Abstract
We prompted to characterize a wastewater bacterium, Pseudoxanthomonas mexicana GTZY, that efficiently transforms toxic mercury and arsenic, explores its bioremediation capability, and reveals their relevant gene resistance operons. The isolated strain was characterized by its phylogenetic, biochemical, and phenotypic properties. The strain GTZY potentially removed 84.3% of mercury and their mercury volatilization (Hg(II) to Hg(0)) was confirmed using the X-ray film method, and its respective merA gene was PCR amplified. In addition, strain GTZY efficiently removed arsenate (68.5%) and arsenite (63.2%), and showed resistance up to > 175 and > 55 mM, respectively. Their genomic annotations disclosed the linkage of Tn2-transposon and int1 in both ends of mer operon (merAPTR). The co-existence of arsP and arsH proteins in its intrinsic ars operon (arsCPRH) was extremely diverse from its ancestral species. We believe that the mercury resistance-conferring mer operon of P. mexicana GTZY presumably derived horizontally from other species in the reactor, while the arsenic resistance-conferring intrinsic ars operon was highly diversified and evolved from its ancestral species. By considering the potential of the strain GTZY to transform heavy metals, this can be used to recover contaminated sites.
Graphical abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Heavy metals are inorganic ill-defined chemical compounds that are ubiquitous in nature. Some heavy metals such as cadmium (Cd), arsenic (As), nickel (Ni), lead (Pb), chromium (Cr), mercury (Hg), copper (Cu), and zinc (Zn) mainly persist in the environment due to the core release of anthropical sources (Loganathan et al. 2015; Govarthanan et al. 2016b, a, 2018, 2019; Jobby et al. 2018; Briffa et al. 2020; Khalid et al. 2020). Metal persistence is considered an emerging situation due to its potential toxicity in the entire ecosystem, including humans, animals, and plants. In humans, it evokes conditions such as neurological and cardiovascular diseases, chronic respiratory disorder, enlarged liver and spleen, cancer, nervous system damage, and hypophosphatemia (Olmedo et al. 2013; Rebelo and Caldas 2016; Luo et al. 2020; Qin et al. 2021). Some heavy metals are considered highly toxic as they could cause human illness even at their relatively low levels. Among those, arsenic and mercury are the prime considerations for their chemical toxicity, persistence, frequency, and potential for human exposure (Hou et al. 2020). According to the national priority list released by the Agency for Toxic Substances and Disease Registry (ATSDR 2018), among the 275 toxic substances, arsenic and mercury were ranked first and third, respectively (Watters and Rayman 2014). The main challenge and limitation posed by these heavy metals is their non-biodegradability nature; therefore, these can not be catabolized by indigenous microorganisms. Alternatively, these heavy metals can be transformed from toxic to non-toxic forms by their nuclear structural changes via microbial activities (Wuana and Okieimen 2011; Jebeli et al. 2017; Wang et al. 2020), but it is narrated only to specific degrading microbes. Another challenge posed by mercury and arsenic contamination is their molecular alteration into toxic methyl mercury (MeHg) and methyl arsenite (MeAs), respectively, which leads to bioaccumulation and biomagnification in the aquatic ecosystem (Christakis et al. 2021; Alabssawy and Hashem 2024). Hence, remediating these heavy metals from the polluted sites is very important. Conventional methods such as chemical treatment, filtration, incineration, excavation, and soil washing are expensive and laborious (Bala et al. 2022). Meanwhile, bioremediation is an eco-friendly natural process to clean up polluted environments by microbes that transform toxic chemical compounds to safe levels (Jeyakumar et al. 2022).
In general, mercury bioremediation mainly occurs by the mechanism of reduction or volatilization, which is governed by mer operon in microorganisms (Boyd and Barkay 2012; Raj and Maiti 2019). In mer operon, merA protein is one of the main constituents along with mercury-binding (merP), regulatory (merR and D), and mercury-transporting proteins (merT and E), and is mainly located in the bacterial chromosome or mobile genetic elements (e.g., plasmids, integrons, or transposons) (Oregaard and Sørensen 2007; Boyd and Barkay 2012; Møller et al. 2014; Naguib et al. 2018). Relying on the availability of toxic mercury (Hg2), merR protein binds specifically into the promoter-operator region and it is followed by regulating the transcription of structural genes. As a result, it produces a key enzyme in the detoxifying system known as mercury reductase (encoded by merA gene), an NAD(P)H-dependent flavin oxidoreductase, reducing toxic mercury (Hg2) into a volatile natured less toxic mercury (Hg0) (Freedman et al. 2012; Dash et al. 2017). Likewise, microbial arsenic bioremediation mainly involves arsenic reduction (arsC), oxidation (AoxAB, arsH), methylation, and intra-cellular bioaccumulation (Pepi et al. 2007; Rahman and Hassler 2014; Govarthanan et al. 2015, 2016a, 2019; Satyapal et al. 2018; Ben Fekih et al. 2018). The ars operon is an energy-gaining detoxifying respiratory system that facilitates the reduction of pentavalent arsenate(V) into trivalent-arsenite(III) (Suhadolnik et al. 2017). The interaction of transcriptional repressor protein (arsR) with the promoter region of ars operon, followed by the binding of arsR protein with arsenate, dissociates the repressor protein from a specific DNA site permitting the ars operon’s transcription (Páez-Espino et al. 2015; Li et al. 2016). The arsC encoding arsenic reductase enzyme governs this organoarsenic reduction process. The ars operons, such as arsRB, arsRBC, arsRDABC, and arsHRBC, are widely demonstrated in gram-negative bacteria (Dey et al. 2016; Ben Fekih et al. 2018; Taran et al. 2019).
The existence and sustainability of these metal resistance operons among the various bacterial communities are usually maintained and transmitted with the help of mobile genetic elements via horizontal gene transfer (Møller et al. 2014; Suhadolnik et al. 2017). In some bacteria, this metal resistance sustainability is preserved intrinsically in the bacterial chromosome, resulting from vertical gene transfer from its ancestral species. Many bacterial species reportedly have arsenic and mercury resistance operons in their chromosome or mobile genetics elements (Møller et al. 2011; Ghaly et al. 2017; Agarwal et al. 2019). However, none among the species in the genus Pseudoxanthomonas have been demonstrated for any heavy metal resistance and their respective operons. The 25 members of this genus are gram-negative, rod-shaped, and non-pathogenic that are long been known for their prevalence in diverse ecological sources, including soil, water, plants, animal, and human tissues (Thierry et al. 2004; Rani et al. 2010; Kittiwongwattana and Thawai 2016; Selvaraj et al. 2018). Although the genus Pseudoxanthomonas is well characterized for its biodegradation capability and surfactant production (Choi et al. 2013; Talwar and Ninnekar 2015; Biswas et al. 2017; Astuti et al. 2019; Lu et al. 2019; Purwasena et al. 2020), it is unremarked in many environmental processes, particularly in terms of its bioremediation capability. This study thereby deals with (i) isolation and characterization of Hg and As resistant strain GTZY from a bioreactor treating antibiotic-containing wastewater, (ii) evaluation of Hg and As removal efficiency of strain GTZY in batch-culture experiments, (iii) disclosure of the mer and ars resistance operon from strain GTZY, and (iv) phylogenetic distribution and relationship of identified mer and ars operon of strain GTZY with other bacterial communities.
Materials and methods
Bacterial strain
The bacterial samples were collected from the aerobic-biofilm wastewater reactor treating aminoglycoside antibiotic, streptomycin (50 mg L−1) (State key laboratory of RCEES, CAS Institute, Beijing, China). The tenfold serial dilution method was used to isolate and enrich bacterial isolate from the collected sludge samples. Inoculated (100 µL aliquots) plates were incubated at 30 °C for 12–18 h. Tryptone soy agar (TSA) and Reasoner’s 2A agar (R2A) media were used for the purpose of bacterial isolation. Gram’s staining was routinely performed to check the microbial purity. Initially, 23 bacterial isolates were harvested from the wastewater reactor. To obtain the potential heavy metal transforming bacteria, the purified isolates were further screened on the growth media amended with sodium arsenate (Na2HAsO4), 1 µM, and mercuric chloride (HgCl2), 10 µM. Among the 23 isolates, strain GTZY was chosen for further studies due to its fast-growing ability and survival capability in the presence of increased concentrations of As(V), 100 µM, and HgCl2, 50 µM. These screened isolates were stored as 40% glycerol stock for further studies.
General characterization of bacterial isolate
The morphological, biochemical, and physiological characterization of isolated bacterial strains was performed based on standard microbiological methods. The optimum temperature and pH were identified by culturing the strain on the nutrient agar media at various temperatures (25−40 °C) and pH (5−8). After 12−18 h incubation, bacterial colony morphology was determined using a single colony grown on the nutrient agar, TSA, LB, and R2A media (Dziurzynski et al. 2023). The hanging drop method was performed to check the motility of isolates under light microscopy. Gram’s staining solutions were manually prepared and isolates were viewed under light microscopy magnifications (× 100). A salt tolerance test was carried out by preparing brain–heart infusion agar plates with the addition of various NaCl concentrations of 0−7% (w/v). Activity of various enzymes (catalase, oxidase, urease, and indole), hydrogen sulfide production, utilization of various carbon sources, nitrate reductability, and hydrolysis of biochemicals (gelatin, starch, and casein) tests were also carried out by the standard microbial methods (Ahn et al. 2019; Batbatan et al. 2022).
16S rRNA gene amplification and phylogenetic tree analysis
The genomic DNA isolation and 16S rRNA gene PCR amplifications were carried out as described in the previous study (Selvaraj et al. 2018). In brief, the genomic DNA was harvested from the bacterial samples using the TIANamp bacteria DNA kit (Tiangen, China), and the isolated DNA was quantified using Nanodrop 1000 spectrophotometer (Nanodrop, USA). The 16S rRNA fragments were amplified using the universal primers (27F-5′-AGAGTTTGATCCTGGCTCAG-3′ and 1492R-5′-AAGGAGGTGATCCAGCCGCA-3′), sanger sequenced (Biobase, China) and interpreted by the Ribosomal Database Project (RDP) and GenBank-National Center for Biotechnology Information (NCBI) database. The MEGA6.06 and Clustal W software were used to construct a phylogenetic tree for isolated strains against its closest 19 bacterial strains. The neighbor-joining method and bootstrap analysis were executed for the bacterial phylogeny and statistical sequence analysis, respectively. In bootstrap analysis, 1000 replicates were applied to determine the confidence values of bacterial phylogenetic tree nodes. Finally, the partial 16S rRNA gene sequence was submitted to the GenBank (Accession No.: MZ242608).
Determination of mercury MIC
After bacterial isolation, the bacterial efficiency on mercury resistance was maintained in LB media by the supplement of mercury chloride solution (10−30 µM; HgCl2). Bacterial aliquots of 1.0 mL of overnight cultures were incubated in 99.0 mL of the Muller Hinton (MH) broth added with 10 µM of HgCl2. Subsequently, those cultures were resuspended with 2−5 mL of Tris-buffered saline (TBS) solution, and absorbance (OD600 nm) was adjusted to 0.100 OD. Further, 125 µL of appropriate bacterial dilutions was inoculated on the 96-well plate containing 125 µL of LB broth (in triplicate) amended with various concentrations (0, 1, 5, 10, 20, 30, 40, 50, 60, 70, and 80 µM) of HgCl2 and incubated at 30 °C for 72 h in the dark. After the incubation, bacterial growth was observed by measuring the absorbance (OD600 nm) using a microplate reader. The inhibition effect of Hg on microbial growth and values of MIC and MIC50 were calculated.
Mercury removal assay
Overnight grown bacterial cultures (in triplicate) added with 10 µM of HgCl2 were subjected to the mercury removal assay. Every 12 h of incubation, 4 mL of culture was centrifuged for 10 min at 7500 rpm, thereby recovered bacterial supernatant was resuspended in 2.8 mL of Tris-buffered saline solution. In the resuspended solution, 0.4 mL of 5-diphenyl-3-thiocarbazone (dithizone) reagent solution (0.50 mg of dithizone mixed with 20 mL of acetone) was added, followed by the addition of 0.4 mL of sulphuric acid (2 N) to maintain the acid pH (pH 2−3), and finally it was added by 0.4 mL of dioxane (Naguib et al. 2019). The acidic pH of the solution enhanced the formation of orange-red color by coupling dithizone and Hg(II). These final solutions were diluted to 5 mL using distilled water and measured mercury removal values at 488 nm in a UV/visible spectrophotometer. Meanwhile, a similar experimental setup was used for the control samples using dead bacterial cells (autoclaved cells). The reference calibration curve was also prepared for standard HgCl2 solutions treated as described above but without bacterial suspension. The detection limit for this method was estimated as < 1 ppb (Nordlander 2024). The actual Hg concentrations during the incubation period were measured and thereby the Hg removal percentage was calculated as follows,
Mercury volatilization assay
A simplified X-ray film method was used to confirm the mercury volatilization from Hg(II) state to Hg(0) state by the strain GTZY. Overnight grown culture which was added with 10 µM; HgCl2 was centrifuged at 6000 rpm for 5 min. Harvested bacterial pellets were resuspended into a 200 µL assay solution containing 0.07 M phosphate buffer (pH 7), 0.2 mM magnesium acetate, 5 mM sodium thioglycolate, and 50 µM HgCl2. The microtiter plate containing the above-mentioned solution was fully covered using an X-ray film (Kodak Scientific Imaging, Hyderabad, India) (Møller et al. 2011, 2014) and incubated at 35 °C in the dark for 60 min. A similar solution mixture without bacterial cells and bacterial suspension without HgCl2 were utilized as control samples.
Determination of arsenic MIC
The bacterial arsenic resistance was evaluated by adding the different concentrations of sodium arsenate (10−400 mM) and sodium arsenite (10−200 mM) into the overnight-grown bacterial culture, while the control samples were added with no arsenic metals. The cultures were resuspended with 2−5 mL of TBS solution with an absorbance (OD600 nm) of 0.100. Further, 125 µL of appropriate bacterial dilutions and 125 µL of MH broth (in triplicate) were added to the 96-well plate containing suspended concentrations of arsenic and incubated at 30 °C in the dark for 48 h (in triplicate). After the incubation, tested bacterial samples were estimated for their microbial growth at 600 nm using a microplate reader. The inhibition effect of arsenic on microbial growth and values of MIC and MIC50 were calculated.
Arsenic removal assay
After the bacterial isolation, the efficiency of arsenic resistance was sustained by adding sodium arsenate (100 mM) and sodium arsenite (100 mM) into the bacterial growth media (NA). To obtain the bacterial assay supernatant, the isolate (1%) was inoculated (in triplicate) in LB broth added with 5 µM sodium arsenate and sodium arsenite solution (as sources for As(V) and As(III), respectively) and incubated at 30 °C for various time intervals (viz. 8, 12, 18, 24, 48, and 72 h). After the definite time points, the respective bacterial biomass was separated from the growth media by 10,000 rpm for 10 min and the derived supernatant was subjected to the bioremediation assay. The actual arsenic utilization by bacterial isolate was determined by the silver diethyldithiocarbamate (SDDC) method (Tahir et al. 2012). The detection limit for this method was evaluated as 1–5 ppb. Briefly, the production of arsine-carrying hydrogen passed through a reduction column impregnated with lead acetate and then into the absorbing solution SDDC dissolved pyridine. The end red-colored product (arsine with SDDC solution) was measured spectrophotometrically at 535 nm.
Arsenic reduction and oxidation assay
The ability of the bacterial isolate to oxidize arsenic (As(III)) and reduce arsenic (As(V)) was checked by the silver nitrate solution (Dey et al. 2016). The bacterial isolates were cultured in the broth media (peptic digest of animal tissue, 5 g/L; yeast extract, 1.5 g/L; and beef extract, 1.5 g/L). After 24 and 48 h of incubation, 100 µL of silver nitrate solution (0.1 M) was added into the grown bacterial culture (100 µL) amended with 1 µM sodium arsenate and sodium arsenite. If the culture broth turns brown, it represents the presence of silver arsenate, while if the culture broth turns yellow, it represents the presence of silver arsenite.
merA and arsC gene sequencing
The extracted and purified genomic DNA was used as a template DNA to screen mercury (merA) and arsenical (arsC) resistance genes. The designed merA (merA-FP-5′-CTACCCCGCGCAACAGGACA-3′ and merA-RP-5′-TGATGGCGCCTTGCGCATTG-3′) and arsC (arsC-FP-5′-TCAGGACGAGGCGCCGATCT-3′ and arsC-RP-3′-ATGGACCGCCCCTACAACCT-5′) primers were used to amplify the partial regions merA and arsC. The purified genomic DNA (50 ng) was used as a template for preparing the PCR reaction. The triplicate PCR mixture (50 μL) comprised of 1 × PCR buffer (with MgCl2+), 2.5 mM of the dNTPs mix, 0.5 μM of each primer, and 1 U of Taq DNA polymerase (Takara, Bio Inc., Shiga, Japan). The PCR conditions were performed as follows: initial denaturation stage at 95 °C for 5 min, followed by 30 cycles of denaturation at 95 °C for 30 s, annealing at 56 °C for 30 s, extension at 72 °C for 2 min, and a final extension at 72 °C for 10 min. The amplified specific PCR products were resolved using 1.2% agarose (Biowest, Hong Kong) gel and sequenced (Sanger sequencing). The gene sequences were analyzed using BLASTN.
Annotations of mer and ars gene operons
The total DNA extraction, DNA quality evaluation, genome sequencing, and genome analysis of strain GTZY were elaborately given in our previous research studies (Selvaraj et al. 2022). Briefly, the extracted total genome of isolated strain GTZY was sequenced using the third-generation single-molecule real-time (SMRT) technology (Pacific Biosciences Inc., USA). The original genome raw reads and obtained filtered subreads were further assembled (3,936,186 bp) using the Hierarchical Genome Assembly Process (HGAP) and the sequenced genome was submitted to the GenBank (Accession No.: CP060028). The whole-genome sequence of Pseudoxanthomonas mexicana strain GTZY was used for the resistance gene cluster’s interpretation. The mer and ars resistance gene clusters related data were initially annotated using the online ORF prediction server, Rapid Annotations using Subsystems Technology (RAST), and their manual annotations were performed using BLAST. A Python application software, Easyfig (ver. 2.1), was used for the linear comparison and visualization of mer and ars resistance gene clusters. In addition, the BLAST comparison between the gene annotated files was manually uploaded, output files (blastn) generated, interactively colored, and enabled a rapid transition using the software, Easyfig. The SnapGene Viewer (ver. 4.1.3) was used for the gene annotation and visualization.
Results
Characterization of bacterial isolate
The isolated bacterial strain was aerobic, gram-negative, rod-shaped, and highly motile. The colonies were pale yellow, circular, convex, and mucoid-natured on the solid nutrient media (Fig. S1). The isolate can grow in the media up to 40 °C and pH 8, while its optimum temperature and pH were 34−37 °C and 7−7.5, respectively. Isolate can tolerate the presence of sodium chloride up to 8% (w/v); however, its optimum concentration was 3.5–5.5%. Their biochemical and phenotypic characterizations are given in Table S1. Biochemically, the isolate turned positive for catalase while remaining negative for oxidase and urease, which in turn ferments glucose and mannitol but not sucrose and lactose. The hydrolysis test turned positive for the gelatin and casein but remained negative for starch. Moreover, it showed negative results for indole production, nitrate reductase, and Voges-Proskauer (VP) test, and positive for the Tween-20 and Methyl Red (MR) test. The length of the 16S rRNA gene sequence was 1424 bp, and it shared the closest relationship (99.39%) with P. mexicana CP4 (Accession No.: MT549102) in the phylogenetic tree by the neighbor-joining method (Fig. 1). Hence, the isolated bacterium was named as Pseudoxanthomonas mexicana strain GTZY and submitted to the GenBank (Accession No.: MZ242608).
A phylogenetic tree showing the clustering of twenty 16S rRNA gene sequences along with an isolated bacterial strain GTZY. The neighbor-joining constructed tree was evaluated by the bootstrap method. The scale bar 0.0002 represents the nucleotide substitution level, and numbers indicate bootstrap values
MIC, volatilization, and bioremediation of mercury
The effect of mercury on microbial growth was studied. Further, MIC and MIC50 were determined by supplementing different Hg(II) concentrations into LB broth inoculated with strain GTZY. After 48 h of incubation, mean values of OD600nm were plotted against 0−80 µM of HgCl2. The MIC value of Hg(II) against strain GTZY was 33.4 µM of HgCl2, and its MIC50 value was 40.8 µM of HgCl2. Microbial growth was notably inhibited by 35.5 µM of HgCl2, and it was completely inhibited (0.21 OD600nm) at 49.7 µM of HgCl2 (Fig. 2A). After 60 h of incubation, the residual concentration of Hg was 1.7 ± 0.2 µM and it was sustained for the remaining periods (Fig. S3A). The mercury-reducing ability of the strain GTZY was confirmed by a mercury volatilization assay conducted at 48 h of incubation. The strain formed a foggy area (dark circle) due to the reaction between Hg(0) and Ag emulsion on the X-ray film (Fig. S2). However, the isolate started to produce foggy formation from 4 h onwards. Hence, the strain GTZY reduced inorganic Hg(II) and formed volatile Hg(0), which reacted with the film. There was no interference of foggy circles observed in the control samples.
Mercury removal by the strain GTZY was determined using the spectrophotometric dithizone method. Initial and residual Hg concentrations were evaluated against the standard calibration curve of various concentrations of Hg. The strain GTZY removed 16.3% of Hg from 4 h onwards, and their removal process was stopped at 48 h of incubation (Fig. 2B). The strain GTZY removed a maximum of 84.3% Hg(II) at 48 h and their removal rate was maintained until 96 h of incubation. At the same time, no mercury removal was found in the control samples.
MIC, reduction, and bioremediation of arsenic metals
The isolate GTZY was resistant to the high concentrations of arsenate and arsenite solutions. Also, it removed the arsenic compound at various periods. Isolate GTZY has efficiently removed arsenate (68.5%) and arsenite (63.2%) at 72 h of incubation (Fig. 3). At 12 h of incubation, the removal rate (%) of arsenate and arsenite accounted for 34.2% and 28.3%. Meanwhile, no notable removal of arsenical compounds was observed in the control samples. The MIC values of isolate towards arsenate and arsenite were > 175 and > 55 mM, and their corresponding MIC50 were 240 and 60.5 mM, respectively. The bacterial growth was completely arrested with 330 mM arsenate and 69.4 mM arsenite (Fig. S4). After 60 h of incubation, the residual concentrations of As(III) and As(V) were 22 ± 2 µM and 27.5 ± 2.5 µM (Fig. S3B); thereafter, it was sustained for the remaining periods. After 24 and 48 h of incubation, 100 µL of silver nitrate solution was added to both bacterial cultures (As(III) and (V)), and the suspensions were turned into a light yellow color, which indicated the presence of As(III). Meanwhile, there was no brownish color observed which indicates the absence of silver arsenate. The result confirmed that isolate GTZY could reduce As(V) to As(III), while the isolate could not oxidize As(III) to As(V).
Percent removal of arsenite(III) and arsenate(V) by the isolate GTZY was estimated using silver diethyldithiocarbamate (SDDC) method. The experimental strain (1%) was inoculated in LB broth amended with 5 µM sodium arsenate and sodium arsenite solution (as sources for As(V) and As(III), respectively) and incubated at 30 °C
Resistance gene sequencing
From the specific gene PCR, arsC (495 bp) and partial merA (550 bp) genes were amplified successfully and this was confirmed by the agarose gel electrophoresis (Fig. S5). The amplified DNA fragments were sequenced and annotated in GenBank. The partial sequence of merA from isolate GTZY was closely similar (coverage, 96%; similarity, 83.37%) to the merA sequence of Lysobacter oculi strain 83–4 (Accession No.: CP029556), which encodes for mercury(II) reductase enzyme. arsC gene of isolate GTZY was closely similar (coverage, 100%; similarity, 99.88%) with the arsC encoding arsenate reductase enzyme of Stenotrophomonas acidaminiphila strain ZAC14D2_NAIMI4 (Accession No.: CP012900).
Mercury gene cluster and its interpretation
From the genome analysis of strain GTZY, an entire set of mercury resistance gene clusters, named merAPTR (2829 bp), was identified (Fig. 4A). The gene cluster possessed 1695 bp of merA (mercury(II) reductase), 279 bp of merP (mercury(II) binding protein), 351 bp of metT (mercuric transport protein), and 408 bp of merR (mercuric(II) resistance transcription regulator) genes. This gene cluster was flanked by a linkage of Tn2-family transposase, ISII (579 bp), and a site-specific integrase, int1 (1080 bp), on both sides. IS-II family transposase was located adjacent to the gene encoding lead, cadmium, zinc, and mercury-transporting ATPase (zntA). The entire mer gene cluster consists of 34 restriction enzyme sites in its respective gene sites.
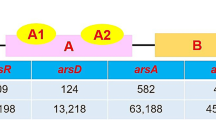
The genetic organization of heavy metal resistance operons from Psudoxanthomonas mexicana GTZY. Arrows indicate the orientation of transcription and its open reading frame. Arrow length indicates the base pair length of particular genes. The particular gene descriptions are given in detail in the text. A Mercury resistance operon and B arsenic resistance operon
Seven different bacterial mercury gene clusters (Thermomonas sp., Stenotrophomonas acidaminiphila, Pseudomonas aeruginosa, Cupriavidus metallidurans, Achromobacter xylosoxidans, Delftia lacustris, and Burkholderia cenocepacia) along with the strain GTZY were chosen for the gene cluster interpretation, and their comparative visualization is given in Fig. 5. The strain GTZY mer gene cluster’s downstream and upstream were highly conserved against three of seven bacterial species. Besides, it had the closest relationship (coverage, 100%; similarity, 99.89%) with the mercury gene cluster of Thermomonas sp. XSG (Accession No.: CP061497) and were followed by the relationship (coverage, 100%; similarity, 99.82%) with Stenotrophomonas acidaminiphila strain ZAC14D2 (Accession No.: AM743169) and Pseudomonas aeruginosa H47921 (Accession No.: CP008861) (coverage, 99%; similarity, 82.17%). From these interpretations, it was evidenced that merA encoding mercury(II) reductase and merP encoding Hg(II) binding protein were invariably conserved in mercury gene clusters of all the bacterial species except Cupriavidus metallidurans CH34 (Accession No.: CP000352). Meanwhile, merR encoding mercury(II) transcription regulator was invariably conserved in all the bacterial species except Achromobacter xylosoxidans FDAARGO 984 (Accession No.: CP066291).
A phylogenetic distribution and genetic organization of mercury resistance operon of Psudoxanthomonas mexicana GTZY. mer gene clusters of different bacterial strains were annotated by BLASTn and compared with the Easyfig tool. The mer operon of strain GTZY has the closest relationship (coverage, 100%; similarity, 99.89%) with the mer operon of Thermomonas sp. XSG (Accession No.: CP061497). The degree of sequence homology (%) between the operons was shown by the grey shades, and shade intensity indicates the relationship level between the mer operons. Arrows indicate the orientation of transcription and its open reading frames. The bar indicates 100 bp lengths of nucleotides
Arsenic gene cluster and its interpretation
From the genome analysis of the strain GTZY, an entire set of arsenic resistance gene clusters, arsCPRH (2791 bp), was identified (Fig. 4B). This gene cluster possessed 495 bp of arsC (arsenate reductase), 1026 bp of arsP (arsenate permease protein), 336 bp of arsR (arsenate repressor protein), and 819 bp arsH (arsenate resistance protein). The identified ars gene cluster comprised 41 restriction enzyme sites in its respective gene sites. Three different bacterial mercury gene clusters (Dokdonella koreensis, Luteimonas sp., and Lysobacter oculi) and the strain GTZY were chosen for the gene cluster interpretation and visualization (Fig. 6). However, strain GTZY arsCPRH gene cluster’s downstream and upstream were more conserved against only to Dokdonella koreensis DS-123 (Accession No.: CP015249) (coverage, 94%; similarity, 79.14%) and least conserved with Lysobacter oculi 83–4 (Accession No.: CP029556) (coverage, 64%; similarity, 77.51%). The arsenic resistance genes arsP, arsR, and arsH were conserved among all four bacterial genera. In the meantime, the arsC gene was observed only in the isolate GTZY and Dokdonella koreensi DS-123. The arsenic gene cluster’s arrangement in Luteimonas sp. (arsPRH) and Lysobacter oculi (arsRPH) was extremely different from the strains GTZY and Dokdonella koreensis (arsCPRH).
A graphical representation of BLASTn comparison of arsenic resistance operon of Psudoxanthomonas mexicana GTZY along with various bacterial species. ars operon of Dokdonella koreensis DS-123 (Accession No.: CP015249) is the only bacterial strain shown the maximum relationship (coverage, 94%; similarity, 79.14%) with ars operon of isolate GTZY and least relationship (coverage, 64%; similarity, 77.51%) with Lysobacter oculi 83–4 (Accession No.: CP029556)
Discussion
Only a few gene clusters encoding toxic chemical degradation have been reported so far in Pseudoxanthomonas species (Choi et al. 2013; Wang et al. 2013). Thereby, in this study, we looked into the bioremediation capability of two different heavy metals (Hg and As) and their related resistance gene operons in the environmental non-pathogenic bacterium, P. mexicana strain GTZY, for the first time. Initially, PCR amplification confirmed the presence of metal resistance genes (merA and arsC). Interestingly, further genome annotation studies revealed the occurrence of corresponding resistance operons set (merAPTR and arsCPRH) in the genome of strain GTZY. In particular, 2829 bp size mercury operon was located initially with mercury ion reducing reductase enzyme (merA), which has been proven to volatilize or reduce the toxic Hg(II) into non-toxic Hg(0); then, it might extrude passively from the microbial cellular environment (Fig. 8) (Sotero-Martins et al. 2008; Agarwal et al. 2019). This detoxification mechanism is similar to some aerobic bacteria and archaea that have previously been reported (Christakis et al. 2021). In particular, Hydrogenobaculum sp, Hydrogenivirga sp, E. coli, Flavobacterium sp, Candidatus sp, Sphingobacterium sp, and Chryseobacterium gleum have shown merA-based resistance mechanisms. Meanwhile, Halobacterium and Candidatus have been reported to reduce Hg(II) to Hg(0) by the unusual merB (encoding organomercury lyase)-based resistance mechanism (Møller et al. 2014; Krout et al. 2022). This volatilization of Hg(II) by methylation mechanism is similar to other enzyme-based heavy metal methylation (e.g., Se and Pb) in Pseudomonas spp., Escherichia spp., Bacillus spp., and Clostridium spp. (Boyd and Barkay 2012).
In merAPTR, periplasmic Hg(II) binding protein-encoding merP and Hg(II) transporting inner membrane protein-encoding merT were exactly located at 10 and 12 bp upstream of the merA gene, respectively. It is followed by the cytoplasmic mercury regulatory protein-encoding merR, located before the transporter gene and 729 bp downstream of the merA gene. The gene cluster orientation and specifications of the strain GTZY’s mer operon were found to be in a close relationship (99.72%) only with mer operons of other gram-negative bacteria, Thermomonas sp. (Accession No.: CP059266) and Stenotrophomonas sp. (Accession No.: AM743169). Despite the orientation (IS2-merA-merP-mer-T-merR-int1) of the mer operon of Pseudomonas sp. and Delftia sp. was similar to the mer operon of strain GTZY, they have only 82.17% and 79.19% of gene similarities, respectively.
More than 80% of bacterial communities were demonstrated for the presence of their mer operon embedded in their chromosomal DNA (Boyd and Barkay 2012) and the strain GTZY does the same, and no plasmids were detected in the strain GTZY. The ancestor species of Pseudoxanthomonas was not detected for any mer operon, excluding the process of strain GTZY which does mercury bioremediation. Besides, another strain Pseudoxanthomonas mexicana GTZY2 (Accession No.: CP060731) isolated from a wastewater reactor (control) not exposed to any antibiotics was not detected for any heavy metal–related operons in its genome. However, the mer operon of the strain GTZY has distinctly flanked both ends by Tn2 family transposase (IS-II) and site-specific integrase (int1). Previously, species of Bacillus, Stenotrophomonas, Pseudomonas, Paenibacillus, Enterobacter, and Exiguobacterium were reported for the mer operon linked with elements of transposons (TnMERI1, Tn5083, Tn6294, and Tn5085) (Ramírez-Díaz et al. 2011; Wang et al. 2013; Matsui et al. 2016; Agarwal et al. 2019) and integron gene (int1) (Ghaly et al. 2017). These two mobile genetic fragments were proven for the transmission of functionally specific genes among the various bacterial communities (Wang et al. 2013; Ghaly et al. 2017; Ben Fekih et al. 2018). Hence, we speculated that the mer operon of strain GTZY is not to be inherited through vertical gene transfer from its ancestral species of Pseudoxanthomonas; instead, it might be received from other related bacterial communities by the horizontal gene transfer mechanisms such as conjugation, transposition, or phage attack with mer operon. Besides, it seems that the strain GTZY can be a perfect fit for the heavy metal selected genome plasticity and interaction with other bacterial communities, which often have consequences with evolved bacterial species in the toxic heavy metal–contaminated sites. Since the strain GTZY was isolated from the high-dose streptomycin-treating reactor, it could have a high fitness cost that leads to the out-competence by wild-type sensitive strains existing in the heavy metal–contaminated sites.
On the other side, the strain GTZY possessed an arsenic gene operon (arsCPRH) in its chromosome with a size of 2791 bp. The identified arsCPRH-based arsenic mechanism (Fig. 8) from the strain GTZY was diverse from previously reported other prokaryotic ars operons such as arsRBC, arsRBCH, and arsRDABC identified in the E. coli, Staphylococcus xylosus, Pseudomonas aeruginosa, Pseudomonas pudita, Pseudomonas fluorescens, Acidiphilium multivorum, and Staphylococcus aureus (Páez-Espino et al. 2015; Chen et al. 2016; Ben Fekih et al. 2018). Like other bacteria, strain GTZY also consists of arsenate reductase encoding the arsC gene, but not permease encoding the arsP gene. The arsenic mechanism and gene cluster orientation of strain GTZY and their gene components had only shown matching with a soil bacterium, Dokdonella koreensis DS-123 (Accession No.: CP015249). The multiple sequence alignment between these two ars operons revealed their maximum gene cluster coverage and base pair similarities accounted for 94% and 79.14%. It is thereby likely that identified ars operon was distinct from other bacterial genera containing ars operons. Besides, the identified ars operon from the strain GTZY has not flanked with any mobile genetic elements in both ends and it is located intrinsically in a chromosome, which might be received from its ancestral strains by the process of vertical gene transfer.
From the genome interpretation study, we confirm the existence of ars operon and mechanism in its ancestral bacterial species, Pseudoxanthomonas suwonensis 11–1 (Accession No.: CP002446), isolated from cotton waste compost (Weon et al. 2006) and Pseudoxanthomonas spadix BD-a59 (Accession No.: CP003093), known for BETX biodegradation (Choi et al. 2013) (Fig. 7). This further reveals the arsenic gene cluster orientation (arsC-arsP-arsR-arsH) of strain GTZY as divergent (< 38% similarity) from their inherited BD-a59 (arsH-arsB-arsC-arsR) and 11–1 (arsR-arsH-arsB-arsC) strains. In particular, arsR and arsH of the strain GTZY were similar with the nucleotides of BD-a59 and 11–1 strains, while arsC was diverse in their ancestral strains, respectively. The genome of strain GTZY is comprised of the membrane permease transporter encoding the arsP gene while BD-a59 and 11–1 strains are located with the most frequent acr3 family efflux transporter encoding arsB gene, which extrudes arsenate(III). Apart from arsCPRH, strain GTZY has also been distributed with additional arsB (963 bp), arsR (219 bp), and arsC (420 bp) genes in its chromosome.
The genomic comparison of intrinsic ars operon of Psudoxanthomonas mexicana GTZY along with its bacterial ancestral species (Easyfig visualization). An extremely diverse relationship (< 38%) was detected against ancestral strains, Pseudoxanthomonas spadix BD-a59 (Accession No.: CP003093), reported for BETX biodegradation (Choi et al. 2013) and Pseudoxanthomonas suwonensis 11–1 (Accession No.: CP002446) isolated from cotton waste compost
The role and cellular mechanisms of mer and ars systems occurred in the Pseudoxanthomonas mexicana strain GTZY. P, periplasmic permease facilitate the cellular uptake of heavy metals (Hg and As); R, regulator initiates the transcription of corresponding genes; T, inner membrane protein transport the Hg(II) to reducing enzyme; A, merA encoding mercury reductase reducing toxic Hg(II) to non-toxic Hg(0); C, arsC encoding arsenic reductase reducing As(V) to As(III); H, arsH encoding arsenical(III) oxidase enzyme transferring toxic MAs(III) to less toxic MAs(V). Reduced or oxidized metals extrude via cell membrane efflux system. The numbers represent the steps of heavy metal detoxification. The reduced As(III) and Hg(0) are further extruded via cell membrane efflux system, arsB encoding Ars3-family transporter and Hg/Cd/Pb transporting protein, respectively
The presence of arsP and its co-existence with the arsH gene in the ars operon of strain GTZY makes them versatile among other known bacterial ars operons. The arsP encoding efflux-related permease system is known for the detoxification or extrudes the toxic organic arsenicals(III) such as roxarsone and monomethylarsonous acid (MMA) (Shen et al. 2014; Chen et al. 2015b). The arsP gene encoding ArsP permease was initially identified in the ars operon of Campylobacter jejuni (Wang et al. 2009), and later it was believed to be a prokaryotic cell membrane pump that could extrude arsenical roxarsone(III) (Shen et al. 2014). The recent genomic interpretation study suggests that the ancestral origin of arsP and their homologs exists in bacteria, archaea, and a few other eukaryotes (Yang et al. 2015). The existence of a gene encoding flavoprotein, arsH, utilizing NADP + to resist and oxidize the toxic methylarsenite (MAs(III)) into MAs(V) was reported in Campylobacter jejuni strain (Li et al. 2016). These arsH enzymes are widely distributed in gram-negative, particularly in Gammaproteobacteria including Pseudomonas putida, Sinorhizobium meliloti, Acidithiobacillus ferrooxidans, Rhodopseudomonas palustris, Ochrobactrum tritici, Thiobacillus ferrooxidans, Herminiimonas arsenicoxydans, and Pseudomonas aeruginosa, but not in gram-positive bacteria (Yang et al. 2005; Chen et al. 2015a; Yang and Rosen 2016; Falgenhauer et al. 2017). The ArsP protein from both P. putida and S. meliloti was evaluated as an organoarsenical oxidase enzyme conferring resistance to trivalent methyl As derivatives. We assumed that arsCPRH of strain GTZY has highly evolved from its ancestral species, in response to the environmental mutations, genetic reshuffling, and recombination with other bacterial communities.
Initially, P. mexicana GTZY isolate was characterized for its phylogeny, morphology, and biochemical properties, and was found specifically relevant to the properties of various reported members of Pseudoxanthomonas. Like other heavy metal–transforming bacteria, strain GTZY did not utilize mercury and arsenic as main carbon sources in the basal media (data not shown). Since these heavy metals are not catabolized by the heavy metal transforming bacteria, their bioremediations were usually evaluated by the removal, reduction, oxidation, and volatilization assays (Wuana and Okieimen 2011; Jebeli et al. 2017; Wang et al. 2020). GTZY has successfully fulfilled the above chemical strategies against the mercury and arsenic metals in the growth media.
The strain GTZY can reduce inorganic Hg(II) and into foggy forming volatile Hg(0) in 4-h incubation. Previously, Mahbub et al. (2017) have proved the ability of Sphingopyxis sp. SE2 to volatilize or remove 44% of Hg(II) in 6-h incubation. This study found that 49.6% of mercury removal was achieved by the strain GTZY in 6 h. However, Sphingobium sp. SA2 was recorded for the maximum Hg(II) volatilization and reduction in 6 h accounts for 79% (Mahbub et al. 2016, 2017). The strain GTZY’s mercury volatilization and reduction ability were further confirmed by the occurrence of the merA gene encoding for mercury(II) reductase. Also, the isolate was proven to resist (> 33.4 µM) and remove (84.3%) the toxic form of Hg(II) in 48-h incubation. The minimum inhibitory concentrations (MIC and MIC50) of strain GTZY against Hg(II) were observed to be 33.4 µM and 40.8 µM, respectively, whereas in other bacterial classes, Alphaproteobacteria, Betaproteobacteria, Flavobacteria, Firmcutes, and Sphingobacteria, MICs were observed in the range of 5–30 µM; Hg(II) (Møller et al. 2011, 2014). After 60-h incubation, the concentration of Hg(II) remained stable at 1.9 ± 0.2 µM along with strain GTZY. In the meantime, thermophilic species Aquificae was proven to be sustainable at 3.0 µM, Hg(II) in the basel media (Freedman et al. 2012). These results represent that the strain GTZY could be useful for preventing the formation of a more toxic form of methyl mercury (MeHg). In contaminated environments, high concentrations and precipitation of Hg(II) always end up with more accumulation of toxic MeHg (Eckley et al. 2020). Hence, this is arguably a better solution to reduce the formation of toxic MeHg, in which a high concentration of Hg(II) should be cleaned up first so that the strain GTZY would be more suitable for detoxification.
On the other side, strain GTZY can remove (68.5%) and resist (> 175 mM) As(V) in the culture media at 72-h incubation. The minimum inhibitory concentrations of strain GTZY towards arsenate and arsenite were > 175 and > 55 mM. Previous studies revealed the arsenate and arsenite MICs of Gammaproteobacteria, Firmcutes, and Kocuria, in the ranges of 75–125 mM and 10–50 mM, respectively, whereas in the Bacillus fusiformis ORAs1, B. thuringiensis ORAs2, Pseudomonas sp. ORAs5, and Aeromonas molluscorum ORAs6, MICs of arsenate and arsenite were > 16.68 mM and > 133.47 mM, respectively (Pepi et al. 2007). In addition, strain GTZY reduced organic As(V) into As(III) form in the liquid solution, and it was further confirmed by the amplification of the arsC gene (495 bp) encoding arsenate reductase. Meanwhile, the strain GTZY was also able to remove (63.2%) and resist (> 55 mM) As(III); however, it could not oxidize organic As(III) into As(V) form in the liquid solution. Similarly, Dey et al. (2016) isolated Bacillus sp. and Aneurinibacillus aneurinilyticus and found that both strains could remove As(V) (51.45–51.99%) and As(III) (50.37–53.29%). Besides, the strains also resisted the arsenate (4500 ppm) and arsenite (550 ppm). However, both strains have oxidized the arsenite, but none of them reduced arsenate into arsenite. Many previous reports suggest different detoxification mechanisms of arsenic resistance bacteria, including arsenic reduction and oxidation, heavy metal efflux pumping, metal adsorption to the cell surface, bacterial cell membrane binding, and complex formation with exopolysaccharides (Zhang et al. 2016; Govarthanan et al. 2016a). Despite the absence of an arsenite oxidase enzyme, we presume that the strain GTZY might be removed from the arsenite(III) by any of the above-proposed detoxification mechanisms.
In the future, the strain GTZY can be used for both real-world in situ and ex situ bioremediation technologies for treating heavy metal–polluted sites. Since the isolated strain is an aerobic and indigenous bacterium, it is suitable for some bioremediation techniques such as biosparging, bioventing, and biostimulation (Bala et al. 2022; Jeyakumar et al. 2022; Alabssawy and Hashem 2024; Helmy and Kardena 2024). Before the real-world implementation of bioremediation, it is better to conduct an in vitro microcosm or model study. This in vitro trial work will help us to understand the preliminary assessment of microbial degradation patterns, environmental parameters, and targeted field issues.
Conclusion
From this study, it is therefore concluded that Pesudoxanthomoas mexicana strain GTZY has potential for the bioremediation of highly toxic heavy metals such as mercury and arsenic. In short, this is the first investigation to open up the resistance mechanism of mer (2.8 kb) and ars (2.7 kb) operons in the chromosome of P. mexicana. The existence of mer operon might be derived via horizontal gene transfer from other bacterial communities, because of the linkage of mobile genetic elements (IS2 and int1) in both terminuses. The intrinsic occurrence of ars operon could be rendered vertically from its ancestral Pseudoxanthomonas species. This intrinsic ars operon was highly diversified and evolved from its ancestral species, which might be due to their mutations, genetic reshuffling, and recombination with other bacterial communities. Previously, strain GTZY was isolated from an aerobic wastewater reactor and characterized for its biochemical, physiological, and phylogeny properties. Further, the isolated strain GTZY showed its effective removal (> 63%) of As(V) and As(III) in 72 h, and its respective arsC gene encoding arsenic reductase was confirmed by the existence of merA gene (495 bp) using target-PCR. Arsenic resistance was observed up to the level of > 175 and > 55 mM, respectively. Meanwhile, the strain removed > 84% of inorganic Hg(II) in 48 h and its reduction into gaseous Hg(0) was exposed by X-ray film method. In addition, it was confirmed by the presence of merA gene (550 bp) encoding mercuric ion reductase. It is thereby claimable that the strain GTZY could be a valuable addition to the Hg and As-contaminated sites for time-effective remediation.
Data availability
The authors confirm that the data supporting the findings of this study are available within the article and on the public database (NCBI-GenBank). The extra information can be obtained by emailing the corresponding author, upon reasonable request.
References
Agarwal M, Rathore RS, Jagoe C, Chauhan A (2019) Multiple lines of evidences reveal mechanisms underpinning mercury resistance and volatilization by Stenotrophomonas sp. MA5 isolated from the Savannah River Site (SRS), USA. Cells 8:309
Ahn Y, Lee UJ, Lee Y-J et al (2019) Oligotrophic media compared with a tryptic soy agar or broth for the recovery of Burkholderia cepacia complex from different storage temperatures and culture conditions. J Microbiol Biotechnol 29:1495–1505
Alabssawy AN, Hashem AH (2024) Bioremediation of hazardous heavy metals by marine microorganisms: a recent review. Arch Microbiol 206. https://doi.org/10.1007/s00203-023-03793-5
Astuti DI, Purwasena IA, Putri RE et al (2019) Screening and characterization of biosurfactant produced by Pseudoxanthomonas sp. G3 and its applicability for enhanced oil recovery. J Pet Explor Prod Technol 9:2279–2289
ATSDR (2018) Division of toxicology and human health sciences (DTHHS). https://www.atsdr.cdc.gov/dthhs/index.html. Accessed 14 Feb 2024
Bala S, Garg D, Thirumalesh BV et al (2022) Recent strategies for bioremediation of emerging pollutants: a review for a green and sustainable environment. Toxics 10:484
Batbatan C, Rosana AR, Fernandez K et al (2022) Screening, characterization, and isolation of pigments from bacteria in mesophotic depths of the Benham Bank Seamount, Philippine rise region. Philipp J Sci 151. https://doi.org/10.56899/151.02.06
Ben Fekih I, Zhang C, Li YP et al (2018) Distribution of arsenic resistance genes in prokaryotes. Front Microbiol 9:2473
Biswas B, Chakraborty A, Sarkar B, Naidu R (2017) Structural changes in smectite due to interaction with a biosurfactant-producing bacterium Pseudoxanthomonas kaohsiungensis. Appl Clay Sci 136:51–57
Boyd ES, Barkay T (2012) The mercury resistance operon: from an origin in a geothermal environment to an efficient detoxification machine. Front Microbiol 3:349
Briffa J, Sinagra E, Blundell R (2020) Heavy metal pollution in the environment and their toxicological effects on humans. Heliyon 6:e04691
Chen J, Bhattacharjee H, Rosen BP (2015a) ArsH is an organoarsenical oxidase that confers resistance to trivalent forms of the herbicide monosodium methylarsenate and the poultry growth promoter roxarsone. Mol Microbiol 96:1042–1052
Chen J, Madegowda M, Bhattacharjee H, Rosen BP (2015b) ArsP: a methylarsenite efflux permease. Mol Microbiol 98:625–635
Chen J, Yoshinaga M, Garbinski LD, Rosen BP (2016) Synergistic interaction of glyceraldehydes-3-phosphate dehydrogenase and ArsJ, a novel organoarsenical efflux permease, confers arsenate resistance. Mol Microbiol 100:945–953
Choi EJ, Jin HM, Lee SH et al (2013) Comparative genomic analysis and benzene, toluene, ethylbenzene, and o-, m-, and p-xylene (BTEX) degradation pathways of Pseudoxanthomonas spadix BD-a59. Appl Environ Microbiol 79:663–671
Christakis CA, Barkay T, Boyd ES (2021) Expanded diversity and phylogeny of mer genes broadens mercury resistance paradigms and reveals an origin for MerA among thermophilic Archaea. Front Microbiol 12. https://doi.org/10.3389/fmicb.2021.682605
Dash HR, Basu S, Das S (2017) Evidence of mercury trapping in biofilm-EPS and mer operon-based volatilization of inorganic mercury in a marine bacterium Bacillus cereus BW-201B. Arch Microbiol 199:445–455
Dey U, Chatterjee S, Mondal NK (2016) Isolation and characterization of arsenic-resistant bacteria and possible application in bioremediation. Biotechnol Rep (amst) 10:1–7
Dziurzynski M, Gorecki A, Pawlowska J et al (2023) Revealing the diversity of bacteria and fungi in the active layer of permafrost at Spitsbergen island (Arctic) – combining classical microbiology and metabarcoding for ecological and bioprospecting exploration. Sci Total Environ 856:159072
Eckley CS, Gilmour CC, Janssen S et al (2020) The assessment and remediation of mercury contaminated sites: a review of current approaches. Sci Total Environ 707:136031
Falgenhauer L, Ghosh H, Guerra B et al (2017) Comparative genome analysis of IncHI2 VIM-1 carbapenemase-encoding plasmids of Escherichia coli and Salmonella enterica isolated from a livestock farm in Germany. Vet Microbiol 200:114–117
Freedman Z, Zhu C, Barkay T (2012) Mercury resistance and mercuric reductase activities and expression among chemotrophic thermophilic Aquificae. Appl Environ Microbiol 78:6568–6575
Ghaly TM, Chow L, Asher AJ et al (2017) Evolution of class 1 integrons: mobilization and dispersal via food-borne bacteria. PLoS One 12:e0179169
Govarthanan M, Lee S-M, Kamala-Kannan S, Oh B-T (2015) Characterization, real-time quantification and in silico modeling of arsenate reductase (arsC) genes in arsenic-resistant Herbaspirillum sp. GW103. Res Microbiol 166:196–204
Govarthanan M, Mythili R, Selvankumar T et al (2016a) Bioremediation of heavy metals using an endophytic bacterium Paenibacillus sp. RM isolated from the roots of Tridax procumbens. 3 Biotech 6:242
Govarthanan M, Shim J, Praburaman L et al (2016b) Isolation of an exopolysaccharide-producing heavy metal-resistant Halomonas sp. MG Arch Microbiol 198:205–209
Govarthanan M, Mythili R, Selvankumar T et al (2018) Myco-phytoremediation of arsenic- and lead-contaminated soils by Helianthus annuus and wood rot fungi, Trichoderma sp. isolated from decayed wood. Ecotoxicol Environ Saf 151:279–284
Govarthanan M, Mythili R, Kamala-Kannan S et al (2019) In-vitro bio-mineralization of arsenic and lead from aqueous solution and soil by wood rot fungus, Trichoderma sp. Ecotoxicol Environ Saf 174:699–705
Helmy Q, Kardena E (2024) Enhancing field-scale bioremediation of weathered petroleum oil-contaminated soil with biocompost as a bulking agent. Case Stud Chem Environ Eng 9:100735
Hou D, O’Connor D, Igalavithana AD et al (2020) Metal contamination and bioremediation of agricultural soils for food safety and sustainability. Nat Rev Earth Environ 1:366–381
Jebeli MA, Maleki A, Amoozegar MA et al (2017) Bacillus flexus strain As-12, a new arsenic transformer bacterium isolated from contaminated water resources. Chemosphere 169:636–641
Jeyakumar P, Debnath C, Vijayaraghavan R, Muthuraj M (2022) Trends in bioremediation of heavy metal contaminations. Environ Eng Res 28:220631–220630
Jobby R, Jha P, Yadav AK, Desai N (2018) Biosorption and biotransformation of hexavalent chromium [Cr(VI)]: a comprehensive review. Chemosphere 207:255–266
Khalid S, Shahid M, Natasha et al (2020) Heavy metal contamination and exposure risk assessment via drinking groundwater in Vehari, Pakistan. Environ Sci Pollut Res Int 27:39852–39864
Kittiwongwattana C, Thawai C (2016) Pseudoxanthomonas helianthi sp. nov., isolated from roots of Jerusalem artichoke (Helianthus tuberosus). Int J Syst Evol Microbiol 66:5034–5038
Krout IN, Scrimale T, Vorojeikina D et al (2022) Organomercurial lyase (MerB)-mediated demethylation decreases bacterial methylmercury resistance in the absence of mercuric reductase (MerA). Appl Environ Microbiol 88. https://doi.org/10.1128/aem.00010-22
Li J, Mandal G, Rosen BP (2016) Expression of arsenic resistance genes in the obligate anaerobe Bacteroides vulgatus ATCC 8482, a gut microbiome bacterium. Anaerobe 39:117–123
Loganathan P, Myung H, Muthusamy G et al (2015) Effect of heavy metals on acdS gene expression in Herbaspirillium sp. GW103 isolated from rhizosphere soil. J Basic Microbiol 55:1232–1238
Lu Z, Sun W, Li C et al (2019) Bioremoval of non-steroidal anti-inflammatory drugs by Pseudoxanthomonas sp. DIN-3 isolated from biological activated carbon process. Water Res 161:459–472
Luo H, Wang Q, Liu Z et al (2020) Potential bioremediation effects of seaweed Gracilaria lemaneiformis on heavy metals in coastal sediment from a typical mariculture zone. Chemosphere 245:125636
Mahbub KR, Krishnan K, Megharaj M, Naidu R (2016) Bioremediation potential of a highly mercury resistant bacterial strain Sphingobium SA2 isolated from contaminated soil. Chemosphere 144:330–337
Mahbub KR, Krishnan K, Naidu R, Megharaj M (2017) Mercury remediation potential of a mercury resistant strain Sphingopyxis sp. SE2 isolated from contaminated soil. J Environ Sci (china) 51:128–137
Matsui K, Yoshinami S, Narita M et al (2016) Mercury resistance transposons in Bacilli strains from different geographical regions. FEMS Microbiol Lett 363:fnw013
Møller AK, Barkay T, Abu Al-Soud W et al (2011) Diversity and characterization of mercury-resistant bacteria in snow, freshwater and sea-ice brine from the High Arctic. FEMS Microbiol Ecol 75:390–401
Møller AK, Barkay T, Hansen MA et al (2014) Mercuric reductase genes (merA) and mercury resistance plasmids in High Arctic snow, freshwater and sea-ice brine. FEMS Microbiol Ecol 87:52–63
Naguib MM, El-Gendy AO, Khairalla AS (2018) Microbial diversity of Mer operon genes and their potential rules in mercury bioremediation and resistance. Open Biotechnol J 12:56–77
Naguib MM, Khairalla AS, El-Gendy AO, Elkhatib WF (2019) Isolation and characterization of mercury-resistant bacteria from wastewater sources in Egypt. Can J Microbiol 65:308–321
Nordlander T (2014) Continuous on-line measuring of HgCl concentration -In “Outotec Mercury Removal Process.” https://www.diva-portal.org/smash/get/diva2:788177/FULLTEXT01.pdf. Accessed 14 Feb 2024
Olmedo P, Pla A, Hernández AF et al (2013) Determination of toxic elements (mercury, cadmium, lead, tin and arsenic) in fish and shellfish samples. Risk assessment for the consumers. Environ Int 59:63–72
Oregaard G, Sørensen SJ (2007) High diversity of bacterial mercuric reductase genes from surface and sub-surface floodplain soil (Oak Ridge, USA). ISME J 1:453–467
Páez-Espino AD, Durante-Rodríguez G, de Lorenzo V (2015) Functional coexistence of twin arsenic resistance systems in Pseudomonas putida KT2440. Environ Microbiol 17:229–238
Pepi M, Volterrani M, Renzi M et al (2007) Arsenic-resistant bacteria isolated from contaminated sediments of the Orbetello Lagoon, Italy, and their characterization. J Appl Microbiol 103:2299–2308
Purwasena IA, Astuti DI, Utami SG (2020) Nitrogen optimization on rhamnolipid biosurfactant production from Pseudoxanthomonas sp. G3 and its preservation techniques. Sains Malays 49:2119–2127
Qin G, Niu Z, Yu J et al (2021) Soil heavy metal pollution and food safety in China: effects, sources and removing technology. Chemosphere 267:129205
Rahman MA, Hassler C (2014) Is arsenic biotransformation a detoxification mechanism for microorganisms? Aquat Toxicol 146:212–219
Raj D, Maiti SK (2019) Sources, toxicity, and remediation of mercury: an essence review. Environ Monit Assess 191:566
Ramírez-Díaz MI, Díaz-Magaña A, Meza-Carmen V et al (2011) Nucleotide sequence of Pseudomonas aeruginosa conjugative plasmid pUM505 containing virulence and heavy-metal resistance genes. Plasmid 66:7–18
Rani A, Sharma A, Adak T, Bhatnagar RK (2010) Pseudoxanthomonas icgebensis sp. nov., isolated from the midgut of Anopheles stephensi field-collected larvae. J Microbiol 48:601–606
Rebelo FM, Caldas ED (2016) Arsenic, lead, mercury and cadmium: toxicity, levels in breast milk and the risks for breastfed infants. Environ Res 151:671–688
Satyapal GK, Mishra SK, Srivastava A et al (2018) Possible bioremediation of arsenic toxicity by isolating indigenous bacteria from the middle Gangetic plain of Bihar, India. Biotechnol Rep (amst) 17:117–125
Selvaraj G-K, Tian Z, Zhang H et al (2018) Culture-based study on the development of antibiotic resistance in a biological wastewater system treating stepwise increasing doses of streptomycin. AMB Express 8:12
Selvaraj G-K, Wang H, Zhang Y et al (2022) Class 1 In-Tn5393c array contributed to antibiotic resistance of non-pathogenic Pseudoxanthomonas mexicana isolated from a wastewater bioreactor treating streptomycin. Sci Total Environ 821:153537
Shen Z, Luangtongkum T, Qiang Z et al (2014) Identification of a novel membrane transporter mediating resistance to organic arsenic in Campylobacter jejuni. Antimicrob Agents Chemother 58:2021–2029
Sotero-Martins A, de Jesus MS, Lacerda M et al (2008) A conservative region of the mercuric reductase gene (merA) as a molecular marker of bacterial mercury resistance. Braz J Microbiol 39:307–310
Suhadolnik MLS, Salgado APC, Scholte LLS et al (2017) Novel arsenic-transforming bacteria and the diversity of their arsenic-related genes and enzymes arising from arsenic-polluted freshwater sediment. Sci Rep 7:11231
Tahir MA, Rasheed H, Malana A (2012) Method development for arsenic analysis by modification in spectrophotometric technique. Drink Water Eng Sci 5:1–8
Talwar MP, Ninnekar HZ (2015) Biodegradation of pesticide profenofos by the free and immobilized cells of Pseudoxanthomonas suwonensis strain HNM. J Basic Microbiol 55:1094–1103
Taran M, Fateh R, Rezaei S, Gholi MK (2019) Isolation of arsenic accumulating bacteria from garbage leachates for possible application in bioremediation. Iran J Microbiol 11:60–66
Thierry S, Macarie H, Iizuka T et al (2004) Pseudoxanthomonas mexicana sp. nov. and Pseudoxanthomonas japonensis sp. nov., isolated from diverse environments, and emended descriptions of the genus Pseudoxanthomonas Finkmann et al. 2000 and of its type species. Int J Syst Evol Microbiol 54:2245–2255
Wang L, Jeon B, Sahin O, Zhang Q (2009) Identification of an arsenic resistance and arsenic-sensing system in Campylobacter jejuni. Appl Environ Microbiol 75:5064–5073
Wang G, Zhao Y, Gao H et al (2013) Co-metabolic biodegradation of acetamiprid by Pseudoxanthomonas sp. AAP-7 isolated from a long-term acetamiprid-polluted soil. Bioresour Technol 150:259–265
Wang L, Hou D, Cao Y et al (2020) Remediation of mercury contaminated soil, water, and air: a review of emerging materials and innovative technologies. Environ Int 134:105281
Watters M, Rayman J (2014) Agency for Toxic Substances and Disease Registry’s don’t mess with mercury initiative. J Environ Health 76:34–36
Weon H-Y, Kim B-Y, Kim J-S et al (2006) Pseudoxanthomonas suwonensis sp. nov., isolated from cotton waste composts. Int J Syst Evol Microbiol 56:659–662
Wuana RA, Okieimen FE (2011) Heavy metals in contaminated soils: a review of sources, chemistry, risks and best available strategies for remediation. ISRN Ecol 2011:1–20
Yang H-C, Rosen BP (2016) New mechanisms of bacterial arsenic resistance. Biomed J 39:5–13
Yang H-C, Cheng J, Finan TM et al (2005) Novel pathway for arsenic detoxification in the legume symbiont Sinorhizobium meliloti. J Bacteriol 187:6991–6997
Yang Y, Wu S, Lilley RM, Zhang R (2015) The diversity of membrane transporters encoded in bacterial arsenic-resistance operons. PeerJ 3:e943
Zhang Z, Yin N, Cai X et al (2016) Arsenic redox transformation by Pseudomonas sp. HN-2 isolated from arsenic-contaminated soil in Hunan, China. J Environ Sci (china) 47:165–173
Acknowledgements
The authors express their gratitude to King Saud University, Researchers Supporting Project (No. RSPD2024R670), King Saud University, Riyadh, Saudi Arabia. We thank Prof. Yang Min and Prof. Yu Zhang (RCEES, CAS Institute, Beijing, China) for kindly providing us P. mexicana strain GTZY.
Funding
The work was supported by the Ministry of Education and State Administration of Foreign Experts Affairs (Grant No. G20190011010), People’s Republic of China. Also, the current work was supported by the SPIHER-Seed money project (Project No. SPIHER/Seed Money/2023–24/06).
Author information
Authors and Affiliations
Contributions
Conceptualization; methodology; software using; data analysis; writing—original draft preparation: Abdul Raheem Nelofer. Project administration, conceptualization: Selvaraj Ganesh-Kumar (corresponding author) and S. Balachandran. Software using: Soorangkattan Saravanan. Software using, data analysis: Karuppanan Kalimuthu. Software using, data analysis: Ganesan Govindarajan. Data analysis: Rajah Dilip Kumar, Shivani Ramamurthy Baluraj. Data analysis, software using, data analysis: Imran Hasan. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
All persons who meet authorship criteria are listed as authors.
Consent for publication
All authors certify that they have participated sufficiently in the work to take public responsibility for the content. Furthermore, each author certifies that this material or similar material has not been and will not be submitted to or published in any other publication.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Robert Duran
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Highlights
• First detailed heavy metals bioremediation study in the genus, Pseudoxanthomonas.
• Isolate has great potential to resist, reduce and remove the toxic Hg(II) and As(V).
• An acquired mobile elements linked mer gene operon was observed in its chromosome.
• An intrinsic ars gene operon was extremely diverse from its ancestral species.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Abdul Raheem, N., Selvaraj, GK., Karuppanan, K. et al. Bioremediation of heavy metals by an unexplored bacterium, Pseudoxanthomonas mexicana strain GTZY isolated from aerobic-biofilm wastewater system. Environ Sci Pollut Res (2024). https://doi.org/10.1007/s11356-024-34602-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11356-024-34602-1