Abstract
Purpose of Review
The pace of identifying cardiomyopathy-associated mutations and advances in our understanding of sarcomere function that underlies many cardiomyopathies has been remarkable. Here, we aim to synthesize how these advances have led to the promising new treatments that are being developed to treat cardiomyopathies.
Recent Findings
The genomics era has identified and validated many genetic causes of hypertrophic and dilated cardiomyopathies. Recent advances in our mechanistic understanding of sarcomere pathophysiology include high-resolution molecular models of sarcomere components and the identification of the myosin super-relaxed state. The advances in our understanding of sarcomere function have yielded several therapeutic agents that are now in development and clinical use to correct contractile dysfunction–mediated cardiomyopathy.
Summary
New genes linked to cardiomyopathy include targets with limited clinical evidence and require additional investigation. Large portions of cardiomyopathy with family history remain genetically undiagnosed and may be due to polygenic disease.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cardiomyopathy encompasses a group of discrete diseases that result in impaired function of cardiac muscle and can be caused by genetic factors, environmental insults to the heart, or a combination of the two [1,2,3]. Genetic cardiomyopathies have been described since the 1950s, with clinical documentation of clear familial inheritance of a range of phenotypes [4, 5]. Basic and clinical research categorized these diseases based on their morphological and functional phenotypes [6]. The most common cardiomyopathies are hypertrophic (HCM) that occurs in 1:200–1:500 people [1], dilated (DCM) that occurs in 1:500 people [7], and others including arrhythmogenic cardiomyopathy [8], syndromic cardiomyopathies, and emerging classification like atrial myopathy [9].
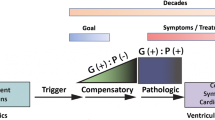
Before the genes that caused cardiomyopathy were identified, cardiomyopathy was a diagnosis of exclusion, defined as pathological myocardial structure and functional changes without an obvious cause (termed idiopathic at the time) [6]. The first genes were identified as causative for HCM and DCM in the 1990s [10,11,12,13,14]. Cardiomyopathy mutations were initially considered to be inherited in a monogenic autosomal dominant manner [5]. However, after the birth of genetic testing of cardiomyopathy patients, it was observed that these mutations showed variable penetrance, with carriers of the same cardiomyopathy-causing mutation showing fulminant disease or asymptomatic presentation [15]. Additionally, cardiomyopathies tend to have a long time of onset (pediatric cardiomyopathies excluded [16]) and are typically identified in the second or third decade of life [1]. Genotype-positive individuals with an asymptomatic phenotype require lifelong vigilance, as the mechanisms that enable the pathological manifestation of mutations are not well understood [17•]. However, the gradual onset of disease also presents a long therapeutic window for the prevention or mitigation of disease [17•].
Mutations in the contractile machinery of the heart (i.e., the sarcomere and myofilaments) cause a majority of familial HCM and account for ~ 30% of familial DCM [17•, 18]. Genomic approaches have yielded a variety of candidate genes in DCM, although the evidence for many of them is still minimal and requires further investigation (Table 1) [17•]. Over the past 30 years, the physiological mechanism for many sarcomere proteins has been established and many pathophysiological consequences linked with cardiomyopathy are well understood (Fig. 1) [18, 19]. This knowledge has recently culminated in the development of small molecule therapeutics that specifically target sarcomere proteins to treat cardiomyopathy [20•, 21••]. In this review, we will cover the recent advances in our understanding of the mechanisms and treatment of cardiomyopathy arising from mutations in sarcomere proteins.
Schematic of the sarcomere proteins linked with cardiomyopathy. A Half sarcomere from Z-disk to M-line with isotropic (I) and anisotropic (A) bands labeled. The C-zone is defined as the ~ 350-nm-wide portion of the myosin thick filament where cMyBP-C is localized in ventricular sarcomeres. The H-zone is the region with no actin overlap and no myosin heads. B Schematic with gene and protein names of the cardiac thin filament with general mechanism of cardiomyopathies listed. C Schematic of the cardiac thick filament. Phosphorylatable regulatory regions are marked with 6-point stars, with yellow stars indicating phosphorylation. Mechanisms of cardiomyopathy-causing mutations listed below the gene names
Classification of Genetic Cardiomyopathies
HCM is a condition where the ventricular walls thicken without obvious cause [1, 5]. HCM was described in 1957 by Dr. Robert Teare from autopsies of eight young patients with asymmetrical hypertrophy of the heart [4], and Dr. Eugene Braunwald provided a comprehensive description of the HCM phenotype in 1964 [5]. HCM is diagnosed via the hemodynamic and morphological nature of the heart [22]. The left ventricle of HCM patients usually has an elevated ejection fraction and a reduced end-systolic volume that can be diagnosed via echocardiogram and exercise/stress testing [22]. HCM patients can be treated with beta-blockers or antihypertensive drugs to alleviate hemodynamic stress [1]. Seventy percent of HCM cases have an intraventricular septal hypertrophy that causes aortic outflow tract obstruction [1]. Surgical resection or septal ablation has been used to restore the ability of blood to exit the ventricle. A majority of HCM patients can live their lives well but they may be limited by progressing disease and the development of heart failure.
In contrast, DCM manifests with thinning and dilation of the ventricle, increased end-diastolic pressure, reduced stroke volume and cardiac output, and dysfunctional filling of the ventricles that often leads to heart failure [23]. DCM has a prevalence of 1:500 [19]. Genetic DCM accounts for approximately 30% of all DCM cases, as DCM is also caused by structural heart disease, valve disorders, hypertension, and other factors [24]. DCM patients are prone to cardiac remodeling and cardiac fibrosis due to the left ventricular dilation. In order to help DCM patients manage their condition, angiotensin-converting enzyme (ACE) inhibitors, beta-blockers, vasodilators, and aldosterone antagonists can be prescribed [25].
Genetic Testing for Cardiomyopathies
When a patient is diagnosed with cardiomyopathy, genetic panel testing is often employed to identify known pathogenic variants in upwards of 100 genes [17•, 26•, 27]. A genetic diagnosis can help inform treatment of the patient and provide information to the extended family regarding who may be at risk of developing disease or passing on a pathogenic mutation [26•, 27]. The first pathogenic mutation linked to HCM was discovered in 1990, with the affected members of two unrelated families sharing the R403Q missense mutation in the MYH7 gene [10]. Currently, 33 genes have been linked to HCM with various levels of evidence, with the 8 most definitive being MYBPC3, MYH7, TNNT2, TNNI3, TPM1, ACTC1, MYL3, and MYL2 (Table 1) [13, 18, 26•, 28,29,30,31,32]. While many mutations have been clearly linked to an autosomal dominant inheritance pattern, the high variability in penetrance of HCM-causing mutations has led to studies that have shown the impact of polygenic modifiers on disease severity [33•].
In the age of genomics research, many variants are initially categorized one way (benign, likely benign, uncertain significance, likely pathogenic, pathogenic), with additional evidence changing the interpretation [26•]. The MYBPC3 c.1224-52G > A intronic splice altering variant occurs in 4% of the South Asian population [34]. This variant is associated with HCM, although it is too prevalent to be a fully penetrant pathogenic mutation. Whole-genome sequencing and clinical phenotyping of a South Asian population revealed minimal association between this variant and HCM features. However, a second MYBPC3 variant, D389V, was found on approximately 10% of the c.1224-52G > A variant alleles and individuals with the D389V variant had significantly more hyperdynamic contractile phenotypes [35]. However, several preclinical models have shown the common c.1224-52G > A to cause sarcomere dysfunction by itself [36]. These findings illustrate the difficulty in understanding the mechanism by which a mutation can impact function.
Genetic DCM is not exclusively caused by mutations in sarcomere genes. Mutations in proteins involved in the cytoskeleton, nuclear envelope, sarcolemma, ion channels, calcium handling, and others are also involved [25, 37]. While the genetic causes of DCM are more varied than of HCM (Table 1), 19–36% of cases are linked to mutations in sarcomere proteins, including mutations in titin that accounts for ~ 20% of familial DCM [38]. The diversity in genetic causes of DCM is remarkable, as is the considerable amount of genetic disease with no known variant associated. In DCM, genetic screening can include over 100 genes but only provides a genetic diagnosis for about 50% of patients [39]. A negative genetic screening can be due to an unknown gene or variant, or a combination of genes that are not pathogenic on their own but combine to ill effect [39]. Variant curation has focused primarily on gene coding regions, while regulatory regions are also capable of contributing to some of the missing genetic linkage for cardiomyopathy [40].
In keeping with the hypothesis that a proportion of DCM is polygenic, patients with DCM-causing mutations in MYH7 have been shown to have significantly more non-synonymous single-nucleotide polymorphisms in approximately 100 cardiomyopathy-related genes than MYH7 mutation carriers with HCM [41•]. This study provides evidence that cumulative, normally benign changes in proteins linked to contractile regulation may influence the severity of disease elicited from a primary pathogenic mutation [41•]. Polygenic modifiers of disease may also account for some of the variable penetrance of HCM and DCM mutations [42•]. There are genes that have enriched expression in the atria including MYL4, MYH6, and MYBPHL that may alter atrial function and exacerbate a mildly pathogenic mutation expressed in ventricular cardiomyocytes [43]. Myosin light chain 4 (MYL4) frameshift mutations have been demonstrated to impair the actomyosin cross-bridge formation and cause a specific atrial cardiomyopathy and atrial remodeling [44].
Sarcomere Proteins in Hypertrophic Cardiomyopathy
β-Myosin Heavy Chain (MYH7)
MYH7 encodes β-myosin heavy chain (β-MHC), the primary myosin motor in the ventricular sarcomere [45]. β-MHC contains an N-terminal motor domain that binds to actin and hydrolyzes ATP to power the cross-bridge cycle and force generation [46]. Mutations can affect the hydrolytic cycle of ATP or the myosin neck region that allows the myosin heads to swing into the interfilament space and interact with actin [47]. MYH7 mutations are more frequent in the motor domain, particularly in the “myosin mesa” region that is critical for actin binding [46, 48]. The molecular biophysical consequences of MYH7 mutations are varied, including increased sarcomere force development, higher ATPase activity, increased actomyosin binding, decrease affinity of the myosin interacting head state, and faster cross-bridge cycling (Table 2) [46, 47, 49•]. Many of these processes are linked, and a single MYH7 mutation can cause a variety of these biophysical changes, but ultimately cause a hypercontractile left ventricle that pathologically hypertrophies [49•, 50•].
There have been several recent advances regarding the so-called super-relaxed state of myosin. The myosin head domain exists in several conformations when not bound to actin, including a disordered relaxed state that resides in the interfilament space, ready to bind to actin [50•, 51, 52]. The super-relaxed state involves the folding back on myosin heads onto the thick filament backbone. Several avenues of research have converged on this mechanism, including cryo-electron microscopy of the myosin head on the thick filament backbone [53], X-ray diffraction data [54••] showing the movement of myosin heads towards the thick filament in association with the low ATPase state, and molecular mechanisms of modulating the super-relaxed state [49•, 51]. Importantly, it has been shown that some MYH7 HCM-causing mutations destabilize the formation of the super-relaxed state and increase the number of disordered relaxed heads ready to generate cross-bridges, therefore increasing ensemble contractility [50•, 51].
Myosin Light Chains (MYL2, MYL3)
Sarcomere myosin is a heteromeric complex consisting of a dimer of myosin heavy chains with a coiled-coil tail integrating into the thick filament and two myosin motor domains in the interfilament space, linked with a flexible “neck” region [55]. Each myosin head is associated with two light chains. In the ventricle, these are the ventricular regulatory (RLC, MYL2) and essential (ELC, MYL3) myosin light chains. The light chains modulate the strength and speed of the myosin heavy chain motor’s ability to generate force [56]. The ELCs bind to the alpha helix of the myosin heavy chain neck and participate in cross-bridge formation [57]. The ventricular RLCs contain calcium-binding EF hand motifs and are phosphorylated [55, 58].
Mutations in the ventricular myosin light chains have been associated with hypertrophic and restrictive cardiomyopathies [30, 59]. The mechanisms for this include altered flexibility of the myosin neck [14] and altered localization of the myosin heads in the interfilament space [57]. The HCM-related MYL2 A57G mutation increases RLC phosphorylation levels and increases the percent of myosin heads in the disordered relaxed state, whereas the restrictive cardiomyopathy-associated MYL2 E143K mutation reduced RLC phosphorylation and promoted the super-relaxed state of myosin [60•]. The ability of the myosin light chains to regulate the myosin super-relaxed and their ability to be phosphorylated and dephosphorylated makes them attractive targets for drug development.
Cardiac Myosin Binding Protein-C (MYBPC3)
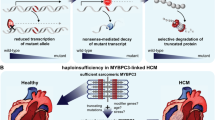
The first MYBPC3 mutations linked to HCM were identified in 1995 [11, 12]. In the following years, it was established that approximately 70% of MYBPC3 mutations associated with HCM were nonsense mutations that truncated portions of the Cʹ-terminal domains of cMyBP-C [61]. MyBP-C is comprised of a linear series of immunoglobulin and fibronectin-like domains, with most of the penultimate Cʹ-terminal domain required for sarcomere incorporation [36, 62]. MYBPC3 truncating mutations cause haploinsufficiency, expressing an insufficient amount of cMyBP-C to maintain normal cross-bridge regulation that results in hypercontractile sarcomeres and HCM [63,64,65,66]. Additional evidence suggests that the truncated allele causes cellular stress by compromising the protein quality control system [67, 68•]. While nonsense mutations in MYBPC3 all result in a similar impairment, the pathogenic effect of MYBPC3 missense mutations depends on the residues affected. These include altering myosin binding affinity [69], regulation of the super-relaxed state [70], ATP hydrolysis [71••], and actin interaction [72], which lead to hypercontractile sarcomeres.
Cardiac MyBP-C can be phosphorylated by a variety of kinase pathways and its function shifts from inhibiting cross-bridge cycling to activating cross-bridge cycling when phosphorylated [73]. Dephosphorylated cMyBP-C promotes the formation of the super-relaxed state whereas phosphorylated cMyBP-C primes myosin heads for cross-bridge formation by promoting the myosin disordered relaxed state [70, 71••]. The myosin S2 region is bound by cMyBP-C to mediate this function, and mutations in MYH7 that alter residues in this region reduce the ability of cMyBP-C to bind to and regulate β-MHC [64]. These interactions between β-MHC and cMyBP-C provide a coherent picture of how both MYH7 and MYBPC3 mutations can converge on a similar HCM phenotype (Table 2).
Thin Filament HCM Mutations
Sarcomere thin filament function consists of a chain of events starting with calcium binding to troponin-C (cTnC, TNNC1) causing a conformational change through troponin T (cTnT, TNNT2) that allows alpha-tropomyosin (α-TM, TPM1) to move, exposing myosin binding sites on actin (ACTC1), allowing cross-bridge formation. This chain of events is regulated by troponin-I (cTnI, TNNI3) and depends on many key residues and protein regions to correctly translate calcium binding to cross-bridge formation. Mutations in thin filament proteins that cause HCM are typically missense mutations that cause an increase in thin filament calcium sensitivity, allowing force development at relatively lower calcium concentrations and resulting in hypercontractility (Table 2) [74,75,76].
The biophysical interaction of the linker between cTnT and α-TM is a major regulatory region and mutations that alter the rigidity or position of the linker dysregulate activation of the thin filaments [77]. Each troponin/tropomyosin regulatory complex spans ~ 38 nm, with neighboring complexes exhibiting cooperativity of activation that is translated through the overlap region of neighboring α-TM molecules [78]. These thin filament mutations also result in hypercontractility due to enhanced cooperativity [79].
Another consequence of HCM-causing thin filament mutations is an increased propensity for arrhythmias due to alteration of the calcium buffering properties of the thin filament. Mutations in the troponin complex can prolong sarcomere calcium release in diastole and promote arrhythmia [75, 80•]. β-Adrenergic signaling causes phosphorylation of cTnI, desensitizing the thin filament to calcium, and increasing the rate of relaxation; thin filament mutations that affect this regulatory effect cause cardiomyopathy [76, 81].
Sarcomere Proteins in Dilated Cardiomyopathy
Titin (TTN)
Titin is a sarcomere protein that interacts at its N-terminus with telethonin (TCAP) and α-actinin in the Z-disk, and then spans nearly 1 micron across a half sarcomere where its C-terminal interacts with myomesin in M-line [82, 83]. Titin is the largest protein, comprised of approximately 17,000–26,000 amino acids depending on isoform and what exons are included in various transcripts. Titin contains a molecular spring region in the I-band that can include an N2B region, common in the ventricle, or the N2B and N2A regions, which are common in the atria. The longer N2B and N2A isoform (N2BA) produces a longer titin molecule that exhibits longer resting sarcomere lengths and reduced sarcomere compliance [84].
DCM has been clearly linked with titin-truncating variants. However, as titin variants were rapidly identified in the early 2010s, an alarming number of titin-truncating variants were identified, enough that it would be impossible for titin truncation alone to cause disease [38]. This mystery was solved by the discovery that titin undergoes high levels of exon exclusion, primarily in exons that encode the I-band of titin, and truncating mutations in these frequently excluded exons are not included in a sufficient amount of transcripts to cause disease [85]. Bona-fide titin-truncating variants are often found in the A-band domains and are generally considered to be pathogenic due to a haploinsufficiency, although fragments of truncated titin have recently been identified, which may cause dysfunction [86, 87].
Titin’s A-band region contains several super-domains composed of 7–11 immunoglobulin-like and fibronectin-like domains that are repeated across the A-band [83]. These repeats provide a molecular ruler that sets some of the thick filament’s periodic protein localization patterns [88]. Because of the staggering size and repetitive nature of A-band titin and our limited understanding of how these domains regulate binding of sarcomere components, missense variants in this region are both plentiful and difficult to interpret [89•]. Understanding the pathogenicity of titin missense mutations will be a challenging new frontier for sarcomere protein research.
β-Myosin Heavy Chain (MYH7)
MYH7 mutations that cause HCM result in hypercontractility, whereas MYH7 mutations that occur in some of the same functional locations cause loss of function and result in DCM. DCM mutations in MYH7 are spread throughout the molecule but are slightly enriched in residues associated with the nucleotide binding pocket on the myosin head [52, 90]. Mechanistically, DCM mutations in MYH7 cause defects in the cross-bridge ATPase cycle [91] resulting in depressed function and hypocontractile outcome [92].
Thin Filaments
HCM-causing thin filament mutations result in calcium sensitization whereas DCM mutations exhibit the opposite effect, with decreased sensitivity of thin filament activation from calcium [93•]. This includes mutations in TNNC1, TNNT2, TNNI3, TPM1, and ACTC1 [78, 93•, 94]. Interestingly, mutations in these genes cause decreased in calcium sensitivity that is uncoupled from the normal phosphorylation-dependent calcium desensitization mediated from cTnI (i.e., the mutations cause structural changes that result in cTnI to adopt its calcium desensitizing interactions) [76, 94]. Cooperativity of thin filament activation is also oppositely impacted, with DCM mutations in TPM1 that compact the α-TM overlap regions resulting in decreased cooperativity [78].
Novel Pharmaceutical Treatments for Cardiomyopathy
The advances in our understanding of the molecular and biophysical mechanisms of disease in sarcomere proteins have led to the rapid advancement of treatments to target these mechanisms. In the early 2010s, several companies set out to develop small molecules that could selectively modulate sarcomere function. Some companies have targeted the hypercontractile mechanism of disease in HCM [101]. A first-in-class compound, originally known as MYK-461, or mavacamten, showed successful reduction in the obstructive phenotype of HCM by reducing myosin contractility without serious adverse effects [21••]. After these promising phase III trial results, mavacamten received FDA approval in April of 2022 for treatment of obstructive HCM, with further trials ongoing. Aficamten is another myosin inhibitor that has shown positive results in phase II clinical trials [102].
Many HCM-causing mutations in MYH7 have been shown to disrupt the super-relaxed state, and HCM-causing MYBPC3-truncating mutations prevent cMyBP-C from promoting the myosin super-relaxed state [50•, 51, 52, 70]. The super-relaxed state can be stabilized or disrupted to sequester or liberate populations of myosin heads to meet the contractile demand [50•, 60•, 103]. This provides a common mechanism of action for mutations in these two proteins. Mavacamten reduces contractility by stabilizing the super-relaxed state, making mavacamten and other small molecules that target this pathway ideal candidates to address a common mechanism of disease [21••, 103, 104].
Myosin activators have also been developed with the goal of increasing contractility. One compound, omecamtiv mecarbil (OM), was designed to increase cardiomyocyte contractility in patients with heart failure with reduced ejection fraction [20•, 105]. OM did not meet its endpoint for improving contractility for heart failure patients in phase III trials [20•] but did significantly improve function for individuals with the lowest contractile function [106•]. While OM failed to gain FDA approval in December of 2022, other myosin activators, such as danicamtiv, are also in development for treating patients with heart failure with reduced ejection fraction [107].
Treating Cardiomyopathy with Genetic Medicine
As titin-truncating variants comprise ~ 20% of genetic DCM, efforts have been made to identify therapeutic approaches to correct these mutations [108•]. Recent advances in exon skipping therapies and CRISPR-dependent reading frame repair, especially in Duchene muscular dystrophy [109], may be applicable to correct TTN-truncating variant expression. Romano et al. observed pathogenic allele dose-dependent decreases in full-length titin protein isoforms for A- and I-band TTN-truncating variants, showing TTN haploinsufficiency. Cardiomyocyte genome editing by SpCas9 restored the TTN protein reading frame, increasing full-length TTN protein levels, and diminishing TTN truncating peptides [108•]. Exon skipping approaches use antisense oligonucleotides to facilitate skipping of one or more exons that contain a mutation and have been used to repair a frameshift mutation in TTN exon 326 given (Ser14450fsX4) [110]. Skipping of TTN exon 326 improved myofibril assembly and stability and ameliorated the DCM phenotype.
Variations on single base editing approaches have also recently been used by two groups to correct a mutation associated with HCM [111, 112]. Adenine base editing (ABE) uses a catalytically dead CRISPR-Cas9, a guide RNA, and a deaminase to convert an adenine to a guanine [113, 114]. The dominant-negative MYH7 R403Q causes HCM via increasing cardiac contractility and is caused by a 1208G > A mutation. These simultaneous publications both showed effective A > G nucleotide editing that restored the MYH7 wild-type sequence, with minimal bystander and off-target editing [111, 112]. These approaches used adeno-associated virus to deliver the editing constructs, and the clinical utility of this approach and base editing are still being evaluated.
Conclusions
Over 30 years have passed since the identification of the first genetic causes of HCM and DCM. In this time, we have acquired a vast, albeit incomplete, understanding of the genes that drive these diseases. Biophysical mechanisms and contractile regulation have matured into therapeutic targets, although successfully translating this wealth of knowledge to the development of therapeutics will require extensive research.
While gene therapy holds great promise for correcting disease-causing mutations directly, the implementation and scalability of these approaches remain to be established. Even the most perfect gene therapy approach would be limited by our identification of all the pathogenic and modifying genes responsible for disease. Therefore, identifying “missing” genetic causes of cardiomyopathy and establishing the role of polygenic factors in the development of disease are crucial.
Basic science research has also delineated the mechanisms responsible for many of the proteins and mutations linked to cardiomyopathy. As demonstrated with therapeutically targeting the myosin super-relaxed state, targeting sarcomere protein interactions can be effective for modulating contractility to counter pathological processes. Additionally, signaling pathways like phosphorylation of ventricular myosin light chains to alter the myosin super-relaxed state provide additional upstream targets that could be leveraged for therapeutic development.
References
Papers of particular interest, published recently, have been highlighted as: • Of importance •• Of major importance
Maron BJ, Desai MY, Nishimura RA, Spirito P, Rakowski H, Towbin JA, et al. Management of hypertrophic cardiomyopathy: JACC state-of-the-art review. J Am Coll Cardiol. 2022;79(4):390–414. https://doi.org/10.1016/j.jacc.2021.11.021.
Pinto YM, Elliott PM, Arbustini E, Adler Y, Anastasakis A, Bohm M, et al. Proposal for a revised definition of dilated cardiomyopathy, hypokinetic non-dilated cardiomyopathy, and its implications for clinical practice: a position statement of the ESC working group on myocardial and pericardial diseases. Eur Heart J. 2016;37(23):1850–8. https://doi.org/10.1093/eurheartj/ehv727.
McNally EM, Barefield DY, Puckelwartz MJ. The genetic landscape of cardiomyopathy and its role in heart failure. Cell Metab. 2015;21(2):174–82. https://doi.org/10.1016/j.cmet.2015.01.013.
Teare D. Asymmetrical hypertrophy of the heart in young adults. Br Heart J. 1958;20(1):1–8. https://doi.org/10.1136/hrt.20.1.1.
Braunwald E, Lambrew CT, Rockoff SD, Ross J Jr, Morrow AG. Idiopathic hypertrophic subaortic stenosis. I A description of the disease based upon an analysis of 64 patients. Circulation. 1964;30(SUPPL 4):3–119. https://doi.org/10.1161/01.cir.29.5s4.iv-3.
Codd MB, Sugrue DD, Gersh BJ, Melton LJ III. Epidemiology of idiopathic dilated and hypertrophic cardiomyopathy. A population-based study in Olmsted County, Minnesota, 1975–1984. Circulation. 1989;80(3):564–72. https://doi.org/10.1161/01.cir.80.3.564.
Pourebrahim K, Marian JG, Tan Y, Chang JT, Marian AJ. A combinatorial oligogenic basis for the phenotypic plasticity between late-onset dilated and arrhythmogenic cardiomyopathy in a single family. J Cardiovasc Aging. 2021;1. https://doi.org/10.20517/jca.2021.15.
Chen K, Rao M, Guo G, Chen X, Chen L, Song J. Sarcomere variants in arrhythmogenic cardiomyopathy: pathogenic factor or bystander? Gene. 2019;687:82–9. https://doi.org/10.1016/j.gene.2018.10.080.
Rivner H, Mitrani RD, Goldberger JJ. Atrial myopathy underlying atrial fibrillation. Arrhythm Electrophysiol Rev. 2020;9(2):61–70. https://doi.org/10.15420/aer.2020.13.
Geisterfer-Lowrance AA, Kass S, Tanigawa G, Vosberg HP, McKenna W, Seidman CE, et al. A molecular basis for familial hypertrophic cardiomyopathy: a beta cardiac myosin heavy chain gene missense mutation. Cell. 1990;62(5):999–1006. https://doi.org/10.1016/0092-8674(90)90274-i.
Bonne G, Carrier L, Bercovici J, Cruaud C, Richard P, Hainque B, et al. Cardiac myosin binding protein-C gene splice acceptor site mutation is associated with familial hypertrophic cardiomyopathy. Nat Genet. 1995;11(4):438–40. https://doi.org/10.1038/ng1295-438.
Watkins H, Conner D, Thierfelder L, Jarcho JA, MacRae C, McKenna WJ, et al. Mutations in the cardiac myosin binding protein-C gene on chromosome 11 cause familial hypertrophic cardiomyopathy. Nat Genet. 1995;11(4):434–7. https://doi.org/10.1038/ng1295-434.
Thierfelder L, Watkins H, MacRae C, Lamas R, McKenna W, Vosberg HP, et al. Alpha-tropomyosin and cardiac troponin T mutations cause familial hypertrophic cardiomyopathy: a disease of the sarcomere. Cell. 1994;77(5):701–12. https://doi.org/10.1016/0092-8674(94)90054-x.
Poetter K, Jiang H, Hassanzadeh S, Master SR, Chang A, Dalakas MC, et al. Mutations in either the essential or regulatory light chains of myosin are associated with a rare myopathy in human heart and skeletal muscle. Nat Genet. 1996;13(1):63–9. https://doi.org/10.1038/ng0596-63.
Lorenzini M, Norrish G, Field E, Ochoa JP, Cicerchia M, Akhtar MM, et al. Penetrance of hypertrophic cardiomyopathy in sarcomere protein mutation carriers. J Am Coll Cardiol. 2020;76(5):550–9. https://doi.org/10.1016/j.jacc.2020.06.011.
Ware SM, Wilkinson JD, Tariq M, Schubert JA, Sridhar A, Colan SD, et al. Genetic causes of cardiomyopathy in children: first results from the Pediatric Cardiomyopathy Genes Study. J Am Heart Assoc. 2021;10(9):e017731. https://doi.org/10.1161/JAHA.120.017731.
• Mazzarotto F, Tayal U, Buchan RJ, Midwinter W, Wilk A, Whiffin N, et al. Reevaluating the genetic contribution of monogenic dilated cardiomyopathy. Circulation. 2020;141(5):387–98. https://doi.org/10.1161/CIRCULATIONAHA.119.037661. This study used cardiomyopathy panel screening on 2538 DCM patients and 912 healthy volunteers and myriads of individuals in a data repository to assess rare variants in 56 commonly tested cardiomyopathy-associated genes. Mutations in TTN, DSP, MYH7, LMNA, BAG3, TNNT2, TNNC1, PLN, ACTC1, NEXN, TPM1, and VCL genes were highly linked with DCM. Other genes may still play a role; they need to be more thoroughly studied, and the authors emphasized the minimal diagnostic utility of other genes in DCM.
Ho CY, Day SM, Ashley EA, Michels M, Pereira AC, Jacoby D, et al. Genotype and lifetime burden of disease in hypertrophic cardiomyopathy: insights from the Sarcomeric Human Cardiomyopathy Registry (SHaRe). Circulation. 2018;138(14):1387–98. https://doi.org/10.1161/CIRCULATIONAHA.117.033200.
Hershberger RE, Hedges DJ, Morales A. Dilated cardiomyopathy: the complexity of a diverse genetic architecture. Nat Rev Cardiol. 2013;10(9):531–47. https://doi.org/10.1038/nrcardio.2013.105.
• Teerlink JR, Diaz R, Felker GM, McMurray JJV, Metra M, Solomon SD, et al. Cardiac myosin activation with omecamtiv mecarbil in systolic heart failure. N Engl J Med. 2021;384(2):105–16. https://doi.org/10.1056/NEJMoa2025797. Findings on this study suggest that the novel myosin activator omecamtiv mecarbil treatment reduced heart failure–associated outcomes. While omecamtiv mecarbil ultimately was not FDA approved, this study provides support for the concept of targeting myosin function to treat heart failure.
•• Olivotto I, Oreziak A, Barriales-Villa R, Abraham TP, Masri A, Garcia-Pavia P, et al. Mavacamten for treatment of symptomatic obstructive hypertrophic cardiomyopathy (EXPLORER-HCM): a randomised, double-blind, placebo-controlled, phase 3 trial. Lancet. 2020;396(10253):759–69. https://doi.org/10.1016/S0140-6736(20)31792-X. Mavacamten is a first-in-class myosin inhibitor for the treatment of HCM. This article reports the results of its phase 3 trial where mavacamten was found to increase exercise capacity, reduce left ventricular outflow tract gradients, and improve heart function in patients with obstructive hypertrophic cardiomyopathy. This is a landmark study for treating cardiac function with myosin modulators.
Marian AJ, Braunwald E. Hypertrophic cardiomyopathy: genetics, pathogenesis, clinical manifestations, diagnosis, and therapy. Circ Res. 2017;121(7):749–70. https://doi.org/10.1161/CIRCRESAHA.117.311059.
Rosenbaum AN, Agre KE, Pereira NL. Genetics of dilated cardiomyopathy: practical implications for heart failure management. Nat Rev Cardiol. 2020;17(5):286–97. https://doi.org/10.1038/s41569-019-0284-0.
Bozkurt B, Colvin M, Cook J, Cooper LT, Deswal A, Fonarow GC, et al. Current diagnostic and treatment strategies for specific dilated cardiomyopathies: a scientific statement from the American Heart Association. Circulation. 2016;134(23):e579–646. https://doi.org/10.1161/CIR.0000000000000455.
Mestroni L, Brun F, Spezzacatene A, Sinagra G, Taylor MR. Genetic causes of dilated cardiomyopathy. Prog Pediatr Cardiol. 2014;37(1–2):13–8. https://doi.org/10.1016/j.ppedcard.2014.10.003.
• Ingles J, Goldstein J, Thaxton C, Caleshu C, Corty EW, Crowley SB, et al. Evaluating the clinical validity of hypertrophic cardiomyopathy genes. Circ Genom Precis Med. 2019;12(2):e002460. https://doi.org/10.1161/CIRCGEN.119.002460. This study investigates the clinical validity of genes that are associated with the development of HCM. Of 33 HCM-causing genes, eight were considered definitive and three had moderate evidence linking them to HCM. The authors urge caution on using mutations in these genes to guide clinical decision-making until further inquiry into the remaining 20 genes provides definitive links with disease.
Dellefave-Castillo LM, Cirino AL, Callis TE, Esplin ED, Garcia J, Hatchell KE, et al. Assessment of the diagnostic yield of combined cardiomyopathy and arrhythmia genetic testing. JAMA Cardiol. 2022;7(9):966–74. https://doi.org/10.1001/jamacardio.2022.2455.
Kimura A, Harada H, Park JE, Nishi H, Satoh M, Takahashi M, et al. Mutations in the cardiac troponin I gene associated with hypertrophic cardiomyopathy. Nat Genet. 1997;16(4):379–82. https://doi.org/10.1038/ng0897-379.
Huang W, Liang J, Yuan CC, Kazmierczak K, Zhou Z, Morales A, et al. Novel familial dilated cardiomyopathy mutation in MYL2 affects the structure and function of myosin regulatory light chain. FEBS J. 2015;282(12):2379–93. https://doi.org/10.1111/febs.13286.
Lee W, Hwang TH, Kimura A, Park SW, Satoh M, Nishi H, et al. Different expressivity of a ventricular essential myosin light chain gene Ala57Gly mutation in familial hypertrophic cardiomyopathy. Am Heart J. 2001;141(2):184–9. https://doi.org/10.1067/mhj.2001.112487.
Prabhakar R, Boivin GP, Grupp IL, Hoit B, Arteaga G, Solaro RJ, et al. A familial hypertrophic cardiomyopathy alpha-tropomyosin mutation causes severe cardiac hypertrophy and death in mice. J Mol Cell Cardiol. 2001;33(10):1815–28. https://doi.org/10.1006/jmcc.2001.1445.
Mogensen J, Klausen IC, Pedersen AK, Egeblad H, Bross P, Kruse TA, et al. Alpha-cardiac actin is a novel disease gene in familial hypertrophic cardiomyopathy. J Clin Invest. 1999;103(10):R39-43. https://doi.org/10.1172/JCI6460.
• Harper AR, Goel A, Grace C, Thomson KL, Petersen SE, Xu X, et al. Common genetic variants and modifiable risk factors underpin hypertrophic cardiomyopathy susceptibility and expressivity. Nat Genet. 2021;53(2):135–42. https://doi.org/10.1038/s41588-020-00764-0. This article reports a genome-wide association study of 2780 HCM cases and 47,486 control that identified 12 loci associated with increased susceptibility to HCM. They found that common single-nucleotide polymorphisms in these regions correlated with increased risk of developing HCM. This provides support for the concept that a portion of HCM has polygenic modifiers.
Dhandapany PS, Sadayappan S, Xue Y, Powell GT, Rani DS, Nallari P, et al. A common MYBPC3 (cardiac myosin binding protein C) variant associated with cardiomyopathies in South Asia. Nat Genet. 2009;41(2):187–91. https://doi.org/10.1038/ng.309.
Viswanathan SK, Puckelwartz MJ, Mehta A, Ramachandra CJA, Jagadeesan A, Fritsche-Danielson R, et al. Association of cardiomyopathy with MYBPC3 D389V and MYBPC3Delta25bp intronic deletion in South Asian descendants. JAMA Cardiol. 2018;3(6):481–8. https://doi.org/10.1001/jamacardio.2018.0618.
Kuster DW, Govindan S, Springer TI, Martin JL, Finley NL, Sadayappan S. A hypertrophic cardiomyopathy-associated MYBPC3 mutation common in populations of South Asian descent causes contractile dysfunction. J Biol Chem. 2015;290(9):5855–67. https://doi.org/10.1074/jbc.M114.607911.
Weisberg A, Winegrad S. Alteration of myosin cross bridges by phosphorylation of myosin-binding protein C in cardiac muscle. Proc Natl Acad Sci U S A. 1996;93(17):8999–9003. https://doi.org/10.1073/pnas.93.17.8999.
Herman DS, Lam L, Taylor MR, Wang L, Teekakirikul P, Christodoulou D, et al. Truncations of titin causing dilated cardiomyopathy. N Engl J Med. 2012;366(7):619–28. https://doi.org/10.1056/NEJMoa1110186.
Pugh TJ, Kelly MA, Gowrisankar S, Hynes E, Seidman MA, Baxter SM, et al. The landscape of genetic variation in dilated cardiomyopathy as surveyed by clinical DNA sequencing. Genet Med. 2014;16(8):601–8. https://doi.org/10.1038/gim.2013.204.
Gacita AM, Fullenkamp DE, Ohiri J, Pottinger T, Puckelwartz MJ, Nobrega MA, et al. Genetic variation in enhancers modifies cardiomyopathy gene expression and progression. Circulation. 2021;143(13):1302–16. https://doi.org/10.1161/CIRCULATIONAHA.120.050432.
• Puckelwartz MJ, Pesce LL, Dellefave-Castillo LM, Wheeler MT, Pottinger TD, Robinson AC, et al. Genomic context differs between human dilated cardiomyopathy and hypertrophic cardiomyopathy. J Am Heart Assoc. 2021;10(7):e019944. https://doi.org/10.1161/JAHA.120.019944. This study used whole-genome sequencing on a cohort of patients with HCM or DCM to evaluate the occurrence of non-synonymous single-nucleotide polymorphisms in cardiomyopathy-associated genes. They found that the increasing numbers of these variants correlated with the likelihood of having DCM compared to HCM. The authors emphasize the importance of the genetic landscape that may alter the development of cardiomyopathy.
• Tadros R, Francis C, Xu X, Vermeer AMC, Harper AR, Huurman R, et al. Shared genetic pathways contribute to risk of hypertrophic and dilated cardiomyopathies with opposite directions of effect. Nat Genet. 2021;53(2):128–34. https://doi.org/10.1038/s41588-020-00762-2. This work reports a genome-wide association study that identifies several loci in HCM and DCM. The loci that were associated with ventricular function showed opposite functional changes in HCM and DCM, supporting the hypothesis that HCM is a hypercontractile disease. The loci were used to create a polygenic risk score that helped explain the genetic basis for patients with variable phenotypes in HCM and DCM.
Pensa AV, Baman JR, Puckelwartz MJ, Wilcox JE. Genetically based atrial fibrillation: current considerations for diagnosis and management. J Cardiovasc Electrophysiol. 2022;33(8):1944–53. https://doi.org/10.1111/jce.15446.
Gudbjartsson DF, Holm H, Sulem P, Masson G, Oddsson A, Magnusson OT, et al. A frameshift deletion in the sarcomere gene MYL4 causes early-onset familial atrial fibrillation. Eur Heart J. 2017;38(1):27–34. https://doi.org/10.1093/eurheartj/ehw379.
Reiser PJ, Portman MA, Ning XH, Schomisch MC. Human cardiac myosin heavy chain isoforms in fetal and failing adult atria and ventricles. Am J Physiol Heart Circ Physiol. 2001;280(4):H1814–20. https://doi.org/10.1152/ajpheart.2001.280.4.H1814.
Nag S, Trivedi DV, Sarkar SS, Adhikari AS, Sunitha MS, Sutton S, et al. The myosin mesa and the basis of hypercontractility caused by hypertrophic cardiomyopathy mutations. Nat Struct Mol Biol. 2017;24(6):525–33. https://doi.org/10.1038/nsmb.3408.
Adhikari AS, Kooiker KB, Sarkar SS, Liu C, Bernstein D, Spudich JA, et al. Early-onset hypertrophic cardiomyopathy mutations significantly increase the velocity, force, and actin-activated ATPase activity of human beta-cardiac myosin. Cell Rep. 2016;17(11):2857–64. https://doi.org/10.1016/j.celrep.2016.11.040.
Yousaf M, Khan WA, Shahzad K, Khan HN, Ali B, Hussain M, et al. Genetic association of beta-myosin heavy-chain gene (MYH7) with cardiac dysfunction. Genes (Basel). 2022. https://doi.org/10.3390/genes13091554.
• Adhikari AS, Trivedi DV, Sarkar SS, Song D, Kooiker KB, Bernstein D, et al. beta-Cardiac myosin hypertrophic cardiomyopathy mutations release sequestered heads and increase enzymatic activity. Nat Commun. 2019;10(1):2685. https://doi.org/10.1038/s41467-019-10555-9. This study evaluated four HCM-causing mutations in b-cardiac myosin. They found that these mutants in the head and neck region of myosin caused release of the myosin heads from their low-energy-associated interacting head motif confirmation. This resulted in hypercontractility and increase ATPase activity and supports the hypothesis that HCM is caused by hypercontractile myosin motors.
• Toepfer CN, Garfinkel AC, Venturini G, Wakimoto H, Repetti G, Alamo L, et al. Myosin sequestration regulates sarcomere function, cardiomyocyte energetics, and metabolism, informing the pathogenesis of hypertrophic cardiomyopathy. Circulation. 2020;141(10):828–42. https://doi.org/10.1161/CIRCULATIONAHA.119.042339. This study showed that HCM-causing myosin heavy chain mutations disrupted the interacting head motif and this also reduced the myosin super-relaxed state, ultimately increasing contractile function arrhythmias. It was also demonstrated that small molecule myosin inhibitor mavacamten stabilized the super-relaxed state and mitigated contractile dysfunction.
McNamara JW, Li A, Dos Remedios CG, Cooke R. The role of super-relaxed myosin in skeletal and cardiac muscle. Biophys Rev. 2015;7(1):5–14. https://doi.org/10.1007/s12551-014-0151-5.
Alamo L, Ware JS, Pinto A, Gillilan RE, Seidman JG, Seidman CE, et al. Effects of myosin variants on interacting-heads motif explain distinct hypertrophic and dilated cardiomyopathy phenotypes. Elife. 2017. https://doi.org/10.7554/eLife.24634.
Hu Z, Taylor DW, Edwards RJ, Taylor KA. Coupling between myosin head conformation and the thick filament backbone structure. J Struct Biol. 2017;200(3):334–42. https://doi.org/10.1016/j.jsb.2017.09.009.
•• Ma W, Henze M, Anderson RL, Gong H, Wong FL, Del Rio CL, et al. The super-relaxed state and length dependent activation in porcine myocardium. Circ Res. 2021;129(6):617–30. https://doi.org/10.1161/CIRCRESAHA.120.318647. This study used X-ray diffraction to evaluate the state of myosin heavy chain heads during length-dependent activation. They found that with stretch, myosin heads were liberated from the super-relaxed state along the thick filament backbone. The myosin inhibitor mavacamten is known to stabilize the super-relaxed state, and this work shows that mavacamten also blunts the stretch activation of myosin heads.
Sheikh F, Lyon RC, Chen J. Functions of myosin light chain-2 (MYL2) in cardiac muscle and disease. Gene. 2015;569(1):14–20. https://doi.org/10.1016/j.gene.2015.06.027.
Lowey S, Waller GS, Trybus KM. Skeletal muscle myosin light chains are essential for physiological speeds of shortening. Nature. 1993;365(6445):454–6. https://doi.org/10.1038/365454a0.
Muthu P, Wang L, Yuan CC, Kazmierczak K, Huang W, Hernandez OM, et al. Structural and functional aspects of the myosin essential light chain in cardiac muscle contraction. FASEB J. 2011;25(12):4394–405. https://doi.org/10.1096/fj.11-191973.
Chang AN, Mahajan P, Knapp S, Barton H, Sweeney HL, Kamm KE, et al. Cardiac myosin light chain is phosphorylated by Ca2+/calmodulin-dependent and -independent kinase activities. Proc Natl Acad Sci U S A. 2016;113(27):E3824–33. https://doi.org/10.1073/pnas.1600633113.
De Bortoli M, Vio R, Basso C, Calore M, Minervini G, Angelini A, et al. Novel missense variant in MYL2 gene associated with hypertrophic cardiomyopathy showing high incidence of restrictive physiology. Circ Genom Precis Med. 2020;13(2):e002824. https://doi.org/10.1161/CIRCGEN.119.002824.
• Yuan CC, Kazmierczak K, Liang J, Ma W, Irving TC, Szczesna-Cordary D. Molecular basis of force-pCa relation in MYL2 cardiomyopathy mice: role of the super-relaxed state of myosin. Proc Natl Acad Sci U S A. 2022;119(8). https://doi.org/10.1073/pnas.2110328119. This study analyzed mutations in the ventricular myosin regulatory light chain (MYL2) mutation that lead to cardiomyopathy. The HCM-associated mutation increased Ca2+ sensitivity of isometric force, disrupted the super-relaxed state of myosin, and moved myosin heads off the thick filament backbone. These data support the ability of mutations in myosin’s resident regulatory proteins to work through a similar mechanism as myosin mutations in the development of cardiomyopathy.
Carrier L. Targeting the population for gene therapy with MYBPC3. J Mol Cell Cardiol. 2021;150:101–8. https://doi.org/10.1016/j.yjmcc.2020.10.003.
Yu B, French JA, Carrier L, Jeremy RW, McTaggart DR, Nicholson MR, et al. Molecular pathology of familial hypertrophic cardiomyopathy caused by mutations in the cardiac myosin binding protein C gene. J Med Genet. 1998;35(3):205–10. https://doi.org/10.1136/jmg.35.3.205.
Marston S, Copeland O, Gehmlich K, Schlossarek S, Carrier L. How do MYBPC3 mutations cause hypertrophic cardiomyopathy? J Muscle Res Cell Motil. 2012;33(1):75–80. https://doi.org/10.1007/s10974-011-9268-3.
Barefield D, Kumar M, Gorham J, Seidman JG, Seidman CE, de Tombe PP, et al. Haploinsufficiency of MYBPC3 exacerbates the development of hypertrophic cardiomyopathy in heterozygous mice. J Mol Cell Cardiol. 2015;79:234–43. https://doi.org/10.1016/j.yjmcc.2014.11.018.
Suay-Corredera C, Pricolo MR, Herrero-Galan E, Velazquez-Carreras D, Sanchez-Ortiz D, Garcia-Giustiniani D, et al. Protein haploinsufficiency drivers identify MYBPC3 variants that cause hypertrophic cardiomyopathy. J Biol Chem. 2021;297(1):100854. https://doi.org/10.1016/j.jbc.2021.100854.
O’Leary TS, Snyder J, Sadayappan S, Day SM, Previs MJ. MYBPC3 truncation mutations enhance actomyosin contractile mechanics in human hypertrophic cardiomyopathy. J Mol Cell Cardiol. 2019;127:165–73. https://doi.org/10.1016/j.yjmcc.2018.12.003.
Bahrudin U, Morisaki H, Morisaki T, Ninomiya H, Higaki K, Nanba E, et al. Ubiquitin-proteasome system impairment caused by a missense cardiac myosin-binding protein C mutation and associated with cardiac dysfunction in hypertrophic cardiomyopathy. J Mol Biol. 2008;384(4):896–907. https://doi.org/10.1016/j.jmb.2008.09.070.
• Helms AS, Tang VT, O’Leary TS, Friedline S, Wauchope M, Arora A, et al. Effects of MYBPC3 loss-of-function mutations preceding hypertrophic cardiomyopathy. JCI Insight. 2020;5(2). https://doi.org/10.1172/jci.insight.133782. This study evaluated cMyBP-C levels in iPSC-CMs with MYBPC3-truncating mutations. It showed a compensation in MYBPC3 protein levels, with reduced rates of both synthesis and degradation. The authors conclude with the potential to regulate the degradation rate of cMyBP-C in cases with reduced levels of cMyBP-C.
Helms AS, Thompson AD, Glazier AA, Hafeez N, Kabani S, Rodriguez J, et al. Spatial and functional distribution of MYBPC3 pathogenic variants and clinical outcomes in patients with hypertrophic cardiomyopathy. Circ Genom Precis Med. 2020;13(5):396–405. https://doi.org/10.1161/CIRCGEN.120.002929.
McNamara JW, Li A, Lal S, Bos JM, Harris SP, van der Velden J, et al. MYBPC3 mutations are associated with a reduced super-relaxed state in patients with hypertrophic cardiomyopathy. PLoS ONE. 2017;12(6):e0180064. https://doi.org/10.1371/journal.pone.0180064.
•• Toepfer CN, Wakimoto H, Garfinkel AC, McDonough B, Liao D, Jiang J, et al. Hypertrophic cardiomyopathy mutations in MYBPC3 dysregulate myosin. Sci Transl Med. 2019;11(476). https://doi.org/10.1126/scitranslmed.aat1199. This study found that reduced levels of cMyBP-C lead to a reduced percentage of myosin heads in the super-relaxed state and an increase in contractility. DCM-causing missense mutations in MYBPC3 had the opposite effect. Importantly, the myosin inhibitor drug mavacamten stabilized the super-relaxed state and returned contractile function to normal levels. This provides evidence that HCM-causing mutations in MYBPC3 can be treated effectively with mavacamten.
Da’as SI, Fakhro K, Thanassoulas A, Krishnamoorthy N, Saleh A, Calver BL, et al. Hypertrophic cardiomyopathy-linked variants of cardiac myosin-binding protein C3 display altered molecular properties and actin interaction. Biochem J. 2018;475(24):3933–48. https://doi.org/10.1042/BCJ20180685.
Barefield D, Sadayappan S. Phosphorylation and function of cardiac myosin binding protein-C in health and disease. J Mol Cell Cardiol. 2010;48(5):866–75. https://doi.org/10.1016/j.yjmcc.2009.11.014.
Karibe A, Tobacman LS, Strand J, Butters C, Back N, Bachinski LL, et al. Hypertrophic cardiomyopathy caused by a novel alpha-tropomyosin mutation (V95A) is associated with mild cardiac phenotype, abnormal calcium binding to troponin, abnormal myosin cycling, and poor prognosis. Circulation. 2001;103(1):65–71. https://doi.org/10.1161/01.cir.103.1.65.
Shafaattalab S, Li AY, Gunawan MG, Kim B, Jayousi F, Maaref Y, et al. Mechanisms of arrhythmogenicity of hypertrophic cardiomyopathy-associated troponin T (TNNT2) variant I79N. Front Cell Dev Biol. 2021;9:787581. https://doi.org/10.3389/fcell.2021.787581.
Gomes AV, Harada K, Potter JD. A mutation in the N-terminus of troponin I that is associated with hypertrophic cardiomyopathy affects the Ca(2+)-sensitivity, phosphorylation kinetics and proteolytic susceptibility of troponin. J Mol Cell Cardiol. 2005;39(5):754–65. https://doi.org/10.1016/j.yjmcc.2005.05.013.
Abdullah S, Lynn ML, McConnell MT, Klass MM, Baldo AP, Schwartz SD, et al. FRET-based analysis of the cardiac troponin T linker region reveals the structural basis of the hypertrophic cardiomyopathy-causing Delta160E mutation. J Biol Chem. 2019;294(40):14634–47. https://doi.org/10.1074/jbc.RA118.005098.
McConnell M, Tal Grinspan L, Williams MR, Lynn ML, Schwartz BA, Fass OZ, et al. Clinically divergent mutation effects on the structure and function of the human cardiac tropomyosin overlap. Biochemistry. 2017;56(26):3403–13. https://doi.org/10.1021/acs.biochem.7b00266.
Bai F, Caster HM, Dawson JF, Kawai M. The immediate effect of HCM causing actin mutants E99K and A230V on actin-Tm-myosin interaction in thin-filament reconstituted myocardium. J Mol Cell Cardiol. 2015;79:123–32. https://doi.org/10.1016/j.yjmcc.2014.10.014.
• Pioner JM, Vitale G, Gentile F, Scellini B, Piroddi N, Cerbai E, et al. Genotype-driven pathogenesis of atrial fibrillation in hypertrophic cardiomyopathy: the case of different TNNT2 mutations. Front Physiol. 2022;13:864547. https://doi.org/10.3389/fphys.2022.864547. This study analyzed the pathological mechanism between two TNNT2 mutations that differentially cause atrial or ventricular dysfunction. R92Q showed an increase in left atrial dilation but no changes in the E163R mutation. The atria from R92Q mice showed an increased propensity for arrhythmia. The findings of this paper illustrate the importance of assessing atrial function in mutations that cause cardiomyopathy.
Tardiff JC, Hewett TE, Palmer BM, Olsson C, Factor SM, Moore RL, et al. Cardiac troponin T mutations result in allele-specific phenotypes in a mouse model for hypertrophic cardiomyopathy. J Clin Invest. 1999;104(4):469–81. https://doi.org/10.1172/JCI6067.
Kruger M, Linke WA. Titin-based mechanical signalling in normal and failing myocardium. J Mol Cell Cardiol. 2009;46(4):490–8. https://doi.org/10.1016/j.yjmcc.2009.01.004.
Granzier HL, Labeit S. The giant protein titin: a major player in myocardial mechanics, signaling, and disease. Circ Res. 2004;94(3):284–95. https://doi.org/10.1161/01.RES.0000117769.88862.F8.
Bennett P, Rees M, Gautel M. The axial alignment of titin on the muscle thick filament supports its role as a molecular ruler. J Mol Biol. 2020;432(17):4815–29. https://doi.org/10.1016/j.jmb.2020.06.025.
Roberts AM, Ware JS, Herman DS, Schafer S, Baksi J, Bick AG, et al. Integrated allelic, transcriptional, and phenomic dissection of the cardiac effects of titin truncations in health and disease. Sci Transl Med. 2015;7(270):270ra6. https://doi.org/10.1126/scitranslmed.3010134.
McAfee Q, Chen CY, Yang Y, Caporizzo MA, Morley M, Babu A, et al. Truncated titin proteins in dilated cardiomyopathy. Sci Transl Med. 2021;13(618):eabd7287. https://doi.org/10.1126/scitranslmed.abd7287.
Hinson JT, Chopra A, Nafissi N, Polacheck WJ, Benson CC, Swist S, et al. HEART DISEASE. Titin mutations in iPS cells define sarcomere insufficiency as a cause of dilated cardiomyopathy. Science. 2015;349(6251):982–6. https://doi.org/10.1126/science.aaa5458.
Tonino P, Kiss B, Strom J, Methawasin M, Smith JE 3rd, Kolb J, et al. The giant protein titin regulates the length of the striated muscle thick filament. Nat Commun. 2017;8(1):1041. https://doi.org/10.1038/s41467-017-01144-9.
• Akinrinade O, Helio T, Lekanne Deprez RH, Jongbloed JDH, Boven LG, van den Berg MP, et al. Relevance of titin missense and non-frameshifting insertions/deletions variants in dilated cardiomyopathy. Sci Rep. 2019;9(1):4093. https://doi.org/10.1038/s41598-019-39911-x. This study investigated non-truncating mutations in titin, which are less well understood compared to titin-truncating variants. While they found rare in-frame mutations in titin, they showed that rare variations in these regions were not more abundant in DCM populations compared to control populations. The authors conclude that many of these non-truncating variants should be classified as likely benign.
van der Linde IHM, Hiemstra YL, Bokenkamp R, van Mil AM, Breuning MH, Ruivenkamp C, et al. A Dutch MYH7 founder mutation, p.(Asn1918Lys), is associated with early onset cardiomyopathy and congenital heart defects. Neth Heart J. 2017;25(12):675–81. https://doi.org/10.1007/s12471-017-1037-5.
Ujfalusi Z, Vera CD, Mijailovich SM, Svicevic M, Yu EC, Kawana M, et al. Dilated cardiomyopathy myosin mutants have reduced force-generating capacity. J Biol Chem. 2018;293(23):9017–29. https://doi.org/10.1074/jbc.RA118.001938.
Schmitt JP, Debold EP, Ahmad F, Armstrong A, Frederico A, Conner DA, et al. Cardiac myosin missense mutations cause dilated cardiomyopathy in mouse models and depress molecular motor function. Proc Natl Acad Sci U S A. 2006;103(39):14525–30. https://doi.org/10.1073/pnas.0606383103.
• Robinson P, Sparrow AJ, Patel S, Malinowska M, Reilly SN, Zhang YH, et al. Dilated cardiomyopathy mutations in thin-filament regulatory proteins reduce contractility, suppress systolic Ca(2+), and activate NFAT and Akt signaling. Am J Physiol Heart Circ Physiol. 2020;319(2):H306–19. https://doi.org/10.1152/ajpheart.00272.2020. The study analyzed DCM-associated mutations in cardiac troponin-T, troponin-I, and a-tropomyosin. They found a shared mechanisms of reduced calcium release, reduction in SR Ca2+2+ ATPase activity, and increase in sodium-calcium exchanger activity.
Memo M, Leung MC, Ward DG, dos Remedios C, Morimoto S, Zhang L, et al. Familial dilated cardiomyopathy mutations uncouple troponin I phosphorylation from changes in myofibrillar Ca(2)(+) sensitivity. Cardiovasc Res. 2013;99(1):65–73. https://doi.org/10.1093/cvr/cvt071.
Sewanan LR, Park J, Rynkiewicz MJ, Racca AW, Papoutsidakis N, Schwan J, et al. Loss of crossbridge inhibition drives pathological cardiac hypertrophy in patients harboring the TPM1 E192K mutation. J Gen Physiol. 2021. https://doi.org/10.1085/jgp.202012640.
• Schwabe FV, Peter EK, Taft MH, Manstein DJ. Assessment of the contribution of a thermodynamic and mechanical destabilization of myosin-binding protein C domain C2 to the pathomechanism of hypertrophic cardiomyopathy-causing double mutation MYBPC3(Delta25bp/D389V). Int J Mol Sci. 2021;22(21). https://doi.org/10.3390/ijms222111949. This study analyzed a recently identified double mutation in MYBPC3 that includes a D389V variant on the highly prevalent delta25bp allele. The findings suggest that the D389V variant may promote the accumulation of misfolded cMyBP-C early in life, followed by a haploinsufficiency later due to the delta25 mutation.
Veltri T, Landim-Vieira M, Parvatiyar MS, Gonzalez-Martinez D, Dieseldorff Jones KM, Michell CA, et al. Hypertrophic cardiomyopathy cardiac troponin C mutations differentially affect slow skeletal and cardiac muscle regulation. Front Physiol. 2017;8:221. https://doi.org/10.3389/fphys.2017.00221.
Schuldt M, Johnston JR, He H, Huurman R, Pei J, Harakalova M, et al. Mutation location of HCM-causing troponin T mutations defines the degree of myofilament dysfunction in human cardiomyocytes. J Mol Cell Cardiol. 2021;150:77–90. https://doi.org/10.1016/j.yjmcc.2020.10.006.
• Singh RR, McNamara JW, Sadayappan S. Mutations in myosin S2 alter cardiac myosin-binding protein-C interaction in hypertrophic cardiomyopathy in a phosphorylation-dependent manner. J Biol Chem. 2021;297(1):100836. https://doi.org/10.1016/j.jbc.2021.100836. This study showed that three HCM-causing myosin heavy chain mutations augment the binding of myosin S2 region with cMyBP-C in HCM. Phosphorylation of cMyBP-C further increased these interactions. Individually, and together with cMyBP-C phosphorylation, these mutants augment the development of force.
• Fomin A, Gartner A, Cyganek L, Tiburcy M, Tuleta I, Wellers L, et al. Truncated titin proteins and titin haploinsufficiency are targets for functional recovery in human cardiomyopathy due to TTN mutations. Sci Transl Med. 2021;13(618):eabd3079. https://doi.org/10.1126/scitranslmed.abd3079. Titin-truncating mutations are highly linked to DCM, although the presence of a truncated product is not known to cause disease. The authors express titin variants with truncations in various regions of the titin molecule in human-induced pluripotent stem cell–derived cardiomyocytes. Impaired contractility due to these mutations was rescued by either fixing the mutation or increasing WT titin levels with proteosome inhibitors.
Green EM, Wakimoto H, Anderson RL, Evanchik MJ, Gorham JM, Harrison BC, et al. A small-molecule inhibitor of sarcomere contractility suppresses hypertrophic cardiomyopathy in mice. Science. 2016;351(6273):617–21. https://doi.org/10.1126/science.aad3456.
Chuang C, Collibee S, Ashcraft L, Wang W, Vander Wal M, Wang X, et al. Discovery of aficamten (CK-274), a next-generation cardiac myosin inhibitor for the treatment of hypertrophic cardiomyopathy. J Med Chem. 2021;64(19):14142–52. https://doi.org/10.1021/acs.jmedchem.1c01290.
Anderson RL, Trivedi DV, Sarkar SS, Henze M, Ma W, Gong H, et al. Deciphering the super relaxed state of human beta-cardiac myosin and the mode of action of mavacamten from myosin molecules to muscle fibers. Proc Natl Acad Sci U S A. 2018;115(35):E8143–52. https://doi.org/10.1073/pnas.1809540115.
Rohde JA, Roopnarine O, Thomas DD, Muretta JM. Mavacamten stabilizes an autoinhibited state of two-headed cardiac myosin. Proc Natl Acad Sci U S A. 2018;115(32):E7486–94. https://doi.org/10.1073/pnas.1720342115.
Cleland JG, Teerlink JR, Senior R, Nifontov EM, Mc Murray JJ, Lang CC, et al. The effects of the cardiac myosin activator, omecamtiv mecarbil, on cardiac function in systolic heart failure: a double-blind, placebo-controlled, crossover, dose-ranging phase 2 trial. Lancet. 2011;378(9792):676–83. https://doi.org/10.1016/S0140-6736(11)61126-4.
• Felker GM, Solomon SD, Claggett B, Diaz R, McMurray JJV, Metra M, et al. Assessment of omecamtiv mecarbil for the treatment of patients with severe heart failure: a post hoc analysis of data from the GALACTIC-HF randomized clinical trial. JAMA Cardiol. 2022;7(1):26–34. https://doi.org/10.1001/jamacardio.2021.4027. This article reported results from the phase three clinical trial of omecamtiv mecarbil for treatment of heart failure. Prior clinical studies show omecamtiv mecarbil did show effective improvements in heart failure. This study identified that a subset of patients with the lowest levels of cardiac function derived significant benefit from omecamtiv mecarbil.
Voors AA, Tamby JF, Cleland JG, Koren M, Forgosh LB, Gupta D, et al. Effects of danicamtiv, a novel cardiac myosin activator, in heart failure with reduced ejection fraction: experimental data and clinical results from a phase 2a trial. Eur J Heart Fail. 2020;22(9):1649–58. https://doi.org/10.1002/ejhf.1933.
• Romano R, Ghahremani S, Zimmerman T, Legere N, Thakar K, Ladha FA, et al. Reading frame repair of TTN truncation variants restores titin quantity and functions. Circulation. 2022;145(3):194–205. https://doi.org/10.1161/CIRCULATIONAHA.120.049997. This study explored TTN-truncating variants and proposed a treatment based on repairing the TTN reading frame to restore TTN protein levels. This technique decreased the TTN-truncating variant levels and increased TTN full-length amount.
Scoto M, Finkel R, Mercuri E, Muntoni F. Genetic therapies for inherited neuromuscular disorders. Lancet Child Adolesc Health. 2018;2(8):600–9. https://doi.org/10.1016/S2352-4642(18)30140-8.
Gramlich M, Pane LS, Zhou Q, Chen Z, Murgia M, Schotterl S, et al. Antisense-mediated exon skipping: a therapeutic strategy for titin-based dilated cardiomyopathy. EMBO Mol Med. 2015;7(5):562–76. https://doi.org/10.15252/emmm.201505047.
Reichart D, Newby GA, Wakimoto H, Lun M, Gorham JM, Curran JJ, et al. Efficient in vivo genome editing prevents hypertrophic cardiomyopathy in mice. Nat Med. 2023;29(2):412–21. https://doi.org/10.1038/s41591-022-02190-7.
Chai AC, Cui M, Chemello F, Li H, Chen K, Tan W, et al. Base editing correction of hypertrophic cardiomyopathy in human cardiomyocytes and humanized mice. Nat Med. 2023;29(2):401–11. https://doi.org/10.1038/s41591-022-02176-5.
Richter MF, Zhao KT, Eton E, Lapinaite A, Newby GA, Thuronyi BW, et al. Phage-assisted evolution of an adenine base editor with improved Cas domain compatibility and activity. Nat Biotechnol. 2020;38(7):883–91. https://doi.org/10.1038/s41587-020-0453-z.
Gaudelli NM, Komor AC, Rees HA, Packer MS, Badran AH, Bryson DI, et al. Programmable base editing of A*T to G*C in genomic DNA without DNA cleavage. Nature. 2017;551(7681):464–71. https://doi.org/10.1038/nature24644.
Funding
This work has been supported by NIH grants NHLBI R00141698 and R56165137 (DYB) and Postdoctoral Fellowship 23POST1023125 (AAA).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Human and Animal Rights and Informed Consent
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Barefield, D.Y., Alvarez-Arce, A. & Araujo, K.N. Mechanisms of Sarcomere Protein Mutation-Induced Cardiomyopathies. Curr Cardiol Rep 25, 473–484 (2023). https://doi.org/10.1007/s11886-023-01876-9
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11886-023-01876-9