Abstract
Leaf rust (LR) is a destructive foliar disease of common wheat (Triticum aestivum L.) worldwide. Growing resistant cultivars with adult plant resistance (APR) are the ideal way to control the disease. The aim of the present study was to map quantitative trait loci (QTL) for leaf rust resistance using 137 doubled-haploid (DH) lines derived from the Pingyuan 50 × Mingxian 169 cross. The same population was previously tested for stripe rust (YR) and powdery mildew (PM) response, and the data were reassessed to determine the presence of QTL with pleiotropic effects. Phenotyping for leaf rust response was conducted in six environments at Baoding in Hebei province and Zhoukou in Henan province. The DH lines were genotyped using the wheat 55K SNP array and additional SSR markers. Eight QTLs for LR resistance were identified in the present study using the inclusive composite interval mapping (ICIM). The flanking sequences of all the SNP probes were subjected to BLAST against the Chinese Spring reference sequence to determine physical positions. Six LR resistance loci were contributed by Pingyuan 50, but QLr.hebau-2DS2 and QLr.hebau-3AL were from Mingxian 169. QLr.hebau-3DS, a new and stable QTL detected in all environments, explained 4.2–20.0% of the phenotypic variation. Another new QTL, QLr.hebau-3AL detected in four environments had a PVE (phenotypic variance explained) of 5.7–10.1%. QLr.hebau-1BL, QLr.hebau-2DS2, and QLr.hebau-7DL are new loci. Two QTLs showed potential pleiotropic effects on disease response. QTL on chromosome 7DL might be pleiotropic to LR and PM, and the QTL on 5DL might confer pleiotropic resistance to LR and YR. Kompetitive allele-specific PCR (KASP) markers for QLr.hebau-3AL and QLr.hebau-3DS were successfully developed and validated. The QTL identified with their flanking markers and in the present study could be useful for fine mapping, candidate gene mining, and marker-assisted selection (MAS).
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Leaf rust, caused by Puccinia triticina (Pt), is one of the most important diseases of common wheat (Triticum aestivum L.) worldwide. The leaf rust pathogen primarily attacks the leaf blades although it can also infect the leaf sheath and glumes in highly susceptible cultivars (Huerta-Espino et al. 2011). This disease can cause yield losses of up to 40% under epidemic conditions mainly through reduction of kernel weight and kernel number per spike (Hershman 1985; Knott 1989). In China, significant epidemics occurred in 1969, 1973, 1975, 1979, and 2012 (Zhou et al. 2013). In those years, climate conditions were apparently favorable for leaf rust (Jin et al. 2017). Although leaf rust can be controlled by fungicides, the use of resistant cultivars is more effective, economic, and environmentally safe.
Resistance to rust is broadly classified into two types, race specific, and race non-specific (Johnson 1988). Race-specific resistance is effective against some but not all pathogen pathotypes. It tends to be controlled by a single gene or some gene combinations and to elicit a hypersensitive response to avirulent pathotypes (Lowe et al. 2011). This type of resistance predominantly applies in the entire growth cycle and therefore has been described as all-stage resistance. Race-non-specific resistance is quantitatively inherited, usually manifested at later growth stages, and often effective against multiple pathogens (Caldwell 1968). This kind of resistance gene is often referred to as adult plant resistance (APR) or slow rusting resistance. The important feature of race-non-specific resistance is its durability (Singh et al. 2005).
Seventy-nine leaf rust resistance genes are currently cataloged (Qureshi et al. 2018). Most of these genes confer race-specific resistance, but eight slow-rusting genes, Lr34 (Dyck 1977), Lr46 (Singh et al. 1998), Lr67 (Herrera-Foessel et al. 2011), Lr68 (Herrera-Foessel et al. 2012), Lr74 (McIntosh et al. 2016), Lr75 (Singla et al. 2017), Lr77 (Kolmer et al. 2018a), and Lr78 (Kolmer et al. 2018b) confer race-non-specific resistance. In addition, a total of 249 QTLs have been described (Pinto da Silva et al. 2018). Identification of new slow-rusting genes is important for breeding cultivars with durable resistance.
A saturated genetic linkage map can improve the precision of QTL mapping, especially those with minor or moderate effectiveness (Stange et al. 2013). High-throughput genotyping platforms are now widely used in genetic mapping of wheat disease resistance genes (Gao et al. 2015; Wen et al. 2017; Zou et al. 2017) include the 9K, 15K, 35K, 55K, 90K, 660K, and 820K SNP arrays (Allen et al. 2017; Cavanagh et al. 2013; Cui et al. 2017; Wang et al. 2014; Winfield et al. 2016). Among them, the wheat 55K SNP array based on the 660K SNP array (http://bioservices.capitalbio.com/index.shtml) was especially developed for wheat by the Chinese Academy of Agricultural Sciences and Affymetrix. The approximately 2600 SNP markers were carefully chosen by considering polymorphism in various wheat collections, and on uniform distribution across all 21 wheat chromosomes.
Pingyuan 50, a leading landrace cultivar in the Yellow and Huai Valley Autumn-sown Wheat Zone of China in the 1950s, has shown APR to leaf rust, stripe rust, and powdery mildew for more than 60 years. In previous studies, QTL for stripe rust and powdery mildew were identified in Pingyuan 50/Mingxian 169 population (Asad et al. 2014; Lan et al. 2010) using SSR markers. In the present study, we used the same Pingyuan 50 × Mingxian 169 DH population and the 55K SNP array genotyping platform to identify QTL for APR to LR and find closely linked molecular markers. We also reanalyzed the earlier stripe rust and powdery mildew data to identify colocated resistance loci.
Materials and methods
Plant materials and Pt pathotypes
The Pingyuan 50 × Mingxian 169 DH population consisted of 137 lines. Zhengzhou 5389 was used as a susceptible control in the test. Three Pt pathotypes THJS, PGTS, and PHTT were used in the field trail. Both Pingyuan 50 and Mingxian 169 showed susceptible infection type (IT) to the three Pt pathotypes at the seedling stage (Fig. S1). In addition, a diverse bread wheat panel including 43 wheat cultivars (lines) from Sichuan province and 54 landraces was used for QTL validation (Table S1).
Field experiments
The 137 DH lines and parents were evaluated for leaf rust response in the field at Baoding in Hebei province (located at 115.47° E longitude and 38.85° N latitude), during the 2010/2011, 2011/2012, 2012/2013, and 2013/2014 cropping seasons, and Zhoukou in Henan province (situated at 114.53° E longitude and 33.80° N latitude) during the 2010/2011 and 2011/2012 cropping seasons (herein referred to as 2011BD, 2012BD, 2013BD, 2014BD, 2011ZK, and 2012ZK, respectively). Both places are hotspot locations for leaf rust infection and spread (Huerta-Espino et al. 2011). The field trials were conducted in randomized complete blocks with three replications (except in 2014BD with a single replication). Field plots were single 1.2 m rows sown with approximately 40 seeds and 30 cm apart between rows. The control Zhengzhou 5389 was planted every tenth row to aid the spread urediniospores in the trial, and it was also planted perpendicular and adjacent to the test rows. Leaf rust infection was initiated by spraying the spreader at tillering stage (Feekes growth stage 5; Large 1954) with a water suspension of an equal urediniospore mixture of Pt pathotypes THJS, PGTS, and PHTT to which a few drops of Tween 20 (0.03%) were added. Avirulence/virulence formulas for three pathotypes were listed as follows: THJS: Lr3ka, 9, 15, 18, 19, 21, 24, 28, 29, 30, 42, 44, 47, 51, 53/Lr1, 2a, 2b, 2c, 3, 3bg, 10, 11, 13, 14a, 14b, 16, 17, 20, 23, 26, 33, 36, 39, 45, B. PGTS: Lr2a, 9, 15, 18, 19, 20, 24, 26, 28, 29, 42, 44, 47, 51, 53/Lr1, 2b, 2c, 3, 3ka, 3bg, 10, 11, 13, 14a, 14b, 16, 17, 21, 23, 30, 33, 36, 39, 45, B. PHTT: Lr2a, 9, 15, 19, 20, 24, 28, 29, 30, 42, 44, 47, 51, 53/Lr1, 2b, 2c, 3, 3ka, 3bg, 10, 11, 13, 14a, 14b, 16, 17, 18, 21, 23, 26, 33, 36, 39, 45, B. Disease severities as percentage leaf area covered with uredinia were scored three times at about 1-week interval with the first scoring 4 weeks after inoculation (Feekes growth stage 10) in each environment according to the modified Cobb scale (Peterson et al. 1948).
Statistical analysis
Phenotypic correlation coefficients between maximum disease severities (MDS) in each environment and analysis of variance (ANOVA) were conducted with SAS (SAS Institute, Cary, NC) software. Broad-sense heritabilities (h2) for leaf rust resistance were calculated by: h2 = σg2/(σg2 + σge2/e + σε2/re), where σg2, σge2, and σε2 were estimates of genotypic, genotype × environment interaction, and error variances, respectively, and e and r were the numbers of environments and replications per environment.
Genotyping
Genomic DNA of parents and DH lines were extracted from non-infected seedling leaves using the CTAB method (Sharp et al. 1988). The 137 DH lines and parents were genotyped with the Affymetrix 55K SNP array (53,064 markers) by CapitalBio Technology Company, Beijing. SNP with monomorphism, markers with > 20% of missing data points, and sites with distorted segregation ratios (P < 1E−5) were removed. In addition, genotypic data of 21 SSR markers (Table S2) from Lan et al. (2010) were also used for further linkage map in the present study. The linkage map was constructed using IciMapping 4.1 (http://www.isbreeding.net/software/?type=detail&id=18, Li et al. 2007). The MapChart 2.3 (http://www.earthatlas.mapchart.com/, Voorrips 2002) was used to draw linkage maps.
QTL detection
QTL mapping of LR MDS in each environment were carried out using ICIM in IciMapping 4.1. YR and PM data kindly provided by Dr. Caixia Lan (Lan et al. 2010) and Dr. Muhammad Azeem Asad (Asad et al. 2014) were reanalyzed using the Affymetrix 55K SNP array. The logarithm of odds (LOD = 3.0) threshold to declare a QTL for each trait was based on 1000 permutation tests. Stepwise regression was used to detect the percentage of variance explained (PVE) by individual QTL and additive effects at LOD peaks. The flanking sequences of all the SNP probes were subjected to BLAST against the Chinese Spring reference sequence (version 1.0 https://urgi.versailles.inra.fr/blast_iwgsc/blast.php, IWGSC 2018) in order to determine physical positions. Factorial ANOVA were conducted to test for the significances of interactions between stable detected resistance loci based on MDS.
KASP marker design and QTL validation
SNP markers highly associated with specific QTL were selected and converted to KASP markers using PolyMarker (Ramirez-Gonzalez et al. 2015). KASP assays were performed in a S1000 Thermal Cycler (Bio-Rad, Hercules, CA). Reaction mixtures consisted of final volumes of 5 μL containing 2.5 μL of 2 × KASP V4.0 Mastermix (LGC Genomics), 0.056 μL of assay primer mix (12 mM of each allele-specific primer and 30 mM of common primer) and 50–100 ng of genomic DNA. The cycling conditions were as follows: 94 °C for 15 min, nine cycles of 94 °C for 20 s, touchdown starting at 65 °C for 60 s (decreasing 0.8 °C per cycle), 32 cycles of 94 °C for 20 s, and 57 °C for 60 s. End-point fluorescence data were visualized with a microplate reader (FLUOstar Omega, BMG LABTECH, Germany) and analyzed by Klustering Caller software (LGC Genomics). The KASP markers were used to detect the population and the diverse bread wheat genotypes. T tests were conducted to compare the two alleles’ effect on LR based on the phenotypic data in the field using the diverse bread wheat panel.
Results
Response to leaf rust
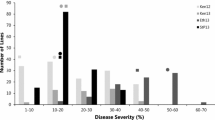
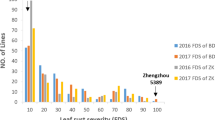
The MDS of susceptible control Zhengzhou 5389 ranged from 90 to 100% across six environments. The mean LR MDS score for Pingyuan 50 and Mingxian 169 was 10.8% and 42.5% across all six environments, respectively, indicating minor genes might be present in both parents. Mean LR severities on the DHs ranged from 12.7 to 30.5% across all trials (Table 1). The frequency distribution of leaf rust MDS in each environment showed a continuous distribution skewed towards resistance (Fig. 1), indicating polygenic inheritance. The MDS in six environments were significantly correlated with Pearson correlation coefficients (r) ranging from 0.35 to 0.78 (P < 0.001) (Table 2). Broad-sense heritability of MDS across environments was 0.81. ANOVA confirmed significant variation among the genotypes, environments, and genotype × environment (Table 3).
Frequency distributions of Pingyuan 50 × Mingxian 169 DH lines for maximum leaf rust severity. Mean values for the parents, Pingyuan 50 and Mingxian 169, are indicated by arrows. 2011BD, 2012BD, 2013BD, 2014BD, 2011ZK, and 2012ZK: the MDS in 2010/2011, 2011/2012, 2012/2013 and 2013/2014 cropping seasons in Baoding, Hebei province, and Zhoukou in Henan during the 2010–2011 and 2011–2012 cropping seasons, respectively
Linkage map construction
A total of 7991 polymorphic markers (including 21 SSR markers) represented by 2382 markers in the A genome, 3852 in the B genome and 1757 in the D genome were used to construct the linkage map. They spanned 1564, 2166, and 1763 cM with marker densities of 0.66, 0.56, and 1.06 cM/marker in the A, B, and D genomes, respectively.
QTL mapping for leaf rust resistance
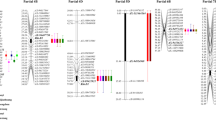
Based on the MDS data, eight QTL for leaf rust resistance were detected. They were designated as QLr.hebau-1BL, QLr.hebau-2DS2, QLr.hebau-3AL, QLr.hebau-3DS, QLr.hebau-4DL, QLr.hebau-5AL, QLr.hebau-5DL, and QLr.hebau-7DL, respectively. The resistance alleles of QTL on 1BL, 3DS, 4DL, 5AL, 5DL, and 7DL were derived from Pingyuan 50, and those on 2DS and 3AL were from Mingxian 169 (Table 4, Fig. 2).
A major QTL for LR resistance, QLr.hebau-3DS, flanked by the markers AX-111490323 and AX-109395143 detected in all tests and explained 4.2–20.0% of the phenotypic variances across all environments (Table 4). The second QTL, QLr.hebau-3AL, flanked by AX-109036576 and AX-111464284, was detected in 2011BD, 2012BD, 2012ZK, and 2013BD, explaining 5.7, 7.6, 10.1, and 9.0% of the phenotypic variances, respectively (Table 4).
QTLs QLr.hebau-4DL, QLr.hebau-5DL, and QLr.hebau-7DL were detected in 2011BD and 2013BD (Table 4). QLr.hebau-4DL, QLr.hebau-5DL, and QLr.hebau-7DL explained 4.6–11.0%, 15.1–22.6%, and 4.9–9.6% of the phenotypic variance, respectively. QLr.hebau-2DS2 flanked by AX-108903243 and AX-110055353 was detected in 2012BD and 2014BD, accounting for 14.6% and 13.2% of the phenotypic variances, respectively.
Two QTLs, QLr.hebau-1BL and QLr.hebau-5AL, with minor effect were each detected in a single environment (Table 4).
Potentially pleiotropic QTL for multiple wheat disease
The stripe rust and powdery mildew MDS data from Lan et al. (2010) and Asad et al. (2014) were reanalyzed to identify potentially pleiotropic QTL conferring resistance to more than one disease. Two QTLs were potentially pleiotropic (Table 4; Fig. 2). Stripe rust locus 5D.3 detected in one environment was at a similar position to QLr.hebau-5DL and that locus might be pleiotropic for LR and YR resistance. QTL on 7DL for powdery mildew resistance detected in single environment was mapped near QLr.hebau-7DL and could be pleiotropic to both LR and powdery mildew resistance factors (Table 4, Table S3).
Interactions between QLr.hebau-3DS and QLr.hebau-3AL for LR
The DHs were divided into four groups (+3AL+3DS, −3AL+3DS, +3AL–3DS, and −3AL–3DS) to analyze the interactions between the two stable QTL (Table 5, Fig. S2). The flanking markers of each QTL were used in determining the presence of parental alleles in the DHs. DHs with combination of QLr.hebau-3AL and QLr.hebau-3DS resistance allele significantly reduced leaf rust MDS compared to those lines that lacked both QTL in all environments (Table 5). The additive and epistasis effects of QLr.hebau-3AL and QLr.hebau-3DS were calculated using the model only considering the effects of the two QTL rather than all the QTL (Table 6). Results showed that the additive effects were the main effects in five environments in addition to 2014BD whose additive effect together with the epistasis effect reducing LR severity (Table 6). Results of AVONA analysis also showed the similar results that there was no significant interaction between the two QTL in five environments and only a marginal interaction was observed in 2014BD (P = 0.01) (Table S4).
KASP markers development and validation
A total of 25 KASP markers were designed and then used to screen the population and the diverse wheat panel. Three and one SNP markers associated with QLr.hebau-3AL and QLr.hebau-3DS, respectively, were successfully converted to KASP markers and successfully genotyped, representing the peak or flanking markers for the identified QTL on these chromosomes (Table S5). The allelic analyses were conducted according to the field phenotype data of the diverse wheat panel. The resistant allele of QLr.hebau-3DS showed significantly difference (at P < 0.05) than the susceptible allele, reducing 4.3–13.5% MDS in the field test. For QLr.hebau-3AL, the resistance allele decreased 2.3–10.5% of leaf rust MDS (Table S6) compared with susceptible allele. These KASP markers can be used in marker-assisted selection (MAS).
Discussion
Comparisons with previous reports of LR QTL
QLr.hebau-3DS
QLr.hebau-3DS was detected in all the environments and explained 4.2–20.0% of the phenotypic variance. To date, four APR genes, QLr.tam-3D/QYr.tam-3D (Basnet et al. 2014), QLr.cim-3DC (Lan et al. 2014), QLr.cim-3DC (Lan et al. 2017), and QLr.cdl-3D (Kolmer et al. 2018c) were mapped on chromosome 3DS. QLr.tam-3D/QYr.tam-3D with pleiotropic resistance to LR and YR from CIMMYT line Quaiu 3 was located at about 118.2 Mb (Basnet et al. 2014) (Table 7). QLr.cim-3DC from Francolin was detected near the centromere of 3D (Lan et al. 2014). Lan et al. (2017) mapped QLr.cim-3DC flanking by gwm341 - barc1119 at a physical position of 109.5–297.5 Mb. QLr.cdl-3D detected in Thatcher*3 × Americano 44d population was linked with K16928 at physical position 5.6 Mb (Table 7). In the study, QLr.hebau-3DS mapped at 40.8–53.2 Mb is different from those mentioned QTL and might be new.
QLr.hebau-3AL
APR QTL QLr.ubo-3A, QLr.sfrs-3A, and QLr.fcu-3AL were reported on chromosome 3A in a durum population of Colosseo × Lloyd (Maccaferri et al. 2008), and common wheat populations Forno × Oberkulmer (Messmer et al. 2000) and TA4152–60 × ND495 (Chu et al. 2009), respectively. QLr.ubo-3A was located on 3AS between 311707 and Xwmc664 (Li et al. 2014). Obviously, this QTL should be different from QLr.hebau-3AL. QLr.sfrs-3A was mapped near 3A centromere (316.9–319.9 Mb) in Forno (Li et al. 2014). QLr.fcu-3AL was mapped to the marker interval Xgwm666 - Xcfa2183. According to the Chinese Spring wheat reference sequence, Xcfa2183 is mapped at 709.4 Mb (Table 7). QLr.hebau-3AL was mapped between AX-109036576 and AX-111464284 with the physical region 556.5–595.9 Mb; hence, these QTLs are different.
QLr.hebau-1BL
Several QTL for LR resistance had been mapped to chromosome 1BL (Li and Bai 2009; Lillemo et al. 2008; Ren et al. 2012; Rosewarne et al. 2008; Rosewarne et al. 2012; Suenaga et al. 2003; William et al. 2003; William et al. 2006). These QTLs have proved to be Lr46 or located at Lr46 region on chromosome 1BL (Li et al. 2014). The physical position of Lr46 at about 670.2 Mb in the Chinese Spring reference map based on the position of csLV46g22 (Table 7). QLr.hebau-1BL.2 was mapped to physical positions of 612.8–624.6 Mb. Moreover, tests with marker csLV46G22 indicated that Lr46 was not present in Pingyuan 50 or Mingxian 169 (data not shown). Hence, QLr.hebau-1BL is different from Lr46.
QLr.hebau-2DS2
To date, APR genes Lr22a (Hiebert et al. 2007) and Lr22b and eight QTLs, viz., QLr.cimmyt-2DS (Rosewarne et al. 2012), QLrlp.osu-2DS (Xu et al. 2005a), QLr.hbau-2DS (Zhang et al. 2009), QLrid.osu-2DS (Xu et al. 2005b), QLr.sfr-2DS (Schnurbusch et al. 2004), QLr.hebau-2DS (Zhang et al. 2017), Qlr.inra-2D (Azzimonti et al. 2014), and QLr.mna-2DS (Tsilo et al. 2014), have been mapped on 2DS. QLr.cimmyt-2DS, QLrlp.osu-2DS, and QLr.hbau-2DS were distally located and linked to SSR marker barc124 at physical position 3.7 Mb (Li et al. 2014, Table 7). QLrid.osu-2DS, QLr.hebau-2DS, and Lr22a were linked with gwm261 at physical position 19.6 Mb (Table 7). QLr.sfr-2DS was mapped at 13.7–23.0 Mb based on the linked markers (Table 7). Qlr.inra-2D (Azzimonti et al. 2014) was linked with cfd36 (14.4 Mb), and QLr.mna-2DS (Tsilo et al. 2014) was linked with wmc453 (56.8 Mb). QLr.hebau-2DS2 is closely linked with AX-108903243 in the study at 49.2 Mb in the Chinese Spring Reference sequence and different from the known region.
QLr.hebau-4DL
The known LR gene, Lr67 (Herrera-Foessel et al. 2011), and two known QTLs, QLr.fcu-4DL (Chu et al. 2009) and QLr.sfrs-4DL (Messmer et al. 2000), were mapped on 4DL. QLr.fcu-4DL and Lr67 were mapped at a similar position of around 412.7 Mb according to the position of closely linked markers (Table 7). In the present study, QLr.hebau-4D closely linked with AX-111002463 in the physical distance 428.6 Mb. QLr.sfrs-4DL was mapped on the distal end of chromosome 4DL, a clearly different location from QLr.hebau-4D (Li et al. 2014). Lr67 also conferred resistance to PM and YR, whereas no effect of QLr.hebau-4D on PM and YR response was detected in the present study. Therefore, QLr.hebau-4D appears to differ from Lr67. The relationship of QLr.hebau-4D and QLr.fcu-4DL needs to be further determined.
QLr.hebau-5AL
Rosewarne et al. (2012) identified QLr.cimmyt-5AL, flanked by wPT-0373 and wPT-0837 in wheat cultivar Avocet. The QTL was mapped at about 2 cM from Vrn-A1 (Rosewarne et al. 2012), and Vrn-A1 was located at 587.0 Mb on the physical map. According to the closely linked markers, QLr.hebau-5AL was mapped on 611.8–631.6 Mb physical distance. Hence, the two QTLs seem to be different.
QLr.hebau-5DL
Messmer et al. (2000) mapped a LR resistant QTL derived from cultivar Oberkulmer flanked by Xpsr906a and Xpsr580a on chromosome 5DL located at about 547.0 Mb (Table 7). QLr.hebau-5DL (mapped at 556.4–561.6 Mb) might be at similar position with Xpsr580a. Soriano and Royo (2015) reported gwm2 was closely linked with the QTL from Oberkulmer. In the study, gwm2 was used to detect the whole population and no polymorphism was observed between Pingyuan 50 and Mingxian 169. QLr.hebau-5DL has a pleiotropic effect to YR and LR resistance but the YR status of the gene in Oberkulmer is unknown.
QLr.hebau-7DL
No known QTL was reported on 7DL at present, so QLr.hebau-7DL should be a novel QTL.
QTL for YR and PM
Lan et al. (2010) mapped three QTL for YR resistance and Asad et al. (2014) mapped four QTL for PM resistance using SSR markers in previous studies. In this study, the earlier stripe rust and powdery mildew data were reanalyzed using the 55 K SNP platform and eight QTL for YR and seven QTL for PM were mapped (Table S3 and Fig. S3). Eight QTLs for YR on chromosomes 1AL, 2AL, 3BS, 5AL, 5D (3), and 6BL were detected. QTL on 1AL and 5D.1 with major effect from Pingyuan 50 were each detected in two environments. The other Yr QTL was detected in one environment. Three QTLs on 1AL, 1DS, and 2DS for powdery mildew resistance were detected in two environments. Four QTLs detected in 3DS, 4AS, 5DL, and 7DL were detected in single environments (Table S3).
In the study, better results have been obtained using high density SNP markers compared with previous reports based only on SSR markers. The QTL results were not consistent with previous results except for QTL on 5AL for YR resistance. This might be due to the following reasons: (a) in the present study, wheat 55K SNP chip with much more polymorphic markers was used for genotyping. However, in previous study, only limited SSR markers on few chromosomes were used for QTL analysis. (b) High LOD score (LOD = 3.0) was set in the present study compared with low LOD score (LOD = 2.0) in previous study. Therefore, some QTLs detected in previous study cannot be detected in this study.
Implications of QTL detected in the population
The Chinese landrace Pingyuan 50 has shown high levels of resistance to leaf rust, stripe rust, and powdery mildew for more than 60 years. In this study, eight QTLs for LR including three potentially pleiotropic QTL and closely linked SNP markers were detected in the Pingyuan 50 × Mingxian 169 DH population. Among these, the new APR QTL QLr.hebau-3DS and QLr.hebau-3AL conferred stable effect for leaf rust. The resistance in both cultivars was conditioned by a number of QTLs that had relatively small effects individually, but acted in an additive manner for a good level of resistance. The SNP markers linked with the two QTLs were successfully converted to KASP assays and will facilitate the pyramiding of rust resistance loci into future cultivars and used in MAS in wheat breeding programs.
References
Allen AM, Winfield MO, Burridge AJ, Downie RC, Benbow HR, Barker GLA, Wilkinson PA, Coghill J, Waterfall C, Davassi A, Scopes G, Pirani A, Webster T, Brew F, Bloor C, Griffiths S, Bentley AR, Alda M, Jack P, Phillips AL, Edwards KJ (2017) Characterization of a wheat breeders’ array suitable for high-throughput SNP genotyping of global accessions of hexaploid bread wheat (Triticum aestivum). Plant Biotechnol J 15:390–401
Asad MA, Bai B, Lan C, Yan J, Xia X, Zhang Y, He Z (2014) Identification of QTL for adult-plant resistance to powdery mildew in Chinese wheat landrace Pingyuan 50. Crop J 2:308–314
Azzimonti G, Marcel TC, Robert O, Paillard S, Lannou C, Goyeau H (2014) Diversity, specificity and impacts on field epidemics of QTLs involved in components of quantitative resistance in the wheat leaf rust pathosystem. Mol Breed 34:549–567
Basnet B, Singh R, Ibrahim A, Herrera-Foessel S, Huerta-Espinot J, Lan C, Rudd J (2014) Characterization of Yr54 and other genes associated with adult plant resistance to yellow rust and leaf rust in common wheat Quaiu 3. Mol Breed 33:385–399
Caldwell RM (1968) Breeding for general and/or specific plant disease resistance. In: Finlay KW, Shepherd KW (eds) Proc 3rd Int Wheat Genet Symp, Australian Academy of Science. Canberra, Australia, pp 263–272
Cavanagh CR, Chao S, Wang S, Huang BE, Stephen S, Kiani S, Forrest K, Saintenac C, Brown-Guedira GL, Akhunova A, See D, Bai G, Pumphrey M, Tomar L, Wong D, Kong S, Reynolds M, Lopez da Silva M, Bockelman H, Talbert L, Anderson JA, Dreisigacker S, Baenziger S, Carter A, Korzun V, Morrell PL, Dubcovsky J, Morell MK, Sorrells ME, Hayden MJ, Akhunov E (2013) Genome-wide comparative diversity uncovers multiple targets of selection for improvement in hexaploid wheat landraces and cultivars. Proc Nati Acad Sci USA 110:8057–8062
Chu CG, Friesen TL, Xu SS, Faris JD, Kolmer JA (2009) Identification of novel QTLs for seedling and adult plant leaf rust resistance in a wheat doubled haploid population. Theor Appl Genet 119:263–269
Cui F, Zhang N, Fan X, Zhang W, Zhao C, Yang L, Pan R, Chen M, Han J, Zhao X, Ji J, Tong Y, Zhang H, Jia J, Zhao G, Li J (2017) Utilization of a Wheat660K SNP array-derived high-density genetic map for high-resolution mapping of a major QTL for kernel number. Sci Rep 7:3788
Dyck PL (1977) Genetics of leaf rust reaction in three introductions of common wheat. Can J Genet Cytol 19:711–716
Gao F, Wen W, Liu J, Rasheed A, Yin G, Xia X, Wu X, He Z (2015) Genome-wide linkage mapping of QTL for yield components, plant height and yield-related physiological traits in the Chinese wheat cross Zhou 8425B/Chinese Spring. Front Plant Sci 6:1099
Herrera-Foessel SA, Lagudah ES, Huerta-Espino J, Hayden MJ, Bariana HS, Singh D, Singh RP (2011) New slow-rusting leaf rust and stripe rust resistance genes Lr67 and Yr46 in wheat are pleiotropic or closely linked. Theor Appl Genet 122:239–249
Herrera-Foessel SA, Singh RP, Huerta-Espino J, Rosewarne GM, Periyannan SK, Viccars L, Calvo-Salazar V, Lan C, Lagudah ES (2012) Lr68: a new gene conferring slow rusting resistance to leaf rust in wheat. Theor Appl Genet 124:1475–1486
Hershman DE (1985) Leaf rust of wheat. Agriculture and Natural Resources Publications 41. https://uknowledge.uky.edu/anr_reports/41/. Accessed May 1985
Hiebert CW, Thomas JB, Somers DJ, McCallum BD, Fox SL (2007) Microsatellite mapping of adult-plant leaf rust resistance gene Lr22a in wheat. Theor Appl Genet 115:877–884
Huerta-Espino J, Singh RP, German S, McCallum BD, Park RF, Chen WQ, Bhardwaj SC, Goyeau H (2011) Global status of wheat leaf rust caused by Puccinia triticina. Euphytica 179:143–160
International Wheat Genome Sequencing Consortium (IWGSC) (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361:eaar7191
Jin XH, Feng GH, Liu DT, Ma HB, Zhang HY (2017) Review on genetic research on leaf rust in wheat. J Triticeae Crops 34:504–512
Johnson R (1988) Durable resistance to yellow (stripe) rust in wheat and its implications in plant breeding. In: Simmonds NW, Rajaram S (eds) Breeding strategies for resistance to the rusts of wheat. CIMMYT, Mexico
Knott DR (1989) The wheat rusts - breeding for resistance. Monographs on Theor Appl genet chapter 12. Springer, Berlin
Kolmer JA, Su Z, Bernardo A, Bai G, Chao S (2018a) Mapping and characterization of the new adult plant leaf rust resistance gene Lr77 derived from Santa Fe winter wheat. Theor Appl Genet 131:1553–1560
Kolmer JA, Bernardo A, Bai G, Hayden MJ, Chao S (2018b) Adult plant leaf rust resistance derived from Toropi wheat is conditioned by Lr78 and three minor QTL. Phytopathology 108:246–253
Kolmer JA, Garvin DF, Hayden M (2018c) Adult plant leaf rust resistance derived from the wheat landrace cultivar Americano 44d is conditioned by interaction of three QTL. Euphytica 214:59
Lan C, Liang S, Zhou X, Zhou G, Lu Q, Xia X, He Z (2010) Identification of genomic regions controlling adult-plant stripe rust resistance in Chinese landrace Pingyuan 50 through bulked segregant analysis. Phytopathology 100:313–318
Lan C, Rosewarne GM, Singh RP, Herrera-Foessel SA, Huerta-Espino J, Basnet BR, Zhang Y, Yang E (2014) QTL characterization of resistance to leaf rust and stripe rust in the spring wheat line Francolin#1. Mol Breed 34:789–803
Lan C, Hale IL, Herrera-Foessel SA, Basnet BR, Randhawa MS, Huerta-Espino J, Dubcovsky J, Singh RP (2017) Characterization and mapping of leaf rust and stripe rust resistance loci in hexaploid wheat lines UC1110 and PI610750 under Mexican environments. Front Plant Sci 8:1450
Large EC (1954) Growth stages in cereals illustration of the Feekes scale. Plant Pathol 3:128–129
Li HH, Ye GY, Wang JK (2007) A modified algorithm for the improvement of composite interval mapping. Genetics 175:361–374
Li T, Bai GH (2009) Lesion mimic associates with adult plant resistance to leaf rust infection in wheat. Theor Appl Genet 119:13–21
Li Z, Lan C, He Z, Singh RP, Rosewarne GM, Chen X, Xia X (2014) Overview and application of QTL for adult plant resistance to leaf rust and powdery mildew in wheat. Crop Sci 54:1907–1925
Lillemo M, Asalf B, Singh RP, Huerta-Espino J, Chen XM, He ZH, Bjørnstad Å (2008) The adult plant rust resistance loci Lr34/Yr18 and Lr46/Yr29 are important determinants of partial resistance to powdery mildew in bread wheat line Saar. Theor Appl Genet 116:1155–1166
Lowe I, Jankuloski L, Chao S, Chen X, See D, Dubcovsky J (2011) Mapping and validation of QTL which confer partial resistance to broadly virulent post-2000 north American races of stripe rust in hexaploid wheat. Theor Appl Genet 123:143–157
Maccaferri M, Mantovani P, Tuberosa R, DeAmbrogio E, Giuliani S, Demontis A, Massi A, Sanguineti MC (2008) A major QTL for durable leaf rust resistance widely exploited in durum wheat breeding programs maps on the distal region of chromosome arm 7BL. Theor Appl Genet 117:1225–1240
McIntosh RA, Dubcovsky J, Rogers WJ, Morris C, Appels RA, and Xia XC (2016) Catalogue of gene symbols for wheat: 2015–2016 supplement. Komugi Wheat Genet. Resour. Database. http://shigen.nig.ac.jp/wheat/komugi/genes/symbolClassList.jsp (accessed 15 Mar. 2017)
Messmer MM, Seyfarth R, Keller M, Schachermayr G, Winzeler M, Zanetti S, Feuillet C, Keller B (2000) Genetic analysis of durable leaf rust resistance in winter wheat. Theor Appl Genet 100:419–431
Peterson RF, Campbell AB, Hannah AE (1948) A diagrammatic scale for estimating rust intensity on leaves and stems of cereals. Can J Res 26:496–500
Pinto da Silva GB, Zanella CM, Martinelli JA, Chaves MS, Hiebert CW, McCallum BD, Boyd LA (2018) Quantitative trait loci conferring leaf rust resistance in hexaploid wheat. Phytopathology 108:1344–1354. https://doi.org/10.1094/Phyto-06-18-0208-RVW
Qureshi N, Bariana H, Kumran VV, Muruga S, Forrest KL, Hayden MJ, Bansal U (2018) A new leaf rust resistance gene Lr79 mapped in chromosome 3BL from the durum wheat landrace Aus26582. Theor Appl Genet 131:1091–1098
Ramirez-Gonzalez RH, Uauy C, Caccamo M (2015) PolyMarker: a fast polyploid primer design pipeline. Bioinformatics 31:2038–2039
Ren Y, Li Z, He Z, Wu L, Bai B, Lan C, Wang C, Zhou G, Zhu H, Xia X (2012) QTL mapping of adult-plant resistances to stripe rust and leaf rust in Chinese wheat cultivar Bainong 64. Theor Appl Genet 125:1253–1262
Rosewarne GM, Singh RP, Huerta-Espino J, Rebetzke GJ (2008) Quantitative trait loci for slow-rusting resistance in wheat to leaf rust and stripe rust identified with multi-environment analysis. Theor Appl Genet 116:1027–1034
Rosewarne GM, Singh RP, Huerta-Espino J, Herrera-Foessel SA, Forrest KL, Hayden MJ, Rebetzke GJ (2012) Analysis of leaf and stripe rust severities reveals pathotype changes and multiple minor QTLs associated with resistance in an Avocet×Pastor wheat population. Theor Appl Genet 124:1283–1294
Schnurbusch T, Paillard S, Schori A, Messmer M, Schachermayr G, Winzeler M, Keller B (2004) Dissection of quantitative and durable leaf rust resistance in Swiss winter wheat reveals a major resistance QTL in the Lr34 chromosomal region. Theor Appl Genet 108:477–484
Sharp PJ, Kreis M, Shewry PR, Gale MD (1988) Location of b-amylase sequence in wheat and its relatives. Theor Appl Genet 75:286–290
Singh RP, Huerta-Espino J, William HM (2005) Genetics and breeding for durable resistance to leaf and stripe rusts in wheat. Turk J Agric For 29:121–127
Singh RP, Mujeeb-Kazi A, Huerta-Espino J (1998) Lr46: a gene conferring slow-rusting resistance to leaf rust in wheat. Phytopathology 88:890–894
Singla J, Lüthi L, Wicker T, Bansal U, Krattinger SG, Keller B (2017) Characterization of Lr75: a partial, broad-spectrum leaf rust resistance gene in wheat. Theor Appl Genet 130:1–12
Soriano JM, Royo C (2015) Dissecting the genetic architecture of leaf rust resistance in wheat by QTL meta-analysis. Phytopathology 105:1585–1593
Stange M, Utz HF, Schrag TA, Melchinger AE, Würschum T (2013) High-density genotyping: an overkill for QTL mapping? Lessons learned from a case study in maize and simulations. Theor Appl Genet 126:2563–2574
Suenaga K, Singh RP, Huerta-Espino J, William HM (2003) Microsatellite markers for genes Lr34/Yr18 and other quantitative trait loci for leaf rust and stripe rust resistance in bread wheat. Phytopathology 93:881–890
Tsilo TJ, Kolmer J, Anderson JA (2014) Molecular mapping and improvement of leaf rust resistance in wheat breeding lines. Phytopathology 104:865–870
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. Heredity 93:77–78
Wang JK (2017) Quantitative genetics. Science Press, China
Wang S, Wong D, Forrest K, Allen A, Chao S, Huang BE, Maccaferri M, Salvi S, Milner SG, Cattivelli L, Mastrangelo AM, Whan A, Stephen S, Barker G, Wieseke R, Plieske J, IWGSC, Lillemo M, Mather D, Appels R, Dolferus R, Brown-Guedira G, Korol A, Akhunova AR, Feuillet C, Salse J, Morgante M, Pozniak C, Luo MC, Dvorak J, Morell M, Dubcovsky J, Ganal M, Tuberosa R, Lawley C, Mikoulitch I, Cavanagh C, Edwards KJ, Hayden M, Akhunov E (2014) Characterization of polyploid wheat genomic diversity using a high-density 90 000 single nucleotide polymorphism array. Plant Biotechnol J 12:787–796
Wen W, He Z, Gao F, Liu J, Jin H, Zhai S, Qu Y, Xia X (2017) A high-density consensus map of common wheat integrating four mapping populations scanned by the 90K SNP array. Front Plant Sci 8:1389
William HM, Singh RP, Huerta-Espino J, Palacios G, Suenaga K (2006) Characterization of genetic loci conferring adult plant resistance to leaf rust and stripe rust in spring wheat. Genome 49:977–990
William M, Singh RP, Huerta-Espino J, Islas SO, Hoisington D (2003) Molecular marker mapping of leaf rust resistance gene Lr46 and its association with stripe rust resistance gene Yr29 in wheat. Phytopathology 93:153–159
Winfield MO, Allen AM, Burridge AJ, Barker GL, Benbow HR, Wilkinson PA, Coghill J, Waterfall C, Davassi A, Scopes G, Pirani A, Webster T, Brew F, Bloor C, King J, West C, Griffiths S, King I, Bentley AR, Edwards KJ (2016) High-density SNP genotyping array for hexaploid wheat and its secondary and tertiary gene pool. Plant Biotechnol J 14:1195–1206
Xu XY, Bai GH, Carver BF, Shaner GE, Hunger RM (2005a) Mapping of QTLs prolonging the latent period of Puccinia triticina infection in wheat. Theor Appl Genet 110:244–251
Xu X, Bai G, Carver BF, Shaner GE, Hunger RM (2005b) Molecular characterization of slow leaf-rusting resistance in wheat. Crop Sci 45:758–765
Zhang L, Li Z, Lillemo M, Xia X, Liu D, Yang W, Luo J, Wang H (2009) QTL mapping for adult-plant resistance to leaf rust in CIMMYT wheat cultivar Saar. Sci Agric Sin 42:388–397
Zhang P, Qi A, Zhou Y, Xia X, He Z, Li Z, Liu D (2017) Quantitative trait loci mapping of adult-plant resistance to leaf rust in a Fundulea 900בThatcher’wheat cross. Plant Breed 136:1–7
Zhou H, Xia X, He Z, Li X, Wang C, Li Z, Liu D (2013) Molecular mapping of leaf rust resistance gene LrNJ97 in Chinese wheat line Neijiang 977671. Theor Appl Genet 126:2141–2147
Zou J, Semagn K, Iqbal M, N’Diaye A, Chen H, Asif M, Navabi A, Perez-Lara E, Pozniak C, Yang RC, Randhawa H, Spaner D (2017) Mapping QTLs controlling agronomic traits in the ‘Attila’בCDC Go’ spring wheat population under organic management using 90K SNP array. Crop Sci 57:365–377
Acknowledgments
We would like to thank Prof. R. A. McIntosh at Plant Breeding Institute, University of Sydney, for the critical review of this manuscript.
Funding
This study was supported by the National Natural Science Foundation of China (31361140367, 31571662, and 31601299).
Author information
Authors and Affiliations
Contributions
PZ phenotyped the population in the field, prepared genotypic data, analyzed the data, and wrote the initial manuscript. CL and MA provided the phenotypic data of stripe rust and powdery mildew and SSR markers. GW modified the language. XX and ZH modified the manuscript and drafted some sections of the manuscript. ZL and DL design this experiment.
Corresponding authors
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 1.57 mb)
Rights and permissions
About this article
Cite this article
Zhang, P., Lan, C., Asad, M.A. et al. QTL mapping of adult-plant resistance to leaf rust in the Chinese landraces Pingyuan 50/Mingxian 169 using the wheat 55K SNP array. Mol Breeding 39, 98 (2019). https://doi.org/10.1007/s11032-019-1004-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-019-1004-5