Abstract
The rapid surge in antibiotic resistance to pathogens has emerged as a grave threat to public health, globally. This multiple drug resistance (MDR) is directly linked to the high rates of morbidity and mortality worldwide due to untreated microbial infections. Therefore, it is inevitable to identify some novel, efficient, and comparatively safer antimicrobial agents to rescue the declining health index. In this regard, nanomaterials with modified structure, size, and infinity have risen as the sole source to tackle the MDR either through ameliorating the efficacy of existing drugs or by triggering entirely new bactericidal mechanisms. Out of all the nanomaterials, metals, and metal oxide nanoparticles with biopolymer-induced reduction have fetched the attention of global researchers due to their significant and promising pathogen-killing ability without any hint of resistance. The current review covers the updated molecular modes of resistance development in Gram-positive and Gram-negative bacteria, comprehensively. This review also highlighted the detailed mode of action of various metallic nanoparticles (silver, zinc, copper, titanium, and cobalt) against MDR pathogens. Moreover, this review article thoroughly discussed the correlation between the mechanisms of resistance and alternative NPs bactericidal modes for better understanding for the readers. Last but not least, toxicity analysis is also explained for safe further use.
Graphical Abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
1 Introduction
Bacterial adherence, proliferation, and infections have become momentous and consequential health and economic problems in everyday life [1,2,3]. This grave issue is posing serious threats to several growing industries like medicine, textile, food packaging, and marine transport, globally [4,5,6]. Antibiotics are fundamental in defense against bacterial infections and were thought of as a magic bullet that would selectively target the disease-causing microbes without affecting the host [7,8,9]. However, there is a continuous decrease in the remedial potential of existing antibiotics. Although antibiotic resistance is a natural response by bacteria, it is greatly accelerated due to its unregulated, over and misuse, leading to the emergence of multiple drug resistance (MDR) [10,11,12,13]: [14,15,16,17]. Antibacterial resistance is also bacteria’s natural response to the selective pressure of an antibiotic. Due to this, a drug is unable to control bacterial growth effectively and they continue to reproduce even in the presence of therapeutic levels of antibiotics [18,19,20]. Microbial resistance against multiple antibiotics is posing serious concerns to the human health index which can significantly contribute to prolonged hospital stays and affect the ratio of mortality and morbidity as a consequence of bacterial diseases [21,22,23].
It is claimed that a post-antibiotic era is imminent due to the speeded evolution of bacterial resistance and a diminished antibiotic pipeline. This would result in common infections being untreatable that were previously fixable [24,25,26]. It is reported that infections caused by MDR bacteria could result in 10 million deaths per year by 2050. This would not only influence the economy dreadfully but could also push 24 million people into extreme poverty by 2030 [27]. Further, the emergence of MDR endorses the concept of innovation, supervision of consumption, and a swift decrease in the misuse of antibiotics [28,29,30,31].
In the dilemma of resistant superbugs, nanoparticles have emerged as the sole hope to tackle the grave issue. Nanoparticles being prepared from chemical and natural reducing agents have been in action against pathogenic bacteria [32]. However, the chemically synthesized NPs pose several indirect and direct toxicological threats due to their chemical origin [33]. On the contrary, biosynthesized NPs got a global interest in various biomedical fields due to the incorporation of biocompatible and biodegradable reducing agents like bacteria, fungi, phytochemicals, and proteins, etc. [14,15,16, 34]: [35]: [14,15,16, 36]. Moreover, the application of chemically fabricated NPs is now discouraged due to their high cost, more time to synthesize, and less biocompatibility [37].
This review will explain the various mechanisms of antibiotic resistance acquisition by the bacteria. It will also narrate the various classes of antibiotics and their route for resistance (Fig. 1). Furthermore, it briefly explains the use of different nanomaterials as potential antimicrobial agents and their modes of action. Last but not least, the toxicity perspective of the biogenic NPs is also narrated in this review.
2 Types of Drug Resistance
There are three possible forms of drug resistance exhibited by bacteria that provide them with advantageous modifications to survive in hostile conditions [38,39,40].
2.1 Intrinsic Resistance
Natural or intrinsic resistance is associated with a change in the structural properties of bacterium. These are inherent characteristics and are not linked to antibiotic selective pressure. For example, the change in permeability of the outer membrane in Gram-negative bacteria makes them insusceptible to glycopeptides [41,42,43]. Other intrinsic antibiotic resistance mechanisms include up-regulation of efflux pumps, activation of drug-altering enzymes, and change in the target site of the antibiotic [43, 44] (Fig. 3).
2.2 Acquired Resistance
Acquired resistance emerges when previously susceptible bacteria gain resistance by alteration in their genetic material. Extrinsic antibiotic resistance mechanisms include mutation in existing genetic features, genetic rearrangement, or acquiring exogenous genetic material through transformation, transduction, and conjugation as shown in Fig. 2 [45,46,47]. Horizontal gene transfer (HGT) is the chief method responsible for the sharing of antibiotic resistance genes between bacteria which has been reportedly increasing due to ongoing antibiotic abuse [48, 49].
2.3 Adaptive Resistance
Adaptive resistance is exhibited as a result of epigenetic changes caused by certain environmental signals e.g., pH, stress, growth rate, levels of antibiotics, and ion concentrations (Fig. 3). However, adaptive resistance is transient and bacteria revert to the non-resistant phenotype, once the external stimulus is removed [39, 40, 50, 51].
3 Mechanisms of Antibiotic Resistance
The major mechanisms of antibiotic resistance include antibiotic inactivation, target site alterations, removal of the drug by active efflux pump, change in a metabolic pathway, alteration of bacterial membrane permeability, and biofilm formation through quorum sensing.
3.1 Antibiotic Inactivation
Enzymatic inactivation of drugs by certain bacteria is an important mechanism for antibiotic resistance. They release drug-degrading enzymes that add an acetyl or phosphate group to the site of the antibiotic, decreasing its ability to attach to the ribosome and causing disruption in protein synthesis [52,53,54]. Genes encoding enzymes may be an intrinsic part of bacterial genome or can also be attained via HGT. Penicillin-resistant bacteria produce the most known enzymes called β-lactamases. They destroy the β-lactam ring of antimicrobial drugs having antimicrobial properties [55, 56]. This type of resistance mechanism has been reported in both Gram-positive and Gram-negative bacteria [57].
3.2 Target Site Alteration
The efficacy of the antibiotic drug depends on its interaction with the target site. To minimize the antibiotic effect of the drug, bacteria modify the target proteins thus affecting the antibiotic-protein interactions [56, 58]. Certain bacteria produce ribosomal protection proteins (RPPs) that bind to the bacterial ribosome resulting in conformational changes in ribosomes. This change in the shape of ribosomes prevents the binding of the antibiotic and the bacteria’s protein-synthesizing machinery remains unaffected [59,60,61]. A well-known example of target site alterations is the mutation in the penicillin-binding proteins (PBPs) in Streptococcus pneumonia leading to resistance against β-lactam antibiotics [62].
3.3 Active Efflux Pumps
Efflux pumps are responsible for expelling the solutes/toxins from inside of the cell. These energy-dependent efflux pumps are present on the plasma membrane and prevent the accumulation of antibiotics by pumping the drug outside the cell and maintaining the internal environment [63,64,65,66]. Resistance against the tetracycline group of antibiotics is developed via antibiotic efflux systems [65, 67, 68].
3.4 Changing the Metabolic Pathway
Certain bacteria overcome the drug effect by altering the metabolic pathway. For example, bacteria become resistant to sulfonamides by changing the metabolic pathway for the synthesis of folic acid. They synthesize folic acid from the environment instead of synthesizing it from Para-aminobenzoic acid (PABA), a precursor involved in folic acid synthesis which is inhibited by sulfonamides [69, 70].
3.5 Alteration of Bacterial Membrane Permeability
Change in the permeability of the internal and external membrane of bacterial cells results in decreased drug uptake. [71, 72]. Transporting proteins and channels i.e., porins in the bacteria’s outer membrane are a major entry route for hydrophilic antibiotics. Change in the number and type of porins in the membrane via mutation in specific porins called OprD, affects the susceptibility of bacteria for the antibiotics such as carbapenems, quinolones, and β -lactams [72,73,74]. Gram-negative bacteria are more resistant to antibiotic drugs as compared to Gram-positive bacteria. The reason lies in the complex outer wall architecture making them more likely to use a reduced permeability mechanism [73, 75] (Fig. 4).
3.6 Biofilm Formation Through Quorum Sensing
In an aqueous environment, certain bacteria build a network of extracellular polymeric substances (EPS). This mainly comprises of polysaccharides, proteins, DNA, and lipids that develop a shielding matrix around them and facilitate their attachment to a solid substrate [76]. Due to the extracellular network and metabolic dormancy, microorganisms covered in the biofilm are difficult to eliminate because it becomes practically more impossible for antibiotics to enter the multilayer structure as compared to a single cell (without biofilm/microcolonies) [77, 78]. It also helps them cling to solid surfaces and acquire a multicellular lifestyle by multiplying inside biofilms which leads to the formation of microcolonies, which are then enclosed in a layer of hydrogel that serves as a barrier between the microbial population and the outside world [79].
Hence, biofilm helps them to infect the host, spread into a new substratum as well as defend them against stress in their environment such as desiccation, high temperature, and action of antibiotics [80,81,82,83]. As a result, bacterial species often establish a resistance to conventional medications by adapting to these challenges [84]. Furthermore, biofilms give rise to the antibiotic degrading enzyme and increase communication between bacteria which makes it easier for mutations to occur by facilitating the transmission of genetic material, which also leads to antibiotic resistance [80,81,82,83].
There are multiple processes involved in the formation of the biofilm, including adherence, cell-to-cell binding, growth, maturity, and dispersion [85]. A signaling pathway known as the quorum sensing (QS) pathway is involved in the development of a coordinated functional community inside a biofilm [86]. It is used by bacteria to communicate with one another within their community through chemical signals called autoinducers. These autoinducers released by one bacterium are also detected by others in a community. When the amount of these molecules reaches a threshold concentration (a quorum), it binds to receptors present in bacteria. The target bacterium subsequently undergoes transduction of the signal into an intracellular biochemical signal and experiences changed gene expression [87]. This auto-signaling system induces cellular processes such as food uptake, movement, exchange of genetic material, and production of secondary metabolites as well as physiological responses such as activation of biofilm formation, and production of antibiotics degrading enzymes [88].
Biofilms contribute to antibiotic resistance by decreased antibiotic penetration due to its hampered diffusion across EPS; an anaerobic environment in inner layers and increased protection due to differentiation and specialization of the bacterial cell [89]. There are three primary QS systems including acyl homoserine lactone QS system (AHL), autoinducing peptide (AIP) QS system and the autoinducer-2 (AI-2) system [90] as explained in Fig. 5.
3.6.1 Quorum Sensing in Gram-Negative Bacteria
3.6.1.1 Acyl Homoserine Lactone (AHL) QS System
It is also termed as “LuxI/LuxRtype” quorum sensing. LuxI is an AHL synthase that synthesizes Acylated homoserine lactones (AHL) from fatty acid [91]. AHLs are autoinducers primarily released by Gram-negative bacteria. This inducing molecule enters bacterial cells and binds LuxR forming LuxR-AHL complex in bacterial cell cytoplasm. luxR (AHL-dependent transcription regulatory protein) undergoes conformational changes and exposes its DNA binding site. This activated protein binds target gene on DNA and transcribes it [92]. There exists a different homolog of LuxI/LuxR signaling system in Gram-negative bacteria. For example, LasI/LasR-RhlI/RhlR virulence system in Pseudomonas aeruginosa, TraI/TraR Virulence System in Agrobacterium tumefaciens, ExpI/ExpR-CarI/CarR Virulence System in Erwinia carotovora and LuxI/LuxR Bioluminescence System in Vibrio fischeri [80,81,82,83].
3.6.2 Quorum Sensing in Gram-Positive Bacteria
3.6.2.1 Autoinducing Peptide (AIP) QS System
Gram-positive bacteria release AIP autoinducer signal for quorum sensing [93]. These are amino acids or processed oligopeptides that are cleaved, modified and transported through ABC (ATP binding cassette) transporter. When the secreted peptide (AIP) reaches threshold concentration, it binds to cell membrane-bounded histidine kinase sensor receptor and phosphorylates it. This phosphoryl group is subsequently transferred to response regulator protein which activates and binds to DNA and causes transcription of QS target gene [94]. Homologs of AIP QS system signaling system in Gram-positive bacteria include ComD/ComE, Competence System in Streptococcus pneumonia, ComP/ComA Competence System in Bacillus subtilis and AgrC/AgrA Virulence System in Staphylococcus aureus [95].
3.7 Resistance Development Variations in Gram-Positive and Gram-Negative Bacteria
3.7.1 Resistance Mechanisms in Gram-Negative Bacteria
Pseudomonas aeruginosa develops resistance against carbapenem and imipenem by porin mutations causing OprD porin deficiency, overexpression of hydrolyzing enzymes such as AmpC, and increased levels of efflux pumps like MexCD-OprJ and alteration of PBPs, which decreases sensitivity to β-lactams and acquire drug degrading enzymes such as β-lactamases and Class B carbapenemases [96]. Macrolide, tetracyclines, and TMP/SMX resistance are similarly regulated by efflux pump production [97]. Its ability to resist colistin is based on altering the negative charge of the outer membrane essential for binding with the positively charged antibiotic. This is accomplished by the production of N4-aminoarabinose, a molecular compound that bind to, and consequently neutralizes the negative charge of lipid A. Moreover, mutations in the RNA polymerase gene cause rifampicin resistance in it [98].
E. coli strains develop resistance by synthesizing a wide range of β-lactamases, particularly AmpC, ESBLs, and class A, B, and D carbapenemases (OXA-48 & NDM) [99]. Furthermore, Enterobacteriaceae and E. coli develop resistance against fluoroquinolones, macrolides, rifampicin, and tetracyclines in a similar fashion [100]. Target site alteration using methylases, macrolide inactivation by phosphotransferases or esterases, and the development of efflux pumps are all modes of development of macrolide resistance and rpoB gene mutations cause Rifampicin resistance [101, 102]. Fluoroquinolone resistance is mainly brought about by polymorphisms of one nucleotide in gyrA gene, whereas tetracycline resistance is mediated by the overproduction of efflux pumps expressed by tet genes and the downregulation of porin [103].
Klebsiella pneumoniae become resistant to penicillins like ampicillin by synthesizing β-lactamases like TEM-1 and SHV-1. Carbapenemases that develop resistance in K. pneumoniae include Ambler Class A, VIM, IMP and NDM of Ambler Class B, Ambler Class D OXA-48 [104]. It acquired resistance against cephalosporins by production of AmpC, ESBLs and wide spectrum β-lactamases [105]. It develops resistance against aminoglycosides by producing efflux pumps, aminoglycoside altering enzymes that acetylate, adenylate or phosphorylate the target antibiotic, or by producing 16S rRNA methylase, which prevents aminoglycoside binding to 30S ribosomal subunit [106]. Fluoroquinolone resistance pathways include mutations in target enzyme gyrA gene, overexpression of acrAB efflux pump, possesses qnr genes on a plasmid, which encode proteins that protects DNA gyrase and topoisomerase IV from fluoroquinolones [107]. Tetracycline resistance is primarily caused by the development of efflux pumps like TetB [108]. Tigecycline resistance arises by upregulation of acrAB efflux pump, which develops either by a lack of inhibition of RamA, its transcription activator, or by disabling mutations in AcrR transcriptional repressor [109]. It also has ability to form biofilms which render antibiotics ineffective [110].
Acinetobacter baumannii acquire resistance through Class B carbapenemases like IMP and VIM, changes in PBPs and porins which give significant degrees of carbapenem resistance [111]. Tetracycline resistance arises by the synthesis of RPPs (ribosomal protection proteins) or the expression of TetA and TetB efflux pumps [112]. Resistance against tigecycline is linked to the expression of the AdeABC efflux pump or decreased uptake by mutations in plsC gene encoding integral membrane protein needed for tigecycline permeability [113]. A. baumannii develops resistance to aminoglycosides by producing all forms of aminoglycoside-altering enzymes, particularly nucleotidyltransferases, aminoglycoside, acetyltransferases and phosphotransferases [114].
Fluoroquinolone resistance is achieved through mutations in the genes encoding topoisomerase IV and DNA gyrase which reduce their susceptibility to fluoroquinolones and by the development of qnr-type protective proteins, which prevent fluoroquinolone binding to DNA gyrase and topoisomerase IV [115]. Furthermore, A. baumannii can also overexpress efflux pumps and inhibit the production of porins, lowering fluoroquinolone intracellular concentrations. The AdeABC efflux pump imparts resistance to many antibiotics simultaneously, including carbapenems, cephalosporins, fluoroquinolones, and aminoglycosides [116].
3.7.2 Resistance Mechanisms in Gram-Positive Bacteria
Gram-positive bacteria can develop resistance through two major mechanisms: the enzyme-mediated degradation of antibiotics through β-lactamases, or by decreasing the receptivity of target site i.e., penicillin-binding protein (PBP). It is done either by acquiring exogenous genetic material or by alterations in PBP genes [117]. VISA (Vancomycin intermediate Staphylococcus aureus) and VRSA (Vancomycin-resistant S. aureus) develop antibiotic resistance through cell wall modification. VISA resistance was associated with D-Ala-D-Ala residues in thicker cell wall than susceptible counterparts that serve as deceptive targets, preventing vancomycin from reaching its actual targets.
VRSA acquires resistant vanA gene containing plasmid from Enterococcus faecalis. It substitutes normal D-alanyl-D-alanine terminal with D-alanyl-D-lactate terminal that decreases binding affinity of vancomycin leading to antibiotic resistance [118]. S. aureus develop resistance to methicillin and β-lactam antibiotics by acquiring resistant gene from heterologous sources that code for PBP2a for which antibiotics have lowest affinity [119].
S. aureus give rise to penicillinase encoded by plasmid, which disintegrates penicillin's β-lactam ring, which is crucial for its antibacterial effect and to become penicillin resistant [120]. Resistance to ciprofloxacin, fluoroquinolones in S. aureus evolved as a result of mutations in genes encoding specific enzymes required for DNA replication i.e., DNA gyrase subunit gyrB and Topoisomerase IV subunit grIA, as well as alterations in drug entry and increased number of NorA efflux pumo [121, 122]. S. aureus aquire resistance against linezolid via variety of mechanisms such as single nucleotide mutations in its binding site V domain of 23S rRNA, inactivating methyltransferase that methylate 23S rRNA and mutations in L3 protein of 50S ribosomal subunit which comes in contact with ribosomal peptidyl transferase [123]. Tetracycline resistance in S. aureus is achieved by ribosomal protection proteins, which dislodge drug from its binding site ribosome, or by developing efflux pumps [124].
Resistance to penicillin in Streptococcus pneumoniae develops as a result of changes in at least one of the six PBPs in cell membrane of S. pneumoniae. This may have arisen by chromosomal mutation or acquired from other bacteria through transformation [125]. Resistance against lincosamide-macrolide-streptogramin in S. pneumoniae is caused by the erm(B) gene, which encodes a methylase, or by the mef(A) gene, which produces an efflux pump [126]. It has been documented that S. epidermidis becomes methicillin quinolones and vancomycin-resistant as a result of the transfer of the resistant mecA gene, which encodes PBP2a [127]. Streptococcus viridans serve as carriers for resistance genes like mel and mef(E) and established resistance to penicillin and other β-lactam antibiotics due to a modification in the penicillin-binding protein [128].
High resistance levels of E. faecium to fluoroquinolones have been documented and is developed by point mutations in parC and gyrA gene which encode topoisomerase IV A and DNA gyrase subunit A and NorA efflux pump [129]. Single nucleotide mutations in the S12 ribosomal protein of E. faecium lead to high-level streptomycin resistance [130]. E. faecium and E. faecalis become resistant to aminoglycosides by making their cell walls impermeable to aminoglycosides. E. faecium synthesizes phosphotransferases and acetyltransferases to disable aminoglycosides enzymatically, including kanamycin, tobramycin and amikacin [131]. Resistance to gentamycin in Enterococcus faecalis is achieved by enzymes that acetylate and phosphorylate the antibiotic, rendering it impossible to bind its target. Rifampicin resistance in both E. faecalis and E. faecium arises through mutations in rpoB gene encoding RNA polymerase [132].
Streptococcus agalactiae resistance to erythromycin and other macrolides is caused by either structural modification in ribosome expressed by erm genes or efflux pump encoded by mefA genes. Furthermore, in Streptococcus agalactiae, ribosomal translocation expressed by linB genes resulted in clindamycin resistance [133]. Bacillus anthracis and Bacillus cereus develop resistance against cephalosporins, ampicillin, penicillin and trimethoprim by producing ß-lactamases [134]. Corynebacterium diphtheria becomes resistant to sulfonamides, chloramphenicol and tetracyclines by acquiring resistant genes such as sul1, tet(W) and cmx via horizantal gene transfer [135]. Resistance to fluoroquinolones and tetracyclines emerged in Listeria monocytogenes by acquiring highly active efflux pump and conjugative transposons, respectively [136].
4 Role of Nanoparticles
To address the escalating problem of antibiotic resistance, there is an urgent need to modulate or even replace the current protocols of using drugs that could assist in lowering drug dependency (DD) [137,138,139,140,141]. Therefore, the search for novel and effective strategies regarding the control of bacterial damages is an urgent exigency and has become a priority for researchers worldwide [4, 6, 142,143,144]. Recently, nanoparticles (NPs) have emerged as promising tools to combat antibiotic resistance of human pathogens against antimicrobial agents [145,146,147,148,149,150]. These varieties of NPs could also be conjugated with various phytochemicals to enhance their respective biomedical potentials [80,81,82,83]. The antimicrobial properties of NPs are attributed to their smaller size and larger surface area to volume ratio which provides greater surface area for NPs contact with microbes [151, 152].
Nanoparticles are known to kill bacteria mainly by the physical rupture of the cell membrane, damaging the intracellular components via Reactive Oxygen Species (ROS) generation and by suppressing the bacterial metabolism [14,15,16, 110, 153, 154].
4.1 Antimicrobial Potential of Silver Nanoparticles (AgNPs)
Silver has been known to old civilizations for its medical-based applications for 2000 years [155]. Since the beginning of the nineteenth century, different compounds retaining silver as a major component have been in use for various antimicrobial purposes [156, 157]. Moreover, it is a fact that silver compounds are toxic to different types of microbes [158,159,160]. Nanoparticles composed of silver are also categorized as potent and viable antimicrobials against a broad spectrum of bacteria including MDR species due to multiple modes of action and high penetration potential [4, 161, 162]. Different properties like the high surface-to-volume ratio of AgNPs make them a more suitable and efficacious antibacterial agent than their counterparts [163,164,165,166]. As a result of this particular factor, AgNPs expressed specific chemical, physical and biological characteristics [167]. On the other hand, the stability of silver nanocomposites is a major concern, which makes its role in the medical and health sector, debatable [4, 168].
In this regard, AgNPs are now being synthesized in different environmental conditions (temperature, light) to assess their stability [169]. Second, human and eco-related toxicity of AgNPs has also emerged as it causes mitochondrial mutilation [170]. To overcome the problem of cytotoxicity, alternative eco-friendly stabilizing and reducing agents like different biopolymers are being used [171,172,173]. Tahir et al. [150] also reported the synthesis and antibacterial activity of AgNPs by using sericin (biopolymer) as a reducing and stabilizing agent. Many other phytochemicals including terpenoids, phenols like eugenol, bacterial probiotics, fungi, algae, proteins, oils and their secondary metabolites have the reducing ability for NPs synthesis as well as individual antimicrobial potential.
4.1.1 Mode of Action of AgNPs
Many scientists have published across the world various mechanisms by which these metallic NPs work against a wide variety of bacteria. Lazar [174] and Periasamy et al. [175] observed and disclosed that the cell membrane deterioration leading to the structural mutations ultimately causes the death of different bacteria when exposed to the AgNPs. Multiple studies against E. coli revealed that AgNPs accumulation in the cell creates gaps in the membranes leading to stability loss and thus microbes collapse [176,177,178]. The efficiency of NPs is inversely dependent upon their size i.e., smaller the size, the greater the penetrance into the bacterial cell and the higher the mortality and vice versa as depicted by several studies conducted by Collins et al. [179], Wu et al. [180], Tamayo et al. [181], Franci et al. [167], Chen et al. [182] and Muchintala et al. [183]. It seems that when NPs with positive potential come into contact with a bacterial surface with a negative charge, attract each other and NPs enter the cell [182].
It is also evident from the studies that resistance against silver is very rare which indicates the multiple and collective microbe-killing mode of actions [184, 185]. Different studies conducted by Rolim et al. [186], Beyth et al. [187] and Lee & Jun, [173] revealed that free radicals are generated when reactive oxygen species are produced by the interaction of bacterial cells with silver-based nanocomposites, causing the mortality. Bury et al. [188], Morones et al. [189] Jung et al. [190], Lee & Jun, [173] and Muchintala et al. [183] revealed that AgNPs can also cause the blockade of protein synthesis and mRNA movement in the cell leading to the genetical collapse depicted in Fig. 6.
Moreover, NPs are responsible formation of free radicals which finally disrupt the membrane potential and bacterial cell lost integrity [191,192,193,194]. It is revealed by recent studies conducted by Elgorban et al. [195] and Shaban et al. [196] that activities of AgNPs i.e., smaller NPs ranging from 1 to 10 nm efficiently damage the membrane and cause respiratory problems than their bulk counterparts. Narware et al. [197] reported that AgNPs kill the bacteria through the oligodynamic mode of action i.e., by inactivation of respiratory enzymes.
4.1.2 Bacterial Strains and Nanoparticles Mode of Action
4.1.2.1 Nanoparticle’s Mode of Action Against Gram-Positive Bacteria
There are non-significant differences in the bactericidal mode of the various NPs against Gram-positive and Gram-negative pathogens [198]. Mostly NPs inhibit the growth of various pathogenic and non-pathogenic bacteria with proportionally similar pattern with minor degree of variations from cell integrity loss to cytotoxicity to DNA damage [199]. Against Staphylococcus aureus, silver NPs cause cytotoxic damage through formation of the reactive oxygen species which ultimately bind with multiple organelles of the cell due to their unstable nature [200,201,202,203].
Moreover, these nanoparticles kill the bacterial pathogens through nucleic acid denaturation and even the damage to the different enzymes which are necessary for their multiple metabolic activities. Furthermore, AgNPs nanocomposites impart irreversible damage to the cells and DNA, ultimately causes the mortality [204,205,206,207]. Wang et al. [208] and Nishu et al. [209] reported that AgNPs inhibit the respiration-related activities of susceptible microorganisms. Overall, different biogenic and metallic NPs have shown various degrees of antibacterial potential against Gram-positive and Gram-negative bacteria. In this regard, AgNPs are proven to be more efficacious against Gram-positive due to their comparatively thin layer of peptidoglycan.
ZnONPs cause severe damage to other Gram-positive rod bacteria including Bacillus, Mycobacterium and Streptomyces causing the charge-dependent toxicity which subsequently destabilizes the membrane potential of the pathogens, thus causes death [210].
Damage to the genetic material (DNA) through inhibition of the replication phenomenon, deterioration of the cytoplasmic membrane or even ATP level modifications through intercalation of the nitrogenous bases are the most common pathways of the copper-based NPs against Staphylococcus species [211]. Certain copper-chitosan conjugated NPs also impart large surface cell wall collapse, cell leakage and wrinkled cell wall which ultimately inhibit the growth of Mycobacterium tuberculosis [212]. Cell cycle arrest against the Mycobacterium followed by cell disintegration through pore formation using selenium NPs is also reported by Estevez et al. [213].
Moreover, different researchers reported the morphological damage, and cytotoxicity by NPs accumulation in the Enterococcus species as a possible mode of bactericidal activity against different Enterococcus species including Enterococcus faecalis [214]. Iron oxide and other NPs also have the ability to counter the various Gram-positive bacteria including Bacillus sp. and Enterococcus sp. by inhibiting the biofilm formation individually as well as in a synergistic manner with various novel synthetic drugs [80,81,82,83,84, 215, 216].
4.1.2.2 Nanoparticle’s Mode of Action Against Gram-Negative Bacteria
AgNPs Block DNA replication and mutilation of the cytoplasmic membrane and abrupt intracellular changes of ATP which lead to bactericidal activity in Salmonella typhi [189, 217]. Biosynthesized NPs from various juice extracts from medicinal plants like Citrus macroptera harbor significant bactericidal potential through inhibition of the biofilm formation against Gram-negative bacteria including Pseudomonas aeruginosa [80,81,82,83].
The bactericidal potential is expressed by NPs as they modulate the permeability of cell membranes and mutate the process of respiration [189, 218, 219]. Morones et al. [189], Mahapatra et al. [220] and Saberpour et al. [221] revealed that nanosilver alters the membrane structure against Vibro cholera. Cell membrane alteration in Klebsiella pneumonia is the leading cause of mortality after exposure to silver-based NPs reported by Manjumeena et al. [222], Tamayo et al. [181], Siddique et al. [223] and Pareek et al. [224]. Lysakowska et al. [225], Salih et al. [226], and Singaravelu et al. [227] depicted in their study that cytoplasmic changes induced by silver nanoparticles are responsible for bactericidal activity against Acinetobacter baumanni.
4.1.3 Molecular Basis of Nanoparticles Mediated ROS Generation
ROS production is a significant byproduct of NP-mediated damage [228]. ROS elements include free radicals such as singlet oxygen (1O2), hydroxyl (HO•), superoxide (O2•–) and non-free radicals like hydrogen peroxide (H2O2), nitric oxide (NO), hypochlorite (OCl–) and hypochlorous acid (HOCl) [229]. They are produced primarily in organelles like the peroxisomes, endoplasmic reticulum (ER), microsomes, cell membrane complexes and, most importantly, mitochondria [230]. Oxygen is utilized for the production of water during oxidative phosphorylation by transfer of electrons via electron transport chain in mitochondria (ETC). Some of these electrons are taken up by molecular oxygen to produce O2−, which can be converted into OH• and other forms of ROS [231].
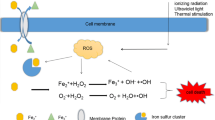
The mechanism behind NPs mediated ROS production differs amongst NPs. The majority of nanoparticles derived from metals can cause free-radical-mediated toxicity via Fenton-type reactions [232]. Metal ions produced by nanoparticles have been proven to interact with chemo-catalysis and redox cycling through the Fenton reaction [Fe2+ + H2O2 → HO− + Fe3+ + •OH] or Fenton-like reaction [H+ + Ag+ H2O2 = Ag+ + H2O + •OH] as shown in Fig. 7 [233]. The isolated metal ion(s) (Ag+) also deactivates cellular enzymes, disrupts membrane structure, disrupts the electron-shuttling process, reduces redox potential levels, lowers mitochondrial membrane potentials (MMP) and increases intracellular ROS accumulation [234]. NPs have also been shown to increase intracellular ROS levels by interfering with the electron transfer process raising the NADP+/NADPH ratio, and interfering with mitochondrial activity [235]. NPs also disrupt the expression of oxidative stress-related genes such as ahpC soxR, soxS, and oxyR, antioxidant genes such as gpx 1 and sod1, and gene-producing NADPH met9. The instabilities in the transcription of oxidative and antioxidant genes triggered by NPs hasten the formation of intracellular ROS [236].
4.1.3.1 Mechanism of ROS Generation
NPs must interact with cell membranes to infiltrate the cell. Nanomaterials are transported into cells through encapsulation in vesicles via endocytosis [237]. Internalization of NPs may or may not depend upon caveolin or clathrin proteins [238]. Metabolic processes that contribute to the formation of intracellular ROS by NPs include binding with mitochondrial components, growth factors activation, and activation of membrane complexes [239].
Mitochondria is a primary organelle in the NPs mediated production of ROS. NPs induce depolarization of the mitochondrial membrane and disrupt electron transport chain by activating NADPH-related enzymes [240]. Hence, After NP exposure, an electron-transport chain is inhibited, raising intracellular concentrations of O2•− through transferring electrons from respiratory complex carriers to O2 [241]. Human fibroblasts and glioblastomas exposed to NPs were shown to accumulate more AgNPs, CuNPs, ZnONPs, etc., in mitochondria which disrupted electron-transport chain and, as a result, high levels of ROS production [242]. In the majority of bacteria, metal ions interacted with NADH dehydrogenase, preventing electron transport to O2 and the production of large amounts of ROS [243].
Moreover, the conversion of GSH into glutathione disulfide, caused by NPs-induced free radical generation, contributes to oxidative stress [244]. The activation of ROS-associated receptors and enzymes by NPs also contributes to the production of intracellular ROS [245]. The ability of NPs to induce DNA damage is due to the formation of the free radical HO•, which binds with DNA and give rise to 8-hydroxyl-2′-deoxyguanosine (8-OHdG), which eventually causes DNA damage [246].
1. NPs are internalized into cells via endocytosis. 2. It leads to production of endocytotic vesicles. 3. Then particle ions are released from endocytotic vesicles into a cell. The primary causes of ROS formation by NPs. 4. Contact with mitochondrial membrane. 5. Attachment to NADPH oxidase which causes loss of electron by oxidizing NADPH and transfer electron to oxygen, converting it into superoxide radical. 6. The majority of nanoparticles cause free-radical-mediated toxicity via Fenton-type reactions. The aforementioned factors cause ROS production and associated effects, which include cell cycle arrest, DNA damage, modification in apoptosis, and cell membrane damage which ultimately leads to cell death.
4.2 Antimicrobial Potential of ZnO NPs
ZnO NPs are fetching considerable attention due to their unique properties like low toxicity, high absorption rate, and biological activities including antibacterial and antioxidant potential [247,248,249,250,251,252]. Several studies have reported the antibacterial properties of ZnO NPs and their non-toxicity to human cells [253,254,255]. Zinc oxide nanoparticles are being used in food packaging as an antimicrobial agent against Salmonella, Staphylococcus aureus, C. jejuni, and E. coli as these are major foodborne bacteria to cause food deterioration [256,257,258].
They show a wide range of antibacterial activities against Gram-positive and Gram-negative bacteria. Moreover, Gram-positive bacteria are found to be more susceptible to ZnO NPs as compared to NPs of other elements of the same group [234, 250, 251]. The antibacterial activity of ZnO NPs is dose and size-dependent,an increase in NPs concentration results in the increased efflux of cytoplasmic content while it decreases with the increase in the size of nanoparticles as revealed by several studies conducted by Tiwari et al. [259], Espitia et al. [260], Abebe et al. [261], and Fahimmunisha et al. [262].
4.2.1 Mode of Action of ZnO NPs Against Different Bacterial Strains
Although the mechanism of toxicity of ZnO is still controversial and requires deep explanation, the proposed mode of action includes contact between ZnO NPs and bacterial cell wall followed by loss of integrity and disruption of the cell wall, hence release of intracellular contents and generation of reactive oxygen species (ROS) [252, 262,263,264,265,266]. Moreover, various studies suggest the dissolution of ZnO NPs into Zn2+ which binds to major biomolecules like proteins and carbohydrates results in the cessation of vital functions of bacteria [234, 267]. Antimicrobial properties of ZnO NPs can also be attributed to the photo-induction process since being a semiconductor ZnO has high photocatalytic efficiency. This photoconductivity is increased when exposed to UV light as it stimulates the interaction between ZnO and bacterial cells [255, 268]. Although Adams et al. [269], Hirota et al. [270], Lakshmi Prasanna &Vijayaraghavan [271], and Jeong et al. [272] provided evidence of ROS generation even under dark conditions [273]. ZnO NPs increase cytotoxicity predominantly by ROS production, which causes oxidative damage and the release of inflammatory chemicals, eventually leading to cell death as depicted in Fig. 7 [274].
The generation of ROS is the most commonly reported mechanism for antibacterial activity which occurs as a result of the formation of different intermediates, the generation of hole pairs and their reaction with oxygen or water as reported by Abebe et al. [261]. Both, direct (inside the bacterial cell) and indirect (outside the bacterial cell) generation of ROS has been observed by Thakur et al. [275]. ROS causes damage to the bacterial cell's biomolecules including proteins and DNA [276] (Fig. 8).
4.2.2 Specific Bacterial Strains and ZnO NPs Mode of Action
4.2.2.1 Campylobacter jejuni
Tiwari et al. [258] reported the formation of cell wall blebs and irregular surfaces in C. jejuni when exposed to ZnO NPs. The disruption in the cell membrane of C. jejuni and overexpression of the genes (ahp C and KatA) that are triggered by oxidative stress was observed by Xie et al. [277]. Altered gene regulation in response to ZnO NPs was also confirmed by Campbell et al. [278] and Cerasi et al. [279]. The formation of ZnO2+ ions from immobilized ZnO NPs also curbed the growth of C. jejuni in raw chicken meat [280].
4.2.2.2 Staphylococcus aureus
The growth of Staphylococcus aureus was inhibited by a complex mechanism involving different metabolic pathways including disruption of sugar metabolism and amino acid biosynthesis instead of ROS generation as described by Kadiyala et al. [281]. Lallo da Silva et al. [282] observed the accumulation and internalization of nanoparticles within the bacterial cell when exposed to ZnONPs.
Another novel mechanism was proposed by Kadiyala (2018) which suggested the killing of Staphylococcus aureus by enhanced expression of pyrimidine biosynthesis and upregulation of carbohydrate degradation rather than ROS generation. However, Navarro-López et al. [283] described the ROS generation as the possible mechanism against Staphylococcus aureus.
4.2.2.3 E. coli
Li et al. [284], Pasquet et al. [285], and Navarro-López et al. [283] reported that cell wall damage, increased membrane permeability, ROS production and release of Zn2+ ions following the dissolution of ZnO NPs led to the toxicity in E. coli when exposed to the ZnO nanoparticles. It was also observed that Zn2+ ions had a smaller effect when compared to the toxicity caused by ROS [286].
4.2.2.4 Staphylococcus epidermidis
Akbar et al. [287] described the morphological and physiological changes in the bacterial cell along with pitted and deformed cell walls in Staphylococcus epidermidis when treated with ZnO NPs. Palanikumar et al. [288] reported the ROS generation and their subsequent accumulation in the cytoplasm as a possible mechanism against Staphylococcus epidermidis.
4.3 Antimicrobial Potential of Copper Nanoparticles (CuNPs)
Copper (Cu) is naturally present in certain food groups in small concentrations and acts as a catalyst for some enzymes. However, when found in high concentration, it was reported to have antimicrobial properties against some major food-borne bacteria including Campylobacter jejunii and Salmonella enterica [289,290,291,292]. Copper has also been recognized as an antimicrobial material by US Environmental Protection Agency (EPA) [293]. The antimicrobial property of copper has led the researchers to synthesize its nanoparticles to enhance its antimicrobial effectiveness associated with high surface area to volume ratio of the NPs. Moreover, the use of CuNPs in many biomedical applications can be cost-effective due to their low cost and ubiquitous availability [294,295,296].
CuNPs have been stated to exhibit broad-spectrum antimicrobial activity [295, 297, 298]. Many researchers have confirmed the antimicrobial properties of CuNPs against a number of Gram-positive and Gram-negative bacteria [299,300,301,302]. They are even reported to have superior antimicrobial properties than AgNPs against B. subtilis and E. coli and can be used as substitutes for expensive AgNPs [300, 303].
However, the formation of oxides by CuNPs on exposure to atmospheric layer can limit the antimicrobial properties [304]. To prevent oxidation during synthesis and storage of CuNPs, an inert atmosphere is required which increases the complexity of the process. Nonetheless, the oxidation of the CuNPs can be avoided by using green synthesis methods [305, 306]
Biomolecules in the plant materials including flavonoids, phenols, and tannin provide stability by capping and prevent oxidation of NPs [307, 308]. Moreover, the use of green synthesis for NPs production is convenient, eco-friendly, and inexpensive [234]. In this regard, Pérez-Alvarez et al. [306] reported the synthesis of CuNPs using cotton. The subsequent NPs formed were found to be stable to oxidation and could be stored for months without any change in the properties.
4.3.1 Mode of Action of Cu Nanoparticles Against Different Bacterial Strains
The information regarding mode of action of CuNPs against microbes is quite limited but proposed mechanisms suggest that the CuNPs release Cu2+ in interaction with cell wall. These ions get adsorbed on the cell wall of bacteria which later leads to formation of pits in the membrane and loss of cell membrane integrity [295, 309,310,311,312]. The affinity of CuNPs towards amines, carboxyl group and sulfhydryl groups in the peptidoglycan layer denatures the proteins of cell membrane [313,314,315]. CuNPs can also penetrate via endocytosis in plasma membrane. Once inside the cell, Cu2+ binds to DNA molecule and disrupt the helical strands by cross-linking, leading to disorganization of nucleic acid molecules [227, 237, 316,317,318].
Metal oxide NPs such as Cr2O3, Co3O4, Mn2O3, Ni2O3, CuNPs, and CoO cause cytochrome c oxidation and oxidation of NADPH into NADP+ leading to oxidative stress [319]. Some photon-activated NPs, for example, TiO2 NPs produce electrons with sufficient amount of energy to convert O2 into 1O2 (singlet oxygen), which causes cellular damage by binding cellular proteins, nucleic acids and lipids as shown in Fig. 7 [320]
The cell may be subjected to oxidative stress due to generation of reactive oxygen species (ROS) as reported by Applerot et al. [321] and Chatterjee et al. [316] as shown in Fig. 9. The increased ROS production disrupts several metabolic pathways through enzyme inactivity and protein dysfunction [322, 323]. The antimicrobial activity of CuNPs is greatly dependent on particle size, composition of bacterial cell wall, and nature of nanoparticles.
4.3.2 Specific Bacterial Strains and CuNPs Mode of Action
4.3.2.1 E. coli
Chatterjee et al. [316] concluded that the mechanism of toxicity of CuNPs against E. coli was found to be the release of Cu2+, followed by loss of integrity of plasma membrane. However, generation of ROS was not reported. While other studies investigated the interaction of CuNPs with cell wall and reported the generation of ROS inside different E. coli strains [303, 311, 324]. Metryka et al. [325] and Sharma et al. [326] stated that the loss of cell membrane integrity, ROS generation and leakage of intracellular content is the cause of bacterial cell death.
4.3.2.2 B. subtilis
The interaction of Cu2+ ions and their subsequent binding with the amines and carboxyl groups in the peptidoglycan layer was observed by Ren et al. [315] against B. subtilis. Li et al. [327] reported that damage to outer membrane and protein disruption are responsible for cellular toxicity. Phan et al. [328] suggested that damage to bacteria was dependent on copper ions (Cu2+) concentration and size/shape of the nano copper.
4.3.2.3 Staphylococcus aureus
Li et al. [329] and Yadav et al. [330] reported the damaged and wrinkled cell wall, increased permeability of plasma membrane, and leakage of intracellular material in Staphylococcus aureus when exposed to Catechin-Cu nanoparticles.
4.4 Antibacterial Properties of Titanium Dioxide (TiO2) Nanoparticles
Titanium dioxide nanoparticles have been explored widely due to their high stability and nontoxic nature. Besides having unique optical properties, TiO2 is an excellent photocatalyst which makes it ideal in antimicrobial applications [331,332,333,334,335]. The antibacterial activity has been explored against a diverse range of Gram-positive and Gram-negative bacteria [336, 337]. Albukhaty et al. [338] prepared TiO2 NPs using sol–gel technique and evaluated their antibacterial potential against S. aureus and E. coli.
Modifications of TiO2 NPs by Garcinia zeylanica and grape seed extract led to enhanced antibacterial activity against S. aureus, Gram-negative (P. aeruginosa) and Gram-positive (S. saprophyticus) respectively [339, 340]. Azizi-Lalabadi [341] further modified the TiO2 NPs by embedding them into 4A zeolite and depicted their increased bactericidal potential against P. fluorescens and E. coli. The antibacterial activity of TiO2 NPs was found to be size and zeta potential dependent i.e., NPs with smaller diameter and more positive zeta potential exhibited higher bactericidal activity [342]. An in vivo investigation found that exposure to Ti, Ag, Cu, and Fe NPs causes genotoxicity by nucleic acid damage. Resulting oxidative stress- stimulates signaling pathways leading to the activation of inflammatory mediators, including interleukins and tumor necrosis factor [343]. Figure 7 shows the molecular basis of ROS production which is also associated with the inflammatory reactions induced by metallic nanoparticles (SiO2 NPs & TiO2 NP).
4.4.1 Mode of Action of TiO2 NPs Against Different Bacterial Strains
Although, TiO2 NPs possess catalytic properties, TiO2 NPs have shown antibacterial activity with and without UV illumination which hints the presence of toxicity mechanisms aside from photolytic reactive oxygen species (ROS) generation [344,345,346,347]. The mechanism of UV-illuminated TiO2 NPs includes production of ROS including superoxide ions (O∙ −) and hydroxyl radicals (OH∙) by reduction of oxygen and oxidation of H2O as illustrated in Fig. 7. This mechanism was found to be in consistent with other photolytic reactions [344, 348, 349].
Generated ROS disrupt the ionic balance, oxidize the cell membrane components and lead to destruction of plasma membrane. After entering the cytosol, TiO2 NPs alter the gene expression by discontinuing the enzyme activity and changing the structure of macromolecules in the bacterial cell [350,351,352]. Sohm et al. [353] investigated the bactericidal action mechanism of TiO2 NPs in a dark environment by exposing E. coli to them. It was illustrated that NPs adsorbed on the cell surface and induced depolarization and instability of plasma membrane. Loss of membrane integrity contributed to leakage of K+ and Mg2+ ions, entrance of Na+ to the cell, and respiratory chain deficiency due to exhaustion of intracellular ATP level. A similar mechanism was proposed by Seil and Webster [335] and Nemattalab et al. [354]. Sohm et al. [353] and Pagnout et al. [355] also concluded that ROS generation had a trivial effect in causing toxicity in dark conditions.
4.4.2 Mode of Action of TiO2 NPs Against Specific Bacterial Strains
4.4.2.1 E. coli
Khan et al. [356], Albukhaty et al. [338] & Nemattalab et al. [354] observed that TiO2 NPs caused ROS generation and phospholipid peroxidation that led to the death of E. coli. Different researcher also documented that TiO2 NPs may also attach to the outer wall of the bacteria and make small pores (porins), which disturb the cell integrity and leads to the death of the bacteria. These NPs have also the ability to deteriorate the activity of various vital enzymes necessary for important functions like pathogenicity [297, 298, 357]
4.4.2.2 Staphylococcus aureus
Abdulrahman et al. [358], Bekele et al. [359], Albukhaty et al. [360] evaluated the bactericidal activity of TiO2 NPs and stated that destruction of the outer membrane and leakage of the cell content was a major cause of bacterial cell death (Fig. 10). Some other researchers also reported that the titanium-based NPs could also generate the non-stabilized reactive oxygen species (ROS) which bind with multiple macromolecules of the bacteria including the nucleic acids, proteins, and lipids thus damaging the DNA machinery and causing cell death [201, 297, 298].
4.4.2.3 Streptococcus
Loss of cell viability and outflow of K + ions were observed in the cells of Streptococcus sobrinus which was followed by slow leakage of intracellular material when exposed to TiO2 NPs [361]. Moreover, Besinis et al. [362] and Pourhajibagher et al. [363] documented that these NPs with size less than 100 nm have the potential to damage the outer morphology of the bacterial body as well as the damage to the DNA. Miron et al. [364] reported the membrane damage in Streptococcus pneumoniae using atomic force microscopy. Khan et al. [365] documented the internalization of the nanoparticles followed by ROS generation and DNA damage.
5 Pseudomonas aeruginosa
Amezaga-Madrid et al. [366] and de Dicastillo et al. [331] documented the impairment of the cell wall and structural damage of the plasma membrane that resulted in subsequent leakage of cellular material of Pseudomonas aeruginosa as shown by SEM and TEM. Rajkumar et al. (2019) reported the antibiofilm activity of the titanium-conjugated NPs which stops the intercellular bacterial communications within colonies and makes them susceptible for cytotoxic damage through direct contact of the NPs. Furthermore, such NPs can also cause mortality through ROS production [367].
5.1 Antimicrobial Potential of Cobalt Nanoparticles (Co NPs)
The transition metal, Cobalt has become of special interest because of the unique physicochemical properties manifested in Co NPs due to the quantum size effects [368]. The nano-sized cobalt particles are known to have catalytic, optical, biomedical and antibacterial properties [369, 370]. They are being explored as an active therapeutic agent against infectious diseases [371]. Synthesis of Co NPs is inexpensive, biocompatible, and does not require any additional stabilizing agent [372]. They are non-toxic to body at lower concentrations but highly effective against bacteria and fungi even at minimal levels of concentration making them favorable for biological applications [373].
The antibiotic potential of Co NPs was found to be stronger as compared to standard antibiotic drug ciprofloxacin [374]. The green method synthesized Co NPs have shown promising antibacterial activity against different bacterial strains including E. coli, S. aureus, and Klebsiella pneumonia [375].
5.1.1 Mode of Action of Co NPs Against Different Bacterial Strains
Igwe and Ekebo [370] reported the possible antibacterial mechanism of cobalt nanoparticles bio-fabricated from the leaf extract of C. odorata against E. coli, K. pneumonia, S. aureus, and S. pyogene. The study suggested the direct contact of Co NPs owing to their small size (20–49 nm) with the outer surface of pathogens led to its rupture and eventually cell death. Metal ions bearing a positive charge are strongly attracted to cell membrane carrying the negative charge that initiates the loss of cell membrane integrity and penetration of ions inside the cell [376]. Kharade et al. [377] also suggested a similar mechanism of CoNPs against Bacillus subtilis and Escherichia coli.
Another mechanism against bacteria was documented by Shriniwas and Subhash [378] involving the destruction of protein synthesis (Fig. 11). Similarly, Guan et al. [379] reported the physiological changes in bacterial cells by disrupting transcriptional regulation caused by Co NPs. Abdal Dayem et al. [235] reported that Co NPs influence oxidative stress through reactions between oxidized metal ions and H2O2 to produce free radicals. This further contributes to DNA damage and protein dysfunction. The interaction of cobalt ions with bacterial enzyme thiol and subsequent imbalance of electron transport chain is another major factor causing bacterial cell death. [380, 381].
5.1.2 Mode of Action of Co NPs Against Specific Bacterial Strains
5.1.2.1 E. coli
Satpathy & Manikandan [382] demonstrated the bactericidal mechanism of E. coli which is caused by binding of Co NPs with cell wall due to increased lipophilicity of metal ions.
5.1.2.2 Staphylococcus aureus
Co NPs showed antibacterial activity against this multi-drug resistant bacteria through electrostatic interaction of Nps with its cell wall [383]. Another possible mechanism could be the release of cobalt ions disrupting the DNA replication machinery and protein inactivation [384].
5.1.2.3 Pseudomonas aeruginosa
Co-NPs exhibited adhesion to the cell wall and altered its permeability. After penetration, ROS production and change in cell signaling caused cell death [385].
5.1.2.4 Bacillus Subtilis
Sivachidambaram et al. [386] suggested that generation of ROS was the key factor in causing bacterial cell death.
6 Discussion
Synthetic drugs/antibiotics have revolutionized the frontline medications for treatment of various life-threatening ailments across the globe but their unregular use has become a nightmare in the form of antimicrobial resistance (AMR) or development of multiple drug resistant (MDR) pathogenic strains [21, 387]. Surprisingly, antimicrobial resistance has paced out the discovery of novel antibiotics to counter the resistance phenomenon. Epidemiological statistics of 204 countries and their peripheries estimated that 4.95 million people with infections due to MDR pathogens in 2019 and more than 1.2 million mortalities due to AMR [388]. Moreover, due to current wave of SARS-CoV-2, worse conditions regarding drugs resistance being anticipated due to administration of a bulk of antibiotics to patients with viral infections [389,390,391] With the passage of time, majority of the bacteria have developed various intrinsic and adaptive/acquired modes to escape from the influence of these conventional therapeutics [50, 53].
During intrinsic antibiotic resistance modes, bacteria get rid of the antibiotics by up-regulation of efflux pumps, activation of drug-altering enzymes and altering the target sites of the antibiotics as depicted in Figs. 6 and 12 [43]. On the other side, in acquired resistance phenomenon, susceptible bacterium develop resistance by altering the genetic makeup. Pathogens may also become resistant through chromosomal mutation, genetic rearrangements or adopting the exogenous genetic material through transformation, transduction and conjugation as shown in Fig. 2 [46, 47]. Although certain new antibiotics are being approved by FDA but there is no guarantee regarding their efficacy [27]. Moreover, these drugs are not cost-efficient and pose serious threats to healthy tissues.
The current dilemma of antibiotic use has urged the whole scientific community to search for some alternative therapeutic solutions with novel modes of action to overcome the AMR [36, 392]. In this regard, nanotechnology has emerged as the sole solution to the problem of antibiotic resistance due to its tunable nature and novel physiochemical and biological properties [393, 394]. Nanoparticles employed the application of entity sizes ranging from 1 to 100 nm with large surface-to-volume ratio, and variable interaction ability to the bacterial surface for efficient bactericidal potential [395, 396]. These nanoparticles are being preferred over the conventional clinical approaches as they have some novel mechanism to kill the bacteria or they have ability to modify the already available mechanisms alongside the existing antibiotics in a synergistic manner.
Nanoparticles are usually conjugated with certain reducing agents of chemical origin for required physiochemical bonding or with bio-constituents of living organisms including plants, animal or bacterial origin. Various metals including silver, copper, zinc, cobalt, titanium, gold and silica, etc., are in use for different biomedical applications including antibacterial [396,397,398,399,400,401]. These NPs in reply of AMR, overcome the problem by adopting the novel pathways including the damage to the outer membranes of the bacteria through pores formation after destabilization of the membranes [402]. As these biogenic nanoparticles are less than 100 nm, they might have the significant ability to cross the cell wall of the bacteria to reach the internal organelles for efficient damage.
Many researchers also reported the high mortality rate of the bacteria after exposure to smallest size NPs as compared to the large-sized NPs of same metal [148, 149, 403]. Mageshwaran et al. [404] also endorsed that NPs with least size have the ability to interact with the genetic material of the pathogen and thus cause mortality. Furthermore, these NPs inhibit the growth of microbes by many other novel modes including the ROS generation which indirectly cause cellular damage by immediate linkage with the bacterial organelles [110, 252, 405]. Similarly, altering the permeability of the outer membrane of the bacteria through ion channel deterioration is another indirect passage of certain NPs to neutralize the pathogens [385] as shown in Fig. 12. It has now been established by the previous studies that mechanism through which pathogens develop resistance against frontline antibiotics are countered by novel NPs mostly by alternative pathways as clearly shown in Figs. 6,8,9,10 and 11.
7 Toxic Nature of Nanoparticles
Despite showing promising results for various biomedical/clinical applications, nanoparticles have been found to be potentially hazardous [406,407,408,409,410]. To minimize the toxicity caused by NPs, dose optimization is critical. Currently, the concentration of NPs being used in vitro for cell damage is quite high which halts its application to humans. Moreover, translating the results from animal studies to specifically human beings may not be applicable [365, 411].
7.1 Nanotoxicology
Nanotoxicology has fetched the attention of health concerns after the emergence of the use of metallic and biogenic nanomaterials in various biomedical ad other health-related fields. This critical class of novel understanding not only highlighted the potential deteriorative health effects of nanoparticles on humans but also on other animals, plants and even the ecotoxicological role by contributing towards air and water pollution [412]. In this regard, Organization for Economic Cooperation and Development in 2005, raised reservations regarding the health risks associated with the use of nanoparticles for the 1st time. As a result of this awareness, various databases across the globe have been established highlighting the potential benefits and risks of the nanoparticles as well as the standard protocols of acute and chronic toxicity testing [413].
7.2 Types of Toxicity
Dose, duration and frequency of exposure and pathway of administration are the critical factors influencing the toxicity [408, 414, 415]. Nanoparticles can reach into the body through skin (subcutaneous, cutaneous) penetrance, inhalation by lungs, orally (GI tract) and intravenously same as the common routes of other drugs administration. The intravenous route is susceptible to highest toxicity due to direct delivery or mixing of any drug/nanomaterial. In this regard, outcomes or the results of the nanomaterials can be desirable, undesirable or both. Undesirable effects are also then referred as negative or adverse effects like allergic reactions [416, 417]. Regarding the types, toxicity can be immediate or slow based on the time of its symptoms, non-reversible or reversible based on whether the adverse impact is permanent or not and localized or systemic due to its local (on the site of administration) or throughout the body [418]. Elimination time, body metabolism response, absorption and distribution of any nanoparticle could also tune the toxicity. Generally, toxicity after single-dose exposure varies from multiple/repeated exposures.
Therefore, on the basis of exposure, toxicity that is observable during the 1st 24 h of administration is termed as acute while, while the response against any nanomaterial after repeated exposure seen within a month is known as subacute and referred to as chronic if the effect is observable after three months of chronic exposure [417, 418].
The purpose of acute toxicity evaluation is to assess the NOEL (non-observable effect level) and MTD (maximum tolerated dose) of the administered nanomaterials. FDP (fixed dose procedure), LD50 (Dose at which 50% mortality occurs) and ATC (acute toxicity category) are the most common methods to evaluate acute toxicity [419, 420]. Moreover, mechanisms of NPs toxicity as well as organ toxicity are elucidated by the acute and sub-acute toxicity [421].
Immediate behavioral, cardiovascular, hematological and neuronal responses, weight fluctuations as well as clinical manifestations including the effect of NPs exposure to respiration, body movements, dilation or constrictions of blood vessels, GI tract function, skin and fur and histopathology of vital organs are being monitored during acute and sub-acute toxicity [416, 422]. To assess the long-term safety of the NPs, neurotoxicity, immunotoxicity, ophthalmological parametric study and heart functioning are being considered. Carcinogenesis, reproductive toxicity, embryotoxicity and genotoxicity are necessary to evaluate the sub-chronic and chronic toxic nature of the NPs [422,423,424].
7.3 Mechanisms of Nanoparticle Toxicity
Physiochemical interactions of the various nanoparticles including silver, titanium, zinc, silica and copper, etc., with any viable cell, are the detriments of their toxic behavior. Variety of toxicity evaluation models like cell lines, zebrafish, rodents, insects (water flea (Daphnia magna) and fruit fly (Drsophila melanogaster) and many others have been used till now [425,426,427]. The exact mechanism of acute or chronic toxicity is still ambiguous but their general mechanisms involve DNA, damage, protein dysfunction and others followed by the generation of reactive species [410, 428].
Furthermore, most of the studies have concluded the ROS generation as the major pathway for NPs toxicity using model organisms [417, 429,430,431]. Yu et al. [432] and Horie & Tabei [433] documented the toxicity of various NPs through generation of ROS in the living cells. In normal physiological state of the body, there is an optimal balance between generation and neutralization of the ROS (singlet oxygen, hypochlorous ions, hydrogen peroxide, superoxide and hydroxyl hydroxyl, etc.) and have a crucial role in cell differentiation, proliferation and death [434, 435]. However, in redox imbalance due to various toxicants, ROS proportion elevates and begins to scavenge the viral macromolecules of the organisms like proteins, carbohydrates, lipids and nucleic acids which ultimately damage the DNA, RNA and various vital enzymes, cytotoxicity and subsequently death [430, 436].
NPs enter the cell through endocytosis and form aggregates. They are considered foreign substances by the body so they may be subjected to phagocytosis by the lysosomes, making their degradation capacity impaired [437]. Further, the toxicity greatly depends on the size, shape, surface area, surface coating and agglomeration. The acute toxicity increases with the decrease in size of NPs [438]. Till now, AgNPs were reported to be most cytotoxic specifically those with the size ≤ 10 nm. They have a tendency to form aggregates in mice organs and induce toxicity, making them a double-edged sword to be used against bacteria [439].
8 Conclusion and Future Perspectives
Metallic biogenic NPs with tunable characteristics like size, shape, and surface area now could produce significant bactericidal potential against a variety of human pathogens. More collaborative work among the scientific community regarding formulation, biosafety, cost-effectiveness, and long-term ecological impact of NPs could lead us to more suitable, economical, and better alternatives to tackle antibiotic resistance with minimum side effects.
Data Availability
All data supporting the findings of this study are available within the review article.
References
S. Bassetti, S. Tschudin-Sutter, A. Egli, M. Osthoff, Optimizing antibiotic therapies to reduce the risk of bacterial resistance. European J. Intern. Med. (2022). https://doi.org/10.1016/j.ejim.2022.01.029
S. Das, R. Samantaray, A. Mallick, S.K. Sahu, S. Sharma, Types of organisms and in-vitro susceptibility of bacterial isolates from patients with microbial keratitis: a trend analysis of 8 years. Indian J. Ophthalmol. 67(1), 49 (2019)
A. Selvaraj, A. Valliammai, C. Sivasankar, M. Suba, G. Sakthivel, S.K. Pandian, Antibiofilm and antivirulence efficacy of myrtenol enhances the antibiotic susceptibility of Acinetobacter baumannii. Sci. Rep. 10(1), 21975 (2020)
A.M. Díez-Pascual, Antibacterial activity of nanomaterials. Nanomaterials 2018(8), 359 (2018)
X. Pang, X. Song, M. Chen, S. Tian, Z. Lu, J. Sun, H.G. Yuk, Combating biofilms of foodborne pathogens with bacteriocins by lactic acid bacteria in the food industry. Compr. Rev. Food Sci. Food Safety 21(2), 1657–1676 (2022)
Q. Xin, H. Shah, A. Nawaz, W. Xie, M.Z. Akram, A. Batool, J.R. Gong, Antibacterial carbon-based nanomaterials. Adv. Mater. 31(45), 1804838 (2019)
G. Dantas, M.O. Sommer, R.D. Oluwasegun, G.M. Church, Bacteria subsisting on antibiotics. Science 320(5872), 100–103 (2008)
M. Lobanovska, G. Pilla, Focus: drug development: Penicillin’s discovery and antibiotic resistance: lessons for the future? Yale J. Biol. Med. 90(1), 135 (2017)
Wuo, M. G., Dulberger, C. L., Brown, R. A., Sturm, A., Ultee, E., Bloom-Ackermann, Z., & Kiessling, L. L. (2022). Antibiotic action revealed by real-time imaging of the mycobacterial membrane. bioRxiv.
M.E. Enany, A.M. Algammal, S.A. Nasef, S.A. Abo-Eillil, M. Bin-Jumah, A.E. Taha, A.A. Allam, The occurrence of the multidrug resistance (MDR) and the prevalence of virulence genes and QACs resistance genes in E. coli isolated from environmental and avian sources. AMB Exp. 9(1), 1–9 (2019)
K. Huang, H. Xia, Y. Zhang, J. Li, G. Cui, F. Li, N. Wu, Elimination of antibiotic resistance genes and human pathogenic bacteria by earthworms during vermicomposting of dewatered sludge by metagenomic analysis. Bioresource Technol. 297, 122451 (2020)
H.H. Kumburu, T. Sonda, M. van Zwetselaar, P. Leekitcharoenphon, O. Lukjancenko, B.T. Mmbaga, G.S. Kibiki, Using WGS to identify antibiotic resistance genes and predict antimicrobial resistance phenotypes in MDR Acinetobacter baumannii in Tanzania. J. Antimicrobial Chemother. 74(6), 1484–1493 (2019)
S. Sanyasi, R.K. Majhi, S. Kumar, M. Mishra, A. Ghosh, M. Suar, L. Goswami, Polysaccharide-capped silver Nanoparticles inhibit biofilm formation and eliminate multi-drug-resistant bacteria by disrupting bacterial cytoskeleton with reduced cytotoxicity towards mammalian cells. Sci. Rep. 6(1), 24929 (2016)
M. Alavi, M.R. Hamblin, J.F. Kennedy, Antimicrobial applications of lichens: secondary metabolites and green synthesis of silver nanoparticles: a review. Nano Micro Biosystems 1(1), 15–21 (2022)
M. Alavi, R. Kowalski, R. Capasso, H. Douglas Melo Coutinho, I. De Rose Alencar Menezes, Various novel strategies for functionalization of gold and silver nanoparticles to hinder drug-resistant bacteria and cancer cells. Micro Nano Bio Aspects 1(1), 38–48 (2022)
M. Alavi, S. Thomas, M. Sreedharan, Modification of silica nanoparticles for antibacterial activities: mechanism of action. Micro Nano Bio Aspects 1(1), 49–58 (2022)
K. Marimuthu, H.K. Gautam, Nanobiotics against antimicrobial resistance: harnessing the power of nanoscale materials and technologies. J. Nanobiotechnol. 20(1), 375 (2022)
D.L. Green, K. Keenan, K.J. Fredricks, S.I. Huque, M.F. Mushi, C. Kansiime, M. Clarkson, The role of multidimensional poverty in antibiotic misuse: a mixed-methods study of self-medication and non-adherence in Kenya, Tanzania, and Uganda. Lancet Glob. Health 11(1), e59–e68 (2023)
V. Kimothi, R.S. Dhariyal, Antibiotic resistance: a review. Int. J. Pharmacy Res. 10(2), 4–12 (2019)
U. Nations, No time to wait: securing the future from drug-resistant infections; report to the secretary-general of the United Nations (WHO, Geneva, 2019)
N. Chakraborty, D. Jha, I. Roy, P. Kumar, S.S. Gaurav, K. Marimuthu, H.K. Gautam, Nanobiotics against antimicrobial resistance: harnessing the power of nanoscale materials and technologies. J. Nanobiotechnol. 20(1), 375 (2022)
R.M. Klevens, M.A. Morrison, J. Nadle, S. Petit, K. Gershman, S. Ray, Active Bacterial Core surveillance (ABCs) MRSA Investigators, Invasive methicillin-resistant Staphylococcus aureus infections in the United States. JAMA 298(15), 1763–1771 (2007)
World Health Organization. Antibiotic resistance (2020). https://www.who.int/news-room/fact-sheets/detail/antibiotic-resistance accessed 11 November 2022).
P.C. Appelbaum, 2012 and beyond: potential for the start of a second pre-antibiotic era? J. Antimicrob. Chemother. 67(9), 2062–2068 (2012)
E. Cox, S. Nambiar, L. Baden, Needed: antimicrobial development. N. Engl. J. Med. 380(8), 783–785 (2019)
O. Pacios, L. Blasco, I. Bleriot, L. Fernandez-Garcia, M. González Bardanca, A. Ambroa, M. Tomás, Strategies to combat multidrug-resistant and persistent infectious diseases. Antibiotics 9(2), 65 (2020)
World Health Organization, Antibacterial Agents in Clinical Development: An Analysis of the Antibacterial Clinical Development Pipeline (World Health Organization, 2019).
D.D. Flannery, K. Chiotos, J.S. Gerber, K.M. Puopolo, Neonatal multidrug-resistant gram-negative infection: epidemiology, mechanisms of resistance, and management. Pediatr. Res. 91(2), 380–391 (2022)
S. Mortazavi-Derazkola, M.A. Ebrahimzadeh, O. Amiri, H.R. Goli, A. Rafiei, M. Kardan, M. Salavati-Niasari, Facile green synthesis and characterization of Crataegus microphylla extract-capped silver nanoparticles (CME@ Ag-NPs) and its potential antibacterial and anticancer activities against AGS and MCF-7 human cancer cells. J. Alloy. Compd. 820, 153186 (2020)
A.A. Moussa, A.F. Md Nordin, R.A. Hamat, A.S. Jasni, High level aminoglycoside resistance and distribution of the resistance genes in Enterococcus faecalis and Enterococcus faecium from teaching hospital in Malaysia. Infection Drug Resist. (2019). https://doi.org/10.2147/IDR.S219544
C.A. Rodriguez, C.D. Mitnick, M.F. Franke, Value of observational data for multidrug-resistant tuberculosis. Lancet Infect. Dis. 19(9), 930–931 (2019)
L. Xu, H.W. Liang, Y. Yang, S.H. Yu, Stability and reactivity: positive and negative aspects for nanoparticle processing. Chem. Rev. 118(7), 3209–3250 (2018)
B. Naseer, G. Srivastava, O.S. Qadri, S.A. Faridi, R.U. Islam, K. Younis, Importance and health hazards of nanoparticles used in the food industry. Nanotechnol. Rev. 7(6), 623–641 (2018)
A.A. Menazea, A.M. Ismail, A. Samy, Novel green synthesis of zinc oxide nanoparticles using orange waste and its thermal and antibacterial activity. J. Inorg. Organomet. Polym. Mater. 31, 4250–4259 (2021)
H. Javid, S. Ahmadi, E. Mohamadian, Therapeutic applications of apigenin and its derivatives: micro and nano aspects. Micro Nano Bio Aspects 2(1), 30–38 (2023)
V. Singh, S. Shrivastava, S.K. Singh, A. Kumar, S. Saxena, Multi-scale temporal convolutional networks and continual learning based in silico discovery of alternative antibiotics to combat multi-drug resistance. Expert Syst. Appl. 215, 119295 (2023)
A. Selmani, D. Kovačević, K. Bohinc, Nanoparticles: From synthesis to applications and beyond. Adv. Coll. Interface. Sci. 303, 102640 (2022)
H. Ge, Y. Wang, X. Zhao, Research on the drug resistance mechanism of foodborne pathogens. Microb. Pathog. 162, 105306 (2022)
J.-H. Lee, Perspectives towards antibiotic resistance: from molecules to population. J. Microbiol. 57(3), 181–184 (2019)
J.H. Lee, Perspectives towards antibiotic resistance: from molecules to population. J. Microbiol. (2019). https://doi.org/10.1007/s12275-019-0718-8
A. Antonoplis, X. Zang, T. Wegner, P.A. Wender, L. Cegelski, Vancomycin–arginine conjugate inhibits growth of carbapenem-resistant E. coli and targets cell-wall synthesis. ACS Chem. Biol. 14(9), 2065–2070 (2019)
H.A. Kadhum, T.H. Hasan, The study of bacillus subtils antimicrobial activity on some of the pathological isolates. Int. J. Drug Deliv. Technol. 9(02), 193–196 (2019)
K. Klobucar, E.D. Brown, New potentiators of ineffective antibiotics: targeting the Gram-negative outer membrane to overcome intrinsic resistance. Curr. Opin. Chem. Biol. 66, 102099 (2022)
A.J. Baylay, L.J. Piddock, M.A. Webber, Molecular mechanisms of antibiotic resistance–Part I, in Bacterial resistance to antibiotics–from molecules to man. (Wiley, Hoboken, 2019), pp.1–26
A.A.J. Aljanaby, I.A.J. Aljanaby, Prevalence of aerobic pathogenic bacteria isolated from patients with burn infection and their antimicrobial susceptibility patterns in Al-Najaf City, Iraq-a three-year cross-sectional study. F1000Research 7(1157), 1157 (2018)
D.I. Andersson, N.Q. Balaban, F. Baquero, P. Courvalin, P. Glaser, U. Gophna, T. Tønjum, Antibiotic resistance: turning evolutionary principles into clinical reality. FEMS Microbiol. Rev. 44(2), 171–188 (2020)
A.H. Holmes, L.S. Moore, A. Sundsfjord, M. Steinbakk, S. Regmi, A. Karkey, L.J. Piddock, Understanding the mechanisms and drivers of antimicrobial resistance. Lancet 387(10014), 176–187 (2016)
I. Álvarez-Rodríguez, L. Arana, B. Ugarte-Uribe, E. Gómez-Rubio, S. Martín-Santamaría, C. Garbisu, I. Alkorta, Type IV coupling proteins as potential targets to control the dissemination of antibiotic resistance. Front. Mol. Biosci. 7, 201 (2020)
A. Magallon, L. Amoureux, T. Garrigos, M. Sonois, V. Varin, C. Neuwirth, J. Bador, Role of AxyABM overexpression in acquired resistance in Achromobacter xylosoxidans. J. Antimicrob. Chemother. 77(4), 926–929 (2022)
F. Han, C. Yu, G. Fu, Warming alters elevation distributions of soil bacterial and fungal communities in alpine grasslands. Global Ecol. Conserv. 39, e02306 (2022)
K.K. Salimiyan Rizi, M. Noghondar, Adaptive antibiotic resistance: overview and perspectives. J. Infect. Dis. Ther 6, 1–3 (2018)
Criswell & Daniel. The “Evolution” of Antibiotic Resistance. Institute for Creation Research. N.p., 2004. Web. 28.)
Y. Liu, Y. Cai, G. Li, W. Wang, P.K. Wong, T. An, Response mechanisms of different antibiotic-resistant bacteria with different resistance action targets to the stress from photocatalytic oxidation. Water Res. 218, 118407 (2022)
F.J. Pérez-Llarena, G. Bou, Proteomics as a tool for studying bacterial virulence and antimicrobial resistance. Front. Microbiol. 7, 410 (2016)
E. Christaki, M. Marcou, A. Tofarides, Antimicrobial resistance in bacteria: mechanisms, evolution, and persistence. J. Mol. Evol. 88(1), 26–40 (2020)
J.M. Munita, C.A. Arias, Mechanisms of antibiotic resistance. Microbiol. Spectrum 4(2), 4 (2016)
M. Cerezales, K. Xanthopoulou, J. Wille, O. Krut, H. Seifert, L. Gallego, P.G. Higgins, Mobile genetic elements harboring antibiotic resistance determinants in. Front. Microbiol. 11, 919 (2020)
J.I. Hwang, J.K. Norsworthy, F. González-Torralva, G.L. Priess, L.T. Barber, T.R. Butts, Non-target-site resistance mechanism of barnyardgrass [Echinochloa crus-galli (L.) P. Beauv] to florpyrauxifen-benzyl. Pest Manag. Sci. 78(1), 287–295 (2022)
Z.T. Laughlin et al., 50S subunit recognition and modification by the Mycobacterium tuberculosis ribosomal RNA methyltransferase TlyA. Proc. Natl. Acad. Sci. 119(14), e2120352119 (2022)
K. Prashanth, T. Vasanth, R. Saranathan, A.R. Makki, S. Pagal, Antibiotic resistance, biofilms and quorum sensing in Acinetobacter species, in Antibiotic resistant bacteria: a continuous challenge in the new millennium. ed. by V. Sjhal (InTech, Orlando, 2012), pp.179–212
M.C. Roberts, Update on acquired tetracycline resistance genes. FEMS Microbiol. Lett. 245(2), 195–203 (2005)
S.B. Southon, S.B. Beres, P. Kachroo, M.O. Saavedra, H. Erlendsdóttir, G. Haraldsson, K.G. Kristinsson, Population genomic molecular epidemiological study of macrolide-resistant Streptococcus pyogenes in Iceland, 1995 to 2016: identification of a large clonal population with a pbp2x mutation conferring reduced in vitro β-lactam susceptibility. J. Clin. Microbiol. 58(9), 10–1128 (2020)
G. Cox, G.D. Wright, Intrinsic antibiotic resistance: mechanisms, origins, challenges and solutions. Int. J. Med. Microbiol. 303(6–7), 287–292 (2013)
X.Z. Li, H. Nikaido, Efflux-mediated drug resistance in bacteria. Drugs 69(12), 1555–1623 (2009)
N.S. Macêdo, Z. de Sousa Silveira, P.P.M. Cordeiro, H.D.M. Coutinho, J.P.S. Júnior, L.J.Q. Júnior, M.V. Da Silva, Inhibition of Staphylococcus aureus efflux pump by O-eugenol and its toxicity in drosophila melanogaster animal model. BioMed Res. Int. (2022). https://doi.org/10.1155/2022/1440996
J. Stephen et al., membrane efflux pumps of pathogenic vibrio species: role in antimicrobial resistance and virulence. Microorganisms 10(2), 382 (2022)
E.B. Breidenstein, C. de la Fuente-Núñez, R.E. Hancock, Pseudomonas aeruginosa: all roads lead to resistance. Trends Microbiol. 19(8), 419–426 (2011)
A. Sharma, R. Sharma, T. Bhattacharyya, T. Bhando, R. Pathania, Fosfomycin resistance in Acinetobacter baumannii is mediated by efflux through a major facilitator superfamily (MFS) transporter—AbaF. J. Antimicrob. Chemother. 72(1), 68–74 (2016)
M. Kok, L. Maton, M. van der Peet, T. Hankemeier, J.C. van Hasselt, Unraveling antimicrobial resistance using metabolomics. Drug Discov. Today (2022). https://doi.org/10.1016/j.drudis.2022.03.015
J. Tan, J. Tay, J. Hedrick, Y.Y. Yang, Synthetic macromolecules as therapeutics that overcome resistance in cancer and microbial infection. Biomaterials 252, 120078 (2020)
S. Santajit, N. Indrawattana, Mechanisms of antimicrobial resistance in ESKAPE pathogens. BioMed Res. Int. (2016). https://doi.org/10.1155/2016/2475067
J. Wang, S. Wang, C. Chen, J. Hu, S. He, Y. Zhou, J. Lin, Treatment of hospital wastewater by electron beam technology: removal of COD, pathogenic bacteria and viruses. Chemosphere 308, 136265 (2022)
C. Aurilio, P. Sansone, M. Barbarisi, V. Pota, L.G. Giaccari, F. Coppolino, M.C. Pace, Mechanisms of action of carbapenem resistance. Antibiotics 11(3), 421 (2022)
L. Fernández, R.E. Hancock, Adaptive and mutational resistance: role of porins and efflux pumps in drug resistance. Clin. Microbiol. Rev. 25(4), 661–681 (2012)
R. Diab, B. Khameneh, O. Joubert, R. Duval, Insights in nanoparticle-bacterium interactions: new frontiers to bypass bacterial resistance to antibiotics. Curr. Pharm. Des. 21(28), 4095–4105 (2015)
H.C. Flemming, E.D. van Hullebusch, T.R. Neu, P.H. Nielsen, T. Seviour, P. Stoodley, S. Wuertz, The biofilm matrix: multitasking in a shared space. Nat. Rev. Microbiol. 21(2), 70–86 (2023)
C. Uruén, G. Chopo-Escuin, J. Tommassen, R.C. Mainar-Jaime, J. Arenas, Biofilms as promoters of bacterial antibiotic resistance and tolerance. Antibiotics 10(1), 3 (2020)
J. Yan, B.L. Bassler, Surviving as a community: antibiotic tolerance and persistence in bacterial biofilms. Cell Host Microbe 26(1), 15–21 (2019)
M.S.A. Khan, M.M. Altaf, I. Ahmad, Chemical nature of biofilm matrix and its significance. Biofilms Plant Soil Health (2017). https://doi.org/10.1002/9781119246329.ch9
M. Majumdar, S.A. Khan, S.C. Biswas, D.N. Roy, A.S. Panja, T.K. Misra, In vitro and in silico investigation of anti-biofilm activity of Citrus macroptera fruit extract mediated silver nanoparticles. J. Mol. Liq. 302, 112586 (2020)
M. Majumdar, S.A. Khan, N.B. Nandi, S. Roy, A.S. Panja, D.N. Roy, T.K. Misra, Green synthesis of iron nanoparticles for investigation of biofilm inhibition property. ChemistrySelect 5(43), 13575–13583 (2020)
M. Majumdar, S. Shivalkar, A. Pal, M.L. Verma, A.K. Sahoo, D.N. Roy, Nanotechnology for enhanced bioactivity of bioactive compounds, in Biotechnological production of bioactive compounds. (Elsevier, Amsterdam, 2020), pp.433–466
S. Majumdar, S. Roy, S. Majumdar, S. Roy, talking about talking microbes, in Microbial communication: mathematical modeling, synthetic biology and the role of noise. (Springer Nature, Berlin, 2020), pp.9–24
D.N. Roy, I. Ahmad, Combating biofilm of ESKAPE pathogens from ancient plant-based therapy to modern nanotechnological combinations, in A complete guidebook on biofilm study. (Academic Press, Cambridge, 2022), pp.59–94
T. Abee, Á.T. Kovács, O.P. Kuipers, S. Van der Veen, Biofilm formation and dispersal in Gram-positive bacteria. Curr. Opin. Biotechnol. 22(2), 172–179 (2011)
C. Solano, M. Echeverz, I. Lasa, Biofilm dispersion and quorum sensing. Curr. Opin. Microbiol. 18, 96–104 (2014)
M.J. Federle, B.L. Bassler, Interspecies communication in bacteria. J. Clin. Investig. 112(9), 1291–1299 (2003)
H.C. Flemming, J. Wingender, U. Szewzyk, P. Steinberg, S.A. Rice, S. Kjelleberg, Biofilms: an emergent form of bacterial life. Nat. Rev. Microbiol. 14(9), 563–575 (2016)
S. Vasudevan, H.A. Joseph, S.S. Swamy, A.P. Solomon, Antibiotic resistance in biofilms, in Introduction to biofilm engineering. (American Chemical Society, Washington, 2019), pp.205–224
X. Zhao, Z. Yu, T. Ding, Quorum-sensing regulation of antimicrobial resistance in bacteria. Microorganisms 8(3), 425 (2020)
W.R. Galloway, J.T. Hodgkinson, S.D. Bowden, M. Welch, D.R. Spring, Quorum sensing in Gram-negative bacteria: small-molecule modulation of AHL and AI-2 quorum sensing pathways. Chem. Rev. 111(1), 28–67 (2011)
Y. Xiao, H. Zou, J. Li, T. Song, W. Lv, W. Wang, S. Tao, Impact of quorum sensing signaling molecules in gram-negative bacteria on host cells: current understanding and future perspectives. Gut Microbes 14(1), 2039048 (2022)
V.S. Bhatt, Quorum sensing mechanisms in gram positive bacteria, in Implication of quorum sensing system in biofilm formation and virulence. (Springer Singapore, Singapore, 2018), pp.297–311
G. Banerjee, A.K. Ray, Quorum-sensing network-associated gene regulation in Gram-positive bacteria. Acta Microbiol. Immunol. Hung. 64(4), 439–453 (2017)
A. Piketh, H. Alam, A. Ahmad, Quorum sensing as an alternative approach to combatting multidrug resistance, in Non-traditional approaches to combat antimicrobial drug resistance. (Springer Nature Singapore, Singapore, 2023), pp.191–220
W.H. Zhao, Z.Q. Hu, β-lactamases identified in clinical isolates of Pseudomonas aeruginosa. Crit. Rev. Microbiol. 36(3), 245–258 (2010)
M. Uwate, Y.K. Ichise, A. Shirai, T. Omasa, T. Nakae, H. Maseda, Two routes of MexS-MexT-mediated regulation of MexEF-OprN and MexAB-OprM efflux pump expression in Pseudomonas aeruginosa. Microbiol. Immunol. 57(4), 263–272 (2013)
Z. Pang, R. Raudonis, B.R. Glick, T.J. Lin, Z. Cheng, Antibiotic resistance in Pseudomonas aeruginosa: mechanisms and alternative therapeutic strategies. Biotechnol. Adv. 37(1), 177–192 (2019)
M.H. Al-Agamy, A. Aljallal, H.H. Radwan, A.M. Shibl, Characterization of carbapenemases, ESBLs, and plasmid-mediated quinolone determinants in carbapenem-insensitive Escherichia coli and Klebsiella pneumoniae in Riyadh hospitals. J. Infect. Public Health 11(1), 64–68 (2018)
A. Stewart, P. Harris, A. Henderson, D. Paterson, Treatment of infections by OXA-48-producing enterobacteriaceae. Antimicrob. Agents Chemother. 62(11), e01195-e1218 (2018)
M.C.P. Nguyen, P.L. Woerther, M. Bouvet, A. Andremont, R. Leclercq, A. Canu, Escherichia coli as reservoir for macrolide resistance genes. Emerg. Infect. Dis. 15(10), 1648 (2009)
M. Xu, Y.N. Zhou, B.P. Goldstein, D.J. Jin, Cross-resistance of Escherichia coli RNA polymerases conferring rifampin resistance to different antibiotics. J. Bacteriol. 187(8), 2783–2792 (2005)
Mounsey, O. J. (2021). Characterization of Relationships Between Fluoroquinolone-Resistant E. coli from Humans, Dogs, and Dairy Cattle Living in South West England (Doctoral dissertation, University of Bristol).
L.S. Tzouvelekis, A. Markogiannakis, M. Psichogiou, P.T. Tassios, G.L. Daikos, Carbapenemases in Klebsiella pneumoniae and other Enterobacteriaceae: an evolving crisis of global dimensions. Clin. Microbiol. Rev. 25(4), 682–707 (2012)
R.L. Ferreira, B.C. Da Silva, G.S. Rezende, R. Nakamura-Silva, A. Pitondo-Silva, E.B. Campanini, M.C.D.S. Pranchevicius, High prevalence of multidrug-resistant Klebsiella pneumoniae harboring several virulence and β-lactamase encoding genes in a Brazilian intensive care unit. Front. Microbiol. 9, 3198 (2019)
Y. Doi, Y. Arakawa, 16S ribosomal RNA methylation: emerging resistance mechanism against aminoglycosides. Clin. Infect. Dis. 45(1), 88–94 (2007)
M.W. Vetting, C.H. Park, S.S. Hegde, G.A. Jacoby, D.C. Hooper, J.S. Blanchard, Mechanistic and structural analysis of aminoglycoside N-acetyltransferase AAC (6′)-Ib and its bifunctional, fluoroquinolone-active AAC (6′)-Ib-cr variant. Biochemistry 47(37), 9825–9835 (2008)
S. Navon-Venezia, K. Kondratyeva, A. Carattoli, Klebsiella pneumoniae: a major worldwide source and shuttle for antibiotic resistance. FEMS Microbiol. Rev. 41(3), 252–275 (2017)
Z.K. Sheng, F. Hu, W. Wang, Q. Guo, Z. Chen, X. Xu, M. Wang, Mechanisms of tigecycline resistance among Klebsiella pneumoniae clinical isolates. Antimicrob. Agents Chemother. 58(11), 6982–6985 (2014)
G. Wang, G. Zhao, X. Chao, L. Xie, H. Wang, The characteristic of virulence, biofilm and antibiotic resistance of Klebsiella pneumoniae. Int. J. Environ. Res. Public Health 17(17), 6278 (2020)
L. Poirel, P. Nordmann, Carbapenem resistance in Acinetobacter baumannii: mechanisms and epidemiology. Clin. Microbiol. Infect. 12(9), 826–836 (2006)
W.F. Penwell, A.B. Shapiro, R.A. Giacobbe, R.F. Gu, N. Gao, J. Thresher, A.A. Miller, Molecular mechanisms of sulbactam antibacterial activity and resistance determinants in Acinetobacter baumannii. Antimicrobial Agents Chemother. 59(3), 1680–1689 (2015)
Q. Chen, X. Li, H. Zhou, Y. Jiang, Y. Chen, X. Hua, Y. Yu, Decreased susceptibility to tigecycline in Acinetobacter baumannii mediated by a mutation in trm encoding SAM-dependent methyltransferase. J. Antimicrob. Chemother. 69(1), 72–76 (2014)
M. Asif, I.A. Alvi, S.U. Rehman, Insight into Acinetobacter baumannii: pathogenesis, global resistance, mechanisms of resistance, treatment options, and alternative modalities. Infection Drug Resist. 11, 1249 (2018)
R. Vázquez-López, S.G. Solano-Gálvez, J.J. Juárez Vignon-Whaley, J.A. Abello Vaamonde, L.A. Padró Alonzo, A. Rivera Reséndiz, T. Barrientos Fortes, Acinetobacter baumannii resistance: a real challenge for clinicians. Antibiotics 9(4), 205 (2020)
S. Jamal, A. Al Atrouni, R. Rafei, F. Dabboussi, M. Hamze, M. Osman, Molecular mechanisms of antimicrobial resistance in Acinetobacter baumannii, with a special focus on its epidemiology in Lebanon. J. Global Antimicrob. Resist. 15, 154–163 (2018)
B. Jubeh, Z. Breijyeh, R. Karaman, Resistance of gram-positive bacteria to current antibacterial agents and overcoming approaches. Molecules 25(12), 2888 (2020)
W.T. Liu, E.Z. Chen, L. Yang, C. Peng, Q. Wang, Z. Xu, D.Q. Chen, Emerging resistance mechanisms for 4 types of common anti-MRSA antibiotics in Staphylococcus aureus: a comprehensive review. Microb. Pathog. 156, 104915 (2021)
A. Sommer, S. Fuchs, F. Layer, C. Schaudinn, R.E. Weber, H. Richard, B. Strommenger, Mutations in the gdpP gene are a clinically relevant mechanism for β-lactam resistance in meticillin-resistant Staphylococcus aureus lacking mec determinants. Microbial Genomics (2021). https://doi.org/10.1099/mgen.0.000623
A. Kumar, M. Kaushal, Progression of β-lactam resistance in Staphylococcus aureus, in Insights into drug resistance in Staphylococcus aureus. (IntechOpen, London, 2021)
M. Afzal, A.K. Vijay, F. Stapleton, M. Willcox, The relationship between ciprofloxacin resistance and genotypic changes in S. aureus ocular isolates. Pathogens 11(11), 1354 (2022)
S. Rajabi, A. Shivaee, M.A. Khosravi, M. Eshaghi, S. Shahbazi, F. Hosseini, Evaluation of multidrug efflux pump expression in clinical isolates of Staphylococcus aureus. Gene Reports 18, 100537 (2020)
I.Y. Yoo, O.K. Kang, H.J. Shim, H.J. Huh, N.Y. Lee, Linezolid resistance in methicillin-resistant Staphylococcus aureus in Korea: high rate of false resistance to linezolid by the VITEK 2 system. Ann. Lab. Med. 40(1), 57–62 (2020)
Y. Zhu, C. Wang, S. Schwarz, W. Liu, Q. Yang, T. Luan, W. Zhang, Identification of a novel tetracycline resistance gene, tet (63), located on a multiresistance plasmid from Staphylococcus aureus. J. Antimicrobial Chemother. 76(3), 576–581 (2021)
T.C. Dewé, J.C. D’Aeth, N.J. Croucher, Genomic epidemiology of penicillin-non-susceptible Streptococcus pneumoniae. Microbial Genomics (2019). https://doi.org/10.1099/mgen.0.000305
C.Y. Wang, Y.H. Chen, C. Fang, M.M. Zhou, H.M. Xu, C.M. Jing, C.H. Zhang, Antibiotic resistance profiles and multidrug resistance patterns of Streptococcus pneumoniae in pediatrics: a multicenter retrospective study in mainland China. Medicine (2019). https://doi.org/10.1097/MD.0000000000015942
A.L. Bloemendaal, E.C. Brouwer, A.C. Fluit, Methicillin resistance transfer from Staphylocccus epidermidis to methicillin-susceptible Staphylococcus aureus in a patient during antibiotic therapy. PLoS ONE 5(7), e11841 (2010)
A. Brenciani, E. Tiberi, E. Tili, M. Mingoia, C. Palmieri, P.E. Varaldo, E. Giovanetti, Genetic determinants and elements associated with antibiotic resistance in viridans group streptococci. J. Antimicrob. Chemother. 69(5), 1197–1204 (2014)
C. Sinel, M. Cacaci, P. Meignen, F. Guérin, B.W. Davies, M. Sanguinetti, V. Cattoir, Subinhibitory concentrations of ciprofloxacin enhance antimicrobial resistance and pathogenicity of Enterococcus faecium. Antimicrobial Agents Chemother. 61(5), 10–1128 (2017)
X. Du, X. Hua, T. Qu, Y. Jiang, Z. Zhou, Y. Yu, Molecular characterization of Rifr mutations in Enterococcus faecalis and Enterococcus faecium. J. Chemother. 26(4), 217–221 (2014)
M. Khodabandeh, M. Mohammadi, M.R. Abdolsalehi, M. Hasannejad-Bibalan, M. Gholami, A. Alvandimanesh, R. Rajabnia, High-level aminoglycoside resistance in Enterococcus faecalis and Enterococcus faecium; as a serious threat in hospitals. Infectious Disorders-Drug 20(2), 223–228 (2020)
M. Georges, E. Odoyo, D. Matano, F. Tiria, C. Kyany’a, D. Mbwika, L. Musila, Determination of Enterococcus faecalis and Enterococcus faecium antimicrobial resistance and virulence factors and their association with clinical and demographic factors in Kenya. J. Pathog. (2022). https://doi.org/10.1155/2022/3129439
W. Wehbeh, R. Rojas-Diaz, X. Li, N. Mariano, L. Grenner, S. Segal-Maurer, J.J. Rahal, Fluoroquinolone-resistant Streptococcus agalactiae: epidemiology and mechanism of resistance. Antimicrob. Agents Chemother. 49(6), 2495–2497 (2005)
Nauta, K. M. (2021). The dissection of Β-lactam resistance in Bacillus anthracis, Bacillus cereus, and Bacillus thuringiensis (Doctoral dissertation, The University of Iowa).
M. Hennart, L.G. Panunzi, C. Rodrigues, Q. Gaday, S.L. Baines, M. Barros-Pinkelnig, S. Brisse, Population genomics and antimicrobial resistance in Corynebacterium diphtheriae. Genome Med. 12, 1–18 (2020)
L.T. Matereke, A.I. Okoh, Listeria monocytogenes virulence, antimicrobial resistance and environmental persistence: a review. Pathogens 9(7), 528 (2020)
R.T. Aruleba, T.A. Adekiya, B.E. Oyinloye, P. Masamba, L.S. Mbatha, A. Pretorius, A.P. Kappo, PZQ therapy: how close are we in the development of effective alternative anti-schistosomal drugs? Infect. Disorders-Drug Targets 19(4), 337–349 (2019)
J. Dey, S.R. Mahapatra, T.K. Raj, T. Kaur, P. Jain, A. Tiwari, M. Suar, Designing a novel multi-epitope vaccine to evoke a robust immune response against pathogenic multidrug-resistant Enterococcus faecium bacterium. Gut Pathogens 14(1), 1–20 (2022)
L.D.R. Dos Santos, J.P.R. Furlan, M.S. Ramos, I.F.L. Gallo, L.V.P. de Freitas, E.G. Stehling, Co-occurrence of mcr-1, mcr-3, mcr-7 and clinically relevant antimicrobial resistance genes in environmental and fecal samples. Arch. Microbiol. 202, 1795–1800 (2020)
A. Haslam, J. Gill, V. Prasad, Estimation of the percentage of US patients with cancer who are eligible for immune checkpoint inhibitor drugs. JAMA Netw. Open 3(3), e200423–e200423 (2020)
S. Roy, I. Hasan, B. Guo, Recent advances in nanoparticle-mediated antibacterial applications. Coord. Chem. Rev. 482, 215075 (2023)
U. Anand, M. Carpena, M. Kowalska-Góralska, P. Garcia-Perez, K. Sunita, E. Bontempi, J. Simal-Gandara, Safer plant-based nanoparticles for combating antibiotic resistance in bacteria: a comprehensive review on its potential applications, recent advances, and future perspective. Sci. Total. Environ. 821, 153472 (2022)
Y. Chen, X. Zheng, Y. Xie, C. Ding, H. Ruan, C. Fan, Anti-bacterial and cytotoxic properties of plasma sprayed silver-containing HA coatings. J. Mater. Sci. - Mater. Med. 19, 3603–3609 (2008)
H. Li, H. Xu, Y.L. Yang, X.L. Yang, Y. Wu, S. Zhang, H.L. Song, Effects of graphite and Mn ore media on electro-active bacteria enrichment and fate of antibiotic and corresponding resistance gene in up flow microbial fuel cell constructed wetland. Water Res. 165, 114988 (2019)
H. Cui, A.L. Smith, Impact of engineered nanoparticles on the fate of antibiotic resistance genes in wastewater and receiving environments: a comprehensive review. Environ. Res. 204, 112373 (2022)
A. Gupta, S. Mumtaz, C.H. Li, I. Hussain, V.M. Rotello, Combatting antibiotic-resistant bacteria using nanomaterials. Chem. Soc. Rev. 48(2), 415–427 (2019)
S. Mumtaz, S. Ali, S.A.R. Kazmi, T.A. Mughal, S. Mumtaz, H.M. Tahir, M.I. Rashid, Analysis of the antimicrobial potential of sericin-coated silver nanoparticles against human pathogens. Microscopy Res. Tech. (2022). https://doi.org/10.1002/jemt.24273
M. Summer, H.M. Tahir, S. Ali, Sonication and heat-mediated synthesis, characterization and larvicidal activity of sericin-based silver nanoparticles against dengue vector (Aedes aegypti). Microsc. Res. Tech. (2023). https://doi.org/10.1002/jemt.24333
M. Summer, H.M. Tahir, S. Ali, R. Abaidullah, S. Mumtaz, S. Nawaz, Bactericidal potential of different size sericin-capped silver nanoparticles synthesized by heat, light, and sonication. J. Basic Microbiol. (2023). https://doi.org/10.1002/jobm.202200632
H.M. Tahir, F. Saleem, S. Ali, Q.U. Ain, A. Fazal, M. Summer, G. Murtaza, Synthesis of sericin-conjugated silver nanoparticles and their potential antimicrobial activity. J. Basic Microbiol. 60(5), 458–467 (2020)
S. Muzammil, S. Hayat, M. Fakhar-E-Alam, B. Aslam, M.H. Siddique, M.A. Nisar, Z. Wang, Nanoantibiotics: future nanotechnologies to combat antibiotic resistance. Front Biosci 10, 352–374 (2018)
X. Zhao, H. Tang, X. Jiang, Deploying gold nanomaterials in combating multi-drug-resistant bacteria. ACS Nano 16(7), 10066–10087 (2022)
M.J. Hajipour, K.M. Fromm, A.A. Ashkarran, D.J. de Aberasturi, IR d. Larramendi, T. Rojo, V. Serpooshan, WJ Parak and M. Mahmoudi. Trends Biotechnol. 30, 499–511 (2012)
E.P. Ivanova, J. Hasan, H.K. Webb, G. Gervinskas, S. Juodkazis, V.K. Truong, R.J. Crawford, Bactericidal activity of black silicon. Nat. Commun. (2013). https://doi.org/10.1038/ncomms3838
S. Medici, M. Peana, V.M. Nurchi, M.A. Zoroddu, Medical uses of silver: history, myths, and scientific evidence. J. Med. Chem. 62(13), 5923–5943 (2019)
A. Hamad, K.S. Khashan, A. Hadi, Silver nanoparticles and silver ions as potential antibacterial agents. J. Inorg. Organomet. Polym. Mater. 30(12), 4811–4828 (2020)
H.H. Lara, L. Ixtepan-Turrent, M. Jose Yacaman, J. Lopez-Ribot, Inhibition of Candida auris biofilm formation on medical and environmental surfaces by silver nanoparticles. ACS Appl. Mater. Interfaces 12(19), 21183–21191 (2020)
T.A.J. de Souza, L.R.R. Souza, L.P. Franchi, Silver nanoparticles: An integrated view of green synthesis methods, transformation in the environment, and toxicity. Ecotoxicol. Environ. Saf. 171, 691–700 (2019)
D.R. Ibraheem, N.N. Hussein, G.M. Sulaiman, H.A. Mohammed, R.A. Khan, O. Al Rugaie, Ciprofloxacin-loaded silver nanoparticles as potent nano-antibiotics against resistant pathogenic bacteria. Nanomaterials 12(16), 2808 (2022)
G.R. Tortella, O. Rubilar, N. Durán, M.C. Diez, M. Martínez, J. Parada, A.B. Seabra, Silver nanoparticles: toxicity in model organisms as an overview of its hazard for human health and the environment. J. Hazard. Mater. 390, 121974 (2020)
S. Ali, S. Perveen, M. Ali, T. Jiao, A.S. Sharma, H. Hassan, Q. Chen, Bioinspired morphology-controlled silver nanoparticles for antimicrobial application. Mater. Sci. Eng. C 108, 110421 (2020)
M. Nilavukkarasi, S. Vijayakumar, S.P. Kumar, Biological synthesis and characterization of silver nanoparticles with Capparis zeylanica L. leaf extract for potent antimicrobial and anti-proliferation efficiency. Mater. Sci. Energy Technol. 3, 371–376 (2020)
L.H. Abdel-Rahman, B.S. Al-Farhan, D. Abou El-ezz, M.A. Abd-El Sayed, M.M. Zikry, A.M. Abu-Dief, Green biogenic synthesis of silver nanoparticles using aqueous extract of Moringa oleifera: access to a powerful antimicrobial, anticancer, pesticidal and catalytic agents. J. Inorg. Organomet. Polym. Mater. 32(4), 1422–1435 (2022)
N. Feroze, B. Arshad, M. Younas, M.I. Afridi, S. Saqib, A. Ayaz, Fungal mediated synthesis of silver nanoparticles and evaluation of antibacterial activity. Microsc. Res. Tech. 83(1), 72–80 (2020)
H. Ji, S. Zhou, Y. Fu, Y. Wang, J. Mi, T. Lu, C. Lü, Size-controllable preparation and antibacterial mechanism of thermo-responsive copolymer-stabilized silver nanoparticles with high antimicrobial activity. Mater. Sci. Eng. C 110, 110735 (2020)
M.J. Sweet, I. Singleton, Silver nanoparticles: a microbial perspective. Adv. Appl. Microbiol. 77, 115–133 (2011)
G. Franci, A. Falanga, S. Galdiero, L. Palomba, M. Rai, G. Morelli, M. Galdiero, Silver nanoparticles as potential antibacterial agents. Molecules 20(5), 8856–8874 (2015)
M. Ramzan, M.I. Karobari, A. Heboyan, R.N. Mohamed, M. Mustafa, S.N. Basheer, B. Zeshan, Synthesis of silver nanoparticles from extracts of wild ginger (Zingiber zerumbet) with antibacterial activity against selective multidrug resistant oral bacteria. Molecules 27(6), 2007 (2022)
P. Korshed, L. Li, D.-T. Ngo, T. Wang, Effect of storage conditions on the long-term stability of bactericidal effects for laser generated silver nanoparticles. Nanomaterials 8, 218 (2018)
W. Ma, L. Jing, A. Valladares, S.L. Mehta, Z. Wang, P. Andy Li, J.J. Bang, Silver nanoparticle exposure induced mitochondrial stress, caspase-3 activation and cell death: Amelioration by sodium selenite. Int. J. Biol. Sci. 11, 860–867 (2015)
P. Aramwit, N. Bang, J. Ratanavaraporn, S. Ekgasit, Green synthesis of silk sericin-capped silver nanoparticles and their potent anti-bacterial activity. Nanoscale Res. Lett. 9(1), 1–7 (2014)
J.M. DeSimone, Practical approaches to green solvents. Science 297(5582), 799–803 (2002)
S.H. Lee, B.H. Jun, Silver nanoparticles: synthesis and application for nanomedicine. Int. J. Mol. Sci. 20(4), 865 (2019)
V. Lazar, Quorum sensing in biofilms—How to destroy the bacterial citadels or their cohesion/power? Anaerobe 17, 280–285 (2011)
S. Periasamy, H.S. Joo, A.C. Duong, T.H.L. Bach, V.Y. Tan, S.S. Chatterjee, M. Otto, How Staphylococcus aureus biofilms develop their characteristic structure. Proc. National Acad. Sci. 109(4), 1281–1286 (2012)
L. Krce, M. Šprung, T. Rončević, A. Maravić, V. ČikešČulić, D. Blažeka, I. Aviani, Probing the mode of antibacterial action of silver nanoparticles synthesized by laser ablation in water: what fluorescence and AFM data tell us. Nanomaterials 10(6), 1040 (2020)
M. Rai, K. Kon, A. Ingle, N. Duran, S. Galdiero, M. Galdiero, Broad-spectrum bioactivities of silver nanoparticles: the emerging trends and future prospects. Appl. Microbiol. Biotechnol. 98(5), 1951–1961 (2014)
A. Salleh, R. Naomi, N.D. Utami, A.W. Mohammad, E. Mahmoudi, N. Mustafa, M.B. Fauzi, The potential of silver nanoparticles for antiviral and antibacterial applications: a mechanism of action. Nanomaterials 10(8), 1566 (2020)
T.L. Collins, E.A. Markus, D.J. Hassett, J.B. Robinson, The effect of a cationic porphyrin on Pseudomonas aeruginosa biofilms. Curr. Microbiol. 61, 411–416 (2010)
D. Wu, W. Fan, A. Kishen, J.L. Gutmann, B. Fan, Evaluation of the antibacterial efficacy of silver nanoparticles against Enterococcus faecalis biofilm. Journal of endodontics 40(2), 285–290 (2014)
L.A. Tamayo, P.A. Zapata, N.D. Vejar, M.I. Azócar, M.A. Gulppi, X. Zhou, M.A. Páez, Release of silver and copper nanoparticles from polyethylene nanocomposites and their penetration into Listeria monocytogenes. Mater. Sci. Eng. C 40, 24–31 (2014)
Z. Chen, Z. Zhang, X. Zhai, Y. Li, L. Lin, H. Zhao, G. Lin, Rapid and sensitive detection of anti-SARS-CoV-2 IgG, using lanthanide-doped nanoparticles-based lateral flow immunoassay. Anal. Chem. 92(10), 7226–7231 (2020)
D. Muchintala, V. Suresh, D. Raju, R.B. Sashidhar, Synthesis and characterization of cecropin peptide-based silver nanocomposites: its antibacterial activity and mode of action. Mater. Sci. Eng. C 110, 110712 (2020)
Y. Cai, D. Wu, X. Zhu, W. Wang, F. Tan, J. Chen, X. Qiao, X. Qiu, Sol-gel preparation of Ag-doped MgO nanoparticles with high efficiency for bacterial inactivation. Ceram. Int. 43(1), 1066–1072 (2017)
S. Silver, Bacterial silver resistance: molecular biology and uses and misuses of silver compounds. FEMS Microbiol. Rev. 27(2–3), 341–353 (2003)
J.P. Rolim, M.A. de Melo, S.F. Guedes, F.B. Albuquerque-Filho, J.R. de Souza, N.A. Nogueira, I.C. Zanin, L.K. Rodrigues, The antimicrobial activity of photodynamic therapy against Streptococcus mutansusing different photosensitizers. J. Photochem. Photobiol. B. 106, 40–46 (2012)
N. Beyth, Y. Houri-Haddad, A. Domb, W. Khan, R. Hazan, Alternative antimicrobial approach: nano-antimicrobial materials. Evidence-Based Complement. Altern. Med. (2015). https://doi.org/10.1155/2015/246012
N.R. Bury, F. Galvez, C.M. Wood, Effects of chloride, calcium, and dissolved organic carbon on silver toxicity: comparison between rainbow trout and fathead minnows. Environ. Toxicol. Chem.: Int. J. 18(1), 56–62 (1999)
J.R. Morones, J.L. Elechiguerra, A. Camacho, K. Holt, J.B. Kouri, J.T. Ramirez, M.J. Yacaman, The bactericidal effect of silver nanoparticles. Nanotechnology 16, 2346–2353 (2005)
W.K. Jung, H.C. Koo, K.W. Kim, S. Shin, S.H. Kim, Y.H. Park, Antibacterial activity and mechanism of action of the silver ion in Staphylococcus aureus and Escherichia coli. Appl. Environ. Microbiol. 74, 2171–2178 (2008)
L. Chun-Nam, C.M. Ho, R. Chen, Q.Y. He, W.Y. Yu, H.P. Sun, K.H. Tam, J.F. Chiu, C.M. Che, Proteomic analysis of the mode of antibacterial action of silver nanoparticles. J. Proteome Res. 5, 916–924 (2006)
P. Jena, M. Bhattacharya, G. Bhattacharjee, B. Satpati, P. Mukherjee, D. Senapati, R. Srinivasan, Bimetallic gold–silver nanoparticles mediate bacterial killing by disrupting the actin cytoskeleton MreB. Nanoscale 12(6), 3731–3749 (2020)
L. Wu, G. Zhu, X. Zhang, Y. Si, Silver nanoparticles inhibit denitrification by altering the viability and metabolic activity of Pseudomonas stutzeri. Sci. Total. Environ. 706, 135711 (2020)
M.S. Yousaf, A. Haider, A. Shahzadi, A. Ul-Hamid, M. Imran, M.A. Khan, M. Ikram, Aggrandized catalytic and bactericidal activity of silver and polyvinylpyrrolidone capped bismuth oxybromide quantum dots: in silico molecular docking studies. J. Inorg. Organomet. Polym. Mater. (2023). https://doi.org/10.1007/s10904-023-02821-7
A.M. Elgorban, A.H. Bahkali, M.A. El-Metwally, M. Elsheshtawi, M.A. Abdel-Wahab, In vitro antifungal activity of some plant essential oils. Int. J. Pharmcol. 11(1), 56–61 (2015)
R.S. Shaban, A.H. Bahkali, M.B. Marwa, Antibacterial activity of biogenic silver nanoparticles produced by Aspergillus terreus. Int. J. Pharmacol. 11, 858–863 (2015)
J. Narware, R.N. Yadav, C. Keswani, S.P. Singh, H.B. Singh, Silver nanoparticle based biopesticides for phytopathogens: scope and potential in agriculture. Nano-Biopesticides Today Future Perspect. 2019, 303–314 (2019)
N. Kaur, A. Singh, W. Ahmad, Microwave assisted green synthesis of silver nanoparticles and its application: a review. J. Inorg. Organomet. Polym. Mater. 33(3), 663–672 (2023)
A. Pal, R. Goswami, D.N. Roy, A critical assessment on biochemical and molecular mechanisms of toxicity developed by emerging nanomaterials on important microbes. Environ. Nanotechnol. Monit. Manag. 16, 100485 (2021)
M.A. Biel, C. Sievert, M. Usacheva, M. Teichert, E. Wedell, N. Loebel, …& Zimmermann, R., Reduction of endotracheal tube biofilms using antimicrobial photodynamic therapy. Lasers Surg. Med. 43(7), 586–590 (2011)
U.F. Gunputh, H. Le, K. Lawton, A. Besinis, C. Tredwin, R.D. Handy, Antibacterial properties of silver nanoparticles grown in situ and anchored to titanium dioxide nanotubes on titanium implant against Staphylococcus aureus. Nanotoxicology 14(1), 97–110 (2020)
J.S. Kim, E. Kuk, K.N. Yu, J.H. Kim, S.J. Park, H.J. Lee, S.H. Kim, Y.K. Park, Y.H. Park, C.Y. Hwang, Antimicrobial effects of silver nanoparticles. Nanomedicine 3, 95–101 (2007)
K. Shameli, M.B. Ahmad, S.D. Jazayeri, P. Shabanzadeh, P. Sangpour, H. Jahangirian, Y. Gharayebi, Investigation of antibacterial properties silver nanoparticles prepared via green method. Chem. Cent. J. 6(1), 73 (2012)
D.A. Kumar, V. Palanichamy, S.M. Roopan, Green synthesis of silver nanoparticles using alternanthera dentata leaf extract at room temperature and their antimicrobialactivity. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 127, 168–171 (2014)
S. Naraginti, A. Sivakumar, Eco-friendly synthesis of silver and gold nanoparticles with enhanced bactericidal activity and study of silver catalyzed reduction of 4-nitrophenol. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 128, 357–362 (2014)
P. Parvekar, J. Palaskar, S. Metgud, R. Maria, S. Dutta, The minimum inhibitory concentration (MIC) and minimum bactericidal concentration (MBC) of silver nanoparticles against Staphylococcus aureus. Biomater. Investigations Dentistry 7(1), 105–109 (2020)
G.R. Salunke, S. Ghosh, R.J. Santosh Kumar, S. Khade, P. Vashisth, T. Kale, S. Chopade, V. Pruthi, G. Kundu, J.R. Bellare, Rapid efficient synthesis and characterization of silver, gold, and bimetallic nanoparticles from the medicinal plant Plumbago zeylanicaand their application in biofilm control. Int. J. Nanomed. 9, 2635–2653 (2014)
S. Wang, R. Su, S. Nie, M. Sun, J. Zhang, D. Wu, N. Moustaid-Moussa, Application of nanotechnology in improving bioavailability and bioactivity of diet-derived phytochemicals. J. Nutr. Biochem. 25(4), 363–376 (2014)
S.D. Nishu, J.H. No, T.K. Lee, Transcriptional response and plant growth promoting activity of Pseudomonas fluorescens DR397 under drought stress conditions. Microbiol. Spectrum 10(4), e00979-e1022 (2022)
A.M. Badawy, R. Silva, B. Morris, K.G. Scheckel, M.T. Suidan, T.M. Tolaymat, Surface charge-dependent toxicity of silver nanoparticles. Environ. Sci. Technol. 45, 283–287 (2011). https://doi.org/10.1021/es1034188
D. Swolana, R.D. Wojtyczka, Activity of silver nanoparticles against Staphylococcus spp. Int. J. Mol. Sci. 23(8), 4298 (2022)
X. Gu, Q. Cheng, P. He, Y. Zhang, Z. Jiang, Y. Zeng, Dihydroartemisinin-loaded chitosan nanoparticles inhibit the rifampicin-resistant mycobacterium tuberculosis by disrupting the cell wall. Front. Microbiol. 12, 735166 (2021)
Estevez, H., Palacios, A., Gil, D., Anguita, J., Vallet-Regi, M., González, B., ... & Luque-Garcia, J. L. (2020). Antimycobacterial effect of selenium nanoparticles on Mycobacterium tuberculosis. Frontiers in microbiology, 11, 800.
S. Djearamane, Z.C. Loh, J.J. Lee, L.S. Wong, R. Rajamani, P.A. Luque, S.X.T. Liang, Remedial aspect of zinc oxide nanoparticles against Serratia marcescens and Enterococcus faecalis. Front. Pharmacol. (2022). https://doi.org/10.3389/fphar.2022.891304
A. Abdelghafar, N. Yousef, M. Askoura, Zinc oxide nanoparticles reduce biofilm formation, synergize antibiotics action and attenuate Staphylococcus aureus virulence in host; an important message to clinicians. BMC Microbiol. 22(1), 1–17 (2022)
L. Shkodenko, I. Kassirov, E. Koshel, Metal oxide nanoparticles against bacterial biofilms: perspectives and limitations. Microorganisms 8(10), 1545 (2020)
A.K. Keshari, R. Srivastava, P. Singh, V.B. Yadav, G. Nath, Antioxidant and antibacterial activity of silver nanoparticles synthesized by Cestrum nocturnum. J. Ayurveda Integrative Med. 11(1), 37–44 (2020)
U. Halder, R.K. Roy, R. Biswas, D. Khan, K. Mazumder, R. Bandopadhyay, Synthesis of copper oxide nanoparticles using capsular polymeric substances produced by Bacillus altitudinis and investigation of its efficacy to kill pathogenic Pseudomonas aeruginosa. Chem. Eng. J. Adv. 11, 100294 (2022)
Y. Zhang, X. Pan, S. Liao, C. Jiang, L. Wang, Y. Tang, L. Chen, Quantitative proteomics reveals the mechanism of silver nanoparticles against multidrug-resistant Pseudomonas aeruginosa biofilms. J. Proteome Res. 19(8), 3109–3122 (2020)
A. Das Mahapatra, C. Patra, J. Mondal, C. Sinha, P. Chandra Sadhukhan, D. Chattopadhyay, Silver nanoparticles derived from Albizia lebbeck bark extract demonstrate killing of multidrug-resistant bacteria by damaging cellular architecture with antioxidant activity. Chem. Select 5(15), 4770–4777 (2020)
M. Saberpour, S. Najar-Peeraye, S. Shams, B. Bakhshi, Effects of chitosan nanoparticles loaded with mesenchymal stem cell conditioned media on gene expression in Vibrio cholerae and Caco-2 cells. Sci. Rep. 12(1), 1–9 (2022)
R. Manjumeena, D. Duraibabu, J. Sudha, P.T. Kalaichelvan, Biogenic nanosilver incorporated reverse osmosis membrane for antibacterial and antifungal activities against selected pathogenic strains: an enhanced eco-friendly water disinfection approach. J. Environ. Sci.: Health ToxicHazard Substance Environ. England 49, 1125–1133 (2014)
M.H. Siddique, B. Aslam, M. Imran, A. Ashraf, H. Nadeem, S. Hayat, U. Qureshi, Effect of silver nanoparticles on biofilm formation and eps production of multidrug-resistant Klebsiella pneumoniae. BioMed. Res. Int. (2020). https://doi.org/10.1155/2020/6398165
Pareek, V., Devineau, S., Sivasankaran, S. K., Bhargava, A., Panwar, J., Srikumar, S., & Fanning, S. (2020). Silver nanoparticles induce a triclosan-like antibacterial action mechanism in multi-drug resistant Klebsiella pneumoniae. bioRxiv.
M.E. Lysakowska, A. Ciebiada-Adamiec, L. Klimek, M. Sienkiewicz, The activity of silver nanoparticles (Axonnite) on clinical and environmental strains of Acinetobacter spp. Burns 41, 364–371 (2015)
T.A. Salih, K.T. Hassan, S.R. Majeed, I.J. Ibraheem, O.M. Hassan, A.S. Obaid, In vitro scolicidal activity of synthesised silver nanoparticles from aqueous plant extract against Echinococcus granulosus. Biotechnol. Rep. 28, e00545 (2020)
D.K. Singaravelu, D.N. Binjawhar, F. Ameen, A. Veerappan, Lectin-fortified cationic copper sulfide nanoparticles gain dual targeting capabilities to treat carbapenem-resistant Acinetobacter baumannii infection. ACS Omega 7(48), 43934–43944 (2022)
S. Sharma, V.K. Singh, A. Kumar, S. Mallubhotla, Effect of nanoparticles on oxidative damage and antioxidant defense system in plants. Mole. Plant Abiotic Stress: Biol. Biotechnol. (2019). https://doi.org/10.1002/9781119463665.ch17
E. Tvrdá, F. Benko, Free radicals: what they are and what they do, in Pathology. (Academic Press, Cambridge, 2020), pp.3–13
N. Ghosh, A. Das, S. Chaffee, S. Roy, C.K. Sen, Reactive oxygen species, oxidative damage and cell death, in Immunity and inflammation in health and disease. (Academic Press, Cambridge, 2018), pp.45–55
R.Z. Zhao, S. Jiang, L. Zhang, Z.B. Yu, Mitochondrial electron transport chain, ROS generation and uncoupling. Int. J. Mol. Med. 44(1), 3–15 (2019)
R. Canaparo, F. Foglietta, T. Limongi, L. Serpe, Biomedical applications of reactive oxygen species generation by metal nanoparticles. Materials 14(1), 53 (2020)
M. Alavi, R. Yarani, ROS and RNS modulation: the main antimicrobial, anticancer, antidiabetic, and antineurodegenerative mechanisms of metal or metal oxide nanoparticles. Nanotheranostics Treat. Nano Micro Biosyst. 2, 22–30 (2023)
K.S. Siddiqi, A. Husen, R.A. Rao, A review on biosynthesis of silver nanoparticles and their biocidal properties. J. Nanobiotechnol. 16(1), 1–28 (2018)
A. Abdal Dayem, M.K. Hossain, S.B. Lee, K. Kim, S.K. Saha, G.M. Yang, S.G. Cho, The role of reactive oxygen species (ROS) in the biological activities of metallic nanoparticles. Int. J. Mol. Sci. 18(1), 120 (2017)
O. Metryka, D. Wasilkowski, A. Mrozik, Evaluation of the effects of Ag, Cu, ZnO and TiO2 nanoparticles on the expression level of oxidative stress-related genes and the activity of antioxidant enzymes in Escherichia coli, Bacillus cereus and Staphylococcus epidermidis. Int. J. Mol. Sci. 23(9), 4966 (2022)
D. Manzanares, V. Ceña, Endocytosis: the nanoparticle and submicron nanocompounds gateway into the cell. Pharmaceutics 12(4), 371 (2020)
J. Mosquera, I. García, L.M. Liz-Marzán, Cellular uptake of nanoparticles versus small molecules: a matter of size. Acc. Chem. Res. 51(9), 2305–2313 (2018)
Q. Chen, N. Wang, M. Zhu, J. Lu, H. Zhong, X. Xue, H. Yin, TiO2 nanoparticles cause mitochondrial dysfunction, activate inflammatory responses, and attenuate phagocytosis in macrophages: a proteomic and metabolomic insight. Redox Biol. 15, 266–276 (2018)
T. Xia, M. Kovochich, J. Brant, M. Hotze, J. Sempf, T. Oberley, A.E. Nel, Comparison of the abilities of ambient and manufactured nanoparticles to induce cellular toxicity according to an oxidative stress paradigm. Nano Lett. 6(8), 1794–1807 (2006)
S.J. Soenen, P. Rivera-Gil, J.M. Montenegro, W.J. Parak, S.C. De Smedt, K. Braeckmans, Cellular toxicity of inorganic nanoparticles: common aspects and guidelines for improved nanotoxicity evaluation. Nano Today 6(5), 446–465 (2011)
P.V. AshaRani, G. Low Kah Mun, M.P. Hande, S. Valiyaveettil, Cytotoxicity and genotoxicity of silver nanoparticles in human cells. ACS Nano 3(2), 279–290 (2009)
K.B. Holt, A.J. Bard, Interaction of silver (I) ions with the respiratory chain of Escherichia coli: an electrochemical and scanning electrochemical microscopy study of the antimicrobial mechanism of micromolar Ag+. Biochemistry 44(39), 13214–13223 (2005)
A. Manke, L. Wang, Y. Rojanasakul, Mechanisms of nanoparticle-induced oxidative stress and toxicity. BioMed Res. Int. (2013). https://doi.org/10.1155/2013/942916
S. Zhang, T. Ouyang, B.M. Reinhard, Multivalent ligand-nanoparticle conjugates amplify reactive oxygen species second messenger generation and enhance epidermal growth factor receptor phosphorylation. Bioconjug. Chem. 33(9), 1716–1728 (2022)
T.C. Dakal, A. Kumar, R.S. Majumdar, V. Yadav, Mechanistic basis of antimicrobial actions of silver nanoparticles. Front. Microbiol. 7, 1831 (2016)
Y. Gao, M.A.V. Anand, V. Ramachandran, V. Karthikkumar, V. Shalini, S. Vijayalakshmi, D. Ernest, Biofabrication of zinc oxide nanoparticles from Aspergillus niger, their antioxidant, antimicrobial and anticancer activity. J. Cluster Sci. 30(4), 937–946 (2019)
K.S. Khashan, G.M. Sulaiman, S.A. Hussain, T.R. Marzoog, M.S. Jabir, Synthesis, characterization and evaluation of anti-bacterial, anti-parasitic and anti-cancer activities of aluminum-doped zinc oxide nanoparticles. J. Inorg. Organomet. Polym. Mater. 30(9), 3677–3693 (2020)
S. Shahid, S.A. Khan, W. Ahmad, U. Fatima, S. Knawal, Size-dependent bacterial growth inhibition and antibacterial activity of Ag-doped ZnO nanoparticles under different atmospheric conditions. Indian J. Pharm. Sci. 80(1), 173–180 (2018)
H.M. Yusof, R. Mohamad, U.H. Zaidan, Abdul Rahman NA Microbial synthesis of zinc oxide nanoparticles and their potential application as an antimicrobial agent and a feed supplement in animal industry: a review. J. Anim. Sci. Biotechnol 10, 57 (2019)
N.A.A. Yusof, N.M. Zain, N. Pauzi, Synthesis of ZnO nanoparticles with chitosan as stabilizing agent and their antibacterial properties against Gram-positive and Gram-negative bacteria. Int. J. Biol. Macromol. 124, 1132–1136 (2019)
X. Zhu, J. Wang, L. Cai, Y. Wu, M. Ji, H. Jiang, J. Chen, Dissection of the antibacterial mechanism of zinc oxide nanoparticles with manipulable nanoscale morphologies. J. Hazard. Mater. 430, 128436 (2022)
S.E. Jin, H.E. Jin, Synthesis, characterization, and three-dimensional structure generation of zinc oxide-based nanomedicine for biomedical applications. Pharmaceutics 11(11), 575 (2019)
E. Sánchez-López, D. Gomes, G. Esteruelas, L. Bonilla, A.L. Lopez-Machado, R. Galindo, E.B. Souto, Metal-based nanoparticles as antimicrobial agents: an overview. Nanomaterials 10(2), 292 (2020)
A. Sirelkhatim, S. Mahmud, A. Seeni, N.H.M. Kaus, L.C. Ann, S.K.M. Bakhori, D. Mohamad, Review on zinc oxide nanoparticles: antibacterial activity and toxicity mechanism. Nano-Micro Lett. 7(3), 219–242 (2015)
M. Moharramnejad, A. Ehsani, S. Salmani, M. Shahi, R.E. Malekshah, Z.S. Robatjazi, H. Parsimehr, Zinc-based metal-organic frameworks: synthesis and recent progress in biomedical application. J. Inorg. Organomet. Polym. Mater. 32(9), 3339–3354 (2022)
M. Pal, Nanotechnology: a new approach in food packaging. J Food Microbiol Safety Hyg 2, 121 (2017)
V. Tiwari, N. Mishra, K. Gadani, P.S. Solanki, N.A. Shah, M. Tiwari, Mechanism of anti-bacterial activity of zinc oxide nanoparticle against carbapenem-resistant Acinetobacter baumannii. Front. Microbiol. 9, 1218 (2018)
D.K. Tiwari, J. Behari, P. Sen, Time and dose-dependent antimicrobial potential of Ag nanoparticles synthesized by top-down approach. Curr. Sci. 95, 647–655 (2008)
P.J.P. Espitia, N.D.F.F. Soares, J.S. dos Reis Coimbra, N.J. de Andrade, R.S. Cruz, E.A.A. Medeiros, Food Bioprocess Tech. 5, 1447–1464 (2012)
B. Abebe, H.A. Murthy, E. Amare, Enhancing the photocatalytic efficiency of ZnO: defects, heterojunction, and optimization. Environ. Nanotechnol. Monit. Manag. 14, 100336 (2020)
B.A. Fahimmunisha, R. Ishwarya, M.S. AlSalhi, S. Devanesan, M. Govindarajan, B. Vaseeharan, Green fabrication, characterization and antibacterial potential of zinc oxide nanoparticles using Aloe socotrina leaf extract: a novel drug delivery approach. J. Drug Deliv. Sci. Technol. 55, 101465 (2020)
B.L. da Silva, B.L. Caetano, B.G. Chiari-Andréo, R.C.L.R. Pietro, L.A. Chiavacci, Increased antibacterial activity of ZnO nanoparticles: Influence of size and surface modification. Colloids Surf. B 177, 440–447 (2019)
K. Dulta, G. Koşarsoy Ağçeli, P. Chauhan, R. Jasrotia, P.K. Chauhan, A novel approach of synthesis zinc oxide nanoparticles by bergenia ciliata rhizome extract: antibacterial and anticancer potential. J. Inorg. Organomet. Polym. Mater. 31, 180–190 (2021)
S.S.N. Fernando, T.D.C.P. Gunasekara, J. Holton, Antimicrobial nanoparticles: applications and mechanisms of action. Sri Lankan J. Infec. Dis. (2018). https://doi.org/10.4038/sljid.v8i1.8167
P.T.L. Huong, N. Van Quang, M.T. Tran, D.Q. Trung, D.T.B. Hop, T.T.H. Tam, V.D. Dao, Excellent visible light photocatalytic degradation and mechanism insight of Co2+-doped ZnO nanoparticles. Appl. Phys. A 128(1), 1–16 (2022)
A. Joe, S.H. Park, K.D. Shim, D.J. Kim, K.H. Jhee, H.W. Lee, E.S. Jang, Antibacterial mechanism of ZnO nanoparticles under dark conditions. J. Ind. Eng. Chem. 45, 430–439 (2017)
K.R. Raghupathi, R.T. Koodali, A.C. Manna, Size-dependent bacterial growth inhibition and mechanism of antibacterial activity of zinc oxide nanoparticles. Langmuir 27(7), 4020–4028 (2011)
L.K. Adams, D.Y. Lyon, P.J. Alvarez, Comparative eco-toxicity of nanoscale TiO2, SiO2, and ZnO water suspensions. Water Res. 40(19), 3527–3532 (2006)
K. Hirota, M. Sugimoto, M. Kato, K. Tsukagoshi, T. Tanigawa, H. Sugimoto, Preparation of zinc oxide ceramics with a sustainable antibacterial activity under dark conditions. Ceram. Int. 36(2), 497–506 (2010)
V. Lakshmi Prasanna, R. Vijayaraghavan, Insight into the mechanism of antibacterial activity of ZnO: surface defects mediated reactive oxygen species even in the dark. Langmuir 31(33), 9155–9162 (2015)
E. Jeong, C.U. Kim, J. Byun, J. Lee, H.E. Kim, E.J. Kim, S.W. Hong, Quantitative evaluation of the antibacterial factors of ZnO nanorod arrays under dark conditions: physical and chemical effects on Escherichia coli inactivation. Sci. Total. Environ. 712, 136574 (2020)
H.T. Hoang, T.T.T. Nguyen, H.M. Do, T.K.N. Nguyen, H.T. Pham, A novel finding of intra-genus inhibition of quorum sensing in Vibrio bacteria. Sci. Rep. 12(1), 15203 (2022)
S. Khanna, D.S. Pardi, C.R. Kelly, C.S. Kraft, T. Dhere, M.R. Henn, E.L. Hohmann, A novel microbiome therapeutic increases gut microbial diversity and prevents recurrent Clostridium difficile infection. J. Infectious Dis. 214(2), 173–181 (2016)
N. Thakur, P. Manna, J. Das, Synthesis and biomedical applications of nanoceria, a redox active nanoparticle. J. Nanobiotechnol. 17(1), 1–27 (2019)
S. Tang, J. Zheng, Antibacterial activity of silver nanoparticles: structural effects. Adv. Healthcare Mater. 7(13), 1701503 (2018)
Y. Xie, Y. He, P.L. Irwin, T. Jin, X. Shi, Antibacterial activity and mechanism of action of zinc oxide nanoparticles against Campylobacter jejuni. Appl. Environ. Microbiol. 77(7), 2325–2331 (2011)
E.A. Campbell, R. Greenwell, J.R. Anthony, S. Wang, L. Lim, K. Das, S.A. Darst, A conserved structural module regulates transcriptional responses to diverse stress signals in bacteria. Mole. Cell 27(5), 793–805 (2007)
M. Cerasi, S. Ammendola, A. Battistoni, Competition for zinc binding in the host-pathogen interaction. Front. Cell. Infect. Microbiol. 3, 108 (2013)
M.J. Hakeem, J. Feng, A. Nilghaz, L. Ma, H.C. Seah, M.E. Konkel, X. Lu, Active packaging of immobilized zinc oxide nanoparticles controls Campylobacter jejuni in raw chicken meat. Appl. Environ. Microbiol. 86(22), e01195-e1220 (2020)
U. Kadiyala, E.S. Turali-Emre, J.H. Bahng, N.A. Kotov, J.S. VanEpps, Unexpected insights into antibacterial activity of zinc oxide nanoparticles against methicillin resistant Staphylococcus aureus (MRSA). Nanoscale 10(10), 4927–4939 (2018)
B. Lallo da Silva, M.P. Abuçafy, E. Berbel Manaia, J.A. Oshiro Junior, B.G. Chiari-Andréo, R.C.R. Pietro, L.A. Chiavacci, Relationship between structure and antimicrobial activity of zinc oxide nanoparticles: an overview. Int. J. Nanomed. (2019). https://doi.org/10.2147/IJN.S216204
D.E. Navarro-López, R. Garcia-Varela, O. Ceballos-Sanchez, A. Sanchez-Martinez, G. Sanchez-Ante, K. Corona-Romero, E.R. López-Mena, Effective antimicrobial activity of ZnO and Yb-doped ZnO nanoparticles against Staphylococcus aureus and Escherichia coli. Mater. Sci. Eng. C 123, 112004 (2021)
M. Li, L. Zhu, D. Lin, Toxicity of ZnO nanoparticles to Escherichia coli: mechanism and the influence of medium components. Environ. Sci. Technol. 45(5), 1977–1983 (2011)
J. Pasquet, Y. Chevalier, E. Couval, D. Bouvier, M.A. Bolzinger, Zinc oxide as a new antimicrobial preservative of topical products: Interactions with common formulation ingredients. Int. J. Pharm. 479(1), 88–95 (2015)
Y. Li, W. Zhang, J. Niu, Y. Chen, Mechanism of photogenerated reactive oxygen species and correlation with the antibacterial properties of engineered metal-oxide nanoparticles. ACS Nano 6(6), 5164–5173 (2012)
A. Akbar, M.B. Sadiq, I. Ali, N. Muhammad, Z. Rehman, M.N. Khan, A.K. Anal, Synthesis and antimicrobial activity of zinc oxide nanoparticles against foodborne pathogens Salmonella typhimurium and Staphylococcus aureus. Biocatal. Agric. Biotechnol. 17, 36–42 (2019)
L. Palanikumar, S.N. Ramasamy, C. Balachandran, Size-dependent antimicrobial response of zinc oxide nanoparticles. IET Nanobiotechnol. 8(2), 111–117 (2014)
M.A. Al-Holy, L.F. Castro, H.M. Al-Qadiri, Inactivation of Cronobacter spp. (Enterobacter sakazakii) in infant formula using lactic acid, copper sulfate and monolaurin. Lett. Appl. Microbiol. 50(3), 246–251 (2010)
G. Faúndez, M. Troncoso, P. Navarrete, G. Figueroa, Antimicrobial activity of copper surfaces against suspensions of Salmonella enterica and Campylobacter jejuni. BMC Microbiol. 4(1), 1–7 (2004)
D.S. Idris, A. Roy, Biogenic synthesis of Ag–CuO nanoparticles and its antibacterial, antioxidant, and catalytic activity. J. Inorg. Organomet. Polym. Mater. (2023). https://doi.org/10.1007/s10904-023-02873-9
L. Weaver, H.T. Michels, C.W. Keevil, Potential for preventing spread of fungi in air-conditioning systems constructed using copper instead of aluminum. Lett. Appl. Microbiol. 50(1), 18–23 (2010)
M. Vincent, R.E. Duval, P. Hartemann, M. Engels-Deutsch, Contact killing and antimicrobial properties of copper. J. Appl. Microbiol. 124(5), 1032–1046 (2018)
M.I. Devi, N. Nallamuthu, N. Rajini, T.S.M. Kumar, S. Siengchin, A.V. Rajulu, N. Ayrilmis, Biodegradable poly (propylene) carbonate using in-situ generated CuNPs coated Tamarindus indica filler for biomedical applications. Mater. Today Commun. 19, 106–113 (2019)
M. Hasanin, M.A. Al Abboud, M.M. Alawlaqi, T.M. Abdelghany, A.H. Hashem, Ecofriendly synthesis of biosynthesized copper nanoparticles with starch-based nanocomposite: antimicrobial, antioxidant, and anticancer activities. Biol. Trace Elem. Res. 200(5), 2099–2112 (2022)
A.P. Ingle, N. Duran, M. Rai, Bioactivity, mechanism of action, and cytotoxicity of copper-based nanoparticles: a review. Appl. Microbiol. Biotechnol. 98(3), 1001–1009 (2014)
G. Wang, W. Jin, A.M. Qasim, A. Gao, X. Peng, W. Li, P.K. Chu, Antibacterial effects of titanium embedded with silver nanoparticles based on electron-transfer-induced reactive oxygen species. Biomaterials 124, 25–34 (2017)
L. Wang, C. Hu, L. Shao, The antimicrobial activity of nanoparticles: present situation and prospects for the future. Int. J. Nanomed. 12, 1227 (2017)
M.F. Gutiérrez, P. Malaquias, V. Hass, T.P. Matos, L. Lourenço, A. Reis, P.V. Farago, The role of copper nanoparticles in an etch-and-rinse adhesive on antimicrobial activity, mechanical properties and the durability of resin-dentine interfaces. J. Dent. 61, 12–20 (2017)
N. Jayarambabu, A. Akshaykranth, T.V. Rao, K.V. Rao, R.R. Kumar, Green synthesis of Cu nanoparticles using Curcuma longa extract and their application in antimicrobial activity. Mater. Lett. 259, 126813 (2020)
Q. Maqbool, S. Iftikhar, M. Nazar, F. Abbas, A. Saleem, T. Hussain, N. Jabeen, Green fabricated CuO nanobullets via Olea europaea leaf extract shows auspicious antimicrobial potential. IET Nanobiotechnol. 11(4), 463–468 (2017)
Y. Yuan, Y. Wu, V. Chinnadurai, M. Saravanan, A. Chinnathambi, S.A. Alharbi, A. Pugazhendhi, In vitro analysis of green synthesized copper nanoparticles using Chloroxylon swietenia leaves for dye degradation and antimicrobial application. Food Chem. Toxicol. 168, 113367 (2022)
K.Y. Yoon, J.H. Byeon, J.H. Park, J. Hwang, Susceptibility constants of Escherichia coli and Bacillus subtilis to silver and copper nanoparticles. Sci. Total. Environ. 373(2–3), 572–575 (2007)
M.S. Usman, M.E. El Zowalaty, K. Shameli, N. Zainuddin, M. Salama, N.A. Ibrahim, Synthesis, characterization, and antimicrobial properties of copper nanoparticles. Int. J. Nanomedicine 8, 4467–4479 (2013)
K. Cheirmadurai, S. Biswas, R. Murali, P. Thanikaivelan, Green synthesis of copper nanoparticles and conducting nanobiocomposites using plant and animal sources. RSC Adv. 4(37), 19507–19511 (2014)
M. Pérez-Alvarez, G. Cadenas-Pliego, O. Pérez-Camacho, V.E. Comparán-Padilla, C.J. Cabello-Alvarado, E. Saucedo-Salazar, Green synthesis of copper nanoparticles using cotton. Polymers 13(12), 1906 (2021)
K.S. Siddiqi, A. Husen, Current status of plant metabolite-based fabrication of copper/copper oxide nanoparticles and their applications: a review. Biomater. Res. 24, 1–15 (2020)
J.Y. Song, H.K. Jang, B.S. Kim, Biological synthesis of gold nanoparticles using Magnolia kobus and Diopyros kaki leaf extracts. Process Biochem. 44(10), 1133–1138 (2009)
K. Gopalakrishnan, C. Ramesh, V. Ragunathan, M. Thamilselvan, Antibacterial activity of Cu2O nanoparticles on E. coli synthesized from Tridax procumbens leaf extract and surface coating with polyaniline. Digest J. Nanomater. Biostruct. 7(2), 833–839 (2012)
M. Raffi, S. Mehrwan, T.M. Bhatti, J.I. Akhter, A. Hameed, W. Yawar, Investigations into the antibacterial behavior of copper nanoparticles against Escherichia coli. Annals of microbiology 60(1), 75–80 (2010)
S. Rajeshkumar, S. Menon, S.V. Kumar, M.M. Tambuwala, H.A. Bakshi, M. Mehta, K. Dua, Antibacterial and antioxidant potential of biosynthesized copper nanoparticles mediated through Cissus arnotiana plant extract. J. Photochem. Photobiol. B: Biol. 197, 111531 (2019)
S. Saleem, B. Ahmed, M.S. Khan, M. Al-Shaeri, J. Musarrat, Inhibition of growth and biofilm formation of clinical bacterial isolates by NiO nanoparticles synthesized from Eucalyptus globulus plants. Microb. Pathog. 111, 375–387 (2017)
T. Foteva, N. Georgieva, Antimicrobial properties of silica/hydroxypropylcellulose hybrids doped with copper ions. J. Chem. Metall. 57(5), 930–936 (2022)
M. Rai, A.P. Ingle, R. Pandit, P. Paralikar, S. Shende, I. Gupta, S.S. da Silva, Copper and copper nanoparticles: role in management of insect-pests and pathogenic microbes. Nanotechnol. Rev. 7(4), 303–315 (2018)
G. Ren, D. Hu, E.W. Cheng, M.A. Vargas-Reus, P. Reip, R.P. Allaker, Characterization of copper oxide nanoparticles for antimicrobial applications. Int. J. Antimicrob. Agents 33(6), 587–590 (2009)
A.K. Chatterjee, R. Chakraborty, T. Basu, Mechanism of antibacterial activity of copper nanoparticles. Nanotechnology 25(13), 135101 (2014)
J.H. Kim, H. Cho, S.E. Ryu, M.U. Choi, Effects of metal ions on the activity of protein tyrosine phosphatase VHR: highly potent and reversible oxidative inactivation by Cu2+ ion. Arch. Biochem. Biophys. 382(1), 72–80 (2000)
Q. Lv, B. Zhang, X. Xing, Y. Zhao, R. Cai, W. Wang, Q. Gu, Biosynthesis of copper nanoparticles using Shewanella loihica PV-4 with antibacterial activity: novel approach and mechanisms investigation. J. Hazard. Mater. 347, 141–149 (2018)
H. Zhang, Z. Ji, T. Xia, H. Meng, C. Low-Kam, R. Liu, A.E. Nel, Use of metal oxide nanoparticle band gap to develop a predictive paradigm for oxidative stress and acute pulmonary inflammation. ACS Nano 6(5), 4349–4368 (2012)
W. Xie, S. Zhang, F. Pan, S. Chen, L. Zhong, J. Wang, X. Pei, Nanomaterial-based ROS-mediated strategies for combating bacteria and biofilms. J. Mater. Res. 36, 822–845 (2021)
G. Applerot, J. Lellouche, A. Lipovsky, Y. Nitzan, R. Lubart, A. Gedanken, E. Banin, Understanding the antibacterial mechanism of CuO nanoparticles: revealing the route of induced oxidative stress. Small 8(21), 3326–3337 (2012)
S. Banerjee, K. Vishakha, S. Das, P.D. Sangma, S. Mondal, A. Ganguli, Oxidative stress, DNA, and membranes targets as modes of antibacterial and antibiofilm activity of facile synthesized biocompatible keratin-copper nanoparticles against multidrug resistant uro-pathogens. World J. Microbiol. Biotechnol. 38(2), 1–16 (2022)
R.K. Swarnkar, J.K. Pandey, K.K. Soumya, P. Dwivedi, S. Sundaram, S. Prasad, R. Gopal, Enhanced antibacterial activity of copper/copper oxide nanowires prepared by pulsed laser ablation in water medium. Appl. Phys. A 122(7), 1–7 (2016)
C. Kaweeteerawat, P. Na Ubol, S. Sangmuang, S. Aueviriyavit, R. Maniratanachote, Mechanisms of antibiotic resistance in bacteria mediated by silver nanoparticles. J. Toxicol. Environ. Health A 80(23–24), 1276–1289 (2017)
O. Metryka, D. Wasilkowski, A. Mrozik, Insight into the antibacterial activity of selected metal nanoparticles and alterations within the antioxidant defence system in Escherichia coli, Bacillus cereus and Staphylococcus epidermidis. Int. J. Mol. Sci. 22(21), 11811 (2021)
P. Sharma, D. Goyal, B. Chudasama, Antibacterial activity of colloidal copper nanoparticles against Gram-negative (Escherichia coli and Proteus vulgaris) bacteria. Lett. Appl. Microbiol. 74(5), 695–706 (2022)
J. Li, K. Rong, H. Zhao, F. Li, Z. Lu, R. Chen, Highly selective antibacterial activities of silver nanoparticles against Bacillus subtilis. J. Nanosci. Nanotechnol. 13(10), 6806–6813 (2013)
D.N. Phan, N. Dorjjugder, Y. Saito, M.Q. Khan, A. Ullah, X. Bie, I.S. Kim, Antibacterial mechanisms of various copper species incorporated in polymeric nanofibers against bacteria. Mater. Today Commun. 25, 101377 (2020)
H. Li, Q. Chen, J. Zhao, K. Urmila, Enhancing the antimicrobial activity of natural extraction using the synthetic ultrasmall metal nanoparticles. Sci. Rep. 5(1), 1–13 (2015)
L. Yadav, R.M. Tripathi, R. Prasad, R.N. Pudake, J. Mittal, Antibacterial activity of Cu nanoparticles against E. coli, Staphylococcus aureus and Pseudomonas aeruginosa. Nano Biomed. Eng 9(1), 9–14 (2017)
C.L. de Dicastillo, M.G. Correa, F.B. Martínez, C. Streitt, M.J. Galotto, Antimicrobial effect of titanium dioxide nanoparticles, in Antimicrobial resistance-a one health perspectivem. (IntechOpen, London, 2020)
A. Kösemen, Z.A. Kösemen, B. Canimkubey, M. Erkovan, F. Başarir, S.E. San, A.V. Tunç, Fe doped TiO2 thin film as electron selective layer for inverted solar cells. Sol. Energy 132, 511–517 (2016)
S. Sagadevan, S. Vennila, P. Singh, J.A. Lett, W.C. Oh, S. Paiman, P.K. Obulapuram, Exploration of the antibacterial capacity and ethanol sensing ability of Cu-TiO2 nanoparticles. J. Exp. Nanosci. 15(1), 337–349 (2020)
M.A. Sebak, T.F. Qahtan, G.M. Asnag, E.M. Abdallah, The role of TiO2 nanoparticles in the structural, thermal and electrical properties and antibacterial activity of PEO/PVP blend for energy storage and antimicrobial application. J. Inorg. Organomet. Polym. Mater. 32(12), 4715–4728 (2022)
J.T. Seil, T.J. Webster, Antimicrobial applications of nanotechnology: methods and literature. Int. J. Nanomed. 7, 2767 (2012)
A. Ansari, V.U. Siddiqui, W.U. Rehman, M.K. Akram, W.A. Siddiqi, A.M. Alosaimi, M.A. Hussein, M. Rafatullah, Green synthesis of TiO2 nanoparticles using acorus calamus leaf extract and evaluating its photocatalytic and in vitro antimicrobial activity. Catalysts 12, 181 (2022). https://doi.org/10.3390/catal12020181
I. Santana, H. Wu, P. Hu, J.P. Giraldo, Targeted delivery of nanomaterials with chemical cargoes in plants enabled by a biorecognition motif. Nat. Commun. 11(1), 1–12 (2020)
S. Albukhaty, L. Al-Bayati, H. Al-Karagoly, S. Al-Musawi, Preparation and characterization of titanium dioxide nanoparticles and in vitro investigation of their cytotoxicity and antibacterial activity against Staphylococcus aureus and Escherichia coli. Animal Biotechnol. (2020). https://doi.org/10.1080/10495398.2020.1842751
M.N. Alomary, M.A. Ansari, proanthocyanin-capped biogenic TiO2 nanoparticles with enhanced penetration, antibacterial and ROS mediated inhibition of bacteria proliferation and biofilm formation: a comparative approach. Chem. European J. 27(18), 5817–5829 (2021)
U.L.N.H. Senarathna, S.S.N. Fernando, T.D.C.P. Gunasekara, M.M. Weerasekera, H.G.S.P. Hewageegana, N.D.H. Arachchi, P.M. Jayaweera, Enhanced antibacterial activity of TiO2 nanoparticle surface modified with Garcinia zeylanica extract. Chem. Central J. 11(1), 1–8 (2017)
M. Azizi-Lalabadi, A. Ehsani, B. Divband, M. Alizadeh-Sani, Antimicrobial activity of Titanium dioxide and Zinc oxide nanoparticles supported in 4A zeolite and evaluation the morphological characteristic. Sci. Rep. 9(1), 1–10 (2019)
C. López de Dicastillo, C. Patiño, M.J. Galotto, J.L. Palma, D. Alburquenque, J. Escrig, Novel antimicrobial titanium dioxide nanotubes obtained through a combination of atomic layer deposition and electrospinning technologies. Nanomaterials 8(2), 128 (2018)
M.F. Song, Y.S. Li, H. Kasai, K. Kawai, Metal nanoparticle-induced micronuclei and oxidative DNA damage in mice. J. Clin. Biochem. Nutr. 50(3), 211–216 (2012)
T.J. Battin, F.V. Kammer, A. Weilhartner, S. Ottofuelling, T. Hofmann, Nanostructured TiO2: transport behavior and effects on aquatic microbial communities under environmental conditions. Environ. Sci. Technol. 43(21), 8098–8104 (2009)
A. Kumar, A.K. Pandey, S.S. Singh, R. Shanker, A. Dhawan, Engineered ZnO and TiO2 nanoparticles induce oxidative stress and DNA damage leading to reduced viability of Escherichia coli. Free Radical Biol. Med. 51(10), 1872–1881 (2011)
C. Pagnout, S. Jomini, M. Dadhwal, C. Caillet, F. Thomas, P. Bauda, Role of electrostatic interactions in the toxicity of titanium dioxide nanoparticles toward Escherichia coli. Colloids Surf. B 92, 315–321 (2012)
J. Rajkumari, C.M. Magdalane, B. Siddhardha, J. Madhavan, G. Ramalingam, N.A. Al-Dhabi, K. Kaviyarasu, Synthesis of titanium oxide nanoparticles using Aloe barbadensis mill and evaluation of its antibiofilm potential against Pseudomonas aeruginosa PAO1. J. Photochem. Photobiol. B: Biol. 201, 111667 (2019)
P. Maheswari, S. Ponnusamy, S. Harish, M.R. Ganesh, Y. Hayakawa, Hydrothermal synthesis of pure and bio modified TiO2: characterization, evaluation of antibacterial activity against gram positive and gram negative bacteria and anticancer activity against KB Oral cancer cell line. Arab. J. Chem. 13(1), 3484–3497 (2020)
M. Ovais, A.T. Khalil, M. Ayaz, I. Ahmad, S.K. Nethi, S. Mukherjee, Biosynthesis of metal nanoparticles via microbial enzymes: a mechanistic approach. Int. J. Mol. Sci. 19(12), 4100 (2018)
J. Hou, L. Wang, C. Wang, S. Zhang, H. Liu, S. Li, X. Wang, Toxicity and mechanisms of action of titanium dioxide nanoparticles in living organisms. J. Environ. Sci. 75, 40–53 (2019)
E.B. Kurutas, The importance of antioxidants which play the role in cellular response against oxidative/nitrosative stress: current state. Nutr. J. 15(1), 1–22 (2015)
S. Sagadevan, S. Imteyaz, B. Murugan, J.A. Lett, N. Sridewi, G.K. Weldegebrieal, W.C. Oh, A comprehensive review on green synthesis of titanium dioxide nanoparticles and their diverse biomedical applications. Green Proc. Synthesis 11(1), 44–63 (2022)
B. Sohm, F. Immel, P. Bauda, C. Pagnout, Insight into the primary mode of action of TiO2 nanoparticles on Escherichia coli in the dark. Proteomics 15(1), 98–113 (2015)
M. Nemattalab, M. Rohani, M. Evazalipour, Z. Hesari, Formulation of Cinnamon (Cinnamomum verum) oil loaded solid lipid nanoparticles and evaluation of its antibacterial activity against multi-drug resistant Escherichia coli. BMC Complement. Med. Therapies 22(1), 1–10 (2022)
C. Pagnout, A. Razafitianamaharavo, B. Sohm, C. Caillet, A. Beaussart, E. Delatour, J.F. Duval, Osmotic stress and vesiculation as key mechanisms controlling bacterial sensitivity and resistance to TiO2 nanoparticles. Commun. Biol. 4(1), 1–15 (2021)
S. Khan, M. Ul-Islam, W.A. Khattak, M.W. Ullah, J.K. Park, Bacterial cellulose-titanium dioxide nanocomposites: nanostructural characteristics, antibacterial mechanism, and biocompatibility. Cellulose 22(1), 565–579 (2015)
S. Shaikh, N. Nazam, S.M.D. Rizvi, K. Ahmad, M.H. Baig, E.J. Lee, I. Choi, Mechanistic insights into the antimicrobial actions of metallic nanoparticles and their implications for multidrug resistance. Int. J. Mol. Sci. 20(10), 2468 (2019)
N.B.A. Abdulrahman, Z.M. Nssaif, Antimicrobial activity of zinc oxide, titanium dioxide and silver nanoparticles against mithicillin-resistant Staphylococcus aureus Isolates. Tikrit J. Pure Sci. 21(3), 49–53 (2016)
E.T. Bekele, B.A. Gonfa, O.A. Zelekew, H.H. Belay, F.K. Sabir, Synthesis of titanium oxide nanoparticles using root extract of Kniphofia foliosa as a template, characterization, and its application on drug resistance bacteria. J. Nanomater. (2020). https://doi.org/10.1155/2020/2817037
S. Albukhaty, L. Al-Bayati, H. Al-Karagoly, S. Al-Musawi, Preparation and characterization of titanium dioxide nanoparticles and in vitro investigation of their cytotoxicity and antibacterial activity against Staphylococcus aureus and Escherichia coli. Anim. Biotechnol. 33(5), 864–870 (2022)
T. Saito, T. Iwase, J. Horie, T. Morioka, Mode of photocatalytic bactericidal action of powdered semiconductor TiO2 on mutans streptococci. J. Photochem. Photobiol. B. 14(4), 369–379 (1992)
A. Besinis, T. De Peralta, R.D. Handy, The antibacterial effects of silver, titanium dioxide and silica dioxide nanoparticles compared to the dental disinfectant chlorhexidine on Streptococcus mutans using a suite of bioassays. Nanotoxicology 8(1), 1–16 (2014)
M. Pourhajibagher, A. Bahador, Synergistic biocidal effects of metal oxide nanoparticles-assisted ultrasound irradiation: antimicrobial sonodynamic therapy against Streptococcus mutans biofilms. Photodiagn. Photodyn. Ther. 35, 102432 (2021)
C. Miron, A. Roca, S. Hoisie, P. Cozorici, L. Sirghi, Photoinduced bactericidal activity of TiO2 thin films obtained by radiofrequency magnetron sputtering deposition. J. Optoelectron. Adv. Mater. 7, 915–919 (2004)
F. Khan, D.T.N. Pham, S.F. Oloketuyi, P. Manivasagan, J. Oh, Y.M. Kim, Chitosan and their derivatives: antibiofilm drugs against pathogenic bacteria. Colloids Surf. B 185, 110627 (2020)
P. Amezaga-Madrid, R. Silveyra-Morales, L. Cordoba-Fierro, G.V. Nevarez-Moorillon, M. Miki-Yoshida, E. Orrantia-Borunda, F.J. Solıs, TEM evidence of ultrastructural alteration on Pseudomonas aeruginosa by photocatalytic TiO2 thin films. J. Photochem. Photobiol. B. 70(1), 45–50 (2003)
S. Arya, H. Sonawane, S. Math, P. Tambade, M. Chaskar, D. Shinde, Biogenic titanium nanoparticles (TiO2 NPs) from Tricoderma citrinoviride extract: synthesis, characterization and antibacterial activity against extremely drug-resistant Pseudomonas aeruginosa. Int. Nano Lett. 11, 35–42 (2021)
W.J. Parak, L. Manna, F.C. Simmel, D. Gerion, P. Alivisatos, Nanoparticles: from theory to application (Wiley-VCH, Weinheim, 2004)
S.J. Hoseyni, M. Manoochehri, M.D. Asli, Synthesis of cobalt nanoparticles by complex demolition method using the reaction between organic ligand Schiff base and cobalt chloride by ultrasonication. Bull. Soc. Roy. Sci. Liège 86, 325–331 (2017)
O.U. Igwe, E.S. Ekebo, Biofabrication of cobalt nanoparticle odorata and their potential. Res. J. Chem. 8(1), 11–17 (2018)
M. Azharuddin, G.H. Zhu, D. Das, E. Ozgur, L. Uzun, A.P. Turner, H.K. Patra, A repertoire of biomedical applications of noble metal nanoparticles. Chem. Commun. 55(49), 6964–6996 (2019)
A.K. Singh, A review on plant extract-based route for synthesis of cobalt nanoparticles: photocatalytic, electrochemical sensing and antibacterial applications. Curr. Res. Green Sustain. Chem. (2022). https://doi.org/10.1016/j.crgsc.2022.100270
N.E. Eleraky, A. Allam, S.B. Hassan, M.M. Omar, Nanomedicine fight against antibacterial resistance: an overview of the recent pharmaceutical innovations. Pharmaceutics 12(2), 142 (2020)
T. Varaprasad, B. Govindh, B.V. Rao, Green synthesized cobalt nanoparticles using Asparagus racemosus root extract & evaluation of antibacterial activity. Int. J. Chem. Tech. Res. 10(9), 339–345 (2017)
C.T. Anuradha, P. Raji, Effect of annealing temperature on antibacterial, antifungal and structural properties of bio-synthesized Co3O4 nanoparticles using Hibiscus Rosa-sinensis. Mater. Res. Exp. 6(9), 095063 (2019)
M.M. Naik, H.B. Naik, G. Nagaraju, M. Vinuth, K. Vinu, R. Viswanath, Green synthesis of zinc doped cobalt ferrite nanoparticles: structural, optical, photocatalytic and antibacterial studies. Nano-Structures & Nano-Objects 19, 100322 (2019)
D. Kharade Suvarta, H. Nikam Gurunath, J. Mane Gavade Shubhangi, R. Patil Sachinkumar, V. Gaikwad Kishor, Biogenic synthesis of cobalt nanoparticles using Hibiscus cannabinus leaf extract and their antibacterial activity. Res. J. Chem. Environ 24(5), 9–13 (2020)
P.P. Shriniwas, T.K. Subhash, Antioxidant, antibacterial and cytotoxic potential of silver nanoparticles synthesized using terpenes rich extract of Lantana camara L. leaves. Biochem. Biophys. Rep 10, 76–81 (2017)
H. Guan, W. Dong, Y. Lu, M. Jiang, D. Zhang, Y. Aobuliaximu, S. Lu, Distribution and antibiotic resistance patterns of pathogenic bacteria in patients with chronic cutaneous wounds in China. Front. Med. 8, 609584 (2021)
M. Hafeez, R. Shaheen, B. Akram, S. Haq, S. Mahsud, S. Ali, R.T. Khan, Green synthesis of cobalt oxide nanoparticles for potential biological applications. Mater. Res. Exp. 7(2), 025019 (2020)
S. Iravani, R.S. Varma, Sustainable synthesis of cobalt and cobalt oxide nanoparticles and their catalytic and biomedical applications. Green Chem. 22(9), 2643–2661 (2020)
G. Satpathy, E. Manikandan, Cobalt nanoparticle as the antibacterial tool. Int. J. Eng. Adv. Technol. (IJEAT) 8, 3684–3687 (2019)
V. Dogra, G. Kaur, S. Jindal, R. Kumar, S. Kumar, N.K. Singhal, Bactericidal effects of metallosurfactants based cobalt oxide/hydroxide nanoparticles against Staphylococcus aureus. Sci. Total. Environ. 681, 350–364 (2019)
S. Chattopadhyay, S.K. Dash, S. Tripathy, B. Das, D. Mandal, P. Pramanik, S. Roy, Toxicity of cobalt oxide nanoparticles to normal cells; an in vitro and in vivo study. Chem. Biol. Interact. 226, 58–71 (2015)
J.S. Ajarem, S.N. Maodaa, A.A. Allam, M.M. Taher, M. Khalaf, Benign synthesis of cobalt oxide nanoparticles containing red algae extract: antioxidant, antimicrobial, anticancer, and anticoagulant activity. J. Clust. Sci. (2021). https://doi.org/10.1007/s10876-021-02004-9
M. Sivachidambaram, J.J. Vijaya, K. Kaviyarasu, L.J. Kennedy, H.A. Al-Lohedan, R.J. Ramalingam, A novel synthesis protocol for Co3O4 nanocatalysts and their catalytic applications. RSC Adv. 7(62), 38861–38870 (2017)
L. Du, S. Ahmad, L. Liu, L. Wang, J. Tang, A review of antibiotics and antibiotic resistance genes (ARGs) adsorption by biochar and modified biochar in water. Sci. Total. Environ. 858, 159815 (2023)
C.J. Murray, K.S. Ikuta, F. Sharara, L. Swetschinski, G.R. Aguilar, A. Gray, N. Tasak, Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 399(10325), 629–655 (2022)
A.R. Mahoney, M.M. Safaee, W.M. Wuest, A.L. Furst, The silent pandemic: emergent antibiotic resistances following the global response to SARS-CoV-2. IScience 24(4), 102304 (2021)
R.F. O’Toole, K.W. Leong, V. Cumming, S.J. van Hal, Vancomycin-resistant Enterococcus faecium and the emergence of new sequence types associated with hospital infection. Res. Microbiol. (2023). https://doi.org/10.1016/j.resmic.2023.104046
J. Tabcheh, J. Vergalli, A. Davin-Régli, N. Ghanem, C. Al-Bayssari, J.M. Brunel, Rejuvenating the activity of usual antibiotics on resistant gram-negative bacteria: recent issues and perspectives. Int. J. Mol. Sci. 24(2), 1515 (2023)
J.M. Morris, K. Mercoulia, M. Valcanis, C.L. Gorrie, N.L. Sherry, B.P. Howden, Hidden resistances: how routine whole-genome sequencing uncovered an otherwise undetected bla NDM-1 gene in vibrio alginolyticus from imported seafood. Microbiol. Spectrum (2023). https://doi.org/10.1128/spectrum.04176-22
E. Altun, M.O. Aydogdu, E. Chung, G. Ren, S. Homer-Vanniasinkam, M. Edirisinghe, Metal-based nanoparticles for combating antibiotic resistance. Appl. Phys. Rev. (2021). https://doi.org/10.1063/5.0060299
A. Frei, A.D. Verderosa, A.G. Elliott, J. Zuegg, M.A. Blaskovich, Metals to combat antimicrobial resistance. Nat. Rev. Chem. (2023). https://doi.org/10.1038/s41570-023-00463-4
A. Chahardoli, M. Mavaei, Y. Shokoohinia, A. Fattahi, Galbanic acid, a sesquiterpene coumarin as a novel candidate for the biosynthesis of silver nanoparticles: in vitro hemocompatibility, antiproliferative, antibacterial, antioxidant, and anti-inflammatory properties. Adv. Powder Technol. 34(1), 103928 (2023)
I.H. Ifijen, M. Maliki, N.U. Udokpoh, I.J. Odiachi, B. Atoe, A concise review of the antibacterial action of gold nanoparticles against various bacteria, in TMS annual meeting & exhibition. (Springer Nature, Cham, 2023), pp.655–664
M. Alherek, O.D. Basu, Impact of low-levels of silver, zinc and copper nanoparticles on bacterial removal and potential synergy in water treatment applications. J. Chem. Technol. Biotechnol. 98(5), 1137–1146 (2023)
A.I. Doulgeraki, C.S. Kamarinou, G.J.E. Nychas, A.A. Argyri, C.C. Tassou, G. Moulas, N. Chorianopoulos, Role of microbial interactions across food-related bacteria on biofilm population and biofilm decontamination by a TiO2-nanoparticle-based surfactant. Pathogens 12(4), 573 (2023)
S.S. Hassan, K.A. Hubeatir, R.M.S. Al-Haddad, Characterization and antibacterial activity of silica-coated bismuth (Bi@ SiO2) nanoparticles synthesized by pulsed laser ablation in liquid. Optik (2023). https://doi.org/10.1016/j.ijleo.2022.170453
P.R. More, S. Pandit, A.D. Filippis, G. Franci, I. Mijakovic, M. Galdiero, Silver nanoparticles: bactericidal and mechanistic approach against drug resistant pathogens. Microorganisms 11(2), 369 (2023)
M. Paesa, C.R. de Ganuza, T. Alejo, C. Yus, S. Irusta, M. Arruebo, G. Mendoza, Elucidating the mechanisms of action of antibiotic-like ionic gold and biogenic gold nanoparticles against bacteria. J. Colloid Interface Sci. 633, 786–799 (2023)
R. Abreu, T. Semedo-Lemsaddek, E. Cunha, L. Tavares, M. Oliveira, Antimicrobial drug resistance in poultry production: current status and innovative strategies for bacterial control. Microorganisms 11(4), 953 (2023)
Y. Zaman, M.Z. Ishaque, S. Ajmal, M. Shahzad, A.B. Siddique, M.U. Hameed, G. Yasin, Tamed synthesis of AgNPs for photodegradation and anti-bacterial activity: effect of size and morphology. Inorg. Chem. Commun. 150, 110523 (2023)
V. Mageshwaran, P. Sivasubramanian, P. Kumar, Y. Nagaraju, Antibacterial response of nanostructured chitosan hybrid materials. Chitosan Nanocomposites: Bionanomech. Appl. (2023). https://doi.org/10.1007/978-981-19-9646-7_7
J. Ge, D. Li, J. Ding, X. Xiao, Y. Liang, Microbial coexistence in the rhizosphere and the promotion of plant stress resistance: a review. Environ. Res. (2023). https://doi.org/10.1016/j.envres.2023.115298
A.K. Halder, A.S. Moura, M.N.D. Cordeiro, Predicting the ecotoxicity of endocrine disruptive chemicals: multitasking in silico approaches towards global models. Sci. Total. Environ. 889, 164337 (2023)
W. Hu, C. Wang, D. Gao, Q. Liang, Toxicity of transition metal nanoparticles: a review of different experimental models in the gastrointestinal tract. J. Appl. Toxicol. 43(1), 32–46 (2023)
D. Nath Roy, R. Goswami, A. Pal, Nanomaterial and toxicity: what can proteomics tell us about the nanotoxicology? Xenobiotica 47(7), 632–643 (2017)
A.M. Schrand, M.F. Rahman, S.M. Hussain, J.J. Schlager, D.A. Smith, A.F. Syed, Metal-based nanoparticles and their toxicity assessment. Wiley Interdiscip. Rev.: Nanomed. Nanobiotechnol. 2(5), 544–568 (2010)
A. Sukhanova, S. Bozrova, P. Sokolov, M. Berestovoy, A. Karaulov, I. Nabiev, Dependence of nanoparticle toxicity on their physical and chemical properties. Nanoscale Res. Lett. 13(1), 44 (2018). https://doi.org/10.1186/s11671-018-2457-x
S. Hua, M.B. De Matos, J.M. Metselaar, G. Storm, Current trends and challenges in the clinical translation of nanoparticulate nanomedicines: pathways for translational development and commercialization. Front. Pharmacol. 9, 790 (2018)
A. Elsaesser, C.V. Howard, Toxicology of nanoparticles. Adv. Drug Deliv. Rev. 64(2), 129–137 (2012)
OECD, Publications in the series on the safety of manufactured nanomaterials, www.oecd.org, 2019
S.J. Choi, J.K. Lee, J. Jeong, J.H. Choy, Toxicity evaluation of inorganic nanoparticles: considerations and challenges. Mol. Cell. Toxicol. 9, 205–210 (2013)
S.C. Gad, Drug safety evaluation (John Wiley, New York, 2002)
V. De Matteis, Exposure to inorganic nanoparticles: routes of entry, immune response, biodistribution and in vitro/in vivo toxicity evaluation. Toxics 5(4), 29 (2017)
R. Mohammadpour, M.A. Dobrovolskaia, D.L. Cheney, K.F. Greish, H. Ghandehari, Subchronic and chronic toxicity evaluation of inorganic nanoparticles for delivery applications. Adv. Drug Deliv. Rev. 144, 112–132 (2019)
C.D. Klaassen, Toxic responses of the respiratory system, in Casarett and & Doull’s toxicology: the basic science of poisons, 5th edn. (McGraw-Hill Companies Inc, New York, 1996), pp.515–534
E.O. Erhirhie, C.P. Ihekwereme, E.E. Ilodigwe, Advances in acute toxicity testing: strengths, weaknesses and regulatory acceptance. Interdiscip. Toxicol. 11, 5–12 (2018)
S. Parasuraman, Toxicological screening. J. Pharmacol. Pharmacother. 2, 74 (2011)
C. Bai, M. Tang, Toxicological study of metal and metal oxide nanoparticles in zebrafish. J. Appl. Toxicol. 40(1), 37–63 (2020)
H. Nehoff, S. Taurin, K. Greish, Toxicological assessment of nanomedicine (Wiley, Hoboken, 2013)
K. Greish, G. Thiagarajan, H. Ghandehari, In vivo methods of nanotoxicology. Methods Mol. Biol. 926, 235–253 (2012)
S. Haldar, Y. Muralidaran, D. Míguez, S.I. Mulla, P. Mishra, Eco-toxicity of nano-plastics and its implication on human metabolism: current and future perspective. Sci. Total. Environ. 861, 160571 (2023)
J. Bahamonde, B. Brenseke, M.Y. Chan, R.D. Kent, P.J. Vikesland, M.R. Prater, Gold nanoparticle toxicity in mice and rats: species differences. Toxicol. Pathol. 46(4), 431–443 (2018). https://doi.org/10.1177/0192623318770608
Y. Cao, S. Li, J. Chen, Modeling better in vitro models for the prediction of nanoparticle toxicity: a review. Toxicol. Mech. Methods 31(1), 1–17 (2021). https://doi.org/10.1080/15376516.2020.1828521
M. Xu, G. Halimu, Q. Zhang, Y. Song, X. Fu, Y. Li et al., Internalization and toxicity: a preliminary study of effects of nanoplastic particles on human lung epithelial cell. Sci. Total. Environ. 694, 133794 (2019). https://doi.org/10.1016/j.scitotenv.2019.133794
P. Khanna, C. Ong, B.H. Bay, G.H. Baeg, Nanotoxicity: an interplay of oxidative stress, inflammation and cell death. Nanomaterials 5(3), 1163–1180 (2015)
P. Makhdoumi, H. Karimi, M. Khazaei, Review on metal-based nanoparticles: role of reactive oxygen species in renal toxicity. Chem. Res. Toxicol. 33(10), 2503–2514 (2020). https://doi.org/10.1021/acs.chemrestox.9b00438
M. Mishra, M. Panda, Reactive oxygen species: The root cause of nanoparticle-induced toxicity in Drosophila melanogaster. Free Radic. Res. 55(6), 919–935 (2021). https://doi.org/10.1080/10715762.2021.1914335
W. Yang, L. Wang, E.M. Mettenbrink, P.L. DeAngelis, S. Wilhelm, Nanoparticle toxicology. Annu. Rev. Pharmacol. Toxicol. 61, 269–289 (2021)
S. Yu, J. Liu, Y. Yin, M. Shen, Interactions between engineered nanoparticles and dissolved organic matter: a review on mechanisms and environmental effects. J. Environ. Sci. 63, 198–217 (2018)
M. Horie, Y. Tabei, Role of oxidative stress in nanoparticles toxicity. Free Radical Res. 55(4), 331–342 (2021)
S.J. Forrester, D.S. Kikuchi, M.S. Hernandes, Q. Xu, K.K. Griendling, Reactive oxygen species in metabolic and inflammatory signaling. Circ. Res. 122(6), 877–902 (2018). https://doi.org/10.1161/circresaha.117.311401
K. Jakubczyk, K. Dec, J. Kaldunska, D. Kawczuga, J. Kochman, K. Janda, Reactive oxygen species - sources, functions, oxidative damage. Pol. Merkur. Lek. 48(284), 124–127 (2020)
P.D. Ray, B.W. Huang, Y. Tsuji, Reactive oxygen species (ROS) homeostasis and redox regulation in cellular signaling. Cell. Signal. 24(5), 981–990 (2012). https://doi.org/10.1016/j.cellsig.2012.01.008
R. Wang, B. Song, J. Wu, Y. Zhang, A. Chen, L. Shao, Potential adverse effects of nanoparticles on the reproductive system. Int. J. Nanomed. 13, 8487 (2018)
C. Egbuna, V.K. Parmar, J. Jeevanandam, S.M. Ezzat, K.C. Patrick-Iwuanyanwu, C.O. Adetunji, C.G. Ibeabuchi, Toxicity of nanoparticles in biomedical application: nanotoxicology. J. Toxicol. 2021, 1–21 (2021)
C. Liao, Y. Li, S.C. Tjong, Bactericidal and cytotoxic properties of silver nanoparticles. Int. J. Mol. Sci. 20(2), 449 (2019)
Funding
Funding was not available for this study.
Author information
Authors and Affiliations
Contributions
MS wrote and edited the manuscript and did the software work. SA edited and evaluated the manuscript. HMT edited and evaluated the manuscript. RA edited the manuscript and help in data acquisition regarding mechanism of NPs action. UF did the graphical and pictorial work. SM did the reference management several times. AH provided the data for manuscript when required. TAM evaluated the data. MAF assisted in software work. HF provided the information regarding antibiotics resistance mechanism.
Corresponding author
Ethics declarations
Competing Interests
There were no competing interests declared unanimously by the authors.
Ethical Approval
Not applicable.
Consent to Participate
All authors have shown consent to participate.
Consent to Publication
All authors showed consent to publish the current review article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Summer, M., Ali, S., Tahir, H.M. et al. Mode of Action of Biogenic Silver, Zinc, Copper, Titanium and Cobalt Nanoparticles Against Antibiotics Resistant Pathogens. J Inorg Organomet Polym 34, 1417–1451 (2024). https://doi.org/10.1007/s10904-023-02935-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10904-023-02935-y