Abstract
Submerged macrophytes and microbial communities are important parts of lake ecosystems. In this study, the bacterial community composition in rhizosphere sediments and water from areas cultivated with (PL) and without (CK) shining pondweed (Potamogeton lucens Linn.) was investigated to determine the effects of P. lucens Linn. on the structure of the bacterial communities in Nansi Lake, China. Molecular techniques, including Illumina MiSeq and qPCR targeting of the 16S rRNA gene, were used to analyze the composition and abundance of the bacterial community. We found that bacterial alpha diversity was higher in PL water than in CK water, and the opposite trend was observed in sediment. In addition, 16S rRNA gene copy number in sediment was lower in PL than in CK. We found 30 (e.g., Desulfatiglans) and 29 (e.g., Limnohabitans) significantly different genera in sediment and water, respectively. P. lucens Linn. can change chemical properties in sediment and water and thereby affect the bacterial community. At the genus level, members of bacterial community clustered according to source (water/sediment) and area (PL/CK). Our study demonstrated that submerged macrophytes can affect the bacterial community composition in both sediment and water, suggesting that submerged macrophytes affect the transportation and cycling of nutrients in lake ecosystems.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
For decades, water eutrophication has been a major source of freshwater pollution in China (Wang et al. 2019) and has caused severe problems such as biodiversity loss and algal blooms (Yin et al. 2020). During the rainy season, agricultural systems release abundant nitrogen (N) and other nutrients into water bodies; therefore, organisms in these water bodies often experience high nutrient supplies (Yan et al. 2018). As the main primary producers of wetland ecosystems, plants (Zhang et al. 2016; Zhao et al. 2019), especially submerged macrophytes, are important participants in lake ecosystems and play an important role in nutrient removal (Qin et al. 2019) and other processes in constructed and natural water bodies. Submerged macrophytes can assimilate a variety of nutrients, such as organic and inorganic N (i.e., NO3−, NO2−, NH3, and NH4+) and phosphorus (P), which can result in chronic toxicity to Hydrilla verticillata at high concentrations (Wang et al. 2010). Yan et al. (2018) reported that P. malaianus, Vallisneria natans, and Hydrilla verticillata had strong nutrient removal effects, indicating that nutrients were easily assimilated by these aquatic plants. Therefore, submerged macrophytes are important for water self-purification systems and for maintaining ecological balance (Han et al. 2018).
Microbial communities are important parts of lake ecosystems (Yan et al. 2018) and play an important role in regulating the water quality of polluted lakes (Zhang et al. 2018). Microbial communities also play vital roles in biogeochemical cycling in the sediments of freshwater lakes (Liu and Yang 2020). Changes in microbial communities may reflect the status of the environment (e.g., water quality) (Liu et al. 2020). Excess N and P in wetlands can be removed through biological, physical, and chemical processes (Ballantine et al. 2014). Sediment N cycling is an important biological process for permanent N removal (Wu et al. 2021). Submerged macrophytes can provide oxygen and appropriate environmental conditions for epiphytic bacterial communities (Bustamante et al. 2011). Wu et al. (2021) reported the direct effects of submerged macrophytes on the bacterial community and their indirect effects through altering sediment C and concluded that a greater development of submerged macrophytes in lakes is associated with greater nitrogen removal from lake sediments. Furthermore, at night, respiration of submerged macrophytes may shift from aerobic to anaerobic because the conditions at night are favorable for anaerobic bacteria (Eriksson 1999). Changes in microbial communities can reflect the stability of effluent and sediment ecosystems. However, it is not clear how the cultivation of submerged plants for purifying water affects the bacterial community in rhizosphere sediments and nearby water.
Nansi Lake, which is located in Shandong Province, covers an area of 1266km2 and is the largest freshwater lake in northern China (34°27′–35°20′N, 116°34′–117°21′E) (Tian et al. 2013); as the main reservoir lake and biodiversity protection area in the east route of the South to North Water Diversion Project, it has an important impact on water quality (Zhang et al. 2021). P. lucens Linn. is an important submerged plant in Nansi Lake and has a good water purification effect. However, how the bacterial communities in water and rhizosphere sediments differ in areas cultivated with P. lucens Linn. (PL) from those in control areas without P. lucens cultivation (CK) remains unclear. In this study, we collected 24 water and sediment samples from PL and CK areas to study the effects of this plant on water quality, nutrients and the microbial community.
Materials and methods
Site description and sampling
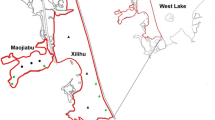
The sampling site was located in Nansi Lake (34°37′N, 117°12′E and altitude 27 m) in Jining, Shandong Province, China (Fig. 1). This region has a temperate monsoon climate with an average annual temperature of 15 °C and a mean annual precipitation of 775 mm. With the development of industry and the increased application of pesticides along the lake area, the industrial and agricultural wastewater and domestic sewage discharged to Nansi Lake are increasing yearly. In 2002, water quality was inferior to class V, which corresponds to “polluted” and “dirty” (Kondrat’eva et al 2009), according to the “China surface water quality standard” (GB3838–2002). It thus has great impacts on agriculture, fisheries, and the domestic water supply. Near Wanzhuang Village (34°37′N, 117°12′E) in Nansi Lake, there is a large area (approximately 1 ha) where only P. lucens Linn. is cultivated (PL) that appears to be in a clear state and an adjacent area (approximately 500 m away) in which the water is muddy and lacks any aquatic plants (CK). P. lucens Linn. was morphologically identified by Dr. Fengyue Shu using taxonomic keys, and voucher specimens were deposited at Qufu Normal University (School of Life Sciences), China. The time of sampling was November 2020, when P. lucens Linn. was in the declining phase. We randomly established three plots (approximately 10 × 10 m) in each of the two areas and selected two sampling sites in each plot. For water samples, 3.0 L water (20 cm depth) was collected at each site and immediately transported to the laboratory at low temperature 0–4 °C. Then, the 1.5 L of each water samples were filtered through 0.22-μm membrane filters (Millipore, USA), and the filtrate was stored at − 80 °C until DNA extraction. The remaining 1.5 L water sample was used for physicochemical analysis (Chao et al. 2021). For sediment samples, we collected surface sediment (0–20 cm) by a Peterson dredger. The rhizosphere sediment was collected from the sediment adhering to the root crowns, where rooting was so dense that all sediment was determined to be under the influence of roots. Sediment was collected by removing a randomly selected plant and associated root crowns to a depth of 20 cm, lightly shaking the plant to remove sediment not associated with roots and then collecting soil attached to roots (Zhou and Fong 2021), All of the sediments from each site were screened, mixed, and packed in polyethylene bags and transported to the laboratory on ice. These samples were collected in the wild environment of Wanzhuang Village, and we obtained permission from the director of the village for our collection. In total, we collected 24 sediment samples [2 individuals × 3 plots × 2 sample sources (water/sediment) × 2 area types (PL/CK)]. The sediments were sieved through a 1.0-mm sieve and stored at –80 °C for further molecular analysis.
Chemical characteristics
For chemical characterization, water samples were filtered through a 0.22-μm microporous filtering film, and sediment samples were air-dried at room temperature and sieved through a 1-mm screen. The pH was determined using a glass combination electrode (Li et al. 2013). The total nitrogen (TN) was determined according to potassium persulfate oxidation-UV spectrophotometry. KCl-extractable NO3− and NH4+ were determined by extraction with 2 M KCl, steam distillation, and titration (Mulvaney 1996). The organic matter (OM) and total potassium (TK) were determined by Nanjing Agricultural University. The total phosphorus (TP) was determined using the perchloric acid-sulfuric acid method (Hedley and Stewart 1982). The content of PO43− in water was analyzed by resin extraction following a protocol modified from Hedley and Stewart (1982).
Total community DNA extraction
Total DNA was extracted from 0.25 g of sediment or microporous filtering film using the Power Soil DNA Isolation Kit (MOBIO Laboratories Inc., Carlsbad, CA, USA) according to the manufacturer’s instructions. The DNA concentration and purity (A260/A280) of the extracts were estimated using a NanoDrop 2000/2000c spectrophotometer. High-quality DNA was stored at –80 °C for subsequent experiments.
Quantitative PCR (qPCR) analysis
The abundance of the bacterial 16S rRNA gene was quantified using a CFX96™ real-time PCR detection system (Bio-Rad, Hercules, CA). The reaction mixture (20 μL) contained FastFire qPCR PreMix (SYBR Green) (Vazyme, China), 10 nM of each primer, ROX Passive Reference Dye, and 1 μL of DNA. Bacterial assays used the primers 515FmodF and 806RmodR (Zhou and Fong 2021) and the following thermal program: 95 °C for 1 min followed by 40 cycles of 95 °C for 10 s and 60 °C for 30 s (Lauber et al. 2013). The standard for measuring the quantity of the 16S rRNA gene was developed from a clone with the correct insert. Plasmid DNA was prepared from the clone using a FastPure Plasmid Mini kit (Vazyme, Nanjing, China).The R2 of the standard curve was > 0.99. The qPCRs were run in quadruplicate with the DNA extracted from each sample.
Pyrosequencing and bioinformatics processing
The primers 515FmodF and 806RmodR (Zhou and Fong 2021) were used to amplify the V4 hypervariable region of the bacterial 16S rRNA gene. The PCR products were sequenced on the Illumina MiSeq PE 300 platform of Majorbio Pharm Technology Co., Ltd. (Shanghai, China). The obtained sequences were submitted to the NCBI Sequence Read Archive under the accession number PRJNA716102.
Paired-end reads were processed using Quantitative Insights into Microbial Ecology (QIIME) software, and presumptive chimeric sequences were screened and discarded using UCHIME (Zhou and Fong 2021). The original sequence data were separated, and the primers were removed (Martin 2011). According to the reading quality profile, the forward reading of the 16S rRNA gene was truncated to 240 bp, and the reverse reading was truncated to 160 bp (Schmidt et al. 2019). All reads were filtered and trimmed using the parameters maxEE = 2 and truncQ = 2. High-quality sequences were refined and resampled according to the lowest number of reads in the sample. To minimize the possibility of retaining OTUs due to sequencing errors, we deleted OTUs if (1) there were fewer than 5 sequences in less than 3 samples in each group or (2) the total number of sequences in all samples was less than 20 using the Silva database (132nd edition; http://www.arb-silva.de). All sequences matching “chloroplast” and “mitochondria” were removed from the dataset.
Statistical analysis
The concentrations of chemical characteristics among samples were determined using one-way analysis of variance (ANOVA), and paired comparisons of treatment means were achieved by Tukey’s procedure at P < 0.05 using SPSS BASE ver. 19.1 statistical software (SPSS, Chicago, IL, USA) (Ahn et al. 2012). Redundancy analysis (RDA) was performed using CANOCO 5.0 to identify the relationships between the bacterial communities and chemical characteristics (Zhang et al. 2015). A phylogenetic tree of the genera showed significant differences between PL and CK using the neighbor-joining method in MEGA v.6.0 and displayed using iTOL (Interactive Tree Of Life, https://itol.embl.de/) (Zhou and Fong 2021).
Results
Chemical characteristics of water and sediment
The chemical characteristics of the sediment and overlying water are shown in Table 1. There was no obvious difference in NH4+, TP, or PO43− in water among the sampling sites. The water concentrations of NO3− and NO2− were significantly lower in PL than in CK. There was no obvious difference in sediment NO3−, NH4+, OM, or TP between PL and CK. The sediment concentration of TK was lower in PL (7.367 g kg−1) than in CK (12.167 g kg−1).
Effects of P. lucens Linn. on the abundance of the 16S rRNA gene
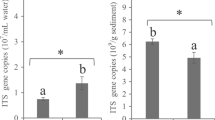
The abundance of the 16S rRNA gene in sediment samples ranged from 1.1 × 108 to 8.2 × 108 copies/g sediment and in water samples ranged from 3.1 × 104 to 1 × 105 copies/mL water (Fig. 2). The abundance of the 16S rRNA gene in sediment samples was significantly lower (P < 0.05) in PL than in CK, whereas the opposite trend was observed in water samples.
Bacterial alpha diversity
A total of 2,879 OTUs (24.97% of the total 11,532) were obtained from the 24 samples. There was a mean of 46,220 classifiable sequences per sample used in the subsequent analysis, with a mean read length of 253 bp. The Good’s coverage values were in the range of 0.96–0.99 at a 97% similarity cutoff, indicating that the current numbers of sequence reads were sufficient for capturing the bacterial diversity in the samples.
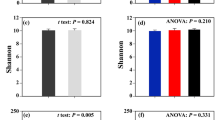
The alpha-diversity indices in the four groups are shown in Table 2. There were significant differences (P < 0.05) Simpson, Chao1, Shannoneven, and Simpsoneven index measures between PL and CK. The Simpson and Chao1 indices were higher, and Shannoneven and Simpsoneven indices were lower in CK water than in water with PL. Furthermore, Ace and Chao1 indices were lower in sediment with PL than in CK sediment. There was no significant difference in the Shannon or Ace index of the bacterial community in water or sediment between PL and CK (P > 0.05).
Phylogenetic analysis of the bacterial community in water and sediment
A total of 702 bacterial and archaeal genera were identified in the 24 samples. At the genus level, strong clustering of bacterial (Fig. 3) communities according to sample source (water/sediment) and area type (PL/CK) was revealed. Samples from sediment clustered into one branch and were divided into group I and group II; samples from water clustered into another branch and were divided into group III and group IV (Fig. 3).
Variation in bacterial communities between water and sediment
A total of 30 genera differentially contributing to the bacterial community in sediment between PL and CK were identified via iTOL, including 17 genera which were significantly more abundant in PL than in CK and 13 genera which showed the opposite trend (Fig. 4a, Table S1).
Bacterial genera statistically different between PLS and CKS (a) and PLW and CKW (b). Colored circles represent the relative abundance of each genus. Taxonomic dendrogram shows the inferred evolutionary relationship of the enriched microbiota of each sample. Total relative abundances of all genera and significant effects across organic and conventional managements are listed in Table S1 and Table S2. PLW, water samples in Potamogeton lucens Linn.; CKW, water samples in CK; PLS, sediment samples in Potamogeton lucens Linn.; CKS, sediment samples in CK
A total of 29 genera differentially contributing to the bacterial community in water between PL and CK were identified via iTOL, including 26 genera which were significantly more abundant in PL than in CK (e.g., Limnohabitans, Algoriphagus, Dinghuibacter) and 3 genera (e.g., Omnitrophus, Terrimonas) which showed the opposite trend (Fig. 4b, Table S2).
Environmental factors influencing bacterial communities’ structure
The RDA results are shown in Fig. 5. The first two axes together explained 84.07% (Fig. 5a) and 84.62% (Fig. 5b) of the variance in bacterial community structure in the sediment and water samples, respectively. The distances between the PL and CK samples were large, indicating that bacterial community structure significantly differed between CK and PL in both sediment (Fig. 5a) and water (Fig. 5b). In the water samples, the diversity of the bacterial community was positively correlated with the TP (P = 0.033) content in PL; such a pattern was not observed in CK (Fig. 5b).
RDA of bacterial communities and environmental factor for individual samples. Environmental factor include TP (total phosphorus), NH4+ (concentration of NH4+), NO3− (concentration of NO3−), TN (total nitrogen), OM (organic matter), and TK (total potassium). a Sediment samples and b water samples. PLW, water samples in Potamogeton lucens Linn.; CKW, water samples in CK; PLS, sediment samples in Potamogeton lucens Linn.; CKS, sediment samples in CK
The bacterial community in PL was affected by TK and NO3− (Fig. 5a). In CK sediment samples, OM and TK were primarily distributed in the same group of taxa in the bacterial communities (Fig. 5a).
Discussion
P. lucens Linn. decreased the concentrations of NO3 − and NO2 − in water and TK in sediment
In this study, P. lucens Linn. removed nutrients from the water, significantly decreasing the concentrations of NO3− and NO2− (P < 0.05) (Table 1). This founding is consistent with the previous finding that P. lucens Linn. can remove N from water (Huo et al. 2010). However, P. lucens Linn. did not substantially reduce N in sediment, which may have resulted from the effect of root exudation on bacterial communities, and P. lucens Linn. was in a declining phase (Yin et al. 2020; He et al. 2020). Furthermore, P. lucens Linn. did not reduce the concentration of TP in water or sediment, which may have been due to the fact that P. lucens Linn. was in a declining phase (He et al. 2020), and their yellow leaves may release more P into the water when we collected samples. Zhang et al. (2019) found that rising temperature significantly increased the growth of P. lucens Linn; we collected our samples in late autumn, so the performance of these plants was reduced. Jin et al. (2017) confirmed that the synergistic purification effect of P. maackianus and four other macrophytes was much greater than the individual uptake effects in water purification. Most likely, the slightly weaker water purification capacity observed in our study was due to the presence of only the single submerged macrophyte species at our sampling sites. Liu and Chen (2018) similarly demonstrated that single plant types show poorer purification effects than several submerged macrophytes in lake systems.
Significant correlation between 16S rRNA gene copy number and P. lucens Linn.
The significant correlation between 16S rRNA gene copy number and PL also indicated that P. lucens Linn. could have a substantial effect on the population sizes of bacteria. The concentration of NO3− in the CK water samples was decreased by 58 times compared with that in the PL water samples. These results may demonstrate that P. lucens Linn. plays an important role in N removal (i.e., NO3− and NO2−) in wetlands (Chang et al. 2006) and that N removal stimulated the growth of some bacterial taxa and subsequently increased the number of 16S rRNA gene copies (Yan et al. 2018). In sediment, the number of 16S rRNA gene copies was obviously different between PL and CK. This result is consistent with the finding that the number of bacteria was negatively correlated with the total organic acid concentration secreted from the roots of P. maackianus, a congener of P. lucens Linn. (Yin et al. 2020).
P. lucens Linn. increased bacterial alpha-diversity in water but decreased it in sediment
Our results showed that P. lucens Linn. can improve bacterial evenness (Shannoneven, Simpsoneven) and decrease bacterial richness (Chao1 index) in water (Table 2). The lower Chao1 index in PL water was likely due to the fact that sampling was conducted during the declining period (November) of this plant (He et al. 2020). A previous study indicated that P. maackianus can release organic acids (Yin et al. 2020). The amount of organic acids has been indicated to be negatively correlated with the diversity of DNA-based bacterial communities (Weisskopf et al. 2008). In contrast, higher organic acid root exudation from some plants (e.g., soybean) has been shown to increase the diversity of the microbial community (Yang et al. 2012). This inconsistency may be due to the different types and amounts of organic acids produced by different plants. In this study, P. lucens Linn. may have produced similar organic acids, leading to a locally weakly acidic environment in the rhizosphere sediment unconducive to the survival of some acid-sensitive bacteria, thus reducing the alpha-diversity of bacterial community. However, this hypothesis needs to be tested in future experiments by in situ GC–MS and related metagenomics techniques to investigate the composition of root exudates.
P. lucens Linn. changed the bacterial community composition in water and sediment
The differences between PL and CK in microbial community structure in water may be explained by the lower concentrations of NO3− and NO2− in PL water than in CK water (Table 1). These results are consistent with previous studies (He et al. 2007; Jorquera et al. 2014). In PL water, there was a higher abundance of Limnohabitans, which are aerobic anoxygenic phototrophs that can supplement their mostly heterotrophic metabolism with harvested light energy (Kasalický et al. 2018). This result may suggest that P. lucens Linn. can purify and improve the light transmittance of the surrounding water, leading to an increase in the number of such microorganisms. Han et al. (2019) reported that Chloroflexi and Bacteroidota played a dominant role when P. malaianus was in the decline period, and we found that the abundances of the phyla Chloroflexi and Bacteroidota (0.04%) were higher in PL water than in CK water (Fig. 4b). These results suggest that P. lucens Linn. may provide suitable conditions for the growth and reproduction of microorganisms in Chloroflexi and Bacteroidota. Desulfatiglans and Ignavibacterium have been shown to contribute to methane oxidation by nitrite and sulfate reduction (Jochum et al. 2018), and Sulfuritalea plays an important role in the degradation of aromatic pollutants (Sperfeld et al. 2019). The higher proportions of these three taxa in PL sediment than in CK sediment indicated that P. lucens significantly promoted carbon metabolism in its rhizosphere sediments. We found higher proportions of Cyanobacteria and Firmicutes, which are abundant under heavy metal stress (Huang et al. 2020), in CK sediment than in PL sediment.
Responses of the bacterial community to environmental conditions in water and sediment
The RDA results showed that N and P were the most important factors related to bacterial community structure. Hu et al. (2020) reported that the concentrations of NH4+ and NO3− were two important factors affecting the abundances of anammox bacteria and denitrifying bacteria, and these two microbial groups compete in many ecological environments. In the sediment samples of this study, we found that these two ions substantially affected the bacterial community, but whether these functional microorganisms are affected at the DNA and RNA levels requires experimental investigation. Yin et al. (2020) showed that the composition of the bacterial community is likely related to variation in NH4 + content and thus the rhizosphere states of aquatic plants. However, in this study, the diversity of the bacterial community in water was negatively correlated with the concentration of NH4+-N.
In this study, the sediment in the unplanted CK area had significantly higher concentrations of TP than PL sediment, and in PL sediment, the diversity of the bacterial community was positively correlated with the concentration of TP (Fig. 5a). These results were similar to those of previous studies (Chen et al. 2014; Dai et al. 2019). In Guanting Reservoir, China, TP concentration was shown to directly affect the number of phosphate-dissolving and/or phosphate-decomposing bacteria in sediment (Li et al. 2005). Furthermore, the TP concentration was found to shape the variation in bacterial community composition within two different drainage areas (Lindström and Bergström 2005). Therefore, the removal of TP from the rhizosphere by P. lucens Linn. strongly affected microorganisms involved in phosphorus metabolism and thus the entire bacterial community.
Conclusion
In this study, we showed that P. lucens Linn. has strong effects on the chemical characteristics and bacterial communities in water and rhizosphere sediment in Nansi Lake, China. P. lucens Linn. can alter the environment by affecting the quality of water, which affects the composition of the bacterial community. The results of this study clarify the effect of P. lucens Linn. in lake ecosystems, especially in structuring the composition of the bacterial community. An optimal purification effect for sewage treatment may be achieved with two or more plant types. However, the relationship between submerged macrophytes and the bacterial community require further study.
Data availability
The datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Ahn JH, Song J, Kim BY, Kim MS, Joa JH, Weon HY (2012) Characterization of the bacterial and archaeal communities in rice field soils subjected to long-term fertilization practices. J Microbiol 50:754–765. https://doi.org/10.1007/s12275-012-2409-6
Ballantine KA, Groffman PM, Lehmann J, Schneider RL (2014) Stimulating nitrate removal processes of restored wetlands. Environ Sci Technol 48:7365–7373. https://doi.org/10.1021/es500799v
Bustamante MA, Mier MV, Estrada JA, Domíguez CD (2011) Nitrogen and potassium variation on contaminant removal for a vertical subsurface flow lab scale constructed wetland. Bioresour Technol 102:7745–7754. https://doi.org/10.1016/j.biortech.2011.06.005
Chang HQ, Yang XE, Fang YY, Pu PM, Li ZK, Rengel Z (2006) In-situ nitrogen removal from the eutrophic water by microbial-plant integrated system. J Zhejiang Univ Sci B 7:521–531. https://doi.org/10.1631/jzus.2006
Chao C, Wang L, Li Y, Yan Z, Liu H, Yu D, Liu C (2021) Response of sediment and water microbial communities to submerged vegetations restoration in a shallow eutrophic lake. Sci Total Environ 801:149701. https://doi.org/10.1016/j.scitotenv.2021.149701
Chen Y, Wen Y, Zhou Q, Vymazal J (2014) Effects of plant biomass on denitrifying genes in subsurface-flow constructed wetlands. Bioresour Technol 157:341–345. https://doi.org/10.1016/j.biortech.2014.01.137
Dai Y, Wu J, Zhong F, Cui NX, Kong LW, Liang W, Cheng SP (2019) Macrophyte identity shapes water column and sediment bacterial community. Hydrobiologia 835:71–82. https://doi.org/10.1007/s10750-019-3930-y
Eriksson PG (1999) An experimental study on effects of submersed macrophytes on nitrification and denitrification in ammonium-rich aquatic systems. Limnol Oceanogr 44:1993–1999
Ghobrial MG, Nassr HS, Kamil AW (2015) Bioactivity effect of two macrophyte extracts on growth performance of two bloom-forming cyanophytes. Egypt J Aquat Res 41:69–81. https://doi.org/10.1016/j.ejar.2015.01.001
Han B, Zhang S, Zhang L, Liu K, Yan L, Wang P, Wang C et al (2018) Characterization of microbes and denitrifiers attached to two species of floating plants in the wetlands of Lake Taihu. PLoS ONE 13:e0207443. https://doi.org/10.1371/journal.pone.0207443
Han B, Addo FG, Mu X, Zhang L, Zhang S, Lv X, Li X et al (2019) Epiphytic bacterial community shift drives the nutrient cycle during Potamogeton malaianus decomposition. Chemosphere 236:124–253. https://doi.org/10.1016/j.chemosphere.2019.06.223
He JZ, Shen JP, Zhang LM, Zhu YG, Zheng YM, Xu MG, Di H (2007) Quantitative analyses of the abundance and composition of ammonia-oxidizing bacteria and ammonia-oxidizing archaea of a Chinese upland red soil under long-term fertilization practices. Environ Microbiol 9:2364–2374. https://doi.org/10.1111/j.1462-2920.2007.01358.x
He D, Ren L, Wu QL (2020) Growing season drives the compositional changes and assembly processes of epiphytic bacterial communities of two submerged macrophytes in Taihu Lake. Fems Microbiol Ecol 96:25. https://doi.org/10.1093/femsec/fiaa025
Hedley MJ, Stewart JWB (1982) Method to measure microbial phosphate in soils. Soil Biol Biochem 14:377–385
Hu J, Zhou Y, Lei Z, Liu G, Hua Y, Zhou W, Wan X et al (2020) Effects of Potamogeton crispus decline in the rhizosphere on the abundance of anammox bacteria and nirS denitrifying bacteria. Environ Pollut 260:114018. https://doi.org/10.1016/j.envpol.2020.114018
Huang S, Song Q, Li Q, Zhang H, Luo X, Zheng Z (2020) Damage of heavy metals to Vallisneria natans (V. natans) and characterization of microbial community in biofilm. Aquat Toxicol 225:105515. https://doi.org/10.1016/j.aquatox.2020.105515
Huo YZ, He WH, Luo K, Wang YY, Zhang YJ, Tian QT, He PM (2010) Bioremediation efficiency of applying Daphnia magna and submerged plants: a case study in Dishui Lake of Shanghai. China Chin J Appl Ecol 21:495
Jin SQ, Zhou JB, Bao WH, Chen J, Li DD, Li Y (2017) Comparison of nitrogen and phosphorus uptake and water purification ability of five submerged macrophytes. Chin J Envir Sci 38:156–161. https://doi.org/10.13227/j.hjkx.201606135
Jochum LM, Schreiber L, Marshall IPG, Jørgensen BB, Schramm A, Kjeldsen KU (2018) Single-cell genomics reveals a diverse metabolic potential of uncultivated desulfatiglans-related deltaproteobacteria widely distributed in marine sediment. Front Microbiol 9:2038. https://doi.org/10.3389/fmicb.2018.02038
Jorquera MA, Martínez OA, Marileo LG, Acuna JJ, Saggar S, Mora ML (2014) Effect of nitrogen and phosphorus fertilization on the composition of rhizobacterial communities of two Chilean andisol pastures. World J Microb Biot 30:99–107. https://doi.org/10.1007/s11274-013-1427-9
Kasalický V, Zeng Y, Piwosz K, Šimek K, Kratochvilová H, Koblížek M (2018) Aerobic anoxygenic photosynthesis is commonly present within the genus Limnohabitans. Appl Environ Microbiol 84(1):e02116-17
Kondrat’eva LM, Fisher NK, Berdnikov NV (2009) Microbiological estimate of water quality in the Amur and Sungari rivers after a technogenic accident in china in 2005. Water Resour 36:548–559. https://doi.org/10.1134/S0097807809050078
Lauber CL, Ramirez KS, Aanderud Z, Lennon J, Fierer N (2013) Temporal variability in soil microbial communities across land-use types. Isme J 7:1641–1650. https://doi.org/10.1038/ismej.2013.50
Li C, Yuan HL, Huang HZ (2005) Vertical distribution of phosphorus and P-dissolving/decomposing bacteria in the sediment of Guanting reservoir. Sci China (Ser D Earth Sci) 48:285–294
Li X, Deng Y, Li Q, Lu C, Wang J, Zhang H, Zhu J et al (2013) Shifts of functional gene representation in wheat rhizosphere microbial communities under elevated ozone. Isme J 7:660–671. https://doi.org/10.1038/ismej.2012.120
Li L, Yue CL, Zhang H, Li HP, Yang L, Wang J (2019) Correlation between water purification capacity and bacterial community composition of different submerged macrophytes. Chin J Envir Sci 40:4962–4970. https://doi.org/10.13227/j.hjkx.201903265
Lindström ES, Bergström AK (2005) Community composition of bacterioplankton and cell transport in lakes in two different drainage areas. Aquat Sci 67:210–219. https://doi.org/10.1007/s00027-005-0769-2
Liu M, Chen KN (2018) Purification effect of submerged macrophyte system with different plants combinations and C/N ratios. Chin J Envir Sci 39:2706–2714. https://doi.org/10.13227/j.hjkx.201710209
Liu TT, Yang H (2020) Comparative analysis of the total and active bacterial communities in the surface sediment of Lake Taihu. Fems Microbiol Ecol 96:fiaa059. https://doi.org/10.1093/femsec/fiaa059.
Liu Z, Iqbal M, Zeng Z, Lian Y, Zheng A, Zhao M, Li Z et al (2020) Comparative analysis of microbial community structure in the ponds with different aquaculture model and fish by high-throughput sequencing. Microb Pathogenesis 142:104101. https://doi.org/10.1016/j.micpath.2020.104101
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. Embnet J 17:10–12. https://doi.org/10.14806/ej.17.1.200
Mulvaney RL (1996) Nitrogen-inorganic forms In: Sparks DL (ed) Method of soil analysis Part 3 chemical methods. Arch Agron Soil Sci:1123–1200
Qin Z, Zhao Z, Xia L, Adam A, Li Y, Chen D, Mela SM et al (2019) The dissipation and risk alleviation mechanism of PAHs and nitrogen in constructed wetlands: the role of submerged macrophytes and their biofilms-leaves. Environ Int 131:104940. https://doi.org/10.1016/j.envint.2019.104940
Schmidt JE, Kent AD, Brisson VL, Gaudin ACM (2019) Agricultural management and plant selection interactively affect rhizosphere microbial community structure and nitrogen cycling. Microbiome 7:146. https://doi.org/10.1186/s40168-019-0756-9
Sperfeld M, Diekert G, Studenik S (2019) Anaerobic aromatic compound degradation in sulfuritalea hydrogenivorans sk43H. Fems Microbiol Ecol 95. https://doi.org/10.1093/femsec/fiy199.
Sun W, Xu MY, Wu WM, Guo J, Xia CY, Sun GP, Wang AJ (2014) Molecular diversity and distribution of anammox community in sediments of the Dongjiang River, a drinking water source of Hong Kong. J Appl Microbiol 116:464–476. https://doi.org/10.1111/jam.12367
Tian C, Pei H, Hu W, Xie J (2013) Phytoplankton variation and its relationship with the environmental factors in Nansi Lake, China. Environ Monit Assess 185:295–310. https://doi.org/10.1007/s10661-012-2554-8
Wang C, Zhang SH, Wang PF, Li W, Lu J (2010) Effects of ammonium on the antioxidative response in Hydrilla verticillata (L.f.) Royle plants. Ecotoxicol Environ Saf 73:189–195. https://doi.org/10.1016/j.ecoenv.2009.08.012
Wang J, Fu Z, Qiao H, Liu F (2019) Assessment of eutrophication and water quality in the estuarine area of Lake Wuli, Lake Taihu, China. Sci Total Environ 650:1392–1402. https://doi.org/10.1016/j.scitotenv.2018.09.137
Weisskopf L, Bayon RCL, Kohler F, Page V, Jossi M, Gobat JM, Martinoia E et al (2008) Spatio-temporal dynamics of bacterial communities associated with two plant species differing in organic acid secretion: a one-year microcosm study on lupin and wheat. Soil Biol Biochem 40:1772–1780
Wu H, Hao B, Cai Y, Liu G, Xing W (2021) Effects of submerged vegetation on sediment nitrogen-cycling bacterial communities in Honghu Lake (China). Sci Total Environ 755:142541. https://doi.org/10.1016/j.scitotenv.2020.142541
Yan LY, Zhang SH, Lin D, Guo C, Yan L, Wang S, He Z (2018) Nitrogen loading affects microbes, nitrifiers and denitrifiers attached to submerged macrophyte in constructed wetlands. Sci Total Environ 622–623:121–126. https://doi.org/10.1016/j.scitotenv.2017.11.234
Yang T, Liu G, Li Y, Zhu S, Zou A, Qi J, Yang Y (2012) Rhizosphere microbial communities and organic acids secreted by aluminum-tolerant and aluminum-sensitive soybean in acid soil. Biol Fert Soils 48:97–108. https://doi.org/10.1007/s00374-011-0608-7
Yin X, Lu J, Wang Y, Liu G, Hua Y, Wan X, Zhao J et al (2020) The abundance of nirS-type denitrifiers and anammox bacteria in rhizospheres was affected by the organic acids secreted from roots of submerged macrophytes. Chemosphere 240:124903. https://doi.org/10.1016/j.chemosphere.2019.124903
Zhang HH, Huang TL, Chen SN, Yang X, Lv K, Sekar R (2015) Abundance and diversity of bacteria in oxygen minimum drinking water reservoir sediments studied by quantitative PCR and pyrosequencing. Microb Ecol 69:618–629. https://doi.org/10.1007/s00248-014-0539-6
Zhang S, Pang S, Wang P, Wang C, Guo C, Addo FG, Li Y (2016) Responses of bacterial community structure and denitrifying bacteria in biofilm to submerged macrophytes and nitrate. Sci Rep 6:36178. https://doi.org/10.1038/srep36178
Zhang HH, Wang Y, Chen SN, Zhao ZF, Feng J, Zhang ZH, Lu KY et al (2018) Water bacterial and fungal community compositions associated with Urban Lakes, Xi’an, China. Int J Environ Res Public Health 15:469. https://doi.org/10.3390/ijerph15030469
Zhang P, Grutters BMC, van Leeuwen CHA, Xu J, Petruzzella A, van den Berg RF, Bakker ES (2019) Effects of rising temperature on the growth, stoichiometry, and palatability of aquatic plants. Front Plant Sci 9:1947. https://doi.org/10.3389/fpls.2018.01947
Zhang X, Wang G, Tan Z, Wang Y, Li Q (2021) Effects of ecological protection and restoration on phytoplankton diversity in impounded lakes along the eastern route of China’s South-to-North Water Diversion Project. Sci Total Environ 795:148870. https://doi.org/10.1016/j.scitotenv.2021.148870
Zhao JW, Peng L, Liu GL, Wan XQ, Hua YM, Zhu DW, David PH (2019) Diversity of anammox bacteria and abundance of functional genes for nitrogen cycling in the rhizosphere of submerged macrophytes in a freshwater lake in summer. J Soil Sediment 19:3648–3656. https://doi.org/10.1007/s11368-019-02340-4
Zhou J, Fong JJ (2021) Strong agricultural management effects on soil microbial community in a non-experimental agroecosystem. Appl Soil Ecol 165:103970. https://doi.org/10.1016/J.APSOIL.2021.103970
Funding
This study was funded by the National Natural Science Foundation of China [Nos. 41807053, 32170530] and the Natural Science Foundation of Shandong Province (ZR2019PB015).
Author information
Authors and Affiliations
Contributions
J.Z., N.Z., and Y.K. conceived the experiments; Z.G., Y.C., W. S., and F.S. conceived and conducted the experiments; Q.W. analyzed the results; and Z.J. wrote the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Responsible Editor: Diane Purchase
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, N., Wu, M., Che, Y. et al. Effects of shining pondweed (Potamogeton lucens) on bacterial communities in water and rhizosphere sediments in Nansi Lake, China. Environ Sci Pollut Res 29, 51665–51673 (2022). https://doi.org/10.1007/s11356-022-19516-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-022-19516-0