Abstract
The improvement of rice leaf morphology is an important goal of rice plant breeding. Studying its molecular development mechanism will facilitate the breeding of a maximally efficient plant type. To further study the developmental mechanism of rice leaves, a dwarf broad-leaf mutant, g44, was obtained by ethyl methanesulfonate mutagenesis of the rice restorer line Jinhui10, and genetic analysis indicated that the trait was controlled by a pair of recessive nuclear gene. The locus was fine mapped on chromosome 6 with 16-kb physical distance. Sequencing revealed that an A-T base substitution occurred in the mutant, resulting in the early termination of LOC_Os06g03710/DLT translation in g44 and yielding only one residual peptide containing 558 amino acids. The phenotypic and sequencing results of the complementary transgenic plants and the allelic mutant g44-1 further confirmed this result. Mutant plants grown under field conditions showed dwarfing, a leaf colour of dark green, leaf widening, and grain enlargement. Paraffin section and cryosection analysis showed that the increase in the number of large vascular bundles and small vascular bundles and the increase in the spacing between adjacent small vascular bundles were the main reasons for the broadening of mutant leaves. In addition, the increase in thickness of the mesophyll cell layer may be the main reason for the increase in leaf colour darkness and net photosynthetic rate. The qPCR results preliminarily predicted that the increase in the number of mutant cells was caused by an enhanced cell division ability, and the DLT/G44 gene was proposed to participate in the regulation of cell division by suppressing the expression of cyclin-related genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Rice is an extremely important food crop. To ensure adequate supply in the face of increasing populations, high yield and superhigh yield are ongoing desirable traits in rice cultivation and breeding. As the main site of photosynthesis in rice, the products of photosynthesis account for more than 90% of the total amount of rice carbohydrates (Li et al. 1998). Increasing leaf area index and efficiency in the use of light energy are great significance for improving yield and quality in rice, with high-efficiency breeding also playing an important role.
In recent decades, with the development of molecular markers and the perfection of map-based cloning technology, several genes related to rice dwarfing and leaf morphological development have been identified; more than 70 genes for rice dwarfing have been reported so far. On the basis of the order in which the genes were identified, they were numbered consecutively. By 2015, the dwarf d series had been expanded to d62, and the semidwarf sd series had been expanded to sd8, in which some dwarf and semidwarf genes are identical genes or alleles, such as d6 and d34, d10, d15 and d16, d11 and d8, sd1 and d47 (Wang et al. 2009). Some genes, such as d41, d43, d46 and d48, have been deleted due to a lack of original materials (Aquino and Jennings Peter 1966). In addition, the linkage positions of 28 dwarf genes have been identified, and they involve nine linkage groups. However, most of their agronomic traits are poor and only a few recessive dwarf genes have been used in production trials, for example, sd1. Due to widespread use of the same dwarf gene, there is a great risk of poor genetic background. The semidwarf gene SD1 material also has a number of shortcomings, such as poor drought resistance and low photosynthetic efficiency (Ye et al. 2015). In consequence, the study of dwarf mutants is still of particular importance. To date, researchers in various countries have made great progress in research on leaf shape regulation in rice, but the interactions between leaf-shaped regulatory genes are still unclear, and the regulatory mechanism of leaf development in rice still needs further study. There are many narrow-leaf mutants identified at home and abroad, such as nal1, nal7, nal9, nrl1, nal3 and nal2. In nal1, the leaves are narrowed and the number of mutant vascular bundles is reduced, with a changed arrangement. In addition, the number of internodes is decreased, which affects the cell division of rice internodes and results in the dwarfing of the plant (Jiang et al. 2015; Zhang et al. 2014; Qi et al. 2008). The leaf width of nal7 is significantly reduced compared to wild type, the NAL7 locus encodes a flavin-containing monooxygenase and inspection of a structural model of NAL7 suggests that the mutation results in an inactive enzyme (Fujino et al. 2008). The leaves of the mutant nal9 are narrowed throughout the whole growth period, and the leaves at the seedling stage are light green. This mutant also shows a high plant height, small spikes and increased tillering (Li et al. 2013). The nrl1 mutant shows a narrowing of the leaves, nal2 and nal3 single mutant leave phenotypes are normal, but the double mutant leaves are narrow and coiled and NAL2 and NAL3 are paralogous, encoding the same OsWOX3A transcriptional activator (Cho et al. 2016; Cho et al. 2013).
Leaf width affects the growth of individual rice plants and the yield per unit area by affecting the light-receiving area and use of light energy. The appropriate leaf width can improve the blade’s erectness, adjust the light-receiving state, increase the leaf area index and increase the photosynthetic product. The mechanism for regulating leaf width in rice is mainly based on broad-leaf mutants. However, there are few reports on the study of broad-leaf mutants in rice. DRL1 encodes a cellulose-like synthase OsCSLD4 and is a major QTL for controlling plant height and leaf width in rice (Ding et al. 2015). The leaves of d1 mutant became shorter and wider, and the plants appeared dwarf (Sun et al. 2014). Overexpression of Osa-miR319b leads to leaf widening and delayed development (Zhang et al. 2016). In the study reported herein, a dwarf broad-leaf mutant, g44, was found in the mutant library of Jinhui 10, an ethyl methanesulfonate (EMS)-mutagenized rice restorer line in the Rice Research Institute of Southwest University. The study identified phenotypes of g44, analysed photosynthetic pigment content, determined photosynthetic parameters and conducted cytological analysis, genetic analysis and map-based cloning, as well as validation and functional analysis of the candidate gene G44. The study enriched the study of broad-leaf mutants and provided materials for further progress in the study of leaf development.
Materials and methods
Plant materials and treatments
The g44 was derived from ethyl methanesulfonate (EMS) mutagenesis of the Indica rice restorer line Jinhui 10, and the allelic mutant g44-1 was derived from the Indica rice maintainer line Xinong 1B, which was grown continuously through multiple generations and was stably inherited. We crossed the Indica rice restorer line ‘Xinong 1B’ and g44. From this cross, the F1 and F2 populations were used for genetic analysis, and the F2 population was used for gene mapping.
Trait measurements
Starting from the heart one leaf stage, the leaf width was measured at different leaf ages until the plants were mature. At the heading stage, we counted the number of large vascular bundles and small vascular bundles, and the number of small vascular bundles between adjacent large vascular bundles in wild-type and g44 inverted leaves. At maturity, 10 plants each from the centre of the wild type and g44 were harvested individually, and the following traits were measured: length of the internode, the length and width of the upper three leaves, the plant height, effective panicle, panicle length, primary branch number, secondary branch number, grain number per panicle, seed setting rate, grain length, grain width and 1000-grain weight.
Determination of photosynthetic pigment content and analysis of photosynthetic parameters
Plants were tested in the heading period. Leaf samples were collected between 7:00 and 8:30 am. The flag leaves, the second upper leaves and the third upper leaves of the wild-type and g44 plants were sampled and used by the following method: the midrib was removed, cut into pieces and approximately 0.05 g weighed out, then placed in a clean centrifuge tube with 20 mL ethanol/acetone (1:1) mixed solution and allowed to stand at room temperature for 24 h. The absorbances of the extracts were measured at 663 nm, 470 nm and 645 nm, and the photosynthetic pigment content in each sample was calculated. The procedure was repeated three times for each sample. Other measurements were taken on sunny days from 9:00 am to 11:00 am. The net photosynthetic rate (Pn), transpiration rate (Tr), stomatal conductance (Gs) and intercellular CO2 concentration (Ci) of the flag leaves, the second upper leaves and the third upper leaves of wild type and g44 were measured using a Li-6400 portable photosynthetic apparatus. Water-use efficiency (WUE) was also calculated.
Cytological observation of leaves
At the heading stage, the fresh mature leaves of the wild type and g44 (unified as the middle of the flag leaf) were taken for paraffin section analysis (Sang et al. 2012). After being fixed with FAA fixation, they were dehydrated with a certain concentration of ethanol, make transparent by xylene, embedded in paraffin and sliced. The slices were subjected to saffron and solid green staining, and the final neutral resin was sealed. Then, they were observed under the microscope and photographed. Fresh rice leaves from wild type and g44 were also selected for cryosection. Samples from the middle part of the blade were treated with OTC frozen section embedding solution before being placed quickly in a − 20 °C refrigerator and frozen. After being sliced with a cryostat, the sections were thawed on a glass slide. The autofluorescence of the leaf-tissue components was observed under a NIKON E6000 microscope to analyse the morphology of the cells. Fresh wild-type and mutant leaves were observed under scanning electron microscopy.
Genetic analysis and map-based cloning of G44
The ten wild-type and mutant leaves in the F2 generation isolates were selected and mixed, and the modified CTAB method (Porebski et al. 1997) was used to extract the genomic DNA of the normal pool and g44 pool for linkage primer selection. Then, g44 plants in the F2 generation segregating population were selected according to a 3:1 separation ratio, and the genomic DNA was extracted for gene mapping. The total PCR reaction system contained 25 μL, comprising 2.5 μL of 10 × PCR buffer, 1.3 μL of 25 mmol L−1 MgCl2, 1.0 μL of 2.5 mmol L−1 dNTPs, 16.0 μL of ddH2O, 2.0 μL of 10 μmol L−1 primers, 2.0 μL of template DNA and 0.2 μL of 5 U μL−1 Taq DNA polymerase. The PCR program was run using 94 °C predenaturation for 5 min and then 35 cycles of 94 °C denaturation for 30 s, 55 °C annealing for 30 s, 72 °C extension for 1 min and 72 °C extension for 10 min. The PCR product was then subjected to electrophoresis using 10% nondenaturing polyacrylamide gel and observed after rapid silver staining.
Using the information about the DNA sequence obtained from the Gramene and NCBI websites, primers were designed to amplify the predicted genes of wild type and g44. The amplified products were recovered and sent for sequencing. Sequence splicing and alignment were performed on Vector NTI Advance 10.1, and BLAST was performed on Gramene and NCBI. All primer information used in this paper is given in Table S4.
Generation of transgenic plants complementing and overexpressing G44
The complementary fragment, the Pcambia1301 vector and the overexpressed fragment and the PTCK303 vector were double-digested. After the gel was recovered, the target fragment and the corresponding vector were ligated with T4 ligase and transferred into an Escherichia coli culture, then preserving the bacterial liquid containing the target gene. All primer information used in this paper is given in Table S4.
Quantitative real-time PCR
Wild-type leaves were taken at the seedling stage, the tillering stage and the heading stage, the target genes were analysed for spatiotemporal expression. In addition, at the tillering stage, the relative expression of sink leaves, whole leaves, and sheaths were analysed. The leaves were placed in a nuclease-free 2-mL centrifuge tube and frozen in liquid nitrogen. RNA was extracted and reverse-transcribed into cDNA, using the kit provided by Tiangen Biotechnology (Beijing) Co., Ltd. QPCR quantitative analysis was then carried out using Takara SYBR Green fluorescent dye. All primer information used in this paper is given in Table S4.
Results
The g44 is a dwarf and broad-leaf mutant
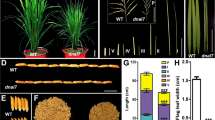
Under field-cultivation conditions, mutant g44 showed dwarfing and a darker green colour than the wild type, and these traits lasted until maturity (Fig. 1a, b). At maturity, mutant g44 showed a shorter length in the flag leaf, the second upper leaf, the third upper leaf, the internodes and the leaves were wider compared to wild type. In addition, it was found that some areas of the blade had more severe folds (Fig. 1c, d) and the grain became wider in mutant (Fig. 1e). At the mature stage, the agronomic traits of wild type and the mutants were investigated and analysed. The results showed that the height of the mutant was only 79 ± 2.16 cm, which was 31.89% shorter than the 116 ± 3.25 cm of wild type, and the mutant plant height was significantly shorter. The panicle length of wild type was 25.57 ± 3.06 cm, while the length of g44 was only 19.18 ± 2.14 cm, which was significantly shorter. In addition, the number of effective panicles, the number of secondary branches, the number of spikelet and the seed setting rate of the mutants were significantly lower than wild type. However, the number of primary branches was not significantly different in the two plant types. There was no difference in the grain length between wild type and the g44, but the grain width of the mutant was 3.53 ± 0.06 mm, which was significantly wider than the 2.96 ± 0.06 mm found in wild type. The grain was broadened, so that the thousand weight of the mutant g44 was 29.41. ±0.77 g, which was 9.6% higher than the 26.83 ± 1.02 g found in wild type. This was a highly significant difference (Table S1).
The G44 is a new allele of DLT
The polymorphic marker was used to analyse the normal gene pool and mutant gene pool. The regulatory gene was located within the range of 16 kb of the physical distance of chromosome 6, and the interval contained three annotated genes (Fig. 2c). Sequencing revealed that the LOC_Os06g03710 gene sequence of the wild type was A at the 1675th base at the coding frame, and the g44 was T. This A-T base substitution resulted in the early termination of LOC_Os06g03710 translation in g44, yielding a residual peptide containing only 558 amino acids, while the wild-type protein contained 617 amino acids (Fig. 2d). Therefore, LOC_Os06g03710 was identified initially as a candidate gene. Sequencing results indicate that G44 was a new allele of DLT. In order to determine whether the candidate gene LOC_Os06g03710 is DLT/G44, the full-length genomic sequence of LOC_Os06g03710 was cloned from the wild type to construct a complementary vector, which was introduced into the mutant, finally we obtained ten complementary lines and the indicators of all complementary transgenic plants were restored, close to those of the wild type (Fig. 2e–g). Thus, the LOC_Os06g03710 is the DLT/G44 gene; in addition, a overexpression vector was constructed, we also obtained ten overexpression lines, GUS staining of plants showed that whole plant could be dyed. But all overexpressed transgenic plants showed lethality (Fig. S1a, b).
Map clone of DLT/G44 gene. (a–d) Fine mapping of DLT/G44. (e–g) Phenotypital analysis of the wild-type, mutant and complementary transgenic plant. (h–i) Analysis of expression pattern of DLT/G44. ST: seeding stages. TT tillering stage, FT heading stage, SL sink leaves at tillering stage, FLS leaf sheaths at tillering stage, FL leaves at tillering stage. The experiments are biologically repeated three times and the data are presented as means ± SE. (e–g) Bar = 15 cm
We used qPCR to study the expression pattern of DLT/G44 gene in the fully-expanded leaves of seedling, tillering, and heading stages. The results showed that the DLT/G44 gene had the highest expression at the seedling stage, but the lowest in the tillering stage (Fig. 2h). The expression level of the DLT/G44 gene of the wild type in the sink leaves, leaves, and leaf sheaths of the tillering stage were analysed. The results showed that the DLT/G44 gene was expressed at very low levels and the gene was hardly expressed in the sink leaves, but the leaves and leaf sheaths showed expression of this gene (Fig. 2i).
A mutant similar to the dlt/g44 phenotype was identified in the mutant library of the EMS mutagenized rice maintainer Xinong1B of the Rice Research Institute of Southwest University and tentatively named g44-1. Compared with the wild-type Xinong1B, the g44-1 plant height was significantly shorter, the internodes were significantly shorter (Fig. S2a, b, e; Table S2) and the leaves were significantly changed, showing a wider, shorter shape and an increased number of vascular bundles (Fig. S2d). The g44-1 plant also showed an increased number of primary branches and secondary branches (Fig. S2f; Table S3), a widening of the grain width and an increased 1000-grain weight (Fig. S2g; Table S3). These traits in the phenotype were consistent with those in that of the mutant g44. Sequencing revealed that a guanine (G) was inserted between the 822th base and the 823th base of the exon of LOC_Os06g03710, and a base substitution of C-T had occurred at the 888th base. These changes resulted in a frameshift mutation from the 275th codon up to the 297th codon, which was a stop codon, causing the translation to terminate prematurely (Fig. S3a, b). The results indicate that the phenotypes of dlt/g44 and g44-1 are caused by mutations in LOC_Os06g03710.
The number of cells increased leads to dlt/g44 leaves become wider and thicker
Compared to the wild type, the leaves of the dlt/g44 broadened throughout the growth period and reached significant or highly significant levels of difference. At maturity, the leaf width of the flag leaf, the second upper leaves and the third upper leaf of the dlt/g44 were 29.6 mm, 25.8 mm and 24.2 mm, respectively, which were significantly wider than the wild-type leaf widths of 22.3 mm, 19.41 mm and 15.4 mm (Fig. 3a–c). Statistical analysis of the number of vascular bundles in the leaves at the heading stage showed that wild type had an average of 7.25 large vascular bundles, whereas dlt/g44 had an average of 7.88, which was significantly higher (Fig. S4e). There were 5.13 small vascular bundles between the two adjacent vascular bundles in the flag leaf of wild type, and dlt/g44 had an average of 6.73. The total number of small vascular bundles of dlt/g44 was 38.77% than in the wild type. Both measures reached a very significant level of difference (Fig. S4d, f). Scanning electron microscopy also showed that the spacing between two adjacent small vascular bundles in the mutant was also significantly higher than in the wild type (Fig. S4a, b, f).
Wild-type and mutant dlt/g44 widened leaves. a Wild-type and mutant flag leaves. b Enlarged view of the flag leaf of the wild type and mutant (middle). c Leaf width of wild-type and mutant whole growth period. An asterisk indicates significant difference at the p < 0.05 by t test. Asterisks indicate significant difference level at the p < 0.01 by t test
To explore further the explanation for the broadening of dlt/g44 leaves, the darkening of the leaf colour and the maintenance of the same chlorophyll content, the wild type and dlt/g44 were observed using paraffin section and frozen section. The paraffin section of the tillering leaves showed that the whole of dlt/g44 leaves was larger, and both the large vascular bundles and the small vasculars bundle were more numerous than wild type (Fig. 4a, b). The enlarged view of the midrib showed that dlt/g44 showed many small vascular bundles on the paraxial surface, whereas wild type had almost none, which caused the midrib of dlt/g44 to be wider than wild type (Fig. 4-1, 2). On both sides of the midrib, the mutant dlt/g44 had a larger vascular bundle than wild type, which further increased the width of the blade (Fig. 4-3, 4).
Cytological observation of the mutant dlt/g44. a, b Cross section of paraffin sections in the middle of wild-type and mutant (dlt/g44) leaves (tillering stage). 1, 2 Magnified view of black box in a and b. 3, 4 A magnified view of the red box in a and b. c-h, Cross section of the middle leaf blades obsearved by frozen section. 5, 6 Magnified view of the white box in e and f. g, h Wild-type and mutant (dlt/g44) large vascular bundle (heading period). i Statistics of cell layers above and below the small vascular bundle. j Statistics of cell numbers between adjacent small vascular bundles. An asterisk indicates significant difference at the p < 0.05 by t test. Asterisks indicate significant difference level at the p < 0.01 by t test. a, b Bar = 0.1 mm. 1–6, e–h Bar = 20 μm. c, d Bar = 0.2 mm
The frozen section of the leaves at the heading stage showed that the leaves of the mutant dlt/g44 were thicker than wild type, and the small vascular bundles between the large vascular bundles were more numerous (Fig. 4c,d). In addition, the spacing between the small vascular bundles was higher in the mutant and the small vascular bundle was larger than in the wild type (Fig. 4e,f). The enlarged view of the mesophyll showed that there was no significant difference in the cell size of the mesophyll cells between dlt/g44 and wild type, but the mesophyll cells of wild type only had about four layers, while dlt/g44 had up to seven. Therefore, the number of mesophyll cell layers of the mutant dlt/g44 was increased significantly (Fig. 4-5, 6, i). The number of cells between adjacent small vascular bundles in each plant type was compared. There was an average of ten cells between the tube bundles in the wild type, and there were as many as thirteen in dlt/g44 (Fig. 4j). The number of cells in the large vascular bundles of the mutants was significantly higher than in the wild type, so the large vascular bundle size of dlt/g44 was larger than that of the wild type (Fig. 4g, h).
DLT/G44 was proposed to participate in the regulation of cell division by suppressing the expression of cyclin-related genes
The number of mesophyll cells, vascular bundle sheath cells, xylem cells, parenchyma cells and other cells in the mutant was increased significantly, which points to the influence of cell division. To determine further whether the mutation of the DLT/G44 gene affects the expression of cyclin related genes, qPCR analysis was conducted. The results show that the expression of CYCA3; 2, CYCD4; 1, CYCP4; 1, CYCT1; 1 and CYCF3; 1 was significantly or very significantly upregulated, including an increase of approximately nine times for CYCD4; 1 and an increase of approximately four times for CYCF3; 1 and CYCT1; 1 (Fig. 5).
The photosynthetic rate of the mutant dlt/g44 is enhanced
Under field conditions, the leaf colour of the mutant dlt/g44 was observed, by the naked eye, to be deeper than that of the wild type throughout the whole growth period. To determine further whether there were differences in photosynthetic pigment content between the leaves of the two plants, the content was determined in three functional leaves of the wild type and dlt/g44 at the heading stage: the flag leaf, the second upper leaf and the third upper leaf. The results showed that there was no significant difference in the chlorophyll a (Chla), chlorophyll b (Chlb) and total chlorophyll content (Chla+Chlb) of Jinhui10 and the mutant dlt/g44 (Fig. 6a–c), with the exception that the carotenoid content (Car) of the mutant dlt/g44 functional leaves was significantly or very significantly lower (Fig. 6d). It was conjectured that another factor was responsible for the deeper leaf colour of mutant dlt/g44.
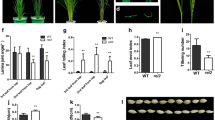
Analysis of photosynthetic pigment content and photosynthetic efficiency of the wild type and the mutant dlt/g44 at the heading stage. a Chlorophyll a. b Chlorophyll b. c Total amount of chlorophyll a and chlorophyll b. d Carotenoid. e Net photosynthetic rate (Pn). f stomatal conductance (Gs). g Transpiration rate (Tr). h Water use efficiency (WUE). (FL is the flag leaf, SL is the second upper leaves, TL is the third upper leaves). An asterisk indicates significant difference at the p < 0.05 by t test. Asterisks indicate significant difference level at the p < 0.01 by t test
Leaf photosynthetic parameters in wild type and dlt/g44 were determined at the heading stage using a Li-6400 portable photosynthetic apparatus. The results showed that mutant dlt/g44 stomatal conductance (Gs) and transpiration rate (Tr) declined significantly compared with Jinhui10 (Fig. 6f, g). In addition, the mutant dlt/g44 net photosynthetic rate (Pn) and water use efficiency (WUE) showed a highly significant improvement, compared with wild type (Fig. 6e, h).
Discussion
In the study reported herein, the mutant dlt/g44 showed plant dwarfing, leaf widening and thickening, leaf erectness, dark green leaf colour and tiller reduction. The regulatory gene was located precisely in the 16-kb physical range on chromosome 6. Sequencing and other tests showed that LOC_Os06g03710 was the DLT gene. It may be concluded that dlt/g44 is a new allele of dlt with functional deletions. Studies on the semidwarf low-tillering mutant dlt have found that DLT encodes a GRAS family protein that is unique to plants and is composed of 617 amino acids. This new member of the protein family is a key positive regulator in the BR signalling pathway of rice and is involved in the feedback inhibition of BR synthesis gene expression (Tong et al. 2009; Hirano et al. 2017). Further studies have found that DLT acts downstream of the BR response, which is regulated by GSK2 and is a direct downstream target of GSK2. GSK2 is a key negative regulator of BR signal transduction and regulates the protein level and activity of DLT by interacting directly with DLT and phosphorylating it (Qiao et al. 2017; Tong et al. 2012). Another allele mutant, d62, showed dwarfing, leaf widening and dark green colouration; these are typical characteristics of ga-related dwarfing mutants. In the mutant d62, the expression level of genes related to the GA metabolic pathway is upregulated significantly, which suggests that d62 genes regulate GA metabolic pathways (Li et al. 2010). The dlt/g44 agrees with previous reports of allelic mutants with similar phenotypes. All show the traits of dwarfism and increased blade width, but in the dlt/g44, the adaxial surface of the blade contains a lot more small vascular bundles than the wild type. In addition to the increased number of vascular bundles, compared with the wild type, the dlt/g44 blade is more upright, and the net photosynthetic rate and water use efficiency of the mutant are significantly higher.
Compared with the wild type, the vascular bundles of the dlt/g44 mutants increased in number significantly. In the paraffin section analysis of the tillering leaves, it was found that dlt/g44 produced many small vascular bundles on the paraxial surface of the midrib. Vascular bundles also occurred inside the midrib, resulting in a larger midline in dlt/g44 compared to the wild type, which increased the blade width to some extent. In addition, the number of large vascular bundles and small vascular bundles in dlt/g44 was significantly higher, the number of mesophyll cell layers in dlt/g44 leaves increased and the number of cells in each leaf structure increased. However, the cells did not differ in size, except for an irregular arrangement, which indicates that the larger dlt/g44 leaves were not caused by changes in cell size. There are two possible reasons for the increase in the number of dlt/g44 cells. The first is that during the initiation of leaf primordium, the frequency of primordial cell division is accelerated, while the critical point of division and differentiation is unchanged, which results in an increase in the number of primordial cells. The second possible cause is that the frequency of primordial basal cell division is unchanged, but the critical point of division and differentiation is delayed, which results in a longer cell division time that leads to an increase in the number of cells. The cell cycle can generally be divided into four periods: G1, S, G2 and M, and is mainly regulated by cyclins. OsCYCP1;1 is expressed in all tissues of rice and plays a negative regulatory role in the rice phosphate starvation signalling pathway (Deng et al. 2014). CycB1;1 affects embryo development by regulating mitosis (Guo et al. 2010). CYCP4;1 affects rice plant type by regulating cell division (La et al. 2006; Sun et al. 2015). In the qPCR analysis results, the gene expression of CYCA3;2 (La et al. 2006; Yu et al. 2003), CYCD4;1 (La et al. 2006; Kono et al. 2003), CYCP4;1, CYCT1;4 (La et al. 2006) and CYCF3;1 (La et al. 2006) was found to be upregulated significantly in the mutant and CYCD4; 1 showed the most significant upregulation. Because CYCD4; 1 encodes cyclin, this upregulation indicates that the mutation of the DLT/G44 gene affects CYCD4;1 expression. CYCD3;1 promotes cell division ability, so it may be conjectured that the increase in the dlt/g44 mesophyll cell layer is caused by an increased capacity for cell division, but in the dlt/g44 mutant, it is uncertain whether the critical point of division and differentiation changes. Therefore, it is considered that an increased capacity for cell division is one of the main reasons for the increase in the number of dlt/g44 leaf cells and that the DLT/G44 gene may participate in the regulation of cell division by inhibiting the expression of CYCD3;1.
Rice is an important food crop, and experts in areas such as physiology, cultivation, breeding, genetics and nutrition attach great importance to high-efficiency breeding. High-efficiency breeding should be based on shaping the ideal rice plant type, so that individual plants can manifest improved light-receiving patterns, and can maximize the efficient use of light energy, thereby increasing the yield of dry matter. Considering leaves as a source of rice and grain as a sink of rice, an increase in dry matter storage can be achieved by increasing the source (Kozak et al. 2007; Yoon et al. 2006; Thomson et al. 2003; Yue et al. 2006). To achieve a greater storage capacity, it is necessary to strengthen the supply capacity of the source. The early construction of the largest leaf area promotes photoautotrophic capacity and promotes plant growth, thereby increasing the accumulation of biomass (Yoshida 1981; Hebert 1990). Under certain environmental conditions, the length, width and thickness of the leaves are important factors that influence the synthesis of photosynthetic products in individual leaves. The morphology and location of the leaves are also important factors in the composition of plant types (Ming et al. 2012). Therefore, one of the important goals when breeding types of rice plant is to improve leaf morphology. The improvement of leaf traits in high-quality rice plants should be focused on achieving leaves that are short and erect and can be widened appropriately, while the second and the third upper leaves can be lengthened and widened (Jang et al. 2017). In short, under the condition of ensuring certain lodging resistance and mutual shading, increasing the leaf area appropriately can also increase the effective photosynthetic area, accordingly, to achieve increased production. The dwarf broad leaf mutant dlt/g44 found in this study was dwarfed and showed broadened leaves from the seedling stage, and these traits continued until maturity. Plant dwarfing is of great significance for high-light efficiency breeding, and the proper widening of leaves also plays a role in increasing yield.
References
Aquino RC, Jennings Peter R (1966) Inheritance and significance of dwarfism in an indica rice variety. Crop Sci 966(6):551–554
Cho SH, Yoo SC, Zhang HT, Pandeya D, Koh HJ, Hwang JY, Kim GT, Paek NC (2013) The rice narrow leaf2 and narrow leaf3 loci encode WUSCHEL-related homeobox 3A (OsWOX3A) and function in leaf, spikelet, tiller and lateral root development. New Phytol 198(4):1071–1084
Cho SH, Kang KY, Lee SH, Lee IJ, Paek NC (2016) OsWOX3A is involved in negative feedback regulation of the gibberellic acid biosynthetic pathway in rice (Oryza sativa). J Exp Bot 67(6):1677–1687
Deng MJ, Hu B, Xu L, Liu Y, Wang F, Zhao HY, Wei XJ, Wang JC, Yi KK (2014) OsCYCP1;1, a PHO80 homologous protein, negatively regulates phosphate starvation signaling in the roots of rice (Oryza sativa L.). Plant Mol Biol 86(6):655–669
Ding ZQ, Lin ZF, Li Q, Wu H, Xiang CY, Wang JF (2015) DNL1, encodes cellulose synthase-like D4, is a major QTL for plant height and leaf width in rice (Oryza sativa L.). Biochem Biophys Res Commun 457(2):133–140
Fujino K, Matsuda Y, Ozawa K, Nishimura T, Koshiba T, Marco WF, Sekiguchi H (2008) NARROW LEAF 7 controls leaf shape mediated by auxin in rice. Mol Gen Genomics 279(5):499–507
Guo J, Wang F, Song J, Sun W, Zhang XS (2010) The expression of Orysa;CycB1;1 is essential for endosperm formation and causes embryo enlargement in rice. Planta 231(2):293–303
Hebert Y (1990) Genetic variation of the rate of leaf appearance in maize: possible yield prediction at the early stage. Euphytica 46(3):237–247
Hirano K, Yoshida H, Aya K, Kawamura M, Hayashi M, Hobo T, Miyako UT, Matsuoka M (2017) SMALL ORGAN SIZE 1 and SMALL ORGAN SIZE 2/DWARF AND LOW-TILLERING form a complex to integrate auxin and brassinosteroid signaling in rice. Mol Plant 10(4):590–604
Jang S, An G, Li HY (2017) Rice leaf angle and grain size are affected by the OsBUL1 transcriptional activator complex. Plant Physiol 173(1):16.01653
Jiang D, Fang JJ, Lou LM, Zhao JF, Yuan SJ, Yin L, Sun W, Peng LX, Guo BT, Li XY (2015) Characterization of a null allelic mutant of the rice NAL1 gene reveals its role in regulating cell division. PLoS One 10(2):e0118169
Kono A, Umedahara C, Lee J, Ito M, Uchimiya H, Umeda M (2003) Arabidopsis D-type cyclin CYCD4;1 is a novel cyclin partner of B2-type cyclin-dependent kinase. Plant Physiol 132(3):1315–1321
Kozak M, Singh PK, Verma MR, Hore DK (2007) Causal mechanism for determination of grain yield and milling quality of lowland rice. Field Crop Res 102(3):178–184
La HG, Li J, Ji ZD, Cheng YJ, Li XL, Jiang SY, Venkatesh PN, Ramachandran S (2006) Genome-wide analysis of cyclin family in rice (Oryza Sativa L.). Mol Gen Genomics 275(4):374–386
Li Z, Pinson SRM, Stansel JW, Paterson AH (1998) Genetic dissection of the source-sink relationship affecting fecundity and yield in rice (shape Oryza sativa L.). Mol Breed 4(5):419–426
Li WQ, Wu JG, Weng SL, Zhang YJ, Zhang DP, Shi CH (2010) Identification and characterization of dwarf 62, a loss-of-function mutation in DLT/OsGRAS-32 affecting gibberellin metabolism in rice. Planta 232(6):1383–1396
Li W, Wu C, Hu GC, Xing L, Qian WJ, Si HM, Sun ZX, Wang XC, Fu YP, Liu WZ (2013) Characterization and fine mapping of a novel rice narrow leaf mutant nal9. J Integr Plant Biol 55(11):1016–1025
Ming L, Chen J, Luo GN, Shao XJ, Wei SQ, Tang ZH, Sheng J, Hu PS (2012) Fine mapping of a major QTL for flag leaf width in rice, qFLW4, which might be caused by alternative splicing of NAL1. Plant Cell Rep 31:863
Porebski S, Bailey LG, Baum BR (1997) Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol Biol Report 15:8–15
Qi J, Qian Q, Bu QY, Li SY, Chen Q, Sun JQ, Liang WX, Zhou YH, Chu CC, Li XG, Ren FG, Palme K, Zhao BG, Chen JF, Chen MS, Li CY (2008) Mutation of the rice narrow leaf1 gene, which encodes a novel protein, affects vein patterning and polar auxin transport. Plant Physiol 147(4):1947–1959
Qiao SL, Sun SY, Wang LL, Wu ZH, Li CX, Li XM, Wang T, Leng LN, Tian WS, Lu TG, Wang XL (2017) The RLA1/SMOS1 transcription factor functions with OsBZR1 to regulate brassinosteroid signaling and rice architecture. Plant Cell 29(2):292–309
Sang XC, Li YF, Luo ZK, Ren DY, Fang LK, Wang N, Zhao FM, Ling YH, Yang ZL, Liu YS, He GH (2012) CHIMERIC FLORAL ORGANS1, encoding a monocot-specific MADS box protein, regulates floral organ identity in rice. Plant Physiol 160:788–807
Sun HY, Qian Q, Wu K, Luo JJ, Wang SS, Zhang CW, Ma YF, Liu Q, Huang XZ, Yuan QB, Han RX, Zhao M, Dong GJ, Guo LB, Zhu XD, Gou ZH, Wang W, Wu YJ, Lin HX, Fu XD (2014) Heterotrimeric G proteins regulate nitrogen-use efficiency in rice. Nat Genet 46(4):652–656
Sun SY, Chen DH, Li XM, Qiao SL, Shi CN, Li CX, Shen HY, Wang XL (2015) Brassinosteroid signaling regulates leaf erectness in Oryza sativa via the control of a specific U-type cyclin and cell proliferation. Dev Cell 34(2):220–228
Thomson MJ, Tai TH, McClung AM, Lai XH, Hinga ME, Lobos KB, Xu Y, Martinez CP, McCouch SR (2003) Mapping quantitative trait loci for yield, yield components and morphological traits in an advanced backcross population between Oryza rufipogon and the Oryza sativa cultivar Jefferson. Theor Appl Genet 107(3):479–493
Tong HN, Jin Y, Liu WB, Li F, Fang J, Yin YH, Qian Q, Zhu LH, Chu CC (2009) DWARF AND LOW-TILLERING, a new member of the GRAS family, plays positive roles in brassinosteroid signaling in rice. Plant J 58(5):803–816
Tong HG, Liu LC, Jin Y, Du L, Yin YH, Qian Q, Zhu LH, Chu CC (2012) DWARF AND LOW-TILLERING acts as a direct downstream target of a GSK3/SHAGGY-like kinase to mediate brassinosteroid responses in rice. Plant Cell 24(6):2562–2577
Wang JY, Nakazaki T, Chen SQ, Chen WF, Saito H, Tsukiyama T, Okumoto Y, Xu ZJ, Tanisaka T (2009) Identification and characterization of the erect-pose panicle gene EP conferring high grain yield in rice ( Oryza sativa L.). Theor Appl Genet 119(1):85–91
Ye H, Feng JH, Zhang LH, Zhang JF, Muhamad S, Mispan CZQ, Beighley DH, Yang JC, Gu XY (2015) Map-based cloning of seed dormancy1-2 identified a gibberellin synthesis gene regulating the development of endosperm-imposed dormancy in rice. Plant Physiol 169(3):2152–2165
Yoon DB, Kang KH, Kim HJ, Ju HG, Kwon SJ, Suh JP, Jeong OY, Ahn SN (2006) Mapping quantitative trait loci for yield components and morphological traits in an advanced backcross population between Oryza grandiglumis and the O. sativa japonica cultivar Hwaseongbyeo. Theor Appl Genet 112(6):1052–1062
Yoshida S (1981) Fundamentals of rice crop science. Int. Rice Res. Inst 1981
Yu Y, Steinmetz A, Meyer D, Brown S, Shen WH (2003) The tobacco A-type cyclin, Nicta;CYCA3;2, at the nexus of cell division and differentiation. Plant Cell 15(12):2763–2777
Yue B, Xue WY, Luo LJ, Xing YZ (2006) QTL analysis for flag leaf characteristics and their relationships with yield and yield traits in rice. Acta Genet Sin 33(9):824–832
Zhang GH, Li SY, Wang L, Ye WJ, Zeng DL, Rao YC, Peng YL, Hu J, Yang YL, Xu J, Ren DY, Gao ZY, Zhu L, Dong GJ, Hu XM, Yan MX, Guo LB, Li CY, Qian Q (2014) LSCHL4 from japonica cultivar, which is allelic to NAL1, increases yield of indica super rice 93-11. Mol Plant 7(8):1350–1364
Zhang C, Ding ZM, Wu KC, Yang L, Li Y, Yang Z, Shi S, Liu XJ, Zhao SS, Yang ZR, Wang Y, Zheng LP, We J, Du ZG, Zhang AH, Miao HQ, Li Y, Wu ZJ, Wu JG (2016) Suppression of jasmonic acid-mediated defense by viral-inducible microRNA319 facilitates virus infection in rice. Mol Plant 9(9):1302–1314
Funding
This work was supported by Chongqing postgraduate research and innovation project (CYS18082), Chongqing Science and Technology Key Project (cstc2016shms-ztzx80007; cstc2017shms-xdny80057) and the Fundamental Research Funds for the Central Universities (XDJK20140010).
Author information
Authors and Affiliations
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOC 2766 kb)
Rights and permissions
About this article
Cite this article
Xie, J., Liao, H., Wang, X. et al. DLT/OsGRAS-32, regulating leaf width and thickness by controlling cell number in Oryza sativa. Mol Breeding 39, 104 (2019). https://doi.org/10.1007/s11032-019-1003-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11032-019-1003-6