Abstract
Glucokinase is an enzyme which is responsible for the conversion of glucose to glucose-6-phosphate through ATP-dependent phosphorylation and has a significant role in glycogen synthesis and hepatic glucose production. Allosteric activators of glucokinase could be an attractive approach for the treatment of T2DM (type 2 diabetes mellitus). Recently, an innovative standard “Index of Ideality of Correlation” has been introduced for the estimation of QSAR (quantitative structural activity relationship) model’s potential. In the present work, QSAR models for activators of glucokinase have been developed with target function TF1 and TF2 using index of ideality of correlation (IIC). Along with this, prediction of calibration sets for different QSAR models generated for different splits is also categorized as correct and wrong. Moreover, dispersion in the different runs of same split is also explained. The values of criteria R2 and IIC for best split prepared with target function TF1 are 0.6554 and 0.7912 and that for TF2 are 0.9531 and 0.9758, respectively. The models developed with index of ideality of correlation are better than the models generated without index of ideality of correlation. The IIC could be a better criteria option for predictability of QSAR model for glucokinase activators.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Increased hepatic glucose production and dysfunction of the pancreatic β-cells are mainly responsible for the whole-body insulin resistance and hyperglycemia, which are related to type 2 diabetes mellitus [1]. It is a chronic metabolic disease influencing about 150 million people throughout the world [2]. In the developing world, it is assumed as one of the primary causes of death, and from recent data of IDF Diabetes Atlas, it is specified as chief obstacle in the universal development [3]. At present, there is not a single oral antidiabetic drug available through which we can achieve permanent glycemic control. In reality, the utilization of combination therapy is assumed as better option than monotherapy, although combination therapy also has several unwanted side effects. Thus, to overcome the crisis related with T2D therapies, the demand of more effective and safe novel antidiabetic drugs is also rising [4].
Glucokinase (GK) can be suggested as a better target option for the treatment of T2D because of having activity in multiple organs which helps in control of whole-body glucose level [5]. It is related to the hexokinase family also known as hexokinase D/hexokinase IV [6]. It is involved in the first step of glycolysis and is accountable for the ATP-dependent phosphorylation of glucose. GK is present in pancreatic β-cells and acts like as detector for secretion of insulin. It maintains the glucose homeostasis due to unique kinetic features [7].
Quantitative structure activity relationships (QSARs)/quantitative structure property relationships (QSPRs) play an important function in screening and development of the novel biomolecules with effectiveness [8]. In QSAR/QSPR, mathematical models are developed through which we can relate the physiochemical or biological property of compounds with their chemical structures [9]. After the development of a QSAR model, actual predictive potential of the QSAR model is corroborated with distinct decisive factors. Development of these criterions is not an easy task. Some matrices have been explained in the literature to explain the predictability. Recently, a new criteria known as the Index of Ideality of Correlation (IIC) has been suggested. The IIC estimates the predictive potential of QSAR model which is not only based on the correlation coefficient but also depends on the residual values of endpoint and arrangement of the dots image related to the diagonal [10]. The purpose of the current research is to compare the IIC with other different well-known criteria of predictive potential of QSAR models for activators of glucokinase in T2D.
Materials and methods
In the present study, a data set consisting of 67 benzamide derivatives was used for QSAR model development. The experimental values for EC50 data were retrieved from literature reports [11,12,13]. Then, these experimental values were changed into negative decimal logarithm (pEC50) which was considered as the dependent variable for QSAR model generation [14]. 3D arrangement of the glucokinase activators were sketched with Marvin Sketch [15], and further, Open Babel [16] was used to convert them into the SMILES depiction. Three different splits were prepared by random distribution of molecules into training, invisible training, calibration, and external validation sets [17]. The training, invisible training sets are like the manufacturer and inspector of the correlation weights, and the external validation set is the indicator of the true predictive potential of the correlation weights [18]. OECD guidelines were precisely followed in QSAR model development [19]. The percentage of the identical distribution of compounds into splits was determined with the well-known method [20], and it is summarized in Table 1. From this table, nonidentical nature of splits can be confirmed.
Optimal descriptors
The CORAL QSAR modeling depends on the concept, described in the following Eq. 1 [21]:
Simplified molecular-input line-entry system (SMILES) notation is regarded as the most suitable depiction for the molecular structures of compounds [22]. In QSAR modeling, molecular optimal descriptor (DCW) is defined as the function of the molecule’s SMILES notation, described in Eq. 2 [23].
Molecular structures of compounds can be shown as SMILES and molecular graph; in several cases, hybrid representation is also used [24]. In hybrid form, both SMILES and molecular graph are employed for model development in QSAR modeling. The CORAL method depends on correlation weights of structural attributes obtained from hydrogen-suppressed graph (HSG), hydrogen-filled graph (HFG), and graph of atomic orbitals (GAO). There are two types of molecular features, named as local and global which are extracted from HSG [25]. The hybrid descriptor based on Monte Carlo simulation of the activators of glucokinase was computed with the following equation [26]:
Index of ideality of correlation (IIC)
In Monte Carlo optimization, sets of correlation weights CW (x) are the coefficients which result in production of the target function with higher value. Different target functions can be calculated for available optimization method by changing the value of parameter WIIC. Here, two versions of the target function were evaluated. The target function is defined as [27]:
where Rtraining and Rinvisible training are the correlation coefficients between the observed and predicted endpoints for the training and invisible training sets, respectively.
The IIC is described as the index of ideality of correlation. The WIIC is an experimental coefficient; generally its value is considered as zero in the Monte Carlo optimization. But in the case of modified version, the value of WIIC is taken as greater than zero, but too large value of WIIC can also ruin the optimization process.
Index of ideality of correlation is defined as:
In Eqs. 7 and 8, the parameters Yobs and Ypred are correspondingly observed and calculated values of pEC50 for the calibration set. If we use IIC as a replacement of the conventional correlation coefficient, the statistical parameters of any inferior models could be improved. Hence, the IIC can be taken as an alternative option to check the characteristic of developed model. The application of the IIC becomes impossible if [27]:
How to rate different criteria of predictive potential as correct or wrong? [28]
(then the rating is given as correct)
And
The rating is given as “correct” if the values of the criteria for both calibration and validation set for model 1 are higher than model 2 or values or the parameters of both calibration and validation set related to model 2 are more than 1. But in comparison of model 1 with model 2, if the value of the X [1] increases for calibration set and the value of R2 decreases for validation set, then rating is given as “wrong.” The XCLB [1] and XCLB [2] demonstrated the values of criteria R2, q2, q2F1, q2F2, Q2F3, Rm2, CCC, and IIC for model 1 and model 2, where models 1 and 2 were prepared with TF1 and TF2 with WIIC values 0 and 0.2, respectively.
In CORAL QSAR modeling, the dispersion in several runs of the same split with same optimization procedure can be explained and calculated with the standard deviation. Along with this, the developed splits could be categorized as correct and uncertain. Firstly, the average values of a criteria X1 for model-1 and X2 for model-2 were calculated, and then their standard deviations ∆1 and ∆2 were determined which were developed with target function TF1 and TF2. On the basis of following inequality, it can be defined as uncertain or correct [29].
Max (∆1, ∆2) = ∆1 > ∆2, otherwise ∆2 is taken
Then, in the standard deviations of a criteria ∆1, ∆2 maximum standard deviation value is determined. Suppose if the value of ∆1 is greater than ∆2, then ∆1 is considered as “Max ∆.” Further, if the difference between the average values of criteria was lower than the max ∆ value, then it is recommended as “uncertain,” and opposite of above statement is supposed to be the “correct.”
Building of CORAL model
Three steps involved in the development of the CORAL QSAR models were [30] the following:
- 1.
The total data set was divided into the training, invisible training, calibration, and validation sets, and different splits were generated by running the CORAL SEA 2019 with the search for preferable number of epochs (N*) and threshold (T); ranges of T and Nepoch were selected from 1 to 10 and 1 to 50, respectively.
- 2.
Then the models were developed with preferable number of threshold (3) and Nepoch (25), and molecular features for all compounds were computed by mean of CORAL.
- 3.
Correlation weights were extracted for all molecular features related to QSAR models.
Figure 1 represents the general scheme used of CORAL model development with Monte Carlo method [31].
Domain of applicability
The statistical defects related to the molecular features depend on allocation of different molecular features into the training and calibration set [28].
where PT(FK) and PC(FK) are probabilities of feature FK to be in training set and calibration set and NT(FK) and NC(FK) are prevalence of feature FK in the training set and calibration sets, respectively.
The defect of the individuals’ SMILES can be calculated as:
The addition of defect of individual SMILES results into the defect of the split related to training, invisible training, calibration, and validation set.
Domain of applicability can be estimated as
where the d \( \overline{\left(\mathrm{SMILES}\right)} \) is the average of the statistical defect of SMILES related to the training set.
Results and discussion
The major purpose behind the use of different criteria to predict the potential of developed QSAR models was to identify that the built models have predictability control or not. The comparison of q2, Rm2, CCC, and IIC provided the satisfactory outcome in terms of the predictive potential of the QSAR model because all the criteria have comparable range from zero to one. Moreover, during comparison of two models, one having larger value of criteria is assumed as superior, and this is true for all above mentioned parameters [10]. Tables 2, 3, and 4 are describing the comparison of different statistical characteristics of three runs of split 1, 2, and 3 of glucokinase activators with Monte Carlo optimization. According to the rating principle, in case of split 1, for IIC, rating was identified as correct in three run, while for R2, CCC, Rm2, and q2 matrices, it was correct only for two runs. In split 2, rating was correct for IIC in all three runs, but for other criteria, it was correct only for two runs, and lastly in split 3, rating was obtained as correct for all the statistical parameters. From the interpretation of above data, it could be observed that the splits prepared with TF2 were better than the TF1 and the first run of split 3 was defined as the best split prepared due to having highest values of R2 (0.9531) and IIC (0.9758). Different QSAR equations of various runs of three splits with target function TF2 are summarized in Table 5, and the rating of recommendations provided by criteria in the three splits of glucokinase is described in Table 6. According to the criteria of standard deviation, splits 1 and 2 were correct or certain for all statistical parameters except Rm2 matrices although split 3 was uncertain for all criteria. The percentage of correct recommendations estimated for different criteria of the predictive potential of QSAR models is listed in Table 7. The percentage of correct recommendations for IIC was calculated as highest 100% followed by q2 matrices with 89% and lastly for the R2, CCC, and Rm2 with 78%. Percentage according to the standard deviation was 67% for all parameters except Rm2 matrices. Figure 2 displays the graphical representation of the IIC versus target function TF1 and TF2.
Mechanistic interpretation
From the data related to the correlation weight of the developed QSAR models, different structural attributes can be framed as stable positive category, stable negative category, and undefined category [32]. Stable positive category is accountable for the enhancement of the calculated endpoint in all prepared splits, while other negative are contradictory of the above statement. Some structural attributes have not a particular role; they have both positive and negative values of descriptors in different runs, and thus, for such attributes, an accurate correlation weight cannot be expressed [33]. Structural attributes extracted from the best split (first run of split 3) are summarized in Table 8 along with their correlation weights, and Fig. 3 shows the SMILES attributes present in one of the glucokinase activators.
Conclusion
The CORAL software provided the robust and predictive QSAR models for the activators of glucokinase containing benzamide moiety. In comparison of the predictive potential of these models, the index of ideality of correlation emerged as a useful criterion. Application of IIC with target functions resulted in improvement of statistical quality of all QSAR models related to different splits. The coefficient WIIC controlled the effect of the index of ideality of correlation in Monte Carlo optimization which is an empirical parameter and depends on the nature of endpoint and compounds diversity of corresponding available data. Hence, IIC can be used for prediction of glucokinase activation in a lucid way.
References
Bebernitz GR, Beaulieu V, Dale BA, Deacon R, Duttaroy A, Gao J, Grondine MS, Gupta RC, Kakmak M, Kavana M, Kirman LC, Liang J, Maniara WM, Munshi S, Nadkarni SS, Schuster HF, Stams T, Denny IS, Taslimi PM, Vash B, Caplan SL (2009) Investigation of functionally liver selective glucokinase activators for the treatment. J Med Chem 52:6142–6152
Bonn P, Brink DM, Fägerhag J, Jurva U, Robb GR, Schnecke V, Svensson A, Waring MJ, Westerlund C (2012) The discovery of a novel series of glucokinase activators based on a pyrazolopyrimidine scaffold. Bioorg Med Chem Lett 22:7302–7305
Charaya N, Pandita D, Grewal AS, Lather V (2018) Design, synthesis and biological evaluation of novel thiazol-2-yl benzamide derivatives as glucokinase activators. Comput Biol Chem 73:221–229
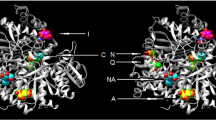
Kumari V, Li C (2008) Comparative docking assessment of glucokinase interactions with its allosteric activators. Curr Chem Genomics 2:76–89
Bertram LS, Black D, Briner PH, Chatfield R, Cooke A, Fyfe MCT, Murray PJ, Rasamison CM, Reynet C, Schofield KL, Shah VK, Spindler F, Taylor A, Turton R, Williams GM, Wong-kai-in P, Yasuda K (2008) Pharmacokinetics, safety and efficacy of glucokinase activating 2-(4-sulfonylphenyl)-N-thiazol-2-ylacetamides : discovery of PSN-GK1. J Med Chem 51:4340–4345
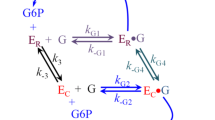
Antoine M, Boutin JA, Ferry G (2009) Binding kinetics of glucose and allosteric activators to human glucokinase reveal multiple conformational states. Biochemistry 48:5466–5482
Bowler JM, Hervert KL, Kearley ML, Miller BG (2013) Small-molecule allosteric activation of human glucokinase in the absence of glucose. ACS Med Chem Lett 4:580–584
Begum S, Achary PGR (2015) Simplified molecular input line entry system-based: QSAR modeling for MAP kinase-interacting protein kinase (MNK1). SAR QSAR Environ Res 26(5):343–361
Begam BF, Kumar JS (2016) Computer assisted QSAR / QSPR approaches – a review. Ind J Sci Tech 9(8). https://doi.org/10.17485/ijst/2016/v9i8/87901
Toropov AA, Toropova AP (2017) The index of ideality of correlation: a criterion of predictive potential of QSPR / QSAR models ? Mutat Res Gen Tox En 819:31–37
Park K, Lee BM, Kim YH, Han T, Yi W, Lee DH, Choi HH, Chong W, Lee CH (2013) Discovery of a novel phenylethyl benzamide glucokinase activator for the treatment of type 2 diabetes mellitus. Bioorg Med Chem Lett 23:537–542
Park K, Lee BM, Hyun KH, Lee DH, Choi HH, Kim H, Chong W, Kim KB, Nam SY (2014) Discovery of 3-(4-methanesulfonylphenoxy)-N-[1-(2-methoxyethoxymethyl)-1H-pyrazol-3-yl]-5-(3-methylpyridin-2-yl)-benzamideas a novel glucokinase activator (GKA) for the treatment of type 2 diabetes mellitus. Bioorg Med Chem 22:2280–2293
Park K, Lee BM, Hyun KH, Han T, Lee DH (2015) Design and synthesis of acetylenyl benzamide derivatives as novel glucokinase activators for the treatment of T2DM. ACS Med Chem Lett 6:296–301
Toropova AP, Toropov AA, Veselinovic JB, Miljkovi FN, Veselinovic AM (2014) QSAR models for HEPT derivates as NNRTI inhibitors based on Monte Carlo method. Eur J Med Chem 77:298–305
Marvin Sketch v.14.11.17.0, (2014) ChemAxon, XhemAxon KFT. Budapest, Hungary
O’Boyle N, Banck M, James CA, Morley C, Vandermeersch T, Hutchison GR (2011) Open babel: an open chemical toolbox. J.Cheminform. 3:33
Kumar P, Kumar A, Sindhu J, Lal S (2019) QSAR models for nitrogen containing monophosphonate and bisphosphonate derivatives as human farnesyl pyrophosphate synthase inhibitors based on Monte Carlo method. Drug Res 69:159–167
Toropova AP, Toropov AA (2017) The index of ideality of correlation: a criterion of predictability of QSAR models for skin permeability ? Sci Total Environ 586:466–472
OECD principles for the validation, for regulatory purposes, of (quantitative) structure-activity relationship models. Available at:http://www.oecd.org/chemicalsafety/risk-assessment/37849783.pdf
Kumar A, Chauhan S (2016) Use of the Monte Carlo method for OECD principles-guided QSAR modeling of SIRT1 inhibitors. Arch Pharm Chem Life Sci 349:1–9
Zivkovic JV, Truti NV, Veselinovic JB, Nikoli GM, Veselinovic AM (2015) Monte Carlo method based QSAR modeling of maleimide derivatives as glycogen synthase kinase-3 β inhibitors. Comp Biol Med. https://doi.org/10.1016/j.compbiomed.2015.07.004
Kumar A, Chauhan S (2016) QSAR differential model for prediction of SIRT1 modulation using Monte Carlo method. Drug Res 67(3):156–162
Sokolović D, Aleksić D, Milenković V, Karaleić S, Mitić D, Kocić J, Mekić B, Veselinović JB, Veselinović AM (2016) QSAR modeling of bis-quinolinium and bis-isoquinolinium compounds as acetylcholine esterase inhibitors based on the Monte Carlo method–the implication for myasthenia gravis treatment. Med Chem Res 25:2989–2998
Manisha, Chauhan S, Kumar P, Kumar A (2019) Development of prediction model for fructose-1,6-bisphosphatase inhibitors using the Monte Carlo method. SAR QSAR Environ Res 30:145–159
Kumar A, Chauhan S (2018) Use of simplified molecular input line entry system and molecular graph based descriptors in prediction and design of pancreatic lipase inhibitors. Future Med Chem 10:1603–1622
Kumar P, Kumar A (2017) Monte Carlo method based QSAR studies of Mer kinase inhibitors in compliance with OECD principles. Drug Res 68(04):189–195
Toropov AA, Carbó-dorca R, Toropova AP (2017) Index of ideality of correlation : new possibilities to validate QSAR : a case study. Struct Chem 29(1):33–38
Toropov AA, Toropova AP (2018) Use of index of ideality of correlation to improve predictive potential for biochemical endpoints. Toxicol Mech Methods. https://doi.org/10.1080/15376516.2018.1506851
Toropova AP, Toropov AA (2019) Does the index of ideality of correlation detect the better model correctly ? Mol Inf. https://doi.org/10.1002/minf.201800157
Toropova AP, Toropov AA (2018) The index of ideality of correlation : improvement of models for toxicity to algae of models for toxicity to algae. Nat Prod Res. https://doi.org/10.1080/14786419.2018.1493591
Toropova AP (2018) The index of ideality of correlation : hierarchy of Monte Carlo models for glass transition temperatures of polymers. J Polym Res. https://doi.org/10.1007/s10965-018-1618-z
Gaikwad R, Ghorai S, Amin SA, Adhikari N, Patel T, Das K, Jha T, Gayen S (2018) Monte Carlo based modelling approach for designing and predicting cytotoxicity of 2-phenylindole derivatives against breast cancer cell line MCF7. Toxicol Vitr 52:23–32
Rescifina A, Floresta G, Marrazzo A, Parenti C, Prezzavento O, Nastasi G, Amata E, Dichiara M, Amata E (2017) Development of a sigma-2 receptor affinity filter through a Monte Carlo based QSAR analysis. Eur J Pharm Sci 106:94–101
Acknowledgments
The authors are highly indebted to Dr. Andrey A. Toropov and Dr. Alla P. Toropova for providing the CORAL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict(s) of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 26 kb)
Rights and permissions
About this article
Cite this article
Nimbhal, M., Bagri, K., Kumar, P. et al. The index of ideality of correlation: A statistical yardstick for better QSAR modeling of glucokinase activators. Struct Chem 31, 831–839 (2020). https://doi.org/10.1007/s11224-019-01468-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11224-019-01468-w