocalamus (hamiltonii and latiflorus) because these two species have roughly same tallness (20 m), same width (16 cm), same internode length (45 cm). Likewise, Gigantochloa (apus and pseudoarundinacea). Not only that, but this attribute is not adequate to compare and distinguish between different genus as (Bambusa bambos and Gigantochloa levis), (Dendrocalamus asper and Bambusa balcooa), (Bambusa vulgaris and Cephalostachyum pergracile), (Bambusa tulda, Gigantochloa atroviolacea and Melocanna baccifera), (Thyrsostachys oliveri and Bambusa cacharensis) and (Bambusa nana, Gigantochloa rostrate and Ochlandra scriptoria) [7].
Therefore, the traditional method of bamboo species identification depends on these attributes is not reliable. In some species, the morphological characters typically used in the conventional taxonomic classification of bamboos show similar affinities. In addition, recognition and grouping of bamboo using anatomical characteristics have failed to prove effective. Various taxonomists have identified the same bamboo species in separate genera or same species which have culminated in several revisions in different genera during the last several decades. Due to the high degree of morphological resemblance, in the field recognition of certain bamboo species is exceedingly challenging even for a taxonomy specialist. Therefore, identifying bamboo species on their physical and morphological characteristics is not reliable.
DNA polymorphism is the substitute for traditional systematic to differentiate individuals or species and classify them according to molecular approach to overcome systematic classification problems. DNA barcoding is best biotechnological approach to subsidise the complications of traditional taxonomy [67]. Short sequence of genome (< 1000 bp) acting as a “DNA barcode’, which could be able to identify a species or a taxonomical group. The study “DNA barcode” expresses that the uniform genetic sequences can categorize taxon in the identical manner as the 11-digit Worldwide merchandise Code recognizes retail merchandises in market [68].
Molecular operational taxonomic unit (MOTU) is a molecular system for identification of various taxa. It is fast and perfect identification tool, and also useful in solving problems in species evolution point of view. Lambert et al., [69] studied diversity of the Earth’s biota by using molecular approach by means of DNA barcoding. Generally, to study DNA barcode includes several key attributes, which include the DNA marker used as a barcode sequence should be appropriate for wide range of taxa, it should able to differentiate among various species and should be conserved within the species, therefore the intraspecific deviation will be non-significant [70, 71]. DNA barcode correspond to the sequence in the form of nucleotides ATCG, which representing adenine, thymine, cytosine and guanine. DNA barcoding could able to serve dual purpose as an advanced device in the new era taxonomists in the enhancement of systematic knowledge as well as being an modern tool for non-taxonomists who require to create a speedy classification [72]. In recent days, there are several researchers are attempting and creating multiple genomic databases, these databases are open for all and any one can use out of it for the classifying the organism/s [73]. The purpose of creating this molecular systematic approach is to facilitate and validate the existing morphological species identifications and support the speeding up of species discoveries and solving the problems in systematic approach through creation of these barcode databases [74]. Molecular database is relatively an easy approach to compare species in a group or identify individual species and to determine distance correlation between the species [75].
Application of DNA Barcoding Technology in Bamboos
Utilization of molecular marker is one of best approaches to overcome the difficulties in identification of genotypes in bamboos [76]. Various DNA sequences were worked out to surmise phylogenetic interactions among the bamboos, however, deviation among the studied DNA sequence was very less or various DNA markers have showed similarity in the studied species [77,78,79,80]. Table 3 summarizes the molecular approaches consequently carry light to the challenging field in the taxonomical classification of several bamboos sp. at various systematic level.
Molecular markers techniques such as SSR, ISSR, RFLP, RAPD, SCoT, AFLT and SNP were in use by several researchers to distinguish the bamboos at species or generic levels. Glimpses of these techniques being used in identification, classification, characterization, cluster analysis or genetic mapping of bamboo species across the world as follows:
Simple Sequence Repeat (SSR)
Microsatellites also known as simple sequence repeats (SSR), which are comprised of short tandem repeated sequence motifs (1–6 bp in length). Genomic SSR markers have become the most common markers of choice for mapping the genes, fingerprinting of genomes and study the genetic diversity with highly reproducible data [112]. In previous studies, by Nayak and Rout, demonstrated the polymorphism of different bamboo species by Locus Ba10 amplified and characterized four bamboo species such as Bamboosa clarata, B. vulgaris, B. vulgaris var striata and B. ventricosa; further, Locus Ba20 was amplified in Dendrocalamus strictus. In recent studies, the species diversity was demonstrated among 803 moso bamboo genotypes from 34 populations with the help of 20 fluorescently labelled SSR markers [56]. Previous studies have developed 15 caespitose bamboo EST-SSR markers to identify the sequence polymorphism across the bamboo species of Bambusa edulis, Bambusa oldhamii, Phyllostachys pubescens and related species [113]. Moreover, the transferability among various bamboo species varies from 30–100%. Characteristics of 20 polymorphic microsatellite loci for Phyllostachys edulis was established with primer sequences, which will be useful in study the population genetics and conservation of the species [114]. Biogeographic history of Bambusa arnhemica from Australia was established using SSR primers [115]. SSR markers will be useful as a tool in understanding the relationship between various intra/inter bamboo species, phylogeny, ecological structure and its diversity. Characteristics of some of SSR markers being used in identification and classification of bamboo species were tabulated in Table 4.
Inter Simple Sequence Repeat (ISSR)
ISSR markers are more resourceful and reliable as it is based on longer primer sequence. ISSR markers have been widely used to identify taxonomic classification of closely resembled species, population genetics studies and phylogenetic characterization. Amom et al. [32] constructed a phylogenetic tree using ten ISSR primers and found similarity matrix between 15 North-East Indian bamboos such as Bambusa tulda, B. nutan, B. mizorameana, B. vulgaris and B. manipureana; Dendrocalamus giganteus, D. hamiltonii, D. sikkimensis, D. hookeri, D. Longispathus and D. Manipureanus; and Schizotachyum dullooa, S. pergracile, S. Munroi and S. fuchsianum. They constructed a genetic relationship between the genus and between the species and expressed disagreement with traditional system of classification. ISSR markers also found beneficial in identifying cross bred of Phyllostachys kwangsiensis and Phyllostachys bambusoides [116]. Mukherjee et al., [60] studied genetic relationships among 22 taxa of bamboo by using twelve ISSR random primers, resulting in amplification of 220 loci and classified several bamboo species (Table 5). Gami et al., [83] collected twenty bamboo accessions from different regions of India and classified bamboo species based on eight ISSR primers. Some known ISSR primers have been used in previous studies to classify bamboo species are given in Table 5.
Random Amplified Polymorphic DNA (RAPD)
RAPD is rapid and efficient technique to screen polymorphism among the species as it is directed with a single, arbitrary and short oligonucleotide primer. In this technique arbitrary primers will be used as a tool for the detection of DNA polymorphism. RAPD technique has showed positive results in identification of genetic relationships between various bamboo genera and species such as, Cephalostachyum, Dendrocalamus, Dinocloa and Sasa using 10 RAPD primers [117]. Several other researchers studied the phylogenetic relationships among the bamboo species viz., Bambusa, Denderocalamus, Melocana, Oxytenanthera, Phyllostachys, Schizostachyum, Teinostachyum, Thyrsostachysby with the help of polymorphic loci of the genomic DNA generated by RAPD [100]. Further, Annisa et al., [118] also studied the genetic relationship between 25 bamboo species of Indonesia such as Bambusa, Dendrocalamus, Dinochloa, Gigantochloa and Schizostachyum by using RAPD technique. Moreover, RAPD technique could be used to draw phylogenetic tree between closely resembled species and to eliminate distantly related species of bamboos [117]. Some characteristics of known RAPD primer sequences have been given in Table 6.
Amplified Fragment Length Polymorphism (AFLP)
AFLP method is a robust and reliable assay to detect genetic polymorphism. Moreover, this technique will be of great use in comparing a large number of genetic loci parallel being functioning under rigorous conditions of experimentation and thereby providing high reproducible results with good precision rate. In previous studies, AFLP was used to prepare cluster analysis of species among bamboo genera such as Bambusa, Dendrocalamus, Gigantochloa and Thyrsostachys [55]. Several researchers are in the opinion that AFLPs will be the best tool in identification of inter/inter specific bamboo species and useful in systematic studies and cluster analysis within the sub tribe. In China, Phyllostachys violascens cultivars were studied by using fifteen primer combinations of AFLP markers to identify genetic diversity among cultivars and found it is more efficient than other genomic techniques [119]. Some known AFLP markers have been presented in Table 7.
Restriction Fragment Length Polymorphism (RFLP)
The technique RFLP uses the combination of restriction enzymes and finds the variation in the banding pattern and reveals the genetic diversity. It is a significant molecular tool in the field of fingerprinting, mapping and paternity testing. Konzen et al., [120] studied variation among species of Bambusa, Dendrocalamus, Guadua and Phyllostachys by using RFLP techniques and constructed genomic library. Previously, Friar and Kochert distinguished 20 species of Phyllostachys by using RFLP and examined the degree of genetic variation, genetic distances and constructed dendrograms for 61 accessions [121].
Start Codon Targeted (SCoT)
SCoT technique will find the polymorphisms using reproducible markers based on the short-conserved region of the genomic sequence surrounding the ATG codon translation start or initiation codon. Previous studies showed that ScoT markers have been useful in discriminating the Bamboo species compared to other molecular techniques [31]. Recently, Amom et al., [2020], used SCoT technique to identify genetic relationship between commercial Bamboo species such as Bambusa cacharensis, B. mizorameana, Dendrocalamus manipureanus, D. hamiltonii and D. Sikkimensis from North-East India. Furthermore, dendrogram analysis based on UPGMA (Unweighted Pair Group Arithmetic Mean Method) demonstrated species-specific clustering of this five bamboo species and found that all molecular techniques used for this study were having close association except RAPD [31].
Single-Nucleotide Nolymorphism (SNP)
These are position in genome where some individuals have one nucleotide (G) and others have a different nucleotide (T) and it is referred as single-nucleotide polymorphism (SNP) or ‘Snips’ [122]. For example, two sequenced DNA fragments from different individuals—CTTAGGTTCGAA and CTGAGGTTCGAA—contain variation in a single nucleotide, that is, G or T. It is genes specific marker and its detection is more rapid because it is based on oligonucleotide hybridization analysis. Species identification was achieved by detecting allelic variations of these type of markers.
Complete phylogeny of bamboo which included bamboo tribes and its subtribes was presented in the form of molecular database by using chloroplast genome of 6.7 kb of coding nucleotide and noncoding nucleotide sequence data and characteristics of 37 microstructure [123]. Classical taxonomy contributes a major role in classifying various bamboo species. However, molecular approach i.e., DNA barcoding can certainly support the traditional system of classification and also make possible identification process more consistent and easier. In India, several researchers are working on preparation of barcodes belong to various plant and animal species (Table 8).
Recognizing bamboo species by DNA barcode is fundamental, since it can precisely distinguish bamboo species in their adolescent stage [81]. Along with these, recognizable proof of the species is useful in bamboo industries as well as in commercial bamboo farming. Surrounding atmosphere plays a critical role in the development and growth of bamboo, hence, this barcoding technology can help the farmer in choosing the species that are the best for growth and development of bamboo in specific climatic conditions [137]. DNA standardized barcode is additionally utilized for building DNA barcode library which further utilized for contrasting and recognizing DNA barcode sequences recuperated from obscure DNA samples [138]. This technology is not only used for identification and classification of species but also has to protect endangered species, preserving natural resources, pharmaceutical, wildlife forensic, authentication of natural health products, illegitimate trading and even amusing activities [74, 90, 139, 140].
Bamboo identification is regularly troublesome because of its sporadic flowering pattern and its long reproducing cycle which changes in each species [141], yet by utilizing DNA barcoding, identification can be made with much simplicity and accuracy. Bamboo has genetic material as DNA, and despite the fact that two closely species have a place with a similar family or genera have contrasted in their DNA [142]. DNA is the genomic material which differs in size from millions to billion according to species and their family, the Moso bamboo’s (Phyllostachys pubescens) whole genome size has been evaluated to be around 2034 Mb [143]. DNA barcoding isn't new, yet the first run through came well known in the year 2003, when Paul Hebert's published a research namely ‘Biological identifications through DNA barcodes’, that presented a global biological distinguishing proof framework for creatures utilizing mitochondrial gene cytochrome c oxidase I (COI) [67]. The COI has been broadly exploited in the animal kingdom and suggested to be an appropriate DNA barcode at several taxonomical levels [144, 145]. But due to its lowering rate of evolution, it is not considered as a suitable DNA marker [146]. A variety of DNA markers have been assessed for their performance as a barcode for a variety of land plants [147]. Numerous combinations of DNA markers have been recommended for the barcoding of plants [148].
The technique of DNA barcoding has been advanced for most of the taxonomic challenging lineages in the Poaceae family. DNA barcoding has been proposed a great technique for the identification of grasses species that can be challenging to recognize at the species level [67]. In 2011 Drumwright et al., [149] demonstrated the identification of grass species with 95% accuracy with the use of matK + rbcL as the main core barcode. The maximum coverage of genes in gene bank to study the species diversity of grass and barcoding included gbss1, matK, ndhF, nrITS, phB, rps16, rpoA, rbcL, rpoc2, and trnL, etc. [150,151,152].
In Bamboo, chloroplast region could be utilized as a marker region for identification. In previous study, seven up-and-comer barcoding loci from chloroplast region used to make barcoding with four coding genes such as matK, rbcL, rpoB and rpoC1, and non-coding inserts such as atpF-atpH, psbA-trnH and psbK-psbI. These 7 barcode loci show diverse oppressive force and their position as rpoC1 < rpoB < atpF-atpH < rbcL < matK < psbK-psbI < psbA-trnH [153]. Consortium for the Barcode of Life (CBOL) mentioned that loci rbcL and matK genes are well-known barcodes for the characterization of terrestrial plant species [71, 148]. Moreover in bamboo species, ITS2 and psbA-trnH barcode would be considered as effective barcodes [154]. In previous study by Cai et al., [93] it is demonstrated that some barcoding loci would be useful in species identification in bamboos viz., ITS2, matK, psbA-trnH, and rbcL. Further, the combination of these loci example rbcL + ITS2 would be better barcodes for identification of bamboo species [88]. Figure 4 illustrates the general procedures of DNA barcoding.
Methodology involved in DNA barcoding
The methodology of DNA barcoding includes the following phases starting from DNA extraction, amplification of DNA by polymerase chain reaction (PCR) with the help of universal barcode primers has been shown in Fig. 5. Combination of several primers has been reported to be useful in identification and discrimination of various taxa at species level as well at genus level in the system of molecular approach (Table 9).
The grouping of the primers sequence from coding region (rbcL, matK and rpoC1) and non-coding region (trnH2-psbA) displayed potential differential insight among the chosen species. Some molecular studies confirmed to identify 48 phylogenetically diverse plant genera by molecular approach i.e. DNA barcoding using nine 9 loci such as, accD, ITS1, matK, rpoC1 ycf5, rbcL-a, rpoB2, trnH-psbA and ndhJ [152, 155]. The DNA barcoding practices involves two fundamental steps: (1) construction of barcode library of identified species and (2) blast the barcode sequence of unidentified species compared to the barcode library intended for identification. Gene Bank has databases of some commercial bamboo species of various genera as Bambusa, Dendrocalamus, Melocanna, Oxytenanthera with their standardized identification for the species-specific region both essential and optional (rbcL, matK and psbA-trnH) (https://www.ncbi.nlm.nih.gov/genbank/). Google has an additional desktop search engine for DNA barcoding. This barcode library can be useful to various government and private agencies for providing agriculture certification to different crops [81, 156,157,158].
Concluding remarks and future prospects
Among the huge diversity of bamboo species, some are more commercial and ecologically important therefore it is necessary to identify such species accurately. Bamboo species recognition, based only on morphology, is a difficult task because it continuously changing and dependent on the environment and mainly due to the absence or long flowering cycle in their entire lifespan. Bamboo plant species characterization with recognizable proof is one of the troublesome errands in agribusiness because of its assortment and distinctive field conditions. Incorrect identification and classification of bamboo species appropriate for various agro-climatic zones can prompt a noteworthy decrease in profitability. It can likewise prompt striking misfortune to farmers if an inappropriate species personality is found a lot later particularly when the species is inadmissible for a particular end-use it was grown for.
DNA barcoding a well-known molecular technique has been observed to be an alternative tool for addressing several taxonomical complexities predominant in bamboo species. DNA barcoding is short and novel for every species of bamboo and its outcome is increasingly accurate, unlike morphological identification. It recognizes species as well as builds up connections between various bamboo species. In addition, it characterizes and recognizes bamboo species without knowing its age, physico-chemical and mechanical properties. This technique has the probability to serve as an innovative method for identifying taxonomically distinct species of bamboo. When, DNA barcode library is well-known and the classification and identification of commercial bamboo species are affordable, it can be used for standardization and certification. DNA barcode can also help to identify the genes which are responsible for quick development, high compressive strength, and high fiber content. DNA barcoding is used to handle the complexity at a taxonomic level not only in the identification of species but also for the selection of planting materials. This technology can act as an innovative molecular tool complementing modern approaches for distinguishing taxonomically difficult species of bamboo. For DNA barcoding, the entire genome sequence and expressive region must be notable ahead of time. Building a DNA barcode library for all bamboo species is expensive and challenging since it has 1575 species and tremendous genome sizes extending from 100 Mbp to 150 Gbp. However, the technology is accurate and reliable, yet expensive because of the inaccessibility of the standardized barcode library for all species. At full scale, this method can be utilized by all bamboo ventures and ranchers for distinguishing species according to their requirements. The government or private agency may give accreditation data modest rate that improves the bamboo trade.
References
Singh, L., Ruprela, N., Dafale, N., & Thul, S. T. (2020). Variation in endophytic bacterial communities associated with the rhizomes of tropical Bamboos. Journal of Sustainable Forestry. https://doi.org/10.1080/10549811.2020.1745655
Akwada, D. R., & Akinlabi, E. T. (2020). Mechanical and Physical Properties of Bamboo Species in Ghana. In M. Awang, S. S. Emamian, & F. Yusof (Eds.), (pp. 423–433). Springer.
Wang, T., Li, Q., Lou, S., Yang, Y., Peng, L., Lin, Z., Hu, Q., & Ma, L. (2019). GSK3/shaggy-like kinase 1 ubiquitously regulates cell growth from Arabidopsis to Moso bamboo (Phyllostachys edulis). Plant Science, 283, 290–300. https://doi.org/10.1016/j.plantsci.2019.03.015
Chaowana, P., & Barbu, M. C. (2017). 13—Bamboo: Potential material for biocomposites. In M. Jawaid, P. Md Tahir, & N. Saba (Eds.), Woodhead publishing series in composites science and engineering (pp. 259–289). Woodhead Publishing. https://doi.org/10.1016/B978-0-08-100959-8.00013-5
Cao, Z. H., Zhou, G. M., & Wong, M. H. (2011). Special issue on Bamboo and climate change in China. The Botanical Review, 77(3), 188. https://doi.org/10.1007/s12229-011-9064-1
Ali, N., Ting, Z., Li, H., Xue, Y., Gan, L., Liu, J., & Long, M. (2015). Heterogeneous expression and functional characterization of cellulose-degrading enzymes from aspergillus niger for enzymatic hydrolysis of alkali pretreated bamboo biomass. Molecular Biotechnology, 57(9), 859–867. https://doi.org/10.1007/s12033-015-9878-x
Sawarkar, A. D., Shrimankar, D. D., Kumar, A., Kumar, A., Singh, E., Singh, L., Kumar, S., & Kumar, R. (2020). Commercial clustering of sustainable bamboo species in India. Industrial Crops and Products, 154, 112693. https://doi.org/10.1016/j.indcrop.2020.112693
Singh, L., Jaiswal, A., Thul, S. T., & Purohit, H. J. (2017). Ecological and economic importance of bamboos. In Advances in Life Sciences (pp. 132–140). S. R. Scientific Publications. Retrieved from http://neeri.csircentral.net/id/eprint/1118
Azeez, M. A., & Orege, J. I. (2018). Bamboo, Its Chemical Modification and Products. Journal of Bamboo Current and Future Prospects. https://doi.org/10.5772/intechopen.76359
Sugesty, S., Kardiansyah, T., & Hardiani, H. (2015). Bamboo as raw materials for dissolving pulp with environmental friendly technology for rayon fiber. Procedia Chemistry, 17, 194–199. https://doi.org/10.1016/j.proche.2015.12.122
Singh, L., Sridharan, S., Thul, S. T., Kokate, P., Kumar, P., Kumar, S., & Kumar, R. (2020). Eco-rejuvenation of degraded land by microbe assisted bamboo plantation. Industrial Crops and Products. https://doi.org/10.1016/j.indcrop.2020.112795
Mali, P. R., & Datta, D. (2018). Experimental evaluation of bamboo reinforced concrete slab panels. Construction and Building Materials, 188, 1092–1100. https://doi.org/10.1016/j.conbuildmat.2018.08.162
Rudradawong, C., & Ruttanapun, C. (2019). High temperature electrical and thermal properties of activated bamboo charcoal/C12A7 mayenite composite prepared by carbon di ff usion process. Materials Chemistry and Physics, 226(January), 296–301. https://doi.org/10.1016/j.matchemphys.2019.01.028
Wang, P., Maliang, H., Wang, C., & Ma, J. (2015). Bamboo charcoal by-products as sources of new insecticide and acaricide. Industrial Crops and Products, 77, 575–581. https://doi.org/10.1016/j.indcrop.2015.09.004
Pant, D., Misra, S., Nizami, A.-S., Rehan, M., van Leeuwen, R., Tabacchioni, S., Goel, R., Sarma, P., Bakker, R., Sharma, N., Kwant, K., Diels, L., & Elst, K. (2019). Towards the development of a biobased economy in Europe and India. Critical Reviews in Biotechnology, 39(6), 779–799. https://doi.org/10.1080/07388551.2019.1618787
Schiavon, M., Ragazzi, M., Rada, E. C., & Torretta, V. (2016). Air pollution control through biotrickling filters: A review considering operational aspects and expected performance. Critical Reviews in Biotechnology, 36(6), 1143–1155. https://doi.org/10.3109/07388551.2015.1100586
Kumar, M., Dutta, S., You, S., Luo, G., Zhang, S., Show, P. L., Sawarkar, A. D., Singh, L., & Tsang, D. C. W. (2021). A critical review on biochar for enhancing biogas production from anaerobic digestion of food waste and sludge. Journal of Cleaner Production. https://doi.org/10.1016/j.jclepro.2021.127143
Hai, L., Choi, E. S., Zhai, L., Panicker, P. S., & Kim, J. (2020). Green nanocomposite made with chitin and bamboo nanofibers and its mechanical, thermal and biodegradable properties for food packaging. International Journal of Biological Macromolecules, 144, 491–499. https://doi.org/10.1016/j.ijbiomac.2019.12.124
Huang, Y., Ji, Y., & Yu, W. (2019). Development of bamboo scrimber: A literature review. Journal of Wood Science, 65(1), 25. https://doi.org/10.1186/s10086-019-1806-4
Muhammad, A., Rahman, M. R., Hamdan, S., & Sanaullah, K. (2019). Recent developments in bamboo fiber-based composites: A review. Polymer Bulletin, 76(5), 2655–2682. https://doi.org/10.1007/s00289-018-2493-9
Fuke, P., Manu, T. M., Kumar, M., Sawarkar, A. D., Pandey, A., & Singh, L. (2021). Role of microbial diversity to influence the growth and environmental remediation capacity of bamboo: A review. Industrial Crops and Products, 167, 113567. https://doi.org/10.1016/j.indcrop.2021.113567
Dransfield, S., & Widjaja, E. A. (1995). Plant resources of South-East Asia. (Vol. 7). Prosea Foundation. Prosea Foundation by Backhuys Publishers. Retrieved from https://edepot.wur.nl/411162
Li, W. (1997). Molecular evolution. Sinauer Associates Incorporated.
Soderstrom, T. R., Ellis, R. P., & Judziewicz, E. J. (1987). The Phareae and Streptogyneae (Poaceae) of Sri Lanka: A morphological-anatomical study. Smithsonian Contributions to Botany.
Janzen, D. H. (1976). Why bamboos wait so long to flower. Annual Review of Ecology and Systematics, 7(1), 347–391.
Ramanayake, S. M. S. D., Meemaduma, V. N., & Weerawardene, T. E. (2007). Genetic diversity and relationships between nine species of bamboo in Sri Lanka, using Random Amplified Polymorphic DNA. Plant Systematics and Evolution, 269(1–2), 55–61. https://doi.org/10.1007/s00606-007-0587-1
Singh, L., Thul, S. T., & Mohan Manu, T. (2021). Chapter 18—Development of bamboo biodiversity on mining degraded lands: A sustainable solution for climate change mitigation. In K. Bauddh, J. Korstad, & P. Sharma (Eds.), Phytorestoration of abandoned mining and oil drilling sites (pp. 439–451). Elsevier. https://doi.org/10.1016/B978-0-12-821200-4.00002-9
Teama, S. (2018). DNA polymorphisms: DNA-based molecular markers and their application in medicine. Genetic Diversity and Disease Susceptibility. https://doi.org/10.5772/intechopen.79517
Kobayashi, M. (1997). Phylogeny of world bamboos analysed by restriction fragment length polymorphisms of chloroplast DNA. The Bamboos, 225–234.
Clark, L. G. (1997). Bamboos: the centerpiece of the grass family. The bamboos, 237–248.
Amom, T., Tikendra, L., Apana, N., Goutam, M., Sonia, P., Koijam, A. S., Potshangbam, A. M., Rahaman, H., & Nongdam, P. (2020). Efficiency of RAPD, ISSR, iPBS, SCoT and phytochemical markers in the genetic relationship study of five native and economical important bamboos of North-East India. Phytochemistry. https://doi.org/10.1016/j.phytochem.2020.112330
Amom, T., Tikendra, L., Rahaman, H., Potshangbam, A., & Nongdam, P. (2018). Evaluation of genetic relationship between 15 bamboo species of North-East India based on ISSR marker analysis. Molecular Biology Research Communications, 7(1), 7–15. https://doi.org/10.22099/mbrc.2018.28378.1303
Maheaswari, R., Kshirsagar, J. T., & Nallasivam, L. (2016). Polymerase chain reaction: A molecular diagnostic tool in periodontology. Journal of Indian Society of Periodontology, 20(2), 128–135. https://doi.org/10.4103/0972-124X.176391
Thakur, J., Dwivedi, M. D., Sourabh, P., Uniyal, P. L., & Pandey, A. K. (2016). Genetic homogeneity revealed using SCoT, ISSR and RAPD markers in micropropagated Pittosporum eriocarpum Royle—An endemic and endangered medicinal plant. PLoS ONE. https://doi.org/10.1371/journal.pone.0159050
Xiong, F., Zhong, R., Han, Z., Jiang, J., He, L., Zhuang, W., & Tang, R. (2011). Start codon targeted polymorphism for evaluation of functional genetic variation and relationships in cultivated peanut (Arachishypogaea L.) genotypes. Molecular Biology Reports, 38(5), 3487–3494. https://doi.org/10.1007/s11033-010-0459-6
Sen, S., Skaria, R., & Muneer, P. M. A. (2010). Genetic diversity analysis in piper species (Piperaceae) using RAPD markers. Molecular Biotechnology, 46(1), 72–79. https://doi.org/10.1007/s12033-010-9281-6
Cichorz, S., Gośka, M., & Litwiniec, A. (2014). Miscanthus: Genetic diversity and genotype identification using ISSR and RAPD markers. Molecular Biotechnology, 56(10), 911–924. https://doi.org/10.1007/s12033-014-9770-0
Aydın, F., Özer, G., Alkan, M., & Çakır, İ. (2020). The utility of iPBS retrotransposons markers to analyze genetic variation in yeast. International Journal of Food Microbiology. https://doi.org/10.1016/j.ijfoodmicro.2020.108647
Cabo, S., Ferreira, L., Carvalho, A., Martins-Lopes, P., Martín, A., & Lima-Brito, J. E. (2014). Potential of Start Codon Targeted (SCoT) markers for DNA fingerprinting of newly synthesized tritordeums and their respective parents. Journal of Applied Genetics, 55(3), 307–312. https://doi.org/10.1007/s13353-014-0211-3
Costa, R., Pereira, G., Garrido, I., Tavares-de-Sousa, M. M., & Espinosa, F. (2016). Comparison of RAPD, ISSR, and AFLP Molecular markers to reveal and classify Orchardgrass (Dactylis glomerata L.) germplasm variations. PLoS ONE, 11(4), e0152972. https://doi.org/10.1371/journal.pone.0152972
Desai, P., Gajera, B., Mankad, M., Shah, S., Patel, A., Patil, G., Narayanan, S., & Kumar, N. (2015). Comparative assessment of genetic diversity among Indian bamboo genotypes using RAPD and ISSR markers. Molecular Biology Reports, 42(8), 1265–1273. https://doi.org/10.1007/s11033-015-3867-9
Etminan, A., Pour-Aboughadareh, A., Mohammadi, R., Ahmadi-Rad, A., Noori, A., Mahdavian, Z., & Moradi, Z. (2016). Applicability of start codon targeted (SCoT) and inter-simple sequence repeat (ISSR) markers for genetic diversity analysis in durum wheat genotypes. Biotechnology & Biotechnological Equipment, 30(6), 1075–1081. https://doi.org/10.1080/13102818.2016.1228478
Fernández, M., Figueiras, A., & Benito, C. (2002). The use of ISSR and RAPD markers for detecting DNA polymorphism, genotype identification and genetic diversity among barley cultivars with known origin. Theoretical and Applied Genetics, 104(5), 845–851. https://doi.org/10.1007/s00122-001-0848-2
Garcia, A. A. F., Benchimol, L. L., Barbosa, A. M. M., Geraldi, I. O., Souza Jr., C. L., & de Souza, A. P. (2004). Comparison of RAPD, RFLP, AFLP and SSR markers for diversity studies in tropical maize inbred lines. Genetics and Molecular Biology. scielo.
Kalendar, R., Antonius, K., Smýkal, P., & Schulman, A. H. (2010). iPBS: a universal method for DNA fingerprinting and retrotransposon isolation. Theoretical and Applied Genetics, 121(8), 1419–1430. https://doi.org/10.1007/s00122-010-1398-2
Kumar, M., Kumar, M., Pandey, A., & Thakur, I. S. (2019). Genomic analysis of carbon dioxide sequestering bacterium for exopolysaccharides production. Scientific Reports, 9(1), 4270. https://doi.org/10.1038/s41598-019-41052-0
Kumar, M., Morya, R., Gnansounou, E., Larroche, C., & Thakur, I. S. (2017). Characterization of carbon dioxide concentrating chemolithotrophic bacterium Serratia sp. ISTD04 for production of biodiesel. Bioresource Technology, 243, 893–897. https://doi.org/10.1016/j.biortech.2017.07.067
Thakur, I. S., Kumar, M., Varjani, S. J., Wu, Y., Gnansounou, E., & Ravindran, S. (2018). Sequestration and utilization of carbon dioxide by chemical and biological methods for biofuels and biomaterials by chemoautotrophs: Opportunities and challenges. Bioresource Technology, 256, 478–490. https://doi.org/10.1016/j.biortech.2018.02.039
Sun, Y., Xia, N., & Stapleton, C. M. A. (2006). Relationships between Bambusa species (Poaceae, Bambusoideae) revealed by random amplified polymorphic DNA. Biochemical Systematics and Ecology, 34(5), 417–423. https://doi.org/10.1016/j.bse.2005.10.015
Goyal, A. K., Pradhan, S., Basistha, B. C., & Sen, A. (2015). Micropropagation and assessment of genetic fidelity of Dendrocalamus strictus (Roxb.) nees using RAPD and ISSR markers. 3 Biotech, 5(4), 473–482. https://doi.org/10.1007/s13205-014-0244-7
Shalini, A., Meena, R. K., Tarafdar, S., & Thakur, S. (2013). Evaluation of genetic diversity in bamboo through DNA marker and study of association with morphological traits. Bulletin of Environment, Pharmacology and Life Sciences, 2(8), 78–83.
Goh, W. L., Chandran, S., Lin, R.-S., Xia, N.-H., & Wong, K. M. (2010). Phylogenetic relationships among Southeast Asian climbing bamboos (Poaceae: Bambusoideae) and the Bambusa complex. Biochemical Systematics and Ecology, 38(4), 764–773. https://doi.org/10.1016/j.bse.2010.07.006
Goh, W. L., Chandran, S., Franklin, D. C., Isagi, Y., Koshy, K. C., Sungkaew, S., Yang, H. Q., Xia, N. H., & Wong, K. M. (2013). Multi-gene region phylogenetic analyses suggest reticulate evolution and a clade of Australian origin among paleotropical woody bamboos (Poaceae: Bambusoideae: Bambuseae). Plant Systematics and Evolution, 299(1), 239–257. https://doi.org/10.1007/s00606-012-0718-1
Das, M., Bhattacharya, S., & Pal, A. (2005). Generation and characterization of SCARs by cloning and sequencing of RAPD products: A strategy for species-specific marker development in bamboo. Annals of Botany, 95(5), 835–841. https://doi.org/10.1093/aob/mci088
Loh, J. P., Kiew, R., Set, O., Gan, L. H., & Gan, Y.-Y. (2000). A Study of Genetic Variation and Relationships within the Bamboo Subtribe Bambusinae using Amplified Fragment Length Polymorphism. Annals of Botany, 85(5), 607–612. https://doi.org/10.1006/anbo.2000.1109
Nayak, S., Rout, G. R., & Das, P. (2003). Evaluation of the genetic variability in bamboo using RAPD markers. Plant Soil and Environment, 49(1), 24–28.
Bennet, S. S. R., Gaur, R. C., & Sharma, P. N. (1990). Thirty seven bamboos growing in India. Retrieved from https://agris.fao.org/agris-search/search.do?recordID=US201300307185
Friar, E., & Kochert, G. (1991). Bamboo germplasm screening with nuclear restriction fragment length polymorphisms. Theoretical and Applied Genetics, 82(6), 697–703. https://doi.org/10.1007/BF00227313
Cai, K., Zhu, L., Zhang, K., Li, L., Zhao, Z., Zeng, W., & Lin, X. (2019). Development and characterization of EST-SSR markers from RNA-Seq data in Phyllostachys violascens. Frontiers in Plant Science. https://doi.org/10.3389/fpls.2019.00050
Mukherjee, A. K., Ratha, S., Dhar, S., Debata, A. K., Acharya, P. K., Mandal, S., Panda, P. C., & Mahapatra, A. K. (2010). Genetic relationships among 22 taxa of bamboo revealed by ISSR and EST-based random primers. Biochemical Genetics, 48(11), 1015–1025. https://doi.org/10.1007/s10528-010-9390-8
Padial, J. M., & De la Riva, I. (2007). Integrative taxonomists should use and produce DNA barcodes. Retrieved from https://www.mapress.com/j/zt/article/view/zootaxa.1586.1.7
Hillis, D. M. (1987). Molecular versus morphological approaches to systematics. Annual review of Ecology and Systematics, 18(1), 23–42.
Dam, J. E. G. Van, Elbersen, H. W., & Montaño, C. M. D. (2018). 6—Bamboo Production for Industrial Utilization. Perennial Grasses for Bioenergy and Bioproducts. Elsevier Inc. https://doi.org/10.1016/B978-0-12-812900-5.00006-0
Kawasaki, M., Yordsri, V., Thanachayanont, C., Junin, C., Asahina, S., Oikawa, T., Saiki, A., & Shiojiri, M. (2017). Structures of green culms and charcoal of Bambusa multiplex. Microscopy and Microanalysis, 23(S1), 1294–1295. https://doi.org/10.1017/S1431927617007139
Banik, R. L. (2016). Silviculture of South Asian priority bamboos. Springer. https://doi.org/10.1007/978-981-10-0569-5
Liese, W., & Köhl, M. (2015). Bamboo: the plant and its uses. Springer.
Hebert, P. D. N., Cywinska, A., Ball, S. L., & DeWaard, J. R. (2003). Biological identifications through DNA barcodes. Proceedings of the Royal Society of London. Series B: Biological Sciences, 270(1512), 313–321. https://doi.org/10.1098/rspb.2002.2218
Lebonah, D. E., Dileep, A., Chandrasekhar, K., Sreevani, S., Sreedevi, B., & Pramoda Kumari, J. (2014). DNA barcoding on bacteria: A review. Advances in Biology. https://doi.org/10.1155/2014/541787
Lambert, D. M., Baker, A., Huynen, L., Haddrath, O., Hebert, P. D. N., & Millar, C. D. (2005). Is a large-scale DNA-based inventory of ancient life possible? Journal of Heredity, 96(3), 279–284. https://doi.org/10.1093/jhered/esi035
Kress, W. J. (2017). Plant DNA barcodes: Applications today and in the future. Journal of Systematics and Evolution, 55(4), 291–307. https://doi.org/10.1111/jse.12254
Vijayan, K., & Tsou, C. H. (2010). DNA barcoding in plants: taxonomy in a new perspective. Current Science, 99, 1530–1541. Retrieved from https://www.jstor.org/stable/24069450
Kaur, S. (2015). DNA barcoding and its applications. International Journal of Engineering Research and General Science, 3(2), 602–604.
Cheifet, B. (2019). Where is genomics going next? Genome Biology, 20(1), 17. https://doi.org/10.1186/s13059-019-1626-2
Hebert, P. D. N., & Gregory, T. R. (2005). The promise of DNA barcoding for taxonomy. Systematic Biology, 54(5), 852–859. https://doi.org/10.1080/10635150500354886
Nater, A., Burri, R., Kawakami, T., Smeds, L., & Ellegren, H. (2015). Resolving evolutionary relationships in closely related species with whole-genome sequencing data. Systematic Biology, 64(6), 1000–1017. https://doi.org/10.1093/sysbio/syv045
Ramakrishnan, M., Yrjälä, K., Vinod, K. K., Sharma, A., Cho, J., Satheesh, V., & Zhou, M. (2020). Genetics and genomics of moso bamboo (Phyllostachys edulis): Current status, future challenges, and biotechnological opportunities toward a sustainable bamboo industry. Food and Energy Security. https://doi.org/10.1002/fes3.229
Wang, X., Ye, X., Zhao, L., Li, D., Guo, Z., & Zhuang, H. (2017). Genome-wide RAD sequencing data provide unprecedented resolution of the phylogeny of temperate bamboos (Poaceae: Bambusoideae). Scientific Reports, 7(1), 1–11. https://doi.org/10.1038/s41598-017-11367-x
Guo, Z.-H., Chen, Y.-Y., Li, D.-Z., & Yang, J.-B. (2001). Genetic variation and evolution of the Alpine Bamboos (Poaceae: Bambusoideae) using DNA sequence data. Journal of Plant Research, 114(3), 315–322. https://doi.org/10.1007/PL00013993
Hodkinson, T. R., Renvoize, S. A., Chonghaile, G. N., Stapleton, C. M. A., & Chase, M. W. (2000). A comparison of ITS nuclear rDNA sequence data and AFLP markers for phylogenetic studies in Phyllostachys (Bambusoideae, Poaceae). Journal of Plant Research, 113(3), 259–269. https://doi.org/10.1007/PL00013936
Zhang, W.-P. (1996). Phylogeny and classification of the bamboos (Poaceae: Bambusoideae) based on molecular and morphological data.
Dev, S. A., Sijimol, K., Prathibha, P. S., Sreekumar, V. B., & Muralidharan, E. M. (2020). DNA barcoding as a valuable molecular tool for the certification of planting materials in bamboo. 3 Biotech, 10(2), 59. https://doi.org/10.1007/s13205-019-2018-8
Ma, P.-F., Zhang, Y.-X., Guo, Z.-H., & Li, D.-Z. (2015). Evidence for horizontal transfer of mitochondrial DNA to the plastid genome in a bamboo genus. Scientific Reports, 5(1), 11608. https://doi.org/10.1038/srep11608
Gami, B., Syed, B. A., & Patel, B. (2015). Assessment of genetic diversity in Bamboo accessions of india using molecular markers. International Journal of Applied Sciences and Biotechnology. https://doi.org/10.3126/ijasbt.v3i2.12587
Zhao, H., Yang, L., Peng, Z., Sun, H., Yue, X., Lou, Y., Dong, L., Wang, L., & Gao, Z. (2015). Developing genome-wide microsatellite markers of bamboo and their applications on molecular marker assisted taxonomy for accessions in the genus Phyllostachys. Scientific Reports, 5(1), 8018. https://doi.org/10.1038/srep08018
Sijimol, K., Dev, S. A., Muralidharan, E. M., & Sreekumar, V. B. (2014). DNA barcoding: An emerging tool for precise identification and certification of planting stock in taxonomically challenging bamboo species. Journal of Bamboo and Rattan, 13(1/2), 29–43.
Lin, Y., Lu, J.-J., Wu, M.-D., Zhou, M.-B., Fang, W., Ide, Y., & Tang, D.-Q. (2014). Identification, cross-taxon transferability and application of full-length cDNA SSR markers in Phyllostachys pubescens. Springerplus, 3(1), 486. https://doi.org/10.1186/2193-1801-3-486
Jethra, G., Mishra, A. K., Pandey, P. S., Choudhary, S., & Chandrasekharan, H. (2014). Phylogenetic and structural scrutiny of matK gene from wheat representing Poaceae family for DNA barcoding. International Journal of Science and Nature, 5(1), 141–146.
YuXiao, Z., YuXing, X., PengFei, M., LiNa, Z., & DeZhu, L. (2013). Selection of potential plastid DNA barcodes for Bambusoideae (Poaceae). Plant Diversity and Resources, 35(6), 743–750.
Waghmare, V. N., & Bagde, U. S. (2013). Random amplified polymorphic DNA based genetic characterization of four important species of Bamboo, found in Raigad district, Maharashtra State, India. African Journal of Biotechnology. https://doi.org/10.5897/AJB2013.12033
Sosa, V., Mejía-Saules, T., Cuéllar, M. A., & Vovides, A. P. (2013). DNA Barcoding in endangered Mesoamerican groups of plants. Botanical Review, 79(4), 469–482. https://doi.org/10.1007/s12229-013-9129-4
Das, M. M., Mahadani, P., Singh, R., Karmakar, K., & Ghosh, S. K. (2013). matK sequence based plant DNA barcoding failed to identify Bambusa (Family: Poaceae) species from Northeast India. Journal of Environment and Sociobiology, 10(1), 49–54.
Nag, A., Gupta, P., Sharma, V., Sood, A., Ahuja, P. S., & Sharma, R. K. (2013). AFLP and RAPD based genetic diversity assessment of industrially important reed bamboo (Ochlandra travancorica Benth). Journal of Plant Biochemistry and Biotechnology, 22(1), 144–149. https://doi.org/10.1007/s13562-012-0114-5
Cai, Z.-M., Zhang, Y.-X., Zhang, L.-N., Gao, L.-M., & Li, D.-Z. (2012). Testing four candidate barcoding markers in temperate woody bamboos (Poaceae: Bambusoideae). Journal of Systematics and Evolution, 50(6), 527–539. https://doi.org/10.1111/j.1759-6831.2012.00216.x
Yang, H.-Q., An, M.-Y., Gu, Z.-J., & Tian, B. (2012). Genetic diversity and differentiation of Dendrocalamus membranaceus (Poaceae: Bambusoideae), a declining Bamboo species in Yunnan, China, as based on inter-simple sequence repeat (ISSR) Analysis. International Journal of Molecular Sciences. https://doi.org/10.3390/ijms13044446
Pattanaik, S., & Hall, J. B. (2011). Molecular evidence for polyphyly in the woody bamboo genus Dendrocalamus (subtribe Bambusinae). Plant Systematics and Evolution, 291(1), 59–67. https://doi.org/10.1007/s00606-010-0380-4
Zhang, Y.-J., Ma, P.-F., & Li, D.-Z. (2011). High-throughput sequencing of six bamboo chloroplast genomes: Phylogenetic implications for temperate woody bamboos (Poaceae: Bambusoideae). PLoS ONE, 6(5), e20596–e20596. https://doi.org/10.1371/journal.pone.0020596
Kumar, P. P., Turner, I. M., Nagaraja Rao, A., & Arumuganathan, K. (2011). Estimation of nuclear DNA content of various bamboo and rattan species. Plant Biotechnology Reports, 5(4), 317–322. https://doi.org/10.1007/s11816-011-0185-0
Yasodha, R. (2011). Characterization of microsatellites in the tribe Bambuseae. Gene Conserve, 10(39).
Bhattacharya, S., Ghosh, J. S., Das, M., & Pal, A. (2009). Morphological and molecular characterization of Thamnocalamus spathiflorus subsp. spathiflorus at population level. Plant Systematics and Evolution, 282(1), 13–20. https://doi.org/10.1007/s00606-008-0092-1
Eevera, T., Rajandran, K., Saradha, S., & Lashmi, A. (2008). Analysis of genetic variation in selected bamboo species using RAPD. Tree and Forestry Science and Biotechnology, 2(1), 54–56.
Sharma, R. K., Gupta, P., Sharma, V., Sood, A., Mohapatra, T., & Ahuja, P. S. (2008). Evaluation of rice and sugarcane SSR markers for phylogenetic and genetic diversity analyses in bamboo. Genome, 51(2), 91–103. https://doi.org/10.1139/G07-101
Gui, Y., Wang, S., Quan, L., Zhou, C., Long, S., Zheng, H., Jin, L., Zhang, X. Y., Ma, N. X., & Fan, L. (2007). Genome size and sequence composition of moso bamboo: A comparative study. Science in China Series C: Life Sciences, 50(5), 700–705. https://doi.org/10.1007/s11427-007-0081-6
Das, M., Bhattacharya, S., Basak, J., & Pal, A. (2007). Phylogenetic relationships among the bamboo species as revealed by morphological characters and polymorphism analyses. Biologia Plantarum, 51(4), 667–672. https://doi.org/10.1007/s10535-007-0140-7
Marulanda, M. L., López, A. M., & Claroz, J. L. (2007). Analyzing the genetic diversity of Guadua spp. in Colombia using rice and sugarcane microsatellites. Crop Breeding and Applied Biotechnology, 7(1).
Nayak, S., & Rout, G. R. (2005). Characterization of microsatellites in Bambusa arundinacea and cross species amplification in other Bamboos. Zeitschrift für Naturforschung C, 60(7–8), 605–610. https://doi.org/10.1515/znc-2005-7-816
Guo, Z.-H., & Li, D.-Z. (2004). Phylogenetics of the Thamnocalamus group and its allies (Gramineae: Bambusoideae): Inference from the sequences of GBSSI gene and ITS spacer. Molecular Phylogenetics and Evolution, 30(1), 1–12. https://doi.org/10.1016/S1055-7903(03)00161-1
Marulanda, M. L., Márquez, P., & Londoño, X. (2002). AFLP analysis of Guadua angustifolia (Poaceae: Bambusoideae) in Columbia with emphasis on the coffee region. J Am Bamboo Soc, 16(1), 32–42.
Kelchner, S. A., & Clark, L. G. (1997). Molecular evolution and phylogenetic utility of the Chloroplastrpl 16 Intron in Chusquea and the Bambusoideae (Poaceae). Molecular Phylogenetics and Evolution, 8(3), 385–397. https://doi.org/10.1006/mpev.1997.0432
Gielis, J. (1997). Genetic variability and relationships in Phyllostachys using random amplified polymorphic DNA. In The Bamboos: Linn. Soc. Symp. Ser. (Vol. 19, pp. 107–124).
Watanabe, M., Ito, M., & Kurita, S. (1994). Chloroplast DNA phylogeny of Asian Bamboos (Bambusoideae, Poaceae) and its systematic implication. Journal of Plant Research, 107(3), 253–261. https://doi.org/10.1007/BF02344252
Hsiao, J.-Y., & Rieseberg, L. H. (1994). Population genetic structure of Yushania niitakayamensis (Bambusoideae, Poaceae) in Taiwan. Molecular Ecology, 3(3), 201–208. https://doi.org/10.1111/j.1365-294X.1994.tb00053.x
Sharma, V., Bhardwaj, P., Kumar, R., Sharma, R. K., Sood, A., & Ahuja, P. S. (2009). Identification and cross-species amplification of EST derived SSR markers in different bamboo species. Conservation Genetics, 10(3), 721–724. https://doi.org/10.1007/s10592-008-9630-1
Saha, M. C., Mian, M. A. R., Eujayl, I., Zwonitzer, J. C., Wang, L., & May, G. D. (2004). Tall fescue EST-SSR markers with transferability across several grass species. Theoretical and Applied Genetics, 109(4), 783–791. https://doi.org/10.1007/s00122-004-1681-1
Jiang, W., Bai, T., Dai, H., Wei, Q., Zhang, W., & Ding, Y. (2017). Microsatellite markers revealed moderate genetic diversity and population differentiation of moso bamboo (Phyllostachys edulis)—A primarily asexual reproduction species in China. Tree Genetics & Genomes, 13(6), 130. https://doi.org/10.1007/s11295-017-1212-2
Kaneko, S., Franklin, D. C., Yamasaki, N., & Isagi, Y. (2008). Development of microsatellite markers for Bambusa arnhemica (Poaceae: Bambuseae), a bamboo endemic to northern Australia. Conservation Genetics, 9(5), 1311–1313. https://doi.org/10.1007/s10592-007-9467-z
Lin, X. C., Lou, Y. F., Liu, J., Peng, J. S., Liao, G. L., & Fang, W. (2010). Crossbreeding of Phyllostachys species (Poaceae) and identification of their hybrids using ISSR markers. Genetics and Molecular Research, 9(3), 1398–1404. https://doi.org/10.4238/vol9-3gmr855
Jiang, W.-X., Zhang, W.-J., & Ding, Y.-L. (2013). Development of polymorphic microsatellite markers for Phyllostachys edulis (Poaceae), an important bamboo species in China. Applications in Plant Sciences, 1(7), 1200012. https://doi.org/10.3732/apps.1200012
Annisa, A., Hafzari, R., Setiawati, T., Irawan, B., & Kusmoro, J. (2019). Evaluation of RAPD markers for molecular identification of five bamboo genera from Indonesia.https://doi.org/10.2478/ffp-2019-0025
Lin, X., Lou, Y., Zhang, Y., Yuan, X., He, J., & Fang, W. (2011). Identification of genetic diversity among cultivars of Phyllostachys violascens using ISSR, SRAP and AFLP markers. The Botanical Review, 77(3), 223–232. https://doi.org/10.1007/s12229-011-9078-8
Konzen, E. R., Peron, R., Ito, M. A., Brondani, G. E., & TSA, S. M. (2017). Molecular identification of bamboo genera and species based on RAPD-RFLP markers. Embrapa Agropecuária Oeste-Artigo em periódico indexado (ALICE). https://doi.org/10.14214/sf.1691
Friar, E., & Kochert, G. (1994). A study of genetic variation and evolution of Phyllostachys (Bambusoideae: Poaceae) using nuclear restriction fragment length polymorphisms. Theoretical and Applied Genetics, 89(2), 265–270. https://doi.org/10.1007/BF00225152
Mittal, B., Chaturvedi, P., & Tulsyan, S. (2013). Restriction Fragment Length Polymorphism. In S. Maloy & K. Hughes (Eds.), Brenner's encyclopedia of genetics (pp. 190–193). Academic Press. https://doi.org/10.1016/B978-0-12-374984-0.01314-0
Kelchner, S. A. (2013). Higher level phylogenetic relationships within the bamboos (Poaceae: Bambusoideae) based on five plastid markers. Molecular Phylogenetics and Evolution, 67(2), 404–413. https://doi.org/10.1016/j.ympev.2013.02.005
Lakra, W. S., Verma, M. S., Goswami, M., Lal, K. K., Mohindra, V., Punia, P., Gopalakrishnan, A., Singh, K. V., Ward, R. D., & Hebert, P. (2011). DNA barcoding Indian marine fishes. Molecular Ecology Resources, 11(1), 60–71. https://doi.org/10.1111/j.1755-0998.2010.02894.x
CCMB. (2019). DNA Barcoding Anurans of India. Retrieved September 13, 2020, from http://tdb.ccmb.res.in/barcode/
Vinitha, M. R., Kumar, U. S., Aishwarya, K., Sabu, M., & Thomas, G. (2014). Prospects for discriminating Zingiberaceae species in India using DNA barcodes. Journal of Integrative Plant Biology, 56(8), 760–773. https://doi.org/10.1111/jipb.12189
Ragupathy, S., Faller, A. C., Shanmughanandhan, D., Kesanakurti, P., Shaanker, R. U., Ravikanth, G., Mathivanan, N., Song, J., Han, J., & Newmaster, S. (2019). Exploring DNA quantity and quality from raw materials to botanical extracts. Heliyon. https://doi.org/10.1016/j.heliyon.2019.e01935
Spots, G. H. (2011). Genetic resources of Phyllanthus in Southern India. Phyllanthus Species: Scientific Evaluation and Medicinal Applications, 97.
Srirama, R., Deepak, H. B., Senthilkumar, U., Ravikanth, G., Gurumurthy, B. R., Shivanna, M. B., Chandrasekaran, C. V., Agarwal, A., & Shaanker, R. U. (2012). Hepatoprotective activity of Indian Phyllanthus. Pharmaceutical Biology, 50(8), 948–953. https://doi.org/10.3109/13880209.2011.649858
Gaikwad, S. S., Ghate, H. V., Ghaskadbi, S. S., Patole, M. S., & Shouche, Y. S. (2012). DNA barcoding of nymphalid butterflies (Nymphalidae: Lepidoptera) from Western Ghats of India. Molecular Biology Reports, 39(3), 2375–2383. https://doi.org/10.1007/s11033-011-0988-7
Xu, S., Li, D., Li, J., Xiang, X., Jin, W., Huang, W., Jin, X., & Huang, L. (2015). Evaluation of the DNA Barcodes in Dendrobium (Orchidaceae) from Mainland Asia. PLoS ONE. https://doi.org/10.1371/journal.pone.0115168
Roy, S., Tyagi, A., Shukla, V., Kumar, A., Singh, U. M., Chaudhary, L. B., Datt, B., Bag, S. K., Singh, P. K., Nair, N. K., Husain, T., & Tuli, R. (2010). Universal plant DNA barcode loci may not work in complex groups: A case study with Indian Berberis Species. PLoS ONE. https://doi.org/10.1371/journal.pone.0013674
Biju, S., Garg, S., Gururaja, K., Shouche, Y., & Walujkar, S. A. (2014). DNA barcoding reveals unprecedented diversity in Dancing Frogs of India (Micrixalidae, Micrixalus): A taxonomic revision with description of 14 new species. Ceylon Journal of Science (Biological Sciences). https://doi.org/10.4038/cjsbs.v43i1.6850
Mathew, G. (2010). Identification of Satyrine Butterflies of Peninsular India through DNA Barcodes. Kerala Forest Research Institute.
Bhagwat, R. M., Dholakia, B. B., Kadoo, N. Y., Balasundaran, M., & Gupta, V. S. (2015). Two new potential barcodes to discriminate Dalbergia species. PLoS ONE, 10(11), e0142965. Retrieved from https://doi.org/10.1371/journal.pone.0142965
Yu, M., Jiao, L., Guo, J., Wiedenhoeft, A. C., He, T., Jiang, X., & Yin, Y. (2017). DNA barcoding of vouchered xylarium wood specimens of nine endangered Dalbergia species. Planta, 246(6), 1165–1176. https://doi.org/10.1007/s00425-017-2758-9
Lobovikov, M., Schoene, D., & Yping, L. (2012). Bamboo in climate change and rural livelihoods. Mitigation and Adaptation Strategies for Global Change, 17(3), 261–276. https://doi.org/10.1007/s11027-011-9324-8
Kress, W. J., & Erickson, D. L. (2012). DNA barcodes: Methods and Protocols. In W. J. Kress & D. L. Erickson (Eds.), (pp. 3–8). Humana Press. https://doi.org/10.1007/978-1-61779-591-6_1
Alacs, E. A., Georges, A., FitzSimmons, N. N., & Robertson, J. (2010). DNA detective: a review of molecular approaches to wildlife forensics. Forensic Science, Medicine, and Pathology, 6(3), 180–194. https://doi.org/10.1007/s12024-009-9131-7
Schindel, D. E., & Miller, S. E. (2005). DNA barcoding a useful tool for taxonomists. Nature, 435(7038), 17–18.
Das, M., Bhattacharya, S., Singh, P., Filgueiras Tarciso S., & Pal, A. B. T.-A. in B. R. (2008). Bamboo taxonomy and diversity in the era of molecular markers. In Incorporating Advances in Plant Pathology (Vol. 47, pp. 225–268). Academic Press. https://doi.org/10.1016/S0065-2296(08)00005-0
Schmidt, L., Fischer, M., & Oja, T. (2018). Two closely related species differ in their regional genetic differentiation despite admixing. AoB Plants, 10(1), ply007–ply007. https://doi.org/10.1093/aobpla/ply007
Gui, Y. J., Wang, S., Quan, L. Y., Zhou, C. P., Long, S. B., Zheng, H. J., Jin, L., Zhang, X. Y., Ma, N. X., & Fan, L. J. (2007). Genome size and sequence composition of moso bamboo: A comparative study. Science in China Series C” Life Sciences, 50(5), 700–705. https://doi.org/10.1007/s11427-007-0081-6
Janzen, D. H., Hajibabaei, M., Burns, J. M., Hallwachs, W., Remigio, E., & Hebert, P. D. N. (2005). Wedding biodiversity inventory of a large and complex Lepidoptera fauna with DNA barcoding. Philosophical Transactions of the Royal Society B: Biological Sciences, 360(1462), 1835–1845. https://doi.org/10.1098/rstb.2005.1715
Smith, M. A., Fisher, B. L., & Hebert, P. D. N. (2005). DNA barcoding for effective biodiversity assessment of a hyperdiverse arthropod group: The ants of Madagascar. Philosophical Transactions of the Royal Society B: Biological Sciences, 360(1462), 1825–1834. https://doi.org/10.1098/rstb.2005.1714
Fazekas, A. J., Burgess, K. S., Kesanakurti, P. R., Graham, S. W., Newmaster, S. G., Husband, B. C., Percy, D. M., Hajibabaei, M., & Barrett, S. C. H. (2008). Multiple multilocus DNA barcodes from the plastid genome discriminate plant species equally well. PLoS ONE. https://doi.org/10.1371/journal.pone.0002802
Liu, J. I. E., Möller, M., Gao, L.-M., Zhang, D.-Q., & Li, D.-Z. (2011). DNA barcoding for the discrimination of Eurasian yews (Taxus L., Taxaceae) and the discovery of cryptic species. Molecular Ecology Resources, 11(1), 89–100. https://doi.org/10.1111/j.1755-0998.2010.02907.x
Hollingsworth, P. M., Forrest, L. L., Spouge, J. L., Hajibabaei, M., Ratnasingham, S., van der Bank, M., D. Bogarin, K. S. Burgess, K. M. Cameron, M. Carine, J. Chacon, A. Clark, J. J. Clarkson, F. Conrad, D. S. Devey, C. S. Ford, T. A.J. Hedderson, M. L. Hollingsworth, B. C. Husband, L. J. …, Little, D. P. (2009). A DNA barcode for land plants. Proceedings of the National Academy of Sciences, 106(31), 12794–12797. https://doi.org/10.1073/pnas.0905845106
Drumwright, A. M., Allen, B. W., Huff, K. A., Ritchey, P. A., & Cahoon, A. B. (2011). Survey and DNA barcoding of Poaceae in flat rock cedar glades and Barrens State Natural Area, Murfreesboro, Tennessee. Castanea, 76(3), 300–310. https://doi.org/10.2179/11-005.1
Hodkinson, T. R., Savolainen, V., Jacobs, S. W. L., Bouchenak-Khelladi, Y., Kinney, M. S., & Salamin, N. (2007). Supersizing: progress in documenting and understanding grass species richness. Reconstructing the tree of life: taxonomy and systematics of species rich taxa, 72, 275–298.
Kikkawa, H. S., Tsuge, K., & Sugita, R. (2016). Real-time PCR quantification of chloroplast DNA supports DNA barcoding of plant species. Molecular Biotechnology, 58(3), 212–219. https://doi.org/10.1007/s12033-016-9918-1
Hajiahmadi, Z., Talebi, M., & Sayed-Tabatabaei, B. E. (2013). Studying Genetic Variability of Pomegranate (Punica granatum L.) Based on Chloroplast DNA and Barcode Genes. Molecular Biotechnology, 55(3), 249–259. https://doi.org/10.1007/s12033-013-9676-2
Hollingsworth, M. L., Andra Clark, A., Forrest, L. L., Richardson, J., Pennington, R. T., Long, D. G., Cowan, R., Chase, M. W., Gaudeu, M., & Hollingsworth, P. M. (2009). Selecting barcoding loci for plants: Evaluation of seven candidate loci with species-level sampling in three divergent groups of land plants. Molecular Ecology Resources, 9(2), 439–457. https://doi.org/10.1111/j.1755-0998.2008.02439.x
Starr, J. R., Naczi, R. F. C., & Chouinard, B. N. (2009). Plant DNA barcodes and species resolution in sedges (Carex, Cyperaceae). Molecular Ecology Resources, 9(SUPPL. 1), 151–163. https://doi.org/10.1111/j.1755-0998.2009.02640.x
Kress, W. J., & Erickson, D. L. (2007). A two-locus global DNA barcode for land plants: The Coding rbcL gene complements the non-coding trnH-psbA spacer region. PLoS ONE. https://doi.org/10.1371/journal.pone.0000508
Singh, D., Pande, A., Kulkarni, S., Kimbahune, S., Hanwate, T., & Sawarkar, A. (2015). Innovation for crop quality certification using ICT. In 2015 7th International Conference on Communication Systems and Networks (COMSNETS) (pp. 1–6). https://doi.org/10.1109/COMSNETS.2015.7098725
Hajibabaei, M., & Singer, G. A. C. (2009). Googling DNA sequences on the World Wide Web. BMC Bioinformatics, 10(14), S4. https://doi.org/10.1186/1471-2105-10-S14-S4
Hanwater, T., Vaidya, M., & Sawarkar, A. (2014). Survey on: Integrated platform for Development Rural Agriculture in India using ICT platform. International Journal of Computer Science and Information Technologies, 5(2), 2030–2032. Retrieved from http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.445.2588&rep=rep1&type=pdf
Acknowledgements
The authors are thankful to Director, CSIR-National Environmental Engineering Research Institute, Nagpur and Department of Computer Science and Engineering, Visvesvaraya National Institute of Technology (VNIT), Nagpur for providing necessary facilities for this work
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This review was approved by research committee of CSIR-NEERI, Nagpur and registered in library.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Sawarkar, A.D., Shrimankar, D.D., Kumar, M. et al. Traditional System Versus DNA Barcoding in Identification of Bamboo Species: A Systematic Review. Mol Biotechnol 63, 651–675 (2021). https://doi.org/10.1007/s12033-021-00337-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12033-021-00337-4