Abstract
Cryptosporidiosis accounts for a surge in infant (<5 years) mortality and morbidity. To date, several drug discovery efforts have been put in place to develop effective therapeutic options against the causative parasite. Based on a recent report, P131 spares inosine monophosphate dehydrogenase (IMPDH) in a eukaryotic model (mouse IMPDH (mIMPDH)) while binding selectively to the NAD+ site in Cryptosporidium parvum (CpIMPDH). However, no structural detail exists on the underlining mechanisms of P131-CpIMPDH selective targeting till date. To this effect, we investigate the selective inhibitory dynamics of P131 in CpIMPDH relative to mIMPDH via molecular biocomputation methods. Pairwise sequence alignment revealed prominent variations at the NAD+ binding regions of both proteins that accounted for disparate P131 binding activities. The influence of these variations was further revealed by the MM/PBSA energy estimations coupled with per-residue energy decomposition which monitored the systematic binding of the compound. Furthermore, relative high-affinity interactions occurred at the CpIMPDH NAD+ site which were majorly mediated by SER22, VAL24, PRO26, SER354, GLY357, and TYR358 located on chain D. These residues are unique to the parasite IMPDH form and not in the eukaryotic protein, highlighting variations that account for preferential P131 binding. Molecular insights provided herein corroborate previous experimental reports and further underpin the basis of CpIMPDH inhibitor selectivity. Findings from this study could present attractive prospects toward the design of novel anticryptosporidials with improved selectivity and binding affinity against parasitic targets.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cryptosporidium is gaining increasing recognition as the most significant protozoan parasite causing diarrhea in animals and humans [1]. Presently, it is reputed to be the principal cause of mortality in children less than 5 years old, claiming 800,000 lives annually and close to three million cases of detected infection [2]. The bulk of the burden of this infection falls in sub-Saharan Africa and South Asia [3,4,5,6]. A joint expert committee from the Food and Agricultural Organization and World Health Organization, undertook a worldwide ranking which placed Cryptosporidium as 5th among the 25 most important foodborne parasites [7, 8]
The infection is self-limiting in immunocompetent individuals and typically would resolve within 2 weeks [9]. This is converse to the outcome in immunodeficient individuals (human immunodeficiency virus/acquired immunodeficiency syndrome (HIV/AIDS)), organ transplant patients, etc., where infection can cause severe and chronic diarrhea, wasting, and most times death [10]. Symptoms are most times aggravated when HIV/AIDS is left untreated, especially in low-resource countries where antiretroviral drugs are not easily accessible and affordable [11].
The causative agent of human cryptosporidiosis is either Cryptosporidium hominis or Cryptosporidium parvum (Cp) [12, 13]. The latter has been found to infect several mammals and therefore responsible for most zoonotic infections [7]. Some risk factors include contaminated water facilities and foods, unhygienic environments, and malnutrition [14]. South Africa is not spared from this menace as Cryptosporidium species are frequently isolated from her surface waters used for domestic purposes [15]. The circumstance poses more problems as HIV/AIDS is endemic in South Africa, with the prevalence rates among the highest in the world [16].
There is a shortage of therapeutic agents employed in the treatment of cryptosporidiosis. Nitazoxanide remains the only Food and Drug Administration (FDA)-approved drug [17]. As it stands, nitazoxanide has an inexplicit mechanism of action, and it is not especially effective in immunocompromised individuals [18]. Nitazoxanide administration is also punctuated with adverse side effects which include hives, decreased liver function, respiratory problems, etc. [19]. Apart from nitazoxanide, several drugs though not FDA approved, have been deployed in the treatment of cryptosporidiosis. However, they also have limited efficacy as they failed in controlled trials in AIDS patients [20, 21]. Examples include CD40 agonist antibody [22], paromomycin [23, 24], spiramycin, azithromycin, and bovine anti-Cryptosporidium immunoglobulin [20, 25, 26], rifamycin [27], rifaximin [28, 29]. Better responses have been documented when nitazoxanide, azithromycin, and/or paromomycin are combined [30,31,32]. In the wake of the ineffectiveness of the drugs, as mentioned above, the discovery of novel anticryptosporidial drugs is pressing.
Inosine monophosphate dehydrogenase (IMPDH) is a crucial enzyme in Cryptosporidium and has been a target for some antibiotic discovery [33,34,35,36]. It is responsible for the synthesis of guanine nucleotides. It catalyzes the initial and rate-limiting step of the oxidation of inosine 5′-monophosphate (IMP) to xanthosine 5′-monophosphate and simultaneously reducing NAD+ to NADH [37]. Its inhibition eventuates the diminution of guanine nucleotide reservoir and consequently, cell death [38]. It is worthy of note that Cryptosporidium though a eukaryote has a prokaryotic IMPDH, which was made possible by a horizontal gene transfer from a bacterium [9, 39]. Prokaryotic and eukaryotic IMPDHs are quite divergent in both their structural and kinetic properties, and these account for the selectivity in action of anticryptosporidial agents [40].
IMPDH are homotetramers with a D4 square symmetry [41]. It is made up of an IMP-binding site, a co-factor site (NAD+-binding site), and a mobile flap [42]. The four IMP and co-factor binding sites are situated close to the subunit interfaces [43]. The IMP site is confined within a monomer, and it is highly conserved in the IMPDH of all organisms [43]. On the other hand, the NAD+-binding site sequence is diverged.
The NAD+-binding site has been consistently targeted for the design of IMPDH inhibitors [33, 43, 44]. Three subdomains make up the NAD+-binding site, namely nicotinamide riboside-binding site (N-subdomain), the pyrophosphate-binding site (P-subdomain); and the adenosine-binding site (A-subdomain) [44]. The N-subdomain lies in the same monomer as IMP, and it is also strongly conserved, being the site of chemical transformation [42]. On the contrary, the adenosine subsite is highly divergent in different organisms [41]. In eukaryotic organism, it is in the same monomer as IMP [45] while in the prokaryote, it interacts with a pocket in the adjacent monomer [41].
A Cryptosporidium drug discovery program targeting IMPDH was undertaken by Hedstrom et al. research group, which led to the synthesis and production of a battery of Cryptosporidium inhibitors [9, 46,47,48,49,50,51,52,53,54,55]. Of all these inhibitors, in a murine model, P131 at a single dose demonstrated an equivalent activity when administered at 250 mg/kg bodyweight in comparison to the control group, which was treated with paromomycin at 2000 mg/kg body weight. Following a thrice-daily administration of both drugs at the same concentration stated earlier, P131 elicited a superior parasiticidal activity when compared to paromomycin. This was followed by a crystallographic determination of the structure of CpIMPDH with P131 bound at the NAD+-binding site [41].
Although the selectivity of anticryptosporidial drugs for Cryptosporidium parvum IMPDH (CpIMPDH) has been established [34, 42], the molecular mechanism of it leaving the hosts’ IMPDH unaffected is quite vague. Herein, we attempted to explain from bioinformatics and molecular biocomputation point of view, the molecular mechanisms associated with the selectivity of P131 toward CpIMPDH. We further compare the structural and conformational dynamics of P131 when bound to the NAD+ site of CpIMPDH and mouse IMPDH (mIMPDH). Findings from this study could present an attractive prospect in informing the design of new and potential inhibitors that possess improved binding, pharmacokinetics, and pharmacodynamics properties and selectivity for parasitic targets that are able to overcome the problems of drug resistance.
Computational Methodology
Sequence Mapping and Alignment
There was a need to determine the disparity among the binding site residues of CpIMPDH and mIMPDH as regards P131. The divergence in these residues may unravel the basis for the differential drug activity. Clustal Omega was used in performing sequence mapping and alignment. The well-suited match for the selected sequences is calculated and presented in a form in which their identities, similarities, and differences can be seen [56]. The FASTA sequences of the IMPDH of the two different hosts were retrieved from UNIPROT. CpIMPDH (ID: Q5CPK7) [57] and mIMPDH2 (ID: P24547) [58]. All these sequences were fed into the Clustal Omega webserver, and the output was retrieved. Residues that make up the P131 binding sites were then investigated for their variations.
Systems Retrieval, Preparation and Molecular Dynamics (MD) Simulations
Two systems were constituted (mIMPDH and CpIMPDH) as investigated in the experimental paper [48]. X-ray crystal structure of the catalytic domain of CpIMPDH co-crystallized with inhibitor P131, was obtained from the RSCB Protein Data Bank (Entry code: 4RV8) [41]. The protein (4RV8) retrieved was a tetramer, however, we only made use of a dimer (Chain A and D) to minimize computational cost and reflect experimental study [41]. The dimer used had 652 residues in total. The crystallized structure of mIMPDH is not available therefore we retrieved the FASTA sequence from UNIPROT (ID: P24547) [58] for homology modeling using the Swiss-Model algorithm [59]. Moreover, the crystal structure of Cricetulus griseus IMPDH in complex with mycophenolic acid (PDB entry: 1JR1.A) [60], which has a 99.22% identity with mIMPDH was used as the template for modeling. The modeled structure was then validated using RAMPAGE, PROCHECK, and VERIFY3D webservers [61,62,63].
Using the Clustal Omega sequence alignment, CpIMPDH and mIMPDH have 35.1% sequence similarity. It is important to note that mIMPDH shares 98.83% sequence similarity to human IMPDH (hIMPDH). The dissimilarities in both sequences hIMPDH and mIMPDH are in ILE192, ASP215, GLN265, ASP292, and ASN296 in hIMPDH, which are replaced by VAL192, ASN215, LEU265, GLU292, and SER296 in mIMPDH. The two proteins also share 100% sequence similarity in the co-factor binding site. This could explain the choice of selection of murine subject as a representative eukaryotic model in the report by Gorla et al. [48]. Although, the CgIMPDH on which the mIMPDH was built shared a higher similarity with hIMPDH (99.6%) than mIMPDH did (98.83%), we made use of the putative structure of mIMPDH rather than CgIMPDH to keep in line with the experimental model used in the investigating the in vivo efficacy of P131 as reported by Gorla et al. [48].
In CpIMPDH, the binding of P131 has been established to span both adenosine and nicotinamide subdomains of the co-factor (NAD+) binding site in IMPDH. Having a prokaryotic antecedent, the binding of P131 at the adenosine subsite interacts with chain D of the IMPDH [41]. The target protein structures (CpIMPDH and mIMPDH) were prepared using the graphical user interface (GUI) of UCSF Chimera [64], which involves the removal of ions, crystal waters, and nonstandard residues. Missing residues were added using MODELLER [65]. The co-crystallized P131 in CpIMPDH (4RV8) was retained and its binding orientation to chains A and D was used as the starting structure for MD simulation. However, ligand P131 retrieved from 4RV8 was docked to mIMPDH using Autodock Vina [66], which has been widely used over other docking algorithms due to its high scoring power [67]. The co-factor binding site of the mIMPDH was defined using the coordinates of co-crystallized P131 in the NAD+-binding site of CpIMPDH achieved by the superimposition of CpIMPDH and mIMPDH. The grid-box was defined as center (X = 2.6487, Y = 21.9974, Z = 77.3995) and size (X = 11.1647, Y = 9.4625, Z = 7.3236). The best docked pose (most negative score) was aligned with the co-crystallized P131 in CpIMPDH to obtain similar binding orientation for P131 in mIMPDH, which was also used as the starting MD structure [64]. The proteins were prepared further by Molegro Molecular Viewer software prior to MD simulation.
Molecular Dynamics Simulation
This was performed by using standard simulation protocol, which has been extensively employed in previous studies [68,69,70]. Each system was further subdivided into apo (IMPDH bound to IMP in its active site) and complexes which comprised of apo bound by P131 in the co-factor site. Afterward, these were set up for MD simulations according to previously reported protocols [71,72,73]. MD simulation was performed using Graphical Processor Unit version of the Particle Mesh Ewald Molecular Dynamics engine in AMBER18 suite coupled with integrated modules [74]. FF14SB forcefield was used in defining protein parameters. P131 parameterization was done using the ANTECHAMBER module, which generated atomic partial charges (Gasteiger—gaff) through the bcc charge scheme [75]. Sequel to this is the generation of topology and parameter files for complexes using the LEAP module, which neutralized the complexes by adding counter ions at a constant pH and solvated them in a 10 Å TIP3P water box. Partial minimization was executed in 2500 steps, using a 500 kcal/mol Å restraint potential followed by full minimization for 5000 steps with no energy restraints. The systems were heated for 50 ps from 0 to 300 K in an NVT canonical ensemble using a Langevin thermostat [76] and a harmonic restraint of 5 kcal/mol Å2. The systems were then equilibrated at 300 k for 1000 ps without energy restraints while the Berendsen barostat was used to maintain atmospheric pressure at 1 bar [77]. The MD production run was carried out for 280 ns with the SHAKE algorithm used in constricting all atomic hydrogen bonds [78]. The trajectories obtained were analyzed by the integrated CPPTRAJ and PTRAJ modules [79]. Origin data analytical tool was used in creating the needed plots [80]. 3D visualization of the structures and corresponding analyses were carried out on the GUI of UCSF Chimera.
Post Dynamic Analysis
Thermodynamics calculations
This calculation was used to probe the selective binding of P131 to CpIMPDH and mIMPDH. The estimation of free binding energy is vital as it gives insight into the binding affinity and stability of ligands in a complex. Molecular Mechanics/Poisson–Boltzmann Surface Area (MM/PBSA) method was used in the estimation of the binding free energy. MM/PBSA is a reliable analytical tool widely used for evaluating the interaction of ligands with biological macromolecules [81,82,83,84]. Mathematically, binding free energy is depicted by the following equation:
From the above equation, ∆Ggas depicts the gas-phase energy while the internal energy is represented as ∆Eint. In the same vein, the coulomb and van der Waals energies are represented as ∆Eele and ∆Evdw, respectively. In addition, ∆Gsol depicts the free energy of solvation while the polar solvation contribution is represented as ∆GPB. On the other hand, ∆Gnp,sol depicts the nonpolar contribution and is determined by the linear relationship between the surface tension proportionality constant (γ), solvent accessible surface area (SASA) (Å2), and a constant β. The surface tension proportionality constant (γ) is given as 0.0072 kcal/(mol Å2) and β as 0.92 kcal/mol, which is estimated by the SASA that is determined by using a water probe of radius 1.4 Å with a surface tension constant, γ of 0.0072 kcal/(mol Å2). Per-residue decomposition analysis was also carried out to obtain the different energies each binding site residues contributed to ligand affinity and stabilization.
Result and Discussion
Structural Validation of the Putative Model of mIMPDH
The modeled structure of mIMPDH retrieved from Swiss-Model webserver was validated and cross-validated on three online platforms—RAMPAGE, PROCHECK, and VERIFY3D. From Ramachandran plot, 85.6% of the total residues fell in the favored region, 12.6% of the residues were in the allowed region, 0.4% was found in the generally allowed region while 1.4% of the total residues were found in the outlier region. This result improved the confidence in the modeled structure. We cross-validated with ProSA and the overall quality of modeled TcHK had a Z-score of −7.43. According to Supplementary Fig. 1, the Z-score is within the range found for other experimentally determined proteins of similar size. Verify 3D estimated 80.52% of the residues to have averaged 3D-1D score ⩾0.2 score which corroborated the good quality of the modeled protein. The graphical representations of the quality of TcHK on VERIFY3D and PROSA webservers are represented in Supplementary Fig. 1.
Relative Sequence Analysis and its Effect on P131 Selectivity
The basis of selectivity has been the subject of numerous researches with the intent to underpin the targetability of small-molecule inhibitors of CpIMPDH. For instance, previous studies by Gollapalli et al., Gorla et al., and MacPherson et al., have established that the selectivity of CpIMPDH inhibitors is based on the extension of the inhibitor binding across the subunit interface into a pocket in the adjacent monomer [34, 41, 47, 55]. These findings therefore serve as the premise for this study (Figs. 1 and 2).
Reaction pathway showing the conversion of inosine monophosphate (IMP) to guanine monophosphate (GMP) and adenosine monophosphate (AMP). IMP dehydrogenase (IMPDH) catalyzes the rate-limiting step in the conversion to GMP [98]
Pairwise sequence alignment between CpIMPDH and mIMPDH revealed variations in their amino acid sequences. Zoning in on the co-factor (NAD+) binding regions of both proteins, we further identified variations among residues involved in direct contact or interaction with P131 upon binding. As observed, most variations unique to CpIMPDH occurred at chain D, which were, SER22, VAL24, LEU25, PRO26, SER354, GLY357, TYR358, LYS73 while some occurred at the main chain (chain A), which were ALA165, HIS166, ASN171, LYS200, ILE213, VAL305, MET326, PRO328, GLU329. Corresponding residues in mIMPDH are shown in Fig. 3.
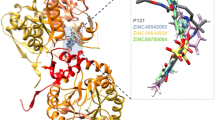
A Superimposed structures of the CpIMPDH and mIMPDH (colored blue and red, respectively). B ClustalW sequence alignment of CpIMPDH and mIMPDH. Red box highlight residues that are identical between both organisms. Binding site amino acid sequences that interacted directly with P131 but where peculiar to only CpIMPDH are highlighted in yellow. The position of nonidentical sequences at both CpIMPDH and mIMPDH are without annotations, while the positions of highly conserved sequences are denoted by (*). Positions of less conserved sequences are denoted by (:) and (.), respectively. These annotations are defined by the ClustalW prediction method (Color figure online)
We further explored the differential binding modes of P131 at the co-factor (NAD+) site of mIMPDH and CpIMPDH as it involved varying residues among both proteins as earlier mentioned. As shown in Fig. 4, using averaged snapshot generated across the simulation period, the molecular interaction profiles of P131-CpIMPDH relative to mIMPDH were dissimilar. These differences in interaction modes could in turn influence the binding affinity and stability of P131 at the NAD+ regions of both proteins.
More specifically, unique CpIMPDH residues such as ALA165, HIS166, GLU329, PRO26, TYR358 (denotes chain D) facilitated strong hydrogen and salt bridge interactions with P131, which were not observed in mIMPDH. These high-affinity and favorable interactions observed in CpIMPDH could explain the preferential binding of P131 over mIMPDH. Also, unfavorable O–O interactions seen in the mIMPDH-P131 complex could further reduce the binding affinity of the compound at the NAD+ site. Thus, this particular sequence variation, with accompanying distinct interactions, could explain the basis of selectivity and favorable binding of P131 toward CpIMPDH. Taken together, these findings also suggest that the interactions of P131 with residues of chain D of the CpIMPDH could be crucial to the selectivity of P131 due to the sequence variation observed on this chain as shown. These finding lend credence to previous reports which posited that interaction of the adenosine subsite with adjacent monomer in prokaryotic IMPDH and the absence of this in eukaryotic IMPDH could underpin basis of selectivity of IMPDH inhibitors [34, 41, 47, 55].
Peculiar Sequence Variation Favors P131 Binding to CpIMPDH
The MM/PBSA method was used to estimate the binding energies (ΔG) of P131 in CpIMPDH, and mIMPDH. As presented in Table 1, P131 had a total ΔG of −37.0 kcal/mol when bound to CpIMPDH and −17.0 kcal/mol in mIMPDH. This notable energy difference could suggest that P131 was bound more favorably to CpIMPDH, which could involve more high-affinity interactions, particularly with residues unique to the NAD+ site of CpIMPDH as earlier indicated in Fig. 4. To quantify the roles of these residues to the binding and stability of P131 in CpIMPDH, we decomposed the overall energy and measured the per-residue contributions compared to mIMPDH.
Per-residue energy plots are presented in Fig. 5, and accordingly, residues with energies > −1 kcal/mol were considered favorable. In CpIMPDH and their respective nonconserved counterparts in mIMPDH, identified residues include CpGLU329 (−3.26 kcal/mol)/mGLN441 (−3.46 kcal/mol), CpALA165 (−2.89 kcal/mol)/mSER276 (−2.63 kcal/mol), CpHIS166 (−1.11 kcal/mol)/mGLN277 (−0.49 kcal/mol). The conserved residual energy contributions in CpIMPDH/mIMPDH are MET302 (−1.82 kcal/mol)/MET414 (−0.6 kcal/mol), GLY303 (−1.63 kcal/mol)/GLY417 (−0.49 kcal/mol). TYR358 (−2.69 kcal/mol) and PRO26 (−1.67 kcal/mol) are present in the chain D of CpIMPDH and have no corresponding residues in mIMPDH as the P131 binding is confined to one chain in the eukaryotes.
A Per-residue energy decomposition analysis of P131 binding site residues in CpIMPDH. B Per-residue energy decomposition analysis of P131 binding site residues in mIMPDH. Both figures represent the energies contributed by the binding sites residues to the total binding free energy of P131 in CpIMPDH and mIMPDH
SER276 and GLN441 were the only residues in mIMPDH that contributed above −1 kcal/mol to the binding of P131. This was different in CpIMPDH where we had seven amino acid residues contributing above −1 kcal/mol to the binding of P131. This disparity could have facilitated the higher binding free energy of P131 in CpIMPDH compared to the weaker binding free energy of P131 in mIMPDH. Interestingly, almost all the residues on chain D of CpIMPDH that interacted with P131 contributed significantly toward the binding, as shown in Fig. 5 (LEU25 = −0.85 kcal/mol, PRO26 = −1.67 kcal/mol, SER354 = −0.7 kcal/mol, and TYR358 = −2.69 kcal/mol). This further suggests they could be critical to P131 selectivity in CpIMPDH.
Peculiar P131 Binding within Parasite IMPDH Active Site Favors Selectivity
The conformational dynamics and orientations of P131 upon binding to either CpIMPDH, or mIMPDH could be crucial in its preferential binding toward CpIMPDH and the resultant therapeutic effect. This is because the varying orientations and the mobility of P131 could somewhat influence the motion of P131 into deep regions of the binding pocket of the enzyme to elicit its therapeutic activity. We, therefore, analyzed the differential binding modes of P131 at the binding site of CpIMPDH over the simulation period, as shown in Fig. 6. The first representative snapshots (P131-mIMPDH and CpIMPDH) were selected at 100 ns, indicative of the starting postequilibrated timeframes. Subsequent snapshots were selected at 220 and 280 ns representing intermediate and final postequilibrated timeframes across both proteins. Likewise, associated binding pocket amino acids that interacted with P131 over the MD simulation were also explored. It was observed that in mIMPDH, a relatively fewer number of residues engaged in interactions with P131 over the simulation period as shown Fig. 6A.
Binding interaction dynamics of P131 at the NAD+ sites of mIMPDH and CpIMPDH. A Time-based analyses of P131 interactions with constituent residues of mIMPDH NAD+ sites at 100, 220, and 280 ns. B Time-based analyses of the interaction of P131 with constituent residues of CpIMPDH NAD+ site at 100, 220, and 280 ns. As shown, binding site residues involved in P131 binding are more in CpIMPDH complex across the simulation timeframes compared to mIMPDH. Legends representing interaction types are also shown
On the contrary, there was a consistently higher number of residues interacting with P131 in CpIMPDH as the simulation proceeded, which could have favored the selective binding of P131 toward CpIMPDH as shown in Fig. 6B. Prominent and recurring residues toward the binding of P131 during the simulation period include ASP163, SER164, ASP252, and MET302. Again, these peculiar residues were unique to only CpIMPDH, suggesting their essential contributions to P131 stability, possibly its eventual selectivity. As shown in Fig. 6B, the peculiar orientations of PI31 and the unique interacting residues likely defined the motion of P131 into the deep regions of CpIMPDH hydrophobic pocket unlike in mIMPDH
The binding of a small-molecule inhibitor can induce a significant conformational change in a protein structure, which could consequently influence its known function [85,86,87,88]. As such, the selective binding of P131 toward CpIMPDH could be affected by peculiar structural dynamics and conformational changes on CpIMPDH. Using known computational techniques, root-mean-square deviation (RMSD), SASA, root-mean-square fluctuation (RMSF), and radius of gyration (RoG), we explore the varying structural dynamics associated with P131 binding to both CpIMPDH and mIMPDH. RMSD calculations allow for the estimation of the structural deviation and stability of an IMPDH structures [89,90,91,92]. From the RMSD plot (Fig. 7A, B), both protein systems had high deviations at starting timeframes of the MD run but attained stability at about 250 ns in both systems. The average RMSD for CpIMPDH-P131 complex and mIMPDH-P131 complex was found to be 2.84 and 2.92 Å, respectively, as shown in Fig. 7. The RMSD plot suggests that the binding of PI31 stabilizes the CpIMPDH structure and leads to a few conformational changes relative to a more unstable structure in the mIMPDH (Fig. 7). The initial jump in average RMSD for both simulated models could be attributed to the initial orientation of P131 in the binding pocket of both CpIMPDH and mIMPDH.
SASA of the bound enzymes were also calculated to ascertain how both enzymes interact with surrounding solvents [93, 94]. SASA gives information about the mobility of the residues and side reorientation as it alternates between the hydrophilic and hydrophobic phases [95]. The repositioning of the side chains from the hydrophilic to the hydrophobic phase signifies protein folding while the converse depicts protein unfolding. The binding of P131 to CpIMPDH increased the SASA values minimally when compared to the apo (Fig. 8). The mean SASA values were 25630 and 24986 Å2. In the mIMPDH model, the binding of p131 did not significantly impact the reorientation and alternation of the protein side chains.
A A 3D surface representation of SASA of the inhibitor binding pocket structure of CpIMPDH upon P131 binding. B A 3D surface representation of SASA of the inhibitor binding pocket structures of mIMPDH. C Comparative solvent accessible surface area of CpIMPDH (blue), and mIMPDH (violet) upon binding of P131 (Color figure online)
We further investigated the local vibrations in both CpIMPDH and mIMPDH upon the binding of P131 by estimating the average residual fluctuations. These were plotted as RMSF as shown in Fig. 9. The RMSF plot highlighted several residual fluctuations in different regions of both CpIMPDH and mIMPDH. Nonetheless, the average fluctuation of mIMPDH in the presence of P131 was relatively higher than the CpIMPDH, suggesting the binding of P131 increased the vibration of individual residues. The lower RMSF in the CpIMPDH models suggests that P131 possibly engaged in stronger and steady interactions, which could have impeded individual residue motions as consistent with the minimal deviations observed in the RMSD calculations. An average RMSF of 8.57 and 38.34 Å was estimated for the CpIMPDH-P131 and mIMPDH-P131 complexes, respectively.
A Comparative RMSF plots of individual residues of bound and unbound conformations of CpIMPDH over the simulation period. B Comparative RMSF plots of individual residues of bound and unbound conformations of mIMPDH over the simulation period. C 3D representation of superimposed post-MD X-ray crystal structures of bound CpIMPDH (red) and mIMPDH (gold) (Color figure online)
To further explore the conformational dynamics of both CpIMPDH and mIMPDH upon binding of P131, we calculated the RoG of each enzyme. RoG provides insights into stability and folding patterns of studied models [96, 97]. Relative to their respective unbound models, the bound CpIMPDH and mIMPDH systems exhibited slightly higher average RoG values throughout the simulation. Here, no conformational shift was observed in the RoG plot as shown in Supplementary Fig. 2, which suggests an insignificant structural deviation or unfolding in both CpIMPDH and mIMPDH upon P131 binding. The apo and complex CpIMPDH exhibited mean RoG values of 27.63 and 28.07 Å, respectively. The average RoG for unbound and bound mIMPDH were also found to be 26.01 and 26.55 Å, respectively. The juxtaposed RoGs of P131-bound CpIMPDH and mIMPDH is represented graphically in Supplementary Fig. 2.
Conclusion
MD simulation, bioinformatics, and advanced post-MD tools were employed in this study to explore the structural basis of P131 selectively toward CpIMPDH rather than mIMPDH as reported by previous in vitro reports, regardless of the substantial structural and sequence similarity of all the organisms. Using MD simulation, Clustal Omega sequence alignment techniques, MM/PBSA analysis of binding free energies, and per-residue decomposition, the structural basis of P131 selectivity was provided. Sequence alignment of the P131 binding sites of both CpIMPDH and mIMPDH revealed variations among constituent amino acids. This allowed the identification of crucial residues within the binding pockets of CpIMPDH that are required binding for P131 binding, however, absent in mIMPDH. Some of the residues though identical in both organisms, it was noted that in mIMPDH, they contributed only little to the binding free energy of P131, suggestive of a weaker binding affinity. Per-residue decomposition plots further consolidated this finding by giving information about the energy contributions of each binding site residue to the affinity and stabilization of P131 within the binding pocket. Amino acid residues such as GLU329 (−3.26 kcal/mol), ALA165 (−2.89 kcal/mol), HIS166A (−1.11 kcal/mol), TYR358 (−2.69 kcal/mol), MET302 (−1.82 kcal/mol), GLY303A (−1.63 kcal/mol), and PRO26 (−1.67 kcal/mol) in the CpIMPDH binding site had higher energy contributions. However, compared to the mIMPDH complex, energy contributions of binding site residues toward P131 binding were relatively lower, consistent with weaker binding. Calculated binding free energy revealed that P131 binds stronger to CpIMPDH than mIMPDH with total binding free energy of −37.00 and −17.01 kcal/mol, respectively. Analysis of the binding modes and orientations of P131 using representative snapshots showed that the orientations of P131 at the binding site of CpIMPDH over the simulation period possibly enhanced its movement into the deep regions of the P131 binding pocket. This was corroborated by the peculiar and higher number of interacting residues observed as the simulation progressed. On the contrary, the movement of P131 appeared to be restricted in mIMPDH binding sites as the simulation progressed, since interacting residues slightly changed and were relatively fewer. Altogether, these findings provide structural insights into the selectivity of P131 toward CpIMPDH. These findings are in line with earlier reports which showed that interaction of the adenosine subsite with adjacent monomer in prokaryotic IMPDH and the absence of this in eukaryotic IMPDH could underpin basis of selectivity and potency of IMPDH inhibitors. Findings could also aid the optimization of P131 with the aim of designing new P131 analogs with improved inhibitory activity and selectivity.
References
Bouzid, M., Kintz, E., & Hunter, P. R. (2018). Risk factors for Cryptosporidium infection in low and middle income countries: a systematic review and meta-analysis. PLoS Neglected Tropical Diseases, 12(6). https://doi.org/10.1371/journal.pntd.0006553.
Fischer Walker, C. L., Aryee, M. J., Boschi-Pinto, C., & Black, R. E. (2012). Estimating diarrhea mortality among young children in low and middle income countries. PLoS ONE. https://doi.org/10.1371/journal.pone.0029151.
Valentiner-Branth, P., Steinsland, H., Fischer, T. K., Perch, M., Scheutz, F., Dias, F., & Sommerfelt, H. (2003). Cohort study of Guinean children: incidence, pathogenicity, conferred protection, and attributable risk for enteropathogens during the first 2 years of life. Journal of Clinical Microbiology, 41(9), 4238–4245. https://doi.org/10.1128/JCM.41.9.4238-4245.2003.
Liu, L., Johnson, H. L., Cousens, S., Perin, J., Scott, S., Lawn, J. E., & Black, R. E. (2012). Global, regional, and national causes of child mortality: an updated systematic analysis for 2010 with time trends since 2000. The Lancet, 379(9832), 2151–2161. https://doi.org/10.1016/S0140-6736(12)60560-1.
Kotloff, K. L., Nataro, J. P., Blackwelder, W. C., Nasrin, D., Farag, T. H., Panchalingam, S., & Levine, M. M. (2013). Burden and aetiology of diarrhoeal disease in infants and young children in developing countries (the Global Enteric Multicenter Study, GEMS): a prospective, case-control study. The Lancet, 382(9888), 209–222. https://doi.org/10.1016/S0140-6736(13)60844-2.
Sow, S. O., Muhsen, K., Nasrin, D., Blackwelder, W. C., Wu, Y., & Farag, T. H. et al. (2016). The burden of cryptosporidium diarrheal disease among children <24 months of age in moderate/high mortality regions of sub-Saharan Africa and South Asia, utilizing data from the global enteric multicenter study (GEMS). PLoS Neglected Tropical Diseases, 10(5), e0004729. https://doi.org/10.1371/journal.pntd.0004729.
Odeniran, P. O., & Ademola, I. O. (2019). Epidemiology of Cryptosporidium infection in different hosts in Nigeria: a meta-analysis. Parasitology International, 71(May), 194–206. https://doi.org/10.1016/j.parint.2019.04.007.
Delahoy, M. J., Omore, R., Ayers, T. L., Schilling, K. A., Blackstock, A. J., Ochieng, J. B., … O’Reilly, C. E. (2018). Clinical, environmental, and behavioral characteristics associated with Cryptosporidium infection among children with moderate-to-severe diarrhea in rural western Kenya, 2008–2012: The Global Enteric Multicenter Study (GEMS). PLoS Neglected Tropical Diseases, 12(7). https://doi.org/10.1371/journal.pntd.0006640.
Umejiego, N. N., Li, C., Riera, T., Hedstrom, L., & Striepen, B. (2004). Cryptosporidium parvum IMP dehydrogenase: identification of functional, structural, and dynamic properties that can be exploited for drug design. Journal of Biological Chemistry, 279(39), 40320–40327. https://doi.org/10.1074/jbc.M407121200.
Putignani, L., & Menichella, D. (2010). Global distribution, public health and clinical impact of the protozoan pathogen cryptosporidium. Interdisciplinary Perspectives on Infectious Diseases, 2010. https://doi.org/10.1155/2010/753512.
O’connor, R. M., Shaffie, R., Kang, G., & Ward, H. D. (2011). Cryptosporidiosis in patients with HIV/AIDS. AIDS, 25(5), 549–60. https://doi.org/10.1097/QAD.0b013e3283437e88.
Chalmers, R. M., & Davies, A. P. (2010). Minireview: clinical cryptosporidiosis. Experimental Parasitology, 124(1), 138–46. https://doi.org/10.1016/j.exppara.2009.02.003.
Kang, G., Sarkar, R., & Desai, N. (2012). Cryptosporidiosis: an under-recognized public health problem. Tropical Parasitology, 2(2), 91 https://doi.org/10.4103/2229-5070.105173.
Mor, S., & Tzipori, S. (2008). Cryptosporidiosis in children in sub-Saharan Africa: A lingering challenge. Clinical Infectious Diseases, 47(7), 915–921. https://doi.org/10.1086/591539.
Aldeyarbi, H. M., Abu El-Ezz, N. M. T., & Karanis, P. (2016). Cryptosporidium and cryptosporidiosis: the African perspective. Environmental Science and Pollution Research, 23(14), 13811–13821. https://doi.org/10.1007/s11356-016-6746-6.
Abdool Karim, S. S., Churchyard, G. J., Karim, Q. A., & Lawn, S. D. (2009). HIV infection and tuberculosis in South Africa: an urgent need to escalate the public health response. Lancet, 374(9693), 921–33. https://doi.org/10.1016/S0140-6736(09)60916-8.
Checkley, W., White Jr, A. C., & Jaganath, D. et al. (2015). A review of the global burden, novel diagnostics, therapeutics, and vaccine targets for cryptosporidium. The Lancet Infectious Diseases, 15(1), 85–94. https://doi.org/10.1016/s1473-3099(14)70772-8.
Mead, J. R. (2002). Cryptosporidiosis and the challenges of chemotherapy. Drug Resistance Updates: Reviews and Commentaries in Antimicrobial and Anticancer Chemotherapy, 5(1), 47–57. http://www.ncbi.nlm.nih.gov/pubmed/12127863.
Lee, S., Harwood, M., Girouard, D., Meyers, M. J., Campbell, M. A., Beamer, G., & Tzipori, S. (2017). The therapeutic efficacy of azithromycin and nitazoxanide in the acute pig model of Cryptosporidium hominis. PLoS ONE, 12(10), e0185906. https://doi.org/10.1371/journal.pone.0185906.
Cabada, M. M., & White, A. C. (2010). Treatment of cryptosporidiosis: do we know what we think we know? Current Opinion in Infectious Diseases, 23(5), 494–9. https://doi.org/10.1097/QCO.0b013e32833de052.
Sparks, H., Nair, G., Castellanos-Gonzalez, A., & White, A. C. (2015). Treatment of Cryptosporidium: what we know, gaps, and the way forward. Current Tropical Medicine Reports, 2(3), 181–187. https://doi.org/10.1007/s40475-015-0056-9.
Fan, X., Upadhyaya, B., Wu, L., Koh, C., Santín-Durán, M., Pittaluga, S., & Jain, A. (2012). CD40 agonist antibody mediated improvement of chronic Cryptosporidium infection in patients with X-linked hyper IgM syndrome. Clinical Immunology, 143(2), 152–161. https://doi.org/10.1016/j.clim.2012.01.014.
Hewitt, R. G., Yiannoutsos, C. T., Higgs, E. S., Carey, J. T., Geiseler, P. J., Soave, R., & Bender, J. F. (2000). Paromomycin: no more effective than placebo for treatment of cryptosporidiosis in patients with advanced human immunodeficiency virus infection. AIDS Clinical Trial Group. Clinical Infectious Diseases, 31(4), 1084–92. https://doi.org/10.1086/318155.
Hussien, S. M. M., Abdella, O. H., Abu-Hashim, A. H., Aboshiesha, G. A., Taha, M. A. A., El-Shemy, A. S., & El-Bader, M. M. (2013). Comparative study between the effect of nitazoxanide and paromomycine in treatment of cryptosporidiosis in hospitalized children. Journal of the Egyptian Society of Parasitology, 43(2), 463–470. http://www.ncbi.nlm.nih.gov/pubmed/24260825.
Allam, A. F., & Shehab, A. Y. (2002). Efficacy of azithromycin, praziquantel and mirazid in treatment of cryptosporidiosis in school children. Journal of the Egyptian Society of Parasitology, 32(3), 969–978. http://www.ncbi.nlm.nih.gov/pubmed/12512828.
Raja, K., Abbas, Z., Hassan, S. M., Luck, N. H., Aziz, T., & Mubarak, M. (2014). Prevalence of cryptosporidiosis in renal transplant recipients presenting with acute diarrhea at a single center in Pakistan. Journal of Nephropathology, 3(4), 127–131. https://doi.org/10.12860/jnp.2014.25.
Holmberg, S. D., Moorman, A. C., Von Bargen, J. C., Palella, F. J., Loveless, M. O., Ward, D. J., & Navin, T. R. (1998). Possible effectiveness of clarithromycin and rifabutin for cryptosporidiosis chemoprophylaxis in HIV disease. HIV Outpatient Study (HOPS) Investigators. JAMA, 279(5), 384–386. https://doi.org/10.1001/jama.279.5.384.
Amenta, M., Dalle Nogare, E. R., Colomba, C., Prestileo, T. S., Di Lorenzo, F., Fundaro, S., & Ferrieri, A. (1999). Intestinal protozoa in HIV-infected patients: effect of rifaximin in Cryptosporidium parvum and Blastocystis hominis infections. Journal of Chemotherapy, 11(5), 391–395. https://doi.org/10.1179/joc.1999.11.5.391.
Gathe, J. C., Mayberry, C., Clemmons, J., & Nemecek, J. (2008). Resolution of severe cryptosporidial diarrhea with rifaximin in patients with AIDS. Journal of Acquired Immune Deficiency Syndromes, 48(3), 363–364. https://doi.org/10.1097/QAI.0b013e31817beb78.
Legrand, F., Grenouillet, F., Larosa, F., Dalle, F., Saas, P., Millon, L., & Rohrlich, P. S. (2011). Diagnosis and treatment of digestive cryptosporidiosis in allogeneic haematopoietic stem cell transplant recipients: a prospective single centre study. Bone Marrow Transplantation, 46(6), 858–862. https://doi.org/10.1038/bmt.2010.200.
Bonatti, H., Barroso, L. F., Sawyer, R. G., Kotton, C. N., & Sifri, C. D. (2012). Cryptosporidium enteritis in solid organ transplant recipients: Multicenter retrospective evaluation of 10 cases reveals an association with elevated tacrolimus concentrations. Transplant Infectious Disease, 14(6), 635–648. https://doi.org/10.1111/j.1399-3062.2012.00719.x.
Krause, I., Amir, J., Cleper, R., Dagan, A., Behor, J., Samra, Z., & Davidovits, M. (2012). Cryptosporidiosis in children following solid organ transplantation. Pediatric Infectious Disease Journal, 31(11), 1135–1138. https://doi.org/10.1097/INF.0b013e31826780f7.
Chen, L., Wilson, D. J., Xu, Y., Aldrich, C. C., Felczak, K., Sham, Y. Y., & Pankiewicz, K. W. (2010). Triazole-linked inhibitors of inosine monophosphate dehydrogenase from human and mycobacterium tuberculosis. Journal of Medicinal Chemistry, 53(12), 4768–4778. https://doi.org/10.1021/jm100424m.
Gollapalli, D. R., MacPherson, I. S., Liechti, G., Gorla, S. K., Goldberg, J. B., & Hedstrom, L. (2010). Structural determinants of inhibitor selectivity in prokaryotic IMP dehydrogenases. Chemistry and Biology, 17(10), 1084–1091. https://doi.org/10.1016/j.chembiol.2010.07.014.
Hedstrom, L., Liechti, G., Goldberg, J. B., & Gollapalli, D. R. (2011). The antibiotic potential of prokaryotic IMP dehydrogenase inhibitors. Current Medicinal Chemistry, 18(13), 1909–18. https://doi.org/10.2174/092986711795590129.
Usha, V., Hobrath, J. V., Gurcha, S. S., Reynolds, R. C., & Besra, G. S. (2012). Identification of novel Mt-Guab2 inhibitor series active against M. tuberculosis. PLoS ONE, 7(3). https://doi.org/10.1371/journal.pone.0033886.
Wang, W., & Hedstrom, L. (1997). Kinetic mechanism of human inosine 5’-monophosphate dehydrogenase type II: random addition of substrates and ordered release of products. Biochemistry, 36(28), 8479–8483. https://doi.org/10.1021/bi970226n.
Zimmermann, A., Gu, J. J., Spychala, J., & Mitchell, B. S. (1996). Inosine monophosphate dehydrogenase expression: transcriptional regulation of the type I and type II genes. In Advances in Enzyme Regulation (Vol. 36, pp. 75–84). Elsevier Ltd. https://doi.org/10.1016/0065-2571(95)00012-7.
Striepen, B., Pruijssers, A. J. P., Huang, J., Li, C., Gubbels, M. J., Umejiego, N. N., & Kissinger, J. C. (2004). Gene transfer in the evolution of parasite nucleotide biosynthesis. Proceedings of the National Academy of Sciences of the United States of America, 101(9), 3154–3159. https://doi.org/10.1073/pnas.0304686101.
Zhang, R. G., Evans, G., Rotella, F. J., Westbrook, E. M., Beno, D., Huberman, E., & Collart, F. R. (1999). Characteristics and crystal structure of bacterial inosine-5’-monophosphate dehydrogenase. Biochemistry, 38(15), 4691–4700. https://doi.org/10.1021/bi982858v.
Kim, Y., Makowska-Grzyska, M., Gorla, S. K., Gollapalli, D. R., Cuny, G. D., Joachimiak, A., & Hedstrom, L. (2015). Structure of Cryptosporidium IMP dehydrogenase bound to an inhibitor with in vivo antiparasitic activity. Acta Crystallographica Section F: Structural Biology Communications, 71, 531–538. https://doi.org/10.1107/S2053230X15000187.
Makowska-Grzyska, M., Kim, Y., Maltseva, N., Osipiuk, J., Gu, M., Zhang, M., & Joachimiak, A. (2015). A novel cofactor-binding mode in bacterial IMP dehydrogenases explains inhibitor selectivity. Journal of Biological Chemistry, 290(9), 5893–5911. https://doi.org/10.1074/jbc.M114.619767.
Hedstrom, L. (2009). IMP dehydrogenase: structure, mechanism, and inhibition. Chemical Reviews, 109(7), 2903–2928. https://doi.org/10.1021/cr900021w.
Felczak, K., Chen, L., Wilson, D., Williams, J., Vince, R., Petrelli, R., & Pankiewicz, K. W. (2011). Cofactor-type inhibitors of inosine monophosphate dehydrogenase via modular approach: Targeting the pyrophosphate binding sub-domain. Bioorganic and Medicinal Chemistry, 19(5), 1594–1605. https://doi.org/10.1016/j.bmc.2011.01.042.
Colby, T. D., Vanderveen, K., Strickler, M. D., Markham, G. D., & Goldstein, B. M. (1999). Crystal structure of human type II inosine monophosphate dehydrogenase: implications for ligand binding and drug design. Proceedings of the National Academy of Sciences of the United States of America, 96(7), 3531–3536. https://doi.org/10.1073/pnas.96.7.3531.
Gorla, S. K., Kavitha, M., Zhang, M., Liu, X., Sharling, L., Gollapalli, D. R., & Cuny, G. D. (2012). Selective and potent urea inhibitors of Cryptosporidium parvum inosine 5′-monophosphate dehydrogenase. Journal of Medicinal Chemistry, 55(17), 7759–7771. https://doi.org/10.1021/jm3007917.
Macpherson, I. S., Kirubakaran, S., Gorla, S. K., Riera, T. V., D’Aquino, J. A., Zhang, M., & Hedstrom, L. (2010). The structural basis of Cryptosporidium-specific IMP dehydrogenase inhibitor selectivity. Journal of the American Chemical Society, 132(4), 1230–1. https://doi.org/10.1021/ja909947a.
Gorla, S. K., McNair, N. N., Yang, G., Gao, S., Hu, M., Jala, V. R., & Hedstrom, L. (2014). Validation of IMP dehydrogenase inhibitors in a mouse model of cryptosporidiosis. Antimicrobial Agents and Chemotherapy, 58(3), 1603–1614. https://doi.org/10.1128/AAC.02075-13.
Johnson, C. R., Gorla, S. K., Kavitha, M., Zhang, M., Liu, X., Striepen, B., & Hedstrom, L. (2013). Phthalazinone inhibitors of inosine-5’-monophosphate dehydrogenase from Cryptosporidium parvum. Bioorganic & Medicinal Chemistry Letters, 23(4), 1004–1007. https://doi.org/10.1016/j.bmcl.2012.12.037.
Kirubakaran, S., Gorla, S. K., Sharling, L., Zhang, M., Liu, X., Ray, S. S., & Cuny, G. D. (2012). Structure-activity relationship study of selective benzimidazole-based inhibitors of Cryptosporidium parvum IMPDH. Bioorganic & Medicinal Chemistry Letters, 22(5), 1985–1988. https://doi.org/10.1016/j.bmcl.2012.01.029.
Maurya, S. K., Gollapalli, D. R., Kirubakaran, S., Zhang, M., Johnson, C. R., Benjamin, N. N., & Cuny, G. D. (2009). Triazole inhibitors of Cryptosporidium parvum inosine 5’-monophosphate dehydrogenase. Journal of Medicinal Chemistry, 52(15), 4623–4630. https://doi.org/10.1021/jm900410u.
Sharling, L., Liu, X., Gollapalli, D. R., Maurya, S. K., Hedstrom, L., & Striepen, B. (2010). A screening pipeline for antiparasitic agents targeting Cryptosporidium inosine monophosphate dehydrogenase. PLoS Neglected Tropical Diseases, 4(8), e794. https://doi.org/10.1371/journal.pntd.0000794.
Umejiego, N. N., Gollapalli, D., Sharling, L., Volftsun, A., Lu, J., Benjamin, N. N., & Hedstrom, L. (2008). Targeting a prokaryotic protein in a eukaryotic pathogen: identification of lead compounds against cryptosporidiosis. Chemistry & Biology, 15(1), 70–77. https://doi.org/10.1016/j.chembiol.2007.12.010.
Sun, Z., Khan, J., Makowska-Grzyska, M., Zhang, M., Cho, J. H., Suebsuwong, C., & Cuny, G. D. (2014). Synthesis, in vitro evaluation and cocrystal structure of 4-oxo-[1]benzopyrano[4,3-c]pyrazole Cryptosporidium parvum inosine 5’-monophosphate dehydrogenase (CpIMPDH) inhibitors. Journal of Medicinal Chemistry, 57(24), 10544–10550. https://doi.org/10.1021/jm501527z.
Gorla, S. K., Kavitha, M., Zhang, M., Chin, J. E. W., Liu, X., Striepen, B., & Cuny, G. D. (2013). Optimization of benzoxazole-based inhibitors of Cryptosporidium parvum inosine 5’-monophosphate dehydrogenase. Journal of Medicinal Chemistry, 56(10), 4028–4043. https://doi.org/10.1021/jm400241j.
Sievers, F., Wilm, A., Dineen, D., Gibson, T. J., Karplus, K., Li, W., … Higgins, D. G. (2011). Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Molecular Systems Biology, 7. https://doi.org/10.1038/msb.2011.75.
Abrahamsen, M. S., Templeton, T. J., Enomoto, S., Abrahante, J. E., Zhu, G., Lancto, C. A., & Kapur, V. (2004). Complete genome sequence of the apicomplexan, Cryptosporidium parvum. Science, 304(5669), 441–445. https://doi.org/10.1126/science.1094786.
Tiedeman, A. A., & Smith, J. M. (1991). Isolation and sequence of a cDNA encoding mouse IMP dehydrogenase. Gene, 97(2), 289–293. https://doi.org/10.1016/0378-1119(91)90065-j.
Waterhouse, A., Bertoni, M., Bienert, S., Studer, G., Tauriello, G., Gumienny, R., & Schwede, T. (2018). SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Research, 46(W1), W296–W303. https://doi.org/10.1093/nar/gky427.
Sintchak, M. D., Fleming, M. A., Futer, O., Raybuck, S. A., Chambers, S. P., Caron, P. R., & Wilson, K. P. (1996). Structure and mechanism of inosine monophosphate dehydrogenase in complex with the immunosuppressant mycophenolic acid. Cell, 85(6), 921–930. https://doi.org/10.1016/S0092-8674(00)81275-1.
Wiederstein, M., & Sippl, M. J. (2007). ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Research, 35(Web Server), W407–W410. https://doi.org/10.1093/nar/gkm290.
Eisenberg, D., Lüthy, R., & Bowie, J. U. (1997). VERIFY3D: assessment of protein models with three-dimensional profiles. Methods in Enzymology, 277, 396–404.
Gopalakrishnan, K., Sowmiya, G., Sheik, S. S., & Sekar, K. (2007). Ramachandran plot on the web (2.0). Protein & Peptide Letters, 14(7), 669–671. https://doi.org/10.2174/092986607781483912.
Pettersen, E. F., Goddard, T. D., Huang, C. C., Couch, G. S., Greenblatt, D. M., Meng, E. C., & Ferrin, T. E. (2004). UCSF Chimera—a visualization system for exploratory research and analysis. Journal of Computational Chemistry, 25(13), 1605–1612. https://doi.org/10.1002/jcc.20084.
Eswar, N., Webb, B., Marti-Renom, M. A., Madhusudhan, M. S., Eramian, D., Shen, M.-Y., & Sali, A. (2006). Comparative protein structure modeling using Modeller. Current Protocols in Bioinformatics, 5, Unit-5.6 https://doi.org/10.1002/0471250953.bi0506s15.
Trott, O., & Olson, A. J. (2010). Software news and update AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization, and multithreading. Journal of Computational Chemistry, 31(2), 455–461. https://doi.org/10.1002/jcc.21334.
Wang, Z., Sun, H., Yao, X., Li, D., Xu, L., Li, Y., & Hou, T. (2016). Comprehensive evaluation of ten docking programs on a diverse set of protein-ligand complexes: the prediction accuracy of sampling power and scoring power. Physical Chemistry Chemical Physics, 18(18), 12964–12975. https://doi.org/10.1039/c6cp01555g.
Machaba, K. E., Mhlongo, N. N., & Soliman, M. E. S. (2018). Induced mutation proves a potential target for TB therapy: a molecular dynamics study on LprG. Cell Biochemistry and Biophysics, 76(3), 345–356. https://doi.org/10.1007/s12013-018-0852-7.
Oguntade, S., Ramharack, P., & Soliman, M. E. (2017). Characterizing the ligand-binding landscape of Zika NS3 helicase-promising lead compounds as potential inhibitors. Future Virology, 12(6), 261–273. https://doi.org/10.2217/fvl-2017-0014.
Olotu, F. A., & Soliman, M. E. S. (2018). From mutational inactivation to aberrant gain-of-function: Unraveling the structural basis of mutant p53 oncogenic transition. Journal of Cellular Biochemistry, 119(3), 2646–2652. https://doi.org/10.1002/jcb.26430.
Agoni, C., Munsamy, G., Ramhrack, P., & Soliman, M. (2020). Human rhinovirus inhibition through capsid “Canyon” perturbation: structural insights into the role of a novel benzothiophene derivative. Cell Biochemistry and Biophysics, 78, 3–13.
Agoni, C., Salifu, E. Y., Munsamy, G., Olotu, F. A., & Soliman, M. (2019). CF3‐pyridinyl substitution on anti‐malarial therapeutics: probing differential ligand binding and dynamical inhibitory effects of a novel triazolopyrimidine‐based inhibitor on Plasmodium falciparum Dihydroorotate dehydrogenase. Chemistry & Biodiversity. https://doi.org/10.1002/cbdv.201900365.
Olotu, F. A., & Soliman, M. E. S. (2019). Dynamic perspectives into the mechanisms of mutation-induced p53-DNA binding loss and inactivation using active perturbation theory: structural and molecular insights toward the design of potent reactivators in cancer therapy. Journal of Cellular Biochemistry, 120(1), 951–966. https://doi.org/10.1002/jcb.27458.
Nair, P. C., & Miners, J. O. (2014). Molecular dynamics simulations: from structure function relationships to drug discovery. In Silico Pharmacology, 2(1). https://doi.org/10.1186/s40203-014-0004-8.
Wang, J., Wolf, R. M., Caldwell, J. W., Kollman, P. A., & Case, D. A. (2004). Development and testing of a general amber force field. Journal of Computational Chemistry, 25(9), 1157–1174. https://doi.org/10.1002/jcc.20035.
Grest, G. S., & Kremer, K. (1986). Molecular dynamics simulation for polymers in the presence of a heat bath. Physical Review A, 33(5), 3628–3631. https://doi.org/10.1103/PhysRevA.33.3628.
Berendsen, H. J. C., Postma, J. P. M., Van Gunsteren, W. F., Dinola, A., & Haak, J. R. (1984). Molecular dynamics with coupling to an external bath. The Journal of Chemical Physics, 81(8), 3684–3690. https://doi.org/10.1063/1.448118.
Ryckaert, J.-P., Ciccotti, G., & Berendsen, H. J. C. (1977). Numerical integration of the cartesian equations of motion of a system with constraints: molecular dynamics of n-alkanes.Journal of Computational Physics, 23(3), 327–341.
Roe, D. R., & Cheatham, T. E. (2013). PTRAJ and CPPTRAJ: software for processing and analysis of molecular dynamics trajectory data. Journal of Chemical Theory and Computation, 9(7), 3084–3095. https://doi.org/10.1021/ct400341p.
Seifert, E. (2014). OriginPro 9.1: scientific data analysis and graphing software-software review. Journal of Chemical Information and Modeling, 54(5), 1552. https://doi.org/10.1021/ci500161d.
Genheden, S., & Ryde, U. (2015). The MM/PBSA and MM/GBSA methods to estimate ligand-binding affinities. Expert Opinion on Drug Discovery, 10(5), 449–461. https://doi.org/10.1517/17460441.2015.1032936.
Hou, T., Wang, J., Li, Y., & Wang, W. (2011). Assessing the performance of the MM/PBSA and MM/GBSA methods. 1. The accuracy of binding free energy calculations based on molecular dynamics simulations. Journal of Chemical Information and Modeling, 51(1), 69–82. https://doi.org/10.1021/ci100275a.
Chaudhary, N., & Aparoy, P. (2017). Deciphering the mechanism behind the varied binding activities of COXIBs through Molecular Dynamic Simulations, MM-PBSA binding energy calculations and per-residue energy decomposition studies. Journal of Biomolecular Structure & Dynamics, 35(4), 868–882. https://doi.org/10.1080/07391102.2016.1165736.
Gupta, A., Chaudhary, N., & Aparoy, P. (2018). MM-PBSA and per-residue decomposition energy studies on 7-Phenyl-imidazoquinolin-4(5H)-one derivatives: identification of crucial site points at microsomal prostaglandin E synthase-1 (mPGES-1) active site. International Journal of Biological Macromolecules, 119, 352–359. https://doi.org/10.1016/j.ijbiomac.2018.07.050.
Beg, A., Khan, F., Lobb, K., Islam, A., Ahmad, F., & Hassan, M. (2019). High throughput screening, docking, and molecular dynamics studies to identify potential inhibitors of human calcium/calmodulin-dependent protein kinase IV. Journal of Biomolecular Structure & Dynamics, 37, 2179–2192.
Dahiya, R., Mohammad, T., Roy, S., Anwar, S., Gupta, P., Haque, A., & Ahmad, F. (2019). Investigation of inhibitory potential of quercetin to the pyruvate dehydrogenase kinase 3: towards implications in anticancer therapy. International Journal of Biological Macromolecules, 136, 1076–1085.
Fatima, S., Mohammad, T., Jairajpuri, D., Rehman, M., Hussain, A., Samim, M., & … Hassan, M. (2019). Identification and evaluation of glutathione conjugate gamma-l-glutamyl-l-cysteine for improved drug delivery to the brain. Journal of Biomolecular Structure & Dynamics, 38(12), 3610–3620.
Gulzar, M., Ali, S., Khan, F., Khan, P., Taneja, P., & Hassan, M. (2019). Binding mechanism of ca eic acid and simvastatin to the integrin linked kinase for therapeutic implications: a comparative docking and MD simulation studies. Journal of Biomolecular Structure & Dynamics, 37, 4327–4337.
Kuzmanic, A., & Zagrovic, B. (2010). Determination of ensemble-average pairwise root mean square deviation from experimental B-factors. Biophysical Journal, 98, 861–871.
Naz, F., Shahbaaz, M., Bisetty, K., Islam, A., Ahmad, F., & Hassan, M. (2015). Designing new kinase inhibitor derivatives as therapeutics against common complex diseases: structural basis of microtubule affinity-regulating kinase 4 (MARK4) inhibition. OMICS, 19, 700–711.
Naz, F., Shahbaaz, M., Khan, S., Bisetty, K., Islam, A., Ahmad, F., & Hassan, M. (2015). PKR-inhibitor binds efficiently with human microtubule affinity-regulating kinase 4. Journal of Molecular Graphics and Modelling, 62, 245–252.
Naz, H., Shahbaaz, M., Haque, M., Bisetty, K., Islam, A., Ahmad, F., & Hassan, M. (2017). Urea-induced denaturation of human calcium/calmodulin-dependent protein kinase IV: a combined spectroscopic and MD simulation studies. Journal of Biomolecular Structure & Dynamics, 35, 463–475.
Menendez, C., Mazola, Y., Guirola, O., Palomares, S., Chinea, G., Hernandez, L., & Musacchio, A. (2015). A comparative molecular dynamics study of thermophilic and mesophilic beta-fructosidase enzymes. Journal of Molecular Modeling, 21, 2772.
Ali, S., Hassan, M., Islam, A., & Ahmad, F. (2014). A review of methods available to estimate solvent-accessible surface areas of soluble proteins in the folded and unfolded states. Current Protein & Peptide Science, 15, 456–476.
Durham, E., Dorr, B., Woetzel, N., Staritzbichler, R., & Meiler, J. (2009). Solvent accessible surface area approximations for rapid and accurate protein structure prediction. Journal of Molecular Modeling, 15(9), 1093–1108. https://doi.org/10.1007/s00894-009-0454-9.
Ali, S., Khan, F., Mohammad, T., Lan, D., Hassan, M., & Wang, Y. (2019). Identification and evaluation of inhibitors of lipase from Malassezia restricta using virtual high-throughput screening and molecular dynamics studies. International Journal of Molecular Sciences, 20, 884.
Khan, F., Shahbaaz, M., Bisetty, K., Waheed, A., Sly, W., Ahmad, F., & Hassan, M. (2016). Large scale analysis of the mutational landscape in beta-glucuronidase: a major player of mucopolysaccharidosis type VII. Gene, 576, 36–44.
Bhagavan, N. V., & Ha, C.-E. (2015). Nucleotide metabolism. In Essentials of medical biochemistry (pp. 465–487). Elsevier. https://doi.org/10.1016/b978-0-12-416687-5.00025-7.
Acknowledgements
Appreciation goes to Center for High Performance Computing, Cape Town, South Africa for providing computational resources.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Omolabi, K.F., Agoni, C., Olotu, F.A. et al. Molecular Basis of P131 Cryptosporidial-IMPDH Selectivity—A Structural, Dynamical and Mechanistic Stance. Cell Biochem Biophys 79, 11–24 (2021). https://doi.org/10.1007/s12013-020-00950-1
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12013-020-00950-1