Abstract
The zebra mussel, Dreissena polymorpha (Pallas, 1771), is a successful invasive bivalve native to the brackish and freshwaters systems of the Ponto-Caspian regions. It is considered one of the world’s 100 worst invasive alien species and has high ecological and economic impacts. Over the past 200 years, it has expanded across the European and North American continents. Since the recent invasion of the Iberian Peninsula in 2001, D. polymorpha has expanded along the Ebro River basin and colonized adjacent basins in Northeastern Iberia. Using 9 microsatellite markers, we analyzed one population from the low Danube and several European and North American introduced locations, including a large number of Iberian sites, to genetically characterize the zebra mussel invasion and to identify putative routes of colonization. Our results reflect a high homogeneity of the Iberian samples, indicating that D. polymorpha colonized the Iberian Peninsula via a single invasion event. The genetic singularity of the populations observed in this region (including 9 unique alleles and significant genetic distances with the rest of analyzed samples) indicates that additional sites from the native and introduced range must be sampled to identify the source of the Iberian invasion.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The zebra mussel, Dreissena polymorpha (Pallas, 1771), is a successful invasive bivalve native to the brackish estuaries and freshwater systems of the Ponto-Caspian region, which includes the Black, Caspian, and Azov Seas, and the lower section of the Danube River drainage in Romania (Müller et al., 2002; Son, 2007; Popa & Murariu, 2009). These natural D. polymorpha populations are characterized by high levels of genetic and morphological variability (Marsden et al., 1995).
This species is considered one of the world’s 100 worst invasive alien species by the Invasive Species Specialist Group (IUCN-ISSG, Lowe et al., 2000). Some biological characteristics [i.e., rapid growth with early sexual maturity, high fecundity (Astanei et al., 2005), dispersal capability (Bij de Vaate et al., 2002)] are related to the species’ invasion potential. In addition, D. polymorpha invasions have been facilitated by human-mediated dispersal mechanisms, such as modifications of river systems and ship-borne trading routes (Bij de Vaate et al., 2002; Havel et al., 2015). Adult D. polymorpha may travel attached to the hulls of commercial and recreational ships (Johnson & Carlton, 1996), and veliger larvae survive in ballast water (Brown & Stepien, 2010).
Due to substantial ecological and economic damage caused by D. polymorpha, the invasions of this species are a focus of public interest. The aggregation of massive colonies of this species produces a significant ecological disequilibrium (Kirsch & Dzialowski, 2012; Havel et al., 2015; Karatayev et al., 2015) with habitat and food competition with autochthonous bivalves that can promote species replacement (Strayer & Malcom, 2007). The socio-economic effects are principally caused by the dense colonies of D. polymorpha that block artificial waterways. In the Ebro River basin, the costs derived from D. polymorpha invasion during the 2005–2009 were estimated to be 11.5 M€, which were invested in the cleaning of water intake pipes in raw water-dependent infrastructures and to the control and prevention of the invasion (Durán et al., 2012).
The current geographic distribution of D. polymorpha includes both the European and North American continents (Karatayev et al., 2015). The Ponto-Caspian region has been the origin of many aquatic invasions in Europe, facilitated by the interconnection of river basins since the 18th century (Bij de Vaate et al., 2002). Three corridors from the Ponto-Caspian region to Europe have been described: a northern route through the Baltic Sea, a central corridor ending at the Rhine River, and a southern corridor that connects the Danube and Rhine Rivers (Bij de Vaate et al., 2002). According to Bij de Vaate et al. (2002), D. polymorpha may have used a combination of the northern and central corridors to expand throughout Europe. Interestingly, additional studies reported genetic evidence suggesting that the southern corridor was also used (Rajagopal et al., 2009; Navarro et al., 2013).
Molecular markers are used to genetically characterize invasive species and to infer the sources and pathways of colonization (Stepien et al., 2005; Sanz et al., 2013; Lopes-Lima et al., 2014). In invasive D. polymorpha, high gene diversity found using distinct molecular markers has been related to large founder populations or multiple colonization events (Gosling et al., 2008). Based on allozyme markers, Marsden et al. (1995) and Lewis et al. (2000) identified two major clusters in American samples that likely originated in the Danube River and in Northern Europe, respectively, as a consequence of multiple colonization events. Stepien et al. (2002, 2005) confirmed this hypothesis by nuclear randomly amplified polymorphic DNA (RAPD) and the cytochrome b (cytb) gene. Gelembuik et al. (2006) and May et al. (2006) using the Cytochrome oxidase subunit I (COI) revealed that all introduced populations in North America and Europe were derived from the Ponto-Caspian Sea basin (May et al., 2006). In Europe, French populations analyzed by COI gene sequences and amplified fragment length polymorphism (AFLP) markers have suggested one continuous linear invasion from northern to southern France despite the opportunities for long-distance dispersal (Tarnowska et al., 2013). Microsatellite markers used to analyze European and North American populations (Müller et al., 2002; Astanei et al., 2005; Brown & Stepien, 2010) have revealed a significant genetic structure in European D. polymorpha attributed to distinct origins and invasion routes with a long invasion history. In North America, the introduced populations had high genetic diversity values at microsatellite loci, confirming multiple European sources with very high numbers of introduced individuals.
The presence of D. polymorpha in the Iberian Peninsula was first reported in the low reaches of the Ebro River with the first sighting in the Ribarroja Reservoir in 2001 (Ruíz-Altaba et al., 2001). Since then, this species has expanded downstream and upstream in the Ebro (Durán & Anadón, 2008; Oscoz et al., 2010; Sanz-Ronda et al., 2014). Dreissena polymorpha has also colonized different Spanish adjacent basins. Larvae and adults were first detected in 2005 in the Mijares River, 105 km south of the Ebro River (Navarro et al., 2013) and in 2011 in the Llobregat River, 215 km north of the Ebro River (Peñarrubia et al., 2015). Rajagopal et al. (2009) used AFLP markers and clustered the Iberian D. polymorpha populations within a wide European genetic group. These authors suggested that the origin of the Iberian invasion was France because the populations from these two countries displayed the highest similarity. The authors also indicated some genetic resemblance between the Iberian and Italian populations. Navarro et al. (2013) analyzed the invasion of D. polymorpha into the Iberian Peninsula using the mitochondrial COI gene and a single microsatellite marker and failed to detect any structure among the introduced populations. This finding was in agreement with our preliminary work that validated the new microsatellite markers used in the current study (Peñarrubia et al., 2015). In both cases, the lack of genetic differentiation detected among Iberian populations may be compatible with a recent invasion from a common source. However, a low number of microsatellites were used in Navarro et al. (2013), and a limited number of individuals were analyzed in Peñarrubia et al. (2015). Then, further analyses increasing the number of markers and individuals are necessary to genetically characterize Iberian D. polymorpha populations and to confirm this invasion pattern.
The objective of the present study was to assess the genetic diversity and population structure of D. polymorpha from introduced locations in the Iberian Peninsula. Additional native and introduced populations outside of the Peninsula were included in the analysis to evaluate putative colonization routes and reconstruct geographical pathways to the Iberian Peninsula. Such information will contribute to a better understanding of the population genetics dynamic following invasion, as well as the dispersal patterns contributing to the species’ spread.
Materials and methods
Collections and DNA extraction and genotyping
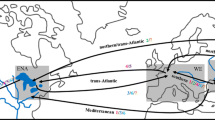
A total of 1,082 D. polymorpha individuals were collected from 2010 to 2013 from 21 sampling points: 18 invaded European sites, 1 location from the low Danube River in Romania (1-Galati) within the native range of the species (Son, 2007; Popa & Murariu, 2009), and 2 from North American locations (Table 1; Fig. 1). Whole mussel bodies without shells were preserved in 96% ethanol prior to processing for the genetic analyses. In addition, because the invasive quagga mussel (D. rostriformis) has been often coexisting and confused with zebra mussel in invaded regions (Marescaux et al., 2015), 10 D. rostriformis individuals collected by A. Bij de Vaate from Ijsselmeer Lake (The Netherlands) were included to check for the presence of this species. DNA extraction was performed using the Real Pure DNA Extraction Kit (Durvitz®, Valencia, Spain) according to Peñarrubia et al. (2015). All sites were genotyped for 9 microsatellite loci [DpolC5, DpolB9, DpolB6, DpolA6, DpolB8 (Naish & Gosling, 2001), Dp31, Dp43, Dp68, and Dp86 (Peñarrubia et al., 2015)] using fluorescent end-labeled forward primers in 2 Multiplex-PCR reactions (see Online Resource 1). The genotypes were assessed using the Applied Biosystems 3130 Genetic analyzer and the GeneMapper 4.0 software (Applied Biosystems, Foster City, CA, U.S.A.).
Geographical locations of D. polymorpha samples analyzed in this study. The single barrier detected by BARRIER software is indicated by a dotted line. See Table 1 for sampling numbers
Data analysis
To test for large allele dropout and null alleles, all microsatellite data were analyzed using MICROCHECKER v 2.2.1 (Van Oosterhout et al., 2004). The Hardy–Weinberg equilibrium (HWE) was measured at each site by the exact probability test (Guo & Thompson, 1992) using GENEPOP software version 4 (Raymond & Rousset, 1995). The unbiased composite-linkage disequilibrium measure D (Weir, 1990) of this program tested possible gametic disequilibria for all pairs of loci in each population. Bonferroni corrections (Rice, 1989) were applied to each single test.
The genetic diversity within each study site was estimated as the number of alleles per locus (N A), the expected heterozygosity (Hs), and the allelic richness (A r) by FSTAT version 2.9.3 (Goudet, 2001). For each location, we tested for recent population bottlenecks using the BOTTLENECK program version 1.2.02 (Cornuet & Luikart, 1996). Following the recommendations for microsatellite data, we used a two-phase model of mutation (TPM; modeled 70% single-step mutations and 30% multi-step changes). The significance of heterozygote excess was estimated using the Wilcoxon rank test because it provided relatively high power with a small number of polymorphic microsatellite loci. Finally, the bottleneck signal was validated by the shape of the allele frequency distribution (‘mode-shift’ indicator, Cornuet & Luikart, 1996).
The pairwise population differentiation (F ST, Weir & Cockerham, 1984) and its significance were calculated using FSTAT version 2.9.3 (Goudet, 2001). The effect of putative null alleles on these values was assessed using the FREENA software (Chapuis & Estoup, 2007). Genetic relationships between populations were further assessed by constructing a neighbor-joining (NJ) tree based on the D A Nei distance (Nei et al., 1983). The confidence of the branching points was estimated by bootstrapping loci with 1,000 replicates of the pairwise distances using the MSA software version 4.05 (Dieringer & Schlötterer, 2003); a subsequent consensus tree was constructed with PHILIP version 3.69 (Felsenstein, 2005).
The number of genetically homogeneous groups (k) was estimated among all sampled collections (native and introduced) using the Bayesian Markov Chain Monte Carlo (MCMC) approach method in the STRUCTURE version 2.3.2 software (Pritchard et al., 2000). Runs for each possible k (1–21) were repeated 10 times. A burn-in period of 20,000 steps followed by 100,000 MCMC replicates was simulated with the model of independent allele frequencies. The optimal number of clusters (k) was selected following the criteria of Evanno et al. (2005) as implemented in STRUCTURE HARVESTER (Earl & von Holdt, 2012). Furthermore, a second group-level Bayesian analysis was estimated using BAPS 5.4 (Corander et al., 2008).
Major genetic discontinuities in the study area were assessed using Monmonier’s algorithm implemented in BARRIER version 2.2 (Manni et al., 2004). This method allows the identification of hidden geographical barriers by detecting abrupt rates of change between pairwise F ST distances for each single microsatellite marker. A barrier was considered robust if it was supported by at least 5 of the 9 microsatellites (Díez-del-Molino et al., 2013). We used the assignment test provided by GENECLASS v.2 (Piry et al., 2004) to assess the putative sources of the introduced D. polymorpha populations. We used a modified version of the assignment test where the population allele frequencies were estimated using a Bayesian approach (Rannala & Mountain, 1997). In addition, we used the ‘simulation option’ of Paetkau et al. (2004) to assess the probability of each individual being rejected from each location. Finally, contemporary migration rates among populations were estimated using the Bayesian inference as implemented in the BAYESASS 3.0 software (Wilson & Rannala, 2003); this method does not assume migration-drift or Hardy–Weinberg equilibrium. According to the author’s recommendations, the collections were grouped by taking into account the clusters from the BAPS analysis and significant F ST values to avoid migration rates higher than 30% (Wilson & Rannala, 2003). A total of 3,000,000 iterations were performed until the MCMC chains reached a stationary distribution. Migration parameters were estimated by sampling 2,000 iterations after a burn-in of 1,000,000 iterations. Delta values were adjusted following the BAYESASS manual recommendations. Ten runs using different starting points were performed, and the results with the highest likelihood were retained.
Results
Gene diversity analysis
All 9 microsatellites used in this study were polymorphic in D. polymorpha and failed to amplify in D. rostriformis. The number of alleles per locus ranged from 3 (Dp86) to 30 (Dp31). According to MICROCHECKER, putative null alleles were detected for single locations in 5 of the 9 loci (DpolB8, DpolA6, Dp31, Dp43, and Dp68). A significant null allele frequency (q ≥ 0.1) was estimated by FREENA at DpolB8 in 1-Galati (q = 0.150), DpolA6 in 6-Mendavia (q = 0.196), 14-Trasimeno (q = 0.155), and 20-Texoma (q = 0.107), Dp31 in 12-Sitjar (q = 0.111), Dp43 in 19-Salford Quays (q = 0.113), and Dp68 in 10-Ebro Delta (q = 0.107). The gene diversity values varied slightly among collections. The lowest number of total alleles (N A), average allelic richness (A r), and genetic diversity (H s) were observed in 14-Trasimeno from central Italy (N A = 51; A r = 4.422; Hs = 0.598), and the highest values were observed in 20-Texoma from North America (N A = 116; A r = 8.574; H s = 0.758) (Table 1). The highest number of private alleles per location was 5 in the native site 1-Galati (Table 1). By country, 9 alleles were distinctive of Spanish locations, six were only detected at French sites, four were detected in Britain, and one was detected at Italian sites. A total of 15 alleles were distinctive for North America. The non-parametric Wilcoxon signed-rank test in FSTAT with 1,000 permutations that was used to compare the genetic diversity among countries (Table 1) indicated that H s and A r were significantly lower at the Spanish and Italian sites compared to the American locations (P < 0.005).
After Bonferroni correction, deviations from HWE were still detected in the native site (1-Galati) at DpolB8 (P < 0.001), in one Spanish location (6-Mendavia) at DpolC5 (P < 0.001) and in 20-Texoma from North America at DpolA6 (P < 0.001) (Table 1). No significant pairwise linkage disequilibrium was observed in any sample. Because significant null alleles and HWE deviations were dispersed at single loci and populations and not associated with linkage disequilibrium, we considered these deviations to not be biologically significant and maintained all loci in the following analyses. According to the BOTTLENECK analyses, excess heterozygosity was significant at two sites in the Ebro River (2-Soborn and 5-St. Vicente). However, the allele-shift model test did not report additional signals for recent bottlenecks at these locations.
Genetic differentiation and population structure analyses
Overall, the genetic differentiation among locations was low (F ST = 0.046) but increased to moderate when this estimator was re-scaled to the theoretical maximum value (G’ ST = 0.170). Additionally, the estimated F ST did not change substantially after correcting for null alleles (F ST = 0.045). Accordingly, we retained all loci in the posterior analyses and computed them based on the original frequencies. The pairwise comparisons ranged from F ST = 0.000 between several Spanish locations to F ST = 0.096 between 20-Texoma (USA) and 11-La Baells (Spain) and between 21-Ray Roberts (USA) and 10-Ebro Delta (Spain) (Table 2). After Bonferroni correction, the pairwise F ST values were mostly significant (P < 0.05) when comparing locations from different countries and not significant when comparing locations within countries (Table 2). Exceptions were observed in most comparisons between French and British locations (sites 15–19), which yielded non-significant F ST values, and the significant genetic differentiation (F ST = 0.051; P < 0.05) found between the two Italian sites (13-Lugano and 14-Trasimeno). In Spain, significant pairwise F ST (P < 0.05) values were detected when comparing the Ebro River sites (2–10) with the 11-Llobregat and the 12-Mijares River (Table 2).
According to the D A distances, the France and Great Britain sites were the most similar introduced locations to the native site (1-Galati), whereas the Spanish and North American locations were the most divergent (Table 2). A neighbor-joining (NJ) tree based on the D A distance and rooted with the native location (1-Galati) also indicated that the highest genetic differentiation was observed among populations from different countries (Fig. 2). All Spanish locations were clustered together with the highest bootstrap support (100%). Within this cluster, a lower structure that separated Mijares and Llobregat from the Ebro River group was also supported (sites 2–10, 74.4%); this structure situated 9-Ribarroja in a basal position, which agreed with the first record of D. polymorpha in this basin. The other European and North American invaded locations grouped together but with very low bootstrap support. Within this cluster, the Italian (sites 13–14) and North American locations (sites 20–21) were clearly isolated (bootstrap values = 94.9% and 100%, respectively) from the rest of the sites.
Neighbor-Joining tree based on the D A distance (Nei et al., 1983). Numbers at the nodes indicate the percentage of trees with the respective node among 1,000 bootstrapping replicates (only values higher than 50% are shown)
Bayesian analyses of the STRUCTURE and Evanno’s method results grouped sites according to a geographical pattern (Fig. 3a and Online Resource 2). The highest probability was obtained with a model of two clusters (k = 2) that distinguished the Spanish locations (sites 2–12) from all remaining native and invaded sites. The next best cluster set detected by STRUCTURE was k = 5. In this model, the first group was characteristic of the native site (1-Galati), France, and Great Britain (sites 15–19) and was partially present in 13-Lugano (Northern Italy). The next two clusters were not population specific and were distributed across all Spanish locations. A fourth cluster corresponded to Italian sites, although it was significantly present (q > 10%) in France and Great Britain (sites 15–19). The last cluster included the two North American locations (sites 20–21) and again was significantly present (q > 10%) in France (16-Metz) and Great Britain (19-Salford Quays) (Fig. 3a and Online Resource 2). The results from the BAPS analyses identified 6 clusters that coincided with the STRUCTURE results in most cases. The two clusters identified in the Iberian Peninsula distinguished the Llobregat River (11-La Baells) from the rest of the Iberian locations. Moreover, the BAPS analysis grouped the Italian site 13-Lugano with the French and British samples (Fig. 3a).
Population clustering Bayesian algorithms implemented in Structure and BAPS depend largely on the properties of the data (i.e., sampling size bias or low F ST values among the clusters) (Latch et al., 2006) and can misestimate the number of groups when there is an overrepresented genetic group in the data set (Frantz et al., 2009). Thus, because the number of sampled Spanish sites might be overrepresented, these Bayesian analyses were reiterated including only three locations from Spain: the Ebro (10-Ebro Delta), Llobregat (11-La Baells), and Mijares (12-Sitjar) Rivers (Fig. 3b). In this case, STRUCTURE yielded the same results with k = 2. Interestingly, at a lower hierarchical level, the STRUCTURE and BAPS results were similar (k = 5) and failed to differentiate the Llobregat River (11-La Baells) from the rest of the Spanish locations. Moreover, the native site (1-Galati) was differentiated from all of the invaded European sites (Fig. 3b).
In the GENECLASS analysis, the samples were assigned correctly to their original locations (P ≥ 75%) in Romania, Italy, and North America, whereas the locations in France and Great Britain presented the lowest self-assignation value (P ≥ 60%) with an alternative assignation to locations from the same countries (Online Resource 3). All Spanish samples (with the exception of Llobregat River, P = 94.12%) had low assignation values for their own location (P ≥ 38.46%) but presented high assignation values for the rest of the Spanish sites (Online Resource 3). Surprisingly, a high percentage of rejection was observed for 13-Lugano from 14-Trasimeno, but the opposing result was not observed (Online Resource 3). No individuals were directly assigned to the native location (1-Galati), but the analyses showed a low percentage of rejection from this sample mainly for the European (but not Spanish) populations (sites 13–19). The North American locations (sites 20–21) were also clearly rejected (rejection average = 88.98%) from the native site.
Finally, the migration rates estimated by BAYESSAS that grouped samples according to the 6 BAPS clusters were extremely low (<1%) with two exceptions. The first was the unidirectional dispersal rate of 9.23% observed from the Ebro and Mijares group (sites 2–10, 12) to the Llobregat River (11-La Baells). The second was also unidirectional and surprisingly flowed from the French-British group (sites 15–19) to the native site (1-Galati) with a dispersal rate of 4.01%. This unexpected directionality could be explained by the higher number of individuals sampled in the French-British group compared to the native population. In addition, Monmonier’s algorithm in BARRIER identified only one barrier supported by at least 5 loci (Fig. 1), which reflected the isolation of the Spanish basins.
Discussion
Colonization history across Europe and North America
Aquatic interconnections of river basins and human-mediated transport (via adult individuals’ attachment to the hulls of ships and larvae in ballast water) have resulted in several corridors for D. polymorpha invasion from the Ponto-Caspian region to Europe (Bij de Vaate et al., 2002; Havel et al., 2015). Our results indicate a lack of significant bottleneck episodes in most European introduced locations, which display high genetic diversity values similar to those within the native range of the species (see 1-Galati in Table 1; Astanei et al., 2005; Brown & Stepien, 2010). This observation agrees with previous studies (Müller et al., 2002; Astanei et al., 2005) and supports the hypothesis that the expected initial loss of gene diversity (produced by founder effect and bottlenecks) cannot occur when an invasive population is established (Brown & Stepien, 2010). In North American locations, high gene diversity values have been explained by multiple invasion events (Brown & Stepien, 2010). Similar and even higher gene diversity in introduced populations relative to the native source has been observed in D. rostriformis, also suspected of having multiple invasion events (Stepien et al., 2002).
In addition to the described high gene diversity values, the overall F ST (G’ ST) and pairwise F ST values were low to moderate. Low genetic differentiation values between populations have been reported in previous studies with D. polymorpha in both invasive (Müller et al., 2002; Astanei et al., 2005; Brown & Stepien, 2010) and native populations (Müller et al., 2002). Nonetheless, a significant population structure was found (Table 2), especially between large geographic areas.
Interestingly, the French and British samples displayed genetic homogeneity among sites and some differences from the native population 1-Galati. The STRUCTURE re-analysis with a reduced number of Spanish samples confirmed this and distinguished these locations (sites 15–19) from the rest of the analyzed samples. According to Bij de Vaate et al. (2002), the first European invasion episode from the native area took place in Northwestern Europe (France and Great Britain) and most likely followed the central geographical corridor. Our sampling design, which focused on Spanish populations, did not allow us to infer the origin and routes of colonization to France and Great Britain. However, based on our results, the localities analyzed in these countries could have been colonized from the same event. Historical records agree with this hypothesis and suggest that D. polymorpha invaded Britain first and from there reached the French locations (Dhur & Massard, 1995; Aldridge et al., 2004).
Dreissena polymorpha populations in Central and Northwestern Europe could serve as a secondary source of invasion to other locations, and a modern southern corridor has been suggested as the path to Italy (Navarro et al., 2013). The genetic relationship between the Italian and the other European locations analyzed and the differences between the two Italian sites may reveal the directionality of the Italian invasion, which started at the northern areas (13-Lugano) and subsequently colonized the central area of Italy (14-Trasimeno). The presence of our Northwestern European cluster in Northern Italy detected by STRUCTURE (Fig. 3) and the results of the GENECLASS assignment test (Online Resource 3) that excluded most Northern Italian individuals from Central Italy (but not vice versa) support this directionality. Moreover, historical records in Italy (Spilinga et al., 2000; Cianfanelli et al., 2010) suggested a progressive spreading of D. polymorpha from north to south. Interestingly, the high genetic differences between the two Italian sites plus the relatively low gene diversity values at these locations could reflect the natural isolation of lacustrine collections and the lack of recent translocations.
In North America, previous genetic studies concluded that the invasion originated from high numbers of founding individuals and/or multiple colonization events from several European locations that resulted in high gene diversity within populations (Astanei et al., 2005; Gosling et al., 2008; Brown & Stepien, 2010). The STRUCTURE and phylogenetic analyses identified North American locations (sites 20–21) as a separate genetic cluster that was more closely related to the French and British locations (sites 15–19). This observation is in agreement with the proposal that Europe is the origin of D. polymorpha in North America. However, our sampling area in North America and Europe (outside of the Iberian Peninsula) was too restricted to yield significant conclusions for this question.
Genetic structure in Iberian basins: origin and routes of colonization
Previous studies using allozyme markers reported that all European samples, including the Spanish sites, belonged to the same genetic group (Rajagopal et al., 2009). This observation would agree with historical records stating that D. polymorpha reached Spain through France from populations introduced to Central Europe. A posterior analysis using the mitochondrial gene COI and a microsatellite marker (Navarro et al., 2013) identified two southern routes of colonization through Central Europe that reached Italy and Spain independently. This putative independent invasion of Spain through France could be supported by the GENECLASS low level of rejection of Spanish introduced locations when assigned to France and could be compatible with our phylogenetic data. Alternatively, the STRUCTURE analysis with three Spanish sites (10–12) indicated a considerable level of mixture within the native (1-Galati) and the introduced locations because a relevant number of individuals from this site could be classified within the Spanish cluster (Fig. 3B with k = 2, and Online Resource 3). Thus, a putative invasion route from Northwestern Europe to Spain or from the low Danube through Central Europe cannot be ruled out.
In both situations, a severe bottleneck and/or a long period of isolation after the invasion episode is necessary to explain the high genetic differences between the putative sources and the introduced Spanish populations. However, our data did not show any signals of a severe bottleneck in Spain, and the gene diversity values were similar to those found in the other invaded and native populations. Additionally, the first record of D. polymorpha in Spain was in 2001, and the elapsed time until sampling would not have been sufficient to allow the genetic divergence observed between the Spanish population and the rest of the sampled sites (both native and invasive). Therefore, a recent invasion of a large and diverse set of individuals from a specific unsampled location would be a more plausible hypothesis. The genetic singularity of the Iberian populations is supported by all of our results (phylogenetic, Bayesian, and F ST-BARRIER) and the 9 private alleles exclusively identified at the Spanish sites. The uniqueness and homogeneity of the Iberian populations reflects a recent single founder event where the origin of the source population should be different from all of the populations analyzed in this study.
Previous studies using microsatellite markers linked the Spanish populations to D. polymorpha along the Caspian Sea region and the upper Danube (Brown & Stepien, 2010). Our results could not test this scenario because only one sample was from the low Danube region. Müller et al. (2002) used microsatellite markers, and Gelembuik et al. (2006) used COI to suggest that the fluctuations and instability of D. polymorpha populations in the Black-Ponto-Caspian Sea basins generated a mixture of different genetic structures in the native range. As a consequence, the identification of the original invasion sources in this species could be increasingly difficult and significantly affected by stochastic sampling effects, even with more extensive analysis of the native range. It is clear that further sampling is needed to discover the source of the Spanish invasion. However, due to the extensive native and invasive distribution of D. polymorpha (Ram et al., 2012), the identification of the original source would be a matter of chance even with an extensive sampling effort.
Dispersion of D. polymorpha in the Iberian Peninsula
The first confirmed populations of D. polymorpha in the Iberian Peninsula were found in Ribarroja and Flix, which are reservoirs located in the middle course of the Iberian Ebro River (Ruiz-Altaba et al., 2001). From there, this species might have spread along the entire river and reached neighboring basins following a step-wise model as described in the French invasion (Tarnowska et al., 2013). Genetic homogeneity among the Ebro samples suggests contact between all Ebro sites that is most likely related to the ability of D. polymorpha larvae (Sanz-Ronda et al., 2014) to passively travel more than 300 km downstream when geographical barriers are not present (Stoeckel et al., 1997). Additionally, our genetic data suggest that the Ebro River may have acted as a source for the recent colonization of the neighboring basins of the Llobregat and Mijares Rivers (11-La Baells and 12-Sitjar, respectively). Although significant pairwise F ST and phylogenetic reconstruction suggested some level of genetic differentiation between these two locations and the Ebro River, all three basins are primarily clustered in a homogeneous genetic group. The first report of this species in the Mijares River was in 2005 (Navarro et al., 2013); thus, natural isolation could explain the incipient level of genetic differentiation detected in the Mijares. However, historical records report that the Llobregat River was colonized just 1 year before sampling, and thus, the elapsed time between invasion and sampling was most likely not sufficient to allow differentiation. These differences might be attributed to a founder event. In fact, gene diversity (H s), allelic richness (A r) and the number of alleles (N A) found in the Llobregat River (11-La Baells) were lower than the diversity values found in the rest of Spain (Table 1).
Interestingly, a lack of bottleneck signals was detected in most of our samples. The consistent absence of bottleneck signals in D. polymorpha is not exclusive to our study and is in fact a recurrent finding in most analyses with this species (Astanei et al., 2005). This phenomenon might be explained by the fact that invasion of this species usually occurs by larval translocation. Due to the high number and the mobility of the larvae, a significant amount of genetic variation is transferred to the new location even when a very small amount of water is transported (Johnson & Carlton, 1996).
Control and management strategies in Spanish basins
Because eradication of D. polymorpha is extremely difficult (Sousa et al., 2014), prevention is the crucial point for invasion control, and efforts should be focused on avoiding the transport of individuals to new locations (Sanz-Ronda et al., 2014). The relatively late invasion in Spain can most likely be explained by the presence of a geographical barrier (the Pyrenees) (Rajagopal et al., 2009) and has allowed the planning of different control and monitoring strategies to prevent large-scale expansion across the country. Since the first invasion of D. polymorpha in 2001, the species has spread along the entire Ebro River, but only two adjacent basins (the Mijares and the Llobregat Rivers) have been infested (Navarro et al., 2013; Peñarrubia et al., 2015). The present study also shows that despite this fast spread that probably originated from a single point of invasion neither multiple invasion events nor large-scale invasion episodes seem to have occurred. It seems then that control and management programs have a partial effectiveness on preventing zebra mussel spreading. Thus, the entry of D. polymorpha to Spain seems to be mostly controlled, but additional measures are likely necessary to avoid further expansion of the species within the country.
Conclusion
The high genetic homogeneity among zebra mussel populations introduced in the Iberian Peninsula supports the hypothesis that mussels spread within Spain from ‘within country’ sources, from a first single invasion event. High gene diversity values and the lack of recent bottleneck signals, without apparent multiple colonization events, suggest that this first invasion took place by a large and diverse set of individuals that escape from the founder event. In spite of the limited sampling outside Spain, our results agree with the routes of invasion previously described through Europe (Spilinga et al., 2000; Bij de Vaate et al., 2002; Brown & Stepien, 2010; Navarro et al., 2013). However, the genetic singularity of zebra mussel in the Iberian Peninsula did not seem to support previous hypothesis about the origin of the Spanish invasion (Rajagopal et al., 2009), and additional sites from the native and introduced range must be sampled to clarify this issue.
References
Aldridge, D. C., P. Elliot & G. D. Moggridge, 2004. The recent and rapid spread of the zebra mussel (Dreissena polymorpha) in Great Britain. Biological Conservation 119: 253–261.
Astanei, I., E. Gosling, J. Wilson & E. Powell, 2005. Genetic variability and phylogeography of the invasive zebra mussel, Dreissena polymorpha (Pallas). Molecular Ecology 14: 1655–1666.
Bij de Vaate, A., K. Jazdzewski, H. A. M. Ketelaars, S. Gollasch & G. Van der Velde, 2002. Geographical patterns in range extension of Ponto-Caspian macroinvertebrate species in Europe. Canadian Journal of Fisheries and Aquatic Sciences 59: 1159–1174.
Brown, J. E. & C. A. Stepien, 2010. Population genetic history of the dreissenid mussel invasions: expansion patterns across North America. Biological Invasions 12: 3687–3710.
Chapuis, M. P. & A. Estoup, 2007. Microsatellite null alleles and estimation of population differentiation. Molecular Biology and Evolution 24: 621–631.
Churchill, C. J., 2013. Spatio-temporal spawning and larval dynamics of a zebra mussel (Dreissena polymorpha) population in a North Texas Reservoir: implications for invasions in the southern United States. Aquatic Invasions 8: 389–406.
Cianfanelli, S., E. Lori & M. Bodon, 2010. Dreissena polymorpha: current status of knowledge about the distribution in Italy. In Van der Velde, G., S. Rajagopal & A. Bij de Vaate (eds), The Zebra Mussel in Europe. Backhuys Publishers, Leiden: 93–100.
Corander, J., P. Marttinen, J. Sirén & J. Tang, 2008. Enhanced Bayesian modelling in BAPS software for learning genetic structures of populations. BMC Bioinformatics 9: 539.
Cornuet, J. M. & G. Luikart, 1996. Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144: 2001–2014.
Dhur, G. & J. A. Massard, 1995. Étude historique et faunistique des Invertébrés immigrés ou introduits dans la Moselle luxembourgeoise et ses affluents. Bulletin de la Société des naturalistes luxembourgeois 96: 127–156.
Dieringer, D. & C. Schlötterer, 2003. Microsatellite analyser (MSA): a platform independent analysis tool for large microsatellite data sets. Molecular Ecology Notes 3: 167–169.
Díez-del-Molino, D., G. Carmona-Catot, R.-M. Araguas, O. Vidal, N. Sanz, E. García-Berthou & J.-L. García-Marín, 2013. Gene flow and maintenance of genetic diversity in invasive Mosquitofish (Gambusia holbrooki). Plos One 8: e82501.
Durán, C. & A. Anadón, 2008. The zebra mussel invasión in Spain and navigation rules. Aquatic Invasions 3: 315–324.
Durán, C., M. Lanao, L. Pérez, C. Chica, A. Anadón & V. Touya, 2012. Estimación de los costes de la invasión del mejillón cebra en la cuenca del Ebro (período 2005-2009). Limnetica 31: 213–230.
Earl, D. A. & B. M. von Holdt, 2012. STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conservation Genetics Resources 4: 359–361.
Evanno, G., S. Regnaut & J. Goudet, 2005. Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Molecular Ecology 14: 2611–2620.
Felsenstein, J., 2005. PHYLIP (Phylogeny Inference Package) version 3.6. Department of Genome Sciences, University of Washington, Seattle.
Frantz, A. C., S. Cellina, A. Krier, L. Schley & T. Burke, 2009. Using spatial Bayesian methods to determine the genetic structure of a continuously distributed population: clusters or isolation by distance? Journal of Applied Ecology 46: 493–505.
Gelembuik, G. W., G. E. May & C. E. Lee, 2006. Phylogeography and systematics of zebra mussels and related species. Molecular Ecology 15: 1033–1050.
Gosling, E., I. Astanei & A. Was, 2008. Genetic variability in Irish populations of the invasive zebra mussel, Dreissena polymorpha: discordant estimates of population differentiation from allozymes and microsatellites. Freshwater Biology 53: 1303–1315.
Goudet, J., 2001. FSTAT: a computer program to estimate and test gene diversities and fixation indices (version 2.9.3) [available on internet at http://www2.unil.ch/popgen/softwares/fstat.htm ].
Guo, S. W. & E. A. Thompson, 1992. Performing the exact test of Hardy-Weinberg proportions for multiple alleles. Biometrics 48: 361–372.
Havel, J. E., K. E. Kovalenko, S. M. Thomaz, S. Amalfitano & L. B. Kats, 2015. Aquatic invasive species: challenges for the future. Hydrobiologia 750: 147–170.
Johnson, L. E. & J. T. Carlton, 1996. Post-establishment spread in large-scale invasions: dispersal mechanisms of the zebra mussel Dreissena polymorpha. Ecology 77: 1686–1690.
Karatayev, A. Y., L. E. Burlakova & D. K. Padilla, 2015. Zebra versus quagga mussels: a review of their spread, population dynamics, and ecosystem impacts. Hydrobiologia 746: 97–112.
Kirsch, K. M. & A. R. Dzialowski, 2012. Effects of invasive zebra mussels on phytoplankton, turbidity, and disolved nutrients in reservoirs. Hydrobiologia 686: 169–179.
Latch, E. K., G. Dharmarajan, J. C. Glaubitz & O. E. Rhodes Jr, 2006. Relative performance of Bayesian clustering software for inferring population substructure and individual assignment at low levels of population differentiation. Conservation Genetics 7: 295–302.
Lewis, K. M., J. L. Feder & G. A. Lamberti, 2000. Population genetics of the zebra mussel Dreissena polymorpha (Pallas): local allozyme differentiation within midwestern lakes and streams. Canadian Journal of Fisheries and Aquatic Sciences 57: 637–643.
Lopes-Lima, M., A. Teixera, E. Froufe, A. Lopes, S. Varandas & R. Sousa, 2014. Biology and conservation of freshwater bivalves: past, present and future perspectives. Hydrobiologia 735: 1–13.
Lowe, S., M. Browne, S. Boudjelas & M. de Poorter, 2000. 100 of the World’s worst invasive alien species: a selection from the Global Invasive Species Database. Aliens 12: 1–12.
Manni, F., E. Guerard & E. Heyer, 2004. Geographic patterns of (genetic, morphologic, linguistic) variation: how barriers can be detected by using Monmonier’s algorithm. Human Bioloby 76: 173–190.
Marescaux, J., P. Boets, J. Lorquet, R. Sablon, K. Van Doninck & J. N. Beisel, 2015. Sympatric Dreissena species in the Meuse River: towards a dominance shift from zebra to quagga mussels. Aquatic Invasions 10: 287–298.
Marsden, J. E., A. Spidle & B. May, 1995. Genetic similarity among zebra mussel populations within North America and Europe. Canadian Journal of Fisheries and Aquatic Sciences 52: 836–847.
May, G. E., G. W. Gelembuik, V. E. Panov, M. I. Orlova & C. E. Lee, 2006. Molecular ecology of zebra mussel invasions. Molecular Ecology 15: 1021–1031.
Müller, J. C., D. Hidde & A. Seitz, 2002. Canal construction destroys the barrier between major European invasion lineages of the zebra mussel. Proceedings of the Royal Society 269: 1139–1142.
Naish, K. A. & E. Gosling, 2001. Trinucleotide microsatellite loci for the zebra mussel Dreissena polymorpha, an invasive species in Europe and North America. Molecular Ecology Notes 1: 286–288.
Navarro, A., J. Sanchez-Fontenla, D. Cordero, M. Faria, J. B. Pena, C. Saavedra, M. Blazquez, O. Ruiz, R. Urena, A. Torreblanca, C. Barata & B. Pina, 2013. Genetic and phenoptypic differentiation of zebra mussel populations colonizing Spanish river basins. Ecotoxicology 22: 915–928.
Nei, M., F. Tajima & Y. Tateno, 1983. Accuracy of estimated phylogenetic trees from molecular data. Journal of Molecular Evolution 19: 153–170.
Oscoz, J., P. Tomás & C. Durán, 2010. Review and new records of non-indigenous freshwater invertebrates in the Ebro River basin (Northeast Spain). Aquatic Invasions 5: 263–284.
Paetkau, D., R. Slade, M. Burden & A. Estoup, 2004. Genetic assignment methods for the direct, real-time estimation of migration rate: a simulation-based exploration of accuracy and power. Molecular Ecology 13: 55–65.
Peñarrubia, L., N. Sanz, C. Pla, O. Vidal & J. Viñas, 2015. Using Massive Parallel Sequencing for the development, validation, and application of population genetics markers in the invasive bivalve zebra mussel (Dreissena polymorpha). PLoS One 10: e0120732.
Piry, S., A. Alapetite, J. M. Cornuet, D. Paetkau, L. Baudouin & A. Estoup, 2004. GENECLASS2: a software for genetic assignment and first-generation migrant detection. Journal of Heredity 95: 536–539.
Pritchard, J. K., M. Stephens & P. Donnelly, 2000. Inference of population structure using multilocus genotype data. Genetics 155: 945–959.
Popa O. P. & D. Murariu, 2009. Freshwater bivalve molluscs invasive in Romania. In: Pyšek P., J. Pergl (eds). Biological Invasions: Towards a Synthesis. Neobiota 8: 123–133.
Rajagopal, S., B. J. A. Pollux, J. L. Peters, G. Cremers, S. Y. Moon-van der Staay, T. van Alen, J. Eygensteyn, A. van Hoek, A. Palau, A. bij de Vaate & G. van der Velde, 2009. Origin of Spanish invasion by the zebra mussel, Dreissena polymorpha (Pallas, 1771) revealed by amplified fragment length polymorphism (AFLP) fingerprinting. Biological Invasions 11: 2147–2159.
Rannala, B. & J. L. Mountain, 1997. Detecting immigration by using multilocus genotypes. Proceedings of the National Academy of Sciences of the United States of America 94: 9197–9201.
Raymond, M. & F. Rousset, 1995. GENEPOP (Version 1.2): population Genetics Software for Exact Tests and Ecumenicism. Journal of Heredity 86: 248–249.
Rice, W. R., 1989. Analyzing tables of statistical tests. Evolution 43: 223–225.
Ruíz-Altaba, C., P. J. Jiménez & M. A. López, 2001. El temido mejillón cebra empieza a invadir los ríos españoles desde el curso bajo del río Ebro. Quercus 188: 50–51.
Sanz-Ronda, F. J., S. Lopez-Saenz, R. San-Martin & A. Palau-Ibars, 2014. Physical habitat of zebra mussel (Dreissena polymorpha) in the lower Ebro River (Northeastern Spain): influence of hydraulic parameters in their distribution. Hydrobiologia 735: 137–147.
Sanz, N., R. M. Araguas, O. Vidal, D. Diez-del-Molino, R. Fernández-Cebrián & J. L. García-Marín, 2013. Genetic characterization of the invasive mosquitofish (Gambusia spp.) introduced to Europe: population structure and colonization routes. Biological Invasions 15: 2333–2346.
Son, M. O., 2007. Native range of the zebra mussel and quagga mussel and new data on their invasions within the Ponto-Caspian Region. Aquatic Invasions 2: 174–184.
Sousa, R., A. Novais, R. Costa & D. Strayer, 2014. Invasive bivalves in fresh waters: impacts from individuals to ecosystems and possible control strategies. Hydrobiologia 735: 233–251.
Spilinga, C., U. Chiappafreddo & Q. Pirisinu, 2000. Dreissena polymorpha (Pallas) al Lago Trasimeno. Rivista di Idrobiologia 39: 145–152.
Stepien, C. A., C. D. Taylor & K. A. Dabrowska, 2002. Genetic variability and phylogeographic patterns of a nonindigenous species invasion: a comparison of exotic versus native zebra and quagga mussel populations. Journal of Evolutionary Biology 15: 314–328.
Stepien, C. A., J. E. Brown, M. E. Neilson & M. A. Tumeo, 2005. Genetic diversity of invasive species in the Great Lakes versus their Eurasian source populations: insights for risk analysis. Risk Analysis 25: 1043–1060.
Stoeckel, J. A., D. W. Schneider, L. A. Soeken, K. D. Blodgett & R. E. Sparks, 1997. Larval dynamics of a riverine metapopulation: implications for zebra mussel recruitment, dispersal and control in a large river system. Journal of the North American Benthological Society 16: 586–601.
Strayer, D. L. & H. M. Malcom, 2007. Effects of zebra mussels (Dreissena polymorpha) on native bivalves: the beginning of the end or the end of the beginning? Journal of the North American Benthological Society 26: 111–122.
Tarnowska, K., C. Daguin-Thiebaut, S. Pain-Devin & F. Viard, 2013. Nuclear and mitochondrial genetic variability of an old invader, Dreissena polymorpha (Bivalvia), in French river basins. Biological Invasions 15: 2547–2561.
Van Oosterhout, C., W. F. Hutchinson, D. P. M. Wills & P. Shipley, 2004. MICRO-CHECKER: software for identifying and correcting genotyping errors in microsatellite data. Molecular Ecology Notes 4: 535–538.
Weir, B. S., 1990. Genetic Data Analysis. Sinauer, Sunderland.
Weir, B. S. & C. C. Cockerham, 1984. Estimating F-statistics for the analysis of population structure. Evolution 38: 1358–1370.
Wilson, G. A. & B. Rannala, 2003. Bayesian inference of recent migration rates using multilocus genotypes. Genetics 163: 1177–1191.
Acknowledgements
We thank many colleagues for D. polymorpha sample collections: R. Miranda (Universidad de Navarra, Spain), C. Alcaraz and M. Pla (IRTA, Government of Catalonia, Spain), R. Mansfield (University of Manchester, UK), B. Gallardo and C. McLaughlan (University of Cambridge, UK), E. Goretti (Università degli Studi di Perugia, Italy), A. Binelli (University of Milan, Italy), J. F. Fruget (Université de Lyon, France), J. N. Beiseland (Université de Lorraine, France), J. Alvarado-Bremer (A&M University at Galveston, U.S.A), and O. Popa and A. M. Krapaland (National Museum of Natural History, Romania). This research was made within the project CGL200909407 of the Spanish Government (MICINN). L. Peñarrubia received a PhD fellowship support from the Spanish MICINN (BES – 2010037446).
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Diego Fontaneto
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Peñarrubia, L., Vidal, O., Viñas, J. et al. Genetic characterization of the invasive zebra mussel (Dreissena polymorpha) in the Iberian Peninsula. Hydrobiologia 779, 227–242 (2016). https://doi.org/10.1007/s10750-016-2819-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10750-016-2819-2