Abstract
Autoimmune diseases occur when the immune system loses tolerance to self-antigens, inducing inflammation and tissue damage. The pathogenesis of autoimmune diseases has not been elucidated. A growing mountain of evidence suggests the involvement of genetic and epigenetic factors in the development of these disorders. Genetic mapping has identified several candidate variants in autoimmune conditions. However, autoimmune diseases cannot be explained by genetic susceptibility alone. The fact that there is only 20 % of concordance for systemic lupus erythematosus (SLE) in homozygotic twins is an indication that epigenetics and environment may also play significant roles. Epigenetics refer to inheritable and potentially reversible changes in DNA and chromatin that regulate gene expression without altering the DNA sequence. The primary mechanisms of epigenetic regulation include DNA methylation, histone modification, and non-coding RNA-mediated regulation. The regulation on gene expression by epigenetics is similar to that by transcription factors (TFs), and the normal execution of biological event is controlled by a combination of epigenetic modifications and TFs. These two mechanisms share similar regulatory logistics and cooperate in part by influencing activity of the binding sites of target genes. In addition, the promoters of TFs have been found themselves to be modified by epigenetic regulators and TFs can also induce epigenetic changes. There is a two-way street in which interplay between epigenetic regulation and TFs plays a role in the pathogenesis of SLE, rheumatoid arthritis, type 1 diabetes, systemic sclerosis, and multiple sclerosis. Understanding of pathogenesis of these autoimmune diseases will help define potential targets for therapeutic strategies.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The immune system is a complex and sophisticated network that protects our bodies from external threats, such as bacteria, fungi, and viruses. Moreover, this network is capable of clearing apoptotic material and undesirable self-components through phagocytosis by dendritic cells and macrophages. This by itself does not induce an immune response, leading to a phenomenon of self-tolerance. However, under certain abnormal conditions, the immune system may lose tolerance to self-materials and attack self-tissues and organs, causing autoimmunity. The pathogenesis of autoimmunity has not been well elucidated [1]; however, studies of dizygotic twins and families have revealed a genetic component to autoimmunity [2]. But, this cannot explain all cases of autoimmunity. Indeed, only about 20 % concordance for systemic lupus erythematosus (SLE) has been found in homozygotic twins, suggesting a role for both environmental and epigenetic factors in the onset of autoimmune disorders [3–5]. Currently, the field of epigenetics has received intensive attention worldwide, because it may serve as a supplementary explanation for genetics in human diseases and because it can be induced by environmental exposures, which in theory, may be relatively easier to change or reverse than genetic hardwiring.
Gene transcription controls all of the actions of a cell, and it can be regulated by epigenetic modifications. Not surprisingly, epigenetic modifications may also occur on the gene loci that encode certain transcription factors (TFs) and thereby serve as an additional regulatory factor for biological processes and cellular function. The interaction between epigenetic modifications and key TFs in regulating the immune system and their roles in the pathogenesis of some autoimmune diseases, such as SLE, rheumatoid arthritis (RA), type 1 diabetes (T1D), systemic sclerosis (SSc), and multiple sclerosis (MS), are critical areas of research and may provide potential therapeutic targets for autoimmune diseases and inspire further research in this field.
Epigenetics
Epigenetic modifications are reversible and potentially heritable changes occurring in genomic DNA and chromatin that do not alter DNA sequence. The types of epigenetic modifications include DNA methylation, histone modification, and microRNA (miRNA)- and long non-coding RNA (lncRNA)-mediated regulation. Currently, epigenetics is a popular and important area of investigation in the pathogenesis of autoimmune disease such as SLE, RA, and T1D [6–9]. In addition to the discordance in the incidence of SLE between homozygotic twins, the influence of environmental triggers (e.g., infection, UV exposure, drugs) and the predominance in females emphasize the importance of aberrant epigenetic modifications in the pathogenesis of SLE [3]. In addition, certain types of drugs, including 5-azacytidine and procainamide [10], which have been reported to induce SLE, can also cause epigenetic changes. Similar observations have been made in other autoimmune diseases; e.g., abnormal gene expression has been observed in RA synovial fibroblasts (RASF) without genetic mutations, suggesting the involvement of epigenetic modifications [11], and sunlight, Epstein-Barr virus (EBV) infection [12, 13], and miRNAs [14, 15] have been implicated in the pathogenesis of MS. These observations strongly suggest that aberrant epigenetic regulation plays an important role in the pathogenesis of autoimmune disorders [16–19].
DNA Methylation and Interactions with Transcription Factors
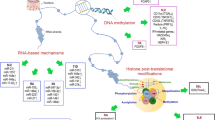
DNA methylation is a biochemical process in which a methyl group is added to a cytosine or adenine at the 5′ position of a CpG dinucleotide, converting the cytosine to a methyl-cytosine [20]. The methyl group represses gene expression when it is present in a gene and permits transcription when it is absent. DNA methylation is involved in many biological processes, including cell development, cell differentiation, and immune responses. The process of DNA methylation is regulated by methyltransferase such as DNA methyltransferase 1 (DNMT1), DNMT3a, and DNMT3b, and each performs specific functions. DNMT1 maintains the methylation status during cell replication, whereas DNMT3a and 3b usually induce de novo methylation [21]. DNA hypermethylation silences gene expression. In contrast, DNA demethylation re-activates the expression of silenced genes, which is also regulated by enzymes, such as ten-eleven translocation methylcytosine dioxygenase 1 (TET1), TET2, and TET3 [22]. In mammalian cells, DNA methylation is restricted to regions of high CpG dinucleotide content, termed CpG islands, which are typically located in promoter regions [23]. The interplay between TFs and DNA methylation consists of four distinct mechanisms (Fig. 1):
CpG Methylation
Methyl groups interfere with the binding of TFs. Many TFs are sensitive to DNA methylation due to their binding sites in genomic DNA containing CpG pairs. When these CpG pairs are methylated, the TFs fail to bind DNA and active transcription processes are blocked. Two basic models have evolved: in the first, DNA methylation can directly repress transcription by blocking transcriptional activators from binding to cognate DNA sequences [24]; in the second, the methyl-CpG-binding domain (MBD) proteins mediate transcriptional suppression by binding to methylated sequences and further altering the chromatin structure by forming a co-repressor complex [25]. This mechanism has been supported by non-systematic experimental evidence that the methylation of E-box (CACGTG) sequences inhibits the binding of N-Myc to the promoter of EGFR [26], and the methylation of PEG3 gene promoter prevents the binding of YY1 [27]. In contrast, methylated cytosine residues can attract both activating [28] and repressing [29] TFs. For example, the methylation of the CRE sequence promotes the DNA binding of C/EBPa and, in turn, activates the transcription of a set of genes involved in adipocyte differentiation [28]. However, recent advances suggest that the ability of the methylation of certain TF binding sites to prevent TF binding is restricted to special cases [30], with most CpG islands remaining nonmethylated regardless of gene expression. Genome-wide studies focusing on CpG islands have uncovered numerous instances of methylation of CpG islands in normal somatic cells. CpG islands in the germline are almost invariably nonmethylated, but a small proportion acquires methylation in somatic tissues [31, 32] (Fig. 1a).
Methylation of Transcription Factors
The promoter region of TFs is methylated, leading to transcriptional repression. For example, RORC is an essential TF gene for Th17 cell differentiation and has been found to be regulated by DNA methylation during the polarization process [33]. In addition, several of the transcription factor genes, such as SPI1, GATA3, TCF-7, Etv5, c-maf, and TBX21, have been shown to be differentially methylated in specific cell lineages and stages of the hematopoietic cascade [34] (Fig. 1b).
Recruitment of DNA Methyltransferases and Ten-Eleven Translocation Methylcytosine Dioxygenases Induced by Transcription Factors
In addition, DNA methyltransferases and demethylases are usually recruited by TFs. For example, TET3 is recruited by REST, a TF that induces gene transcription [35]. Some TFs can interact with DNMT1 to induce the recruitment of DNMT1 on DNA sites usually bound by TFs and promote the DNA methylation maintenance of CpG located on or in the vicinity of these sites. However, the findings published by Hervouet et al. demonstrate that DNMT1 interacts with TFs to promote the inheritance of site-specific DNA methylation, while the DNMT1-PCNA-UHRF1 complex promotes the inheritance of DNA methylation without site preference. Fifty-eight TFs, including NF-kappa B (NF-κB)-p65 and STAT1, have been identified that interact with recombinant DNMT1 [36] (Fig. 1c).
Transcription Factor Promoted Transcription of DNA Methyltransferases and Ten-Eleven Translocation Methylcytosine Dioxygenases
TFs promote the expression of methyltransferases and demethylases. Another group reported that STAT3 promotes DNMT1 expression by binding promoter 1 and enhancer 1 of the DNMT1 gene in malignant T lymphocytes [37]. This finding was supported by treatment of the malignant T lymphocytes with STAT3 siRNA, which abrogated expression of DNMT1, inhibited cell growth, and induced programmed cell death (Fig. 1d).
Histone Modifications and Interactions with Transcription Factors
Histone modifications are another important epigenetic mechanisms for regulating gene expression. DNA is packaged into the nucleus as chromatin, and the nucleosome is the basic subunit of the chromatin. Each nucleosome is formed by 146 base pairs (bp), or two turns, of DNA wrapped around a histone core and contains two copies each of H2A, H2B, H3, and H4. The histones present small protein tails that protrude from the nucleosome and are accessible to modifications, including methylation, acetylation, and ubiquitination [38]. Each modification has specific functions. For example, the acetylation of histone H3K9 enhances transcription, whereas the methylation of histone H3K9 suppresses transcription. Among these modifications, acetylation has been the most intensively studied. Acetylation is regulated by histone acetyltransferases (HATs) and histone deacetylases (HDAC). HAT transfer acetyl groups to lysine residues which lead to gene activation; HDAC removes acetyl groups, resulting in gene silencing [39]. Unlike acetylation, histone methylation occurs on either arginine or lysine residues and is regulated by histone methyltransferases (HMTs) and histone demethylases (HDMs). The effects of methylation are modulated by both the position of the modified residue and the number of methyl groups. It is well known that the H3K4me3 modification enhances gene expression, whereas H3K9me3 and H3K27me3 modification leads to gene repression [40, 41].
The interaction between histone modifications and TFs can also be divided into four distinct categories (Fig. 2):
Four forms of interplay between histone modification and TFs. a To form heterochromatin, the histone deacetylation of histone tails caused by HDAC enzymes in association with DNA methylation (M) confers a dense configuration of DNA that prevents its transcription. In the euchromatic state, histone tails are acetylated by HAT enzymes in association with DNA demethylation to promote gene expression. b TFs can be either modified before binding to histones. c TFs regulate the transcription of HDACs or HATs. d HDACs can bind to TFs and regulate transcription together
-
1.
Histone modifications interfere with the binding of TFs to their target DNAs. For example, histone deacetylation confers a heterochromatic configuration that prevents TFs from binding to the DNA. In contrast, histone acetylation causes a euchromatic configuration and promotes the affinity of DNA and TFs [42] (Fig. 2a).
-
2.
Histone modification enzymes regulate TFs. This mechanism is supported by the findings that HDAC3 interacts with and regulates GCMa, which is a TF that regulates development [43] (Fig. 2b).
-
3.
TFs interact with HATs and HDACs directly and recruit these enzymes to their target DNA loci to regulate gene transcription, for example, YY1, a sequence-specific DNA binding transcription factor that activates and represses many genes through acetylation by p300 and deacetylation by HDACs [44] (Fig. 2c).
-
4.
TFs regulate the expression of histone modification enzymes. For example, NF-κB was recently found to regulate the expression of SIRT1 [45] (Fig. 2d).
miRNAs and Interactions with Transcription Factors
miRNAs are small, non-coding RNAs (21–23 bp long) and function as posttranscriptional and posttranslational regulators of gene expression. miRNAs perform their functions by binding to the 3′-untranslated region (UTR) of the messenger RNA (mRNA) of a target gene, causing mRNA cleavage, translational repression, or translational arrest [46, 47]. Over 2000 miRNAs are registered in the human miRNA databases, and up to 1000 of them control one third of the transcriptome and regulate cell differentiation, cell cycle, apoptosis, and immune responses [48–50]. Based on their functions, it is not surprising that they play important roles in the innate and adaptive immune systems and thus may be involved in autoimmune disorders [51, 52].
miRNA transcripts are generated by RNA polymerase II in the nucleus to compose primary miRNAs (pri-miRNAs), which are then recognized by nuclear enzymes, such as Drosha, and form ∼70-nucleotide hairpin precursor miRNAs (pre-mRNAs). The mature miRNAs are cleaved from pre-mRNAs by the enzyme Dicer and form a duplex 18–23 nucleotides in length. One of these two strands with lower stability in the 5′ end will be associated with the RNA-induced silencing complex (RISC), the place where miRNAs bind to the mRNA targets. The predominant regulatory effect of miRNAs is to repress their target mRNAs [53, 54].
miRNAs and Transcription Factors
In complex multicellular organisms, TFs do not work alone but together in a cooperative network [55]. There is increasing evidence to support the hypothesis that miRNA is one of these cooperators, and miRNAs are the principal class of gene regulators together with TFs. Gene transcription occurs when TFs bind to cis-regulatory sites, which are usually situated upstream of protein-coding genes. Then, one or more miRNAs bind to the cis-regulatory sites as well, often in the 3′ UTR of the mRNA, and repress protein translation [56]. Thus, TFs regulate gene expression at the transcriptional level and miRNA function at the posttranscriptional level. However, recent studies observed that miRNAs provided genetic switch mechanisms to essentially repress target gene expression by altering TF function and TF-mediated actions [57]. For example, miR-493-5p, miR-124/506, and TF SP1 have been found to be involved in a TF-miR co-regulation network [22].
In addition, miRNAs and TFs have been found to form autoregulatory feedback loops, in which the expression of one affects either the presence or the absence of the other. These loops consist of unilateral or reciprocal negative feedback and double-negative feedbacks [58] (Fig. 3). In a unilateral negative feedback loop, expression of TFs is negatively controlled by the miRNAs, while the miRNAs are positively regulated by TFs. In a reciprocal negative feedback loop, the expression of the TF is repressed by the miRNAs and the miRNAs themselves are inhibited by the TFs. In double-negative feedback loops, the TF-regulated miRNAs are directly responsible for transcriptional activation and inactivation, while the miRNAs themselves are regulated by TFs [56].
Long Non-Coding RNAs and Interactions with Transcription Factors
With the recent technical advances in genome-wide studies, it is becoming increasingly obvious that the majority (over 98 %) of the human genome is transcribed into non-protein-coding RNAs (ncRNAs) [59, 60]. In addition to miRNAs, a large number of long ncRNAs, greater than 200 nt in length, have been identified, but only a minority of them have been assigned functions. Based on their genomic proximity to protein-coding genes, lncRNAs are divided into five types: sense, antisense, intronic, intergenic, and bidirectional lncRNAs [61]. Unlike miRNAs, lncRNAs can both negatively and positively regulate gene expression and may function by forming lncRNA/RNA, lncRNA/protein, or lncRNA/chromatin interactions [62, 63]. lncRNAs are currently the focus of intense research because multiple lines of evidence suggest that lncRNAs contribute to a range of human diseases, from neurodegeneration to cancer, by altering their primary structure, secondary structure, and expression levels [64, 65]. However, only a few lncRNAs have been found to regulate immune responses [65–69], including T cell differentiation, dendritic cell functions, and cytokine production, and therefore play a role in autoimmune disorders in a cell-type specific manner [70].
The mechanisms by which lncRNAs regulate gene expression are still incompletely understood. Recent advances suggest that lncRNAs regulate gene expression at the pre-transcriptional, transcriptional, and posttranscriptional level. For example, lncRNAs can mediate epigenetic changes by recruiting chromatin-remodeling complexes, including PRC1, PRC2, G9a, and MLL, to specific locations in the genome [71, 72]. lncRNAs regulate dendritic cell differentiation by binding directly to STAT3 in the cytoplasm, which promotes STAT3 phosphorylation on tyrosine-705 by preventing STAT3 from further binding to and being dephosphorylated by SHP1 [67]. At the transcriptional level, some lncRNAs, such as 7SK [73], have been revealed to directly affect the loading and activity of either RNA polymerase II or general TFs and to further influence the general output of mRNAs. In addition, lncRNAs, such as NRON [74], serve as either co-factors or inhibitors to regulate the activity of particular TFs. At the posttranscriptional level, antisense lncRNAs are capable of modulating mRNA editing, transport, translation, and degradation and can mediate alternative splicing of the mRNA by forming RNA duplexes [75, 76] (Fig. 4).
lncRNAs regulate gene expression and the interaction of lncRNAs between TFs. a lncRNAs can recruit chromatin-modifying complexes to specific genomic loci to regulate target gene expression. b lncRNAs influence the general output of mRNAs by directly affecting the loading and activity of RNAPII (right) or general TFs (left). In addition, lncRNAs can act as co-factors or inhibitors to regulate the activity of a particular TF. lncRNA Evf2 acts as a co-activator of the TF DLX2 to regulate Dlx5 and Dlx6 gene transcription
lncRNAs are involved in many biological process and various diseases. In this review, we focus on their actions in the immune system, which is an area where their function is only just beginning to be studied. A recent study revealed highly dynamic and cell-specific expression patterns for lncRNAs during T cell differentiation. Many lncRNAs are found to be bound and controlled by the key TFs T-bet, GATA-3, STAT4, and STAT6, which are the principal TFs for Th1 and Th2 differentiation [68].
The Interaction of Transcription Factors and Aberrant Epigenetic Modifications in Autoimmune Diseases
Systemic Lupus Erythematosus
SLE is a multisystem autoimmune disorder that predominately affects women (the female to male ratio is 9 to 1) during their reproductive age [77, 78]. It is characterized by diverse autoantibodies in the blood circulation [79], together with autoreactive T and B lymphocytes [80, 81]. Although the direct cause of SLE remains unidentified, many factors are believed to contribute to autoimmunity in SLE, including genetic susceptibility, epigenetics, hormones, and environmental factors [82–85]. SLE occurs when an individual with genetic susceptibility to lupus encounters environmental triggers such as sunlight, drugs, or infection. The role of DNA methylation in SLE first attracted worldwide attention in the 1960s and has since become the subject of intense research [86]. In humans, DNA demethylation has been found in SLE CD4+ T cells, but not in either CD8+T cells or peripheral blood mononuclear cells (PBMCs) [87, 88]. However, the global DNA methylation and histone modification status may not reflect real gene expression but may instead reflect the activation status of the cells.
Although epigenetic modifications such as DNA methylation, histone modifications [89], miRNAs [90, 91], and lncRNA expression [92] are well documented as critical players in the pathogenesis of SLE, only a few studies have focused on the interaction between TFs and epigenetic modifications [88]. Recently, our colleagues Zhao et al. have observed that regulatory factor X-box 1 (RFX1), a TF belonging to the regulation factor for X-box protein family, is decreased in SLE CD4+ T cells and regulates DNA methylation in CD4+ T cells [93]. In this study, we found that RFX1 can recruit DNMT1, HDAC1, and suppressor of variegation 3–9 (Drosophila) homolog 1 (SUV39H1) to target gene promoter. RFX1 downregulation contributes to DNA hypomethylation and histone H3 hyperacetylation and decreased H3K9 tri-methylation in CD11a and CD70 promoter regions in lupus CD4+ T cells, which leads to CD11a and CD70 overexpression, thereby triggering an autoimmune response [93, 94].
One of our important findings is the role of TF E4BP4 in SLE and its epigenetic mechanism. We observed that E4BP4 directly regulates CD40L expression by binding to the promoter region and altering the histone acetylation and methylation of the CD40L loci. The effect of E4BP4 has been proven by its overexpression in lupus CD4+ T cells which inhibited the activation and self-reactivity of the T cells [95]. In addition, Tsokos et al. also reported that the TF CREMα recruited DNMT3a to the IL2 promoter and favored a permissive chromatin conformation at the IL17A locus. These findings led to the increased expression of IL-2 and IL-17 in naive, central memory, and effector memory CD4+ T cells, which might contribute to the pathogenesis of SLE [96]. More recently, our group observed a role for miR-1246 in the regulation of TF Early B cell factor 1 (EBF1), which regulated the development, activation, and proliferation of B cells by activating the AKT signaling pathway, suggesting a regulatory role for miR-1246 in the development of SLE [97].
Rheumatoid Arthritis
RA is a chronic and systemic inflammatory autoimmune condition that primarily affects the joints and is characterized by the progressive destruction of joints [98]. The synovial fibroblasts (SF) have been identified as the main player in the initiation of the disease [99]. Epigenetic regulation is also a novel field of research in RA, but there are many lines of evidence supporting its role in the pathogenesis of this disease [100]. These epigenetic mechanisms include DNA hyper-methylation [101], aberrant histone modification [102], and differentially expressed miRNAs [103] and lncRNAs [104]. However, most of the evidence regarding the interaction between TFs and these modifications is restricted to indirect evidence from the NF-κB pathway.
SIRT6, a member of the HDAC sirtuin family, has been found to interact with the NF-κB subunit RelA, to suppress NF-κB-dependent gene expression by deacetylating H3K9 [105], and to further inhibit the activity of NF-κB target gene-related immune responses that may contribute to RA [100]. There is additional indirect evidence from the study of death receptor 3 (DR-3), a protein that causes apoptosis and activation of NF-κB. The DR-3 promoter was found to be hyper-methylated in RA, causing the synovial cells to be resistant to apoptosis [106, 107]. However, as the interplay between TFs and epigenetics has been well studied in T cell differentiation [108] and RA is a Th1- and Th17-associated disease [109], increasing studies may be conducted on this relationship in near future.
Type 1 Diabetes
T1D is a T cell-mediated autoimmune disorder in which T cells cannot distinguish self-pancreatic cells, especially the beta cells, from dangerous pathogens and consequently destroy the pancreas [110]. As with many other autoimmune diseases, T1D develops in genetically susceptible individuals whose immune function is modulated by environmental factors [111, 112]. Epigenetic mechanisms partially explain the influence of environment agents, especially the diet, on T1D [113, 114]. It has been reported that hypomethylation of the transcription factor HOXA9 contributes to T1D [115], and a recent study on discordant monozygotic twins showed that global DNA hypomethylation within gene promoter regions may contribute to T1D [114, 116]. In addition, the TF NF-κB is also upregulated by H3K4 methyltransferase and causes an increase in inflammatory gene expression in diabetic mice [117]. Enhanced NF-κB-p65 gene expression also resulted from increased H3K4me1 and reduced H3K9 methylation [118]. Moreover, DNA methylation was found to block the binding of IRF7 to Foxp3, which reduced the number of regulatory T cells and contributed to the pathogenesis of T1D [119, 120]. In an autoimmune diabetes mouse model, Foxp3 was found to be unable to interact with the HAT Tip60, the histone deacetylase HDAC7, and the Ikaros family zinc finger 4, Eos, which led to reduced Foxp3 acetylation and enhanced K48-linked polyubiquitylation, contributing to Treg cell insufficiency that subsequently enables autoimmunity [121].
Systemic Sclerosis
SSc is a rare, connective tissue disease of unknown etiology that is characterized by the accumulation of collagen deposits in the skin and other tissues with a progressive vasculopathy [122]. Similar to other rheumatoid diseases, SSc has been reported to result in part from epigenetic modifications [123] based on evidence including the downregulation of miR-29a in SSc fibroblasts [123, 124] and abnormal DNA methylation level on CD4+ T cells and on certain autoimmune-related genes, such as Foxp3, CD40L, and CD11a in CD4+ T cells from SSc patients [125–128]. In SSc, hypermethylated CpG islands are found in the Fli1 promoter, which is a transcription factor that inhibits collagen production. The reduced expression of Fli1 increases collagen synthesis and promotes collagen accumulation and the tissue fibrosis that is a characteristic feature of SSc [129, 130]. Moreover, increased DNA methylation is observed at the Foxp3 locus, which is the key TF that regulates Treg cell generation, contributing to the reduced number of Treg cells in SSc [127]. However, hyper-methylation of the Foxp3 locus has also been reported in SSc and is thought to be regulated by X chromosomal inactivation [131]. In addition, the HDAC inhibitor trichostatin A (TSA) reportedly inhibits the TGF-β-induced activation of the TFs SMAD3 and SMAD4, which are the downstream of TGF-β, and influences the signaling pathways involved in fibrosis [132], indicating the involvement of this interaction in SSc.
Multiple Sclerosis
MS is an inflammatory condition characterized by immune system reactivity against myelin in the central nervous system that results in varying degrees of either relapsing or progressive neurological degeneration. Epigenetic mechanisms are implicated in the pathogenesis of MS [15, 133]. In epigenetic studies, it is reported that the HDAC sirtuin 1 deacetylates interferon regulatory factor 1 (IRF1), leading to fewer of the Th17 cells that play a critical role in MS [134]. Increased DNA methylation is also observed at the Foxp3 locus, decreasing Treg activity and further contributing to MS [135]. In addition, both hypomethylation at the Il17A/Infg loci and increased methylation at the Il4/Foxp3 loci are found and contribute to the imbalance of Th1 and Th2 responses in MS [136]. Interestingly, miR-155 deficiency in Treg cells results in enhanced suppressor of cytokine signaling 1 (SOCS1) expression, with impaired activation of signal transducer and activator of transcription 5 (STAT5) TF in response to limiting amounts of IL-2 [137], dampening the Treg activity in MS.
Conclusions
The field of epigenetics is growing and providing novel insights into the pathogenesis of autoimmune diseases. This exciting progress in epigenetic research has enabled us to explore new explanations for the etiology of these diseases. Increasing evidence supports the involvement of abnormal epigenetic regulation mediated by TFs in the pathogenesis of autoimmune diseases, and technological advances enable epigenomic analysis on a large scale and the investigation of interaction between epigenetic mechanisms and TFs in genome-wide studies. Although a lot of evidence on certain epigenetic regulation in diseases has been reported in recent decades, not many reports have explored the upstream and downstream players of these epigenetic regulations, not to mention the interplay of TFs and epigenetic modifications (recent progress is summarized in Table 1). Moreover, to date, not much progress has been made on understanding this dynamic interplay of TFs and epigenetic modifications in the context of autoimmune conditions. Further study is needed to better understand the regulation of TFs and epigenetic mechanisms in the pathogenesis of autoimmune diseases and identify the optimal therapeutic targets.
References
Lu Q (2014) Unmet needs in autoimmunity and potential new tools. Clin Rev Allergy Immunol 47:111–8
Wandstrat A, Wakeland E (2001) The genetics of complex autoimmune diseases: non-MHC susceptibility genes. Nat Immunol 2:802–9
Rhodes B, Vyse TJ (2008) The genetics of SLE: an update in the light of genome-wide association studies. Rheumatology (Oxford) 47:1603–11
Alarcon-Riquelme ME (2007) Recent advances in the genetics of autoimmune diseases. Ann N Y Acad Sci 1110:1–9
Floreani A, Leung PS, Gershwin ME (2015) Environmental basis of autoimmunity Clin Rev Allergy Immunol
Jeffries MA, Sawalha AH (2011) Epigenetics in systemic lupus erythematosus: leading the way for specific therapeutic agents. Int J Clin Rheumatol 6:423–39
Ballestar E (2010) Epigenetics lessons from twins: prospects for autoimmune disease. Clin Rev Allergy Immunol 39:30–41
Brown CC, Wedderburn LR (2015) Genetics: mapping autoimmune disease epigenetics: what’s on the horizon? Nat Rev Rheumatol 11:131–2
Jeffries MA, Sawalha AH (2015) Autoimmune disease in the epigenetic era: how has epigenetics changed our understanding of disease and how can we expect the field to evolve? Expert Rev Clin Immunol 11:45–58
Quddus J, Johnson KJ, Gavalchin J, Amento EP, Chrisp CE, Yung RL et al (1993) Treating activated CD4+ T cells with either of two distinct DNA methyltransferase inhibitors, 5-azacytidine or procainamide, is sufficient to cause a lupus-like disease in syngeneic mice. J Clin Invest 92:38–53
Sanchez-Pernaute O, Ospelt C, Neidhart M, Gay S (2008) Epigenetic clues to rheumatoid arthritis. J Autoimmun 30:12–20
Kragt J, van Amerongen B, Killestein J, Dijkstra C, Uitdehaag B, Polman C et al (2009) Higher levels of 25-hydroxyvitamin D are associated with a lower incidence of multiple sclerosis only in women. Mult Scler 15:9–15
Oksenberg JR, Baranzini SE, Sawcer S, Hauser SL (2008) The genetics of multiple sclerosis: SNPs to pathways to pathogenesis. Nat Rev Genet 9:516–26
Koch MW, Metz LM, Kovalchuk O (2013) Epigenetics and miRNAs in the diagnosis and treatment of multiple sclerosis. Trends Mol Med 19:23–30
Kucukali CI, Kurtuncu M, Coban A, Cebi M, Tuzun E (2014) Epigenetics of multiple sclerosis: an updated review. Neuromolecular Med
Strickland FM, Li Y, Johnson K, Sun Z, Richardson BC (2015) CD4(+) T cells epigenetically modified by oxidative stress cause lupus-like autoimmunity in mice. J Autoimmun 62:75–80
Bao Y, Cao X (2015) Epigenetic control of B cell development and B-cell-related immune disorders. Clin Rev Allergy Immunol
Renauer P, Coit P, Sawalha AH (2015) Epigenetics and vasculitis: a comprehensive review. Clin Rev Allergy Immunol
Saito Y, Saito H, Liang G, Friedman JM (2014) Epigenetic alterations and microRNA misexpression in cancer and autoimmune diseases: a critical review. Clin Rev Allergy Immunol 47:128–35
Bernstein BE, Meissner A, Lander ES (2007) The mammalian epigenome. Cell 128:669–81
Denis H, Ndlovu MN, Fuks F (2011) Regulation of mammalian DNA methyltransferases: a route to new mechanisms. EMBO Rep 12:647–56
Abdel-Wahab O, Mullally A, Hedvat C, Garcia-Manero G, Patel J, Wadleigh M et al (2009) Genetic characterization of TET1, TET2, and TET3 alterations in myeloid malignancies. Blood 114:144–7
Bird A (2002) DNA methylation patterns and epigenetic memory. Genes Dev 16:6–21
Klose RJ, Bird AP (2006) Genomic DNA methylation: the mark and its mediators. Trends Biochem Sci 31:89–97
Fan S, Zhang X (2009) CpG island methylation pattern in different human tissues and its correlation with gene expression. Biochem Biophys Res Commun 383:421–5
Perini G, Diolaiti D, Porro A, Della Valle G (2005) In vivo transcriptional regulation of N-Myc target genes is controlled by E-box methylation. Proc Natl Acad Sci U S A 102:12117–22
Kim J, Kollhoff A, Bergmann A, Stubbs L (2003) Methylation-sensitive binding of transcription factor YY1 to an insulator sequence within the paternally expressed imprinted gene, Peg3. Hum Mol Genet 12:233–45
Chatterjee R, Vinson C (1819) CpG methylation recruits sequence specific transcription factors essential for tissue specific gene expression. Biochim Biophys Acta 2012:763–70
Nan X, Ng HH, Johnson CA, Laherty CD, Turner BM, Eisenman RN et al (1998) Transcriptional repression by the methyl-CpG-binding protein MeCP2 involves a histone deacetylase complex. Nature 393:386–9
Medvedeva YA, Khamis AM, Kulakovskiy IV, Ba-Alawi W, Bhuyan MS, Kawaji H et al (2014) Effects of cytosine methylation on transcription factor binding sites. BMC Genomics 15:119
Ali I, Seker H (2010) A comparative study for characterisation and prediction of tissue-specific DNA methylation of CpG islands in chromosomes 6, 20 and 22. Conf Proc IEEE Eng Med Biol Soc 2010:1832–5
Ghosh S, Yates AJ, Fruhwald MC, Miecznikowski JC, Plass C, Smiraglia D (2010) Tissue specific DNA methylation of CpG islands in normal human adult somatic tissues distinguishes neural from non-neural tissues. Epigenetics 5:527–38
Cohen CJ, Crome SQ, MacDonald KG, Dai EL, Mager DL, Levings MK (2011) Human Th1 and Th17 cells exhibit epigenetic stability at signature cytokine and transcription factor loci. J Immunol 187:5615–26
Ivascu C, Wasserkort R, Lesche R, Dong J, Stein H, Thiel A et al (2007) DNA methylation profiling of transcription factor genes in normal lymphocyte development and lymphomas. Int J Biochem Cell Biol 39:1523–38
Perera A, Eisen D, Wagner M, Laube SK, Kunzel AF, Koch S et al (2015) TET3 is recruited by REST for context-specific hydroxymethylation and induction of gene expression. Cell Rep
Hervouet E, Vallette FM, Cartron PF (2010) Dnmt1/transcription factor interactions: an alternative mechanism of DNA methylation inheritance. Genes Cancer 1:434–43
Zhang Q, Wang HY, Woetmann A, Raghunath PN, Odum N, Wasik MA (2006) STAT3 induces transcription of the DNA methyltransferase 1 gene (DNMT1) in malignant T lymphocytes. Blood 108:1058–64
Rothbart SB, Strahl BD (1839) Interpreting the language of histone and DNA modifications. Biochim Biophys Acta 2014:627–43
Peserico A, Simone C (2011) Physical and functional HAT/HDAC interplay regulates protein acetylation balance. J Biomed Biotechnol 2011:371832
Renaudineau Y, Youinou P (2011) Epigenetics and autoimmunity, with special emphasis on methylation. Keio J Med 60:10–6
Black JC, Van Rechem C, Whetstine JR (2012) Histone lysine methylation dynamics: establishment, regulation, and biological impact. Mol Cell 48:491–507
Gregory PD, Wagner K, Horz W (2001) Histone acetylation and chromatin remodeling. Exp Cell Res 265:195–202
Chuang HC, Chang CW, Chang GD, Yao TP, Chen H (2006) Histone deacetylase 3 binds to and regulates the GCMa transcription factor. Nucleic Acids Res 34:1459–69
Yao YL, Yang WM, Seto E (2001) Regulation of transcription factor YY1 by acetylation and deacetylation. Mol Cell Biol 21:5979–91
Katto J, Engel N, Abbas W, Herbein G, Mahlknecht U (2013) Transcription factor NFkappaB regulates the expression of the histone deacetylase SIRT1. Clin Epigenetics 5:11
Chen CZ, Li L, Lodish HF, Bartel DP (2004) MicroRNAs modulate hematopoietic lineage differentiation. Science 303:83–6
Fabian MR, Sonenberg N, Filipowicz W (2010) Regulation of mRNA translation and stability by microRNAs. Annu Rev Biochem 79:351–79
Inui M, Martello G, Piccolo S (2010) MicroRNA control of signal transduction. Nat Rev Mol Cell Biol 11:252–63
Bartel DP (2009) MicroRNAs: target recognition and regulatory functions. Cell 136:215–33
Baltimore D, Boldin MP, O’Connell RM, Rao DS, Taganov KD (2008) MicroRNAs: new regulators of immune cell development and function. Nat Immunol 9:839–45
O’Connell RM, Rao DS, Chaudhuri AA, Baltimore D (2010) Physiological and pathological roles for microRNAs in the immune system. Nat Rev Immunol 10:111–22
Yan S, Yim LY, Lu L, Lau CS, Chan VS (2014) MicroRNA regulation in systemic lupus erythematosus pathogenesis. Immune Netw 14:138–48
Johnson SM, Lin SY, Slack FJ (2003) The time of appearance of the C. elegans let-7 microRNA is transcriptionally controlled utilizing a temporal regulatory element in its promoter. Dev Biol 259:364–79
Biemar F, Zinzen R, Ronshaugen M, Sementchenko V, Manak JR, Levine MS (2005) Spatial regulation of microRNA gene expression in the Drosophila embryo. Proc Natl Acad Sci U S A 102:15907–11
Olson EN (2006) Gene regulatory networks in the evolution and development of the heart. Science 313:1922–7
Arora S, Rana R, Chhabra A, Jaiswal A, Rani V (2013) miRNA-transcription factor interactions: a combinatorial regulation of gene expression. Mol Genet Genomics 288:77–87
Chen CY, Chen ST, Fuh CS, Juan HF, Huang HC (2011) Coregulation of transcription factors and microRNAs in human transcriptional regulatory network. BMC Bioinformatics 12(Suppl 1):S41
Krol J, Loedige I, Filipowicz W (2010) The widespread regulation of microRNA biogenesis, function and decay. Nat Rev Genet 11:597–610
Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A et al (2012) Landscape of transcription in human cells. Nature 489:101–8
International Human Genome Sequencing C (2004) Finishing the euchromatic sequence of the human genome. Nature 431:931–45
Rinn JL, Chang HY (2012) Genome regulation by long noncoding RNAs. Annu Rev Biochem 81:145–66
Kretz M, Siprashvili Z, Chu C, Webster DE, Zehnder A, Qu K et al (2013) Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 493:231–5
Johnsson P, Ackley A, Vidarsdottir L, Lui WO, Corcoran M, Grander D et al (2013) A pseudogene long-noncoding-RNA network regulates PTEN transcription and translation in human cells. Nat Struct Mol Biol 20:440–6
Wapinski O, Chang HY (2011) Long noncoding RNAs and human disease. Trends Cell Biol 21:354–61
Li Z, Chao TC, Chang KY, Lin N, Patil VS, Shimizu C et al (2014) The long noncoding RNA THRIL regulates TNFalpha expression through its interaction with hnRNPL. Proc Natl Acad Sci U S A 111:1002–7
Carpenter S, Aiello D, Atianand MK, Ricci EP, Gandhi P, Hall LL et al (2013) A long noncoding RNA mediates both activation and repression of immune response genes. Science 341:789–92
Wang P, Xue Y, Han Y, Lin L, Wu C, Xu S et al (2014) The STAT3-binding long noncoding RNA lnc-DC controls human dendritic cell differentiation. Science 344:310–3
Hu G, Tang Q, Sharma S, Yu F, Escobar TM, Muljo SA et al (2013) Expression and regulation of intergenic long noncoding RNAs during T cell development and differentiation. Nat Immunol 14:1190–8
Heward JA, Lindsay MA (2014) Long non-coding RNAs in the regulation of the immune response. Trends Immunol 35:408–19
Hrdlickova B, Kumar V, Kanduri K, Zhernakova DV, Tripathi S, Karjalainen J et al (2014) Expression profiles of long non-coding RNAs located in autoimmune disease-associated regions reveal immune cell-type specificity. Genome Med 6:88
Wang KC, Yang YW, Liu B, Sanyal A, Corces-Zimmerman R, Chen Y et al (2011) A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 472:120–4
Nagano T, Mitchell JA, Sanz LA, Pauler FM, Ferguson-Smith AC, Feil R et al (2008) The Air noncoding RNA epigenetically silences transcription by targeting G9a to chromatin. Science 322:1717–20
Chen R, Yang Z, Zhou Q (2004) Phosphorylated positive transcription elongation factor b (P-TEFb) is tagged for inhibition through association with 7SK snRNA. J Biol Chem 279:4153–60
Sharma S, Findlay GM, Bandukwala HS, Oberdoerffer S, Baust B, Li Z et al (2011) Dephosphorylation of the nuclear factor of activated T cells (NFAT) transcription factor is regulated by an RNA-protein scaffold complex. Proc Natl Acad Sci U S A 108:11381–6
Beltran M, Puig I, Pena C, Garcia JM, Alvarez AB, Pena R et al (2008) A natural antisense transcript regulates Zeb2/Sip1 gene expression during Snail1-induced epithelial-mesenchymal transition. Genes Dev 22:756–69
Annilo T, Kepp K, Laan M (2009) Natural antisense transcript of natriuretic peptide precursor A (NPPA): structural organization and modulation of NPPA expression. BMC Mol Biol 10:81
D’Cruz DP, Khamashta MA, Hughes GR (2007) Systemic lupus erythematosus. Lancet 369:587–96
Yu C, Gershwin ME, Chang C (2014) Diagnostic criteria for systemic lupus erythematosus: a critical review. J Autoimmun 48–49:10–3
Tan EM, Kunkel HG (1966) Characteristics of a soluble nuclear antigen precipitating with sera of patients with systemic lupus erythematosus. J Immunol 96:464–71
Takeno M, Nagafuchi H, Kaneko S, Wakisaka S, Oneda K, Takeba Y et al (1997) Autoreactive T cell clones from patients with systemic lupus erythematosus support polyclonal autoantibody production. J Immunol 158:3529–38
Santulli-Marotto S, Retter MW, Gee R, Mamula MJ, Clarke SH (1998) Autoreactive B cell regulation: peripheral induction of developmental arrest by lupus-associated autoantigens. Immunity 8:209–19
Gatto M, Zen M, Ghirardello A, Bettio S, Bassi N, Iaccarino L et al (2013) Emerging and critical issues in the pathogenesis of lupus. Autoimmun Rev 12:523–36
Ghodke-Puranik Y, Niewold TB (2015) Immunogenetics of systemic lupus erythematosus: a comprehensive review. J Autoimmun 64:125–36
Kuhn A, Wenzel J, Weyd H (2014) Photosensitivity, apoptosis, and cytokines in the pathogenesis of lupus erythematosus: a critical review. Clin Rev Allergy Immunol 47:148–62
Meroni PL, Penatti AE (2015) Epigenetics and systemic lupus erythematosus: unmet needs. Clin Rev Allergy Immunol
Cannat A, Seligmann M (1968) Induction by isoniazid and hydrallazine of antinuclear factors in mice. Clin Exp Immunol 3:99–105
Javierre BM, Fernandez AF, Richter J, Al-Shahrour F, Martin-Subero JI, Rodriguez-Ubreva J et al (2010) Changes in the pattern of DNA methylation associate with twin discordance in systemic lupus erythematosus. Genome Res 20:170–9
Zhao M, Liu S, Luo S, Wu H, Tang M, Cheng W et al (2014) DNA methylation and mRNA and microRNA expression of SLE CD4+ T cells correlate with disease phenotype. J Autoimmun 54:127–36
Zhou Y, Qiu X, Luo Y, Yuan J, Li Y, Zhong Q et al (2011) Histone modifications and methyl-CpG-binding domain protein levels at the TNFSF7 (CD70) promoter in SLE CD4+ T cells. Lupus 20:1365–71
Zhao S, Wang Y, Liang Y, Zhao M, Long H, Ding S et al (2011) MicroRNA-126 regulates DNA methylation in CD4+ T cells and contributes to systemic lupus erythematosus by targeting DNA methyltransferase 1. Arthritis Rheum 63:1376–86
Pan W, Zhu S, Yuan M, Cui H, Wang L, Luo X et al (2010) MicroRNA-21 and microRNA-148a contribute to DNA hypomethylation in lupus CD4+ T cells by directly and indirectly targeting DNA methyltransferase 1. J Immunol 184:6773–81
Shi L, Zhang Z, Yu AM, Wang W, Wei Z, Akhter E et al (2014) The SLE transcriptome exhibits evidence of chronic endotoxin exposure and has widespread dysregulation of non-coding and coding RNAs. PLoS One 9, e93846
Zhao M, Sun Y, Gao F, Wu X, Tang J, Yin H et al (2010) Epigenetics and SLE: RFX1 downregulation causes CD11a and CD70 overexpression by altering epigenetic modifications in lupus CD4+ T cells. J Autoimmun 35:58–69
Zhao M, Wu X, Zhang Q, Luo S, Liang G, Su Y et al (2010) RFX1 regulates CD70 and CD11a expression in lupus T cells by recruiting the histone methyltransferase SUV39H1. Arthritis Res Ther 12:R227
Zhao M, Liu Q, Liang G, Wang L, Luo S, Tang Q et al (2013) E4BP4 overexpression: a protective mechanism in CD4+ T cells from SLE patients. J Autoimmun 41:152–60
Hedrich CM, Crispin JC, Rauen T, Ioannidis C, Apostolidis SA, Lo MS et al (2012) cAMP response element modulator alpha controls IL2 and IL17A expression during CD4 lineage commitment and subset distribution in lupus. Proc Natl Acad Sci U S A 109:16606–11
Luo S, Liu Y, Liang G, Zhao M, Wu H, Liang Y et al (2015) The role of microRNA-1246 in the regulation of B cell activation and the pathogenesis of systemic lupus erythematosus. Clin Epigenetics 7:24
Kourilovitch M, Galarza-Maldonado C, Ortiz-Prado E (2014) Diagnosis and classification of rheumatoid arthritis. J Autoimmun 48–49:26–30
Zhu X, Song Y, Huo R, Zhang J, Sun S, He Y et al (2015) Cyr61 participates in the pathogenesis of rheumatoid arthritis by promoting proIL-1beta production by fibroblast-like synoviocytes through an AKT-dependent NF-kappaB signaling pathway. Clin Immunol 157:187–97
Klein K, Gay S (2015) Epigenetics in rheumatoid arthritis. Curr Opin Rheumatol 27:76–82
Kuchen S, Seemayer CA, Rethage J, von Knoch R, Kuenzler P, Beat AM et al (2004) The L1 retroelement-related p40 protein induces p38delta MAP kinase. Autoimmunity 37:57–65
Grabiec AM, Tak PP, Reedquist KA (2008) Targeting histone deacetylase activity in rheumatoid arthritis and asthma as prototypes of inflammatory disease: should we keep our HATs on? Arthritis Res Ther 10:226
Nakamachi Y, Kawano S, Takenokuchi M, Nishimura K, Sakai Y, Chin T et al (2009) MicroRNA-124a is a key regulator of proliferation and monocyte chemoattractant protein 1 secretion in fibroblast-like synoviocytes from patients with rheumatoid arthritis. Arthritis Rheum 60:1294–304
Muller N, Doring F, Klapper M, Neumann K, Schulte DM, Turk K et al (2014) Interleukin-6 and tumour necrosis factor-alpha differentially regulate lincRNA transcripts in cells of the innate immune system in vivo in human subjects with rheumatoid arthritis. Cytokine 68:65–8
Kawahara TL, Michishita E, Adler AS, Damian M, Berber E, Lin M et al (2009) SIRT6 links histone H3 lysine 9 deacetylation to NF-kappaB-dependent gene expression and organismal life span. Cell 136:62–74
Takami N, Osawa K, Miura Y, Komai K, Taniguchi M, Shiraishi M et al (2006) Hypermethylated promoter region of DR3, the death receptor 3 gene, in rheumatoid arthritis synovial cells. Arthritis Rheum 54:779–87
Bull MJ, Williams AS, Mecklenburgh Z, Calder CJ, Twohig JP, Elford C et al (2008) The Death Receptor 3-TNF-like protein 1A pathway drives adverse bone pathology in inflammatory arthritis. J Exp Med 205:2457–64
Suarez-Alvarez B, Rodriguez RM, Fraga MF, Lopez-Larrea C (2012) DNA methylation: a promising landscape for immune system-related diseases. Trends Genet 28:506–14
Kosmaczewska A, Ciszak L, Swierkot J, Szteblich A, Kosciow K, Frydecka I (2015) Exogenous IL-2 controls the balance in Th1, Th17, and Treg cell distribution in patients with progressive rheumatoid arthritis treated with TNF-alpha inhibitors. Inflammation 38:765–74
Xie Z, Chang C, Zhou Z (2014) Molecular mechanisms in autoimmune type 1 diabetes: a critical review. Clin Rev Allergy Immunol 47:174–92
Burgio E, Lopomo A, Migliore L (2015) Obesity and diabetes: from genetics to epigenetics. Mol Biol Rep 42:799–818
Noble JA (2015) Immunogenetics of type 1 diabetes: a comprehensive review. J Autoimmun 64:101–12
Dang MN, Buzzetti R, Pozzilli P (2013) Epigenetics in autoimmune diseases with focus on type 1 diabetes. Diabetes Metab Res Rev 29:8–18
Stefan M, Zhang W, Concepcion E, Yi Z, Tomer Y (2014) DNA methylation profiles in type 1 diabetes twins point to strong epigenetic effects on etiology. J Autoimmun 50:33–7
Miao F, Smith DD, Zhang L, Min A, Feng W, Natarajan R (2008) Lymphocytes from patients with type 1 diabetes display a distinct profile of chromatin histone H3 lysine 9 dimethylation: an epigenetic study in diabetes. Diabetes 57:3189–98
Elboudwarej E, Cole M, Briggs FB, Fouts A, Fain PR, Quach H et al (2016) Hypomethylation within gene promoter regions and type 1 diabetes in discordant monozygotic twins. Journal of autoimmunity
Li Y, Reddy MA, Miao F, Shanmugam N, Yee JK, Hawkins D et al (2008) Role of the histone H3 lysine 4 methyltransferase, SET7/9, in the regulation of NF-kappaB-dependent inflammatory genes. Relevance to diabetes and inflammation. J Biol Chem 283:26771–81
Brasacchio D, Okabe J, Tikellis C, Balcerczyk A, George P, Baker EK et al (2009) Hyperglycemia induces a dynamic cooperativity of histone methylase and demethylase enzymes associated with gene-activating epigenetic marks that coexist on the lysine tail. Diabetes 58:1229–36
Wang Z, Zheng Y, Hou C, Yang L, Li X, Lin J et al (2013) DNA methylation impairs TLR9 induced Foxp3 expression by attenuating IRF-7 binding activity in fulminant type 1 diabetes. J Autoimmun 41:50–9
Tan T, Xiang Y, Chang C, Zhou Z (2014) Alteration of regulatory T cells in type 1 diabetes mellitus: a comprehensive review. Clin Rev Allergy Immunol 47:234–43
Bettini ML, Pan F, Bettini M, Finkelstein D, Rehg JE, Floess S et al (2012) Loss of epigenetic modification driven by the Foxp3 transcription factor leads to regulatory T cell insufficiency. Immunity 36:717–30
Hudson M, Fritzler MJ (2014) Diagnostic criteria of systemic sclerosis. J Autoimmun 48–49:38–41
Ciechomska M, van Laar JM, O’Reilly S (2014) Emerging role of epigenetics in systemic sclerosis pathogenesis. Genes Immun 15:433–9
Maurer B, Stanczyk J, Jungel A, Akhmetshina A, Trenkmann M, Brock M et al (2010) MicroRNA-29, a key regulator of collagen expression in systemic sclerosis. Arthritis Rheum 62:1733–43
Lei W, Luo Y, Lei W, Luo Y, Yan K, Zhao S et al (2009) Abnormal DNA methylation in CD4+ T cells from patients with systemic lupus erythematosus, systemic sclerosis, and dermatomyositis. Scand J Rheumatol 38:369–74
Lian X, Xiao R, Hu X, Kanekura T, Jiang H, Li Y et al (2012) DNA demethylation of CD40l in CD4+ T cells from women with systemic sclerosis: a possible explanation for female susceptibility. Arthritis Rheum 64:2338–45
Wang YY, Wang Q, Sun XH, Liu RZ, Shu Y, Kanekura T et al (2014) DNA hypermethylation of the forkhead box protein 3 (FOXP3) promoter in CD4+ T cells of patients with systemic sclerosis. Br J Dermatol 171:39–47
Wang Y, Shu Y, Xiao Y, Wang Q, Kanekura T, Li Y et al (2014) Hypomethylation and overexpression of ITGAL (CD11a) in CD4(+) T cells in systemic sclerosis. Clin Epigenetics 6:25
Kubo M, Czuwara-Ladykowska J, Moussa O, Markiewicz M, Smith E, Silver RM et al (2003) Persistent down-regulation of Fli1, a suppressor of collagen transcription, in fibrotic scleroderma skin. Am J Pathol 163:571–81
Wang Y, Fan PS, Kahaleh B (2006) Association between enhanced type I collagen expression and epigenetic repression of the FLI1 gene in scleroderma fibroblasts. Arthritis Rheum 54:2271–9
Broen JC, Coenen MJ, Radstake TR (2011) Deciphering the genetic background of systemic sclerosis. Expert Rev Clin Immunol 7:449–62
Huber LC, Distler JH, Moritz F, Hemmatazad H, Hauser T, Michel BA et al (2007) Trichostatin A prevents the accumulation of extracellular matrix in a mouse model of bleomycin-induced skin fibrosis. Arthritis Rheum 56:2755–64
Hollenbach JA, Oksenberg JR (2015) The immunogenetics of multiple sclerosis: a comprehensive review. J Autoimmun 64:13–25
Yang H, Lee SM, Gao B, Zhang J, Fang D (2013) Histone deacetylase sirtuin 1 deacetylates IRF1 protein and programs dendritic cells to control Th17 protein differentiation during autoimmune inflammation. J Biol Chem 288:37256–66
Liu B, Tahk S, Yee KM, Fan G, Shuai K (2010) The ligase PIAS1 restricts natural regulatory T cell differentiation by epigenetic repression. Science 330:521–5
Guan H, Nagarkatti PS, Nagarkatti M (2011) CD44 Reciprocally regulates the differentiation of encephalitogenic Th1/Th17 and Th2/regulatory T cells through epigenetic modulation involving DNA methylation of cytokine gene promoters, thereby controlling the development of experimental autoimmune encephalomyelitis. J Immunol 186:6955–64
Lu LF, Thai TH, Calado DP, Chaudhry A, Kubo M, Tanaka K et al (2009) Foxp3-dependent microRNA155 confers competitive fitness to regulatory T cells by targeting SOCS1 protein. Immunity 30:80–91
Acknowledgments
This work was supported by the National Natural Science Foundation of China (grant no. 81220108017, no. 81430074, no. 81373205, and no. 81270024), the Specialized Research Fund for the Doctoral Program of Higher Education (grant no. 20120162130003), and Hunan Natural Science Funds for Distinguished Young Scientists (No. 14JJ1009).
Author information
Authors and Affiliations
Corresponding author
Additional information
Haijing Wu and Ming Zhao contributed equally to this work.
Rights and permissions
About this article
Cite this article
Wu, H., Zhao, M., Yoshimura, A. et al. Critical Link Between Epigenetics and Transcription Factors in the Induction of Autoimmunity: a Comprehensive Review. Clinic Rev Allerg Immunol 50, 333–344 (2016). https://doi.org/10.1007/s12016-016-8534-y
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12016-016-8534-y







 ); inhibition (
); inhibition ( )
)