Abstract
About half of the globular proteins are composed of regular secondary structures, α-helices, and β-sheets, while the rest are constituted of irregular secondary structures, such as turns or coil conformations. Other regular secondary structures are often ignored, despite their importance in biological processes. Among such structures, the polyproline II helix (PPII) has interesting behaviours. PPIIs are not usually associated with conventional stabilizing interactions, and recent studies have observed that PPIIs are more frequent than anticipated. In addition, it is suggested that they may have an important functional role, particularly in protein–protein or protein–nucleic acid interactions and recognition. Residues associated with PPII conformations represent nearly 5% of the total residues, but the lack of PPII assignment approaches prevents their systematic analysis. This short review will present current knowledge and recent research in PPII area. In a first step, the different methodologies able to assign PPII are presented. In the second step, recent studies that have shown new perspectives in PPII analysis in terms of structure and function are underlined with three cases: (1) PPII in protein structures. For instance, the first crystal structure of an oligoproline adopting an all-trans polyproline II (PPII) helix had been presented; (2) the involvement of PPII in different diseases and drug designs; and (3) an interesting extension of PPII study in the protein dynamics. For instance, PPIIs are often linked to disorder region analysis and the precise analysis of a potential PPII helix in hypogonadism shows unanticipated PPII formations in the patient mutation, while it is not observed in the wild-type form of KISSR1 protein.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Protein sequences encompass the information needed to provide the right protein folding pathways to the biologically active protein fold. Nonetheless, it is the protein functions at atomistic level that directs their structures, i.e., the biological functions need to find the proper set of local protein conformations to perform its activity. Three-dimensional structure information is usually described as a simple succession of repetitive structures (see Fig. 1), namely, the α-helix and the β-sheet, connected by “random” coil (Eisenberg 2003; Pauling and Corey 1950). Helical structures are locally stabilized by hydrogen bond patterns of backbone atoms (between residues i and i + 4) (Pauling et al. 1951), while extended structures are also maintained by hydrogen bonds but at longer distances (Pauling and Corey 1951a). They represent 1/3rd and 1/5th of the total residues, respectively. A third defined state, called β-turns, is characterized by the reversal of polypeptide chain and is stabilized by a hydrogen bond between the first and last residues (Richardson 1981; Rose 1978; Venkatachalam 1968). 25% of the residues are associated with such structures (Bornot and de Brevern 2006).
Structural characteristics of three secondary structures. a Right-handed α-helix, b left-handed PPII, and c three β-strands forming sheet. The cartoon representation highlights the structural geometry, while ball and stick represent the atomic arrangements of the three secondary structures. The proline rings can be observed in (b), and the comparison of oxygen (red) and nitrogen (blue) clearly indicates the absence of intra H-bonding in PPII. In a and c, the close proximity of oxygen and nitrogen atoms makes it favourable for intra H-bonding. High helical rise of the PPII and lack of intra H-bonding make its backbone highly solvent accessible. Visualization is done with the PyMOL software (Delano 2013) (color figure online)
However, another common repetitive conformation exists, characterized before the β-turns in the 1950s, but often forgotten, namely, Poly-l-proline-II helices II (PPII) helix (Cowan et al. 1955; Pauling and Corey 1951b) (see Fig. 1b). It can be characterized as a left-handed helical structure with dihedral angle characteristic to that of β-strands and with an overall shape resembling a triangular prism (Arnott and Dover 1968; Sasisekharan 1959) (see Fig. 2 for a comparison with other local structure conformations). The PPII helix has distinct trans-isomers of peptide bonds with dihedral angles of [−75°, +150°]. The rise per residue of PPII helix is 3.1 Å with three residues per turn. Thus, this distinct helical structure rises at 9.3 Å per turn compared to 6.0 Å pitch of a 310 helix. The primary reason for such open and relatively elongated geometry of PPII is the absence of H-donor atoms due to the cyclic side chain of proline residues. Therefore, the PPII conformation is highly acceptable of H-donor atoms from its environment or third party moieties enhancing its solvation energy. PPII is observed commonly in the collagen triple helix and hence was deemed confined to fibrous proteins (Bochicchio and Tamburro 2002; Soman and Ramakrishnan 1983; Sreerama and Woody 1994, 2003). It would be found through circular dichroïsm studies that PPII is present in folded proteins and in other structural folding contexts as well. Later, Creamer et al. (Whittington et al. 2005) demonstrated the existence of PPII in denatured proteins, while NMR studies (Toal and Schweitzer-Stenner 2014) established PPII as a favoured local structure over α-helices in denatured states. Interestingly, the presence of proline residues is not a strict requirement for a PPII and that indeed establishes PPII as a distinct class in secondary structures. Rather, it has been advocated since 1993 (Adzhubei and Sternberg 1993) to include PPII in mainstream secondary structures, such as α-helices and β-sheets. A striking fact is that residues associated with PPII conformations represent nearly 5% of the total residues in a structure (Mansiaux et al. 2011), but the lack of popular PPII assignment approaches prevents their systematic analysis.
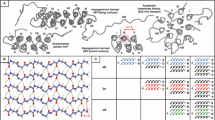
Orientation and structural organization of the different helices. a α-helix: right handed with a spherical coiling. b 310 –helix, c π-helix, and d polyproline helix: left handed with a triangular prism coiling. Proline residues are marked in yellow. e PPII helix with minimum residues possible. Only three residues can adopt a PPII conformation. In this example, none of the residue is proline. The proline rings can be observed in (d). High helical rise of the PPII can be clearly seen. Visualisation done with the PyMOL software (Delano 2013) (color figure online)
The structural properties of PPII make it highly suitable for partnered interactions. Since the backbone of PPII lacks any intra-hydrogen bonding, it requires external partners for hydration. This unique property is a reason for PPII conformation to interact with SH3 domain and thus playing a regulating role in crucial signalling pathways and cell recognition involving SH3. The distinctive structural properties, such as open, elongated structure, suggest PPII to be involved in interaction with nucleic acids. PPII has also been observed to be involved in amyloid fibrillar pathologies, such as Parkinson’s pathology. Since the PPII helix is relatively small and flexible, it is highly useful in design of cell-penetrating peptides (CPP). The current review aims at covering the different definitions of PPII based on contexts and the various methodologies that assign PPII helix. Later, it also reviews the role of PPII in protein–protein and protein–DNA interactions, involvement of PPII conformation in pathologies, and recent advances made in PPII scaffold applications.
Developments in PPII structural assignment
PPII dihedral angles are quite particular. The most classical way to analyse them is to use Ramachandran map (1963), as shown in Fig. 3. The map is based on calculations of dihedral angles between the two adjacent planes of protein backbone, hinged at C α atoms. The dihedral rotation of the planes is restricted by the steric clashes that define the disallowed regions on the map. Therefore, the map is a very powerful tool to assess the stability of a structure based on the local analysis of degrees of freedom for dihedral planes. Further evolution of the map leads to the marking of areas for specific secondary structures, namely, α-helix, β-strands, and later β-turns (see Fig. 3a). Lately, allowed region for PPII was assigned from the north-western quadrant of the map, allowed for β-strands (see Fig. 3b). A recent review catalogues the evolution of Ramachandran map very efficiently (Carugo and Djinovic-Carugo 2013). It is, however, very distinctive observation that Prof. Ramachandran incepted the idea based on the collagen hydrogen bonding argument (Bella et al. 1994; Rich and Crick 1955), which arose due to the presence of hydroxyproline.
More than 20 secondary structure assignment methods (SSAM) had been published in 30 years (Aksianov and Alexeevski 2012; Cao et al. 2016; Carter et al. 2003; Cubellis et al. 2005b; Dupuis et al. 2004; Fodje and Al-Karadaghi 2002; Frishman and Argos 1995; Hosseini et al. 2008; Hutchinson and Thornton 1996; Kabsch and Sander 1983; King and Johnson 1999; Kneller and Hinsen 2015; Labesse et al. 1997; Law et al. 2014; Majumdar et al. 2005; Martin et al. 2005; Oluwatobi Salawu 2016; Parisien and Major 2005; Park et al. 2011; Richards and Kundrot 1988; Sklenar et al. 1989; Zacharias and Knapp 2014). They have been defined with various criteria (Offmann et al. 2007): the most popular SSAM uses backbone hydrogen bonding pattern-based methods (Carter et al. 2003; Fodje and Al-Karadaghi 2002; Frishman and Argos 1995; Kabsch and Sander 1983; Zhang and Sagui 2015).
Nonetheless, very few SSAM assigns PPII to the protein coordinates. Only five SSAMs, to be more precise, include the assignment of PPII conformations. The first available approach was XTLSSTR (King and Johnson 1999), where a structure is assigned based on a simple approach similar to the visual inspection of secondary structures. It calculates three distances and two angles based on the backbone geometry and then searches for amide–amide interactions. It successfully assigns α-Helix, 310 Helix, Extended β-strand, hydrogen bonded and non-hydrogen bonded turns, and polyproline (type-II) helices.
SEGNO (Cubellis et al. 2005b) makes assignment based on distance and torsion angle calculation. For assigning PPII, it uses dihedral angles between the two-peptide planes separated by one and two residues, respectively, named diheco and diheco2. An important observation is that PPII is assigned when a residue is not defined as β-strand by SEGNO and lies within predefined values of Φ and Ψ angles. Later, taking into account the range of the four diheco angles (220–270 and 100–140), the PPII helical conformation is assigned to the residue. These thresholds are relaxed for the termini of PPII with a minimum length of the helix to be three residues and the overall shape of PPII is deemed to be like a triangular prism.
PROSS (Srinivasan and Rose 1999) uses the concept of mesostates from a torsional grid for the assignments. The grid is described as the unit squares covering all areas in a Ramachandran plot. The grids are of two kinds based on their unit area: smaller unit square: fine grid and broader unit square: coarse grid. Based on the type, each unit grid is referred to as a coarse/fine mesostate. Therefore, in principle, the Ramachandran plot is converted into a Φ/Ψ grid with marked regions (allowed, favourable, and disallowed) covering more than one mesostates. In a very similar approach related to SEGNO, PROSS also does not directly assign PPII conformation rather resolute it out after β-strand leftovers.
DSSP-PPII (Mansiaux et al. 2011) is an extension of DSSP with included dihedral angle parameters for PPII assignment, thus isolating PPII from coils. Kabsch and Sander’s DSSP (Kabsch and Sander 1983) has been the most widely used method. It is based on detection of hydrogen bonds defined under an electrostatic criterion. It makes an elaborate eight state SSA: α-Helix, 310 Helix, π-helix, β-turn, bend, extended strand, β-bridge, and coil. DSSP has been implemented in numerous databases and softwares, e.g., PDB (Berman et al. 2000; Bernstein et al. 1977) and GROMACS (Pronk et al. 2013; Van Der Spoel et al. 2005). Although being widely used and treated as a gold standard methodology, DSSP does not assign PPII. DSSP-PPII (Mansiaux et al. 2011) uses dihedral space (Φ and Ψ, −75° and +145°) to define the core of PPII while increasing by ε radiating out at 1 degree. The value of ε is chosen as an equilibrium between the number of amino acids assigned as PPII by the three previous approaches (with an extra constraints, two consecutive dihedral angles should be assigned as PPII. One of the major features of this method is to use DSSP that is already an established and trusted method for other secondary structure elements (SSE). Therefore, the code can be adapted to apparently any other assignment method, if and when required. A specific database had been proposed to the scientific community (Chebrek et al. 2014).
ASSP (Kumar and Bansal 2015), an extension of helical geometry calculation program, HELANAL-plus (Bansal et al. 2000) that is used to calculate the local helical structure parameters: twist, rise, virtual torsion, and radii. ASSP uses the difference between these parameters calculated over two or more adjacent Cα windows of four residues. Later, in the protocol, the overlaps are resolved based on the established minimum lengths of helices: α(4), 310(3), π(5), and PPII(3). Therefore, PPII conformations are assigned based on the helical geometry of the local region. Since it uses HELANAL, which further is based on Sugeta and Miyazawa, and Shakarji methods for helical geometry, ASSP tends to assign β-sheets with less efficiency (Shakarji (1998); Sugeta and Miyazawa 1967). They applied their SSAM to analyse in detail the PPII (Kumar and Bansal 2016) and found that near 3/4 of PPIIs occur in conjunction with α-helices and β-strands, and serve as linkers as well. They also underline a large number of CH···OH-bonds.
All these methods are well designed for PPII assignments. However, the number of PPII assignment approaches is still limited compared to SSAM for other secondary structure elements, and remains a limitation for the use by scientific community.
Survey of amino acids in PPII conformation
The Adzhubei and Sternberg paper in 1993 (1993) had refreshed the interest in PPII as mainstream secondary structures, such as α-helices and β-sheets, but also underlined the non-obligation of PPII to be constituted with only proline residues. Numerous mutational studies, e.g., SH3 domain—PPII peptide binding analysis provided a desired assertion that PPII conformations are favourable in denatured space (Creamer 1998; Ferreon and Hilser 2003). Impact of residue level mutations on PPII concludes that PPII conformation is retained even after successive changes of proline with alanine or glycine residues, implying that PPIIs are not constituted by a succession of proline residues alone. Therefore, PPII should rather be understood as a structural conformation found with different residue propensities in folded and unfolded states. Others experiments further establish PPII as a separate structural class (Adzhubei et al. 2013; Stapley and Creamer 1999).
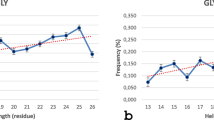
Apart from these studies, restricted coiled library analysis performed by Jha et al. explores the influence of neighbours on the residues having favourable PPII propensities (Jha et al. 2005). Examination of bias-free coiled library sets reveals dominant PPII conformation for ten of amino acid residues (Pro, Ala, Met, Glu, Leu, Asn, Cys, Gln, Lys, Gly, and Tyr). Another proposal of similar propensities comes from Cubellis and coworkers which analyse position specific propensities in 5700 PPII helices and classified data with peptide lengths (Cubellis et al. 2005a). Thus, residues, such as Ala, Met, Lys, Thr, and Leu, favour PPII conformation in longer peptides, while Asp, Ile, and Glu adopt the conformation in shorter peptides (<3 res). Trp, Phe, and Gly do not favour PPII; however, interestingly, Gly is present in a repetitive motif in collagen triple helix, while Trp and Phe have been crystallized in interaction with PPII–hydrophobic motif interactions. Thus, supposedly, these residues could stabilize and mark the terminus of a PPII helix (Cubellis et al. 2005a). In the most recent survey, Kumar and Bansal show that 40% of PPIIs contain no Pro residues. Besides, aromatic amino acids are avoided within the helix, while Gly, Asn, and Asp residues are preferred in the proximal flanking regions (Kumar and Bansal 2016).
Based on hard-sphere Monte Carlo simulations, the propagation of the PPII helix is logically explained by the interaction between the prolyl ring and the backbone (Cβ) of the previous residue. However, this logic breaks when a poly-Alanine adopts a PPII conformation, and therefore, a better explanation could be the neighbouring environment and the presence of polar residues (Creamer 1998). PPII does not have characteristic main chain H-bonding pattern; thus arguably, Ser, Thr, Gln, and other polar residues can stabilize the PPII helix by non-local hydrogen bonding with the backbone (Creamer 1998; Cubellis et al. 2005a). The overall survey of amino acid propensities reveals that propensities of amino acids in PPII are highly context based. They seem to deviate according to the presence of PPII in fibrous or globular protein context.
Role of PPII in protein–protein (PPI) and DNA–protein interactions
The distinct feature of polyproline helices is that unlike other SSE, they do not have intra-hydrogen bonding, making the backbone, as well as the side chains, highly solvent accessible. Such conformations would be hankering for finding partners for hydrogen bonding and stabilization. Therefore, the sequence and structural characteristics of PPII make it worth to be probed for partnered interactions. One of the important tools to study the PPII role in protein–protein and DNA–protein interactions is the SH3 domain models. SH3 (Src homology 3) domains are small yet important structural domains in proteins involved in cell signalling and regulation, e.g., Tyrosine kinases. SH3 domains are also well known to interact with PPII conformations (Agrawal and Kishan 2002). Hence, host-pathogen models designed with SH3 domains are critical to understand interaction space of PPII conformation with respect to proteins and/or nucleic acids. Many such studies focusing on signal transduction and cell–cell recognition have been explored for potential PPII–protein and PPII–nucleic acid interactions (Hicks and Hsu 2004; Williamson 1994). For instance, C-terminus of Synapsin-I, a protein regulating synaptic vesicle transport in neurons, is proline-rich region. Synapsin-I interacts with the cytoplasmic polyproline region of membrane protein, vesicle-associated membrane protein 1(VAMP-I) (Williamson 1994). Phosphorylation of a serine residue upstream of C-terminus PPII helix regulates the secretion of a synaptic vesicle, while VAMP-I helps in recognition. Similarly, in Ras-GTP signalling pathway, the SH3 domains of the adaptor protein bind to the polyproline region of SoS protein (xPxxPPPψxPx) leading to exchange of GTP. Another set of interactions (Booker et al. 1992) is in vacuolar sorting, where SH3 domain of phosphatidylinositol-3 kinase binds to the GTP-binding protein dynamic. Structurally, it is acknowledged that the PPII helix-binding region of SH3 domain is a smooth hydrophobic surface flanked by conserved charged residues (Booker et al. 1992). The PPII interactions also have a significant structural–functional role in transcription, as many transcription factors have proline-rich terminals (Koleske et al. 1992). This could also indicate points to the role of PPII interactions in multimeric complex formation during transcription. A well-characterized case of PPII–protein interaction is the RNA polymerase II (RNApolII). C-terminus of RNApolII has multiple copies of conserved motif YSPTSPS, which further is a two-fold SPXX motif. SPXX is a DNA binding motif found in DNA binding domains (Suzuki 1989; Suzuki et al. 1990). Furthermore, Hicks and Hsu (2004) investigated the structural aspects of PPII in DNA binding and recognition (Hicks and Hsu 2004). Exemplifying with three DNA interacting proteins; viz. third K homology domain of NOVA-2 [see Fig. 4 (Lewis et al. 2000)], the Epstein–Barr nuclear antigen-1, and the Drosophila paired protein homeodomain, they quantify the binding of PPII to the nucleotides’ minor groove and underline the specificity and non-specificity of recognition. The optimal size and specific recognition offered by PPII backbone residues strongly advocate to recognize PPII as a nucleic acid binding motif (Hicks and Hsu 2004).
Interaction of Nova protein K homology domain with RNA hairpin [PDB id: 1ec6_A (Lewis et al. 2000)]. The conserved motif of the variable loop is colour in yellow. The two PPII helices are coloured in magenta. The occurrence of C-term helix is reported to be the difference between RNA bound and unbound form. Visualisation is done with the PyMOL software (Delano 2013) (color figure online)
Functional role of polyproline in diseases
Role of PPII in protein–protein and DNA–protein interactions, and role in sorting and transport mechanisms have been investigated for its involvement in pathologies and diseases. KISS-1 Receptor (KISS1R) has in its intracellular domain three triplets of Proline–Arginine–Arginine (PRR). The addition of a fourth triplet induces the formation of a PPII, and inhibits KISS1R presentation on cell membrane. The retention of KISS1R in cytoplasm ceases the interaction with kisspeptin and thus abolishes the secretion of GnRH leading to Hypogonadotropic hypogonadism (Chevrier et al. 2013). Besides, several studies using ROA (Raman optical activity) and VBD (vibrational circular dichroism) structural visualization techniques confirm the presence of PPII conformation in pathological fibrillar aggregates (Adzhubei et al. 2013; Blanch et al. 2000; Bochicchio and Tamburro 2002). Conversion of PPII to β-sheet conformation in amyloidogenic precursor of human lysozyme may indicate a highly potential role of PPII in numerous amyloid-based conformational disorders (Blanch et al. 2000). For instance, phosphorylation of a threonine flanked by a PPII in Tau protein leads to the misfolding and aggregation of microtubular proteins in Alzheimer’s disease (Syme et al. 2002). A similar role of PPII has been found in α-synuclein, responsible for aggregation in Alzheimer’s and Parkinson’s pathologies (Adzhubei et al. 2013). Taken together, this emphasizes a deeper understanding of its structural features (Adzhubei et al. 2016).
Recent advances in polyproline research
The growing interest in physico-chemical and structural properties of PPII, especially their short extended-helical structure attracted the attention of pharmaceutical companies. Very recently, cell-penetrating vector approaches are designed based on PPII scaffold (Eiriksdottir et al. 2010; Foged and Nielsen 2008; Franz et al. 2016; Geisler and Chmielewski 2009; Ruzza et al. 2004; Yamashita et al. 2016). As explained above, PPII backbone has a high solvent accessibility and thus is highly hydrated in solvents. Therefore, use of PPII for cell penetration poses a challenge for hydrating the PPII-based moiety and their convenient uptake in hydrophobic membranes (Franz et al. 2016). Chmielewski’s group (Fillon et al. 2005) addressed this by designing and introducing cationic and hydrophobic moieties on the PPII backbone and observed no structural change. The compactness and inherent flexibility of the PPII conformation is the key to their adaptability and accompanied by cationic and hydrophobic moieties; they becomes highly suitable for a cell-penetrating vector (Foged and Nielsen 2008). The study observes a tremendous increase in PPII-based Cell-Penetrating Peptide (CPP) uptake compared to the traditional ones. Another important difference is the claimed reduction in toxicity. This is based on the observations that PPII scaffold-based CPP: Sweet Arrow Peptides—SAP(E)—obtain a net negative charge unlike the traditional CPP which are positively charged (Franz et al. 2016; Geisler and Chmielewski 2009; Li et al. 2010).
Conclusion and perspectives
Polyproline II helix is arguably a distinct member in secondary structure elements, based on its geometry, sequence, and structure. PPII has a left-handed geometry compared to right-handedness of popular protein helices (see Fig. 2). Its sequence composition varies based on the presence in a globular or fibrous protein environment. It is quite an interesting observation that proline, a major α-helix breaker/kink, when in succession adapts a distinct helical form itself. Moreover, it dominates the α-helical form in denatured space. Such examples can be appreciated in light of the expanse of the second structural space. Although PPII conformation represents only 5% of the conformational space, we highly advocate for it to be considered in regular secondary structures. Besides, its representation is equivalent if not more than the 310 helices. The involvement of PPII–protein and PPII–nucleic acid interactions in different pathologies, structural applications, and drug carriers makes it even more viable candidate to be included in the main regular secondary structures. Its potential role in Alzheimer’s and Parkinson’s could not be ignored, given recent publications on the subject. The presence of PPII in regular, ordered, and disordered regions while establishes that its distinctiveness is not sufficient to seize the complete structural space of PPII conformations. Therefore, more assignment approaches and coiled library experiments are needed to explore such conformations. This review addresses the neglect on conformations, such as PPII and bias towards “regular” secondary structures. Figure 5 shows the number of publications about PPII since 1968. The increase is clear, but remains limited. The number of papers had never been higher than 100 papers/per year. In regards to the interest of this “lost” secondary structure, we can expect a better representation in the future.
Yearwise publications trends on polyproline II helices. The bars depict the number of publications corresponding the year on x-axis. An exponential function is represented in blue curve. Dark bars show the sudden surge in publications compared to the previous year. Visualisation is done with the R software (R Core Team 2013)
References
Adzhubei AA, Sternberg MJ (1993) Left-handed polyproline II helices commonly occur in globular proteins. J Mol Biol 229:472–493. doi:10.1006/jmbi.1993.1047
Adzhubei AA, Sternberg MJ, Makarov AA (2013) Polyproline-II helix in proteins: structure and function. J Mol Biol 425:2100–2132. doi:10.1016/j.jmb.2013.03.018
Adzhubei AA, Anashkina AA, Makarov AA (2016) Left-handed polyproline-II helix revisited: proteins causing proteopathies. J Biomol Struct Dyn. doi:10.1080/07391102.2016.1229220
Agrawal V, Kishan KV (2002) Promiscuous binding nature of SH3 domains to their target proteins. Protein Pept Lett 9:185–193
Aksianov E, Alexeevski A (2012) SheeP: a tool for description of beta-sheets in protein 3D structures. J Bioinform Comput Biol 10:1241003. doi:10.1142/S021972001241003X
Arnott S, Dover SD (1968) The structure of poly-l-proline II. Acta Crystallogr B 24:599–601
Bansal M, Kumar S, Velavan R (2000) HELANAL: a program to characterize helix geometry in proteins. J Biomol Struct Dyn 17:811–819. doi:10.1080/07391102.2000.10506570
Bella J, Eaton M, Brodsky B, Berman HM (1994) Crystal and molecular structure of a collagen-like peptide at 1.9 A resolution. Science 266:75–81
Berman HM et al (2000) The Protein Data Bank. Nucleic Acids Res 28:235–242
Bernstein FC et al (1977) The protein data bank: a computer-based archival file for macromolecular structures. J Mol Biol 112:535–542
Blanch EW, Morozova-Roche LA, Cochran DA, Doig AJ, Hecht L, Barron LD (2000) Is polyproline II helix the killer conformation? A Raman optical activity study of the amyloidogenic prefibrillar intermediate of human lysozyme. J Mol Biol 301:553–563. doi:10.1006/jmbi.2000.3981
Bochicchio B, Tamburro AM (2002) Polyproline II structure in proteins: identification by chiroptical spectroscopies, stability, and functions. Chirality 14:782–792. doi:10.1002/chir.10153
Booker GW, Breeze AL, Downing AK, Panayotou G, Gout I, Waterfield MD, Campbell ID (1992) Structure of an SH2 domain of the p85 alpha subunit of phosphatidylinositol-3-OH kinase. Nature 358:684–687. doi:10.1038/358684a0
Bornot A, de Brevern AG (2006) Protein beta-turn assignments. Bioinformation 1:153–155
Cao C, Wang G, Liu A, Xu S, Wang L, Zou S (2016) A new secondary structure assignment algorithm using calpha backbone fragments. Int J Mol Sci 17:333. doi:10.3390/ijms17030333
Carter P, Andersen CA, Rost B (2003) DSSPcont: continuous secondary structure assignments for proteins. Nucleic Acids Res 31:3293–3295
Carugo O, Djinovic-Carugo K (2013) Half a century of Ramachandran plots. Acta Crystallogr D Biol Crystallogr 69:1333–1341. doi:10.1107/S090744491301158X
Chebrek R, Leonard S, de Brevern AG, Gelly JC (2014) PolyprOnline: polyproline helix II and secondary structure assignment database Database (Oxford) 2014 doi:10.1093/database/bau102
Chevrier L, de Brevern A, Hernandez E, Leprince J, Vaudry H, Guedj AM, de Roux N (2013) PRR repeats in the intracellular domain of KISS1R are important for its export to cell membrane. Mol Endocrinol 27:1004–1014. doi:10.1210/me.2012-1386
Cowan PM, McGavin S, North AC (1955) The polypeptide chain configuration of collagen. Nature 176:1062–1064
Creamer TP (1998) Left-handed polyproline II helix formation is (very) locally driven. Proteins 33:218–226
Cubellis MV, Caillez F, Blundell TL, Lovell SC (2005a) Properties of polyproline II, a secondary structure element implicated in protein-protein interactions. Proteins 58:880–892. doi:10.1002/prot.20327
Cubellis MV, Cailliez F, Lovell SC (2005b) Secondary structure assignment that accurately reflects physical and evolutionary characteristics. BMC Bioinform 6(Suppl 4):S8. doi:10.1186/1471-2105-6-S4-S8
Delano WL (2013) The PyMOL molecular graphics system on World Wide Web. http://www.pymol.org. Accessed 3 Jan 2017
Dupuis F, Sadoc JF, Mornon JP (2004) Protein secondary structure assignment through Voronoi tessellation. Proteins 55:519–528. doi:10.1002/prot.10566
Eiriksdottir E, Konate K, Langel U, Divita G, Deshayes S (2010) Secondary structure of cell-penetrating peptides controls membrane interaction and insertion. Biochim Biophys Acta 1798:1119–1128. doi:10.1016/j.bbamem.2010.03.005
Eisenberg D (2003) The discovery of the alpha-helix and beta-sheet, the principal structural features of proteins. Proc Natl Acad Sci USA 100:11207–11210
Ferreon JC, Hilser VJ (2003) The effect of the polyproline II (PPII) conformation on the denatured state entropy. Protein Sci 12:447–457. doi:10.1110/ps.0237803
Fillon YA, Anderson JP, Chmielewski J (2005) Cell penetrating agents based on a polyproline helix scaffold. J Am Chem Soc 127:11798–11803. doi:10.1021/ja052377g
Fodje MN, Al-Karadaghi S (2002) Occurrence, conformational features and amino acid propensities for the pi-helix. Protein Eng 15:353–358
Foged C, Nielsen HM (2008) Cell-penetrating peptides for drug delivery across membrane barriers. Expert Opin Drug Deliv 5:105–117. doi:10.1517/17425247.5.1.105
Franz J, Lelle M, Peneva K, Bonn M, Weidner T (2016) SAP(E)—a cell-penetrating polyproline helix at lipid interfaces. Biochim Biophys Acta 1858:2028–2034. doi:10.1016/j.bbamem.2016.05.021
Frishman D, Argos P (1995) Knowledge-based protein secondary structure assignment. Proteins 23:566–579. doi:10.1002/prot.340230412
Geisler I, Chmielewski J (2009) Cationic amphiphilic polyproline helices: side-chain variations and cell-specific internalization. Chem Biol Drug Des 73:39–45. doi:10.1111/j.1747-0285.2008.00759.x
Hicks JM, Hsu VL (2004) The extended left-handed helix: a simple nucleic acid-binding motif. Proteins 55:330–338. doi:10.1002/prot.10630
Hosseini SR, Sadeghi M, Pezeshk H, Eslahchi C, Habibi M (2008) PROSIGN: a method for protein secondary structure assignment based on three-dimensional coordinates of consecutive C(alpha) atoms. Comput Biol Chem 32:406–411. doi:10.1016/j.compbiolchem.2008.07.027
Hutchinson EG, Thornton JM (1996) PROMOTIF—a program to identify and analyze structural motifs in proteins. Protein Sci 5:212–220. doi:10.1002/pro.5560050204
Jha AK, Colubri A, Zaman MH, Koide S, Sosnick TR, Freed KF (2005) Helix, sheet, and polyproline II frequencies and strong nearest neighbor effects in a restricted coil library. Biochemistry 44:9691–9702. doi:10.1021/bi0474822
Kabsch W, Sander C (1983) Dictionary of protein secondary structure: pattern recognition of hydrogen-bonded and geometrical features. Biopolymers 22:2577–2637. doi:10.1002/bip.360221211
King SM, Johnson WC (1999) Assigning secondary structure from protein coordinate data. Proteins 35:313–320
Kneller GR, Hinsen K (2015) Protein secondary-structure description with a coarse-grained model. Acta Crystallogr D Biol Crystallogr 71:1411–1422. doi:10.1107/S1399004715007191
Koleske AJ, Buratowski S, Nonet M, Young RA (1992) A novel transcription factor reveals a functional link between the RNA polymerase II CTD and TFIID. Cell 69:883–894
Kumar P, Bansal M (2015) Identification of local variations within secondary structures of proteins. Acta Crystallogr D Biol Crystallogr 71:1077–1086. doi:10.1107/S1399004715003144
Kumar P, Bansal M (2016) Structural and functional analyses of PolyProline-II helices in globular proteins. J Struct Biol 196:414–425. doi:10.1016/j.jsb.2016.09.006
Labesse G, Colloc’h N, Pothier J, Mornon JP (1997) P-SEA: a new efficient assignment of secondary structure from C alpha trace of proteins. Comput Appl Biosci 13:291–295
Law SM, Frank AT, Brooks CL 3rd (2014) PCASSO: a fast and efficient Calpha-based method for accurately assigning protein secondary structure elements. J Comput Chem 35:1757–1761. doi:10.1002/jcc.23683
Lewis HA, Musunuru K, Jensen KB, Edo C, Chen H, Darnell RB, Burley SK (2000) Sequence-specific RNA binding by a Nova KH domain: implications for paraneoplastic disease and the fragile X syndrome. Cell 100:323–332
Li L, Geisler I, Chmielewski J, Cheng JX (2010) Cationic amphiphilic polyproline helix P11LRR targets intracellular mitochondria. J Control Release 142:259–266. doi:10.1016/j.jconrel.2009.10.012
Majumdar I, Krishna SS, Grishin NV (2005) PALSSE: a program to delineate linear secondary structural elements from protein structures. BMC Bioinformatics 6:202. doi:10.1186/1471-2105-6-202
Mansiaux Y, Joseph AP, Gelly JC, de Brevern AG (2011) Assignment of polyproline II conformation and analysis of sequence–structure relationship. PLoS One 6:e18401. doi:10.1371/journal.pone.0018401
Martin J, Letellier G, Marin A, Taly JF, de Brevern AG, Gibrat JF (2005) Protein secondary structure assignment revisited: a detailed analysis of different assignment methods. BMC Struct Biol 5:17. doi:10.1186/1472-6807-5-17
Offmann B, Tyagi M, de Brevern AG (2007) Local protein structures. Curr Bioinform 3:165–202
Oluwatobi Salawu E (2016) RaFoSA: random forests secondary structure assignment for coarse-grained and all-atom protein systems. Cogent Biol 2:1214061
Parisien M, Major F (2005) A new catalog of protein beta-sheets proteins 61:545–558. doi:10.1002/prot.20677
Park SY, Yoo MJ, Shin J, Cho KH (2011) SABA (secondary structure assignment program based on only alpha carbons): a novel pseudo center geometrical criterion for accurate assignment of protein secondary structures. BMB Rep 44:118–122. doi:10.5483/BMBRep
Pauling L, Corey RB (1950) Two hydrogen-bonded spiral configurations of the polypetide chain. J Am Chem Soc 72:5349
Pauling L, Corey RB (1951a) The pleated sheet, a new layer configuration of polypeptide chains. Proc Natl Acad Sci USA 37:251–256
Pauling L, Corey RB (1951b) The structure of fibrous proteins of the collagen-gelatin group. Proc Natl Acad Sci USA 37:272–281
Pauling L, Corey RB, Branson HR (1951) The structure of proteins; two hydrogen-bonded helical configurations of the polypeptide chain. Proc Natl Acad Sci USA 37:205–211
Pronk S et al (2013) GROMACS 4.5: a high-throughput and highly parallel open source molecular simulation toolkit. Bioinformatics 29:845–854. doi:10.1093/bioinformatics/btt055
R Core Team (2013) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Ramachandran GN, Ramakrishnan C, Sasisekharan V (1963) Stereochemistry of polypeptide chain configurations. J Mol Biol 7:95–99
Rich A, Crick FH (1955) The structure of collagen. Nature 176:915–916
Richards FM, Kundrot CE (1988) Identification of structural motifs from protein coordinate data: secondary structure and first-level supersecondary structure. Proteins 3:71–84. doi:10.1002/prot.340030202
Richardson JS (1981) The anatomy and taxonomy of protein structure. Adv Protein Chem 34:167–339
Rose GD (1978) Prediction of chain turns in globular proteins on a hydrophobic basis. Nature 272:586–590
Ruzza P, Calderan A, Guiotto A, Osler A, Borin G (2004) Tat cell-penetrating peptide has the characteristics of a poly(proline) II helix in aqueous solution and in SDS micelles. J Pept Sci 10:423–426. doi:10.1002/psc.558
Sasisekharan V (1959) Structure of poly-l-proline II. Acta Crystallogr 12:897–903
Shakarji CM (1998) Least-squares fitting algorithms of the NIST algorithm testing system. J Res Natl Inst Stand Technol 103:633
Sklenar H, Etchebest C, Lavery R (1989) Describing protein structure: a general algorithm yielding complete helicoidal parameters and a unique overall axis. Proteins 6:46–60. doi:10.1002/prot.340060105
Soman KV, Ramakrishnan C (1983) Occurrence of a single helix of the collagen type in globular proteins. J Mol Biol 170:1045–1048
Sreerama N, Woody RW (1994) Poly(pro)II helices in globular proteins: identification and circular dichroic analysis. Biochemistry 33:10022–10025
Sreerama N, Woody RW (2003) Structural composition of betaI- and betaII-proteins. Protein Sci 12:384–388. doi:10.1110/ps.0235003
Srinivasan R, Rose GD (1999) A physical basis for protein secondary structure. Proc Natl Acad Sci USA 96:14258–14263
Stapley BJ, Creamer TP (1999) A survey of left-handed polyproline II helices. Protein Sci 8:587–595. doi:10.1110/ps.8.3.587
Sugeta H, Miyazawa T (1967) General method for calculating helical parameters of polymer chains from bond lengths, bond angles, and internal-rotation angles. Biopolymers 5:673–679
Suzuki M (1989) SPXX, a frequent sequence motif in gene regulatory proteins. J Mol Biol 207:61–84
Suzuki M, Sohma H, Yazawa M, Yagi K, Ebashi S (1990) Histone H1 kinase specific to the SPKK motif. J Biochem 108:356–364
Syme CD, Blanch EW, Holt C, Jakes R, Goedert M, Hecht L, Barron LD (2002) A Raman optical activity study of rheomorphism in caseins, synucleins and tau. New insight into the structure and behaviour of natively unfolded proteins. Eur J Biochem 269:148–156
Toal S, Schweitzer-Stenner R (2014) Local order in the unfolded state: conformational biases and nearest neighbor interactions. Biomolecules 4:725–773. doi:10.3390/biom4030725
Van Der Spoel D, Lindahl E, Hess B, Groenhof G, Mark AE, Berendsen HJ (2005) GROMACS: fast, flexible, and free. J Comput Chem 26:1701–1718. doi:10.1002/jcc.20291
Venkatachalam CM (1968) Stereochemical criteria for polypeptides and proteins. V. Conformation of a system of three linked peptide units. Biopolymers 6:1425–1436
Whittington SJ, Chellgren BW, Hermann VM, Creamer TP (2005) Urea promotes polyproline II helix formation: implications for protein denatured states. Biochemistry 44:6269–6275. doi:10.1021/bi050124u
Williamson MP (1994) The structure and function of proline-rich regions in proteins. Biochem J 297(Pt 2):249–260
Yamashita H et al (2016) Development of a cell-penetrating peptide that exhibits responsive changes in its secondary structure in the cellular environment. Sci Rep 6:33003. doi:10.1038/srep33003
Zacharias J, Knapp EW (2014) Protein secondary structure classification revisited: processing DSSP information with PSSC. J Chem Inf Model 54:2166–2179. doi:10.1021/ci5000856
Zhang Y, Sagui C (2015) Secondary structure assignment for conformationally irregular peptides: comparison between DSSP, STRIDE and KAKSI. J Mol Graph Model 55:72–84. doi:10.1016/j.jmgm.2014.10.005
Acknowledgements
We would like to thank Catherine Etchebest for fruitful discussions. This work was supported by grants from the Ministry of Research (France), University Paris Diderot, Sorbonne, Paris Cité (France), National Institute for Blood Transfusion (INTS, France), National Institute for Health and Medical Research (INSERM, France), and labex GR-Ex. The labex GR-Ex, reference ANR-11-LABX-0051 is funded by the program “Investissements d’avenir” of the French National Research Agency, reference ANR-11-IDEX-0005-02. TjrN, NSr, and AdB acknowledge to Indo-French Centre for the Promotion of Advanced Research/CEFIPRA for collaborative grant (number 5302-2). NSh acknowledges support from ANRT (CIFRE convention number 2015/0832). NSr acknowledges for J.C. Bose fellowship and general support from DBT. AMV is supported by Allocations Regionales de Researche grant from Region Reunion.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: J. D. Wade.
Rights and permissions
About this article
Cite this article
Narwani, T.J., Santuz, H., Shinada, N. et al. Recent advances on polyproline II. Amino Acids 49, 705–713 (2017). https://doi.org/10.1007/s00726-017-2385-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00726-017-2385-6