Abstract
Nutrition and pollution stress stimulate genetic adaptation in microorganisms and assist in evolution of diverse metabolic pathways for their survival on several complex organic compounds. Persistent organic pollutants (POPs) are highly lipophilic in nature and cause adverse effects to the environment and human health by biomagnification through the food chain. Diverse microorganisms, harboring numerous plasmids and catabolic genes, acclimatize to these environmentally unfavorable conditions by gene duplication, mutational drift, hypermutation, and recombination. Genetic aspects of some major POP catabolic genes such as biphenyl dioxygenase (bph), DDT 2,3-dioxygenase, and angular dioxygenase assist in degradation of biphenyl, organochlorine pesticides, and dioxins/furans, respectively. Microbial metagenome constitutes the largest genetic reservoir with miscellaneous enzymatic activities implicated in degradation. To tap the metabolic potential of microorganisms, recent techniques like sequence and function-based screening and substrate-induced gene expression are proficient in tracing out novel catabolic genes from the entire metagenome for utilization in enhanced biodegradation. The major endeavor of today’s scientific world is to characterize the exact genetic mechanisms of microbes for bioremediation of these toxic compounds by excavating into the uncultured plethora. This review entails the effect of POPs on the environment and involvement of microbial catabolic genes for their removal with the advanced techniques of bioremediation.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
The new era of accelerating urbanization has enabled mankind to exploit natural resources leading to the emergence of various industrial centers and increase in industrial escalation have brought rise in air, water, and soil pollution to an alarming level. The environmental pollution is caused by many persistent pollutants like alkanes, antibiotics, cyanides, dioxins, phenols, polychlorinated biphenyls (PCBs), polycyclic aromatic hydrocarbons (PAHs), pesticides, synthetic azo dyes, polyaromatic, chlorinated, and nitro-aromatic compounds. Toxic chemicals which exist persistently in the environment for several years before getting totally mineralized are known as persistent organic pollutants (POPs) (UNEP 2006).
POPs include aldrin, dieldrin, endrin, chlordane, dichlorodiphenyltrichloroethane (DDT), heptachlor, mirex, toxaphene, polychlorinated biphenyl, hexachlorobenzene, polychlorinated dibenzo-p-dioxin, and polychlorinated dibenzofuran. The Stockholm (2001) under the support of United Nation Environmental Program (UNEP) also specified a set of POPs, considered as potential endocrine-disrupting chemicals (EDCs) (ecoestrogens) such as phenols, biphenyl compounds, phthalates, etc., in the environment (Nagao 1998; Hutz 1999; Borgeest et al. 2002). Recently, nine compounds have been listed under POPs through Stockholm Convention (2010), namely, chlordecone, hexabromobiphenyl, α-hexachlorocyclohexane, β-hexachlorocyclohexane, lindane, tetrabromo diphenyl ether, pentachlorobenzene, perfluorooctane sulfonic acid, and hexabromo diphenyl ether. In addition to the wide range of POPs, new emerging contaminants are also gaining concerns due to the huge toxic effects based on their toxic equivalency factor. These contaminants include pharmaceuticals, personal care products, steroids, hormones, surfactants, abused drugs, flame retardants, and industrial additives (La Farre et al. 2008; Covaci et al. 2011; Richardson and Ternes 2011). Among these emerging contaminants, polybromodiphenyl ether (PBDEs), perfluorinated compounds (PFCs), short-chained chlorinated paraffins (SCCPs), and hexabromocyclododecane (HBCDs) groups are highly toxic and cause serious damages to the ecosystem. However, the Stockholm Convention (2009) has included only perfluorooctane sulfonic acid (PFOS) from PFCs, commercial pentabromodiphenyl ether (penta-BDE), and commercial octabromodiphenyl ether (octa-BDE) from PBDEs into the list of POPs (Li et al. 2014).
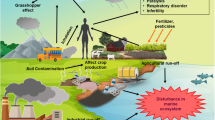
Entry of the toxic POPs in the environment occurs through different sources like intensive agriculture, pharmaceuticals, pulp, paper, and mining industries. They cause behavioral abnormalities and birth defects in fish, insects, and mammals. In case of humans, adverse effects are more prominent with behavioral, developmental, endocrine, immunologic, neurologic, and reproductive changes causing cancer, diminished intelligence, attention deficits, learning disorders, poor gross, and motor coordination disabilities (Agency for toxic substances and disease registry ATSDR 2002; Kumar 2004). POPs in mother blood can be readily transferred through the placenta to the offsprings. People having a diet of fish, shell fish, and wild food rich in fat are more prone to POP exposure (United States Environmental Protection Agency USEPA 2009). The lipophilic nature of POPs render their deposition in the fatty tissue of living beings which after biomagnification leads to acute and chronic toxicities in different trophic level of the food chain. In the coastal environment, POPs bind to various sediment particles which help them in crossing estuaries (USEPA 2009). Hence, POPs stuck in the sediments are absorbed by benthic fauna and also can get volatilized through the water column. POPs are considered to have “grasshopper effect” due to their semi volatile nature and ability to travel great distances through cycles of evaporation, atmospheric cycling, and deposition (Gouin et al. 2004). At the present scenario, deciphering a compatible mode of biological attempt to resolve the problem of environmental pollution is the major global issue (Gienfrada and Rao 2008).
Natural communities of versatile and diverse microorganisms harbor a remarkable physiological adaptability and catabolic potential for the breakdown of massive organic molecules. Organic contaminants present in the environment are degraded by them releasing simpler products such as carbon dioxide and water by the process of biomineralization (Das et al. 2016). With the advent of time, stress induced by pollution and nourishment deficit is escalating the diversity of microbial metabolic pathways. The movement of mobile genetic elements such as plasmids and transposons by horizontal gene transfer delivers a harmless and economic substitute for organic contaminant degrdadation (Pieper and Reineke 2000; Sinha et al. 2009). Although numerous investigations are taking place, many facets of microbial assisted bioremediation still remains uncharted. Gene rearrangements and mutation followed by catabolic gene expression in the microbes might be beneficial for adaptation in complex contaminated sites. Therefore, tracing out the exact genetic system of bacteria and their metabolic map in degrading toxic persistent organic pollutants will provide a broad insight for enhanced bioremediation. In addition, their evolutionary potential and genetic flexibility traced through advanced techniques from the uncultured plethora will reveal the generation of new catabolic traits for detoxification or degradation of these compounds. This review provides brief account of the molecular perspectives of biodegradation of POPs and their bioremediation aspects.
Microbial degradation of POPs
Microbial population having the ability to utilize a wide range of POPs includes many aerobic and anaerobic bacteria. These POPs are degraded in the oxic zone by various microbes which are chemoorganotroph in nature (Fig. 1). Bacterial genera showing chemoorganotrophy for POP degradation include Aeromicrobium, Bacillus, Brevibacterium, Burkholderia, Desulfotomaculum, Desulfovibrio, Dietzia, Escherichia, Gordonia, Methanoseata, Methanospirillum, Micrococcus, Moraxella, Mycobacterium, Pandoraea, Pelatomaculum, Pseudomonas, Rhodococcus, Sphingobium, and Syntrophobacter (Chowdhury et al. 2008). Aerobic Gram-negative rods Pseudomonas fluorescens and Pseudomonas putida have the highest biodegradation potential owing to their catabolic enzymes (Houghton and Shanley 1994). Catabolic enzymes in a catabolic pathway are specific protein molecules which catalyze the degradation of complex molecules into simpler ones and release the chemical energy stored in the bonds of those molecules. Therefore, understanding the evolution of modern catabolic pathways and their respective catabolic enzymes is an effective means to determine enhanced cleanup of pollutants (Mrozik et al. 2003). Gram-positive bacteria Corynebacterium, Mycobacterium, Nocardia, and Rhodococcus are highly effective in biodegradation and are commonly described as biodegraders of four ring polyaromatic hydrocarbons (Nzila 2013). Cometabolism exhibiting strains include Achromobacter, Alcaligenes, Arthrobacter, Aspergillus, Azotobacter, Bacillus, Brevibacterium, Flavobacterium, Hydrogenomonas, Methylosinus trichosporium, Microbacterium, Micrococcus, Nocardia, Pseudomonas, Rhodococcus chlorophenolicus, Streptomyces, Trichoderma, Vibrio, and Xanthomonas (Fritsche and Hofrichter 2000). These microorganisms have the ability to degrade a pollutant without using it as a growth substrate, while sustaining its own growth by assimilating a different substrate (Nzila 2013). Different genera of microorganisms develop numerous enzymatic modifications in the metabolic pathways to actively metabolize various POPs (Table 1). Reactions like reduction, oxidation, hydrolysis, dehalogenation, and methylation are basically performed by several aerobic and anaerobic microorganisms.
Janibacter sp. has been reported to utilize dibenzofuran, while certain other bacterial species like Bacillus cereus, Bacillus vireti, and Sphingomonas yanoikuyae JAR02 have been documented to effectively degrade benzo[a]pyrene (Yamazoe et al. 2004; Rentz et al. 2008; Rout et al. 2012). Ralstonia sp. SA-5 and Pseudomonas sp. SA-6 were also recognized to substantially metabolize 2,3,4,5-tetrachlorobiphenyl (2,3,4,5-tetraCB) and 2,3′,4′,5-tetraCB (Adebusoye et al. 2008). A marine bacterium Pseudomonas aeruginosa JP-11 was isolated from the coastal sediments of Odisha and was found to metabolize biphenyl within 72 h when supplied as the sole source of carbon (Chakraborty and Das 2016). Yeasts such as Aureobasidium pullulans, Candida maltosa, Exophiala jeanselmei, Rhodotorula glutinis, and Trichosporon cutaneum could efficiently utilize aromatic pollutants like acetophenone, benzoic acid, ortho-cresol, para-cresol, and phenol (Fritsche and Hofrichter 2004). Molds such as Aspergillus fumigatus, Aspergillus niger, Fusarium flocciferum, Penicillium frequentens, and Penicillium simplicissimum utilized substitutes of phenols and benzoic acid (Hofrichter et al. 1994). A novel aerobic dieldrin-degrading bacterium, Pseudonocardia sp. strain KSF27 was isolated from an enrichment culture of soil-charcoal perfusion system capable of degrading aldrin trans-diol, dieldrin, and other persistent organochlorine pesticides, such as endosulfan sulfate, α-endosulfan, β-endosulfan, heptachlor, heptachlor epoxide, and chlordecone (Sakakibara et al. 2011). Two predominant bacteria Burkholderia sp. strain MED-7 and Cupriavidus sp. MED 5 were reported to degrade dieldrin and endrin in aerobic conditions, when grown on 1,2-epoxycyclohexane (ECH). Hexachlorobenzene, a highly recalcitrant environmental pollutant, was aerobically degraded by Nocardioides sp. strain PD653 (Matsumoto et al. 2008; Takagi et al. 2009). Likewise, 2, 2′,4,4′-tetrabromodiphenyl ether (BDE-47) was degraded by Pseudomonas stutzeri strain BFR01 isolated from the polluted soil of a brominated flame retardant production industry (Zhang et al. 2013).
Desulfitobacterium and Dehalococcoides are promising anaerobic dehalorespiring bacteria. Anaerobic bacteria using chloroethenes as final electron acceptor include Dehalobacter, Dehalococcoides, Dehalococcoides ethenogenes, Desulfitobacterium, Desulfuromonas, Geobacter, and Sulfurospirillum (Futagami et al. 2008). Clostridium, Desulfobacterium, Desulfovibrio, Methanococcus, and Methanosarcina are also capable of reductive dehalogenation and are ubiquitous in nature (Zhang and Bennett 2005; Bedard 2008). Mixed consortia of numerous species in aerobic and anaerobic conditions are extremely useful in biodegradation of complex organic compounds to form carbon dioxide and water. Due to restricted substrate biodegradation by single species, miscellaneous assemblages of bacterial populations are employed with extensive enzymatic potential for enhanced biodegradation (Ghazali et al. 2004).
POPs remediation in soil and water can be undertaken by the collective use of plants and bacteria in the system as the plants transport nutrients to their associated rhizosphere. Endophytic bacteria, reported to degrade POPs, also maintain plant growth by siderophore production, 1-aminocyclopropane-1-carboxylic acid (ACC) deaminase production, and nitrogen fixation (Arslan et al. 2015). Moreover, bacteria also help by reducing the toxicity of the pollutants in the environment by evapotranspiration (Afzal et al. 2014). This is an ecofriendly approach as it utilizes plants to transform, sequester, extract, and detoxify pollutants present in sediments, soil, groundwater, surface water, and atmosphere for restoration of contaminated sites (Samardjieva et al. 2015).
Adaptation strategies of POPs degrading microorganisms
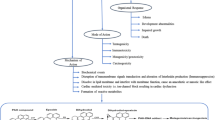
Gene mutation, gene rearrangement, and differential regulation of genes in microbes help in their survival in various unfavorable conditions (Thomas and Nielsen 2005). Microbes undergo numerous genetic permutation and combination to ensure a metabolically active life (Fig. 2). A cellular mechanism known as hypermutation helps the immune system to adapt to foreign elements. Therefore, in stressful environments, hypermutability in microorganisms aids in faster adaptation than normo-mutators (Marcobal et al. 2008). However, in stable environments or newly adapted environments, this has no value and is a fitness disadvantage. This effect can be reduced by the restoration of normal mutability by horizontal gene transfer of good mismatch repair (MMR) genes from heterologous sources. This can be achieved by homeologous recombination (i.e., recombination between largely homologous but nonidentical DNA sequences) (Rayssiguier et al. 1989; Jayaraman 2011).
Adaptation strategies of microorganism degrading POPs. On nutrition and pollution stress, several types of modifications occur in metabolic genes of microorganisms. Some of which includes frameshift mutation, gene duplication, and genetic recombination coding new functional proteins for adaptation in unfavourable conditions
Increasing chemical pollution of organic compounds induces error-prone DNA replication due to base mutation and nucleotide replacement accelerating mutational drift (Black 1999). Gene duplication results in the independency of selective pressure on the extra gene copy enhancing speedy mutations. Insertion elements play an imperative role in DNA rearrangements, gene transmission, and activation/inactivation of silent genes. DNA replication and repair may occasionally incorporate single site mutations at random, which might not be the prime reason for catabolic enzyme diversity in microbes. Sometimes, gene conversion or slipped-strand mispairing can cause natural alterations in DNA sequences (Niedle et al. 1988). However, metabolic engineering-based approach can be utilized for gene conversion to enable degradation of POPs. In a study by Yan et al. (2006), cytochrome P-450cam variant coded by the gene cassette (cam A+ cam B+ cam C) was incorporated into the nonessential pcpM gene of Sphingobium chlorophenolicum ATCC 39723 (a pentachlorophenol degrader) by homologous recombination. This recombinant strain could degrade hexachlorobenzene at an increased rate of 0.67 nmol mg (dry weight)−1 h−1.
Phase variation is an inherited reversible form of gene regulation in bacteria. DNA slippage is potentially a significant source of genetic diversity for catabolic genes and is one of the primary mechanisms of phase variation. This occurs by slipped-strand mispairing (SSM), which is generated between the mother and daughter strand during DNA replication, recruiting significantly diverse genes for the catabolism of persistent organic compounds (Levinson and Gutman 1987; Henderson et al. 1999). Short sequence repeat (SSR) microsatellites (short, contiguous homogenous or heterogeneous repetitive DNA sequence of 6 bp or less) are the susceptible regions for SSM (van Belkum et al. 1998). This can change the number of repeat units leading to the alteration of gene expression at transcriptional and translational level. Thus, bacteria inhabiting a restricted and unfavorable environment might benefit from SSM by unregulated phenotypic diversity (Richardson and Stojiljkovic 2001; Torres-Cruz and van der Woude 2003). In a study, Vallaeys et al. (1999) reported genetic diversity in the catabolic pathway of 2,4-dichlorophenoxyacetic acid (2,4-D). The genes involved for 2,4-D degradation include tfdA, tfdB, and tfdC. However, sequence comparison indicated a wide divergence, demonstrating an early origin of tfd genes. Burkholderia sp. strain TFD2 harboring a tfdA gene was closely related to that of Burkholderia sp. strain RASC while its tfdB gene was grouped with the tfdB gene of Sphingomonas sp. strain TFD26. There was also a lack of variance in the partial nucleotide sequences of the tfdB genes from Burkholderia sp. TFD2 and Sphingomonas sp. TFD26. Therefore, SSM might be the significant source for exhibiting independent recruitment of varied gene cassettes in POP degradation.
Environmental changes and biotic evolution leading to new functional niches permit relentless gene families diversification and existence of new lineage of genes (Francino 2012). Natural spontaneous generation strategies contributing to microbial evolution include minor indigenous changes in the nucleotide sequence of the genome, intragenomic reshuffling, and procurement of DNA sequence from other organisms. Systems of site-specific recombination, DNA repair, and restriction modification are examples which modulate the genetic variation frequency (Arber 2000). Mobile genetic elements like plasmids and transposons accomplishing horizontal gene transfer and patchwork assembly are involved in pollution stress adaptation (Top and Springael 2003). Three mechanisms of gene transfer-transformation, conjugation, and transduction lead to vertical and horizontal gene transfer (HGT) within the microbes (Thomas and Nielsen 2005; Heuer and Smalla 2012). Among them, conjugation is the primary mechanism of horizontal catabolic gene transfer leading to bacterial adaptation to the changing environment (Sørensen et al. 2005). The plasmids, transposons, integrons, and insertion sequences (IS) are imperative mobile genetic elements involved in adaptation mechanism (Schlüter et al. 2007). HGT is beneficial in bioremediation shifting microbial communities in favor of the degradation of POPs. Another strategy for POP degradation includes genetic bioaugmentation which indicates the introduction of small fraction of bacteria harboring genes for enzymes that mineralize the contaminant of interest and stimulate in situ HGT of those degradative mobile genetic elements to the native bacterial community. The model HGT system TOL plasmid, a diverse family of degradative plasmids, was described to be involved in the degradation of xylenes, toluene, and related species (Ikuma 2011).
Another facet is the microbial biofilm which has been considered as the potential agent for bioremediation. A potent biofilm forming marine bacterium Pseudomonas mendocina NR802 was isolated from Rushukulya, Odisha, India, capable of utilizing phenanthrene (Mangwani et al. 2014). Another marine bacterium Stenotrophomonas acidaminihila NCW-702, isolated from Chilika Lake, Odisha, India, was found to degrade 71.1 ± 3.1 and 40.2 ± 2.4 % of phenanthrene and pyrene, respectively (Mangwani et al. 2015). The multilayered, three dimensional structures of biofilm encapsulated in a hydrated extracellular polymeric substances (EPS) on a substratum provides a perfect environment for horizontal gene transfer of mobile genetic elements due to close proximity and quick spread of plasmid DNA by conjugation of the competent bacteria (Cvitkovitch et al. 2003). This enables rapid phage spreading and plasmid uptake by competent bacteria (Jefferson 2004). Therefore, application of high biofilm-forming microbial communities can effectively provide a stable environment for gene transfer as well as enhance speedy biodegradation of complex persistent compounds such as pyrene, phenanthrene, polychlorobiphenyl, and pesticides (Molin and Nielsen 2003).
Cell membrane modification by microbes is also an adaptation strategy for their survival in toxic environmental conditions (de Carvalho et al. 2009). In this regard, certain surface-active compounds like biosurfactants are produced and also the presence of membrane bound efflux pumps help in the removal of toxic compounds such as biphenyl, benzene, and polychlorobiphenyl outside the cells (Ron and Rosenberg 2002; Van Hamme et al. 2003; Chakraborty and Das 2014. Chakraborty and Das 2016). Bioaugmentation with multicomponent system by the addition of previously adapted pure bacterial strain and consortium with biodegradation relevant genes enveloped in a vector can be transported by conjugation into indigenous microorganisms. This forms a better way for in situ bioremediation of chlorinated solvents, herbicides, ethylbenzene, xylenes, emerging contaminants, etc. (El Fantroussi and Agathos 2005; Morgante et al. 2010).
Involvement of the catabolic genes
Pollutant degradation is dependent on the adaptive response of microbial communities to the pollutant, which differs between selective enrichment of genes and genetic changes (Sinha et al. 2011). The rapid exchange of novel catabolic activities occurs by the transmission of genes between microbes, frequently by broad host range plasmids. This leads to the unique catabolic function of a single recombinant strain in biodegradation, which is not present in either strain individually (Pemberton and Schmidt 2001). The location of these catabolic genes varies in different organisms (Table 2). Genes may be situated on plasmid or present on genomic DNA consisting of single operon system, as found in phenol degradation (Khan et al. 2001). Sometimes, they may be present on plasmid genome organized in two operon systems as found in degradation of polychlorinated biphenyls, or organized on two or more operon systems for degradation (Seo et al. 2009). Frequently, genes may be located on transposons, as found in dibenzo-p-dioxin and dibenzofuran degradation by Sphingomonas sp. RW1 which contains a Tn-5 lacZ mini transposon (Megharaj et al. 2011).
Most of the catabolic genes are constitutively expressed at a low level, but on exposure to the desired compound, transcription of specific genes are activated. In some cases, simple operon system having all necessary enzymes for absolute degradation of a particular metabolite is activated, whereas in complex systems, the metabolites are transformed into varying forms with the activation of intricate regulatory mechanisms of numerous semi-independent operons (Pemberton and Schmidt 2001). In Pseudomonas sp. P51, chlorobenzene catabolic gene tcbAB was located in the transposon Tn5280 (van der Meer et al. 1991). Similarly, in the isolate Alcaligenes BR60, chlorobenzoate catabolic gene cbaABC was located in the transposon Tn5271 (Nakatsu et al. 1991). Biphenyl degrading gene bph/cbdABCD in Ralstonia eutropha A5 was present as in the conjugative catabolic transposon Tn4371. An organochlorine herbicide, 2,4 dichlorophenoxyacetic acid was effectively catabolized by Alcaligenes eutrophus harboring the plasmid pJP4 (Ledger et al. 2006). Rhodococcus erythropolis and Sphingomonas sp. have plasmids and gene clusters on their chromosome capable of catalyzing PCBs, polybrominated biphenyls (PBBs), and chlorinated dibenzo-p-dioxins, respectively. Presence of a linear plasmid pBD2 in R. erythropolis helped in trichloroethylene degradation (Pemberton and Schmidt 2001). Thus, the catabolic genes for degradation of POPs have been reported in plasmids, transposons, and genomes organized on one or multiple operon systems.
Microbial degradation process of most toxic POPs
Stability of POPs in environment depends on their electrophilicity, nucleophilicity, photodegradability, and biodegradability, which determine the reactivity and favorability of a chemical reaction. Electrophilicity index, a specific property of a chemical species, is the square of its electronegativity divided by its chemical hardness (Chattaraj et al. 2006). The electrophilicity index of polychlorinated biphenyls and benzidine was analyzed which was adequate enough to describe their toxicity (Roy et al. 2006). Compounds with increasing substitutions of halogens have high recalcitrance. In nucleophilic aromatic substitution, aromatic hydrodehalogenation occurs by hydride transfer. This hydride transfer to an aromatic substrate weakens the carbon-halogen bond rendering their heterolytic cleavage (Sadowsky et al. 2014). Nucleophilic aromatic substitution has been of biological importance as both glutathione-S-transferases and 4-chlorobenzoyl-CoA dehalogenases catalyze hydrodehalogenation (Zheng and Ornstein 1997; Xu et al. 2004). Likewise, light-induced chemical transformation of an organic contaminant is known as photodegradation which has an advantage of good selectivity, low reaction temperature, and complete degradation (Ray, 2000). The stability of POPs is also dependent on the photodegradable ability of the organic contaminants. For example, the chloro-substituted aromatic compounds and DDT have been demonstrated to be photomineralized in water; however, biphenyls, dioxins, and furans are comparatively slow to photooxidize (Wenzel et al. 1999).
POP uptake by plants and microbes depends on a number of physicochemical characteristics such as octanol-water partition coefficient (log Kow), acidity constant (pKa), aqueous solubility (Sw), octanol solubility (So), and the concentration of the pollutant (Admire et al. 2014). Some of the microorganisms are unable to mineralize these pollutants because they may not be recognized as substrate by the existing enzymes used for degradation. Sometimes, they might be chemically and biologically very stable containing substitution groups like amino, carbamyl, halogens, methoxy, nitro, and sulphonate (Jha et al. 2015). Moreover, in some cases, the compounds might be insoluble in water which could remain adsorbed to the external matrices of soil. Furthermore, large molecular size of POPs and absence of permease in the environment might reduce their transport into microbial cells. Therefore, there may be numerous reasons for unproductive biodegradation in toxic contaminated sites (Isken and de Bont 1998).
According to the United Nations Environment Programme (UNEP 2006), the 12 POPs with the nine new POPs fall into three broad categories, namely, pesticides, industrial chemicals and by-products, and emerging contaminants. A representative from each group is discussed in the further section.
1,1,1-Trichloro-2,2-bis-(4′-chlorophenyl) ethane (DDT)
DDT is a persistent, chlorinated organic insecticide present in the environment. Having a biological half-life of 8 years, it is not metabolized very rapidly by animals and is deposited and stored in their fatty tissues (Joesten et al. 2006). The USA have banned the use of DDT, but due to its wide insecticidal applications on extensive diverse insects, it is still used worldwide. Presence of chlorine atom contributes to their low solubility, preferentially partitioning them into lipophilic phase and making them highly toxic. It renders thinning of eggshells in many birds especially falcons enlisting them into the endangered category. It adversely affects hormone balance and also causes cancer. However, some microorganisms attack the aromatic and alicyclic moieties of DDT (Nadeau et al. 1994; Hay and Focht 1998; Quensen et al. 1998). With the aerobic bacterium Ralstonia eutropha A5, DDT is converted to cis-2,3 dihydrodiol DDT using the enzyme DDT 2,3-dioxygenase. This is followed by the formation of 6-oxo-2-hydroxy-7-(4′-chlorophenyl)-3,8,8,8,-tetrachloroocta-2Z, 4Z-dienote, which is finally recruited to 2-,4-dichlorobenzoate pathway in the aerobic mode of metabolism. However, with the anaerobic bacterium Klebsiella pneumonia, DDT dehydrochlorinase enzyme cleave DDT to 1,1-dichloro-2,2-bis (4′-chlorophenyl) ethylene (DDE), which enters into the aerobic pathway as mentioned above. In a consortium, composed of Bacillus sp., Staphylococcus sp., and Stenotrophomonas sp., the degradation of DDT formed l,l-dichloro-2,2-bis(p-chlorophenyl)ethane (Mwangi et al. 2010). DDT-degrading bacteria include Bacillus circulans, Bacillus pumilus, Enterobacter aerogenes, Enterobacter cloacae, Escherichia coli, Flavobacterium sp., Hydrogenomonas, Klebsiella pneumonia, Micrococcus, Pseudomonas aeruginosa, Pseudoxanthomonas jiangsuensis, Pseudomonas putida, and fungi such as Phanerochaete chrysosporium, Saccharomyces cerevisiae, and Trichoderma viridae (Sharma et al. 1987; Beunink and Rehm 1988; Wang et al. 2011).
Microbial reductive dechlorination (RD) is the foremost mechanism of conversion from DDT isomers to dichlorodiphenyldichloroethane (DDD) under reducing conditions. It substitutes the aliphatic chlorine for a hydrogen atom with the involvement of one proton and two electrons (Fries et al. 1969). The sequential two-electron transfer reaction causes dissociation of chloride anion forming p, p-dichloro-diphenyl-dichloroethyl radical. This is followed by second electron transfer reaction protonating the radical to form DDD (Bylaska et al. 2004). This cumulative process is termed dehalogenation, which results in chlorine removal and hydrogen ion addition to the compound. Thus, susceptibility to oxidation and formation of less toxic products are augmented. DeWeerd et al. (1990) identified Desulfomonile tiedjei, an anaerobic bacterium, which coupled the reductive dechlorination of 3-chlorobenzoate. Cleavage of carbon–chlorine bond is the primary mechanism for DDT mineralization (Haggblom and Bossert 2004). Dechlorination of DDT under reducing conditions forms DDD, which on aerobic conditions, further degrades to form the polar product, DDOH (Mwangi et al. 2010). The mutualistic anaerobic microbial communities favor RD of this compound (Mohn and Tiedje 1992). Usually, DDT degradation follows this pathway; however, aerobic bacterial degradation was first reported by Nadeau et al. (1994) using Alcaligenes eutrophus A5, which oxidized DDT in presence of dioxygenase to form 2,3-dihydrodiol-DDT which further degraded to 4-chlorobenzoic acid (Fig. 3). In a study by Kamanavalli and Ninnekar (2004), the growth of Pseudomonas sp. was reported on biphenyl supplemented with 0.05 % w/v DDT. The degradation produced 2,3-dihydroxy DDT via the meta-cleavage pathway forming 4-chlorobenzoic acid as the dead end-product. Another Gram-negative, strictly aerobic bacterium, Pseudoxanthomonas jiangsuensis, was isolated from a DDT-contaminated soil, which predominantly degraded DDT to diphosphatidylglycerol, phosphatidylethanolamine, and phosphatidylglycerol (Wang et al. 2011).
Aerobic and anaerobic degradation pathway of DDT. DDT (A) is acted upon by the enzymes DDT 2,3-dioxygenase and DDT dehydrochlorinase recruiting them to aerobic and anaerobic pathways respectively. The primary product of the aerobic pathway is cis-2,3-dihydrodiol DDT (B) which is acted upon by the enzyme cis-2,3-dihydrodiol DDT dehydrogenase to form 2,3-dihydroxy DDT (C). C is further converted to 6-oxo-2 hydroxy-7-(4′-chlorophenyl)-3,8,8,8,8 tetrachloroocta-2Z, 4Z-dienote (D) finally forming 4-chlorobenzoate (K). The primary product of the anaerobic pathway is 1,1-dichloro-2,2-bis (4′-chlorophenyl)-ethylene (DDE) (E). E is cleaved to 1-chloro-2,2-bis-(4′-chlorophenyl)-ethylene (DDMU) (F) and 1,1-dichloro-2-(dihydroxy-4′-chlorophenyl)-2-(4-chlorophenyl) ethylene (G) by the enzyme DDE dehalogenase. F advances to the anaerobic pathway to form 4-chlorophenyl acetate (H) which leads to the formation of 4-chlorobenzoate (K). Similarly, on action of dioxygenase enzyme, G is converted to 6-oxo-2-hydroxy-7-(4′-chlorophenyl)-3,8,8,8-trichloroocta-2Z,4Z,7-trienoate (H). By the action of hydrolase, H forms 4- chlorobenzaldehyde (J) forming 4-chlorobenzoate (K) which is then mineralized by entering into the citric acid cycle via the intermediate 4-hydroxybenzoate (L)
Polychlorobiphenyl (PCB)
PCBs, the representative of industrial chemicals, are a class of chemicals consisting of 209 compounds, collectively known as congeners. The degree of dechlorination and the locus of the chlorinated sites mark the differences in the compounds. PCBs are widely used in insulator fluid for transformers and extender of insecticides. Due to chemical complexity and high toxicity on human and wild life, they were banned in the mid-1980s, but they are omnipresent globally due to their resistance to degradation (Kidd et al. 2012).
The microorganisms follow the meta-cleavage pathway for PCB degradation to produce Krebs cycle intermediates and chlorobenzoates (Pieper 2005). In aerobic biodegradation, the primary step is the dioxygenation of PCB congeners, by the biphenyl dioxygenase enzyme (Fig. 4). It catalyzes the incorporation of two hydroxyl groups in the aromatic ring of the PCB congener, which becomes more susceptible to enzymatic ring fission reactions (Bruhlmann and Chen 1999). Biphenyl dioxygenase is a multicomponent enzyme consisting of a terminal dioxygenase (large alpha and small beta subunit), ferredoxin, and ferredoxin reductase encoded by bph operon (Erickson and Mondello 1992). bphA gene, encoding the large alpha subunit of the enzyme, functions in substrate recognition (Fig. 5). This BphA enzyme is substrate specific and varies from organism to organism. Presence of bphA gene been reported in Pseudomonas aeruginosa JP-11 isolated from the Paradip port, Odisha, India, with the function of 98.86 ± 2.29 % of biphenyl degradation (Chakraborty and Das 2016). Its presence was also observed in Burkholderia cepacia and Pseudomonas pseudoalcaligenes having sequence similarity in their bph operon. However, in Burkholderia cepacia, biphenyl dioxygenase favorably deoxygenates ortho-substituted PCBs, while in Pseudomonas pseudoalcaligenes KF707, this enzyme significantly deoxygenates para-substituted PCBs (Erickson and Mondello 1993; Gibson et al. 1993). DNA shuffling experiments in Burkholderia cepacia, Comamonas testosteroni, and Rhodococcus globerulus obtained variants of enzyme with better degradation capabilities for PCBs (Furukawa 2000). This method incorporated in vitro random recombination of bphA genes by fragmentation and PCR reassembly. It was seen that hybrid BphA, II-9 (hybrid of B. cepacia and C. testosteroni) was able to oxygenate 2,6 dichlorobiphenyl up to 58 % after 18 h, while the parental enzymes did the same reaction with 10 % less efficiency (Barriault et al. 2002). Utilizing a rational design approach, biphenyl dioxygenase of P. pseudoalcaligenes KF707 was three-dimensionally modeled. This was based on the crystallographic analysis of naphthalene dioxygenase enzyme from Pseudomonas sp. (Suenaga et al. 2002). A clear picture of key positions near the active site of the enzyme was visualized which were utilized for site-directed mutagenesis. Hence, mutants of Pseudomonas 1335F, T376N, and F377L very efficiently degraded 2,5,2′,5′-tetrachlorobiphenyl that was not mineralized by the wild-type biphenyl dioxygenase enzyme (Ang et al. 2005).
Genes involved in catabolic degradation of PCBs. Biphenyl dioxygenase encoded by bph A1A2A3A4 gene results in dioxygenation of the dual ring. The bph B codes for enzyme dehydrogenase involved in the formation of 2,3-dihydroxybiphenyl. This is further acted upon by Bph C (bph C) forming 2-hydroxy-6-oxonated-6 phenylhexa-2,4-dienoic acid (HOPDA). The gene product of bph D forms 2-hydroxypenta-2,4-dienoate, which acts as a substrate for the enzymes encoded by bph EFG forming acetyl CoA that enters into the Krebs cycle
bph operon model conserved among PCB-degrading bacteria: A1, terminal dioxygenase large subunit; A2, terminal dioxygenase small subunit; A3, ferrodoxin; A4, ferrodoxin reductase; B, dihydrodiol dehydrogenase; C, 2,3-dihyrodoxybiphenyl dihydrogenase; D, 2-hydroxy-6-oxo-6-phenylhexa-2,4-dienoic acid hydrolase; E, 2-hydroxypenta 2,4-dienote hydratase; F, 4-hydroxy-2-oxovalerate aldolase; G, acetaldehyde dehydrogenase
Mostly the primary product of PCB degradation is chlorobenzoate, which requires certain catabolic plasmids from other microorganisms for its cleavage. Pseudomonas aeruginosa harboring the plasmid pE43 contains the oxygenolytic ortho-dechlorination ohb gene, whereas Arthrobacter globiformis containing the plasmid pPC3 carried the hydrolytic para-dechlorination fcb gene. Comamonas testosteroni transformation by these recombinant plasmids was capable of utilizing ortho- and para-chlorobiphenyl as their sole carbon source (Hrywna et al. 1999). Likewise, 4,4′-dichlorobiphenyl (double para-replaced congeners) was easily mineralized by P. pseudoalcaligenes KF707 but 2,5,2′,5,-tetrachlorobiphenyl could not be degraded. Similarly, B. cepacia LB400 has a broad substrate specificity attacking 2,5,2′,5′-tetrachlorobiphenyl via 3,4-dioxygenation (Suenaga et al. 2001). Amino acid mutations in enzymes also determine change in substrate specificity. In BphA enzyme, asparagine (Asn) at the 376 position renders broad substrate specificity, whereas enzymes with narrow specificity contain threonine (Thr) at this site. The replacement of Thr-376 with Asn in P. pseudoalcaligenes KF707 BphA led to the introduction of 3,4-dioxygenase activity for 2,5,4′-trichlorobiphenyl and 2,5,2′,5′-tetrachlorobiphenyl degradation (Kimura et al. 1997). In naphthalene dioxygenase, this amino acid corresponded to Thr-351 in the large subunit. However, the alteration of Thr-351 to Asn in this enzyme showed an insignificant effect on product formation (Parales et al. 2000). Earlier report stated that four amino acid substitutions in P. pseudoalcaligenes KF707 BphA increase its degradation ability for single aromatic compounds such as alkyl benzenes, benzene, and toluene (Suenaga et al. 1999). Therefore, site-directed mutagenesis-based structural modeling of enzymes can deliver their steric information with altered specificity and function.
Degradation of PCBs has been reported to carry out by bacterial species like Acidovorax, Bacillus, Burkholderia, Comamonas, Corynebacterium, Cupriavidus, Pseudomonas, Rhodococcus, and Sphingomonas (Furukawa and Fujihara 2008; Seeger et al. 2009). Burkholderia xenovorans LB400 was able to degrade a broad range of PCBs and therefore was considered as a model bacterium for PCB degradation (Seeger et al. 2010). Rhodococcus jostii RHA1 was another potent PCB-degrading soil bacterium (McLeod et al. 2006).
In presence of biphenyls and chlorobiphenyls, PCB-degrading bacteria accumulate polyphosphates (polyP) in their exponential phase as an adaptive mechanism. Pseudomonas strain B4 and B. fungorum LB400 accumulated polyP for their augmented survival in presence of chlorobiphenyl (Chavez et al. 2004). Thus, it is concluded that pollution stress drives the synthesis of certain stress-resistant adaptations in bacteria for their survival in toxic contaminated sites. Additionally, designing of engineered proteins with modified active sites can also help in enhanced bioremediation of toxic organic pollutants.
Polychlorinated dibenzo-p-dioxins (PCDDs) and dibenzofurans (PCDFs)
Polychlorinated dibenzo-p-dioxins (PCDDs) and dibenzofurans (PCDFs) are the representative industrial organochlorine by-products having toxic manifestations. They are released as impurities in trace amounts from the chemicals like chlorinated phenols and their derivatives, chlorinated biphenyl ethers, pentachlorophenol, and PCBs. They are also the by-products from combustion of iron and steel production. The pulp and paper industries carrying out chlorine bleaching, chlorine-alkali plants using graphite electrodes, exhausts of car from petrol having chlorinated solvents, sewage sludge, etc. release PCDDs into the environment (Rappe 1991). PCDD/Fs have a high tendency to adsorb into soil, sediments, and bioaccumulate in higher organisms, which render acute and chronic toxicities in them.
Three decades ago, Bumpus et al. (1985) first reported the degradation of 2,3,7,8-tetrachlorodibenzo-p-dioxin (TCDD) by the white rot fungus Phanerochaete chrysosporium. Later, Bunge et al. (2001) reported an anaerobic bacterium Dehalococcoides sp. strain CBDB1 which was capable of converting 1,2,3,7,8-penta CDD to 2,7 or 2,8-DCDD. Biodegradation of chlorinated dioxins by aerobic means is initiated by the angular dioxygenase (Habe et al. 2001; Nojiri and Omori 2002). Strains like Brevibacterium sp., Terrabacter sp., Serratia marsescens, Sphingomonas sp., and S. wittichii efficiently metabolizes dioxin and furans (Jaiswal and Thakur 2007). The angular dioxygenase attacks the ring adjacent to the ether oxygen bridging them (Fig. 6). Different dioxygenase have been isolated from various microorganisms like carbazole dioxygenase from Pseudomonas sp., dibenzofuran dioxygenase from Terrabacter sp., and dibenzo-p-dioxygenase from Sphingomonas sp. (Habe et al. 2001). The dioxygenase further catalyzes the formation of diols forming chlorinated 2,2′,3-trihydroxybiphenyl from PCDD and PCDF. The hydroxylated ring is oxidized at meta-position by the dioxygenases. The ring-opened products are metabolized further yielding chlorinated catechol and chlorinated salicylates from PCDD and PCDF, respectively. PCDF having chloro groups on both rings results in formation of chlorinated 2-methyl-4H-chroman-4-ones (Keim et al. 1999; Fukuda et al. 2002). Dioxygenase of some bacterial strains such as Beijerinckia sp. and Alcaligenes sp. results in deoxygenation at lateral position, which leads to dihydrodiol and dihydroxy-2-CDD metabolite formation (Klecka and Gibson 1980; Parsons and Storms 1989). Bacillus megaterium producing cytochrome P450 causes monooxygenation of 2,3-DCDD and 2,3,7-TCDD, producing its hydroxylated metabolites (Sulistyaningdyah et al. 2004).
Catabolic biodegradation pathway of dibenzo-p-dioxin and dibenzofuran. Dibenzo-p-dioxin and dibenzofuran are acted upon by angular dioxygenase (DbfA) producing 2,2′,3-trihydroxybiphenyl. It is acted upon by 2,2′,3-trihydroxybiphenyl dioxygenase (DbfB) to form 2,6 dioxo-6-phenylhexa-3-enoate. It is further mineralized by hydrolase to form catechol and salicylaldehyde, respectively. These intermediates forms acetyl Co-A and enters into the Krebs cycle
Anaerobic biodegradation of PCDD/Fs to lower chlorinated dioxin occurs by an electron-donating substrate by the process of reductive dechlorination (Bunge et al. 2003). Due to their hydrophobic nature, these compounds are poorly bioavailable to the environment and surroundings. As reported by Kao et al. (2001), the aqueous solubility and logarithm value of octanol-water coefficient (logP) of 2,3,7,8-TeCDD is 0.019 ppb and 6.5, respectively. Thus, microbial derived surfactants can be an effective measure for increasing the bioavailability of these compounds into microbial cells. For increased access of pollutants to the microbes, an approach involving membrane protein modification assists transformation of hydrophobic POPs into the cells. For example, Sphingomonas wittichii strain RW1 was genetically engineered to express super channel membrane proteins which enabled macromolecule diffusion into the cells (Aso et al. 2006). The overexpression of super channels on this bacterial membrane improved the metabolism of dibenzofuran. This suggested that the channeling proteins can be genetically recruited for dibenzofuran degradation. Abiotic redox reaction combined with microbial catabolic actions also augments the bioremediation potential of POPs (Jeon et al. 2013).
Emerging POPs and degradative enzymes
The emerging contaminants in the environment are of major concern with myriad toxic effects. They act as endocrine-disrupting chemicals by mimicking and blocking the function of hormones in the endocrine system. This affects the health of human and animal species. Therefore, microbial degradation is the natural way of treating these emerging contaminants. Ying et al. (2008) studied the degradation of bisphenol A (BPA), 4-tert-octylphenyl, 4-n-nonylphenol and natural, synthetic estrogens in various environmental matrices such as groundwater and surface water. A similar study conducted by Hernandez-Raquet et al. (2007) showed β-estradiol degradation of nonylphenol is more favorable in aerobic than anaerobic conditions. However, there are several other enzymes utilized for the detoxification and elimination of these hydrophobic xenobiotics in the environment.
The enzyme cytochrome P450s (CYPs) are extensively dispersed hemoproteins helping in biosynthesis of endogenous compounds. They aid in oxidative detoxification and elimination of hydrophobic xenobiotics including pollutants, drugs, and pesticides (Omura 1999). It helps in detoxification of trichloroethene (TCE), 1,1,1-trichloroethane (TCA), chloroform, and benzene. These CYPs help in microbial natural product synthesis, as biocatalysts, for bioremediation and also as drug and agrochemical targets. It has been reported to commence the initial oxidation of carbon sources such as alkanes by Candida sp. Bacterial enzyme CYP177A1 has also been reported to detoxify hexahydro-1,3.5-trinitro-1,3.5-triazine (RDX) followed by phytoremediation (Rylott et al. 2006). White and brown rot fungi are the major microbes harboring these enzymes. Use of several polycyclic aromatic compounds as inducer was examined by determining the CYP gene expression pattern in fungus (Syed et al. 2010). These fungi are of further interest in commercial bioremediation as they also degrade lignin (Matsuzaki and Wariishi 2005). Likewise, CYP249 enzyme was found in Rhodococcus and Gordonia which assisted the degradation of petrol additives such as methyl tert-butyl ether, ethyl tert-butyl ether, and tert-amyl butyl ether (Malandain et al. 2010). Cytochrome P450 alkane hydroxylase constitutes a superfamily of ubiquitous heme-thiolate monooxygenases with an important role in the microbial degradation of oil, chlorinated hydrocarbons, fuel additives, and many other compounds (Van Beilen and Funhoff 2007). Cytochrome P450 enzyme system are useful for biodegradation of petroleum hydrocarbons. Yeasts such as Candida maltosa, Candida tropicalis, and Candida apicola also synthesize these cytochrome P450 enzymes (Scheller et al. 1998).
Another important enzyme required for degradation of hydrophobic toxic contaminants is the glutathione-S-transferase. Bacterial glutathione-S-transferases (GSTs) are part of an enzyme superfamily that plays a crucial role in cellular detoxification (Rossjohn et al. 1998). Several classes of prokaryotes have these enzyme implicated in biodegradation of xenobiotics, and protection against chemical and oxidative stresses and antimicrobial drug resistance (Allocati et al. 2009). In addition to that, bacterial GSTs aid in biotransformation of dichloromethane, degradation of lignin, atrazine (ATZ), and reductive dechlorination of pentachlorophenol. The electrophilic groups of a wide range of hydrophobic toxic compounds are attacked by the tripeptide glutathione (GSH) (catalyzing nucleophilicity) which promote the excretion of these toxic compunds from the cell (Hayes et al. 2005). In a study by McGuiness et al. (2007), a specific bacterial glutathione-S-transferase (GST) BphKLB400 [wild type and mutant (Ala180Pro)] was expressed by Burkholderia xenovorans LB400, which could dehalogenate toxic chlorinated organic pesticides. This mechanism protected the inoculated pea plants from the effects of a chlorinated organic pesticide, chloromequat chloride (McGuinness et al. 2007). Besides these, the huge uncultivated overabundance of microbial genome can be explored by discovering novel enzymes and proteins for bioremediation of toxic organic contaminants.
Molecular approaches to study microorganisms for bioremediation
Microorganisms encompass the overwhelming majority of life forms having miscellaneous functions. Explorations by environmental microbiologists estimate that less than 2 % of bacteria have been cultured in the laboratory. Thus, bacterial diversity on earth is still a domain to be disclosed further (Wade 2002). The uprising field of metagenomics has revolutionized microbiology by offering a glance on the massive and unknown world of microbes. DNA directly extracted from environmental samples endeavors in combination of different techniques for genetic characterization and diversity analysis. DNA-DNA hybridization technique is a major approach for illuminating the genomic imprints of unculturable bacteria and their gene expression analysis in presence of toxic compounds in the environment (Saylor and Layton 1990; Leadbetter 2003). The technique of direct genomic cloning is an approach to discover unknown sequences and their function in an ecosystem. Construction of a metagenomic library is a modified approach for deciphering the genetic fingerprint for elucidation of novel catabolic genes (Kakirde et al. 2010). Rhee et al. (2004) developed a comprehensive 50-mer-based oligonucleotide microarray to analyze naphthalene-amended enrichment and soil microcosms. This consisted of the 2402 known genes and pathways involved in biodegradation and metal resistance containing 1662 unique and group-specific probes with <85 % similarity to their nontarget sequences. Rhodococcus was dominant in naphthalene-degrading enrichments whereas Ralstonia, Comamonas, and Burkholderia were most abundant in the soil microcosms.
Function-driven screening based on the functional activity screening of cloned genes is a simple, successful method utilized for discovering potent catabolic genes (Rondon et al. 2000). A novel functional screening system of metagenome extract thin-layer chromatography (META) was developed based on the principle of high-performance thin-layer chromatography (HPTLC). This was used as a functional screening method for the rapid detection of glycosyltransferase (GT) and many other flavonoid-modifying enzymes from the metagenomic clone libraries (Rabausch et al. 2013). On contrary, it has certain limitations as expression of the functional gene of interest in numerous host cells is a laborious process. Therefore, an alternative technique was discovered known as the sequence-driven approach for the identification of genes based on their conserved regions in varied microbial genome (Schloss and Handelsman 2003). Hybridization probes and PCR primers are designed from the sequence database for clone library screening. Gene targeting by gene-specific primers and genome walking is effective for screening novel genes irrespective of gene expression (Culligan et al. 2014). However, this screening-based quantitation sometimes leads to prediction of DNA consensus which might be nonfunctional as catabolic genes. Due to lack in acquisition of full-length genes or full gene clusters, the desired product might not be obtained (Tyson et al. 2004; Venter et al. 2004). Both the screening strategies are helpful in selecting novel catabolic genes, but they are labor-intensive due to low frequency hits of clones (Henne et al. 2000).
Considering above shortcomings, another technique known as substrate-induced gene-expression (SIGEX) has been developed. It involves the screening of a metagenome library for acquiring environmental stimuli-based catabolic genes. The basic principle is the catabolic gene expression induced by specific compounds (substrates or metabolites). Sometimes, it might be regulated by elements adjoining the catabolic genes. SIGEX is a widely accepted technique, which not only facilitates cloning of various catabolic genes in short time span but also explores novel genes that are otherwise intricate to track (Uchiyama and Watanabe 2007). This technique was successfully used for screening aromatic hydrocarbon-induced genes from a constructed groundwater metagenome library as well as characterization of a phenol degradation operon from Ralstonia eutropha, an organism isolated from sludge (Uchiyama et al. 2005). In a study, 384 putative aromatic-inducible clones were recovered from the polycyclic aromatic hydrocarbon metagenomic library using SIGEX. Of which, 96 clones were highly similar in sequence with the aromatic-degrading genes or operons from the genus Pseudomonas (Meier 2014). Therefore, metagenome is defined as all the genetic material present in an environmental sample, consisting of genomes of many individual organisms. It provides genetic information on potential novel catalysts or enzymes, genomic linkages, and phylogenetic relation of uncultured organisms (Thomas et al. 2012).
Stable isotope probing (SIP) is another imperative technique, which links metabolic potential to phylogenetic and metagenomic information within a community (Abram 2015). It tracks isotopically labeled substances into phylogenetic and functional biomarkers. This tool identifies active members of the microbial community with essential functional potential. SIP integrated metagenomics provides a deeper insight into the application of genes useful for biodegradation of naphthalene, polychlorobiphenyl, benzene, etc. (Wang et al. 2012). The frequency of clones bearing target gene is increased fourfold when SIP is succeeded by metagenomic library construction (Uhlik et al. 2013). SIP was first used by Dumont and Murrell (2005) with the combination of function and sequence-based metagenomic library screening. Soil was incubated with 13CH4, followed by construction of a 13C-DNA metagenomic library using a bacterial artificial chromosome (BAC). Library screening resulted in the discovery of a clone carrying pmoCAB operon, encoding methane monooxygenase for trichloroethylene degradation.
SIP meta-transcriptomics is another approach providing high sensitivity functional information of genes useful in biodegradation of toxic organic contaminants. This was demonstrated by presence of naphthalene dioxygenase and methane monooxygenase gene in unculturable Pseudomonas sp. and Acidovorax sp. having a key role in naphthalene biodegradation (Huang et al. 2009). Therefore, unculturable microorganisms could play active roles in biodegradation in the ecosystem. Another combined technique of RNA SIP-Raman-fluorescence in situ hybridization is noteworthy for resolving ulcultivable microbial ecology, functionality, and niche specialization vested in the natural environment (Huang et al. 2009).
Fluorescence in situ hybridization (FISH) uses rRNA-targeted oligonucleotide probes for taxonomic identification of microbial cells (Wagner et al. 2003). FISH combined with SIP associates microbial phylogeny to metabolic activity at the single-cell level. Thus, unraveling the potent catabolic genes from the entire metagenome by these numerous techniques can help in deducing novel gene expression for enhanced bioremediation.
POPs degrading genes from the metagenome
Microbial metagenome constitutes the largest reservoir of genes with diverse enzymatic activities implicated in degradation (Galvão et al. 2005). Extensive research is yet to be conducted to discover various microbial species from the environment having functional roles in POP biodegradation (Hill et al. 2010). Detection of a particular gene of interest by studying community metagenomics is improbable due to vast diversity and gene abundance in the microbial ecosystem. This shortcoming can be surmounted by targeting metagenomics to specific subpopulations that may have the probability to contain the gene of interest (Schloss and Handelsman 2003). Investigation of common effluent treatment plant (CETP) metagenome has enabled the linkage of taxonomic and catabolic diversity in discovering novel biodegradation genes/pathways of toxic organic effluents (More et al. 2014). Oxygenase is the primary enzyme for biodegradation of toxic organic pollutants belonging to the oxidoreductase group of enzymes. Their primary function includes oxidation of reduced substrates by transferring molecular oxygen (O2) utilizing FAD/NADH/NADPH as a cosubstrate (Karigar and Rao 2011). Jadeja et al. (2014) have explored and documented the actual abundance of oxygenases in a target CETP from already reported metagenome sequence toward degradation of naphthalene, anthracene, phenol, biphenyl and o-toluidine. Metagenomics unlocking the black-box of potent catabolic genes illustrated the presence of homogentisate 1,2-dioxygenase, phenylacetate Co-A oxygenase, phenol dioxygenase, benzene, and toluene dioxygenase.
Three metagenomic datasets of sewage effluent treatment plant used were from biological phosphorus removal treatment plant and tannery waste metagenomes of Hong Kong, Denmark, and China, respectively (Jadeja et al. 2014). Iwai et al. (2009) used gene-targeted-metagenomics and pyrosequencing method for obtaining a better understanding of ecology and sequence depth of biphenyl dioxygenase genes. The substrate-specific primers for this gene yielded 2000 and 604 sequences from 5′ and 3′ ends of the PCR products, respectively. Complete linkage clustering determined that 95 % and 41 % of the valid sequences were allocated to 22 and 3 novel clusters. Exploring further, metagenomic libraries were constructed from Turban Basin elucidating the presence of thermostable pyrethroid-hydrolyzing enzyme (Fan et al. 2012). Moreover, current metagenomic survey on a highly contaminated hexachlorocyclohexane (HCH) dumpsite revealed the enrichment of Sphingomonadaceae, as well as lin genes (used in HCH degradation), plasmids, and transposons with increasing HCH contamination. These metagenomic data were used to construct ancestral genotype reconstructs delivering the linkage of genes from the ancestors (Sangwan et al. 2012, 2014). Sites contaminated with chlorinated pesticides were also evaluated for the analysis of microbial communities which detected dehydrodechlorinase (linA) gene variants involved in gamma-hexachlorocyclohexane (c-HCH, lindane) degradation. This linA gene could be cloned, expressed in desirable hosts having further utility in enzymatic bioremediation.
Construction of a fosmid library was reported by means of the metagenomic DNA from aerobic and anaerobic enrichments of a biodegraded petroleum sample. Hexadecane screening from the library identified 72 positive clones from a total of 5000 fosmid clones out of which five were able to degrade 70 % of hexadecane in chromatographic assays. The sequencing of the genes unraveled novel arrangements of hexadecane degradation genes (Sierra-García et al. 2014). Additionally, metagenomic DNA characterization from freshwater and marine sediments illustrated the presence and diversity of potent DDT, HCH, and ATZ degrading genes such as hdt, hdg, and atzB genes, encoding hydratase, dehalogenase, and ethylaminohydrolase activity, respectively (Fang et al. 2014). Another novel 2,4-dichlorophenol hydroxylase (TfdB, EC 1.14.13.20) gene (tfdB-JLU) was discovered by functional screening from a polychlorinated biphenyl-contaminated soil metagenome. The gene could effectively degrade ortho-substituted dichlorophenols, 2-chlorophenol, and 3-chlorophenol with respect to 2,4-dichlorophenol (Lu et al. 2011). Thus, the recent techniques and ideas in exploring metagenome can serve to reveal the secret lives of unculturable microorganisms which are also essential to the functioning of our natural environment.
Conclusion
Growing industrial activities and urbanization cause environmental pollution which release toxic POPs such as pesticides, polychlorobiphenyl, dioxins, and other emerging contaminants affecting the ecosystem stability. Several natural communities of aerobic and anaerobic microorganisms such as Aeromicrobium, Bacillus, Brevibacterium, Burkholderia, Desulfotomaculum, Desulfovibrio, Dietzia, Escherichia, Gordonia, Mycobacterium, Pseudomonas, Rhodococcus, Sphingobium, and Syntrophobacter exist with adaptable catabolic potential triggering the breakdown of these POPs. Microbial bioremediation is the paramount cost-effective approach in today’s world for biotransformation of POPs to their nontoxic forms. They harbor catabolic genes in their genome, plasmid, and transposon expressing diverse aromatic monooxygenase, dioxygenase, cytochrome P450, and glutathione-S-transferase enzymes for the degradation and removal of POPs from the environment. However, most of the microorganisms are still uncultured in the laboratory and the concept of metagenomics appeared to study this unculturable microorganisms. Metagenomics is opening the access to the world of uncultivated microorganisms by finding the unique genes from varied environmental resources. Advancements in gene sequencing and sequence-driven screening have facilitated the illustration of novel genes from various environmental samples. In addition, technique of SIGEX has also emerged to address the bioremediation of POPs. However, further development of innovative strategies is essentially required for opening the black box of hidden microbes with untapped potential. Further exploration of techniques and microbes will provide platform for the development of new groundbreaking screening methods that will be indispensable to expand the range of accessible microbes and genes in environmental metagenomes.
References
Abram F (2015) Systems-based approaches to unravel multi-species microbial community functioning. Comput Struc Biotechnol J 13:24–32
Adebusoye SA, Ilori MO, Picardal FW, Amund OO (2008) Cometabolic degradation of polychlorinated biphenyls (PCBs) by axenic cultures of Ralstonia sp. strain SA-5 and Pseudomonas sp. strain SA-6 obtained from Nigerian contaminated soils. World J Microb Biot 24:61–68
Admire B, Lian B, Yalkowsky SH (2014) Estimating the physicochemical properties of polyhalogenated aromatic and aliphatic compounds using UPPER: Part 2. Aqueous solubility, octanol solubility and octanol–water partition coefficient. Chemosphere 119:1441–1446
Afzal M, Khan QM, Sessitsch A (2014) Endophytic bacteria: prospects and applications for the phytoremediation of organic pollutants. Chemosphere 117:232–242
Agency for toxic substances and disease registry (ATSDR) (2002) Public Health Statement for DDT, DDE, and DDD. http://www.atsdr.cdc.gov/phs/phs.asp?id=79&tid=20. Accessed 21 Jan 2015
Allocati N, Federici L, Masulli M, Di Ilio C (2009) Glutathione transferases in bacteria. Febs J 276:58–75
Ang EL, Zhao H, Obbard JP (2005) Recent advances in the bioremediation of persistent organic pollutants via biomolecular engineering. Enzyme Microb Technol 37:487–496
Arber W (2000) Genetic variation: molecular mechanisms and impact on microbial evolution. FEMS Microbiol Rev 24:1–7
Arslan M, Imran A, Khan QM, Afzal M (2015) Plant–bacteria partnerships for the remediation of persistent organic pollutants. Environ Sci Pollut Res 1–15, [Epub ahead of print]
Aslanzadeh J, Hedrick HG (1985) Search for Mirex-degrading soil microorganisms. Soil Science 139:369–374
Aso Y, Miyamoto Y, Harada KM, Momma K, Kawai S, Hashimoto W (2006) Engineered membrane superchannel improves bioremediation potential of dioxin-degrading bacteria. Nat Biotechnol 24:188–9
Bala K, Sharma P, Lal R (2010) Sphingobium quisquiliarum sp. nov., a hexachlorocyclohexane (HCH) degrading bacterium isolated from an HCH-contaminated soil. IJSEM 60:429–43
Barriault D, Plant MM, Sylvestre M (2002) Family shuffling of a targeted bphA region to engineer biphenyl dioxygenase. J Bacteriol 184:794–800
Bedard DL (2008) A case study for microbial biodegradation: anaerobic bacterial reductive dechlorination of polychlorinated biphenyls-from sediment to defined medium. Annual Rev Microbiol 62:253–270
Beunink J, Rehm HJ (1988) Synchronous anaerobic and aerobic degradation of DDT by an immobilized mixed culture system. Appl Microbiol Biotechnol 29:72–80
Black JG (1999) Bioremediation. En: Microbiology: Principles and Explorations, 4th edn. Prentice Hall Publishers, Upper Saddle River, pp 751–752
Borgeest C, Greenfeld C, Tomic D, Flaws JA (2002) The effects of endocrine disrupting chemicals on the ovary. Front Biosci 7:1941–1948
Bruhlmann F, Chen W (1999) Turning biphenyl dioxygenase for extended substrate specificity. Biotechnol Bioeng 63:544–51
Bumpus JA, Tien M, Wright D, Aust SD (1985) Oxidation of persistent environmental pollutants by a white rot fungus. Science 228:1434–1436
Bunge M, Adrian L, Kraus A, Lorenz WG, Andreesen JR, Gorisch H, Lechner U (2003) Reductive dehaogenation of chlorinated dioxins by the anaerobic bacterium Dehalococcoides ethenogenes genes sp. strain CBDBI. Nature 421:357–360
Bunge M, Ballerstedt H, Lechner U (2001) Regiospecific dechlorination of spiked tetra and trichlorodibenzo-p-dioxins by anaerobic bacteria from PCDD/F-contaminated Spittelwasser sediments. Chemosphere 43:675–681
Bylaska EJ, Dixon DA, Felmy AR, Aprà E, Windus TL, Zhan CG, Tratnyek PG (2004) The energetics of the hydrogenolysis, dehydrohalogenation, and hydrolysis of 4, 4′-dichloro-diphenyl-trichloroethane from ab initio electronic structure theory. J Phys Chem A 108(27):5883–5893
Chanama M, Chanama S (2011) Expression of pentachlorophenol-degradative genes of Sphingobium chlorophenolica ATCC39723 in Escherichia coli. Asia J Public Health 2:78–83
Chakraborty J, Das S (2014) Biosurfactant based bioremediation of toxic metals. In: Microbial Biodegradation and Bioremediation. Elsevier, pp 167–201
Chakraborty J, Das S (2016) Characterization of the metabolic pathway and catabolic gene expression in biphenyl degrading marine bacterium Pseudomonas aeruginosa JP-11. Chemosphere 144:1706–1714
Chattaraj PK, Sarkar U, Roy DR (2006) Electrophilicity index. Chem Rev 106:2065–2091
Chaudhry GR, Chapalamadugu S (1991) Biodegradation of halogenated organic compounds. Microbiol Rev 55:59–79
Chavez FP, Lunsdorf H, Jerez CA (2004) Growth of polychlorinated biphenyl degrading bacteria in the presence of biphenyl and chlorobiphenyls generates oxidative stress and massive accumulation of inorganic polyposhphate. Appl Environ Microbiol 70:3064–3072
Chowdhury A, Pradhan S, Saha M, Sanyal N (2008) Impact of pesticides on soil microbiological parameters and possible bioremediation strategies. Indian J Microbiol 48:114–127
Covaci A, Harrad S, Abdallah MAE, Ali N, Law RJ, Herzke D, de Wit CA (2011) Novel brominated flame retardants: a review of their analysis, environmental fate and behaviour. Environ Int 37:532–556
Culligan EP, Sleator RD, Marchesi JR, Hill C (2014) Metagenomics and novel gene discovery: promise and potential for novel therapeutics. Virulence 5:399–412
Cvančarová M, Křesinová Z, Filipová A, Covino S, Cajthaml T (2012) Biodegradation of PCBs by ligninolytic fungi and characterization of the degradation products. Chemosphere 88:1317–23
Cvitkovitch DG, Li YH, Ellen RP (2003) Quorum sensing and biofilm formation in Streptococcal infections. J Clin Invest 112:1626–1632
Das S, Dash HR, Chakraborty J (2016) Genetic basis and importance of metal resistant genes in bacteria for bioremediation of contaminated environments with toxic metal pollutants. Appl Microbiol Biotechnol 100:2967–2984
de Carvalho CCCR, Wick LY, Heipieper HJ (2009) Cell wall adaptations of planktonic and biofilm Rhodococcus erythropolis cells to growth on C5 to C16 n-alkane hydrocarbons. Appl Microbiol Biotechnol 82:311–320
De J, Ramaiah N, Sarkar A (2006) Aerobic degradation of highly chlorinated polychlorobiphenyls by a marine bacterium, Pseudomonas CH07. World J Microbiol Biotechnol 22:1321–1327
DeWeerd KA, Mandelco L, Tanner RS, Woese CR, Suflita JM (1990) Desulfomonile tiedjei gen nov and a novel anaerobic dehalogenating sulfate-reducing bacterium. Arch Microbiol 154:23–30
Dileep K (2008) Biodegradation and bioremediation of pesticide in soil: concept, method and recent developments. Indian J Microbiol 48:35–40
Don RH, Pemberton JM (1981) Properties of six pesticides degradation plasmids isolated from Alcaligenes paradoxus and Alcaligenes eutrophus. J Bacteriol 145:681–686
Dumont MG, Murrell JC (2005) Stable isotope probing-linking microbial identity to function. Nat Rev Microbiol 3:499–504
El Fantroussi S, Agathos SN (2005) Is bioaugmentation a feasible strategy for pollutant removal and site remediation? Curr Opin Microbiol 8:268–275
Erickson BD, Mondello FJ (1992) Nucleotide sequencing transcriptional mapping of the genes encoding biphenyl dioxygenase, a multicomponent polychlorinated biphenyl degrading enzyme in Pseudomonas strain LB400. Appl Environ Microbiol 174:2903–12
Erickson BD, Mondello FJ (1993) Enhanced biodegradation of polychlorinated-biphenyls after site-directed mutagenesis of a biphenyl dioxygenase gene. Appl Environ Microbiol 59:3858–62
Fan X, Liu X, Huang R, Liu Y (2012) Identification and characterization of a novel thermostable pyrethroid-hydrolyzing enzyme isolated through metagenomic approach. Microb Cell Fact 11:1
Fang H, Cai L, Yang Y, Ju F, Li X, Yu Y, Zhang T (2014) Metagenomic analysis reveals potential biodegradation pathways of persistent pesticides in freshwater and marine sediments. Sci Total Environ 470:983–992
Francino MP (2012) The ecology of bacterial genes and the survival of the new. Int J Evol Biol 2012:394026
Fries GR, Marrow GS, Gordon CH (1969) Metabolism of o, p’-and p, p’-DDT by rumen microorganism. J Agri Food Chem 17:860–862
Fritsche W and Hofrichter M (2000) Aerobic degradation by microorganisms, in environmental processes. Soil decontamination J Klein Ed. Wiley-VCH Weinheim Germany pp. 146–155
Fritsche W and Hofrichter M (2004) Aerobic degradation of recalcitrant organic compounds by microorganisms. Environmental Biotechnology. Concepts and Applications. (eds HJ Jördening and J Winter), Wiley-VCH Verlag GmbH & Co. KGaA, Weinheim, FRG
Fukuda K, Nagata S, Taniguchi H (2002) Isolation and characterization of dibenzofuran-degrading bacteria. FEMS Microbiol Lett 208:179–185
Furukawa K (2000) Engineering dioxygenases for efficient degradation of environmental pollutants. Curr Opin Biotechnol 11:244–9
Furukawa K, Fujihara H (2008) Microbial degradation of polychlorinated biphenyls: biochemical and molecular features. J Biosc Bioeng 105:433–449
Futagami T, Goto M, Furukawa K (2008) Biochemical and genetic bases of dehalorespiration. The Chem Rec 8:1–12
Galvão TC, Mohn WW, de Lorenzo V (2005) Exploring the microbial biodegradation and biotransformation gene pool. Trends Biotechnol 23:497–506
Garg N, Bala K, Lal R (2012) Sphingobium lucknowense sp. nov. a hexachlorocyclohexane (HCH)-degrading bacterium isolated from HCH-contaminated soil. IJSEM 62:618–623
George SE, Claxton LD (1988) Biotransformation of chlordecone by Pseudomonas species. Xenobiotica 18:407–416
Ghazali FM, Rahman RNZA, Salleh AB, Basri M (2004) Biodegradation of hydrocarbons in soil by microbial consortium. Int Biodeter Biodegr 54:61–67
Gibson DT, Cruden DL, Haddock JD, Zylstra GJ, Brand JM (1993) Oxidation of polychlorinated-biphenyls by Pseudomonas sp. strain LB400 and Pseudomonas pseudoalcaligenes KF707. J Bacteriol 175:4561–4564
Gienfrada L, Rao MA (2008) Interactions between xenobiotics and microbial and enzymatic soil activity. Crit Rev Environ Sci Technol 38:269–310
Gooch JW, Matsumura F (1985) Evaluation of the toxic components of toxaphene in Lake Michigan lake trout. J Agric Food Chem 33:844–848
Gouin T, Mackaya D, Jones KC, Harner T, Meijer SN (2004) Evidence for the “grasshopper” effect and fractionation during long-range atmospheric transport of organic contaminants. Environ Pollut 128:139–148
Habe H, Chung JS, Lee JH, Kasuga K, Yoshida T, Nojiri H, Omori T (2001) Degradation of chlorinated dibenzofurans and dibenzo-p-dioxins by two types of bacteria having angular dioxygenases with different features. Appl Environ Microbiol 67:3610–3617
Haggblom, MM, Bossert ID (2004) Halogenated organic compounds-a global perspective. In Dehalogenation Springer US pp. 3–29
Hay AG, Focht DD (1998) Cometabolism of 1, 1-dichloro-2, 2-bis (4-chlorophenyl)ethylene by Pseudomonas acidovorans M3GY grown on biphenyl. Appl Environ Microbiol 64:2141–6
Hayes JD, Flanagan JU, Jowsey IR (2005) Glutathione transferases. Annu Rev Pharmacol Toxicol 45:51–88
Henderson IR, Owen P, Nataro JP (1999) Molecular switches—the ON and OFF of bacterial phase variation. Mol Microbiol 33:919–932
Henne A, Schmitz RA, Bomeke M, Gottschalk G, Daniel R (2000) Screening of environmental DNA libraries for the presence of genes conferring lipolytic activity on Escherichia coli. Appl Environ Microbiol 66:3113–3116
Hernandez-Raquet G, Soef A, Delgenès N, Balaguer P (2007) Removal of the endocrine disrupter nonylphenol and its estrogenic activity in sludge treatment processes. Water Res 41:2643–2651
Heuer H, Smalla K (2012) Plasmids foster diversification and adaptation of bacterial populations in soil. FEMS Microbiol Rev 36:1083–1104
Hill JE, Fernando WU, Zello GA, Tyler RT, Dahl WJ, Van Kessel AG (2010) Improvement of the representation of bifidobacteria in fecal microbiota metagenomic libraries by application of the cpn60 universal primer cocktail. Appl Environ Microbiol 76:4550–4552
Hiraishi A, Yoshida N, Takahashi N (2005) Phylogenetic characterization of a polychlorinated-dioxin dechlorinating microbial community by use of microcosm studies. Appl Environ Microbiol 71:4325–4334
Hofrichter M, Bublitz F, Fritsche W (1994) Unspecific degradation of halogenated phenols by the soil fungus Penicillium frequentans Bi7/2. J Basic Microbiol 34:163–172
Houghton JE, Shanley MS (1994) Catabolic potential of Pseudomonads: a regulatory perspective. In: Chaudhry GR (ed) Biological degradation and bioremediation of toxic chemicals. Dioscorides press, Portland pp, pp 11–32
Hrywna Y, Tsoi TV, Maltseva OV, Quensen JF, Tiedje JM (1999) Construction and characterization of two recombinant bacteria that grow on ortho- and para-substituted chlorobiphenyls. Appl Environ Microbiol 65:2163–2169
Huang WE, Ferguson A, Singer AC, Lawson K, Thompson IP, Kalin RM, Whiteley AS (2009) Resolving genetic functions within microbial populations: in situ analyses using rRNA and mRNA stable isotope probing coupled with single-cell Raman-fluorescence in situ hybridization. Appl Environ Microbiol 75:234–41
Huang Y, Wang J (2013) Degradation and mineralization of DDT by the ectomycorrhizal fungi Xerocomus chrysenteron. Chemosphere 92:760–4
Hutz RJ (1999) Reproductive endocrine disruption by environmental xenobiotics that modulate the estrogen-signaling pathway, particularly tetrachlorodibenzo-p-dioxin (TCDD). J Reprod Dev 45:1–12
Ikuma K (2011) The effect of select biological and environmental factors on the horizontal gene transfer and functionality of the TOL plasmid: a case study for genetic bioaugmentation. Duke University, Doctoral dissertation
Isken S, de Bont JA (1998) Bacteria tolerant to organic solvents. Extremophiles 2:229–238
Iwai S, Chai B, Sul WJ, Cole JR, Hashsham SA, Tiedje JM (2009) Gene-targeted-metagenomics reveals extensive diversity of aromatic dioxygenase genes in the environment. ISME J 4:279–285
Jadeja NB, More RP, Purohit HJ, Kapley A (2014) Metagenomic analysis of oxygenases from activated sludge. Bioresour Technol 165:250–256
Jaiswal PK, Thakur IS (2007) Isolation and characterization of dibenzofuran-degrading Serratia marcescens from alkalophilic bacterial consortium of the chemostat. Curr Microbiol 55:447–454
Jayaraman R (2011) Hypermutation and stress adaptation in bacteria. J Genetics 90:383–391
Jefferson KK (2004) What drives bacteria to produce a biofilm? FEMS Microbiol Lett 236:163–173
Jeon JR, Murugesan K, Nam IH, Chang YS (2013) Coupling microbial catabolic actions with abiotic redox processes: a new recipe for persistent organic pollutants (POPs) removal. Biotechnol Adv 31:246–256
Jha SK, Jain P, Sharma HP (2015) Xenobiotic degradation by bacterial enzymes. Int J Curr Microbiol App Sci 4:48–62
Joesten MD, Hogg JL, Castellion ME (2006) The world of chemistry: essentials: essentials. Cengage Learning
Juhasz A, Naidu R (2000) Enrichment and isolation of non-specific aromatic degraders from unique uncontaminated (plant and fecal material) sources and contaminated soils. J Appl Microbiol 89:642–650
Kaiya S, Utsunomiya S, Suzuki S, Yoshida N, Futamata H, Yamada T, Hiraishi A (2012) Isolation and functional gene analyses of aromatic-hydrocarbon-degrading bacteria from a polychlorinated-dioxin-dechlorinating process. Microbes Environ 27:127–35
Kakirde KS, Parsley LC, Liles MR (2010) Size does matter: application-driven approaches for soil metagenomics. Soil Biol Biochem 42:1911–1923
Kamanavalli CM, Ninnekar HZ (2004) Biodegradation of DDT by a Pseudomonas sp. Curr Microbiol 48:10–13
Kao CM, Chen SC, Liu JK, Wu MJ (2001) Evaluation of TCDD biodegradability under different redox conditions. Chemosphere 44:1447–1454
Karigar CS, Rao SS (2011) Role of microbial enzymes in the bioremediation of pollutants: a review. Enz Res 805187
Kataoka R, Takagi K, Kamei I, Kiyota H, Sato Y (2010) Biodegradation of dieldrin by a soil fungus isolated from a soil with annual endosulfan applications. Environ Sci Technol 44:6343–9
Keim T, Francke W, Schmidt S, Fortnagel P (1999) Catabolism of 2, 7 dichloro and 2, 4, 8-trichlorodibenzofuran by Sphingomonas sp. strain RW1. J Ind Microbiol Biotechnol 23:359–363
Khan AA, Wang RF, Cao WW, Doerge DR, Wennerstrom D, Cerniglia CE (2001) Molecular cloning, nucleotide sequence and expression of genes encoding a polycyclic ring dioxygenase from Mycobacterium sp. strain PYR-1. Appl Environ Microbiol 67:3577–3585
Kidd KA, Becher G, Bergman Å, Muir DC, Woodruff TJ (2012) Human and wildlife exposures to EDCs. State of the Science of Endocrine Disrupting Chemicals 1–261
Kimbara K (2005) Recent developments in the study of microbial aerobic degradation of polychlorinated biphenyls. Microbes Environ 20:127–134
Kimura N, Nishi M, Goto M, Furukawa K (1997) Functional analyses of a variety of chimeric dioxygenases constructed from two biphenyl dioxygenases that are similar structurally but different functionally. J Bacteriol 179:3936–3943
Kitagawa W, Takami S, Miyauchi K, Masai E, Kamagata Y, Tiedje JM, Fukuda M (2002) Novel 2, 4-dichlorophenoxyacetic acid degradation genes from oligotrophic Bradyrhizobium sp. strain HW13 isolated from a pristine environment. J Bacteriol 184:509–18
Klecka GM, Gibson DT (1980) Metabolism of dibenzo-para-dioxin and chlorinated dibenzo-para-dioxins by a Beijerinckia species. Appl Environ Microbiol 39:288–296
Kumar S (2004) Occupational exposure associated with reproductive dysfunction. J Occup Health 46:1–19
Kumari R, Subudhi S, Suar M, Dhingra G, Raina V, Dogra C, Lal R (2002) Cloning and characterization of lin genes responsible for the degradation of hexachlorocyclohexane isomers by Sphingomonas paucimobilis strain B90. Appl Environ Microbiol 68:6021–6028
La Farre M, Pérez S, Kantiani L, Barceló D (2008) Fate and toxicity of emerging pollutants, their metabolites and transformation products in the aquatic environment. TrAC Trend Anal Chem 27:991–1007
Lacayo-Romero M, Quillaguamán J, Van Bavel B, Mattiasson B (2005) A toxaphene-degrading bacterium related to Enterobacter cloacae, strain D1 isolated from aged contaminated soil in Nicaragua. Syst Appl Microbiol 28:632–639
Leadbetter JR (2003) Cultivation of recalcitrant microbes: cells are alive, well and revealing their secrets in the 21st century laboratory. Curr Opin Microbiol 6:274–281
Ledger T, Pieper DH, Gonzalez B (2006) Chlorophenol hydroxylases encoded by plasmid pJP4 differentially contribute to chlorophenoxyacetic acid degradation. Appl Environ Microbiol 72:2783–2792
Levinson G, Gutman GA (1987) Slipped-strand mispairing: a major mechanism for DNA sequence evolution. Mol Biol Evol 4:203–221
Li X, Gao Y, Wang Y, Pan Y (2014) Emerging persistent organic pollutants in Chinese Bohai Sea and its coastal regions. Sci World J 608231
Lu XX, Chen CQ, Zhang S, Ou Y, Yin L, Wu W (2012) Microbial degradation of 2, 2′,4, 4′-tetrabrominated diphenyl ether under anaerobic condition. Huan Jing Ke Xue 33:1000–1007
Lu Y, Yu Y, Zhou R, Sun W, Dai C, Wan P, Ren H (2011) Cloning and characterization of a novel 2, 4-dichlorophenol hydroxylase from a metagenomic library derived from polychlorinated biphenyl-contaminated soil. Biotechnol Lett 33:1159–1167
Malandain C, Fayolle-Gulchard F, Vogel TM (2010) Cytochromes P450-mediated degradation of fuel oxygenates by environmental isolates. FEMS Microbiol Ecol 72:289–296
Mangwani N, Shukla SK, Rao TS, Das S (2014) Calcium-mediated modulation of Pseudomonas mendocina NR802 biofilm influences the phenanthrene degradation. Colloids Surf B 114:301–309
Mangwani N, Kumari S, Das S (2015) Involvement of quorum sensing genes in biofilm development and degradation of polycyclic aromatic hydrocarbons by a marine bacterium Pseudomonas aeruginosa N6P6. Appl Microbiol Biotechnol 99:10283–10297
Marcobal AM, Sela DA, Wolf YI, Makarova KS, Mills DA (2008) Role of hypermutability in the evolution of the genus Oenococcus. J Bacteriol 190:564–570
Matsumoto E, Kawanaka Y, Yun SJ, Oyaizu H (2008) Isolation of dieldrin and endrin degrading bacteria using 1, 2 epoxycyclohexane as a structural analog of both compounds. Appl Microbiol Biotechnol 80:1095–1103
Matsuzaki F, Wariishi H (2005) Molecular characterization of cytochrome P450 catalyzing hydroxylation of benzoates from the white-rot fungus Phanerochaete chrysosporium. Biochem Biophys Res Comm 334:1184–1190
McGuinness MC, Mazurkiewicz V, Brennan E, Dowling DN (2007) Dechlorination of pesticides by a specific bacterial glutathione S-transferase, BphKLB400: potential for bioremediation. Eng Life Sci 7:611–615
McLeod MP, Warren RL, Hsiao WW, Araki Myhre NM, Fernandes C, Miyazawa D, Wong W, Lillquist AL, Wang D, Dosanjh Hara MH, Petrescu A, Morin RD, Yang G, Stott JM, Schein JE, Shin H, Smailus D, Siddiqui AS, Marra MA, Jones SJ, Holt R, Brinkman FS, Miyauchi K, Fukuda M, Davies JE, Mohn WW, Eltis LD (2006) The complete genome of Rhodococcus sp. RHA1 provides insights into a catabolic powerhouse. Proc Natl Acad Sci USA 103:15582–15587
Megharaj M, Ramakrishnan B, Venkateswarlu K, Sethunathan N, Naidu R (2011) Bioremediation approaches for organic pollutants: a critical perspective. Environ Int 37:1362–1375
Meier MJ (2014) Metagenomic analysis of contaminated soils. Doctoral dissertation, Carleton University Ottawa
Merlin C, Devers M, Crouzet O, Heraud C, Steinberg C, Mougin C, Martin-Laurent F (2014) Characterization of chlordecone-tolerant fungal populations isolated from long-term polluted tropical volcanic soil in the French West Indies. Environ Sci Pollut Res 21:4914–4927
Miyauchi K, Sukda P, Nishida T, Ito E, Matsumoto Y, Masai E, Fukuda M (2008) Isolation of dibenzofuran-degrading bacterium, Nocardioides sp. DF412, and characterization of its dibenzofuran degradation genes. J Biosci Bioeng 105:628–635
Mohn WW, Tiedje JM (1992) Microbial reductive dehalogenation. Microbiol Rev 56:482–507
Molin S, Nielsen TT (2003) Gene transfer occurs with enhanced efficiency in biofilms and induces stabilization of the biofilm structure. Curr Opin Biotechnol 14:255–261
More RP, Mitra S, Raju SC, Kapley A, Purohit HJ (2014) Mining and assessment of catabolic pathways in the metagenome of a common effluent treatment plant to induce the degradative capacity of biomass. Bioresour Technol 153:137–146
Morgante V, López-López A, Flores C, González M, González B, Vásquez M, Seeger M (2010) Bioaugmentation with Pseudomonas sp. strain MHP41 promotes simazine attenuation and bacterial community changes in agricultural soils. FEMS Microbiol Ecol 71:114–126
Mougin C, Pericaud C, Malosse C, Laugero C, Ashter M (1999) Biotransformation of the insecticide lindane by the white-rot basidiomycete Phanerochaete chrysosporium. Pestic Sci 41:1–59
Mrozik A, Piotrowska-Seget Z, Labuzek S (2003) Bacterial degradation and bioremediation of polycyclic aromatic hydrocarbons. Pol J Environ Stud 12(1)
Mwangi K, Boga HI, Muigai AW, Kiiyuikia C, Tsanuo MK (2010) Degradation of dichlorodiphenyltrichloroethane (DDT) by bacterial isolates from cultivated and uncultivated soil. Afr J Microbiol Res 4:185–196
Nadeau LJ, Menn FM, Breen A, Sayler GS (1994) Aerobic degradation of 1,1,1-trichloro-2,2-bis(4-chlorophenyl)ethane (DDT) by Alcaligenes eutrophus A5. Appl Environ Microbiol 60:51–55
Nagao T (1998) Health effects of endocrine disruptors. JPN J Tox Environ Health 44:151–167
Nakatsu C, Ng J, Singh R, Straus N, Wyndham C (1991) Chlorobenzoate catabolic transposon Tn5271 is a composite class I element with flanking class II insertion sequences. Pro Nat Acad Sci 88:8312–8316
Niedle EL, Harnett C, Bonitz S, Ornston LN (1988) DNA sequence of Acinetobacter calcoaceticus catechol 1, 2-dioxygenase-I structural gene catA: evidence for evolutionary divergence of intradiol dioxygenases by acquisition of DNA sequence repetitions. J Bacteriol 170:4874–4880
Nojiri H, Omori T (2002) Molecular bases of aerobic bacterial degradation of dioxins: involvement of angular dioxygenation. Biosci Biotechnol Biochem 66:2001–2016
Nzila A (2013) Update on the cometabolism of organic pollutants by bacteria. Environ Pollut 178:474–482
Omura T (1999) Forty years of cytochrome P450. Biochem Biophys Res Comm 266:690–698
Othsubo Y, Nagata Y, Kimbara K, Takagi M, Ohta A (2000) Expression of the bph genes involved in biphenyl/PCB degradation in Pseudomonas sp. KKS102 induced by the biphenyl degradation intermediate, 2-hydroxy-6-oxo-6-phenylhexa-2, 4-dienoic acid. Gene 256:223–228
Pal R, Bala S, Dadhwal M, Kumar M, Dhingra G, Prakash O, Lal R (2005) Hexachlorocyclohexane-degrading bacterial strains Sphingomonas paucimobilis B90A, UT26 and Sp+, having similar lin genes, represent three distinct species, Sphingobium indicum sp. Nov., Sphingobium japonicum sp. nov. and Sphingobium francense sp. nov., and reclassification of [Sphingomonas] chungbukensis as Sphingobium chungbukense comb. nov. Int J Syst Evol Microbiol 55:1965–1972
Parales RE, Lee K, Resnick SM, Jiang H, Lessner DJ, Gibson DT (2000) Substrate specificity of naphthalene dioxygenase: effect of specific amino acids at the active site of the enzyme. J Bacteriol 182:1641–1649
Parsons JR, Storms MCM (1989) Biodegradation of chlorinated dibenzo-para-dioxins in batch and continuous cultures of strain JB1. Chemosphere 19:1297–1308
Patil KC, Matsumura F, Boush GM (1970) Degradation of Endrin, Aldrin, and DDT by soil microorganisms. J Appl Microbiol 19:879–881
Pemberton JM, Schmidt R (2001) Catabolic plasmids. Encyclopaedia Life Science, University of Queensland, Australia
Pieper DH, Reineke W (2000) Engineering bacteria for bioremediation. Curr Opin Biotechnol 11:262–27
Pieper DH (2005) Aerobic degradation of polychlorinated biphenyls. Appl Microbiol Biotechnol 67:170–191
Qi Y, Zhao L, Olusheyi OZ, Tan X (2007) Isolation and preliminary characterization of a 3-chlorobenzoate degrading bacteria. J Environ Sci (China) 19:332–7
Quensen JF, Mueller SA, Jain MK, Tiedje JM (1998) Reductive dechlorination of DDE to DDMU in marine sediment microcosms. Science 280:722–724
Rabausch U, Juergensen J, Ilmberger N, Böhnke S, Fischer S, Schubach B, Streit WR (2013) Functional screening of metagenome and genome libraries for detection of novel flavonoid-modifying enzymes. Appl Environ Microbiol 79:4551–4563
Ramesh C, Atur K, Ajay S, Owen P (2004) Bioremediation of pesticide contaminated soils, Applied Bioremediation and Phytoremediation. Springer-verlag Berlin Heidelberg pp. 35–55
Ray MB (2000) Photodegradation of the volatile organic compounds in the gas phase: a review. Dev Chem Eng Min Proc 8:405–439
Rappe C (1991) Sources of and human exposure to PCDDs and PCDFs. In: Gallo MA et al (eds) Biological basis for risk assessment of dioxins and related compounds. Cold Spring Harbor, New York, pp 121–129
Rayssiguier C, Thaler D, Radman M (1989) The barrier to recombination between Escherichia coli and Salmonella typhimurium is disrupted in mismatch repair mutants. Nature 342:396–401
Rentz JA, Alvarez PJ, Schnoor JL (2008) Benzo[a]pyrene degradation by Sphingomonas yanoikuyae JAR02. Environ Pollut 151:669–677
Rhee SK, Liu X, Wu L, Chong SC, Wan X, Zhou J (2004) Detection of genes involved in biodegradation and biotransformation in microbial communities by using 50-mer oligonucleotide microarrays. Appl Environ Microbiol 70:4303–4317
Richardson AR, Stojiljkovic I (2001) Mismatch repair and the regulation of phase variation in Neisseria meningitidis. Mol Microbiol 40:645–655
Richardson SD, Ternes TA (2011) Water analysis: emerging contaminants and current issues. Anal Chem 83:4614–4648
Romero ML, Terrazas E, Van Bavel B, Mattiasson B (2006) Degradation of toxaphene by Bjerkandera sp. strain BOL13 using waste biomass as a cosubstrate. Appl Microbiol Biotechnol 71(4):549–554
Ron EZ, Rosenberg E (2002) Biosurfactants and oil bioremediation. Curr Opin Biotechnol 13:249–252
Rondon MR, August PR, Bettermann AD, Brady SF, Grossman TH, Liles MR, Loiacono KA, Lynch BA, MacNeil IA, Minor C, Tiong CL, Gilman M, Osburne MS, Clardy J, Handelsman J, Goodman RM (2000) Cloning the soil metagenome: a strategy for accessing the genetic and functional diversity of uncultured microorganisms. Appl Environ Microbiol 66:2541–2547
Rossjohn J, Polekhina G, Feil SC, Allocati N, Masulli M, Di Ilio C, Parker MW (1998) A mixed disulfide bond in bacterial glutathione transferase: functional and evolutionary implications. Structure 6:721–734
Rout P, Jiwal S, Sasikala C, Mohandass R (2012) Biodegradation of Benzo[a]pyrene by the mixed culture of Bacillus cereus and Bacillus vireti isolated from the petrochemical industry. J Environ Biol 33:985–989
Roy DR, Parthasarathi R, Subramanian V, Chattaraj PK (2006) An electrophilicity based analysis of toxicity of aromatic compounds towards Tetrahymena pyriformis. QSAR Comb Sci 25:114–122
Rylott EL, Jackson RG, Edwards J, Womack GL, Seth-Smith HM, Rathbone DA, Bruce NC (2006) An explosive-degrading cytochrome P450 activity and its targeted application for the phytoremediation of RDX. Nature Biotechnol 24:216–219
Sadowsky D, McNeill K, Cramer CJ (2014) Dehalogenation of aromatics by nucleophilic aromatic substitution. Environ Sci Technol 48:10904–10911
Sahu SK, Patnaik KK, Sethunathan N (1992) Dehydrochlorination of δ-isomer of hexachlorocyclohexane by a soil bacterium, Pseudomonas sp. Bull Environ Contam Toxicol 48:265–268
Sakakibara F, Takagi K, Kataoka R, Kiyota H, Sato Y, Okada S (2011) Isolation and identification of dieldrin-degrading Pseudonocardia sp. strain KSF27 using a soil-charcoal perfusion method with aldrin trans-diol as a structural analog of dieldrin. Biochem Biophys Res Commun 411:76–81
Samardjieva K, Tavares F, Pissarra J (2015) Histological and ultrastructural evidence for zinc sequestration in Solanum nigrum L. Protoplasma 252:345–357
Sangwan N, Lata P, Dwivedi V, Singh A, Niharika N, Kaur J, Lal R (2012) Comparative metagenomic analysis of soil microbial communities across three hexachlorocyclohexane contamination levels. PLoS One 7:e46219
Sangwan N, Verma H, Kumar R, Negi V, Lax S, Khurana P, Lal R (2014) Reconstructing an ancestral genotype of two hexachlorocyclohexane-degrading Sphingobium species using metagenomic sequence data. ISME J 8:398–408
Sato A, Watanabe T, Watanabe Y, Harazono K, Fukatsu T (2002) Screening for basidiomycetous fungi capable of degrading 2, 7-dichlorodibenzo-p-dioxin. FEMS Microbiol Lett 213:213–217
Saylor GS, Layton AC (1990) Environmental application of nucleic acid. Annu Rev Microbiol 44:625–648
Scheller U, Zimmer T, Becher D, Schauer F, Schunck WH (1998) Oxygenation cascade in conversion of n-alkanes to α, ω-dioic acids catalyzed by cytochrome P450 52A3. J Biol Chem 273:32528–32534
Schloss PD, Handelsman J (2003) Biotechnological prospects from metagenomics. Curr Opin Biotechnol 14:303–310
Schlüter A, Szczepanowski R, Pühler A, Top EM (2007) Genomics of IncP-1 antibiotic resistance plasmids isolated from wastewater treatment plants provides evidence for a widely accessible drug resistance gene pool. FEMS Microbiol Rev 31:449–477
Seeger M, Saavedra JM, Acevedo Gonzalez FM (2009) Mineralization of PCBs by the genetically modified strain Cupriavidus necator JMS34 and its application for bioremediation of PCBs in soil. Appl Environ Microbiol 87:1543–1554
Seeger M, Hernández M, Méndez V, Ponce B, Córdova M, González M (2010) Bacterial degradation and bioremediation of chlorinated herbicides and biphenyls. J Soil Sci Plant Nut 10:320–332
Seo JS, Keum YS, Li QX (2009) Bacterial degradation of aromatic compounds. Int J Environ Res Pub Health 6:278–309
Sharma SK, Sadasivam KV, Dave JM (1987) DDT degradation by bacteria from activated sludge. Environ Int 13:183–190
Shields MS, Hooper SW, Sayler GS (1985) Plasmid-mediated mineralization of 4-chlorobiphenyl. J Bacteriol 163:882–889
Sierra-García IN, Alvarez JC, de Vasconcellos SP, de Souza AP, de Oliveira VM, de Oliveira VM (2014) New hydrocarbon degradation pathways in the microbial metagenome from Brazilian petroleum reservoirs. PloS One 9:e90087
Singh BK, Kuhad RC (1999) Biodegradation of lindane (γ-hexachlorocyclohexane) by the white-rot fungus Trametes hirsutus. Lett Appl Microbiol 28:238–241
Singh BK, Kuhad RC (2000) Degradation of the insecticide lindane (γ-HCH) by white-rot fungi Cyathus bulleri and Phanerochaete sordida. Pest Manage Sci 56:142–146
Sinha S, Chattopadhyay P, Pan I, Chatterjee S, Chanda P, Bandyopadhyay D, Sen SK (2009) Microbial transformation of xenobiotics for environmental bioremediation. Afr J Biotechnol 8:6016–6027
Sinha S, Chattopadhyay P, Pan I, Chatterjee S, Chanda P, Bandyopadhyay D, Sen SK (2011) Microbial transformation of xenobiotics for environmental bioremediation. Afr J Biotechnol 8:22
Sørensen SJ, Bailey M, Hansen LH, Kroer N, Wuertz S (2005) Studying plasmid horizontal transfer in situ: a critical review. Nat Rev Microbiol 3:700–710
Stockholm Convention (2001) Persistent Organic Pollutants and the Stockholm Convention: A Resource Guide Prepared by Resource Futures International for the World Bank and CIDA
Stockholm Convention (2009) Stockholm convention on persistent organic pollutants (POPs). Secretariat of the Stockholm Convention on Persistent Organic Pollutants, United Nations Environment Programme (UNEP), United Nations Treaty, Stockholm
Stockholm Convention (2010) The new POPs under the Stockholm Convention. http://chm.pops.int/Implementation/NewPOPs/TheNewPOPs/tabid/672/Default.aspx. C.N.681.2015.TREATIES-XXVII. Accessed 15 Dec 2015
Suenaga H, Nishi A, Watanabe T, Sakai M, Furukawa K (1999) Engineering a hybrid Pseudomonad to acquire 3,4-dioxygenase activity for polychlorinated biphenyls. J Biosci Bioeng 87:430–435
Suenaga H, Goto M, Furukawa K (2001) Emergence of multifunctional oxygenase activities by random priming recombination. J Biol Chem 276:22500–22506
Suenaga H, Watanabe T, Sato M, Furukawa KN (2002) Alteration of regiospecificity in biphenyl dioxygenase by active-site engineering. J Bacteriol 184:3682–8
Sulistyaningdyah WT, Ogawa J, Li QS, Shinkyo R, Sakaki T, Nouye K, Schmid RD, Shimizu S (2004) Metabolism of polychlorinated dibenzo-p-dioxins by cytochrome P450BM-3 and its mutant. Biotechnol Lett 26:1857–1860
Syed K, Doddapaneni H, Subramanian V, Lam YW, Yadav JS (2010) Genome to function characterization of novel fungal P450 monooxygenases oxidizing polycyclic aromatic hydrocarbons (PAHs). Biochem Biophys Res Commun 399:492–497
Takagi K, Iwasaki A, Kamei I, Satsuma K, Yoshioka Y, Harada N (2009) Aerobic mineralization of hexachlorobenzene by newly isolated pentachloronitrobenzene-degrading Nocardioides sp. strain PD653. Appl Environ Microbiol 75:4452–4458
Tas N, Eekert MHA, Wagner A, Schraa G, Vos W, Smidt MD (2011) Role of “Dehalococcoides” sp. in the anaerobic transformation of hexachlorobenzene in European rivers. Appl Environ Microbiol 77:4437–4445
Thomas CM, Nielsen KM (2005) Mechanisms of, and barriers to, horizontal gene transfer between bacteria. Nat Rev Microbiol 3:711–721
Thomas T, Gilbert J, Meyer F (2012) Metagenomics—a guide from sampling to data analysis. Microb Inform Exp 2:1
Top EM, Springael D (2003) The role of mobile genetic elements in bacterial adaptation to xenobiotic organic compounds. Curr Opin Biotechnol 14:262–269
Torres-Cruz J, van der Woude MW (2003) Slipped-strand mispairing can function as a phase variation mechanism in Escherichia coli. J Bacteriol 185:6990–6994
Tyson GW, Chapman J, Hugenholtz P, Allen EE, Ram RJ, Richardson PM, Solovyev VV, Rubin EM, Rokhsar DS, Banfield JF (2004) Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature 428:37–43
Uchiyama T, Watanabe K (2007) The SIGEX scheme: high throughput screening of environmental metagenomes for the isolation of novel catabolic genes. Biotechnol Genet Eng 24:107–116
Uchiyama T, Abe T, Ikemura T, Watanabe K (2005) Substrate-induced gene-expression screening of environmental metagenome libraries for isolation of catabolic genes. Nat Biotechnol 23:88–93
Uhlik O, Leewis MC, Strejcek M, Musilova L, Mackova M, Leigh MB, Macek T (2013) Stable isotope probing in the metagenomics era: a bridge towards improved bioremediation. Biotechnol Adv 31:154–165
UNEP (2006) Global Programme of Action (GPA) for the Protection of the Marine Environment from Land-based Activities Ecosystem based management. United Nations Environment Programme, The Hague. ISBN 92-807-2707-9
United States Environmental Protection Agency (USEPA) (2009) Persistent Organic Pollutants: A Global Issue, A Global Response. http://www.epa.gov
Vallaeys T, Courde L, Mc Gowan C, Wright AD, Fulthorpe RR (1999) Phylogenetic analyses indicate independent recruitment of diverse gene cassettes during assemblage of the 2, 4-D catabolic pathway. FEMS Microbiol Ecol 28:373–382
van Belkum A, Scherer S, van Alphen L, Verbrugh H (1998) Short sequence DNA repeats in prokaryotic genomes. Microbiol Mol Biol Rev 62:275–293
van der Meer JR, Zehnder AJ, de Vos WM (1991) Identification of a novel composite transposable element, Tn5280, carrying chlorobenzene dioxygenase genes of Pseudomonas sp. strain P51. J Bacteriol 173:7077–7083
Van Beilen JB, Funhoff EG (2007) Alkane hydroxylases involved in microbial alkane degradation. Appl Microbiol Biotechnol 74:13–21
Van Hamme JD, Singh A, Ward OP (2003) Recent advances in petroleum microbiology. Microbiol Mol Biol Rev 67:503–549
Venter JC, Remington K, Heidelberg JF, Halpern AL, Rusch D, Eisen JA, Wu D, Paulsen I, Nelson KE, Nelson W, Fouts DE, Levy S, Knap AH, Lomas MW, Nealson K, White O, Peterson J, Hoffman J, Parsons R, Baden-Tillson H, Pfannkoch C, Rogers YH, Smith HO (2004) Environmental genome shotgun sequencing of the Sargasso Sea. Science 304:66–74
Verma A, Ali D, Farooq M, Pant AB, Ray RS, Hans RK (2011) Expression and inducibility of endosulfan metabolizing gene in Rhodococcus strain isolated from earthworm gut microflora for its application in bioremediation. Bioresour Technol 102:2979–2984
Vitali VMV, Machado KMG, Andrea MMD, Bononi VLR (2006) Screening mitosporic fungi for organochlorides degradation. Brazilian J Microbiol 37:256–261
Wade W (2002) Unculturable bacteria-the uncharacterized organisms that cause oral infections. J R Soc Med 95:81–83
Wagner M, Horn M, Daims H (2003) Fluorescence in situ hybridisation for the identification and characterisation of prokaryotes. Curr Opin Microbiol 6:302–9
Wang GL, Bi M, Liang B, Jiang J, Li S (2011) Pseudoxanthomonas jiangsuensis sp. a DDT degrading bacterium isolated from a long-term DDT-polluted soil. Curr Microbiol 62:1760–1766
Wang Y, Chen Y, Zhou Q, Huang S, Ning K, Xu J, Huang WE (2012) A culture-independent approach to unravel uncultured bacteria and functional genes in a complex microbial community. PLoS One 7:e47530
Weir KM, Sutherland TD, Horne I, Russell RJ, Oakeshott JG (2006) A single monooxygenase, ese, is involved in the metabolism of the organochlorides endosulfan and endosulfate in an Arthrobacter sp. Appl Environ Microbiol 72:3524–30
Wenzel A, Gahr A, Niessner R (1999) TOC-removal and degradation of pollutants in leachate using a thin-film photoreactor. Water Res 33:937–946
Xiao P, Mori T, Kamei I, Kondo R (2011) Metabolism of organochlorine pesticide heptachlor and its metabolite heptachlor epoxide by white rot fungi, belonging to genus Phlebia. FEMS Microbiol Lett 314:140–146
Xu D, Wei Y, Wu J, Dunaway-Mariano D, Guo H, Cui Q, Gao J (2004) QM/MM studies of the enzyme-catalyzed dechlorination of 4-chlorobenzoyl-CoA provide insight into reaction energetics. J Am Chem Soc 126:13649–13658
Yamazoe A, Yagi O, Oyaizu H (2004) Degradation of polycyclic aromatic hydrocarbons by a newly isolated dibenzofuran-utilizing Janibacter sp. strain YY-1. Appl Microbiol Biotechnol 65:211–8
Yan DZ, Liu H, Zhou NY (2006) Conversion of Sphingobium chlorophenolicum ATCC 39723 to a hexachlorobenzene degrader by metabolic engineering. Appl Environ Microbiol 72:2283–2286
Ying GG, Toze S, Hanna J, Yu XY, Dillon PJ, Kookana RS (2008) Decay of endocrine-disrupting chemicals in aerobic and anoxic groundwater. Water Res 42:1133–1141
Zhang C, Bennett GN (2005) Biodegradation of xenobiotics by anaerobic bacteria. Appl Microbiol Biotechnol 67:600–618
Zhang S, Xia X, Xia N, Wu S, Gao F, Zhou W (2013) Identification and biodegradation efficiency of a newly isolated 2, 2′, 4, 4′-tetrabromodiphenyl ether (BDE-47) aerobic degrading bacterial strain. Int Biodeter Biodegr 76:24–31
Zheng YJ, Ornstein RL (1997) Mechanism of nucleophilic aromatic substitution of 1-chloro-2, 4-dinitrobenzene by glutathione in the gas phase and in solution. Implications for the mode of action of glutathione S-transferases. J Am Chem Soc 119:648–655
Acknowledgments
The authors would like to acknowledge the authorities of NIT, Rourkela, for providing facilities. J.C. gratefully acknowledges the receipt of fellowship from Ministry of Human Resource Development, Government of India from NIT, Rourkela, for doctoral research. S.D. thanks the Department of Biotechnology, Ministry of Science and Technology, Government of India, for research grants on microbial bioremediation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Responsible editor: Robert Duran
Rights and permissions
About this article
Cite this article
Chakraborty, J., Das, S. Molecular perspectives and recent advances in microbial remediation of persistent organic pollutants. Environ Sci Pollut Res 23, 16883–16903 (2016). https://doi.org/10.1007/s11356-016-6887-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11356-016-6887-7