Abstract
The SbNHX1 gene encodes a vacuolar Na+/H+ antiporter that involved in the maintenance of ion homeostasis and compartmentalization of excess Na+ or K+ ions into the vacuole. Transgenic castor plants were developed by an improved method of Agrobacterium mediated genetic transformation using spermidine (1 mM) along with acetosyringone (200 μM), which enhanced the transformation efficiency about twofolds from 2.76 to 5.91 %. Transgenic plants were confirmed by PCR using gene (SbNHX1, hptII and gus) specific primers. The single gene integration event was confirmed by RTqPCR and Southern hybridization. Transgenic lines CL7 and CL13 showed high expression of the SbNHX1 gene compared to CL6 and CL12, therefore selected for physio-biochemical analyses, which were carried out under varying NaCl concentrations. Higher chlorophyll, RWC, K+ content, K+/Na+ ratio and lower electrolytic leakage, proline, MDA, Na+ contents compared to WT confirmed that ectopic expression of the SbNHX1 gene enhances salt tolerance of transgenic plants by modulating physiological process under stress condition. Though transgenic lines were affected under stress conditions but performed better compared to WT plants. The present study is the first report of engineering salt tolerance in castor, so far. Transgenic castor may be utilized for the cultivation in marginal salty land and thus open up the possibility of releasing arable land, which is presently under castor cultivation.
Similar content being viewed by others

Avoid common mistakes on your manuscript.
Introduction
Castor (Ricinus communis L.), a member of the Euphorbiaceae family, is a non-edible oilseed crop, widely cultivated in tropical and subtropical regions of the world, such as India, Brazil, China, Thailand, Ethiopia and Philippines. India is the largest producer of castor with an annual production of about 2.6 million tonnes. Castor seeds contain high oil content ranging between 42 and 58 %. Castor oil is unsuitable for edible purpose because of high ratio (84–90 %) of a monounsaturated hydroxy fatty acid ricinoleate (12-hydroxy-octadeca cis-9-enoic acid; 18:1-OH) (Weiss 2000). In recent years, castor oil has been used extensively for medicinal and industrial purposes, such as laxative, paint, varnishes, nylon type synthetic polymer, cosmetics, resins, textile dyeing, hydraulic fluid, caulks and high quality lubricants for high speed aero-engines and jet turbine engines (Weiss 2000). Castor bean is a potential source of biodiesel, however extremely high viscosity and hygroscopicity of ricinoleic acid fascinate moisture and produce high water content that hampers the use of castor oil for biodiesel production (Conceicao et al. 2007). Rojas-Barros et al. (2004) reported a natural castor mutant that can be used for biodiesel production because it contains high oleic acid and low ricinoleic acid. Recent studies and genetic improvements have increased the oil content in castor, but plant growth and production intend to decline because of abiotic and biotic stresses, such as high salinity, drought, pathogens and insect pests (Dange et al. 2005; Severino et al. 2012).
Plant development and productivity are adversely affected due to increase in salinization of lands. It is one of the irresistible forms of stress that limits the distribution of fertile land around the world. Over 6 % of land throughout the world is affected by salt, which is approximately 22 % of the total agricultural land (FAO 2008). The threshold level of salinity for castor growth and survival is 4–7 dS m−1 (40–70 mM), thereafter plants show reduction in growth, photosynthesis and chlorophyll content with increasing salinity (Zhou et al. 2010). Thus, salinity is one of the major challenges for increased production of castor (Severino et al. 2012).
Plants adopt numerous molecular mechanisms to prevent themselves from the negative impact of salt stress. The simplest strategy that plants employ for salt tolerance is sodium exclusion mediated by Na+/H+ antiporters and sodium compartmentalization in large vacuoles for maintaining low level of sodium in the cytosol (Apse et al. 1999). All NHX1 proteins are localized in the vacuolar membrane and involved in ion homeostasis by using vacuolar membrane H+-ATPase and H+-pyrophosphatases driven proton gradient (Apse et al. 1999). Over-expression of Arabidopsis thaliana AtNHX1 conferred enhanced salt tolerance in Arabidopsis (Apse et al. 1999) and several other plant species, such as cotton (He et al. 2005), buckwheat (Chen et al. 2008) and poplar (Jiang et al. 2012). Na+/H+ antiporter has been isolated from different halophytes, such as Mesembryanthemum crystallinum (Chauhan et al. 2000), Atriplex gmelini (Hamada et al. 2001), Thellungiella halophila (Wu et al. 2009), Salicornia brachiata (Jha et al. 2011a), Karelinia caspica (Liu et al. 2012), Leptochloa fusca (Rauf et al. 2014) and showed high expression under varying salt stress concentrations.
An efficient, rapid and reproducible in vitro regeneration and genetic transformation system is a prerequisite for developing a transgenic plant of any species. Previous studies on regeneration of castor have been reported from many meristematic explants (Sujatha and Reddy 1998). Agrobacterium and microprojectile bombardment mediated gene transfer methods are widely used for genetic transformation. Till date, there were few reports of genetic transformation via Agrobacterium (Sujatha and Sailaja 2005) and particle bombardment (Sailaja et al. 2008) in castor.
Salicornia brachiata is an extreme halophyte and extensively explored for abiotic stress responsive genes (Jha et al. 2011b; Chaturvedi et al. 2012; Tiwari et al. 2014; Udawat et al. 2014). The plant requires NaCl for tissue culture (Joshi et al. 2012), contains unique oligosaccharides (Mishra et al. 2013), metabolites (Mishra et al. 2015), sulphur rich seed storage proteins (Jha et al. 2012) and eaten as salad green. Thus S. brachiata is a potential indigenous candidate as the gene resource to be used for developing abiotic stress tolerant transgenic plants (Jha et al. 2011b; Joshi et al. 2013; Chaturvedi et al. 2014; Singh et al. 2014a, b). In this study, Agrobacterium mediated genetic transformation of castor using spermidine along with acetosyringone (for the vir gene induction) was reported and salt tolerant transgenic castor was developed using the SbNHX1 gene, cloned from an extreme halophyte S. brachiata. It was observed that over-expression of the SbNHX1 gene enhances salt stress by modulating physiological processes of the transgenic plant.
Materials and methods
Plant material and explant preparation
Decoated mature castor seeds of a local cultivar (GCH-7) were surface sterilized with 0.1 % (w/v) aqueous mercuric chloride solution for 10 min and subsequently rinsed 6 times with sterile distilled water. Embryo explants were aseptically dissected from endosperm and papery cotyledons were removed carefully. Explants were pre-cultured on solid Murashige and Skoog (MS; Murashige and Skoog 1962) medium (pH 5.8) supplemented with 0.88 µM BA, 0.7 % (w/v) agar and 3 % (w/v) sucrose. Embryos were incubated at 25 °C for 1 week in the dark at controlled laboratory conditions.
Genetic transformation and plant regeneration
Agrobacterium infection and co-cultivation
Genetic transformation was carried out via Agrobacterium tumefaciens (EHA105 strain) harbouring the plasmid construct pCAMBIA1301-SbNHX1, which contains the gusA reporter gene and selectable hptII gene, both controlled by cauliflower mosaic virus (CaMV) 35S promoter (Jha et al. 2011a). Agrobacterium culture was grown in Luria–Bertani (LB) medium containing 25 mg L−1 rifampicin and 50 mg L−1 kanamycin until the absorbance reached up to 0.6 at 600 nm (OD600 = 0.6). Bacterial cells were pelleted by centrifugation at 2700×g for 10 min at 4 °C. The pellet was resuspended in ½ MS media (50 ml) containing 2 % sucrose, 200 µM acetosyringone (AS) and 1 mM spermidine (SPD). The radicle part was removed from the pre-cultured explants, embryo axes were injured by sterile needle in the meristematic region and infected with 50 ml Agrobacterium suspension for 30 min, including a gentle shaking at 10 g to improve the infection. Excess Agrobacterium cells were blotted out and explants were co-cultivated on MS medium in the dark at 25 °C for 3 days.
Selection and regeneration of transformants
Following co-cultivation, infected explants were washed four times with half strength (½) MS medium containing 400 mg L−1 cefotaxime for 5 min each and further rinsed in ½ MS medium for 10 min to remove the antibiotic. Explants were dried on sterile filter paper and transferred to MS media supplemented with 0.88 µM BA + 2.27 µM TDZ and 400 mg L−1 cefotaxime. After 10 days, explants were subjected to three rounds of selection on selection media (SM; MS media supplemented with 2.22 µM BA + 2.32 µM KN + Cefotaxime 400 mg L−1) with 20, 25 or 30 mg L−1 hygromycin, respectively. After selection, putative transformed shoots were transferred to shoot induction medium (SIM; MS medium supplemented with 2.22 µM BA + 1.14 µM TDZ and 400 mg L−1 cefotaxime) for 15 days. Regenerated shoots were transferred to shoot proliferation medium (SPM; MS medium supplemented with 2.22 µM BA + 2.32 µM KN and 400 mg L−1 cefotaxime) for 30 days. Proliferated shoots were transferred to shoot elongation medium (SEM; MS medium supplemented with 1.33 µM BA + 2.32 µM KN and 400 mg L−1 cefotaxime) for further 30 days. Elongated shoots were transferred to root induction medium (RIM; ½ MS salts with B5 vitamins + 1.47 μM IBA + 0.6 mg L−1 AgNO3 + 20 mg L−1 PVP and 400 mg L−1 cefotaxime) for rooting. After 30–40 days, rooted plants were transferred to plastic pots (5 × 10 cm) containing autoclaved soil and vermiculite mixture (1:1). The pots were covered with transparent plastic bags and plantlets were maintained under high humidity for 2 weeks and then gradually exposed to culture room conditions for acclimatization. All cultures were maintained under controlled laboratory conditions; 16/8 h (light/dark) photoperiod at 25 ± 2 °C with cool white fluorescent lamp of 35 μmol m−2 s−1 light intensity.
Histochemical GUS assay
Transient β-glucuronidase gene expression was observed in transformed embryo axes after 24 h of transformation. Regenerated putative transgenic shoots (T0 plants) were assessed for constitutive gus expression after 3, 6 and 9 weeks of selection. Histochemical GUS assay was performed by incubating tissues in GUS assay buffer (1 g L−1 X-Gluc with 0.05 M Na2HPO4, 0.5 mM K3Fe(CN)6, 0.5 mM K4Fe(CN)6, 10 mM EDTA and 0.1 % (v/v) Triton X-100) for 12 h at 37 °C. The tissue was subsequently de-stained in 70 % alcohol and GUS spots were examined under a binocular stereomicroscope (Olympus, Japan).
Molecular analysis
PCR analysis
The genomic DNA of putative transgenic lines of castor was isolated using CTAB method (Doyle and Doyle 1987). Transformation was confirmed by PCR using SbNHX1 (F: 5′-ATG TGG TCA CAG TTG AGC TC-3′ and R: 5′-CTA TGT TCT GTC TAG CAA AT TG-3′), reporter gene gus (F: 5′-GAT CGC GAA AAC TGT GGA AT-3′ and R: 5′-TGA GCG TCG CAG AAC ATT AC-3′) and hygromycin selection marker hptII gene specific primers (F: 5′-TTC TTT GCC CTC GGA CGA GTG-3′ and R: 5′-ACA GCG TCT CCG ACC TGA TG-3′) with an initial denaturation temperature of 94 °C for 5 min, subsequent 35 cycles of 94 °C denaturation for 1 min, 60 °C annealing for 1 min, 72 °C extension for 2 min followed by final extension at 72 °C for 7 min.
Real time quantitative PCR
The real time quantitative PCR (RTqPCR) assay was carried out in the Real-Time iQ5 Cycler (Bio-Rad, USA) to calculate gene copy number. Genomic DNA concentration was determined by NanoDrop Spectrophotometer and diluted to 1, 10 and 100 ng μL−1 concentrations. The RcFAH gene exists as single copy in the castor genome (Chan et al. 2010), therefore selected as internal control and standard in RTqPCR for the determination of numbers of trans-gene. RTqPCR conditions were optimized for gus gene using primers (GusRTF: 5′-CTA TGC CGG AAT CCA TCG-3′, GusRTR: 5′-GCA TCA CGC AGT TCA ACG-3′) and the RcFAH gene (GenBank: XM002528081) using primers (OHXF: 5′-CGT ATG TCG CTT GGC TGG T-3′, OHXR: 5′-TGT TAG AAT GGT GGC GGC-3′). The RTqPCR reaction was performed using 0.25 μM primers (gus or RcFAH) in a 20 μl reaction mixture using QuantiFast SYBR Green PCR reaction kit (Qiagen, USA). The optimized RTqPCR conditions were an initial denaturation at 95 °C for 5 min; subsequent 40 cycles at 94 °C for 30 s, 60 °C for 30 s and 72 °C for 45 s followed by a cycle at 95 and 60 °C for 1 min each. The RTqPCR specificity was checked through a melt curve analysis and 1.5 % agarose gel electrophoresis. The experiment was repeated three times independently with two replicates each time. Standard curve was plotted using threshold cycle (Ct) value of the endogenous gene (RcFAH) and the transgene (gus) to determine RTqPCR efficiencies (Shepherd et al. 2009) as follows:
The efficiency values were put in the following formula (Shepherd et al. 2009) to determine the copy number ratio of the gus gene to RcFAH gene.
Southern hybridization
Gene copy number and integration were further analyzed by Southern blot analysis. For this, genomic DNA (30 μg) was isolated from transformants, digested with HindIII and BamHI, separated by a 0.7 % agarose gel electrophoresis and transferred to Hybond N+ membrane (Amersham Pharmacia, UK) by the capillary method using alkaline transfer buffer (0.4 N NaOH, 1 M NaCl). The purified pCAMBIA1301-SbNHX1 plasmid and DNA from a non-transformed plant were used as a positive and negative control, respectively. The membrane was air-dried and DNA transferred to the membrane was fixed by UV cross-linking using 56 mJ cm−2 energy for 1 min in a UVC 500 cross-linker (Amersham Biosciences, UK). The blot was hybridized with PCR-generated probe for the SbNHX1 gene, labelled with DIG-11-dUTP and amplified from the plasmid construct pCAMBIA1301-SbNHX1 using PCR DIG probe synthesis kit (Roche, Germany). The hybridized membrane was detected by using CDP-Star chemiluminescent as a substrate, following the manufacturer’s instructions (Roche, Germany) and signals were visualized with X-ray film after 20 min.
Subcellular localization of a SbNHX1:RFP translational fusion protein
The full length SbNHX1 cDNA was amplified with gene specific primers (F: 5′-CACC ATG TGG TCA CAG TTG AGC TC-3′ and R: 5′-CTA TGT TCT GTC TAG CAA AT TG-3′) and fused with RFP (red fluorescent protein) using Gateway technology (Walhout et al. 2000). The amplified blunt-end PCR product (SbNHX1) was first cloned into pENTER/D-TOPO entry vector (Invitrogen, USA) followed by further cloning into destination vector pSITE-4CA by the LR recombination reaction using LR Clonase II enzyme mix (Invitrogen, USA). Fusion gene construct (RFP:SbNHX1) was transformed into onion epidermal cells by particle bombardment (PDS-1000/He Biolistic, Biorad, USA). The pSITE-4CA (RFP) vector was used as control. Transformed onion epidermal cells were observed for transient expression of RFP using an epifluorescence microscope (Axio Imager, Carl Zeiss AG, Germany).
Over-expression analysis
The total RNA was isolated from WT and transgenic plant samples (CL6, CL7, CL12 and CL13) and over-expression of the SbNHX1 gene was confirmed by semi-quantitative Reverse transcriptase-PCR (Rt-PCR) analysis. The cDNA was prepared using ImProm-II Reverse Transcription System kit (Promega, USA). Semi-quantitative Rt-PCR analysis was performed using cDNA (1 µl) as a template, SbNHX1 gene as a target and RcFAH gene as an internal control. The Rt-PCR was performed using SbNHX1 (Rt-NHX1F: 5′-ATG GTG TTT GGG TTG CTG A-3′ and Rt-NHX1R; 5′-CTG CTT CGT CTT GGT TGT CC-3′) and RcFAH gene specific primers (OHXF and OHXR) and the amplification products were analysed with 1.2 % agarose gel.
Physio-biochemical analyses
Leaf disc assay and chlorophyll estimation
Leaf senescence assay was carried out to analyze transgenic plants for their salt tolerance (Joshi et al. 2013). Healthy leaves from transgenic lines (T0 generation) and WT of similar age were detached, leaf discs of 5 mm diameter were punched out and floated in 5 ml sterilized distilled water containing different concentrations of sodium chloride (0, 50, 100, 150 or 200 mM) for 8 days. Leaf discs were maintained under 16/8 h light/dark photoperiod with white fluorescent lamp of 35 μmol m−2 s−1 light intensity at 25 ± 2 °C. The effect of salt concentrations on leaf discs was assessed by observing phenotypic changes.
Leaf discs of WT and transgenic lines treated with different concentrations of NaCl were homogenized in 80 % acetone and incubated for 1–2 h with shaking at 15 g in the dark. The homogenate was centrifuged at 1000×g for 5 min. The absorption of the supernatant was recorded (at 645 and 665 nm) and chlorophyll content was calculated per gram fresh weight of the tissue (Arnon 1949) as follows:
Salt stress assay
Transgenic lines (CL7 and CL13) and WT plants were assessed for salt tolerance. For this, plants were grown in plastic pot, irrigated with 200 mM NaCl at 2 days interval and effect of salt was observed after 7 days. Leaves were harvested from treated and untreated plants (WT and transgenic lines, CL7 and CL13) and different physio-biochemical analyses (relative water, proline, malondialdehyde and ion contents, and electrolyte leakage) were performed.
Electrolyte leakage analysis
Leaves were harvested from treated and untreated plants (WT and transgenic lines, CL7 and CL13) and washed thoroughly with deionised water to remove surface-adhered electrolytes. Samples were immersed in close vials containing 10 ml deionized water and incubated at 25 °C on a rotary shaker for 24 h. Subsequently, the initial electrical conductivity (EC) of the solution (Lt) was measured using conductivity meter (SevenEasy, Mettler Toledo AG 8603, Switzerland). Leaf Samples were autoclaved at 120 °C for 20 min, cooled up to 25 °C and final electrical conductivity (L0) of the solution was measured. The electrolyte leakage was determined as follows:
Relative water content analysis
Similar size of leaves was used for the determination of relative water content (RWC). Leaves were harvested from treated and untreated plants (WT and transgenic lines, CL7 and CL13) and their fresh weight (FW) were recorded immediately. Leaves were submerged into the deionised water for overnight and turgid weight (TW) was recorded. Finally, leaves were kept at 80 °C for 48 h and dry weight (DW) was recorded. RWC was estimated in percentage as follows (Barrs and Weatherley 1962):
Proline content analysis
Free proline contents of treated and untreated leaves (WT and transgenic lines, CL7 and CL13) were extracted in 3 % sulphosalicylic acid by centrifugation at 5000×g for 30 min. The extract was treated with acetic acid and ninhydrin followed by boiling for 1 h. Proline content was calculated by a standard curve, prepared against known concentration of proline measured at 520 nm absorbance (Bates et al. 1973).
Malondialdehyde content analysis
Lipid peroxidation was estimated by quantifying MDA contents. Leaf samples (2 g) from WT and transgenic lines (CL7 and CL13) were homogenized in 2 ml trichloroacetic acid (0.1 %, TCA), centrifuged at 10,000×g for 10 min and supernatant were collected. Two set of solution were prepared. Solution 1 (+TBA) contained Thiobarbituric acid (0.65 % w/v) prepared in TCA (20 % w/v), whereas solution 2 (−TBA) was 20 % w/v TCA (TBA was excluded). A measure of 1 ml solution (1 or 2) was added to 1.0 ml extract. The mixtures were incubated at 95 °C for 30 min and samples were cooled on ice, centrifuged at 10,000×g for 5 min. The absorbance of supernatants was measured at 440, 532 and 600 nm. Lipid peroxidation was estimated by determining the concentration of malondialdehyde (MDA) produced by TBA reaction using following formula (Hodges et al. 1999):
Ion content analysis
For ion content analysis, treated and untreated leaves tissues (WT and transgenic lines, CL7 and CL13) were dried at 70 °C for 48 h in hot air oven and dry weight was measured. Dried tissues were digested with 4 ml perchloric acid and nitric acid solution (3:1) for 8–10 h. Digested samples were dried on the hot plate, diluted to 25 ml with deionised water and filtered through 0.2-mm filter. Ion contents were measured by inductively coupled plasma optical emission spectrometer (Optima 2000DV, Perkin Elmer, Germany).
Statistical analysis
All data were expressed as mean ± SE and subjected to analysis of variance (ANOVA) to determine the significance of difference between the means of WT and transgenic plants of each treatment group. A Tukey HSD multiple comparison of mean test was used, significant differences were considered at P < 0.01 and indicated by similar letters.
Results
Genetic transformation and regeneration of transgenic plants
Transgenic castor plants were developed by Agrobacterium mediated genetic transformation using embryo axes as explants. The effect of AS alone or with SPD on the transformation was assessed by means of transient gus expression efficiency. Out of 20 explants per treatment, number of explants showing transient gus expression (high, medium or low) was calculated (Fig. 1 and Table S1). Among different AS concentrations, 15 explants out of 20, showed gus expression, in which 9, 2 and 4 explants showed high, medium and low gus intensity, respectively. The maximum gus expression efficiency (75 %) was observed with 200 μM AS after 72 h. Therefore, 200 μM AS was selected for experiments with different concentrations of SPD. About 100 % gus expression was observed with a combination of 200 μM AS and 1 mM SPD, co-cultivated for 72 h, in which 15, 4 and 1 explants showed high, medium and low gus intensity, respectively (Figure S1 and Table S1). A total of 1016 embryo axis explants were used for the genetic transformation and the transformation efficiency was calculated as the number of GUS and PCR positive plants, survived after third round of selection with respect to total explants transformed (Table 1). Average transformation efficiency of 2.76 % was calculated, when transformation was done with 200 μM AS alone, whereas 5.91 % efficiency was achieved with the combination of AS and SPD (200 μM AS + 1 mM SPD) (Table 1).
Agrobacterium mediated transformation of castor using embryo axis as explant: a embryo axis explant used for transformation; transient gus expression of explants with b high, c medium and d low intensity. Selection of transformants after e 1st round (30 days old), f 2nd round (50 days old) and g 3rd round (75 days old) on hygromycin. Regeneration of transformants; h shoot induction (90 days old), i multiple shoot proliferation (120 days old), j shoot elongation (180 days old), k rooting (220 days old) and l hardening (270 days old) of transgenic plant in plastic pot. m GUS assay of whole leaf of transgenic plant
After co-cultivation, transformants were regenerated to transgenic plants after hygromycin selection (Fig. 1). Regenerated leaves from transgenic lines were assayed for the constitutive expression of gus gene and 50 % of the total regenerated transgenic plants were found GUS positive (Fig. 1). Putative transgenic plants were transferred to plastic pots for hardening and total 4 transgenic lines (CL6, Cl7, CL12 and CL13) were obtained after hardening.
Molecular analysis of transgenic plants
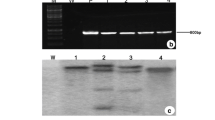
The gene integration in regenerated transgenic plants was confirmed by the presence of SbNHX1, selectable marker hptII and reporter gus genes (Fig. 2a) using PCR amplification of expected bands of sizes 1.68, 0.96 and 1.28 kb, respectively (Fig. 2b–d). Copy number of the integrated gene was determined by RTqPCR using RcFAH gene as an internal control (Figure S2) and single copy of gene insertion was observed in all transgenic lines (CL6, CL7, CL12 and CL13 obtained after hardening). The melt curve analyses of genes RcFAH and gus showed single peaks that confirmed the absence of any dimer or other contamination (Figure S2). Single band amplification of the gus gene as well as control gene was confirmed with 1.2 % agarose gel electrophoresis. Copy number of the gene was further confirmed by Southern blot analysis and single gene copy insertion was detected in all transgenic lines CL6, CL7, CL12 and CL13 (Fig. 2e, f). The over-expression of the SbNHX1 gene was confirmed by reverse transcriptase PCR (Rt-PCR). Transgenic lines (CL7 and CL13) showed high expression of the gene, whereas low expression was observed in CL6 and CL12 transgenic lines (Fig. 2g).
Molecular analysis of transgenic plants. a Schematic map of plant expression gene construct pCAMBIA1301-SbNHX1. PCR confirmation of the b SbNHX1 gene (1.68 kb), c gus gene (1.28 kb) and d hptII gene (0.96 kb). Lane M 1 kb molecular weight marker ladder; lane PC positive control; lane WT wild type plant and lane CL transgenic lines. Southern hybridization of putative transgenic plants using genomic DNA digested with e HindIII and f BamHI. All the four selected lines showed single copy insertion of the gene. g Over-expression of the SbNHX1 gene in transgenic lines compared to wild-type control plants analyzed by semi-quantitative Rt-PCR, where the RcFAH gene was used as internal gene control
Subcellular localization of SbNHX1:RFP fusion protein
The subcellular localization study was performed with pSITE-4CA construct (control) expressing RFP alone and the RFP:SbNHX1 fusion construct (Fig. 3a) by transient expression assays using onion epidermal cells (Fig. 3b). In vivo subcellular localization analysis revealed that the SbNHX1 protein is a vacuolar NHX (Fig. 3b). Red fluorescence signals were observed uniformly distributed in the entire cell region of the onion cells transformed with RFP alone, whereas vacuolar membrane localized signals were detected in cells which are transformed with RFP:SbNHX1 fusion construct (Fig. 3b).
Physio-biochemical analyses of transgenic plants
Based on the SbNXH1 gene expression level (Fig. 2g), two independent transgenic lines; CL7, and CL13 were selected for the further physio-biochemical analyses.
Leaf disc assay and chlorophyll content
Leaf disc assay showed salt tolerance of WT and T0 transgenic lines (CL7 and CL13) up to 50 mM NaCl. At high salt concentration, transgenic lines were also affected but showed improved growth and enhanced salt tolerance compared to WT (Fig. 4a). Similar effect of salt concentration was also observed on chlorophyll content (Fig. 4b). The chlorophyll content of WT and transgenic lines were not affected up to 50 mM NaCl concentration, thereafter reduced significantly in WT with increasing salt concentrations. The chlorophyll content did not reduce drastically in transgenic lines compared to WT. Furthermore, transgenic line CL13 showed enhanced salt tolerance compared to transgenic line CL7.
Leaf disc assay and chlorophyll content analysis. Leaf disc assay of transgenic castor for assessing salt tolerance (A). Leaf discs of WT and transgenic lines CL7 and CL13 were floated in a–c 0 mM, d–f 50 mM, g–i 100 mM, j–l 150 mM and m–o 200 mM NaCl solution for 8 days. Chlorophyll content analysis (B). Chlorophyll content of WT and transgenic lines CL7 and CL13 were estimated at 0, 50, 100, 150 and 200 mM NaCl. Graph represents mean and SE of chlorophyll content in the leaves
Salt stress assay
Transgenic lines (CL7 and CL13) and WT plants were assessed for salt tolerance in soil condition for which, both transgenic and WT plants were irrigated with 200 mM NaCl solutions (at 2 days interval). Transgenic lines (CL7 and CL13) showed enhanced tolerance at 200 mM NaCl compared to WT plants (Fig. 5). The result shows a positive effect of the Na+/H+ antiporter on salt endurance.
Electrolyte leakage and relative water content
The electrolyte leakage and relative water content (RWC) were found almost similar in transgenic and WT plants under unstressed condition (0 mM NaCl), however significant (P > 0.01) reduction in electrolyte leakage was observed in transgenic lines (CL7 and CL13) during salt stress (200 mM NaCl) compared to WT plant (Fig. 6a). Similarly, transgenic lines showed higher RWC compared to WT plants under salt stress condition (Fig. 6b).
Physiological analyses of transgenic castor lines under different stress condition. Comparison of a electrolyte leakage, b relative water content, c proline and d MDA contents of wild type (WT) and transgenic lines (CL7 and CL13) treated with salt (200 mM NaCl). Graph represents the mean ± SE followed by similar letters are significantly different according to Tukey HSD at P < 0.01
Proline and MDA content
Both transgenic lines (CL7 and CL13) and WT plants exhibited approximately similar proline and MDA contents at unstressed conditions (0 mM NaCl). Transgenic lines showed lower accumulation of proline content under stress condition compared to WT plants (Fig. 6c). Lower content of MDA is an indicative of reduced oxidative damage, which was observed in transgenic lines compared to WT plants (Fig. 6d).
Ion content analysis
Transgenic (CL7 and CL13) and WT plants exhibited almost similar Na+ and K+ contents as well as K+/Na+ ratio under controlled conditions (i.e. unstressed; 0 mM NaCl). There was significant increase of Na+ in WT plants compared to transgenic plants under stress treatment. Though high Na+ content was also detected in transgenic plants (CL7 and CL13) under stress condition compared to their respective control plants (transgenic plant under unstressed condition; 0 mM NaCl), but the increase was insignificant compared to increase of Na+ accumulation in WT (Fig. 7a). The result indicates that transgenic lines accumulated less Na+ compared to WT. A significant accumulation of K+ was observed in transgenic lines and WT plants under stress conditions, however among plants, CL13 showed maximum K+ content followed by CL7 and WT plants (Fig. 7b). In the case of K+/Na+ ratio, all transgenic lines and WT showed high ratio at unstressed condition (0 mM NaCl). However, at 200 mM NaCl, ratio decreased in both transgenic and WT plants but transgenic lines exhibited higher ratio than WT (Fig. 7c).
Discussion
Antiporters are attributed to ion homeostasis and the strategy to accumulate Na+ or K+ inside vacuoles is used by many plants, especially halophytes like Salicornia brachiata to survive under salt stress condition. The SbNHX1 gene was found actively involved in the regulation of ion homeostasis (Jha et al. 2011b; Joshi et al. 2013). In this study, salt tolerant transgenic castor was developed using the SbNHX1 gene cloned from extreme halophyte S. brachiata. Physiological analyses of transgenic plants were used to understand the efficacy of an integrated gene and its involvement to modulate the physiological traits.
Transgenic castor plants, over-expressing the SbNHX1 gene were developed via A. tumefaciens mediated genetic transformation using embryo axes as explants (Fig. 1). In this study, juvenile embryo axis was chosen as explant because it is acquiescent to higher transformation efficiency (Sujatha and Reddy 1998; Sujatha and Sailaja 2005; Sailaja et al. 2008). Optimization of an efficient regeneration and genetic transformation protocol is a key step before proceeding to further biotechnological programmes of any crop (Singh et al. 2010; Joshi et al. 2011; Pandey et al. 2013; Tiwari et al. 2015). Different parameters have been used earlier for castor transformation (Sujatha and Sailaja 2005). The condition optimized for transformation in this study was Agrobacterium cell culture of 0.6 OD600, 200 µM AS, 1 mM SPD, a co-cultivation time of 72 h and wounding using a needle (Figure S1). SPD is a polyamine and reported to enhance the Agrobacterium tumefaciens transformation frequency by inducing the vir genes (Kumar and Rajam 2005). SPD is commonly used in microprojectile bombardment mediated genetic transformation. Recently, 200 μM AS was used along with 1 mM SPD during Agrobacterium infection to enhance the transformation efficiency in banana (Chong-Pérez et al. 2012).
There were some reports on the improvement of disease tolerance in castor (Dange et al. 2005), but limited information is available for the improvement of abiotic stress tolerance. Salt stress is one of the major factor among different abiotic stresses, which negatively affects the growth of castor plant (Zhou et al. 2010). Salt stress tolerance of a plant is a multigenic trait, which is controlled by the coordinated action of several genes. But it is also reported that over-expression of a single gene can confer salt tolerance to the plant (Apse et al. 1999; Jha et al. 2011b; Chaturvedi et al. 2014; Singh et al. 2014a, b). It may take into account that the protein produced in transgenic plants by over-expression of only one gene probably interacts with other proteins of plant, contributing for tolerance to salt stress, which is a very complex process. In the present study, we have successfully developed salt tolerant transgenic castor over-expressing the salt-responsive SbNHX1 gene via optimized Agrobacterium mediated transformation method. This protocol yielded transgenic plants with 5.91 % transformation efficiency, which is higher than previous reports (Sujatha and Sailaja 2005; Sailaja et al. 2008) for castor. RTqPCR is recently being used to study functional gene abundance (Yousuf et al. 2012; Keshri et al. 2013; Yousuf et al. 2014a, b; Keshri et al. 2015) and as an alternative to Southern hybridization to determine copy numbers of integrated genes (Joshi et al. 2013). Four transgenic lines CL6, CL7, CL12 and CL13 showed single copy gene insertion analyzed by RTqPCR, which was further confirmed by Southern analysis (Fig. 2). The SbNHX1 was classified as a vacuolar NHX gene based protein topology and the presence of functional domains (Jha et al. 2011a). However, a direct evidence of the vacuolar localization was shown in the present study (Fig. 3). It was observed that AtNHX1 and AtNHX2 were localize to the vacuole (Bassil et al. 2011). Recently, transient expression of AtNHX3 and AtNHX4 displayed expression at the vacuolar membrane (McCubbin et al. 2014).
Chlorophyll content, electrolyte leakage, RWC, proline and MDA contents are considered notable indicators for the physiological status of plants. Physiological response of transgenic lines CL7 and CL13 was assessed under salt stress in a dose-dependent manner and compared with WT plants (Figs. 4, 5, 6). Higher chlorophyll content, potassium ion content, K+/Na+ and water retention capacity (RWC) of transgenic plants exhibited better physiology of plants under stressed conditions. Moreover, lower accumulation of proline and MDA contents revealed that transgenic castor lines have higher osmoprotectants and low lipid peroxidation, respectively compared to WT plants under stress condition. These results evident the better physiological status of transgenic plants compared to WT under stress. Additionally, significant increase of Na+ and reduction of K+ in WT plants were observed compared to transgenic plants, over-expressing the SbNHX1 gene under stress condition (Fig. 7). Accumulation of less Na+ but more K+ in transgenic plants than WT plants under salt stress suggested that salinity resistance could be due to the optimised K+ compartmentalization. It was observed that AtNHX1 protein can transport Na+ and K+ with equal affinity and it transported K+ for osmoregulation under saline stress (Venema et al. 2002). It was also found that AtNHX1 and AtNHX2 were actively involved in K+ homeostasis than Na+ sequestration (Bassil et al. 2011). Similarly, AtNHX1 gene over-expression provided NaCl tolerance to tomato by accumulating K+ in vacuoles (Barragán et al. 2012). Previously, Jatropha and poplar over-expressing SbNHX1 and AtNHX1, respectively showed similar results, moreover WT Jatropha also showed bleaching in leaves under stress treatments compared to transgenic plants (Jiang et al. 2012; Joshi et al. 2013). Similar results have been reported in buckwheat (Chen et al. 2008) and cotton (He et al. 2005) over-expressing the AtNHX1 gene during salt stress. It was also observed that antiporters AtNHX1 and AtNHX2 together account for the Na+, K+/H+ antiport activity, involved in K+ homeostasis and Arabidopsis NHX double mutant shows increased Na+ content compared to WT under 100 mM NaCl stress (Barragán et al. 2012).
Furthermore, transgenic lines (CL7 and CL13) and WT plants were also assessed for salt tolerance in soil condition for which, both transgenic and WT plants were irrigated with 200 mM NaCl solutions (at 2 days interval). The result further confirmed that transgenic plants, over-expressing the SbNHX1 gene had better growth compared to WT plant and showed salt endurance (Fig. 5). The PgNHX1 gene from Pennisetum glaucum has been reported to confer salinity tolerance when over-expressed in Brassica juncea and transgenic plants could tolerate higher salt stress than the WT in a leaf disc assay and in pot (Rajagopal et al. 2007). Salt tolerance of a halophyte Karelinia caspica was shown to decrease by RNAi silencing of KcNHX1 (Liu et al. 2012). Recently, it was observed that over-expression of TaNHX3 enhances salt stress tolerance in tobacco by improving physiological processes (Lu et al. 2014).
Conclusion
In conclusion, introgression of the SbNHX1 gene modulates physiology of transgenic plants and thus imparts enhanced stress endurance. Transgenic castor plants were developed using optimized and improved protocol with 5.91 % transformation efficiency. Transgenic lines CL7 and CL13 contained higher chlorophyll, RWC, K+ contents and K+/Na+ ratio, furthermore lower electrolytic leakage, proline, MDA and Na+ contents were observed in transgenic lines compared to WT under stress condition. In the present study, transgenic lines showed improved growth and enhanced salt tolerance compared to control plants at 200 mM NaCl stress. The transgenic castor with improved salt tolerance may be utilized for the cultivation in salt-affected marginal areas for sustainable land management, oil production and other industrial applications.
References
Apse MP, Aharon GS, Snedden WA, Blumwald E (1999) Salt tolerance conferred by over expression of a vacuolar Na+/H+ antiport in Arabidopsis. Science 285:1256–1258
Arnon DI (1949) Copper enzymes in isolated chloroplasts polyphenol oxidase in Beta vulgaris. Plant Physiol 24:1–15
Barragán V, Leidi EO, Andrés Z, Rubio L, De Luca A, Fernández JA, Cubero B, Pardo JM (2012) Ion exchangers NHX1 and NHX2 mediate active potassium uptake into vacuoles to regulate cell turgor and stomatal function in Arabidopsis. Plant Cell 24:1127–1142
Barrs HD, Weatherley PE (1962) A re-examination of the relative turgidity technique for estimating water deficits in leaves. Aust J Biol Sci 15:413–428
Bassil E, Tajima H, Liang YC, Ohto MA, Ushijima K, Nakano R, Esumi T, Coku A, Belmonte M, Blumwald E (2011) The Arabidopsis Na+/H+ antiporters NHX1 and NHX2 control vacuolar pH and K+ homeostasis to regulate growth, flower development, and reproduction. Plant Cell 23:3482–3497
Bates LS, Waldern R, Teare ID (1973) Rapid determination of free proline for water stress studies. Plant Soil 39:205–207
Chan AP, Crabtree J, Zhao Q, Lorenzi H, Orvis J, Puiu D (2010) Draft genome sequence of the oilseed species Ricinus communis. Nat Biotechnol 28:951–956
Chaturvedi AK, Mishra A, Tiwari V, Jha B (2012) Cloning and transcript analysis of type 2 metallothionein gene (SbMT-2) from extreme halophyte Salicornia brachiata and its heterologous expression in E. coli. Gene 499:280–287
Chaturvedi AK, Patel MK, Mishra A, Tiwari V, Jha B (2014) The SbMT-2 gene from a halophyte confers abiotic stress tolerance and modulates ROS scavenging in transgenic tobacco. PLoS ONE 9(10):e111379
Chauhan S, Forsthoefel N, Ran Y, Quigley F, Nelson DE, Bohnert HJ (2000) Na+/myo-inositol symporters and Na+/H+ antiport in Mesembryanthemum crystallinum. Plant J 24:511–522
Chen LH, Zhang B, Xu ZQ (2008) Salt tolerance conferred by overexpression of Arabidopsis vacuolar Na+/H+ antiporter gene AtNHX1 in common buckwheat (Fagopyrum sculentum). Transgenic Res 17:121–132
Chong-Pérez B, Reyes M, Rojas L, Ocaña B, Pérez B, Kosky RG, Angenon G (2012) Establishment of embryogenic cell suspension cultures and Agrobacterium-mediated transformation in banana cv. ‘Dwarf Cavendish’ (Musa AAA): effect of spermidine on transformation efficiency. Plant Cell, Tissue Organ Cult 111:79–90
Conceicao MM, Fernandez VJ, Bezerra AF, Silva MCD, Santos IMG, Silva FC (2007) Dynamic kinetic calculations of castor oil biodiesel. J Therm Anal Calorim 87:865–869
Dange SRS, Desal AG, Patel SI (2005) Diseases of castor. In: Saharan GS, Mehta N, Sangwan MS (eds) Diseases of oilseed crops. Indus Publishing Company, New Delhi, pp 211–234
Doyle JJ, Doyle JL (1987) A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19:11–15
FAO (2008) FAO land and plant nutrition management service. http://www.fao.org/ag/agl/agll/spush
Hamada A, Shona M, Xia T, Ohta M, Hayashi Y, Tanaka A, Hayakawa T (2001) Isolation and characterization of a Na+/H+ antiporter gene from the halophyte Atriplex gmelini. Plant Mol Biol 46:35–42
He C, Yan J, Shen G, Fu L, Holaday AS, Auld D, Blumwald E, Zhang H (2005) Expression of an Arabidopsis vacuolar sodium/proton antiporter gene in cotton improves photosynthetic performance under salt conditions and increases fiber yield in the field. Plant Cell Physiol 46:1848–1854
Hodges DM, DeLong JM, Forney CF, Prange RK (1999) Improving the thiobarbituric acid-reactive-substances assay for estimating lipid peroxidation in plant tissues containing anthocyanin and other interfering compounds. Planta 207:604–611
Jha A, Joshi M, Yadav NS, Agarwal PK, Jha B (2011a) Cloning and characterization of the Salicornia brachiata Na+/H+ antiporters gene SbNHX1 and its expression by abiotic stress. Mol Biol Rep 38:1965–1973
Jha B, Sharma A, Mishra A (2011b) Expression of SbGSTU (tau class glutathione S-transferase) gene isolated from Salicornia brachiata in tobacco for salt tolerance. Mol Biol Rep 2011(38):4823–4832
Jha B, Singh NP, Mishra A (2012) Proteome profiling of seed storage proteins reveals the nutritional potential of Salicornia brachiata Roxb., an extreme halophyte. J Agric Food Chem 60:4320–4326
Jiang C, Zheng Q, Liu Z, Xu W, Liu L, Zhao G, Long X (2012) Overexpression of Arabidopsis thaliana Na+/H+ antiporter gene enhanced salt resistance in transgenic poplar (Populus euramericana ‘Neva’). Trees 26:685–694
Joshi M, Mishra A, Jha B (2011) Efficient genetic transformation of Jatropha curcas L. by microprojectile bombardment using embryo axes. Ind Crops Prod 33:67–77
Joshi M, Mishra A, Jha B (2012) NaCl plays a key role for in vitro micropropagation of Salicornia brachiata, an extreme halophyte. Ind Crops Prod 35:313–316
Joshi M, Jha A, Mishra A, Jha B (2013) Developing transgenic jatropha using the SbNHX1 gene from an extreme halophyte for cultivation in saline wasteland. PLoS ONE 8(8):e71136
Keshri J, Mishra A, Jha B (2013) Microbial population index and community structure in saline-alkaline soil using gene targeted metagenomics. Microbiol Res 168:165–173
Keshri J, Yousuf B, Mishra A, Jha B (2015) The abundance of functional genes, cbbL, nifH, amoA and apsA and bacterial community structure of intertidal soil from Arabian Sea. Microbiol Res Online. doi:10.1016/j.micres.2015.02.007
Kumar SV, Rajam MV (2005) Polyamines enhance Agrobacterium tumefaciens vir gene induction and T-DNA transfer. Plant Sci 168:475–480
Liu L, Zeng Y, Pan X, Zhang F (2012) Isolation, molecular characterization, and functional analysis of the vacuolar Na+/H+ antiporter genes from the halophyte Karelinia caspica. Mol Biol Rep 9:7193–7202
Lu W, Guo C, Li X, Duan W, Ma C, Zhao M et al (2014) Overexpression of TaNHX3, a vacuolar Na+/H+ antiporter gene in wheat, enhances salt stress tolerance in tobacco by improving related physiological processes. Plant Physiol Biochem 76:17–28
McCubbin T, Bassil E, Zhang S, Blumwald E (2014) Vacuolar Na+/H+ NHX-type antiporters are required for cellular K+ homeostasis, microtubule organization and directional root growth. Plants 3(3):409–426
Mishra A, Joshi M, Jha B (2013) Oligosaccharide mass profiling of nutritionally important Salicornia brachiata, an extreme halophyte. Carbohydr Polym 92:1942–1945
Mishra A, Patel MK, Jha B (2015) Non-targeted metabolomics and scavenging activity of reactive oxygen species reveal the potential of Salicornia brachiata as a functional food. J Funct Foods 13:21–31
Murashige T, Skoog F (1962) A revised medium for rapid growth and bio-assays with tobacco tissue cultures. Physiol Plant 15:473–497
Pandey S, Mishra A, Patel MK, Jha B (2013) An efficient method for Agrobacterium-mediated genetic transformation and plant regeneration in cumin (Cuminum cyminum L.). Appl Biochem Biotechnol 171:385–392
Rajagopal D, Agarwal P, Tyagi W, Singla-Pareek SL, Reddy MK, Sopory SK (2007) Pennisetum glaucum Na+/H+ antiporters confers high level of salinity tolerance in transgenic Brassica juncea. Mol Breed 19:1–9
Rauf M, Shahzad K, Ali R, Ahmad M, Habib I, Mansoor S et al (2014) Cloning and characterization of Na+/H+antiporter (LfNHX1) gene from a halophyte grass Leptochloa fusca for drought and salt tolerance. Mol Biol Rep 41:1669–1682
Rojas-Barros P, De Haro A, Muñoz J, Fernández-Martínez JM (2004) Isolation of a natural mutant in castor bean (Ricinus communis L.) with high oleic/low ricinoleic acid content. Crop Sci 44:76–80
Sailaja M, Tarakeswari M, Sujatha M (2008) Stable genetic transformation of castor (Ricinus communis L.) via particle gun-mediated gene transfer using embryo axes from mature seeds. Plant Cell Rep 27:1509–1519
Severino LS, Cordoba-Gaona OJ, Zanotto MD, Auld DL (2012) The influence of the caruncle on the germination of castor seed under high salinity or low soil water content. Seed Sci Technol 40:139–143
Shepherd CT, Lauter ANM, Scott MP (2009) Determination of transgene copy number by real-time quantitative PCR. In: Scott MP (ed) Methods in molecular biology: transgenic maize. Humana, New York, pp 129–134
Singh N, Mishra A, Joshi M, Jha B (2010) Microprojectile bombardment mediated genetic transformation of embryo axes and plant regeneration in cumin (Cuminum cyminum L.). Plant Cell Tissue Org Cult 103:1–6
Singh N, Mishra A, Jha B (2014a) Over-expression of the peroxisomal ascorbate peroxidase (SbpAPX) gene cloned from halophyte Salicornia brachiata confers salt and drought stress tolerance in transgenic tobacco. Mar Biotechnol 16:321–332
Singh N, Mishra A, Jha B (2014b) Ectopic over-expression of peroxisomal ascorbate peroxidase (SbpAPX) gene confers salt stress tolerance in transgenic peanut (Arachis hypogaea). Gene 547:119–125
Sujatha M, Reddy TP (1998) Differential cytokinin effects on the stimulation of in vitro shoot proliferation from meristematic explants of castor (Ricinus communis L.). Plant Cell Rep 17:561–566
Sujatha M, Sailaja M (2005) Stable genetic transformation of castor (Ricinus communis L.) via Agrobacterium tumefaciens-mediated gene transfer using embryo axes from mature seeds. Plant Cell Rep 23:803–810
Tiwari V, Chaturvedi AK, Mishra A, Jha B (2014) The transcriptional regulatory mechanism of the peroxisomal ascorbate peroxidase (pAPX) gene cloned from an extreme halophyte, Salicornia brachiata. Plant Cell Physiol 55:201–217
Tiwari V, Chaturvedi AK, Mishra A, Jha B (2015) An efficient method of Agrobacterium-mediated genetic transformation and regeneration in local Indian cultivar of groundnut (Arachis hypogaea) using grafting. Appl Biochem Biotechnol 175:436–453
Udawat P, Mishra A, Jha B (2014) Heterologous expression of an uncharacterized universal stress protein gene (SbUSP) from the extreme halophyte, Salicornia brachiata, which confers salt and osmotic tolerance to E. coli. Gene 536:163–170
Venema K, Quintero FJ, Pardo JM, Donaire JP (2002) The Arabidopsis Na+/H+ exchanger AtNHX1 catalyzes low affinity Na+ and K+ transport in reconstituted liposomes. J Biol Chem 277:2413–2418
Walhout A, Temple G, Brasch M, Hartley J, Lorson MA, van den Heuvel S, Vidal M (2000) GATEWAY recombinational cloning: application to the cloning of large numbers of open reading frames or ORFeomes. Methods Enzymol 328:575–592
Weiss A (2000) Oilseed crops. Blackwell Science, Oxford
Wu CX, Gao XH, Kong XQ, Zhao YX, Zhang H (2009) Molecular cloning and functional analysis of a Na+/H+ antiporter gene ThNHX1 from a halophytic plant Thellungiella halophila. Plant Mol Biol Rep 27:1–12
Yousuf B, Keshri J, Mishra A, Jha B (2012) Application of targeted metagenomics to explore abundance and diversity of CO2-fixing bacterial community using cbbL gene from the rhizosphere of Arachis hypogaea. Gene 506:18–24
Yousuf B, Kumar R, Mishra A, Jha B (2014a) Differential distribution and abundance of diazotrophic bacterial communities across different soil niches using gene targeted clone library approach. FEMS Microbiol Lett 360:117–125
Yousuf B, Kumar R, Mishra A, Jha B (2014b) Unravelling the carbon and sulphur metabolism in coastal soil ecosystems using comparative cultivation-independent genome-level characterisation of microbial communities. PLoS ONE 9(9):e107025
Zhou G, Ma BL, Li J, Feng C, Lu J, Qin P (2010) Determining salinity threshold level for castor bean emergence and stand establishment. Crop Sci 50:2030–2036
Acknowledgments
CSIR-CSMCRI Communication No. PRIS-035/2013. The financial support received from Council of Scientific and Industrial Research (CSIR), New Delhi [SIMPLE; BSC0109] is thankfully acknowledged. Dr. Santlal Jaiswar is duly acknowledged for providing technical support in the microscopic analysis of localization study. Authors are grateful to Arabidopsis Biological Resource Centre, Ohio State University, USA for providing the pSITE-4CA vector.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Patel, M.K., Joshi, M., Mishra, A. et al. Ectopic expression of SbNHX1 gene in transgenic castor (Ricinus communis L.) enhances salt stress by modulating physiological process. Plant Cell Tiss Organ Cult 122, 477–490 (2015). https://doi.org/10.1007/s11240-015-0785-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11240-015-0785-4