Abstract
Light-driven water oxidation in photosynthesis occurs at the oxygen-evolving center (OEC) of photosystem II (PSII). Chloride ions (Cl−) are essential for oxygen evolution by PSII, and two Cl− ions have been found to specifically bind near the Mn4CaO5 cluster in the OEC. The retention of these Cl− ions within the OEC is critically supported by some of the membrane-extrinsic subunits of PSII. The functions of these two Cl− ions and the mechanisms of their retention both remain to be fully elucidated. However, intensive studies performed recently have advanced our understanding of the functions of these Cl− ions, and PSII structures from various species have been reported, aiding the interpretation of previous findings regarding Cl− retention by extrinsic subunits. In this review, we summarize the findings to date on the roles of the two Cl− ions bound within the OEC. Additionally, together with a short summary of the functions of PSII membrane-extrinsic subunits, we discuss the mechanisms of Cl− retention by these extrinsic subunits.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Photosystem II (PSII) is a multi-subunit pigment-protein complex found in the thylakoid membranes of oxygenic photosynthetic organisms (Debus 1992). Using light energy, PSII catalyzes the oxidation of water to molecular oxygen, producing most of the atmospheric oxygen on Earth (Dismukes et al. 2001; Barber 2016). The unique water-oxidizing reaction takes place in the oxygen-evolving center (OEC) of PSII, which is formed by several membrane-intrinsic and membrane-extrinsic subunits (Shen et al. 2021).
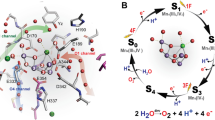
The OEC contains an inorganic Mn4CaO5 cluster, consisting of four Mn ions, one Ca2+ ion, and five oxygen atoms arranged in a “distorted chair” form (Umena et al. 2011; Suga et al. 2015). In the water-oxidizing reaction, two water molecules are converted into one molecular oxygen, four protons, and four electrons in a stepwise manner through the S-state cycle (Si; i = 0–4) of the OEC (Kok et al. 1970). S1 is the most dark-stable state, and flash illumination advances each Si (i = 0–3) state to the next Si+1 state. Molecular oxygen is released in the final step from the transient, most oxidized S4 state to the most reduced S0 state. Several hydrogen-bonded water channels surround the Mn4CaO5 cluster, and may be involved in the inlet of substrate water molecules or the exit of produced protons (Murray and Barber 2007; Ho and Styring 2008; Gabdulkhakov et al. 2009; Umena et al. 2011; Vassiliev et al. 2012; Nagao et al. 2017c; Kern et al. 2018; Kaur et al. 2021). Additionally, two Cl− ions specifically bind within the OEC, near the Mn4CaO5 cluster (Murray et al. 2008; Kawakami et al. 2009; Umena et al. 2011). They are related to some of the hydrogen-bond networks (Saito et al. 2013, 2015; Sakashita et al. 2017a).
Cl− has long been known to be an essential cofactor for oxygen evolution by photosynthesis (Warburg and Lüttgens 1944; Arnon and Whatley 1949; Bové et al. 1963; for review, see Homann 2002), and was discovered to play a role in the water-oxidizing reaction of PSII (Heath and Hind 1969; Izawa et al. 1969). However, it had been challenging to determine the detailed functions of Cl− and also the locations and number of binding sites (Debus 1992; van Gorkom and Yocum 2005; Yocum 2008). Some studies predicted one Cl− binding site (Lindberg et al. 1993; Lindberg and Andréasson 1996), whereas other studies proposed two Cl− binding sites (Boussac 1995; van Vliet and Rutherford 1996). Cl− was implicated as a ligand to the Mn cluster in several studies (Critchley 1985; Rutherford 1989), while some studies using extended X-ray absorption fine structure (EXAFS) spectroscopy suggested the possibility that the Cl− ions were not ligated to the Mn cluster (Yachandra et al. 1986a, b; Haumann et al. 2006). The binding sites of Cl− were finally determined by X-ray crystallography; two Cl− binding sites were found near the Mn4CaO5 cluster, and the Cl− ions were not directly ligated to the Mn4CaO5 cluster (Murray et al. 2008; Kawakami et al. 2009; Umena et al. 2011) (Fig. 1).
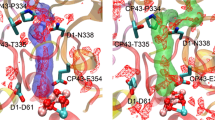
Locations of the two Cl− binding sites in the OEC of PSII (PDB ID: 3WU2). D1, D2, and CP43 are shown in green, blue, and yellow cartoon views, respectively. The Cl− ions are shown as pink spheres, the water molecules nearby the Cl− ions are shown as orange spheres, and the amino acid residues associated to the Cl− ions are shown as stick models
Studies using green plant PSII showed that the membrane-extrinsic subunits are important for the retention of Cl− ions required for optimal oxygen-evolving activity (Akabori et al. 1984; Andersson et al. 1984; Miyao and Murata 1985). As will be described in a later section (“Extrinsic subunits of PSII retaining Cl− in the OEC”), differences are observed in the compositions of extrinsic subunits among the various oxygenic phototrophs (De Las Rivas et al. 2004; Bricker and Burnap 2005; Enami et al. 2005, 2008; Roose et al. 2007b, 2016; Bricker et al. 2012; Ifuku 2015; Ifuku and Noguchi 2016; Ifuku and Nagao 2021). However, extrinsic subunits in other species were also found to play a role in Cl− retention (Shen et al. 1997, 1998; Enami et al. 1998; Nagao et al. 2010a). The detailed mechanism of Cl− retention by the extrinsic subunits remains to be fully elucidated.
Since the report of the cyanobacterial PSII crystal structure at 1.9 Å resolution (Umena et al. 2011), dramatic progress has been made in understanding the functions of Cl− ions bound within the OEC. The functions of extrinsic subunits have been extensively studied as well. The recently reported PSII structures of various species (Ago et al. 2016; Wei et al. 2016; Su et al. 2017; Nagao et al. 2019; Pi et al. 2019) have facilitated a deeper understanding of their functions.
This review mainly focuses on the roles of the two OEC-bound Cl− ions in the water-oxidizing reaction of PSII, as well as the structural mechanisms of Cl− retention by membrane-extrinsic subunits. The roles of the Cl− ions and the characteristics of extrinsic subunits are summarized, followed by a detailed discussion on the functional regions of extrinsic subunits that could be important for Cl− retention in PSII. There have been excellent, comprehensive reviews that provide further information on the water-oxidizing reaction of PSII (McEvoy and Brudvig 2006; Vinyard et al. 2013; Yano and Yachandra 2014; Shen 2015; Lubitz et al. 2019).
Roles of Cl− ions in the OEC of PSII
Effect of Cl− ions on the water-oxidizing reaction by the Mn4CaO5 cluster
The two Cl− binding sites in the OEC of PSII are called the Cl-1 and Cl-2 binding sites (Murray et al. 2008; Kawakami et al. 2009; Umena et al. 2011) (Fig. 1). The Cl-1 binding site, which had also been confirmed by Guskov et al. (2009), is surrounded by the amino group of D2-Lys317, the backbone nitrogen of D1-Glu333, and the side chain of D1-Asn181. The Cl-2 binding site is surrounded by the backbone nitrogen of D1-Asn338, D1-Phe339, and CP43-Glu354. The Cl− ions are not directly ligated to the Mn4CaO5 cluster, but D1-Glu333 and CP43-Glu354 are both directly involved in the coordination of the Mn4CaO5 cluster, with D1-Glu333 serving as a bidentate ligand to Mn3 and Mn4, and CP43-Glu354 as a bidentate ligand to Mn2 and Mn3 (Umena et al. 2011).
Electron paramagnetic resonance (EPR) spectroscopy analyses, performed to study the structure and the magnetic properties of the Mn4CaO5 cluster (Debus 1992), demonstrated that Cl− affects the Mn4CaO5 cluster in the S2 state (Dismukes and Siderer 1981; Pokhrel et al. 2011). A characteristic g = 2 multiline signal was observed in the S2 state in the presence of Cl− (Dismukes and Siderer 1981). Depletion of Cl− significantly lowered the above multiline signal intensity, while giving rise to a different g = 4.1 signal (van Vliet and Rutherford 1996). The EPR properties were recovered by resupplying Cl− (Ono et al. 1986). Similar effects of Cl− on the g = 2 multiline EPR signal in the S2 state were observed by comparing untreated PSII and extrinsic subunit-depleted PSII under different Cl− conditions. The S2 multiline signal was not significantly affected by the surrounding Cl− concentration of untreated PSII (Franzén et al. 1985), whereas in NaCl-washed PSII, lacking extrinsic subunits (PsbP and PsbQ) required for Cl− retention, the multiline signal was diminished by depletion of Cl− from the buffer solution (Toyoshima et al. 1984; De Paula et al. 1986; Imaoka et al. 1986). Studies including Fourier transform infrared (FTIR) spectroscopy further showed that the Cl-1 site is structurally coupled with the Mn4CaO5 cluster (Service et al. 2013; Suzuki et al. 2013).
It has been suggested that the g = 2 and g = 4.1 forms indicate two different conformations (open-cubane and closed-cubane) of the Mn4CaO5 cluster in the S2 state (Isobe et al. 2012; Pantazis et al. 2012; Bovi et al. 2013). It has been proposed that their stabilities are determined by the hydrogen-bond network around Cl-1, especially by proton transfer between D1-Asp61 and W1, a water molecule ligand to Mn4 of the Mn4CaO5 cluster (Pokhrel and Brudvig 2014; Amin et al. 2016; Yang et al. 2021). Recent theoretical studies have suggested that Cl-1 inhibits deprotonation of W2, another water molecule ligand to Mn4, and facilitates proton transfer from W1 via D1-Asp61 (Saito et al. 2020a; Mandal et al. 2022). Removal of Cl-1 has been shown to inhibit electron transfer from S2 to YZ (D1-Tyr161) by increasing the redox potential (Em) of S2/S3 (Mandal et al. 2022). The removal also leads to the formation of a salt bridge between D1-Asp61 and D2-Lys317, which would hinder the release of protons from D1-Asp61 via the water channel (Cl-1 channel) passing through this location (Rivalta et al. 2011; Amin et al. 2016; Mandal et al. 2022). Recently, it has been suggested that Cl-1 also affects the Mn4CaO5 cluster photoassembly in a similar way (Vinyard et al. 2019; Russell and Vinyard 2021).
The functions of Cl− have also been studied using a combination of Cl− substitution with other anions, and EPR spectroscopy or FTIR spectroscopy. Some of the most frequently used anions for Cl− substitution are bromide (Br−), iodide (I−), fluoride (F−), and nitrate (NO3−) ions. The oxygen-evolving activity of Cl−-depleted PSII is greatly recovered by Br−, partially by I− and NO3−, and barely by F−. The order of effectiveness in supporting the water-oxidizing reaction is Cl− ≥ Br− > I− ≥ NO3− > F−, with the effectiveness of I− and NO3− inverted at higher concentrations (Hind et al. 1969; Kelley and Izawa 1978; Ono et al. 1987; Homann 1988a; Hasegawa et al. 2002). EPR studies have shown that while the S2 multiline signal can be observed in Br−-substituted PSII, it is remarkably diminished by substitution with other anions, such as F− (Casey and Sauer 1984; Damoder et al. 1986; Yachandra et al. 1986b; Ono et al. 1987). FTIR studies have investigated the effects of Cl− on the conformational change of the OEC, mostly upon the S1-to-S2 transition. The overall features of the FTIR spectra were similar among untreated PSII and PSII with Br−, NO3−, or I− substituted for Cl−, although that of F−-substituted PSII showed significant suppression of different bands (Hasegawa et al. 2002). However, the spectral features in the amide I region (1700–1600 cm−1) that was changed by Cl− depletion and recovered by Cl− reconstitution were not restored by anion substitution with Br−, NO3−, and I− (Hasegawa et al. 2002; Kondo and Noguchi 2018). The effects on the amide I bands have been attributed to conformational changes, including those of the polypeptide chains surrounding the Cl− binding site(s) (Kondo and Noguchi 2018). The X-ray crystallographic studies mentioned earlier confirmed that Br− and I− can bind to the Cl− binding site (Murray et al. 2008; Kawakami et al. 2009; Umena et al. 2011). The collective findings suggest that, in addition to the negative charge, other properties such as the size of the Cl− ion also affect the function of Cl− in the water-oxidizing reaction.
Several studies have probed for the roles of Cl− by introducing mutations at residues D2-Lys317 and D1-Asn181, both associated to Cl-1 with their side chains. In the D2-K317R mutant of Synechocystis sp. PCC 6803, Cl-1 remained bound, although with lowered affinity (Pokhrel et al. 2013; Suzuki et al. 2013). From the changes observed in the FTIR spectra (Pokhrel et al. 2013; Suzuki et al. 2013), EPR spectra, flash-induced O2 yield patterns, and O2-release kinetics (Pokhrel et al. 2013) by this mutation, and Cl−/NO3− replacement in the wild-type (WT) strain and D2-K317R mutant (Suzuki et al. 2013), it was suggested that D2-Lys317 and Cl-1 have structural effects on the Mn4CaO5 cluster, and that they affect proton transfer through the Cl-1 channel. The D2-K317A, D2-K317Q, and D2-K317E mutants were also studied, and PSII with the D2-K317A mutation was found to lose Cl− binding at the Cl-1 site while remaining active for oxygen evolution, although at a lower rate than WT PSII (Pokhrel et al. 2013). It was suggested that Cl-1 is required when the residue at the D2-Lys317 position is positively charged, in order to prevent the formation of a salt bridge with D1-Asp61. The lack of Cl− requirement for oxygen-evolving activity in D2-K317A PSII may also suggest that binding of Cl-2 has little effect on the activity in D2-K317A PSII. Although D1-Asn181 is another residue associated to Cl-1, the D1-N181A and D1-N181S mutations showed some different effects compared to mutations at D2-Lys317 (Pokhrel et al. 2015). In both D1-N181A and D1-N181S PSII, Cl-1 remained bound, but the steady-state oxygen-evolving activities were lower than half of that of WT PSII. While the midfrequency S2-minus-S1 FTIR difference spectra were similar to that with the D2-K317A mutation, flash-induced O2 yield measurements revealed normal S-state cycling with a miss factor similar to that of WT PSII, although the O2-release kinetics were slow. The authors concluded that the D1-N181A and D1-N181S mutations did not have significant effects on proton transfer, but slowed O–O bond formation by affecting the geometry of a substrate water. While a number of studies, including those mentioned above, have emphasized the importance of Cl-1 for the water-oxidizing reaction of PSII, to date, none have clarified whether or not Cl-2 is essential for the reaction.
Banerjee et al. (2018) showed that compared to cyanobacterial PSII, spinach PSII is more sensitive to Cl− depletion, and that the D1-N87A mutation in cyanobacterial PSII, which replicates residue 87 of D1 in spinach PSII, leads to similar Cl−-binding properties as spinach PSII. This D1-(Asn/Ala)87 residue had been found to be the most significant difference in D1 residues of cyanobacterial PSII and land plant PSII near the water channels (Vogt et al. 2015). In contrast to D1-N87A, the Cl− effect in D1-N87D mutant cyanobacterial PSII was similar to that of WT cyanobacterial PSII, while the S-state cycling was impaired unlike in D1-N87A (Banerjee et al. 2018). In the study, the authors concluded that the effect of Cl− (Cl-1) on the oxygen-evolving activity of PSII can be modulated by residue 87 of D1 by affecting the hydrogen-bond network that interconnects this residue with the Cl-1 site.
Effect of Cl− ions on water channels
As previously mentioned, the Mn4CaO5 cluster is surrounded by several networks of hydrogen-bonded water molecules (Umena et al. 2011). These water channels, some of which are affected by the Cl− ions as had been proposed by Olesen and Andréasson (2003), could be the pathways for the inlet of substrate water molecules or for the exit of protons. There are five main water channels: the O1, YZ-N298, Cl-1, O4, and Cl-2 channels (Nagao et al. 2017c; Sakashita et al. 2017a; Suga et al. 2019). The O1 channel, also referred to as the “large channel” (Ho and Styring 2008), proceeds from a large water cluster interacting with YZ (D1-Tyr161), D1-His190, and D1-Asn298, and passes through the O1 site of the Mn4CaO5 cluster and the D1-Glu329 residue (Murray and Barber 2007; Sakashita et al. 2017a, b; Suga et al. 2019). It should be noted, however, that while the O1 channel is observed in PSII structures from thermophilic cyanobacteria, this channel was recently reported to be blocked in PSII from Synechocystis sp. PCC 6803, a mesophilic cyanobacteria (Gisriel et al. 2022). The YZ-N298 channel starts from the same water cluster as the O1 channel but passes through D1-Asn332 (Umena et al. 2011; Nakamura et al. 2014; Shen 2015; Nagao et al. 2017c). The Cl-1 channel, also referred to as the “E65/E312 channel” (Sakashita et al. 2017a) or the “broad channel” (Ho and Styring 2008), proceeds from D1-Asp61 through the Cl-1 binding site and the D1-Glu65/D2-Glu312 pair (Ferreira et al. 2004; Ishikita et al. 2006; Umena et al. 2011; Saito et al. 2013). The O4 channel, also referred to as the “narrow channel” (Ho and Styring 2008), starts from the O4 site of the Mn4CaO5 cluster and passes through D1-Asn338 (Saito et al. 2015; Takaoka et al. 2016; Sakashita et al. 2017a; Suga et al. 2019; Shimada et al. 2020; Kuroda et al. 2021). The Cl-2 channel consists of a hexagonal hydrogen-bonded network formed by Cl-2 and five water molecules near the Mn4CaO5 cluster (Sakashita et al. 2017a), and a water channel proceeding from the lumenal bulk surface through PsbU-Asp96 and PsbU-Asn99, ending near the hexagonal hydrogen-bonded network (Umena et al. 2011). This channel is different from the aforementioned channels in that it does not connect the Mn4CaO5 cluster with the bulk surface because of interruption by the polypeptide backbone of the C-terminal region of D1 (Sakashita et al. 2017a). Despite extensive studies on these channels, no consensus has been established regarding the functions of each of them.
Of the five channels mentioned above, Cl-1 is involved in the Cl-1 channel (Fig. 2a). Although the Cl− binding sites had not yet been determined at that time, Ferreira et al. (2004) identified the channel corresponding to the Cl-1 channel in the PSII crystal structure that they obtained. The possibility that this channel functions as a proton-exit pathway or the water-inlet pathway was proposed (Barber et al. 2004; De Las Rivas and Barber 2004; Ferreira et al. 2004). A theoretical study by Ishikita et al. (2006) supported the view that this channel could function as a proton-exit pathway. Later, when Cl-1 was found to be located in this channel, it was proposed that Cl-1 could maintain the structure of this channel and/or proton transfer through this channel (Murray et al. 2008; Kawakami et al. 2009). Numerous theoretical and experimental studies have further investigated the function of the Cl-1 channel. Many studies have suggested that this channel is a proton-release pathway (Gabdulkhakov et al. 2009; Pokhrel et al. 2011; Rivalta et al. 2011; Ghosh et al. 2019; Ibrahim et al. 2020; Saito et al. 2020a; Hussein et al. 2021; Kaur et al. 2021; Kuroda et al. 2021). However, the channel may also play a role in the inlet of water molecules (Vassiliev et al. 2010, 2012; Sakashita et al. 2017a; Reiss et al. 2019; de Lichtenberg et al. 2021). As previously mentioned, Cl-1 has been suggested to be important for the structure of this channel by preventing the formation of a salt bridge between D1-Asp61 and D2-Lys317 (Rivalta et al. 2011; Pokhrel et al. 2013; Amin et al. 2016; Mandal et al. 2022), and for proton transfer through this channel (Kawashima et al. 2018; Saito et al. 2020a; Mandal et al. 2022).
Water channels found near the two Cl− binding sites in the OEC of PSII (PDB ID: 3WU2). The two Cl− ions are shown as green spheres, the four water molecules ligand to the Mn4CaO5 cluster (W1–W4) are shown as light blue spheres, and some of the residues nearby water channels are shown as yellow stick models. The arrows show the direction to the thylakoid lumen. a Cl-1 channel passing through the Cl-1 binding site. The water molecules participating in the Cl-1 channel are shown as dark blue spheres. b, c Water channels related to Cl-2. b Cl-2 channel (hexagonal Cl-2–water network) consisting of Cl-2 and five water molecules, and O1 channel that could connect to the Cl-2 channel. Water molecules in the Cl-2 channel and the O1 channel are shown as blue and pink spheres, respectively. c O4 channel and Cl-2 channel. Water molecules in the O4 channel and the Cl-2 channel are shown as orange and blue spheres, respectively
In contrast to Cl-1, few studies have focused on the role of Cl-2 in hydrogen-bond networks. Of the five aforementioned channels, Cl-2 is related to the O4 channel and the Cl-2 channel (Fig. 2b, c). The O4 channel was first identified by Ho and Styring (2008) through solvent-accessibility simulations. The authors proposed that this channel could function as a proton-exit pathway. Many studies have suggested that the O4 channel plays a role in the release of protons (Gabdulkhakov et al. 2009; Saito et al. 2015, 2020b; Takaoka et al. 2016; Suga et al. 2017, 2019; Shimizu et al. 2018; Reiss et al. 2019; Ibrahim et al. 2020; Sakashita et al. 2020; Shimada et al. 2020; Yamamoto et al. 2020; Okamoto et al. 2021), whereas others have proposed its role in the inlet of water molecules (Vassiliev et al. 2010, 2012; Askerka et al. 2016; Retegan and Pantazis 2016; Retegan et al. 2016; Pantazis 2019). The most recent reports, including FTIR studies and theoretical studies taking the protein environment into account, support proton release through the O4 channel in the S0-to-S1 transition (Saito et al. 2020a, b; Shimada et al. 2020; Yamamoto et al. 2020). Residues D1-Asn338 and CP43-Glu354, both coordinated with Cl-2, interact with water molecules involved in the O4 channel (Fig. 2c). CP43-Glu354 seems especially important for this channel, since it interacts with two of the four water molecules that are arranged in single file from O4 (Sakashita et al. 2020). Therefore, it can be assumed that Cl-2 is required to sustain the optimal structure of the O4 channel. It should be noted that while the O4 channel is suggested to be the proton-exit pathway upon the S0-to-S1 transition (Saito et al. 2015), studies on the Cl− requirement of the S-state cycle indicated that the S0 state can advance to the S1 state in the absence of Cl− (Itoh et al. 1984; Wincencjusz et al. 1997). Cl-2 may be an efficiency-enhancing factor for proton release through the O4 channel.
In the Cl-2 channel, the other channel affected by Cl-2, Cl-2 participates in the hydrogen-bonded network itself (Fig. 2b, c). Nevertheless, in contrast to the intensively studied O4 channel, very few studies have focused on the Cl-2 channel. The Cl-2 channel was reported in the high-resolution crystal structure of PSII by Umena et al. (2011), and was initially proposed to function as a proton-exit channel or a water-inlet channel. However, the polypeptide backbone of the C-terminal region of D1 prevents the Cl-2 channel from connecting the Mn4CaO5 cluster to the bulk surface (Sakashita et al. 2017a). Until recently, no studies had explicitly investigated the function of the Cl-2 channel. A recent theoretical study based on hybrid ab initio quantum mechanics/molecular mechanics (QM/MM) molecular dynamics (MD) simulations proposed proton release through this channel by a novel mechanism in which unidirectional proton transfer occurs through a peptide bond (Nakamura et al. 2019). However, this proposed mechanism has not been supported or refuted in other studies. Sakashita et al. (2017a) showed through MD simulations using the PSII structure of thermophilic cyanobacteria that the water molecules constructing the hexagonal hydrogen-bonded network together with Cl-2 originate from the O1 channel. The authors suggested that this hexagonal hydrogen-bonded network could be connected to the O1 channel by reorientation of the C-terminal D2-Leu352 residue. This could be the same in green plant PSII in which these channels have been found to be conserved (Sakashita et al. 2017b), although it may be different in mesophilic cyanobacterial PSII in which the O1 channel was observed to be blocked (Gisriel et al. 2022). Further investigation is required to determine the roles of the Cl-2 channel or the hexagonal Cl-2–water network.
Extrinsic subunits of PSII retaining Cl− in the OEC
Compositions and binding sites
The membrane-extrinsic subunits bind to the lumenal side of PSII and surround the Mn4CaO5 cluster to form the OEC. Although the mechanism of water oxidation and the basic subunit structure of the PSII core are highly conserved among oxygenic photosynthetic organisms, ranging from cyanobacteria to diverse photosynthetic eukaryotes, drastic evolutionary changes have been observed in the composition of the extrinsic subunits (De Las Rivas et al. 2004; Bricker and Burnap 2005; Enami et al. 2005, 2008; Roose et al. 2007b, 2016; Bricker et al. 2012; Ifuku 2015; Ifuku and Noguchi 2016; Ifuku and Nagao 2021). Green plants, including land plants and green algae, have a set of three extrinsic subunits: PsbO, PsbP, and PsbQ (Åkerlund and Jansson 1981; Yamamoto et al. 1981; Kuwabara and Murata 1982). Only PsbO is always observed among all oxyphototrophs. In cyanobacterial PSII, PsbV and PsbU are present instead of PsbP and PsbQ (Bowes et al. 1983; Stewart et al. 1985; Shen et al. 1992; Shen and Inoue 1993), although they possess CyanoP and CyanoQ, which are homologs of PsbP and PsbQ, respectively (Kashino et al. 2002; De Las Rivas et al. 2004; Thornton et al. 2004). Both CyanoP and CyanoQ are absent in the PSII structures of thermophilic cyanobacteria. However, a recently reported cryo-electron microscopy (cryo-EM) structure of PSII from a mesophilic cyanobacterium revealed the binding of CyanoQ in addition to PsbO, PsbV, and PsbU (Gisriel et al. 2022). Red algal PSII and diatom PSII bind PsbQ´, a homolog of CyanoQ, in addition to PsbO, PsbV, and PsbU (Enami et al. 1995, 1998; Ohta et al. 2003; Nagao et al. 2007). Diatom PSII also binds an additional extrinsic subunit, Psb31 (Nagao et al. 2007, 2010b; Okumura et al. 2008).
X-ray crystallography and cryo-EM have revealed the binding sites of the membrane-extrinsic proteins in cyanobacterial (Umena et al. 2011; Suga et al. 2015; Gisriel et al. 2022), red algal (Ago et al. 2016), green plant (Wei et al. 2016; Su et al. 2017; Sheng et al. 2019), and diatom PSII (Nagao et al. 2019; Pi et al. 2019) (Fig. 3). PsbO binds to a similar location in different species. The binding sites of PsbV and PsbU are identical among the PSII of cyanobacteria, red algae, and diatoms, and are spatially replaced by PsbP in green plant PSII. The PsbQ binding site of green plant PSII is either vacant or occupied by CyanoQ in cyanobacterial PSII, while it is occupied by PsbQ´ in red algal and diatom PSII.
Binding sites of extrinsic subunits in PSII from different species. PSII structures from a cyanobacteria (Thermosynechococcus vulcanus; PDB ID: 3WU2), b green plants (Spinacia oleracea; PDB ID: 3JCU), c red algae (Cyanidium caldarium; PDB ID: 4YUU), and d diatoms (Chaetoceros gracilis; PDB ID: 6JLU) are shown in cartoon view with the extrinsic subunits colored; PsbO (blue), PsbV (red), PsbU (yellow), PsbP (orange), PsbQ and PsbQ´ (pink), and Psb31 (green). The two Cl− ions bound in the OEC are shown as green spheres, and the Mn4CaO5 cluster is shown with red, purple, and black spheres. Note that Cl-1 is not observed in (c) and (d), presumably due to insufficient resolution or loss of the ion during experimental procedures
Fundamental functions
Although the composition of PSII extrinsic subunits varies among different species, the sets of these subunits seem to share fundamental functions. A major role of the extrinsic subunits is to form a functional, stable OEC that is shielded from the bulk solution. As can be seen in the PSII structures, the extrinsic subunits interact with lumenal domains of the PSII core subunits, covering up the Mn4CaO5 cluster and Cl− ions bound at the lumenal surface of the membrane-intrinsic subunits (Umena et al. 2011; Ago et al. 2016; Wei et al. 2016; Nagao et al. 2019). Studies on green plant PSII have shown that depletion of extrinsic subunits leads to an exposed Mn4CaO5 cluster, enabling access of exogenous reductants (Ghanotakis et al. 1984a, c; Ifuku et al. 2005b). Similarly, it has been suggested that depletion of extrinsic subunits in cyanobacterial PSII destabilizes the OEC structure (Shen et al. 1995a, 1998; Inoue-Kashino et al. 2005). A recent study using high-speed atomic force microscopy (HS-AFM) on green plant PSII illustrated that extrinsic subunits are required to fix the flexible CP43 lumenal domain (Pro304–Pro400), including residues directly interacting with the Mn4CaO5 cluster and Cl-2 (Tokano et al. 2020). Meanwhile, cryo-EM structures of assembly intermediates of cyanobacterial PSII, associated with assembly factors but not with the extrinsic subunits, suggested that even with the assembly factors bound, parts within the CP43 lumenal domain become flexible (Huang et al. 2021; Zabret et al. 2021). Furthermore, FTIR studies in different species have shown that the extrinsic subunits are required for the proper conformational change of the OEC during the S1-to-S2 transition (Tomita et al. 2009; Uno et al. 2013; Nagao et al. 2015). While shielding the OEC, the extrinsic subunits also participate in the construction of water-filled channels, allowing the inlet of substrate water molecules to the Mn4CaO5 cluster, as well as the release of protons from it (Bondar and Dau 2012; Vassiliev et al. 2012; Nagao et al. 2017c). A theoretical study showed that cyanobacterial PSII and green plant PSII have structurally conserved channels that pass through the different extrinsic subunits (Sakashita et al. 2017b).
Individual characteristics and functions
Numerous studies have focused on the structural and functional properties of individual extrinsic subunits. The data have been extensively summarized in previous reviews (Seidler 1996; Bricker and Burnap 2005; Roose et al. 2007b, 2016; Enami et al. 2008; Ifuku et al. 2008, 2011; Fagerlund and Eaton-Rye 2011; Bricker et al. 2012; Ifuku 2015; Ifuku and Noguchi 2016; Sasi et al. 2018; Ifuku and Nagao 2021). In this section, therefore, the characteristics and functions of each extrinsic subunit are described briefly.
PsbO, also designated OEC33, is the extrinsic subunit essential for the high oxygen-evolving activity of PSII in all oxygenic photosynthetic organisms. This subunit can solely bind to the PSII intrinsic proteins, and has a critical role in stabilizing the Mn4CaO5 cluster (Metz et al. 1980; Ghanotakis et al. 1984a; Miyao and Murata 1984; Kuwabara et al. 1985). FTIR studies have shown the requirement of PsbO for proper conformational change of the OEC upon the S1-to-S2 transition (Nagao et al. 2015). PsbO also participates in the Cl-1 channel (Barber et al. 2004; De Las Rivas and Barber 2004; Ferreira et al. 2004; Ishikita et al. 2006; Ifuku and Nagao 2021), and partially in the O4 channel (Takaoka et al. 2016).
PsbV is a cytochrome (Cyt), designated as Cyt c550 (Shen et al. 1995b), but it has not been clarified whether it has redox functions in PSII (Guerrero et al. 2014; Khorobrykh et al. 2018). Cyanobacterial ΔPsbV mutant strains lacking PsbV were unable to grow photoautotrophically in the absence of Cl− or Ca2+ (Shen et al. 1998; Katoh et al. 2001; Kirilovsky et al. 2004). A major function of PsbV is to critically support the retention of Cl− and Ca2+ within the OEC, and this function is conserved among the PsbV proteins of cyanobacteria, red algae, and diatoms (Shen and Inoue 1993; Enami et al. 1998; Shen et al. 1998; Nagao et al. 2010a). Although cyanobacterial and red algal PsbV have differences in their binding properties (Enami et al. 2003) and their detailed effects on the conformational change of the OEC, FTIR studies have shown that they both have a role in arranging the structure of the OEC together with PsbO (Uno et al. 2013; Nagao et al. 2015). It has been suggested that PsbV is involved in the O1 channel and the YZ-N298 channel (Umena et al. 2011; Nagao et al. 2017c; Xiao et al. 2020; Ifuku and Nagao 2021).
PsbU is an extrinsic subunit with functions similar to those of PsbV; its binding is important for the retention of Cl− and Ca2+ in cyanobacterial, red algal, and diatom PSII (Shen and Inoue 1993; Shen et al. 1997; Enami et al. 1998; Inoue-Kashino et al. 2005; Nagao et al. 2010a). Cyanobacterial ΔPsbU mutant strains lacking PsbU showed similar photoautotrophic growth rates as WT strains when grown in normal BG-11 growth medium. However, growth was reported to be partially impaired when Cl− was depleted, and even more so when both Cl− and Ca2+ were depleted (Shen et al. 1997; Inoue-Kashino et al. 2005). Additionally, FTIR studies have shown that the binding of PsbU together with PsbO and PsbV contributes to the proper OEC structure in cyanobacterial and red algal PSII (Uno et al. 2013; Nagao et al. 2015). PsbU is also involved in the O4 channel (Saito et al. 2015; Takaoka et al. 2016; Ifuku and Nagao 2021).
PsbP, also designated OEC23 or OEC24, is an extrinsic subunit unique to green plant PSII (Åkerlund et al. 1982). In land plants, RNA interference (RNAi)-mediated knockdown of PsbP led to dramatically diminished photoautotrophy, emphasizing the importance of this subunit (Ifuku et al. 2005b; Yi et al. 2007). PsbP, which spatially replaces PsbV and PsbU in the other types of PSII (Wei et al. 2016; Ifuku and Nagao 2021) (Fig. 3), has functional roles similar to those of PsbV/PsbU. Binding of PsbP crucially supports Cl− and Ca2+ retention (Andersson et al. 1984; Ghanotakis et al. 1984b; Miyao and Murata 1985, 1986; Homann 1987), and many different mutations in PsbP lead to a decreased Cl− retaining ability (Miyao et al. 1988; Ifuku and Sato 2002; Ifuku et al. 2005a; Kakiuchi et al. 2012; Nishimura et al. 2014). FTIR studies have suggested that PsbP is essential for inducing proper conformational changes in the OEC (Tomita et al. 2009), and mutations in PsbP can perturb these conformational changes (Tomita et al. 2009; Ido et al. 2012; Kakiuchi et al. 2012; Nishimura et al. 2014). The FTIR study by Kondo and Noguchi (2018) further indicated that PsbP affects the conformational structures around the Cl− ion(s). PsbP is also suggested to be involved in the O1 and O4 channels of green plant PSII (Sakashita et al. 2017b). In addition to these functions that resemble those of PsbV and PsbU, the N-terminal region of PsbP has been shown to interact with the α-subunit of Cyt b559 (PsbE) (Ido et al. 2012, 2014; Su et al. 2017) and modulate the redox potential of this cytochrome (Nishimura et al. 2016). Cyt b559 is indicated to be involved in an alternative electron flow (cyclic electron flow) within PSII, suppressing photoinhibition by accepting electrons from the reducing side of PSII and donating electrons to P680+ (Radmer and Kok 1975; Shinopoulos and Brudvig 2012; Chu and Chiu 2016; Takagi et al. 2019). Therefore, PsbP is likely to have a role in fine-tuning the balance between the oxidation of water and the reduction of plastoquinone by PSII (Ifuku and Nagao 2021). The functional regions of PsbP will be further described and discussed in a later section (“Mechanisms of Cl− retention by PsbP”) of this review.
PsbQ, also designated OEC16, OEC17, or OEC18, is the other extrinsic subunit unique to green plant PSII (Åkerlund et al. 1982). In contrast to PsbP-RNAi plants, PsbQ-RNAi plants exhibited no apparent phenotypes when grown under normal light conditions (Ifuku et al. 2005b; Yi et al. 2009). However, PsbQ-RNAi plants showed impaired autotrophy when grown under low light conditions (Yi et al. 2006). In vitro, the binding of PsbQ together with PsbP supports the oxygen-evolving activity of PSII under very low Cl− conditions (Akabori et al. 1984; Imaoka et al. 1984; Miyao and Murata 1985). FTIR analyses have shown that PsbQ has little influence on the conformational changes of OEC when WT-PsbP is bound to PSII, while it induced a proper OEC conformation that had been perturbed by mutations in PsbP (Kakiuchi et al. 2012). An auxiliary role of PsbQ in supporting PsbP binding has therefore been suggested, and the cryo-EM structure of green plant PSII supports this possibility (Wei et al. 2016). PsbQ also partially participates in the O1 channel of green plant PSII (Sakashita et al. 2017b). The domains of PsbQ will be further described in a later section (“Mechanisms of Cl− retention by PsbQ”).
PsbQ´, found in red algal and diatom PSII (Enami et al. 1995, 1998; Nagao et al. 2007), is an extrinsic subunit that has relatively low sequence similarity with PsbQ (e.g., 18.8% (40.9%) sequence identity (similarity) between PsbQ of Spinacia oleracea and PsbQ´ of the red alga, Cyanidium caldarium) (Ohta et al. 2003; Nagao et al. 2013). Green algal PsbQ is more closely related to PsbQ´ of red algae and diatoms than PsbQ of land plants; therefore, it can also be classified as PsbQ´ (Yabuta et al. 2010; Ishikawa et al. 2020). While land plant PsbQ requires PsbO and PsbP for functional binding to PSII (Miyao and Murata 1989), PsbQ´ can directly bind to PSII (Enami et al. 1998; Suzuki et al. 2003; Nagao et al. 2010a). FTIR analyses of red algal PSII suggested that binding of PsbQ´ does not directly lead to a proper conformational change in the OEC upon the S1-to-S2 transition (Uno et al. 2013). However, PsbQ´ participates in supporting the effective binding of PsbV and PsbU to PSII (Enami et al. 1998; Nagao et al. 2010a).
Psb31 is the extrinsic subunit unique to diatom PSII (Nagao et al. 2007; Okumura et al. 2008). This subunit is structurally similar to PsbQ and PsbQ´ (Nagao et al. 2013), but its binding site is different from those of PsbQ and PsbQ´ (Nagao et al. 2017b; Pi et al. 2019) (Fig. 3). Reconstitution of Psb31 to PSII with no other extrinsic subunits, resulted in partial recovery of oxygen-evolving activity, suggesting that Psb31 has unique functions (Nagao et al. 2010a). It had been proposed that the flexible C-terminal region of Psb31 extends towards the OEC to affect the oxygen-evolving reaction (Nagao et al. 2017a, b), and a recently determined cryo-EM structure of diatom PSII confirmed that this C-terminal region reaches toward the OEC and contacts the YZ-N298 channel (Pi et al. 2019).
CyanoP and CyanoQ are lipoproteins found in cyanobacteria (Kashino et al. 2002; De Las Rivas et al. 2004; Thornton et al. 2004; De Las Rivas and Roman 2005). CyanoP has been suggested to be the ancestral homolog of PsbP (Ishihara et al. 2007), while CyanoQ has been suggested to be that of PsbQ and PsbQ´ (De Las Rivas and Roman 2005). Until recently, neither CyanoP nor CyanoQ had been observed in any of the crystal structures of cyanobacterial PSII. However, a recent cryo-EM structure of PSII from the mesophilic cyanobacterium Synechocystis sp. PCC 6803 revealed the binding of CyanoQ to PSII at a similar location as PsbQ or PsbQ´ in PSII of other species (Gisriel et al. 2022). CyanoQ had been indicated to bind to PSII and optimize the oxygen-evolving activity, stabilizing the binding of PsbV (Kashino et al. 2006; Roose et al. 2007a). In contrast, CyanoP, which has not been found in cyanobacterial PSII structures, has been suggested to be an assembly factor of PSII (Cormann et al. 2014; Knoppová et al. 2016).
In terms of the detailed mechanisms of Cl− retention by extrinsic subunits, this has been extensively studied through site-directed mutagenesis of PsbP of green plant PSII. However, most of these investigations were performed before the cryo-EM structures of green plant PSII were revealed. Thus, in the following sections, the functional regions of the Cl− retention-related green plant PSII extrinsic subunits, PsbP and PsbQ, are discussed and compared with the extrinsic subunits in the other types of PSII, based on a combination of the results of previous studies and observations from the recently reported PSII structures.
Mechanisms of Cl− retention by PsbP
As previously mentioned, in green plant PSII, PsbP is critical for Cl− retention, contributing to the proper conformational structures around the Cl− ion(s). However, which regions of PsbP are important for this function remains unclear. Release-reconstitution experiments with various mutated PsbP proteins have been performed to determine the functional regions for Cl− retention by PsbP. Additionally, although the resolutions were not high enough to determine side-chain conformations, the recently revealed green plant PSII cryo-EM structures enable further discussions on the regions important for PsbP functions. This section discusses the roles of two candidate functional regions, the well-studied N-terminal region, and a most likely important loop named Loop 4, with a particular focus on the retention of Cl− (Fig. 4).
N-terminal region of PsbP
The N-terminal region is the most intensively studied site within PsbP. This region consists of an N-terminal tail (Ala1–(Glu/Asp)17) and a subsequent β-hairpin structure located near the C-terminal helix (Fig. 4). Various deletions of nine to nineteen N-terminal residues of PsbP, decreased the oxygen-evolving activity of PSII and the binding affinity of PsbP to PSII (Miyao et al. 1988; Ifuku et al. 2005a, 2008; Kakiuchi et al. 2012). Even when amounts similar to WT-PsbP were bound to PSII, PsbP proteins depleted of fifteen and nineteen N-terminal residues (Δ15- and Δ19-PsbP) showed a remarkably decreased ability to retain Cl− and Ca2+. FTIR analysis of PSII reconstituted with Δ15-PsbP revealed that the N-terminal residues were required for proper conformational changes around the Mn4CaO5 cluster (Tomita et al. 2009). On the other hand, mutation N15D decreased the oxygen-evolving activity, but did not have an apparent effect on the binding affinity of PsbP (Ifuku and Sato 2001). Furthermore, mutations K11A, K13A, and P20C had minor effects on the oxygen-evolving activity and PsbP binding affinity (Nishimura et al. 2014; Asada et al. 2018). Considering the cryo-EM structure (Su et al. 2017), residues in the Ala1–Phe9 region seem to be capable of interacting with PsbE, while residues in the Lys13–Thr16 and (Ala/Ser)32–Asn35 regions could contact a loop region of D1.
The above observations suggest that multiple interactions between the N-terminal region of PsbP and the membrane-intrinsic subunits are necessary for the functional binding of PsbP, while several particular residues could influence Cl− retention and the oxygen-evolving activity of PSII. A D1-loop region found close to the N-terminal region of PsbP is located at the N-terminal side of a lumenal helical structure of D1. Adjacent to the C-terminal side of this helix is the Cl-1 binding site (Fig. 5). Therefore, the interactions between the N-terminal region of PsbP and this loop of D1 could affect the binding of Cl-1. Kondo and Noguchi (2018) also pointed out that these interactions, as well as the possible interactions of PsbP-Asn52 and PsbP-Phe53 with the D1 lumenal helical structure, could affect the Cl-1 binding site via this helical region.
Regions in PsbP nearby the lumenal helical structure of D1 (PDB ID: 5XNL). Cl− ions are shown as pink spheres, and D1 and PsbP are shown in green and orange cartoon views, respectively. Several PsbP residues that could interact with the D1 lumenal helix are shown as stick models, and regions Lys13–Thr16, Ala32–Asn35, and Asn52–Phe53 of PsbP, and the region including the C-terminal lumenal helix of D1 are highlighted
The recently reported structures of diatom PSII show that the N-terminal region of PsbQ´ binds along the interface of PsbV and membrane-intrinsic subunits, overlapping with the N-terminal region of PsbP in green plant PSII (Pi et al. 2019) (Fig. 6). It is possible that the N-terminal regions of PsbP and PsbQ´ have similar functions. This could partially explain why, unlike PsbQ (Miyao and Murata 1989), PsbQ´ can solely bind to PSII (Enami et al. 1998; Suzuki et al. 2003; Nagao et al. 2010a), and why both PsbQ´ and PsbO are required for full binding of PsbV (Enami et al. 1998; Nagao et al. 2010a), whereas PsbP only requires PsbO (Miyao and Murata 1989). In addition, although irrelevant to Cl− retention, it is noteworthy that both PsbP and PsbQ´ affect the redox properties of QA, even though these extrinsic subunits and QA bind to the other sides of the thylakoid membrane (Ono and Inoue 1986; Yi et al. 2007; Ido et al. 2009; Roose et al. 2010; Semin et al. 2018; Yamada et al. 2018; Kato and Noguchi 2021). This effect on QA could be related to the photoprotection of PSII (Kato and Noguchi 2022). The mechanisms of such transmembrane effects have not been clarified, but could be related to the interactions of the N-terminal regions of PsbP and PsbQ´ with membrane-intrinsic subunits, such as PsbE, and possibly PsbJ. In fact, PsbJ has been suggested to affect the redox potentials of QA and/or QB (Regel et al. 2001; Ohad et al. 2004; Boussac et al. 2021; Shen et al. 2021). Furthermore, as mentioned earlier, the interaction of the N-terminal region of PsbP with PsbE (the α-subunit of Cyt b559) led to a transmembrane effect regulating the redox potential of Cyt b559 (Nishimura et al. 2016). Further studies are required to confirm whether the N-terminal regions of PsbP and PsbQ´ have common functions.
Locations of the N-terminal region of PsbP in green plant PSII (PDB ID: 5XNL) and the N-terminal region of PsbQ´ in diatom PSII (PDB ID: 6JLU). Location of a PsbP in green plant PSII and b PsbQ´ in diatom PSII. c Superposition of a and b with the N-terminal regions of PsbP/PsbQ´ enlarged. PsbE, PsbJ, PsbP, and PsbQ´ are shown in blue, green, orange, and red cartoon views, respectively
A study on the N-terminus-depleted PsbP proteins, Δ15- and Δ19-PsbP, also revealed the role of the N-terminal region in the tight folding of PsbP (Ifuku et al. 2005a). Compared to Δ15-PsbP, Δ19-PsbP proteins required lower temperatures for expression in Escherichia coli in a soluble form, and were easily digested by trypsin treatment. These findings suggest that residues 16 to 19 (Thr16–Met19 in spinach PsbP) may be important for the structural integrity of the PsbP protein. PSII cryo-EM structures (Wei et al. 2016; Su et al. 2017) and PsbP crystal structures (Ifuku et al. 2004; Kopecky Jr et al. 2012; Cao et al. 2015) show that of the above four residues, (Asp/Glu)17 and Phe18 could interact with the C-terminal helix (Gly171–Phe183) of PsbP. In the tobacco PsbP crystal structure (Ifuku et al. 2004), the carboxyl group of Glu17 is located 2.7 Å from the amino group of the conserved Lys174, and the side chain of Phe18 is located less than 4 Å from multiple residues (including the conserved Lys174 and Phe175) of the C-terminal helix (Fig. 7a). It is possible that these interactions are important for maintaining the overall structure of PsbP. In fact, the C-terminal helix of PsbP has been indicated to be critical for the overall structure of the protein. Depletion of the C-terminal helix of PsbP led to a drastic conformational change, hindering the import of PsbP to the thylakoid lumen, and stimulating its degradation (Roffey and Theg 1996). Interestingly, the crystal structure of DcrB, a PsbP-like-fold-containing lipoprotein found in Enterobacteriaceae, with a truncated N-terminal region revealed the formation of an N-terminal domain swapped crystallographic homodimer (Rasmussen et al. 2018). The swapped domain was the N-terminal β-hairpin, which should interact with the C-terminal helix if it were within full-length PsbP (Fig. 7b). This indicates the potential flexibility of the N-terminal region of PsbP, and supports the possibility that perturbation of interactions between the N- and C-terminal regions destabilizes the conformation of PsbP. The cooperation of the N- and C-terminal regions of PsbP in supporting the oxygen-evolving activity of PSII (Ido et al. 2012) may also be related to this structural linkage between them.
a N-terminal β-hairpin structure (beige) and C-terminal α-helix (blue) of PsbP (PDB ID: 1V2B). Several residues that could have a role in the interaction between the N- and C-terminal regions are shown as stick models. b Comparison of the structure of PsbP (upper molecule; PDB ID: 5XNL) and DcrB (lower molecule; PDB ID: 6E8A). PsbP and DcrB are shown in rainbow-colored cartoon view with the N-terminus in blue and the C-terminus in red
Loop 4 region of PsbP
The Loop 4 region of PsbP (consisting of residues Thr135–Gly142) is found near the Cl-2 binding site, close to the C-terminal region of D2 and the C-terminal loop of D1. The distances between the polypeptide backbones (alpha carbons) of PsbP-Loop 4 and the C-terminal regions of D1 and D2 are, at the shortest, approximately 5 Å and 4 Å, respectively (i.e., 4.7 Å for PsbP-Glu140 – D1-Pro340, and 4.0 Å for PsbP-Asp139 – D2-Ala352). Although the side-chain conformations are not clear with the resolutions of the current green plant PSII cryo-EM structures, it is likely that these regions interact, or at least affect the conformations of each other, as has been proposed previously (Kondo and Noguchi 2018). Considering that Cl-2 interacts with the polypeptide backbones of D1-Asn338 and D1-Phe339, which are residues in the C-terminal region of D1, the PsbP-Loop 4 would almost directly affect the Cl-2 binding site. Therefore, PsbP-Loop 4 seems to be particularly important for Cl− retention.
In cyanobacterial, red algal, and diatom PSII, the position of PsbP-Loop 4 is occupied by the C-terminus of PsbU (Fig. 8). Interestingly, previous studies using red algal PSII have indicated that this C-terminal region of PsbU is crucial in Cl− retention (Okumura et al. 2001, 2007; Sano et al. 2008). These authors showed that the aromatic ring of the conserved tyrosine residue PsbU-Tyr92 (corresponding to PsbU-Tyr103 in cyanobacterial PSII) is important for optimizing Cl− availability. The authors predicted that this attribute was related to the interaction of this conserved tyrosine residue with D1-Pro340, which was later revealed to be located near Cl-2. Taking this into consideration, PsbP-Loop 4 may have both spatially and functionally replaced the C-terminal region of PsbU.
Comparison of the structure around a PsbP-Loop 4 of green plant PSII (PDB ID: 5XNL) and b PsbU C-terminus of cyanobacterial PSII (PDB ID: 3WU2). c Superposition of (a) and (b). D1, D2, CP47, PsbP, and PsbU are shown in green, blue, pink, beige, and yellow cartoon views, respectively. The Cl− ions are shown as pink spheres. CP47-Arg384, PsbU-Tyr103, and PsbP-Asp139 (found in green plant PSII at the location corresponding to the cyanobacterial PsbU-Tyr103) are shown as stick models
Near PsbP-Loop 4 and the C-terminal region of D2, another conserved residue, CP47-Arg384, was found. The CP47-R384G mutant strain of cyanobacteria showed phenotypes, such as decreased oxygen-evolving activity, increased S2 lifetime, and increased susceptibility to photoinhibition, although its growth rate was similar to that of the control strain (Putnam-Evans et al. 1996). The effects of this mutation and relevant mutations (e.g., R384G/R385G, R384E/R385E, and Δ(R384-V392)) have been summarized (Bricker and Frankel 2002; Eaton-Rye and Putnam-Evans 2005; Morris et al. 2016). A recent study showed that the conformation of the loop region formed by CP47-Arg384 and CP47-Arg385 was changed by the binding of an assembly factor instead of extrinsic proteins (Huang et al. 2021). It is possible that the observed effects of the CP47-R384G mutation in cyanobacteria were, to some extent, due to its effect on the conformation of the C-terminal regions of PsbU and D2.
Mechanisms of Cl− retention by PsbQ
In vitro, PsbQ supports the oxygen-evolving activity of green plant PSII under very low Cl− conditions. However, in contrast to PsbP, PsbQ is located distant from the reaction center of PSII, and does not interact with regions near the Cl− binding sites of PSII. Additionally, FTIR studies have shown that PsbQ has little influence on the OEC structure, at least during the S1-to-S2 transition. Therefore, it remains unclear how PsbQ affects the Cl− dependence of the oxygen-evolving activity.
Studies on the structure of PsbQ have shown that PsbQ can be divided into two regions: the core region folded as a four-helix bundle structure and the flexible N-terminal region (Balsera et al. 2003a, b, 2005; Calderone et al. 2003; Rathner et al. 2015) (Fig. 9). This is very similar to the structure of PsbQ´ (Nagao et al. 2019). The possible roles of each region in supporting Cl− retention are discussed below.
Four-helix bundle core of PsbQ
The four-helix bundle region of PsbQ binds to the lumenal surface of CP43, placing the lumenal domain of CP43 between PsbQ and the OEC (Wei et al. 2016; Su et al. 2017; Sheng et al. 2019) (Fig. 9). PsbQ´, the PsbQ homolog found in red algae and diatoms, and CyanoQ, the PsbQ homolog found in cyanobacteria, have four-helix bundle cores similar to that of PsbQ, which bind to PSII at the same location as the PsbQ core (Ago et al. 2016; Nagao et al. 2019; Pi et al. 2019; Gisriel et al. 2022) (Fig. 3). Interestingly, Psb27, an assembly factor involved in the biogenesis and repair of PSII (Mabbitt et al. 2014), also binds to this location with its four-helix bundle structure (Huang et al. 2021; Zabret et al. 2021; Gisriel and Brudvig 2022). Psb27 is associated with the lumenal domain (Loop E) of CP43 (Liu et al. 2011, 2013; Komenda et al. 2012; Huang et al. 2021; Zabret et al. 2021), and it has been suggested that this interaction may stabilize the position of CP43-Loop E (Avramov et al. 2020). As previously mentioned, the lumenal domain (Loop E) of CP43 is important for the OEC structure. Therefore, it is possible that the interaction of the PsbQ core or the PsbQ´ core with CP43-Loop E has some influence on the structure of the OEC. However, FTIR studies have shown that, at least during the S1-to-S2 transition, both PsbQ in green plant PSII and PsbQ´ in red algal PSII have little effect on the structure of the OEC (Tomita et al. 2009; Uno et al. 2013). Additionally, depletion of PsbQ in green plant PSII does not affect the oxygen-evolving activity under Cl− sufficient conditions (Miyao and Murata 1985; Homann 1988b). Moreover, binding of only PsbQ to green algal PSII, either with no extrinsic subunits or with only PsbO bound, does not recover the oxygen-evolving activity (Suzuki et al. 2003). Thus, the effect of the interaction between PsbQ and CP43-Loop E is more likely to be insignificant. Alternatively, the four-helix bundled core may be important for the binding of PsbQ to PSII (Meades et al. 2005), and bringing the N-terminal region to the right location.
N-terminal region of PsbQ
The N-terminal region of PsbQ binds along the lumenal surface of PsbP. The N-terminal end of PsbQ is located near the PsbP–CP47 interface and extends towards the PsbP–CP43 interface (Wei et al. 2016; Sheng et al. 2019) (Fig. 9). In land plant PsbQ, the N-terminal region contains a hydrophobic part rich in proline and glycine residues (Gly8–Gly19 in spinach PsbQ) (Balsera et al. 2003b) (Fig. 10). A prolyl endoproteinase specifically cleaves the N-terminal region of land plant PsbQ at the C-terminal side of Pro12 (Kuwabara et al. 1986; Kuwabara 1992; Kuwabara and Suzuki 1994). Release-reconstitution experiments were performed with the obtained PsbQ protein lacking the first twelve residues of the N-terminal region (Kuwabara et al. 1986). The N-terminal truncated PsbQ protein bound poorly to PSII and had lost its ability to support the oxygen-evolving activity at low Cl− concentrations (Kuwabara et al. 1986). These findings illustrate the importance of the N-terminal region of PsbQ for the binding and function of this subunit. The first twelve residues of the N-terminal region of PsbQ include the proline-rich region (Pro9–Pro12 in spinach PsbQ). It has been suggested by a high-resolution PsbQ crystal structure that this region forms a polyproline type II (PPII) structure (Balsera et al. 2005), which could play a role in protein–protein interactions (Adzhubei et al. 2013; Narwani et al. 2017). However, this structure was not confirmed by nuclear magnetic resonance (NMR) spectroscopy (Rathner et al. 2015), and the green plant PSII cryo-EM structure does not have sufficient resolution needed for confirmation (Wei et al. 2016).
Amino acid sequence alignment of PsbQ from land plants and green algae. Amino acids matching the consensus sequence are shown in blue (land plants) or gray (green algae) backgrounds. The asterisks (*), colons (:), and dots (.) indicate identical, conserved, and semi-conserved amino acids, respectively, and are colored blue for within only land plants, gray for within only green algae, and white with a black background for both land plants and green algae. The box with black solid lines indicates (Gln/Glu)25–Asp28 (numbered based on spinach PsbQ), the region well conserved among PsbQ of land plants and green algae, and the box with black dotted lines indicates the proline- and glycine-rich regions of land plant PsbQ
As described previously, the N-terminal region of PsbP is important for its function and structure, as well as for binding to PSII. Δ15-PsbP, which is depleted of fifteen residues at the N-terminal end, does not functionally bind to PSII (Ifuku et al. 2005a; Tomita et al. 2009; Kakiuchi et al. 2012). However, PsbQ can restore the functional binding of Δ15-PsbP to PSII (Kakiuchi et al. 2012). In the presence of PsbQ, Δ15-PsbP showed a binding ability almost identical to that of WT-PsbP, and its ability to stimulate the oxygen-evolving activity was partially recovered. Similarly, while the H144A mutation of PsbP resulted in reduced oxygen-evolving activity and Cl− retention ability (Ido et al. 2012), these functional defects of H144A-PsbP were compensated for by the binding of PsbQ (Kakiuchi et al. 2012). FTIR analysis confirmed that the binding of PsbQ supports the correct conformational change of the OEC (Kakiuchi et al. 2012) which was perturbed in PSII reconstituted with only Δ15-PsbP and H144A-PsbP (Tomita et al. 2009; Ido et al. 2012; Kakiuchi et al. 2012). These results indicate that PsbQ helps the binding of PsbP to PSII and supports the functional conformation of PsbP. The N-terminal region of PsbQ is closely associated with PsbP (Wei et al. 2016; Sheng et al. 2019). For example, the residues at the N-terminal end (Glu1–Pro9 in spinach PsbQ) are associated with a long flexible loop of PsbP (Wei et al. 2016), and some conserved residues such as (Gln/Glu)25 and Asp28 are associated with the C-terminal region of PsbP. Interactions between PsbP and the N-terminal region of PsbQ are likely to support the binding of mutated PsbP proteins to PSII. Moreover, interactions between the C-terminal helix of PsbP and the N-terminal region of PsbQ could be important in maintaining the proper linkage between the N- and C-terminal regions of PsbP, which was previously discussed as being critical for the overall conformation of PsbP.
The exact mechanism by which PsbQ supports the oxygen-evolving activity of WT PSII under low Cl− conditions remains to be clarified. One conceivable hypothesis, based on the above discussion, is that the Cl− retention ability of PsbP can be maximized by the N-terminal region of PsbQ supporting the binding and optimal conformation of PsbP. This effect of PsbQ could only be observed under low Cl− conditions, possibly because the Cl− retention ability of PsbP on its own is high enough for Cl− retention under sufficient Cl− conditions. Although green algal PsbQ does not contain the previously mentioned proline-rich hydrophobic region within its N-terminal region (Balsera et al. 2003b) (Fig. 10), it interacts with PsbP at locations very similar to that of land plant PsbQ (Sheng et al. 2019). Therefore, it is possible that the green algal PsbQ supports PsbP in a similar way as in land plant PSII.
In addition to the above hypothesis, another possible mechanism of the function of PsbQ can be proposed. In this mechanism, PsbQ regulates Cl− accessibility through its effect on water channels within PSII. Sakashita et al. (2017b) showed that several water channels, including the O1 channel, are structurally conserved between thermophilic cyanobacterial PSII and green plant PSII. PsbQ-Asp28 (one of the few residues in the N-terminal region that is well conserved among land plant and green algal PsbQ (Balsera et al. 2003a, 2005) (Fig. 10)) and PsbP residues around the C-terminal helix of PsbP have been suggested to be located at the lumenal end of the O1-PsbP channel in green plant PSII (Sakashita et al. 2017b). In a different report, the water molecules in the Cl-2 cavity were suggested to originate from the O1 channel (Sakashita et al. 2017a). Therefore, the O1 channel is the water channel that may connect the Cl-2 binding site to the bulk solution, at least in thermophilic cyanobacterial PSII and green plant PSII. Thus, considering that PsbQ supports the retention of Cl− under low Cl− conditions, the N-terminal region of PsbQ may have some effect on the structure of the O1-PsbP channel, and could affect the diffusion of Cl− ions through this channel. This proposed mechanism is not mutually exclusive with the first hypothesis, and may explain why the PsbQ N-terminal region (Glu/Gln)25–Asp28 is well conserved in both land plants and green algae, whose PSII bind PsbP (Fig. 10), but not in CyanoQ of cyanobacteria or PsbQ´ of red algae and diatoms, whose PSII bind PsbV and PsbU instead of PsbP (Balsera et al. 2005; Gisriel and Brudvig 2022).
Conclusions
As summarized in this review, much progress has been made in understanding the roles of Cl− ions within the OEC. Success in obtaining a high-resolution PSII crystal structure (Umena et al. 2011) has allowed detailed investigations on the roles of each of the two Cl− ions bound near the Mn4CaO5 cluster. The importance of Cl-1 has been well studied, and it is becoming evident that its effect on the surrounding protein structure and hydrogen-bond network influences the water-oxidizing reaction. Additionally, the PSII structures from various species have provided valuable information, providing insights into the mechanisms of Cl− retention by extrinsic subunits.
However, unanswered questions remain. As very few studies have focused on the functions of Cl-2, it is still unclear whether Cl-2 affects the oxygen-evolving activity of PSII, and if so, how. In terms of the mechanisms of Cl− retention, some predictions can be made based on previous reports and PSII structures. However, further experimental evidence and PSII structures with higher resolutions (except for cyanobacterial PSII) would be required to determine these details.
Abbreviations
- Cryo-EM:
-
Cryo-electron microscopy
- Cyt:
-
Cytochrome
- EPR:
-
Electron paramagnetic resonance
- FTIR:
-
Fourier transform infrared
- OEC:
-
Oxygen-evolving center
- PSII:
-
Photosystem II
- RNAi:
-
RNA interference
References
Adzhubei AA, Sternberg MJE, Makarov AA (2013) Polyproline-II helix in proteins: structure and function. J Mol Biol 425:2100–2132. https://doi.org/10.1016/j.jmb.2013.03.018
Ago H, Adachi H, Umena Y et al (2016) Novel features of eukaryotic photosystem II revealed by its crystal structure analysis from a red alga. J Biol Chem 291:5676–5687. https://doi.org/10.1074/jbc.M115.711689
Akabori K, Imaoka A, Toyoshima Y (1984) The role of lipids and 17-kDa protein in enhancing the recovery of O2 evolution in cholate-treated thylakoid membranes. FEBS Lett 173:36–40. https://doi.org/10.1016/0014-5793(84)81012-1
Åkerlund H-E, Jansson C (1981) Localization of a 34000 and a 23000 Mr polypeptide to the lumenal side of the thylakoid membrane. FEBS Lett 124:229–232. https://doi.org/10.1016/0014-5793(81)80143-3
Åkerlund H-E, Jansson C, Andersson B (1982) Reconstitution of photosynthetic water splitting in inside-out thylakoid vesicles and identification of a participating polypeptide. Biochim Biophys Acta Bioenergy 681:1–10. https://doi.org/10.1016/0005-2728(82)90271-7
Amin M, Pokhrel R, Brudvig GW et al (2016) Effect of chloride depletion on the magnetic properties and the redox leveling of the oxygen-evolving complex in photosystem II. J Phys Chem B 120:4243–4248. https://doi.org/10.1021/acs.jpcb.6b03545
Andersson B, Critchley C, Ryrie IJ et al (1984) Modification of the chloride requirement for photosynthetic O2 evolution: the role of the 23 kDa polypeptide. FEBS Lett 168:113–117. https://doi.org/10.1016/0014-5793(84)80217-3
Arnon DI, Whatley FR (1949) Is chloride a coenzyme of photosynthesis? Science 110:554–556. https://doi.org/10.1126/science.110.2865.554
Asada M, Nishimura T, Ifuku K, Mino H (2018) Location of the extrinsic subunit PsbP in photosystem II studied by pulsed electron-electron double resonance. Biochim Biophys Acta 1859:394–399. https://doi.org/10.1016/j.bbabio.2018.03.002
Askerka M, Wang J, Vinyard DJ et al (2016) S3 state of the O2-evolving complex of photosystem II: insights from QM/MM, EXAFS, and femtosecond X-ray diffraction. Biochemistry 55:981–984. https://doi.org/10.1021/acs.biochem.6b00041
Avramov AP, Hwang HJ, Burnap RL (2020) The role of Ca2+ and protein scaffolding in the formation of nature’s water oxidizing complex. Proc Natl Acad Sci USA 117:28036–28045. https://doi.org/10.1073/pnas.2011315117
Balsera M, Arellano JB, Gutiérrez JR et al (2003a) Structural analysis of the PsbQ protein of photosystem II by Fourier transform infrared and circular dichroic spectroscopy and by bioinformatic methods. Biochemistry 42:1000–1007. https://doi.org/10.1021/bi026575l
Balsera M, Arellano JB, Pazos F et al (2003b) The single tryptophan of the PsbQ protein of photosystem II is at the end of a 4-α-helical bundle domain. Eur J Biochem 270:3916–3927. https://doi.org/10.1046/j.1432-1033.2003.03774.x
Balsera M, Arellano JB, Revuelta JL et al (2005) The 1.49Å resolution crystal structure of PsbQ from photosystem II of Spinacia oleracea reveals a PPII structure in the N-terminal region. J Mol Biol 350:1051–1060. https://doi.org/10.1016/j.jmb.2005.05.044
Banerjee G, Ghosh I, Kim CJ et al (2018) Substitution of the D1-Asn87 site in photosystem II of cyanobacteria mimics the chloride-binding characteristics of spinach photosystem II. J Biol Chem 293:2487–2497. https://doi.org/10.1074/jbc.M117.813170
Barber J (2016) Photosystem II: the water splitting enzyme of photosynthesis and the origin of oxygen in our atmosphere. Q Rev Biophys 49:e14. https://doi.org/10.1017/s0033583516000093
Barber J, Ferreira K, Maghlaoui K, Iwata S (2004) Structural model of the oxygen-evolving centre of photosystem II with mechanistic implications. Phys Chem Chem Phys 6:4737–4742. https://doi.org/10.1039/B407981G
Bondar A-N, Dau H (2012) Extended protein/water H-bond networks in photosynthetic water oxidation. Biochim Biophys Acta 1817:1177–1190. https://doi.org/10.1016/j.bbabio.2012.03.031
Boussac A (1995) Exchange of chloride by bromide in the manganese photosystem-II complex studied by cw- and pulsed-EPR. Chem Phys 194:409–418. https://doi.org/10.1016/0301-0104(94)00419-B
Boussac A, Sellés J, Hamon M, Sugiura M (2021) Properties of photosystem II lacking the PsbJ subunit. Photosynth Res. https://doi.org/10.1007/s11120-021-00880-w
Bové JM, Bové C, Whatley FR, Arnon DI (1963) Chloride requirement for oxygen evolution in photosynthesis. Zeitschrift Für Naturforsch B 18:683–688. https://doi.org/10.1515/znb-1963-0902
Bovi D, Narzi D, Guidoni L (2013) The S2 state of the oxygen-evolving complex of photosystem II explored by QM/MM dynamics: spin surfaces and metastable states suggest a reaction path towards the S3 state. Angew Chemie Int Ed 52:11744–11749. https://doi.org/10.1002/anie.201306667
Bowes JM, Stewart AC, Bendall DS (1983) Purification of photosystem II particles from Phormidium laminosum using the detergent dodecyl-β-D-maltoside. Properties of the purified complex. Biochim Biophys Acta 725:210–219. https://doi.org/10.1016/0005-2728(83)90241-4
Bricker TM, Frankel LK (2002) The structure and function of CP47 and CP43 in photosystem II. Photosynth Res 72:131. https://doi.org/10.1023/A:1016128715865
Bricker TM, Burnap RL (2005) The extrinsic proteins of photosystem II. In: Wydrzynski TJ, Satoh K, Freeman JA (eds) Photosystem II: the light-driven water: plastoquinone oxidoreductase. Springer, Dordrecht, pp 95–120. https://doi.org/10.1007/1-4020-4254-X_6
Bricker TM, Roose JL, Fagerlund RD et al (2012) The extrinsic proteins of photosystem II. Biochim Biophys Acta 1817:121–142. https://doi.org/10.1016/j.bbabio.2011.07.006
Calderone V, Trabucco M, Vujicić A et al (2003) Crystal structure of the PsbQ protein of photosystem II from higher plants. EMBO Rep 4:900–905. https://doi.org/10.1038/sj.embor.embor923
Cao P, Xie Y, Li M et al (2015) Crystal structure analysis of extrinsic PsbP protein of photosystem II reveals a manganese-induced conformational change. Mol Plant 8:664–666. https://doi.org/10.1016/j.molp.2015.01.002
Casey JL, Sauer K (1984) EPR detection of a cryogenically photogenerated intermediate in photosynthetic oxygen evolution. Biochim Biophys Acta 767:21–28. https://doi.org/10.1016/0005-2728(84)90075-6
Chu H-A, Chiu Y-F (2016) The roles of cytochrome b559 in assembly and photoprotection of photosystem II revealed by site-directed mutagenesis studies. Front Plant Sci 6:1261. https://doi.org/10.3389/fpls.2015.01261
Cormann KU, Bartsch M, Rögner M, Nowaczyk MM (2014) Localization of the CyanoP binding site on photosystem II by surface plasmon resonance spectroscopy. Front Plant Sci 5:595. https://doi.org/10.3389/fpls.2014.00595
Critchley C (1985) The role of chloride in photosystem II. Biochim Biophys Acta 811:33–46. https://doi.org/10.1016/0304-4173(85)90004-7
Damoder R, Klimov VV, Dismukes GC (1986) The effect of Cl− depletion and X− reconstitution on the oxygen-evolution rate, the yield of the multiline managanese EPR signal and EPR Signal II in the isolated photosystem-II complex. Biochim Biophys Acta 848:378–391. https://doi.org/10.1016/0005-2728(86)90214-8
De Las Rivas J, Balsera M, Barber J (2004) Evolution of oxygenic photosynthesis: genome-wide analysis of the OEC extrinsic proteins. Trends Plant Sci 9:18–25. https://doi.org/10.1016/j.tplants.2003.11.007
De Las Rivas J, Barber J (2004) Analysis of the structure of the PsbO protein and its implications. Photosynth Res 81:329–343. https://doi.org/10.1023/B:PRES.0000036889.44048.e4
De Las Rivas J, Roman A (2005) Structure and evolution of the extrinsic proteins that stabilize the oxygen-evolving engine. Photochem Photobiol Sci 4:1003–1010. https://doi.org/10.1039/B506874F
de Lichtenberg C, Kim CJ, Chernev P et al (2021) The exchange of the fast substrate water in the S2 state of photosystem II is limited by diffusion of bulk water through channels – implications for the water oxidation mechanism. Chem Sci. https://doi.org/10.1039/D1SC02265B
De Paula JC, Li PM, Miller AF et al (1986) Effect of the 17- and 23-kilodalton polypeptides, calcium, and chloride and electron transfer in photosystem II. Biochemistry 25:6487–6494. https://doi.org/10.1021/bi00369a022
Debus RJ (1992) The manganese and calcium ions of photosynthetic oxygen evolution. Biochim Biophys Acta 1102:269–352. https://doi.org/10.1016/0005-2728(92)90133-M
Dismukes GC, Siderer Y (1981) Intermediates of a polynuclear manganese center involved in photosynthetic oxidation of water. Proc Natl Acad Sci USA 78:274–278. https://doi.org/10.1073/pnas.78.1.274
Dismukes GC, Klimov VV, Baranov SV et al (2001) The origin of atmospheric oxygen on Earth: the innovation of oxygenic photosynthesis. Proc Natl Acad Sci USA 98:2170–2175. https://doi.org/10.1073/pnas.061514798
Eaton-Rye JJ, Putnam-Evans C (2005) The CP47 and CP43 core antenna components. In: Wydrzynski TJ, Satoh K, Freeman JA (eds) Photosystem II: the light-driven water:plastoquinone oxidoreductase. Springer, Dordrecht, pp 45–70. https://doi.org/10.1007/1-4020-4254-X_4
Enami I, Murayama H, Ohta H et al (1995) Isolation and characterization of a photosystem II complex from the red alga Cyanidium caldarium: association of cytochrome c-550 and a 12 kDa protein with the complex. Biochim Biophys Acta 1232:208–216. https://doi.org/10.1016/0005-2728(95)00122-0
Enami I, Kikuchi S, Fukuda T et al (1998) Binding and functional properties of four extrinsic proteins of photosystem II from a Red Alga, Cyanidium caldarium, as studied by release−reconstitution experiments. Biochemistry 37:2787–2793. https://doi.org/10.1021/bi9724624
Enami I, Iwai M, Akiyama A et al (2003) Comparison of binding and functional properties of two extrinsic components, Cyt c550 and a 12 kDa protein, in cyanobacterial PSII with those in Red Algal PSII. Plant Cell Physiol 44:820–827. https://doi.org/10.1093/pcp/pcg106
Enami I, Suzuki T, Tada O et al (2005) Distribution of the extrinsic proteins as a potential marker for the evolution of photosynthetic oxygen-evolving photosystem II. FEBS J 272:5020–5030. https://doi.org/10.1111/j.1742-4658.2005.04912.x
Enami I, Okumura A, Nagao R et al (2008) Structures and functions of the extrinsic proteins of photosystem II from different species. Photosynth Res 98:349–363. https://doi.org/10.1007/s11120-008-9343-9
Fagerlund RD, Eaton-Rye JJ (2011) The lipoproteins of cyanobacterial photosystem II. J Photochem Photobiol B Biol 104:191–203. https://doi.org/10.1016/j.jphotobiol.2011.01.022
Ferreira KN, Iverson TM, Maghlaoui K et al (2004) Architecture of the photosynthetic oxygen-evolving center. Science 303:1831–1838. https://doi.org/10.1126/science.1093087
Franzén L-G, Hansson Ö, Andréasson L-E (1985) The roles of the extrinsic subunits in photosystem II as revealed by EPR. Biochim Biophys Acta 808:171–179. https://doi.org/10.1016/0005-2728(85)90040-4
Gabdulkhakov A, Guskov A, Broser M et al (2009) Probing the accessibility of the Mn4Ca cluster in photosystem II: channels calculation, noble gas derivatization, and cocrystallization with DMSO. Structure 17:1223–1234. https://doi.org/10.1016/j.str.2009.07.010
Ghanotakis DF, Babcock GT, Yocum CF (1984a) Structural and catalytic properties of the oxygen-evolving complex. Correlation of polypeptide and manganese release with the behavior of Z+ in chloroplasts and a highly resolved preparation of the PS II complex. Biochim Biophys Acta 765:388–398. https://doi.org/10.1016/0005-2728(84)90180-4
Ghanotakis DF, Topper JN, Babcock GT, Yocum CF (1984b) Water-soluble 17 and 23 kDa polypeptides restore oxygen evolution activity by creating a high-affinity binding site for Ca2+ on the oxidizing side of photosystem II. FEBS Lett 170:169–173. https://doi.org/10.1016/0014-5793(84)81393-9
Ghanotakis DF, Topper JN, Yocum CF (1984c) Structural organization of the oxidizing side of photosystem II. Exogenous reductants reduce and destroy the Mn-complex in photosystems II membranes depleted of the 17 and 23 kDa polypeptides. Biochim Biophys Acta 767:524–531. https://doi.org/10.1016/0005-2728(84)90051-3
Ghosh I, Khan S, Banerjee G et al (2019) Insights into proton-transfer pathways during water oxidation in photosystem II. J Phys Chem B 123:8195–8202. https://doi.org/10.1021/acs.jpcb.9b06244
Gisriel CJ, Brudvig GW (2022) Comparison of PsbQ and Psb27 in photosystem II provides insight into their roles. Photosynth Res. https://doi.org/10.1007/s11120-021-00888-2
Gisriel CJ, Wang J, Liu J et al (2022) High-resolution cryo-electron microscopy structure of photosystem II from the mesophilic cyanobacterium, Synechocystis sp. PCC 6803. Proc Natl Acad Sci USA 119:e2116765118. https://doi.org/10.1073/pnas.2116765118
Guerrero F, Zurita JL, Roncel M et al (2014) The role of the high potential form of the cytochrome b559: Study of Thermosynechococcus elongatus mutants. Biochim Biophys Acta 1837:908–919. https://doi.org/10.1016/j.bbabio.2014.02.024
Guskov A, Kern J, Gabdulkhakov A et al (2009) Cyanobacterial photosystem II at 2.9-Å resolution and the role of quinones, lipids, channels and chloride. Nat Struct Mol Biol 16:334–342. https://doi.org/10.1038/nsmb.1559
Hasegawa K, Kimura Y, Ono T (2002) Chloride cofactor in the photosynthetic oxygen-evolving complex studied by Fourier transform infrared spectroscopy. Biochemistry 41:13839–13850. https://doi.org/10.1021/bi026595n
Haumann M, Barra M, Loja P et al (2006) Bromide does not bind to the Mn4Ca complex in its S1 state in Cl−-depleted and Br−-reconstituted oxygen-evolving photosystem II: evidence from X-ray absorption spectroscopy at the Br K-edge. Biochemistry 45:13101–13107. https://doi.org/10.1021/bi061308r
Heath RL, Hind G (1969) The role of Cl− in photosynthesis II. The effect of Cl− upon fluorescence. Biochim Biophys Acta 172:290–299. https://doi.org/10.1016/0005-2728(69)90071-1
Hind G, Nakatani HY, Izawa S (1969) The role of Cl− in photosynthesis I. The Cl− requirement of electron transport. Biochim Biophys Acta 172:277–289. https://doi.org/10.1016/0005-2728(69)90070-X
Ho FM, Styring S (2008) Access channels and methanol binding site to the CaMn4 cluster in photosystem II based on solvent accessibility simulations, with implications for substrate water access. Biochim Biophys Acta 1777:140–153. https://doi.org/10.1016/j.bbabio.2007.08.009
Homann PH (1987) The relations between the chloride, calcium, and polypeptide requirements of photosynthetic water oxidation. J Bioenergy Biomembr 19:105–123. https://doi.org/10.1007/BF00762720
Homann PH (1988a) Structural effects of Cl− and other anions on the water oxidizing complex of chloroplast photosystem II. Plant Physiol 88:194–199. https://doi.org/10.1104/pp.88.1.194
Homann PH (1988b) Chloride relations of photosystem II membrane preparations depleted of, and resupplied with, their 17 and 23 kDa extrinsic polypeptides. Photosynth Res 15:205–220. https://doi.org/10.1007/BF00047353
Homann PH (2002) Chloride and calcium in photosystem II: from effects to enigma. Photosynth Res 73:169–175. https://doi.org/10.1023/A:1020486729283
Huang G, Xiao Y, Pi X et al (2021) Structural insights into a dimeric Psb27-photosystem II complex from a cyanobacterium Thermosynechococcus vulcanus Proc Natl Acad Sci USA 118:e2018053118. https://doi.org/10.1073/pnas.2018053118
Hussein R, Ibrahim M, Bhowmick A et al (2021) Structural dynamics in the water and proton channels of photosystem II during the S2 to S3 transition. Nat Commun 12:6531. https://doi.org/10.1038/s41467-021-26781-z
Ibrahim M, Fransson T, Chatterjee R et al (2020) Untangling the sequence of events during the S2 → S3 transition in photosystem II and implications for the water oxidation mechanism. Proc Natl Acad Sci USA 117:12624–12635. https://doi.org/10.1073/pnas.2000529117
Ido K, Ifuku K, Yamamoto Y et al (2009) Knockdown of the PsbP protein does not prevent assembly of the dimeric PSII core complex but impairs accumulation of photosystem II supercomplexes in tobacco. Biochim Biophys Acta 1787:873–881. https://doi.org/10.1016/j.bbabio.2009.03.004
Ido K, Kakiuchi S, Uno C et al (2012) The conserved His-144 in the PsbP protein is important for the interaction between the PsbP N-terminus and the Cyt b559 subunit of photosystem II. J Biol Chem 287:26377–26387. https://doi.org/10.1074/jbc.M112.385286
Ido K, Nield J, Fukao Y et al (2014) Cross-linking evidence for multiple interactions of the PsbP and PsbQ proteins in a higher plant photosystem II supercomplex. J Biol Chem 289:20150–20157. https://doi.org/10.1074/jbc.M114.574822
Ifuku K (2015) Localization and functional characterization of the extrinsic subunits of photosystem II: an update. Biosci Biotechnol Biochem 79:1223–1231. https://doi.org/10.1080/09168451.2015.1031078
Ifuku K, Sato F (2001) Importance of the N-terminal sequence of the extrinsic 23 kDa polypeptide in photosystem II in ion retention in oxygen evolution. Biochim Biophys Acta 1546:196–204. https://doi.org/10.1016/S0167-4838(01)00139-X
Ifuku K, Sato F (2002) A truncated mutant of the extrinsic 23-kDa protein that absolutely requires the extrinsic 17-kDa protein for Ca2+ retention in photosystem II. Plant Cell Physiol 43:1244–1249. https://doi.org/10.1093/pcp/pcf136
Ifuku K, Noguchi T (2016) Structural coupling of extrinsic proteins with the oxygen-evolving center in photosystem II. Front Plant Sci 7:84. https://doi.org/10.3389/fpls.2016.00084
Ifuku K, Nagao R (2021) Evolution and function of the extrinsic subunits of photosystem II. In: Shen J-R, Satoh K, Allakhverdiev SI (eds) Photosynthesis: molecular approaches to solar energy conversion. Springer, Cham, pp 429–446. https://doi.org/10.1007/978-3-030-67407-6_16
Ifuku K, Nakatsu T, Kato H, Sato F (2004) Crystal structure of the PsbP protein of photosystem II from Nicotiana tabacum. EMBO Rep 5:362–367. https://doi.org/10.1038/sj.embor.7400113
Ifuku K, Nakatsu T, Shimamoto R et al (2005a) Structure and function of the PsbP protein of photosystem II from higher plants. Photosynth Res 84:251–255. https://doi.org/10.1007/s11120-004-7160-3
Ifuku K, Yamamoto Y, Ono TA et al (2005b) PsbP protein, but not PsbQ protein, is essential for the regulation and stabilization of photosystem II in higher plants. Plant Physiol 139:1175–1184. https://doi.org/10.1104/pp.105.068643
Ifuku K, Ishihara S, Shimamoto R et al (2008) Structure, function, and evolution of the PsbP protein family in higher plants. Photosynth Res 98:427–437. https://doi.org/10.1007/s11120-008-9359-1
Ifuku K, Ido K, Sato F (2011) Molecular functions of PsbP and PsbQ proteins in the photosystem II supercomplex. J Photochem Photobiol B Biol 104:158–164. https://doi.org/10.1016/j.jphotobiol.2011.02.006
Imaoka A, Yanagi M, Akabori K, Toyoshima Y (1984) Reconstitution of photosynthetic charge accumulation and oxygen evolution in CaCl2-treated PS II particles. FEBS Lett 176:341–345. https://doi.org/10.1016/0014-5793(84)81193-X
Imaoka A, Akabori K, Yanagi M et al (1986) Roles of three lumen-surface proteins in the formation of S2 state and O2 evolution in photosystem II particles from spinach thylakoid membranes. Biochim Biophys Acta 848:201–211. https://doi.org/10.1016/0005-2728(86)90042-3
Inoue-Kashino N, Kashino Y, Satoh K et al (2005) PsbU provides a stable architecture for the oxygen-evolving system in cyanobacterial photosystem II. Biochemistry 44:12214–12228. https://doi.org/10.1021/bi047539k
Ishihara S, Takabayashi A, Ido K et al (2007) Distinct functions for the two PsbP-like proteins PPL1 and PPL2 in the chloroplast thylakoid lumen of Arabidopsis. Plant Physiol 145:668–679. https://doi.org/10.1104/pp.107.105866
Ishikawa N, Yokoe Y, Nishimura T et al (2020) PsbQ-like protein 3 functions as an assembly factor for the chloroplast NADH dehydrogenase-like complex in Arabidopsis. Plant Cell Physiol 61:1252–1261. https://doi.org/10.1093/pcp/pcaa050
Ishikita H, Saenger W, Loll B et al (2006) Energetics of a possible proton exit pathway for water oxidation in photosystem II. Biochemistry 45:2063–2071. https://doi.org/10.1021/bi051615h
Isobe H, Shoji M, Yamanaka S et al (2012) Theoretical illumination of water-inserted structures of the CaMn4O5 cluster in the S2 and S3 states of oxygen-evolving complex of photosystem II: full geometry optimizations by B3LYP hybrid density functional. Dalt Trans 41:13727–13740. https://doi.org/10.1039/C2DT31420G
Itoh S, Yerkes CT, Koike H et al (1984) Effects of chloride depletion on electron donation from the water-oxidizing complex to the photosystem II reaction center as measured by the microsecond rise of chlorophyll fluorescence in isolated pea chloroplasts. Biochim Biophys Acta 766:612–622. https://doi.org/10.1016/0005-2728(84)90122-1
Izawa S, Heath RL, Hind G (1969) The role of chloride ion in photosynthesis III. The effect of artificial electron donors upon electron transport. Biochim Biophys Acta 180:388–398. https://doi.org/10.1016/0005-2728(69)90123-6
Kakiuchi S, Uno C, Ido K et al (2012) The PsbQ protein stabilizes the functional binding of the PsbP protein to photosystem II in higher plants. Biochim Biophys Acta 1817:1346–1351. https://doi.org/10.1016/j.bbabio.2012.01.009
Kashino Y, Lauber WM, Carroll JA et al (2002) Proteomic analysis of a highly active photosystem II preparation from the cyanobacterium Synechocystis sp. PCC 6803 reveals the presence of novel polypeptides. Biochemistry 41:8004–8012. https://doi.org/10.1021/bi026012+
Kashino Y, Inoue-Kashino N, Roose JL, Pakrasi HB (2006) Absence of the PsbQ protein results in destabilization of the PsbV protein and decreased oxygen evolution activity in cyanobacterial photosystem II. J Biol Chem 281:20834–20841. https://doi.org/10.1074/jbc.M603188200
Kato Y, Noguchi T (2021) Effects of stromal and lumenal side perturbations on the redox potential of the primary quinone electron acceptor QA in photosystem II. Biochemistry 60:3697–3706. https://doi.org/10.1021/acs.biochem.1c00624
Kato Y, Noguchi T (2022) Redox properties and regulatory mechanism of the iron-quinone electron acceptor in photosystem II as revealed by FTIR spectroelectrochemistry. Photosynth Res. https://doi.org/10.1007/s11120-021-00894-4
Katoh H, Itoh S, Shen J-R, Ikeuchi M (2001) Functional analysis of psbV and a novel c-type cytochrome gene psbV2 of the thermophilic cyanobacterium Thermosynechococcus elongatus strain BP-1. Plant Cell Physiol 42:599–607. https://doi.org/10.1093/pcp/pce074
Kaur D, Zhang Y, Reiss KM et al (2021) Proton exit pathways surrounding the oxygen evolving complex of photosystem II. Biochim Biophys Acta 1862:148446. https://doi.org/10.1016/j.bbabio.2021.148446
Kawakami K, Umena Y, Kamiya N, Shen J-R (2009) Location of chloride and its possible functions in oxygen-evolving photosystem II revealed by X-ray crystallography. Proc Natl Acad Sci USA 106:8567–8572. https://doi.org/10.1073/pnas.0812797106
Kawashima K, Takaoka T, Kimura H et al (2018) O2 evolution and recovery of the water-oxidizing enzyme. Nat Commun 9:1247. https://doi.org/10.1038/s41467-018-03545-w
Kelley PM, Izawa S (1978) The role of chloride ion in photosystem II I. Effects of chloride ion on photosystem II electron transport and on hydroxylamine inhibition. Biochim Biophys Acta 502:198–210. https://doi.org/10.1016/0005-2728(78)90042-7
Kern J, Chatterjee R, Young ID et al (2018) Structures of the intermediates of Kok’s photosynthetic water oxidation clock. Nature 563:421–425. https://doi.org/10.1038/s41586-018-0681-2
Khorobrykh AA, Yanykin DV, Klimov VV (2018) Photooxidation and photoreduction of exogenous cytochrome c by photosystem II preparations after various modifications of the water-oxidizing complex. Photosynthetica 56:244–253. https://doi.org/10.1007/s11099-017-0762-8
Kirilovsky D, Roncel M, Boussac A et al (2004) Cytochrome c550 in the cyanobacterium Thermosynechococcus elongatus: STUDY OF REDOX MUTANTS. J Biol Chem 279:52869–52880. https://doi.org/10.1074/jbc.M408206200
Knoppová J, Yu J, Konik P et al (2016) CyanoP is involved in the early steps of photosystem II assembly in the cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol 57:1921–1931. https://doi.org/10.1093/pcp/pcw115
Kok B, Forbush B, McGloin M (1970) Cooperation of charges. Photochem Photobiol 11:457–475. https://doi.org/10.1111/j.1751-1097.1970.tb06017.x
Komenda J, Knoppová J, Kopečná J et al (2012) The Psb27 assembly factor binds to the CP43 complex of photosystem II in the cyanobacterium Synechocystis sp. PCC 6803. Plant Physiol 158:476–486. https://doi.org/10.1104/pp.111.184184
Kondo J, Noguchi T (2018) PsbP-induced protein conformational changes around Cl− ions in the water oxidizing center of photosystem II. Photosynthetica 56:178–184. https://doi.org/10.1007/s11099-017-0749-5
Kopecky V Jr, Kohoutova J, Lapkouski M et al (2012) Raman spectroscopy adds complementary detail to the high-resolution X-ray crystal structure of photosynthetic PsbP from Spinacia oleracea. PLoS ONE 7:e46694. https://doi.org/10.1371/journal.pone.0046694
Kuroda H, Kawashima K, Ueda K et al (2021) Proton transfer pathway from the oxygen-evolving complex in photosystem II substantiated by extensive mutagenesis. Biochim Biophys Acta 1862:148329. https://doi.org/10.1016/j.bbabio.2020.148329
Kuwabara T (1992) Characterization of a prolyl endopeptidase from spinach thylakoids. FEBS Lett 300:127–130. https://doi.org/10.1016/0014-5793(92)80179-K
Kuwabara T, Murata N (1982) Inactivation of photosynthetic oxygen evolution and concomitant release of three polypeptides in the photosystem II particles of spinach chloroplasts. Plant Cell Physiol 23:533–539. https://doi.org/10.1093/oxfordjournals.pcp.a076378
Kuwabara T, Suzuki K (1994) A prolyl endoproteinase that acts specifically on the extrinsic 18-kDa protein of photosystem II: purification and further characterization. Plant Cell Physiol 35:665–675. https://doi.org/10.1093/oxfordjournals.pcp.a078642
Kuwabara T, Miyao M, Murata T, Murata N (1985) The function of 33-kDa protein in the photosynthetic oxygen-evolution system studied by reconstitution experiments. Biochim Biophys Acta 806:283–289. https://doi.org/10.1016/0005-2728(85)90107-0
Kuwabara T, Murata T, Miyao M, Murata N (1986) Partial degradation of the 18-kDa protein of the photosynthetic oxygen-evolving complex: a study of a binding site. Biochim Biophys Acta 850:146–155. https://doi.org/10.1016/0005-2728(86)90019-8
Lindberg K, Andréasson L-E (1996) A one-site, two-state model for the binding of anions in photosystem II. Biochemistry 35:14259–14267. https://doi.org/10.1021/bi961244s
Lindberg K, Vänngård T, Andréasson L-E (1993) Studies of the slowly exchanging chloride in photosystem II of higher plants. Photosynth Res 38:401–408. https://doi.org/10.1007/BF00046767
Liu H, Huang RY-C, Chen J et al (2011) Psb27, a transiently associated protein, binds to the chlorophyll binding protein CP43 in photosystem II assembly intermediates. Proc Natl Acad Sci USA 108:18536–18541. https://doi.org/10.1073/pnas.1111597108
Liu H, Chen J, Huang RY-C et al (2013) Mass spectrometry-based footprinting reveals structural dynamics of loop E of the chlorophyll-binding protein CP43 during photosystem II assembly in the cyanobacterium Synechocystis 6803. J Biol Chem 288:14212–14220. https://doi.org/10.1074/jbc.M113.467613
Lubitz W, Chrysina M, Cox N (2019) Water oxidation in photosystem II. Photosynth Res 142:105–125. https://doi.org/10.1007/s11120-019-00648-3
Mabbitt PD, Wilbanks SM, Eaton-Rye JJ (2014) Structure and function of the hydrophilic photosystem II assembly proteins: Psb27, Psb28 and Ycf48. Plant Physiol Biochem 81:96–107. https://doi.org/10.1016/j.plaphy.2014.02.013
Mandal M, Saito K, Ishikita H (2022) Requirement of chloride for the downhill electron transfer pathway from the water-splitting center in natural photosynthesis. J Phys Chem B 126:123–131. https://doi.org/10.1021/acs.jpcb.1c09176
McEvoy JP, Brudvig GW (2006) Water-splitting chemistry of photosystem II. Chem Rev 106:4455–4483. https://doi.org/10.1021/cr0204294
Meades GD, McLachlan A, Sallans L et al (2005) Association of the 17-kDa extrinsic protein with photosystem II in higher plants. Biochemistry 44:15216–15221. https://doi.org/10.1021/bi051704u
Metz JG, Wong J, Bishop NI (1980) Changes in electrophoretic mobility of a chloroplast membrane polypeptide associated with the loss of the oxidizing side of photosystem II in low fluorescent mutants of Scenedesmus. FEBS Lett 114:61–66. https://doi.org/10.1016/0014-5793(80)80861-1
Miyao M, Murata N (1984) Role of the 33-kDa polypeptide in preserving Mn in the photosynthetic oxygen-evolution system and its replacement by chloride ions. FEBS Lett 170:350–354. https://doi.org/10.1016/0014-5793(84)81342-3
Miyao M, Murata N (1985) The Cl− effect on photosynthetic oxygen evolution: interaction of Cl− with 18-kDa, 24-kDa and 33-kDa proteins. FEBS Lett 180:303–308. https://doi.org/10.1016/0014-5793(85)81091-7
Miyao M, Murata N (1986) Light-dependent inactivation of photosynthetic oxygen evolution during NaCl treatment of photosystem II particles: the role of the 24-kDa protein. Photosynth Res 10:489–496. https://doi.org/10.1007/BF00118315
Miyao M, Murata N (1989) The mode of binding of three extrinsic proteins of 33 kDa, 23 kDa and 18 kDa in the photosystem II complex of spinach. Biochim Biophys Acta 977:315–321. https://doi.org/10.1016/S0005-2728(89)80086-6
Miyao M, Fujimura Y, Murata N (1988) Partial degradation of the extrinsic 23-kDa protein of the photosystem II complex of spinach. Biochim Biophys Acta 936:465–474. https://doi.org/10.1016/0005-2728(88)90024-2
Morris JN, Eaton-Rye JJ, Summerfield TC (2016) Environmental pH and the requirement for the extrinsic proteins of photosystem II in the function of cyanobacterial photosynthesis. Front Plant Sci 7:1135. https://doi.org/10.3389/fpls.2016.01135
Murray JW, Barber J (2007) Structural characteristics of channels and pathways in photosystem II including the identification of an oxygen channel. J Struct Biol 159:228–237. https://doi.org/10.1016/j.jsb.2007.01.016
Murray JW, Maghlaoui K, Kargul J et al (2008) X-ray crystallography identifies two chloride binding sites in the oxygen evolving centre of photosystem II. Energy Environ Sci 1:161–166. https://doi.org/10.1039/B810067P
Nagao R, Ishii A, Tada O et al (2007) Isolation and characterization of oxygen-evolving thylakoid membranes and photosystem II particles from a marine diatom Chaetoceros gracilis. Biochim Biophys Acta 1767:1353–1362. https://doi.org/10.1016/j.bbabio.2007.10.007
Nagao R, Kato K, Suzuki T et al (2019) Structural basis for energy harvesting and dissipation in a diatom PSII–FCPII supercomplex. Nat Plants 5:890–901. https://doi.org/10.1038/s41477-019-0477-x
Nagao R, Moriguchi A, Tomo T et al (2010a) Binding and functional properties of five extrinsic proteins in oxygen-evolving photosystem II from a marine centric diatom, Chaetoceros gracilis. J Biol Chem 285:29191–29199. https://doi.org/10.1074/jbc.M110.146092
Nagao R, Tomo T, Noguchi E et al (2010b) Purification and characterization of a stable oxygen-evolving photosystem II complex from a marine centric diatom, Chaetoceros gracilis. Biochim Biophys Acta 1797:160–166. https://doi.org/10.1016/j.bbabio.2009.09.008
Nagao R, Suga M, Niikura A et al (2013) Crystal structure of Psb31, a novel extrinsic protein of photosystem II from a marine centric diatom and implications for its binding and function. Biochemistry 52:6646–6652. https://doi.org/10.1021/bi400770d
Nagao R, Tomo T, Noguchi T (2015) Effects of extrinsic proteins on the protein conformation of the oxygen-evolving center in cyanobacterial photosystem II as revealed by Fourier transform infrared spectroscopy. Biochemistry 54:2022–2031. https://doi.org/10.1021/acs.biochem.5b00053
Nagao R, Suzuki T, Dohmae N et al (2017a) Functional role of Lys residues of Psb31 in electrostatic interactions with diatom photosystem II. FEBS Lett 591:3259–3264. https://doi.org/10.1002/1873-3468.12830
Nagao R, Suzuki T, Okumura A et al (2017b) Electrostatic interaction of positive charges on the surface of Psb31 with photosystem II in the diatom Chaetoceros gracilis. Biochim Biophys Acta 1858:779–785. https://doi.org/10.1016/j.bbabio.2017.06.002
Nagao R, Ueoka-Nakanishi H, Noguchi T (2017c) D1-Asn-298 in photosystem II is involved in a hydrogen-bond network near the redox-active tyrosine Y(Z) for proton exit during water oxidation. J Biol Chem 292:20046–20057. https://doi.org/10.1074/jbc.M117.815183
Nakamura S, Nagao R, Takahashi R, Noguchi T (2014) Fourier transform infrared detection of a polarizable proton trapped between photooxidized tyrosine YZ and a coupled histidine in photosystem II: relevance to the proton transfer mechanism of water oxidation. Biochemistry 53:3131–3144. https://doi.org/10.1021/bi500237y
Nakamura A, Kang J, Terada R et al (2019) Novel mechanism of Cl-dependent proton dislocation in photosystem II (PSII): hybrid ab initio quantum mechanics/molecular mechanics molecular dynamics simulation. J Phys Soc Japan 88:84802. https://doi.org/10.7566/JPSJ.88.084802
Narwani TJ, Santuz H, Shinada N et al (2017) Recent advances on polyproline II. Amino Acids 49:705–713. https://doi.org/10.1007/s00726-017-2385-6
Nishimura T, Uno C, Ido K et al (2014) Identification of the basic amino acid residues on the PsbP protein involved in the electrostatic interaction with photosystem II. Biochim Biophys Acta 1837:1447–1453. https://doi.org/10.1016/j.bbabio.2013.12.012
Nishimura T, Nagao R, Noguchi T et al (2016) The N-terminal sequence of the extrinsic PsbP protein modulates the redox potential of Cyt b559 in photosystem II. Sci Rep 6:21490. https://doi.org/10.1038/srep21490
Ohad I, Dal Bosco C, Herrmann RG, Meurer J (2004) Photosystem II proteins PsbL and PsbJ regulate electron flow to the plastoquinone pool. Biochemistry 43:2297–2308. https://doi.org/10.1021/bi0348260
Ohta H, Suzuki T, Ueno M et al (2003) Extrinsic proteins of photosystem II. Eur J Biochem 270:4156–4163. https://doi.org/10.1046/j.1432-1033.2003.03810.x
Okamoto Y, Shimada Y, Nagao R, Noguchi T (2021) Proton and water transfer pathways in the S2 → S3 transition of the water-oxidizing complex in photosystem II: time-resolved infrared analysis of the effects of D1–N298A mutation and NO3– substitution. J Phys Chem B 125:6864–6873. https://doi.org/10.1021/acs.jpcb.1c03386
Okumura A, Ohta H, Inoue Y, Enami I (2001) Identification of functional domains of the extrinsic 12 kDa protein in red algal PSII by limited proteolysis and directed mutagenesis. Plant Cell Physiol 42:1331–1337. https://doi.org/10.1093/pcp/pce170
Okumura A, Sano M, Suzuki T et al (2007) Aromatic structure of tyrosine-92 in the extrinsic PsbU protein of red algal photosystem II is important for its functioning. FEBS Lett 581:5255–5258. https://doi.org/10.1016/j.febslet.2007.10.015
Okumura A, Nagao R, Suzuki T et al (2008) A novel protein in photosystem II of a diatom Chaetoceros gracilis is one of the extrinsic proteins located on lumenal side and directly associates with PSII core components. Biochim Biophys Acta 1777:1545–1551. https://doi.org/10.1016/j.bbabio.2008.09.004
Olesen K, Andréasson L-E (2003) The function of the chloride ion in photosynthetic oxygen evolution. Biochemistry 42:2025–2035. https://doi.org/10.1021/bi026175y
Ono T-A, Nakayama H, Gleiter H et al (1987) Modification of the properties of S2 state in photosynthetic O2-evolving center by replacement of chloride with other anions. Arch Biochem Biophys 256:618–624. https://doi.org/10.1016/0003-9861(87)90619-9
Ono T, Inoue Y (1986) Effects of removal and reconstitution of the extrinsic 33, 24 and 16 kDa proteins on flash oxygen yield in photosystem II particles. Biochim Biophys Acta 850:380–389. https://doi.org/10.1016/0005-2728(86)90194-5
Ono T, Zimmermann JL, Inoue Y, Rutherford AW (1986) EPR evidence for a modified S-state transition in chloride-depleted photosystem II. Biochim Biophys Acta 851:193–201. https://doi.org/10.1016/0005-2728(86)90125-8
Pantazis DA (2019) The S3 state of the oxygen-evolving complex: overview of spectroscopy and XFEL crystallography with a critical evaluation of early-onset models for O-O bond formation. Inorganics 7:1–30. https://doi.org/10.3390/inorganics7040055
Pantazis DA, Ames W, Cox N et al (2012) Two interconvertible structures that explain the spectroscopic properties of the oxygen-evolving complex of photosystem II in the S2 State. Angew Chemie Int Ed 51:9935–9940. https://doi.org/10.1002/anie.201204705
Pi X, Zhao S, Wang W et al (2019) The pigment-protein network of a diatom photosystem II–light-harvesting antenna supercomplex. Science 365:4406. https://doi.org/10.1126/science.aax4406
Pokhrel R, Brudvig GW (2014) Oxygen-evolving complex of photosystem II: correlating structure with spectroscopy. Phys Chem Chem Phys 16:11812–11821. https://doi.org/10.1039/C4CP00493K
Pokhrel R, McConnell IL, Brudvig GW (2011) Chloride regulation of enzyme turnover: application to the role of chloride in photosystem II. Biochemistry 50:2725–2734. https://doi.org/10.1021/bi2000388
Pokhrel R, Service RJ, Debus RJ, Brudvig GW (2013) Mutation of lysine 317 in the D2 subunit of photosystem II alters chloride binding and proton transport. Biochemistry 52:4758–4773. https://doi.org/10.1021/bi301700u
Pokhrel R, Debus RJ, Brudvig GW (2015) Probing the effect of mutations of asparagine 181 in the D1 subunit of photosystem II. Biochemistry 54:1663–1672. https://doi.org/10.1021/bi501468h
Putnam-Evans C, Burnap R, Wu J et al (1996) Site-directed mutagenesis of the CP 47 protein of photosystem II: alteration of conserved charged residues in the domain 364E–444R. Biochemistry 35:4046–4053. https://doi.org/10.1021/bi952661s
Radmer R, Kok B (1975) Energy capture in photosynthesis: photosystem II. Annu Rev Biochem 44:409–433. https://doi.org/10.1146/annurev.bi.44.070175.002205
Rasmussen DM, Soens RW, Davie TJ et al (2018) The structure of DcrB, a lipoprotein from Salmonella enterica, reveals flexibility in the N-terminal segment of the Mog1p/PsbP-like fold. J Struct Biol 204:513–518. https://doi.org/10.1016/j.jsb.2018.10.005
Rathner P, Rathner A, Horničáková M et al (2015) Solution NMR and molecular dynamics reveal a persistent alpha helix within the dynamic region of PsbQ from photosystem II of higher plants. Proteins Struct Funct Bioinforma 83:1677–1686. https://doi.org/10.1002/prot.24853
Regel RE, Ivleva NB, Zer H et al (2001) Deregulation of electron flow within photosystem II in the absence of the PsbJ protein. J Biol Chem 276:41473–41478. https://doi.org/10.1074/jbc.M102007200
Reiss K, Morzan UN, Grigas AT, Batista VS (2019) Water network dynamics next to the oxygen-evolving complex of photosystem II. Inorganics 7:39. https://doi.org/10.3390/inorganics7030039
Retegan M, Pantazis DA (2016) Interaction of methanol with the oxygen-evolving complex: atomistic models, channel identification, species dependence, and mechanistic implications. Chem Sci 7:6463–6476. https://doi.org/10.1039/C6SC02340A
Retegan M, Krewald V, Mamedov F et al (2016) A five-coordinate Mn(IV) intermediate in biological water oxidation: spectroscopic signature and a pivot mechanism for water binding. Chem Sci 7:72–84. https://doi.org/10.1039/C5SC03124A
Rivalta I, Amin M, Luber S et al (2011) Structural-functional role of chloride in photosystem II. Biochemistry 50:6312–6315. https://doi.org/10.1021/bi200685w
Roffey RA, Theg SM (1996) Analysis of the import of carboxyl-terminal truncations of the 23-kilodalton subunit of the oxygen-evolving complex suggests that its structure is an important determinant for thylakoid transport. Plant Physiol 111:1329–1338. https://doi.org/10.1104/pp.111.4.1329
Roose JL, Kashino Y, Pakrasi HB (2007a) The PsbQ protein defines cyanobacterial photosystem II complexes with highest activity and stability. Proc Natl Acad Sci USA 104:2548–2553. https://doi.org/10.1073/pnas.0609337104
Roose JL, Wegener KM, Pakrasi HB (2007b) The extrinsic proteins of photosystem II. Photosynth Res 92:369–387. https://doi.org/10.1007/s11120-006-9117-1
Roose JL, Frankel LK, Bricker TM (2010) Documentation of significant electron transport defects on the reducing side of photosystem II upon removal of the PsbP and PsbQ extrinsic proteins. Biochemistry 49:36–41. https://doi.org/10.1021/bi9017818
Roose JL, Frankel LK, Mummadisetti MP, Bricker TM (2016) The extrinsic proteins of photosystem II: update. Planta 243:889–908. https://doi.org/10.1007/s00425-015-2462-6
Russell BP, Vinyard DJ (2021) Chloride facilitates Mn(III) formation during photoassembly of the photosystem II oxygen-evolving complex. Photosynth Res. https://doi.org/10.1007/s11120-021-00886-4
Rutherford AW (1989) Photosystem II, the water-splitting enzyme. Trends Biochem Sci 14:227–232. https://doi.org/10.1016/0968-0004(89)90032-7
Saito K, Rutherford AW, Ishikita H (2013) Mechanism of tyrosine D oxidation in photosystem II. Proc Natl Acad Sci USA 110:7690–7695. https://doi.org/10.1073/pnas.1300817110
Saito K, William Rutherford A, Ishikita H (2015) Energetics of proton release on the first oxidation step in the water-oxidizing enzyme. Nat Commun 6:8488. https://doi.org/10.1038/ncomms9488
Saito K, Mandal M, Ishikita H (2020a) Energetics of ionized water molecules in the H-bond network near the Ca2+ and Cl– binding sites in photosystem II. Biochemistry 59:3216–3224. https://doi.org/10.1021/acs.biochem.0c00177
Saito K, Nakagawa M, Ishikita H (2020b) pKa of the ligand water molecules in the oxygen-evolving Mn4CaO5 cluster in photosystem II. Commun Chem 3:89. https://doi.org/10.1038/s42004-020-00336-7
Sakashita N, Watanabe HC, Ikeda T et al (2017a) Origins of water molecules in the photosystem II crystal structure. Biochemistry 56:3049–3057. https://doi.org/10.1021/acs.biochem.7b00220
Sakashita N, Watanabe HC, Ikeda T, Ishikita H (2017b) Structurally conserved channels in cyanobacterial and plant photosystem II. Photosynth Res 133:75–85. https://doi.org/10.1007/s11120-017-0347-1
Sakashita N, Ishikita H, Saito K (2020) Rigidly hydrogen-bonded water molecules facilitate proton transfer in photosystem II. Phys Chem Chem Phys 22:15831–15841. https://doi.org/10.1039/d0cp00295j
Sano M, Okumura A, Suzuki T et al (2008) Identification of functional domains of PsbU in red algal PSII by site-directed mutagenesis. In: Allen JF, Gantt E, Golbeck JH, Osmond B (eds) Photosynthesis. Energy from the sun. Springer, Dordrecht, pp 487–490. https://doi.org/10.1007/978-1-4020-6709-9_110
Sasi S, Venkatesh J, Daneshi RF, Gururani MA (2018) Photosystem II extrinsic proteins and their putative role in abiotic stress tolerance in higher plants. Plants (basel, Switzerland) 7:100. https://doi.org/10.3390/plants7040100
Seidler A (1996) The extrinsic polypeptides of photosystem II. Biochim Biophys Acta 1277:35–60. https://doi.org/10.1016/S0005-2728(96)00102-8
Semin BK, Davletshina LN, Mamedov MD (2018) Effect of different methods of Ca2+ extraction from PSII oxygen-evolving complex on the QA− oxidation kinetics. Photosynth Res 136:83–91. https://doi.org/10.1007/s11120-017-0441-4
Service RJ, Yano J, Dilbeck PL et al (2013) Participation of glutamate-333 of the D1 polypeptide in the ligation of the Mn4CaO5 cluster in photosystem II. Biochemistry 52:8452–8464. https://doi.org/10.1021/bi401339f
Shen J-R (2015) The structure of photosystem II and the mechanism of water oxidation in photosynthesis. Annu Rev Plant Biol 66:23–48. https://doi.org/10.1146/annurev-arplant-050312-120129
Shen J-R, Burnap RL, Inoue Y (1995a) An independent role of cytochrome c-550 in cyanobacterial photosystem II as revealed by double-deletion mutagenesis of the psbO and psbV genes in Synechocystis sp. PCC 6803. Biochemistry 34:12661–12668. https://doi.org/10.1021/bi00039a023
Shen J-R, Vermaas W, Inoue Y (1995b) The role of cytochrome c-550 as studied through reverse genetics and mutant characterization in Synechocystis sp. PCC 6803. J Biol Chem 270:6901–6907. https://doi.org/10.1074/jbc.270.12.6901
Shen J-R, Ikeuchi M, Inoue Y (1992) Stoichiometric association of extrinsic cytochrome c550 and 12 kDa protein with a highly purified oxygen-evolving photosystem II core complex from Synechococcus vulcanus. FEBS Lett 301:145–149. https://doi.org/10.1016/0014-5793(92)81235-E
Shen JR, Inoue Y (1993) Binding and functional properties of two new extrinsic components, cytochrome c-550 and a 12-kDa protein, in cyanobacterial photosystem II. Biochemistry 32:1825–1832. https://doi.org/10.1021/bi00058a017
Shen J-R, Ikeuchi M, Inoue Y (1997) Analysis of the psbU gene encoding the 12-kDa extrinsic protein of photosystem II and studies on its role by deletion mutagenesis in Synechocystis sp. PCC 6803. J Biol Chem 272:17821–17826. https://doi.org/10.1074/jbc.272.28.17821
Shen J-R, Qian M, Inoue Y, Burnap RL (1998) Functional characterization of Synechocystis sp. PCC 6803 ΔpsbU and ΔpsbV mutants reveals important roles of cytochrome c-550 in cyanobacterial oxygen evolution. Biochemistry 37:1551–1558. https://doi.org/10.1021/bi971676i
Shen J-R, Nakajima Y, Akita F, Suga M (2021) Structure, electron transfer chain of photosystem II and the mechanism of water splitting. In: Shen J-R, Satoh K, Allakhverdiev SI (eds) Photosynthesis: molecular approaches to solar energy conversion. Springer, Cham, pp 3–38. https://doi.org/10.1007/978-3-030-67407-6_1
Sheng X, Watanabe A, Li A et al (2019) Structural insight into light harvesting for photosystem II in green algae. Nat Plants 5:1320–1330. https://doi.org/10.1038/s41477-019-0543-4
Shimada Y, Kitajima-Ihara T, Nagao R, Noguchi T (2020) Role of the O4 channel in photosynthetic water oxidation as revealed by Fourier transform infrared difference and time-resolved infrared analysis of the D1–S169A mutant. J Phys Chem B 124:1470–1480. https://doi.org/10.1021/acs.jpcb.9b11946
Shimizu T, Sugiura M, Noguchi T (2018) Mechanism of proton-coupled electron transfer in the S0-to-S1 transition of photosynthetic water oxidation as revealed by time-resolved infrared spectroscopy. J Phys Chem B 122:9460–9470. https://doi.org/10.1021/acs.jpcb.8b07455
Shinopoulos KE, Brudvig GW (2012) Cytochrome b559 and cyclic electron transfer within photosystem II. Biochim Biophys Acta 1817:66–75. https://doi.org/10.1016/j.bbabio.2011.08.002
Stewart AC, Ljungberg U, Åkerlund H-E, Andersson B (1985) Studies on the polypeptide composition of the cyanobacterial oxygen-evolving complex. Biochim Biophys Acta 808:353–362. https://doi.org/10.1016/0005-2728(85)90145-8
Su X, Ma J, Wei X et al (2017) Structure and assembly mechanism of plant C2S2M2-type PSII-LHCII supercomplex. Science 357:815–820. https://doi.org/10.1126/science.aan0327
Suga M, Akita F, Hirata K et al (2015) Native structure of photosystem II at 1.95 Å resolution viewed by femtosecond X-ray pulses. Nature 517:99–103. https://doi.org/10.1038/nature13991
Suga M, Akita F, Sugahara M et al (2017) Light-induced structural changes and the site of O=O bond formation in PSII caught by XFEL. Nature 543:131–135. https://doi.org/10.1038/nature21400
Suga M, Akita F, Yamashita K et al (2019) An oxyl/oxo mechanism for oxygen-oxygen coupling in PSII revealed by an X-ray free-electron laser. Science 366:334–338. https://doi.org/10.1126/science.aax6998
Suzuki T, Minagawa J, Tomo T et al (2003) Binding and functional properties of the extrinsic proteins in oxygen-evolving photosystem II particle from a green alga, Chlamydomonas reinhardtii having His-tagged CP47. Plant Cell Physiol 44:76–84. https://doi.org/10.1093/pcp/pcg010
Suzuki H, Yu J, Kobayashi T et al (2013) Functional roles of D2-Lys317 and the interacting chloride ion in the water oxidation reaction of photosystem II as revealed by Fourier transform infrared analysis. Biochemistry 52:4748–4757. https://doi.org/10.1021/bi301699h
Takagi D, Ifuku K, Nishimura T, Miyake C (2019) Antimycin A inhibits cytochrome b559-mediated cyclic electron flow within photosystem II. Photosynth Res 139:487–498. https://doi.org/10.1007/s11120-018-0519-7
Takaoka T, Sakashita N, Saito K, Ishikita H (2016) pKa of a proton-conducting water chain in photosystem II. J Phys Chem Lett 7:1925–1932. https://doi.org/10.1021/acs.jpclett.6b00656
Thornton LE, Ohkawa H, Roose JL et al (2004) Homologs of plant PsbP and PsbQ proteins are necessary for regulation of photosystem II activity in the cyanobacterium Synechocystis 6803. Plant Cell 16:2164–2175. https://doi.org/10.1105/tpc.104.023515
Tokano T, Kato Y, Sugiyama S et al (2020) Structural dynamics of a protein domain relevant to the water-oxidizing complex in photosystem II as visualized by high-speed atomic force microscopy. J Phys Chem B 124:5847–5857. https://doi.org/10.1021/acs.jpcb.0c03892
Tomita M, Ifuku K, Sato F, Noguchi T (2009) FTIR evidence that the PsbP extrinsic protein induces protein conformational changes around the oxygen-evolving Mn cluster in photosystem II. Biochemistry 48:6318–6325. https://doi.org/10.1021/bi9006308
Toyoshima Y, Akabori K, Imaoka A et al (1984) Reconstitution of photosynthetic charge accumulation and oxygen evolution in CaCl2-treated PS II particles. FEBS Lett 176:346–350. https://doi.org/10.1016/0014-5793(84)81194-1
Umena Y, Kawakami K, Shen J-R, Kamiya N (2011) Crystal structure of oxygen-evolving photosystem II at a resolution of 1.9 Å. Nature 473:55–60. https://doi.org/10.1038/nature09913
Uno C, Nagao R, Suzuki H et al (2013) Structural coupling of extrinsic proteins with the oxygen-evolving center in red algal photosystem II as revealed by light-induced FTIR difference spectroscopy. Biochemistry 52:5705–5707. https://doi.org/10.1021/bi4009787
van Gorkom HJ, Yocum CF (2005) The calcium and chloride cofactors. In: Wydrzynski TJ, Satoh K, Freeman JA (eds) Photosystem II: the light-driven water: plastoquinone oxidoreductase. Springer, Dordrecht, pp 307–327. https://doi.org/10.1007/1-4020-4254-X_14
van Vliet P, Rutherford AW (1996) Properties of the chloride-depleted oxygen-evolving complex of photosystem II studied by electron paramagnetic resonance. Biochemistry 35:1829–1839. https://doi.org/10.1021/bi9514471
Vassiliev S, Comte P, Mahboob A, Bruce D (2010) Tracking the flow of water through photosystem II using molecular dynamics and streamline tracing. Biochemistry 49:1873–1881. https://doi.org/10.1021/bi901900s
Vassiliev S, Zaraiskaya T, Bruce D (2012) Exploring the energetics of water permeation in photosystem II by multiple steered molecular dynamics simulations. Biochim Biophys Acta 1817:1671–1678. https://doi.org/10.1016/j.bbabio.2012.05.016
Vinyard DJ, Ananyev GM, Charles Dismukes G (2013) Photosystem II: the reaction center of oxygenic photosynthesis. Annu Rev Biochem 82:577–606. https://doi.org/10.1146/annurev-biochem-070511-100425
Vinyard DJ, Badshah SL, Riggio MR et al (2019) Photosystem II oxygen-evolving complex photoassembly displays an inverse H/D solvent isotope effect under chloride-limiting conditions. Proc Natl Acad Sci USA 116:18917–18922. https://doi.org/10.1073/pnas.1910231116
Vogt L, Vinyard DJ, Khan S, Brudvig GW (2015) Oxygen-evolving complex of photosystem II: an analysis of second-shell residues and hydrogen-bonding networks. Curr Opin Chem Biol 25:152–158. https://doi.org/10.1016/j.cbpa.2014.12.040
Warburg O, Lüttgens W (1944) Weitere Experimente zur Kohlensäureassimilation. Naturwissenschaften 32:301. https://doi.org/10.1007/BF01475363
Wei X, Su X, Cao P et al (2016) Structure of spinach photosystem II–LHCII supercomplex at 3.2 Å resolution. Nature 534:69–74. https://doi.org/10.1038/nature18020
Wincencjusz H, van Gorkom HJ, Yocum CF (1997) The photosynthetic oxygen evolving complex requires chloride for its redox state S2→S3 and S3→S0 transitions but not for S0→S1 or S1→S2 transitions. Biochemistry 36:3663–3670. https://doi.org/10.1021/bi9626719
Xiao Y, Zhu Q, Yang Y et al (2020) Role of PsbV-Tyr137 in photosystem II studied by site-directed mutagenesis in the thermophilic cyanobacterium Thermosynechococcus vulcanus. Photosynth Res 146:41–54. https://doi.org/10.1007/s11120-020-00753-8
Yabuta S, Ifuku K, Takabayashi A et al (2010) Three PsbQ-like proteins are required for the function of the chloroplast NAD(P)H dehydrogenase complex in Arabidopsis. Plant Cell Physiol 51:866–876. https://doi.org/10.1093/pcp/pcq060
Yachandra VK, Guiles RD, McDermott A et al (1986a) The state of manganese in the photosynthetic apparatus: 4. Structure of the manganese complex in photosystem II studied using EXAFS spectroscopy. The S1 state of the O2-evolving photosystem II complex from spinach. Biochim Biophys Acta 850:324–332. https://doi.org/10.1016/0005-2728(86)90188-X
Yachandra VK, Guiles RD, Sauer K, Klein MP (1986b) The state of manganese in the photosynthetic apparatus: 5. The chloride effect in photosynthetic oxygen evolution. Is halide coordinated to the EPR-active manganese in the O2-evolving complex? Studies of the substructure of the low-temperature multiline EPR signal. Biochim Biophys Acta 850:333–342. https://doi.org/10.1016/0005-2728(86)90189-1
Yamada M, Nagao R, Iwai M et al (2018) The PsbQ´ protein affects the redox potential of the QA in photosystem II. Photosynthetica 56:185–191. https://doi.org/10.1007/s11099-018-0778-8
Yamamoto M, Nakamura S, Noguchi T (2020) Protonation structure of the photosynthetic water oxidizing complex in the S0 state as revealed by normal mode analysis using quantum mechanics/molecular mechanics calculations. Phys Chem Chem Phys 22:24213–24225. https://doi.org/10.1039/D0CP04079G
Yamamoto Y, Doi M, Tamura N, Nishimura M (1981) Release of polypeptides from highly active O2-evolving photosystem-2 preparation by this treatment. FEBS Lett 133:265–268. https://doi.org/10.1016/0014-5793(81)80520-0
Yang KR, Lakshmi KV, Brudvig GW, Batista VS (2021) Is deprotonation of the oxygen-evolving complex of photosystem II during the S1 → S2 transition suppressed by proton quantum delocalization? J Am Chem Soc 143:8324–8332. https://doi.org/10.1021/jacs.1c00633
Yano J, Yachandra V (2014) Mn4Ca cluster in photosynthesis: where and how water is oxidized to dioxygen. Chem Rev 114:4175–4205. https://doi.org/10.1021/cr4004874
Yi X, Hargett SR, Frankel LK, Bricker TM (2006) The PsbQ protein is required in Arabidopsis for photosystem II assembly/stability and photoautotrophy under low light conditions. J Biol Chem 281:26260–26267. https://doi.org/10.1074/jbc.M603582200
Yi X, Hargett SR, Liu H et al (2007) The PsbP protein is required for photosystem II complex assembly/stability and photoautotrophy in Arabidopsis thaliana. J Biol Chem 282:24833–24841. https://doi.org/10.1074/jbc.M705011200
Yi X, Hargett SR, Frankel LK, Bricker TM (2009) The PsbP protein, but not the PsbQ protein, is required for normal thylakoid architecture in Arabidopsis thaliana. FEBS Lett 583:2142–2147. https://doi.org/10.1016/j.febslet.2009.05.048
Yocum CF (2008) The calcium and chloride requirements of the O2 evolving complex. Coord Chem Rev 252:296–305. https://doi.org/10.1016/j.ccr.2007.08.010
Zabret J, Bohn S, Schuller SK et al (2021) Structural insights into photosystem II assembly. Nat Plants 7:524–538. https://doi.org/10.1038/s41477-021-00895-0
Acknowledgements
This research was supported by JSPS KAKENHI (Grant No. JP20H031160) to KeI and Akira Yoshino Research Grant of the Chemical Society of Japan to KeI.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Imaizumi, K., Ifuku, K. Binding and functions of the two chloride ions in the oxygen-evolving center of photosystem II. Photosynth Res 153, 135–156 (2022). https://doi.org/10.1007/s11120-022-00921-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11120-022-00921-y