Abstract
This study tests population genetic patterns across the Eurasian dreissenid mussel invasions of North America—encompassing the zebra mussel Dreissena polymorpha (1986 detection) and the quagga mussel D. rostriformis bugensis (detected in 1990, which now has largely displaced the former in the Great Lakes). We evaluate their source-spread relationships and invasion genetics using 9–11 nuclear microsatellite loci for 583 zebra mussels (21 sites) and 269 quagga mussels (12 sites) from Eurasian and North American range locations, with the latter including the Great Lakes, Mississippi River basin, Atlantic coastal waterways, Colorado River system, and California reservoirs. Additionally, mtDNA cytochrome b gene sequences are used to verify species identity. Our results indicate that North American zebra mussels originate from multiple non-native northern European populations, whereas North American quagga mussels trace to native estuaries in the Southern Bug and Dnieper Rivers. Invasive populations of both species show considerable genetic diversity and structure (zebra F ST = 0.006–0.263, quagga F ST = 0.008–0.267), without founder effects. Most newer zebra mussel populations have appreciable genetic diversity, whereas quagga mussel populations from the Colorado River and California show some founder effects. The population genetic composition of both species changed over time at given sites; with some adding alleles from adjacent populations, some losing them, and all retaining closest similarity to their original composition. Zebra mussels from Kansas and California appear genetically similar and assign to a possible origin from the St. Lawrence River, whereas quagga mussels from Nevada and California assign to a possible origin from Lake Ontario. These assignments suggest that overland colonization pathways via recreational boats do not necessarily reflect the most proximate connections. In conclusion, our microsatellite results comprise a valuable baseline for resolving present and future dreissenid mussel invasion pathways.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Genetic studies using high-resolution microsatellite DNA loci offer means to elucidate the pathways and vectors of invasive species, and assess their temporal population dynamics. This is the first comprehensive population genetic analysis of the Eurasian dreissenid mussel (Mollusca: Bivalvia: Dreissenidae) invasion across North America, which has been one of the most ecologically and economically important aquatic introductions. The zebra mussel Dreissena polymorpha appeared in the Great Lakes in 1986, where it was introduced via ballast water (Hebert et al. 1989; Carlton 2008), followed by the quagga mussel D. rostriformis bugensis in ~1989 (May and Marsden 1992; Mills et al. 1996). The zebra mussel had previously considerably expanded its original Ponto-Eurasian range via central European canals in the Black Sea and Baltic Sea drainages (Morton 1993; Bij de Vaate et al. 2002), whereas the Great Lakes was one of the quagga mussel’s first documented expansions (Pligin 1979; Karatayev et al. 2008). The zebra mussel also spread from the Baltic Sea drainage to Italy (1960s), Ireland (1997), and Spain (2001) by recreational boats (Bij de Vaate et al. 2002; Aldridge et al. 2004; Gosling et al. 2008).
Dreissenid mussels widely disperse via their planktonic larvae, whereas juveniles and adults adhere to hard substrates with byssal threads (summarized by Ackerman et al. 1994). These life history traits distinguish dreissenids from most native North American freshwater bivalves, which lack a planktonic larval stage and do not attach to hard substrates (Johnson and Padilla 1996). The spread of dreissenids was facilitated by their life history traits, including unintentional transport of larvae in ships’ ballast and bilge water, and by attachment of their juveniles and adults to engines, hulls, and anchors (Ricciardi et al. 1995; Mackie and Schloesser 1996; Johnson et al. 2001). As they have spread, dreissenid mussels have exerted both economic and ecological impacts. Notably, their fouling of power generation and water treatment facilities in the United States and Canada has cost $161–$467 million/year (Connelly et al. 2007). They filter feed on phytoplankton with high efficiency that has shifted the basis of Great Lakes’ aquatic food chains from pelagic to benthic (Berg et al. 1996; Zhu et al. 2006). Spread of dreissenid “mats” on the lake floors—where they host an associated community of plants and invertebrates in their shells and interstices—has further increased benthification, markedly changing the ecosystem (Zhu et al. 2006; Hebert et al. 2009; Stewart et al. 2009). Dreissenids likely facilitated the invasion of the round goby Neogobius melanostomus (Vanderploeg et al. 2002), their natural predator that became established throughout the Great Lakes, whose populations have high genetic diversity and show no founder effect (Stepien and Tumeo 2006; Brown and Stepien 2009).
Upon establishment in North America, the zebra mussel spread rapidly, expanding throughout the Great Lakes and into the Mississippi and Hudson River drainages within 5 years of detection (Nalepa and Schloesser 1993; Allen and Ramcharan 2001; Kelly et al. 2009). The quagga mussel spread more slowly (Kelly et al. 2009), gradually replacing zebra mussels in regions of the St. Lawrence River (Ricciardi and Whoriskey 2004) and the lower Great Lakes (Mills et al. 1999; Jarvis et al. 2000; Wilson et al. 2006), and is now doing the same in the upper Lakes (Nalepa et al. 2009). Researchers and managers attempted to stop westward spread at the 100th meridian, to prevent colonization of the heavily dammed and diverted waterways of the western United States (Mangin 2001; Britton and McMahon 2005; Bossenbroek et al. 2007, 2009). However, in 2007 the quagga mussel was detected in Lake Mead, Nevada—a reservoir on the Colorado River—and then discovered downstream (Stokstad 2007). Dreissenids since have spread into many southern California reservoirs via the extensive canal system radiating from the Colorado River (http://nas.er.usgs.gov/queries/FactSheet.asp?speciesID=95). In 2008, zebra mussels were detected in San Justo Reservoir in central California (http://nas.er.usgs.gov/queries/FactSheet.asp?speciesID=5). It is important that we understand the population sources and pathways of these recent expansions so that we can evaluate the efficacy of our control efforts and attempt to avoid further expansions.
Role of genetics
Modern genetics can play a crucial role to aid invasive species identification and evaluate their population linkages. Hebert et al. (1989), May and Marsden (1992), and Stepien et al. (1999) used genetics to determine the identity of dreissenid invaders in the Great Lakes. Additional genetic research has examined portions of the invasive ranges for zebra (Elderkin and Klerks 2001; Stepien et al. 2001, 2003, 2005; Gosling et al. 2008) and quagga mussels (Stepien et al. 2001, 2003, 2005), illustrating the need for more rapidly-evolving reproducible genetic markers to understand their origin and spread, compare genetic variation, determine population genetic structure, and predict their expansion pathways.
The genetic composition of an introduced population can be substantially altered from its native source(s), including the amount of overall variation and its geographic partitioning. Genetic variation comprises the raw material for adaptive evolution, which can play a critical role governing invasive success (Blossey and Notzold 1995; Bossdorf et al. 2005). Selectively neutral molecular markers can serve as a proxy for estimating quantitative genetic variation in phenotypic traits that underlies adaptive evolution (Merila and Crnokrak 2001; Reed and Frankham 2001; Chun et al. 2009). Population genetic data also are useful for elucidating the number and sources of an introduction (Stepien et al. 2005; Rosenthal et al. 2008; Brown and Stepien 2009), as well as determining whether changes occur during the time course of its establishment and spread (Andreakis et al. 2009; Henry et al. 2009). Such information can aid control efforts. For example, determining the genetic origin of the invasive climbing fern Lygodium microphyllum led to a biological control agent that was matched to its native genotypes (Goolsby et al. 2004).
The genetic variation of an introduced population may be reduced, increased, or unchanged relative to native populations with reduction as the most common outcome (Nei et al. 1975; Ficetola et al. 2008; Henry et al. 2009). Three processes may reduce genetic diversity of the introduction and preclude significant variation among local population groups. First, most introductions are founded by only a few individuals that represent only a small part of native variation, and subsequent bottlenecks and Allee effects may further reduce it (DeWalt and Hamrick 2004; Grapputo et al. 2005; Griffen and Drake 2008). Second, introduction and spread likely lead to high gene flow that homogenizes population genetic structure (Viard et al. 2006; Kim et al. 2009). Lastly, a recent introduction may lack sufficient time for local adaptation or drift to generate local population genetic structure via stochastic processes in the new range (Maron et al. 2004; Whitney and Gabler 2008). Any new genetic variation would need to arise from mutations and recombinations, before population structure could develop (Perez et al. 2008). Selection also might lead to reduced genetic diversity; since when an introduced genotype is more fit, its numbers will increase relative to others, resulting in a net loss of genetic variability (Kliber and Eckert 2005). If these processes predominate during an introduction, the new population should be genetically depauperate in comparison to native populations (Amsellam et al. 2000; DeWalt and Hamrick 2004; Grapputo et al. 2005).
If an introduction is founded from multiple genetically differentiated native populations, its resultant genetic diversity can be higher than that characterizing native populations, therefore possibly increasing heterozygosity and/or generating novel allelic combinations (Ellstrand and Schierenbeck 2000). Several studies have shown patterns indicating multiple introductions and high genetic diversity in invaders (Kolbe et al. 2004; Williams et al. 2005; Stepien et al. 2005; Brown and Stepien 2009). Because increased genetic diversity likely also increases potential for adaptive evolution, an invader with high diversity may evolve quickly in its new range. Another possible evolutionary outcome for an introduction is no significant change in genetic diversity between the native and introduced ranges. This is possible if large population size is maintained during an invasion (Brown and Marshall 1981) or regained shortly following introductions (Zenger et al. 2003).
Understanding the origin of population genetic structure, or lack thereof, in an invasive species also can provide valuable information on transport pathways from source populations. Correctly identifying transport pathways and source populations is critical to interpreting an invader’s relative ecological success in introduced versus native ranges and for targeting management efforts to shut down invasion pathways (Amsellam et al. 2000; Kang et al. 2007). Like the zebra and quagga mussels, 73% of Great Lakes invaders entered via ballast water exchange from oceanic vessels (Holeck et al. 2004; Ricciardi 2006; Kelly et al. 2009) and 70% of those from 1985 to 2000 traced to the Eurasian Ponto-Caspian region (Ricciardi and MacIsaac 2000). Three shipping routes extend from the Ponto-Caspian region, across the Atlantic, and into the Great Lakes, including: (1) from the Black Sea through the Mediterranean Sea, (2) from the Danube River into the North Sea, or (3) from the Dnieper River into the Baltic Sea (Ricciardi and MacIsaac 2000; Grigorovich et al. 2002). These pathways are evaluated in our study for the zebra and quagga mussel introductions.
Relatively few Great Lakes-introduced species have been genetically assessed to date. Both the spiny water flea Bythotrephes longimanus (Berg et al. 2002) and the ruffe Gymnocephalus cernuus (Stepien et al. 1998, 2004, 2005) showed evidence of single-source colonization. The water flea originated from Lake Ladoga in Russia (Berg et al. 2002) and the ruffe from the Elbe River system in Germany (Stepien et al. 2004, 2005). All the Elbe River ruffe genotypes are present in the upper Great Lakes in similar frequencies, and likely were introduced into the upper Great Lakes via increased US shipping trade with Germany near the time of its reunification (Stepien et al. 2004, 2005). In contrast, Chinese mitten crab Eriocheir sinensis introductions into the Great Lakes traced to multiple European ports rather than native Asian populations (Tepolt et al. 2007); however, this species does not reproduce in freshwater and thus failed to establish (Herborg et al. 2007). The Great Lakes introduction of the freshwater tubenose goby Proterorhinus semilunaris has been characterized by levels of genetic diversity that are similar to native river source populations (Stepien et al. 2005; Stepien and Tumeo 2006; Neilson and Stepien 2009). Invasive Ponto-Caspian round goby Neogobius melanostomus populations in the Great Lakes have considerable genetic variation and population structure in comparison with native populations (Dillon and Stepien 2001; Stepien et al. 2005; Stepien and Tumeo 2006; Brown and Stepien 2008), and the Dnieper River population was identified as its primary source (Brown and Stepien 2009).
Genetic studies of the dreissenid mussel invasion by Marsden et al. (1995) and Stepien et al. (1999, 2002, 2005) separately described relatively high levels of genetic diversity, suggesting little overall loss of genetic diversity in comparison with putative source populations. Allozyme markers by Marsden et al. (1995) appeared to suggest a single source for the zebra mussel invasion, yet a later allozyme study showed significant genetic heterogeneity among populations in the Great Lakes and a complex of inland lakes, suggesting an important role for stochastic processes governing their population genetic structure (Lewis et al. 2000). Subsequent genetic studies using mitochondrial and nuclear DNA markers similarly found high genetic diversity and implicated multiple founding sources, pointing to western central European sources for the zebra mussel (Stepien et al. 2002, 2005). In this new study, we analyze the invasion genetics of the zebra mussel and the quagga mussel in North America and Eurasia using high-resolution nuclear microsatellite markers for the first genetic study across their expanded geographic range.
We test the following hypotheses underlying the genetic history of the dreissenid invasions in North America:
-
1.
Each species was founded by a single versus multiple source populations.
-
2.
The invasive populations are genetically homogenous/heterogeneous across their ranges.
-
3.
Genetic diversity levels reflect overall founder effects/show no founder effects.
-
4.
Populations outside the Great Lakes have reduced/similar/greater genetic diversity compared to those within the Great Lakes.
-
5.
Once established, populations remain genetically static/change over time.
Materials and methods
Sampling, DNA extraction, and amplification
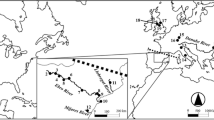
We sampled 24 zebra mussel (N = 583) and 13 quagga mussel (N = 269) population locations across North America and Eurasia that were collected from 1991 to 2008 (Fig. 1, Table 1). Five locations (three for zebra mussels and two for quagga mussels) had samples from more than 1 year, which were compared to test for temporal population changes. Samples were collected by hand or by fishery agency trawls, and were either immediately frozen, or placed in 95% ethanol and stored at room temperature. Genomic DNA was extracted from mantle tissue using Qiagen DNeasy kits (Qiagen, Inc.; Valencia, CA), eluted in 100 μl of water, stored at 4°C until used for amplification, and then archived at −80°C.
Maps showing current range and sampling locations for zebra mussels and quagga mussels. A North America (USGS; http://nas.er.usgs.gov/taxgroup/mollusks/zebramussel/), B Eurasia (DAISIE; http://www.europe-aliens.org/)
Genetic variation was assessed from eight microsatellite (μsat) loci for zebra mussels (identified as Dpo) and six for quagga mussels (Dbu) developed by Feldheim et al. (in review), and three additional previously published loci per species (Table 2; Wilson et al. 1999; Naish and Boulding 2001; Feldheim et al. in review). PCR amplifications were performed in our laboratory using 10 μl reaction volumes of 0.6 units Taq, 50 μM nucleotides, 50 mM KCl, 2.5 mM MgCl2, 10 mM Tris–HCl, 0.5 μM of each primer, and 30 ng of template; with a mineral oil overlay to maintain reaction volume. A thermal cycle of 2 min at 94°C for initial denaturation was followed by 35 cycles of denaturation at 94°C for 30 s, annealing for 1 min at 52–58°C (primer-specific; Table 2), and extension at 72°C for 30 s; capped by a final extension at 72°C for 5 min. Amplification products were diluted 1:50, of which 1 μl was added to 13 μl of formamide and ABI (Applied Biosystems, Inc.; Foster City, California) Gene Scan 500 size standard, and loaded onto a 96-well plate for analysis on an ABI 3130xl Genetic Analyzer using GeneMapper v4.0. Output profiles were checked to confirm allelic size variants.
A portion of the mtDNA cyt b gene was amplified from randomly selected representative individuals to confirm species identity at sites using polymerase chain reactions (PCR) of 25 μl, including 1 unit Taq polymerase, 200 μM dNTPs, 50 mM KCl, 2 mM MgCl2, 10 mM Tris–HCl, 0.5 μM of each primer (151F, 270R; Merritt et al. 1998), and 30 ng of template. Amplification on a MJR DYAD thermalcycler (Bio-Rad Laboratories, Hercules, CA) consisted of an initial denaturation step at 94°C for 2 min, followed by 35 cycles of denaturation at 94°C for 45 s, annealing at 52°C for 30 s, and extension at 72°C for 1 min; concluded by a final 72°C extension for 3 min.
Sequencing of the mitochondrial (mt) cytochrome (cyt) b gene (336 bp) used the primers 151F and 270R, and was outsourced to the Cornell University Life Sciences Core Laboratories Center (http://cores.lifesciences.cornell.edu/brcinfo, Ithaca, NY). Sequences then were trimmed by us to remove primer sequences, aligned with CLUSTAL X v2.0 (Larkin et al. 2007), and adjusted using BIOEDIT v7.0 (Hall 1999, 2004). These sequences are deposited in GenBank as accession numbers GQ988724-32 for D. polymorpha and GQ988733-45 for D. rostriformis bugensis.
Genetic analyses
ARLEQUIN (v3.11; http://cmpg.unibe.ch/software/arlequin3; Excoffier et al. 2005) was used to assign individuals to the correct mtDNA cyt b haplotype; and allelic frequencies, number of private alleles, conformance of μsats to Hardy–Weinberg (HW) equilibrium expectations, and linkage disequilibrium were evaluated in GENEPOP v4.0 (http://kimura.univ-montp2.fr/%7Erousset/Genepop.htm; Rousset 2008). HW deviations were tested for heterozygosity deficiency or excess, and for the presence of null alleles using MICRO-CHECKER v2.23 (http://www.microchecker.hull.ac.uk; van Oosterhout et al. 2004, 2006). Levels of significance for all tests were adjusted using Bonferroni correction (Sokal and Rohlf 1995).
Microsatellite genetic composition of the samples was analyzed to identify true populations (i.e., those distinguished by significantly divergent gene pools) using the F-statistic analog θ ST (Weir and Cockerham 1984) and contingency tests (Raymond and Rousset 1995; Goudet et al. 1996). Relationships among recently diverged samples, such as those tested here, have been shown to be better resolved in models using contingency tests (see Balloux and Lugon-Moulin 2002), which are independent of HW equilibrium assumptions, non-parametric, and unaffected by sample size (Raymond and Rousset 1995; Goudet et al. 1996). Our use of the F-statistic analog facilitated direct comparisons with other studies (see Brown and Stepien 2008, 2009). Probability values of both tests were adjusted using sequential Bonferroni corrections (Rice 1989). In order to further analyze the relationships among population sites, pairwise genetic distances were calculated based on the microsatellite data using Cavalli-Sforza chord distances (Cavalli-Sforza and Edwards 1967) and neighbor-joining trees (Saitou and Nei 1987) were constructed in PHYLIP v.3.68 (http://www.phylip.com/; Felsenstein 1989). The trees were rooted to their respective close relatives D. stankovici and D. rostriformis grimmi (see Stepien et al. 2003, 2005 for phylogeny) using microsatellite data from Feldheim et al. (in review). Three-dimensional factorial correspondence analysis (3D-FCA; Benzecri 1973) was used to examine population divisions in GENETIX v.4.05 (http://www.genetix.univ-montp2.fr/genetix/genetix.htm; Belkhir et al. 2004) via a visual representation of the relationships among populations without a priori assumptions about populations, evaluating variation within and among sites.
Populations that experience a genetic bottleneck usually exhibit a decrease in the number of alleles at polymorphic microsatellite loci faster than their respective heterozygosity declines (Luikart et al. 1998a). In order to detect such a signal, a test for heterozygosity excess was carried out using the program BOTTLENECK 1.2.02 (http://www.ensam.inra.fr/URLB/bottleneck/bottleneck.html; Cornuet and Luikart 1996; Piry et al. 1999). This test compared the observed gene diversity (Nei 1987) to the expected equilibrium gene diversity, which is calculated from the observed number of alleles assuming a constant-size (i.e., equilibrium) population (Luikart et al. 1998b). Wilcoxon tests using a stepwise mutation model were used to establish whether the number of loci showing heterozygosity excess was significantly greater than that expected in populations at equilibrium, with distribution of expected gene diversity at equilibrium estimated from 10,000 simulations (Cornuet and Luikart 1996).
The Bayesian model-based methods of Rannala and Mountain (1997) in STRUCTURE v 2.3.1 (http://pritch.bsd.uchicago.edu/structure.html; Pritchard et al. 2000; Pritchard and Wen 2004) and GENECLASS2 (http://www.ensam.inra.fr/URLB/GeneClass2/Installation.htm; Piry et al. 2004) use the individual as the unit, assigning it to the most likely population group(s) regardless of geographic origin. STRUCTURE was used to assign individuals to population groups, ranging from K = 1 (a single population group, i.e., the null hypothesis of panmixia) to K = N (the total N of sampling groups), with the relative frequency of individual membership per group totaling 1.00. Ten independent runs for each K were used with burn-ins of 100,000 replicates and 1,000,000 replicates. We complemented this analysis with assignment tests using GENECLASS2, which assigned each individual a probability of membership among each sampled location using simulated population sizes of 10,000 individuals per sampling site and a 0.01 rejection level (Cornuet et al. 1999).
Results
We recovered 386 zebra mussel alleles from 11 loci and 228 quagga mussel alleles using nine loci. All populations for both species conformed to Hardy–Weinberg equilibrium expectations, all loci were unlinked, and there was no evidence of null alleles.
Our results support that both dreissenid introductions to North America originated from multiple Eurasian sources. For the zebra mussel, the 3D-FCA (Fig. 2A) and the neighbor-joining tree (Fig. 3A) analyses indicate at least 2 different introductions, with Lake Erie samples from both years being distinct from other North American samples. The majority of the North American samples group with introduced locations in northern Europe and the Volga River, but the Lake Erie samples do not group with any locations we tested here.
Three-dimensional factorial correspondence analysis (GENETIX v.4.05; Belkhir et al. 2004) for A zebra mussels and B quagga mussels. A Zebra mussels in L. Erie were distinct from all other samples. B Quagga mussel samples separate into 3 distinct groups, with western North American samples clustering with those from L. Ontario. Years are given for samples from the same location, but from different years
Genetic distance trees among A zebra mussel and B quagga mussel population sites constructed in PHYLIP v3.6 (Felsenstein 1989) with Cavalli-Sforza chord distances (Cavalli-Sforza and Edwards 1967) based on microsatellite data. Bootstrap percentage values are from 1000 pseudoreplicates. Branch lengths are proportional to genetic divergences. Years are given for samples from the same location, but from different years
The 3D-FCA (Fig. 2B) and the neighbor-joining tree (Fig. 3B) for quagga mussels are largely congruent, highlighting distinct introduction events—one that contributed the majority of the diversity found in North American samples, and a second that influenced diversity in Lake Erie (evident in samples from both years). Quagga mussels that were introduced to the North American Great Lakes appear to trace to the general area of the Southern Bug and Dnieper River estuaries, which are geographically proximate estuaries (~60 km).
Invasive population structure
All introduced zebra mussel populations appear significantly different from each other in pairwise tests (Tables 3A, 4A), showing a high degree of genetic heterogeneity across their ranges, with the exception of a single comparison between the Kansas locations. These two locations do not significantly differ from each other, but diverge from all other populations (Table 3A). Similarly, the only population sites with low self-assignment in the GENECLASS test were the Kansas samples, which had high assignments to each other (Table 5A). Bayesian assignment analysis reveals 5 major groups within the zebra mussel data set (Fig. 4B). One (colored blue in Fig. 4A) links most of the introduced Eurasian populations (sites 16–19) with the Volga River Russia samples (site 21). A second group (colored pink) differentiates the Lake Erie (site 10, ‘02, ‘04) population from all others. The third group (colored tan) links the Dnieper River (site 20) with samples from the Mississippi River drainage (sites 4–6), Oneida Lake (site 13) and the Hudson River (site 15, 2003 only). The fourth group (colored green) links samples from the upper Great Lakes (sites 7–9) and the earlier Hudson River sampling date (site 15, 1991). The final group (colored brown) assigns zebra mussel locations in the western United States (site 1–3) to a group that includes the Lake Ontario (site 11) and St. Lawrence River (site 14) samples, suggesting that the eastern Great Lakes may have served as the source for this westward expansion. This grouping also has strong support in the 3D-FCA (Fig. 2A) and the neighbor-joining tree (Fig. 3A).
Bayesian STRUCTURE analysis (Pritchard et al. 2000; Pritchard and Wen 2004) for A zebra mussels and B quagga mussels. Analysis discerned 5 groups (posterior probability = 0.91) for zebra mussels and 3 groups (posterior probability = 0.89) for quagga mussels. Both species show evidence of multiple introductions. Recent populations established in western North America link to the eastern portions of the Great Lakes. Years are given for samples from the same location, but from different years
All quagga mussel populations significantly differ from one another in the pairwise tests (Table 3B). Their AMOVA groupings also are significant (Table 4A), and the populations show high self-assignment (Table 5B). The 3D-FCA (Fig. 2B), neighbor-joining tree (Fig. 3B), and Bayesian assignment test (Fig. 4B) are congruent in recovering 3 population groups. The first grouping links quagga mussel populations from the upper Great Lakes (sites C–D) with the invasive Volga River sample (site L). The second group comprises samples from Oneida Lake (H), the St. Lawrence River (I), the Dnieper River (site K), and some individuals from Lake Erie (E) in 2007. Quagga mussel sampling locations in the western United States (sites A, B) group with the Lake Ontario (F, G) samples in the 3D-FCA (Fig. 2B), the neighbor-joining tree (Fig. 3B), and the Bayesian assignment analysis (Fig. 4B); these suggests that Lake Ontario was a likely source for the quagga mussels in the Colorado River and southern California—similar to the pattern shown in zebra mussels.
Genetic diversity levels
Individual zebra mussel sampling locations have 65–151 total alleles (Table 1). Loci Dpo04 and Dpo272 have the fewest alleles (13), whereas locus DpolB8 has the most (58; Table 2A). The introduced population site in Spain (map location 16) possesses the fewest alleles, and the native Dnieper River (site 20) population has the greatest number (Fig. 1A). Great Lakes samples average 90 alleles, whereas native Eurasian populations average 113, which is a significant difference (t-test t = 2.168, df = 13, P = 0.0492). Samples from long established locations in the Mississippi River Basin (sites 4–6) average 105 alleles and the recent western expansion locations (1–3) average 106 alleles. Both are significantly higher than the average for the Great Lakes locations (P = 0.042 and P = 0.002, respectively). Zebra mussels show a decrease in average number of alleles per location from Eurasia to the Great Lakes, but subsequent expansion areas have higher numbers of alleles. However, these consist mostly of low frequency alleles (<0.01%) that also are found in various Great Lakes locations. Great Lakes zebra mussel samples have average observed heterozygosity (H O) values that are slightly lower than those found in the native Eurasian range (0.67 ± 0.08 vs. 0.74 ± 0.09), which is significant (t = 2.700, df = 13, P = 0.018), although the heterozygosity distribution does not suggest a genetic bottleneck for the North American introduction (P = 0.735) in the BOTTLENECK analysis. Zebra mussel populations outside the Great Lakes have somewhat lower H O (0.67 ± 0.08) than those within the Great Lakes (0.73 ± 0.04), which is not significant (t = −1.789, df = 16, P = 0.093). However, testing for genetic bottlenecks indicates that the three western expansion locations experienced bottlenecks (P = 0.0001 in each). In contrast, this analysis approach suggests that populations from Lake Superior (p = 0.011) and Lake Ontario (p = 0.005) underwent continuous population expansions.
Quagga mussel samples have 39–110 total alleles (Table 1), with loci Dbu75 and Dbu92 having the fewest alleles (13) and locus Dbug05 having the most (40; Table 2B). The introduced site in California (site A) has the fewest number of alleles, and the introduced population in Lake Huron (site D) possesses the most. Great Lakes sites average 78 alleles and native locations average 74; whose means significantly differ (t = 4.763, df = 9, P = 0.001). The recently established location in California and Nevada averages 48 alleles, which is significantly lower than the Great Lakes locations (t = 4.014, df = 8, P = 0.004). Great Lakes populations of quagga mussels have an average H O that appears slightly lower than in their native Eurasian range (0.76 ± 0.08 vs. 0.74 ± 0.05), but is not significant (t = −0.709, df = 9, P = 0.496). Quagga mussel samples outside the Great Lakes have lower H O than those found within (0.76 ± 0.08 vs. 0.60 ± 0.18), which is significant (t = 4.007, df = 8, P = 0.039). Great Lakes populations of quagga mussels have a greater than average number of alleles, but do not differ in observed heterozygosity from native Eurasian populations, with their heterozygosity distribution refuting a genetic bottleneck (P = 0.338). However, recent western expansions of the quagga mussel have lower average numbers of alleles and lower observed heterozygosities, supporting a founder effect. Moreover, heterozygosity excesses detected by the BOTTLENECK analysis for Lake Matthews CA (P = 0.018) and Lake Mead NV (P = 0.040) samples support a genetic bottleneck in the western expansion of the quagga mussel. The reduction appears greatest in the California sample, which traces to an origin from Lake Mead via the Colorado River aqueduct. Two samples show signs of population expansion detectable in the BOTTLENECK analysis: Lake Huron (P = 0.003) and western Lake Ontario (P = 0.024); these suggest rapid, continuous population growth following introduction.
Temporal genetic change
Temporal comparisons among three zebra mussel sampling locations reveal significant differences among years using both F ST and contingency tests (Table 3A), with strong self-assignment using GENECLASS (Table 5A). Thus all samples are most similar to their original population compositions, despite some temporal changes. Temporal comparisons of zebra mussel samples from Lake Erie (−16 alleles; 17%; θST = 0.091, P < 0.0001) and Lake Superior (−22 alleles; 21%; θST = 0.154, P < 0.0001) show a net loss of alleles (Table 1), whereas the Hudson River population reveals a net gain of alleles (+28 alleles; 33%; θST = 0.140, P < 0.0001). According to the BOTTLENECK analysis, the Lake Superior population lost the signal of rapid population expansion between 1995 and 2006. None of the other time pair comparisons show demographic signals in the BOTTLENECK analysis, suggesting that those changes did not result from changes in the size of the reproducing population.
For quagga mussels, only the Lake Erie location was tested for temporal change, and the two sampling years show a significant difference, with a large gain in alleles (+41 alleles; 84%; θST = 0.124, P < 0.0001). This renders the 2007 Lake Erie samples more similar to those from the St. Lawrence River and from the Black Sea, possibly due to population mixing caused by additional introductions into Lake Erie (Figs. 2B, 3B).
Discussion
Multiple founding sources contributed to significant genetic structure among dreissenid mussel populations of both species across their North American invasive ranges, likely reflecting some maintenance of native population differences due to differential colonization and expansion. In addition, the relatively small founder effects shown in the invasive dreissenid populations highlight the ease of introducing these organisms in large numbers—especially during their veliger larval stages. In both species, samples from Lake Erie appear genetically distinct, suggesting the likelihood of separate introductions from Eurasia. This appears especially likely given that both Mills et al. (1996) and Carlton (2008) found evidence of Lake Erie samples of both species (including shells on beaches, reports from divers on natural gas wellheads in northern Lake Erie, on water treatment filters in Ontario, and on the hull of a fishing vessel) preceding the formal discovery of dreissenid mussels in the Great Lakes. Boileau and Hebert (1993) also found that the Lake Erie population of the zebra mussel appeared genetically distinct from other Great Lakes and European populations using allozymes (which may have been influenced by the “cryptic” presence of the quagga mussel). Like Boileau and Hebert (1993), the Lake Erie zebra mussels we sampled do not show strong connections with any of the analyzed European locations in the present study. However, Stepien et al. (2002) found that Lake Erie samples were most similar to those from Poland and the Netherlands using randomly amplified nuclear polymorphic DNA. Our samples from those regions grouped with the other Great Lakes populations, and not with Lake Erie. Additional sampling and analyses from possible source regions thus is warranted.
Previously invaded locations often serve as hubs for further spread of an invader (Bossenbroek et al. 2009; Floerl et al. 2009; Kelly et al. 2009), which appears likely for the zebra mussel. These genotypes may be pre-adapted for invasive success. For example, the bloody red shrimp Hemimysis anomala is another Ponto-Caspian invader whose previously invaded Northern European locations served as sources for their establishment in the North American Great Lakes (Audzijonyte et al. 2008). Likewise, the Chinese mitten crab has been introduced to the Great Lakes from several European ports, but has failed to establish (Tepolt et al. 2007). Similarly, the Eurasian ruffe Gymnocephalus cernuus invaded the Great Lakes from the Elbe River via the Baltic Sea region (Stepien et al. 2005). The Great Lakes populations then served as such a hub for dreissenid expansion into the rest of North America (this study), and are predicted to serve as hubs for future further expansion into other parts of the world (Kelly et al. 2009).
In contrast to zebra mussels, North American quagga mussels appear to have arrived directly from their native Ponto-Caspian region, with their invasive populations tracing to the Southern Bug and Dnieper Rivers. This is congruent with results from Stepien et al. (2002), who analyzed nuclear RAPD markers to discern that quagga mussels from western Lake Erie were distinct from other North American quagga mussels, with all tracing back to a Dnieper River source (that study preceded our collections from the Southern Bug River that we analyzed here). Similarly, the North American round goby Neogobius melanostomus invasion traced back to the Dnieper River (Brown and Stepien 2009). The Dnieper River watershed thus served as a common source population for invasive species that became successfully established in the Great Lakes. The geographic difference in founding sources discerned between the two dreissenid species’ suggests that zebra and quagga mussels each arrived in the Great Lakes in unrelated introduction events—the zebra mussel originating on ships from the Baltic Sea and northern Europe, and the quagga mussel on ships from the Black Sea. These correspond to two of the main invasion pathways out of the Ponto-Caspian identified previously (Ricciardi and MacIsaac 2000; Grigorovich et al. 2002).
The observed heterozygosities we found for sampling locations of the North American quagga mussel (average H O = 0.73) appears higher than the diversity Therriault et al. (2005) discerned for invasive quagga mussel locations along the Volga River in Europe (average H O = 0.50; although the latter used fewer microsatellite markers than we did—6 compared to our 9). The number of alleles per population is similar between the two studies when populations are adjusted for sample size, suggesting that introductions in North America and the Volga River each had large founding populations.
Neither species experienced a genetic bottleneck when it was introduced to North America suggesting that a large number of propagules were introduced into the Great Lakes, which likely contributed to their successful establishment. Likewise, the brown mussel Perna perna introduction into the Gulf of Mexico did not show a bottleneck and successfully expanded its range following ballast water introduction from multiple sources in South America and southern Africa (Holland 2001). Similarly, the round goby did not undergo a genetic bottleneck during its introduction into the Great Lakes (Stepien and Tumeo 2006; Brown and Stepien 2009). All of these invasion examples thus were founded by large numbers of successful colonists. The high volumes of ship ballast tanks facilitate massive introductions (Kelly et al. 2009), providing early life history stages of species introduced by this vector a good chance of establishment (Bax et al. 2003; Drake and Lodge 2004; Holeck et al. 2004).
We also detected fine scale patterns of divergences among North American populations of both dreissenid mussel species. Their distinct population genetic structures likely resulted from long distance “jump” dispersal patterns from divergent sources via ballast water or recreational boating, followed by local spread. Many of these populations have high genetic variation, likely due to large founding numbers that may have been supplemented by continued migration events via commercial and recreational boating traffic. The genetic bottlenecks shown in the western populations of both species suggest that they likely were founded by relatively small numbers of individuals, with little subsequent gene flow. This fits the model of low-frequency transport via recreational boats predicted for dreissenid expansions into the western regions of North America (Johnson et al. 2001; Bossenbroek et al. 2007). In contrast, our samples from the edges of the Great Lake show evidence for having experienced periods of rapid growth, likely fueled by large introductions via commercial shipping. We discern significantly different genetic compositions of zebra mussel populations between the northern and southern Mississippi River sites, corroborating the gradient observed from allozyme data by Elderkin and Klerks (2001) and from nuclear AFLPs (amplified frequency length polymorphisms) by Elderkin et al. (2004). This genetic structure is attributed to boat-mediated introductions and the difficulty for free-spawning mussels to maintain consistent recruitment in lotic systems (Elderkin and Klerks 2001; Elderkin et al. 2004). Stepien et al. (2002) also found a marked genetic difference between the southern Mississippi River samples of the zebra mussel versus those from the Great Lakes. That neutral genetic difference may be accompanied by adaptive variation of genotypes in the south versus the north, which merits further analysis of quantitative adaptive traits and their heritability.
The significant temporal change in genetic structure of the tested populations suggests some allelic turnover at those locations, due to contribution by some recruits from other areas. However, the colonies retain their original population genetic structure, thus exhibiting genetic “resilience”. Haag and Garton (1995) found significant genetic differences among zebra mussel larvae and adults from western Lake Erie (the same location as our Lake Erie samples), indicating possible selection. Additionally, the temporal differences observed in our zebra mussel samples from the Hudson River likely are linked to the cycling population dynamics observed in that system, where large year classes dominate recruitment on a 2–4 year cycle, with population size fluctuating over an order of magnitude (Strayer and Malcolm 2006). We detected evidence for rapid population expansion in our 1995 Lake Superior samples, but by 2006 the population had stabilized, and no such signal remained. These signals are ephemeral (Cornuet and Luikart 1996) and would be predicted to fade after a decade, as found here. Although we did not test for temporal sampling regimes in other populations, additional evidence indicates that 10 years is sufficient for zebra mussel populations to return to equilibrium after demographic events. When Boileau and Hebert (1993) examined the newly established Oneida Lake zebra mussel population, it had much lower heterozygosity than their other North American samples. A decade later, our Oneida Lake sample has a heterozygosity level that is very similar to those characterizing Great Lakes sites, suggesting that its initial bottleneck faded with time and was augmented by new recruitment. Similarly, the invasion of New Zealand by Drosophila pseudoobscura revealed a founder effect, followed by a return to equilibrium values within two decades, although that invasive population—unlike dreissenids—retained low genetic diversity (Reiland et al. 2002).
The 100th Meridian Initiative
Western expansion populations of both dreissenid species did not originate from the closest geographic sources—which would have been either the Mississippi River basin or the southwestern Great Lakes. Instead, they appear to trace to population origins in the eastern Great Lakes. This represents a possible mixed success for the 100th Meridian Initiative (Bossenbroek et al. 2009), with our study results suggesting that few individuals colonized from the Mississippi River drainage or the western Great Lakes, where 100th Meridian control resources were concentrated (Mangin 2001). Instead, it appears that the successful colonists originated from areas much further east—in Lake Ontario and the St. Lawrence River. This sort of long-distance dispersal event is hard to predict and observe (Buchan and Padilla 1999; Leung et al. 2006), but can be detected using genetic methods, as shown in our study.
Under the 100th Meridian Initiative, more easterly population areas may not have received the program’s intensive education programs that focused on the western frontier of the zebra mussel’s distribution, rather than towards its core distribution in the Great Lakes. The effectiveness of those education programs is supported by the apparent lack of spread from targeted regions, as shown by our findings and by the slowing spread of infestations to inland lakes in areas with intense education campaigns (Johnson et al. 2006; J. Bossenbroek, personal communication, 2009). Thus, education efforts carried out to prevent the spread of dreissenid mussels likely were effective (Johnson et al. 2001) in those target areas, but should have been more comprehensive.
Conclusions
The dreissenid mussel introductions to North America show some similarities between the two species and with other Great Lakes invaders, as well as some key differences. Notably, both quagga and zebra mussels were introduced from multiple source locations in Europe. North American populations of zebra mussels originated from multiple previously-invaded locations within northern Europe and the Baltic Sea regions. In contrast, quagga mussels colonized directly from native populations originating from estuaries of the Southern Bug and Dnieper Rivers. The genetic differences of these historic colonization patterns led to population structure observed in the Great Lakes for both species, which are genetically heterogeneous across their ranges and show considerable population genetic divergence across North America. Genetic diversity levels in both species reflect no overall founder effects in comparisons of the Great Lakes populations to likely Eurasian source locations, indicating very high numbers of introduced individuals. Zebra mussel populations outside of and within the Great Lakes have similar levels of genetic diversity, whereas quagga mussel populations show reduced genetic diversity in newly established western populations. Once established, the genetic composition of given populations genetically changed over time, but retained their original genetic signatures. External recruitment and long distance gene flow contributed to population genetic diversity, which increased over time. Since the genetic signatures of these populations remain distinctive, showing closest assignment with themselves, the genotypes that became established first retained “genetic population resilience” over time, leading to differentiation among sites. These population factors thus significantly shaped the genetic identity of the dreissenid mussel invasions and provided the raw material governing their genetic adaptations.
References
Ackerman JD, Sim B, Nichols SJ, Claudi R (1994) A review of the early life history of zebra mussels (Dreissena polymorpha): comparisons with marine bivalves. Can J Zool 72:1169–1179
Aldridge DC, Elliott P, Moggridge GD (2004) The recent and rapid spread of the zebra mussel (Dreissena polymorpha) in Great Britain. Biol Conserv 119:253–261
Allen YC, Ramcharan CW (2001) Dreissena distribution in commercial waterways of the US: using failed invasions to identify limiting factors. Can J Fish Aquat Sci 58:898–907
Amsellam L, Noyer JL, Le Bourgeois T, Hossaert-McKey M (2000) Comparison of genetic diversity in the invasive weed Rubus alceifolius Poir. (Rosaceae) in its native range and in areas of introduction, using amplified fragment length polymorphism (AFLP) markers. Mol Ecol 9:443–455
Andreakis N, Kooistra WHCF, Procaccini G (2009) High genetic diversity and connectivity in the polyploidy invasive seaweed Asparagopsis taxiformis (Bonnemaisoniales) in the Mediterranean, explored with microsatellite alleles and multilocus genotypes. Mol Ecol 18:212–226
Audzijonyte A, Wittmann KJ, Vainola R (2008) Tracing recent invasions of the Ponto-Caspian mysid shrimp Hemimysis anomala across Europe and to North America with mitochondrial DNA. Divers Distrib 14:179–186
Balloux F, Lugon-Moulin N (2002) The estimation of population differentiation with microsatellite markers. Mol Ecol 11:155–165
Bax N, Williamson A, Aguero M, Gonzalez E, Geeves W (2003) Marine invasive alien species: a threat to global biodiversity. Mar Policy 27:313–323
Belkhir K, Borsa P, Chikhi L, Raufaste N, Catch F (2004) GENETIX 4.05, Population genetics software for Windows TM. Université de Montpellier II. Montpellier
Benzecri JP (1973) L’analyse des donnees. Tome II: L’analyse des correspondances. Dunod, Paris-Bruxelles-Montreal
Berg DJ, Fisher SW, Landrum PF (1996) Clearance and processing of algal particles by zebra mussels (Dreissena polymorpha). J Great Lakes Res 22:779–788
Berg DJ, Garton DW, MacIsaac HJ, Panov VE, Telesh IV (2002) Changes in the genetic structure of the North American Bythotrephes populations following invasion from Lake Ladoga, Russia. Freshw Biol 47:275–282
Bij de Vaate A, Jazdzewski K, Ketelaars HAM, Gollasch S, Van der Velde G (2002) Geographical patterns in range extension of Ponto-Caspian macroinvertebrate species in Europe. Can J Fish Aquat Sci 59:1159–1174
Blossey B, Notzold R (1995) Evolution of increased competitive ability in invasive nonindigenous plants: a hypothesis. J Ecol 83:887–889
Boileau MG, Hebert PDN (1993) Genetics of the zebra mussel (Dreissena polymorpha) in populations from the Great Lakes region and Europe. In: Nalepa TF, Schloesser DW (eds) Zebra mussels: biology, impacts, and control. Lewis Publishers, Boca Raton, pp 227–238
Bossdorf O, Auge H, Lafuma L, Rogers WE, Siemann E, Prati D (2005) Phenotypic and genetic differentiation between native and introduced plant populations. Oecologia 144:1–11
Bossenbroek JM, Johnson LE, Peters B, Lodge DM (2007) Forecasting the expansion of zebra mussels in the United States. Conserv Biol 21:800–810
Bossenbroek JM, Finnoff DC, Shogren JF, Warziniack TW (2009) Advances in ecological and economic analyses of invasive species: dreissenid mussels as a case study. In: Keller RP, Lodge DM, Lewis MA, Shogren JF (eds) Bioeconomics of invasive species: integrating ecology, economics, policy, and management. Oxford University Press, New York, pp 244–265
Britton DK, McMahon RF (2005) Analysis of trailered boat traffic and the potential westward spread of zebra mussels across the 100th Meridian. Am Malacol Bull 20:147–159
Brown AHD, Marshall DR (1981) Evolutionary changes accompanying colonization in plants. In: Scudder GEC, Reveal JL (eds) Evolution today, proceedings of the second international congress of systematic and evolutionary biology. Hunt Institute for Botanical Documentation, Carnegie-Mellon University, Pittsburgh
Brown JE, Stepien CA (2008) Ancient divisions, recent expansions: phylogeography and population genetics of the round goby Apollonia melanostoma across Eurasia. Mol Ecol 17:2598–2615
Brown JE, Stepien CA (2009) Invasion genetics of the Eurasian round goby in North America: tracing sources and spread patterns. Mol Ecol 18:64–79
Buchan LAJ, Padilla DK (1999) Estimating the probability of long-distance overland dispersal of invading aquatic species. Ecol Appl 9:254–265
Carlton JT (2008) The zebra mussel Dreissena polymorpha found in North America in 1986 and 1987. J Great Lake Res 34:770–773
Cavalli-Sforza LL, Edwards AWF (1967) Phylogenetic analysis: models and estimation procedures. Am J Hum Gen 19:526–533
Chun YJ, Nason JD, Moloney KA (2009) Comparison of quantitative and molecular genetic variation of native vs. invasive populations of purple loosestrife (Lythrum salicaria L., Lythraceae). Mol Ecol 18:3020–3035
Connelly NA, O’Neill CR, Knuth BA, Brown TL (2007) Economic impacts of zebra mussels on drinking water treatment and electric power generation facilities. Environ Manage 40:105–112
Cornuet JM, Luikart G (1996) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144:2001–2014
Cornuet JM, Piry S, Luikart G, Estoup A, Solignac M (1999) New methods employing multilocus genotypes to select or exclude populations as origins of individuals. Genetics 153:1989–2000
DeWalt SJ, Hamrick JL (2004) Genetic variation of introduced Hawaiian and native Costa Rican populations of an invasive tropical shrub, Clidemia hirta (Melastomataceae). Am J Bot 91:1155–1162
Dillon AK, Stepien CA (2001) Genetic and biogeographic relationships of the invasive round (Neogobius melanostomus) and tubenose (Proterorhinus marmoratus) gobies in the Great Lakes versus Eurasian populations. J Great Lakes Res 27:267–280
Drake JM, Lodge DM (2004) Global hot spots of biological invasions: evaluating options for ballast-water management. Proc R Soc Lond B Biol Sci 271:575–580
Elderkin CL, Klerks PL (2001) Shifts in allele and genotype frequencies in zebra mussels Dreissena polymorpha along the latitudinal gradient formed by the Mississippi River. J North Am Benthol Soc 20:595–605
Elderkin CL, Perkins EJ, Leberg PL, Klerks PL, Lance RF (2004) Amplified fragment length polymorphism (AFLP) analysis of the genetic structure of the zebra mussel Dreissena polymorpha in the Mississippi River. Freshw Biol 49:1487–1494
Ellstrand NC, Schierenbeck KA (2000) Hybridization as a stimulus for the evolution of invasiveness in plants? Proc Natl Acad Sci USA 97:7043–7050
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes—application to human mitochondrial DNA restriction data. Genetics 131:479–491
Excoffier L, Laval G, Schneider S (2005) ARLEQUIN ver. 3.0: An integrated software package for population genetics data analysis. Evol Bioinformatics Online 1:47–50. http://cmpg.unibe.ch/software/arlequin3/ v3.11
Feldheim KA, Brown JE, Murphy DJ, Stepien CA (in review) Microsatellite loci for Dreissenid Mussels (Mollusca: Bivalvia) and relatives: markers for assessing invasive and native species. Mol Ecol Res
Felsenstein J (1989) PHYLIP—phylogeny inference package (version 3.2). Cladistics 5:164–166
Ficetola GF, Bonin A, Miaud C (2008) Population genetics reveals origin and number of founders in a biological invasion. Mol Ecol 17:773–782
Floerl O, Inglis GJ, Dey K, Smith A (2009) The importance of transport hubs in stepping stone invasions. J Appl Ecol 46:37–45
Goolsby JA, Makinson JR, Hartley DM, Zonneveld R, Wright AD (2004) Pre-release evaluation and host range testing of Floracarus perrepae (Eriophyidae) genotypes for biological control of Old World climbing fern. In: Cullen JM, Briese DT, Kriticos DJ, Lonsdale WM, Morin L, Scott JK (eds) Proceedings of the XI international symposium on biological control of weeds. CSIRO Entomology, Canberra, Australia
Gosling E, Astanei I, Was A (2008) Genetic variability in Irish populations of the invasive zebra mussel Dreissena polymorpha: discordant estimates of population differentiation from allozymes and microsatellites. Freshw Biol 53:1303–1315
Goudet J, Raymond M, de Meeus T, Rousset F (1996) Testing differentiation in diploid populations. Genetics 144:1933–1940
Grapputo A, Boman S, Lindstrom L, Lyytinen A, Mappes J (2005) The voyage of an invasive species across continents: genetic diversity of North American and European Colorado potato beetle populations. Mol Ecol 14:4207–4219
Griffen BD, Drake JM (2008) A review of extinction in experimental populations. J Anim Ecol 77:1274–1287
Grigorovich IA, MacIsaac HJ, Shadrin NV, Mills EL (2002) Patterns and mechanisms of aquatic invertebrate introductions in the Ponto-Caspian region. Can J Fish Aquat Sci 59:1189–1208
Haag WR, Garton DW (1995) Variation in genotype frequencies during the life history of the bivalve, Dreissena polymorpha. Evolution 49:1284–1288
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95
Hall T (2004) BioEdit v. 7.0.0. (Available online at www.mbio.ncsu.edu/BioEdit/)
Hebert PDN, Muncaster BW, Mackie GL (1989) Ecological and genetic studies on Dreissena polymorpha (Pallas)—a new mollusk in the Great Lakes. Can J Fish Aquat Sci 46:1587–1591
Hebert CE, Weseloh DVC, Idrissi A, Arts MT, Roseman E (2009) Diets of aquatic birds reflect changes in the Lake Huron ecosystem. Aquat Ecosyst Health Manage 12:37–44
Henry P, Le Lay G, Goudet J, Guisan A, Jahodova S, Besnard G (2009) Reduced genetic diversity, increased isolation and multiple introductions of invasive giant hogweed in the western Swiss Alps. Mol Ecol 18:2819–2831
Herborg LM, Weetman D, Van Oosterhout C, Hanfling B (2007) Genetic population structure and contemporary dispersal patterns of a recent European invader, the Chinese mitten crab, Eriocheir sinensis. Mol Ecol 16:231–242
Holeck KT, Mills EL, MacIsaac HJ, Dochoda MR, Colautti RI, Ricciardi A (2004) Bridging troubled waters: biological invasions, transoceanic shipping, and the Laurentian Great Lakes. Bioscience 54:919–929
Holland BS (2001) Invasion without a bottleneck: microsatellite variation in natural and invasive populations of the brown mussel Perna perna (L). Mar Biotechnol 3:407–415
Jarvis P, Dow J, Dermott R, Bonnell R (2000) Zebra (Dreissena polymorpha) and quagga mussel (Dreissena bugensis) distribution and density in Lake Erie, 1992–1998. Canadian Technical Report of Fisheries and Aquatic Sciences 2304. Department of Fisheries and Oceans, Canada Centre for Inland Waters, Burlington, Ontario
Johnson LE, Padilla DK (1996) Geographic spread of exotic species: ecological lessons and opportunities from the invasion of the zebra mussel Dreissena polymorpha. Biol Conserv 78:23–33
Johnson LE, Ricciardi A, Carlton JT (2001) Overland dispersal of aquatic invasive species: a risk assessment of transient recreational boating. Ecol Appl 11:1789–1799
Johnson LE, Bossenbroek JM, Kraft CE (2006) Patterns and pathways in the post-establishment spread of non-indigenous aquatic species: the slowing invasion of North American inland lakes by the zebra mussel. Biol Invasions 8:475–489
Kang M, Buckley YM, Lowe AJ (2007) Testing the role of genetic factors across multiple independent invasions of the shrub Scotch broom (Cytisus scoparius). Mol Ecol 16:4662–4673
Karatayev AY, Mastitsky SE, Burlakova LE, Olenin S (2008) Past current and future of the central European corridor for aquatic invasions in Belarus. Biol Invasions 10:215–232
Kelly DW, Lamberti GA, MacIsaac HJ (2009) The Laurentian Great Lakes as a case study of biological invasion. In: Keller RP, Lodge DM, Lewis MA, Shogren JF (eds) Bioeconomics of invasive species: integrating ecology, economics, policy, and management. Oxford University Press, New York, pp 205–225
Kim KS, Bagley MJ, Coates BS, Hellmich RL, Sappington TW (2009) Spatial and temporal genetic analyses show high gene flow among European corn borer (Lepidoptera: Crambidae) populations across the Central US Corn Belt. Environ Entomol 38:1312–1323
Kliber A, Eckert CG (2005) Interaction between founder effect and selection during biological invasion in an aquatic plant. Evolution 59:1900–1913
Kolbe JJ, Glor RE, Schettino LRG, Lara AC, Larson A, Losos JB (2004) Genetic variation increases during biological invasion by a Cuban lizard. Nature 431:177–181
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Leung B, Bossenbroek JM, Lodge DM (2006) Boats pathways and aquatic biological invasions: estimating dispersal potential with gravity models. Biol Invasions 8:241–254
Lewis KM, Feder JL, Lamberti GA (2000) Population genetics of the zebra mussel Dreissena polymorpha (Pallas): local allozyme differentiation within midwestern lakes and streams. Can J Fish Aquat Sci 57:637–643
Luikart G, Allendorf FW, Cornuet J-M, Sherwin WB (1998a) Distortion of allele frequency distributions provides a test for recent population bottlenecks. J Hered 89:238–247
Luikart G, Sherwin WB, Steele BM, Allendorf FW (1998b) Usefulness of molecular markers for detecting population bottlenecks via monitoring genetic change. Mol Ecol 7:963–974
Mackie GL, Schloesser DW (1996) Comparative biology of zebra mussels in Europe and North America: an overview. Am Zool 36:244–258
Mangin S (2001) The 100th Meridian Initiative: a strategic approach to prevent the westward spread of zebra mussels and other aquatic nuisance species. U.S. Fish and Wildlife Service
Maron JL, Vila M, Bommarco R, Elmendorf S, Beardsley P (2004) Rapid evolution of an invasive plant. Ecol Monograph 74:261–280
Marsden JE, Spidle A, May B (1995) Genetic similarity among zebra mussel populations within North America and Europe. Can J Fish Aquat Sci 52:836–847
May B, Marsden JE (1992) Genetic identification and implications of another invasive species of dreissenid mussel in the Great Lakes. Can J Fish Aquat Sci 49:1501–1506
Merila J, Crnokrak P (2001) Comparison of genetic differentiation at marker loci and quantitative traits. J Evol Biol 14:892–903
Merritt TJS, Shi L, Chase MC, Rex MA, Etter RJ, Quattro JM (1998) Universal cytochrome b primers facilitate intraspecific studies in molluscan taxa. Mol Mar Biol Biotechnol 7:7–11
Mills EL, Rosenberg G, Spidle AP, Ludyanskiy M, Pligin Y, May B (1996) A review of the biology and ecology of the quagga mussel (Dreissena bugensis) a second species of freshwater dreissenid introduced to North America. Am Zool 36:271–286
Mills EL, Chrisman JR, Baldwin B, Owens RW, O’Gorman R, Howell T, Roseman EF, Raths MK (1999) Changes in the dreissenid community in the lower Great Lakes with emphasis on southern Lake Ontario. J Great Lakes Res 25:187–197
Morton B (1993) The anatomy of Dreissena polymorpha and the evolution and success of the heteromyarian form in the Dreissenoidea. In: Nalepa TF, Schloesser DW (eds) Zebra mussels: biology, impacts, and control. Lewis Publishers, Boca Raton, pp 185–215
Naish KA, Boulding EG (2001) Trinucleotide microsatellite loci for the zebra mussel Dreissena polymorpha, an invasive species in Europe and North America. Mol Ecol Notes 1:286–288
Nalepa TF, Schloesser DW (eds) (1993) Zebra mussels: biology, impacts, and control. Lewis/CRC Press, Inc., Boca Raton
Nalepa TF, Fanslow DL, Lang GA (2009) Transformation of the offshore benthic community in Lake Michigan: recent shift from the native amphipod Diporeia spp. to the invasive mussel Dreissena rostriformis bugensis. Freshw Biol 54:466–479
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York
Nei M, Maruyama T, Chakraborty R (1975) The bottleneck effect and genetic variability in populations. Evolution 29:1–10
Neilson ME, Stepien CA (2009) Escape from the Ponto-Caspian: evolution and biogeography of the neogobiin species flock (Gobiidae: Teleostei). Mol Phys Evol 52:84–102
Perez JE, Afonsi C, Nirchio M, Salazar SK (2008) Bioinvaders: the acquisition of new genetic variation. Interciencia 33:935–940
Piry S, Luikart G, Cornuet J-M (1999) BOTTLENECK: a computer program for detecting recent reductions in the effective population size using allele frequency data. J Hered 90:502–503
Piry S, Alapetite A, Cornuet J-M, Paetkau D, Baudouin L, Estoup A (2004) GENECLASS2: a software for genetic assignment and first-generation migrant detection. J Hered 95:536–539. http://www1.montpellier.inra.fr/URLB/
Pligin YV (1979) Areal extension of the area of Dreissena bugensis Andr. In: Sixth meeting on the investigation of molluscs. Molluscs. Main results of their study. Abstract of communications. Nauka, Leningrad, pp 222–224
Pritchard JK, Wen W (2004) Documentation for STRUCTURE software: Vers. 2. Department of Human Genetics, University of Chicago, Chicago, IL. http://pritch.basd.uchicago.edu
Pritchard JK, Stephens M, Donelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Rannala B, Mountain JL (1997) Detecting immigration by using multilocus genotypes. Proc Natl Acad Sci USA 94:9197–9201
Raymond M, Rousset F (1995) An exact test for population differentiation. Evolution 49:1280–1283
Reed DH, Frankham R (2001) How closely correlated are molecular and quantitative measures of genetic variation? A meta-analysis. Evolution 55:1095–1103
Reiland J, Hodge S, Noor MAF (2002) Strong founder effect in Drosophila pseudoobscura colonizing New Zealand from North America. J Hered 93:415–420
Ricciardi A (2006) Patterns of invasion of the Laurentian Great Lakes in relation to changes in vector activity. Divers Distrib 12:425–433
Ricciardi A, MacIsaac HJ (2000) Recent mass invasion of the North American Great Lakes by Ponto-Caspian species. Trends Ecol Evol 15:62–65
Ricciardi A, Whoriskey FG (2004) Exotic species replacement: shifting dominance of dreissenid mussels in the Soulanges Canal, upper St. Lawrence River, Canada. J North Am Benth Soc 23:507–514
Ricciardi A, Serrouya R, Whoriskey FG (1995) Aerial exposure tolerance of zebra and quagga mussels (Bivalvia, Dreissenidae)—implications for overland dispersal. Can J Fish Aquat Sci 52:470–477
Rice WR (1989) Analyzing tables of statistical tests. Evolution 43:223–225
Rosenthal DM, Ramakrishnan AP, Cruzan MB (2008) Evidence for multiple sources of invasion and intraspecific hybridization in Brachypodium sylvaticum (Hudson) Beauv. in North America. Mol Ecol 17:4657–4669
Rousset F (2008) GENEPOP007: a complete re-implementation of the GENEPOP software for Windows and Linux. Mol Ecol Resour 8:103–106
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sokal RR, Rohlf FJ (1995) Biometry, 3rd edn. W. H. Freeman, New York
Stepien CA, Tumeo MA (2006) Invasion genetics of Ponto-Caspian gobies in the Great Lakes: a “cryptic” species, absence of founder effects, and comparative risk analysis. Biol Invasions 8:61–78
Stepien CA, Dillon AK, Chandler MD (1998) Genetic identity, phylogeography, and systematics of ruffe Gymnocephalus in the North American Great Lakes and Eurasia. J Great Lakes Res 24:361–378
Stepien CA, Hubers AN, Skidmore JL (1999) Diagnostic genetic markers and evolutionary relationships among invasive dreissenoid and corbiculoid bivalves in North America: phylogenetic signal from mitochondrial 16S rDNA. Mol Phys Evol 13:31–49
Stepien CA, Morton B, Dabrowska KA, Guarnera RA, Radja T, Radja B (2001) Genetic diversity and evolutionary relationships of the troglodytic ‘living fossil’ Congeria kusceri (Bivalvia: Dreissenidae). Mol Ecol 10:1873–1879
Stepien CA, Taylor CD, Dabrowska KA (2002) Genetic variability and phylogeographic patterns of a nonindigenous species invasion: a comparison of exotic versus native zebra and quagga mussel populations. J Evol Biol 15:314–328
Stepien CA, Taylor CD, Grigorovich IA, Shirman SV, Wei R, Korniushin AV, Dabrowska KA (2003) DNA and systematic analysis of invasive and native dreissenid mussels: is Dreissena bugensis really D. rostriformis? Aquat Invaders 14:8–18
Stepien CA, Ford AM, Dillon-Klika AK, Tumeo MA (2004) Risk analysis and genetic identity of the Eurasian source population for the ruffe (Gymnocephalus cernuus) invasion in the Great Lakes. In: Barry TP, Malison JA (eds) Proceedings of Percis III, the 3rd International Symposium on Percid Fishes, University of Wisconsin Sea Grant Institute, Madison, WI. 91–92
Stepien CA, Brown JE, Neilson ME, Tumeo MA (2005) Genetic diversity of invasive species in the Great Lakes versus their Eurasian source populations: insights for risk analysis. Risk Anal 25:1043–1060
Stewart TJ, Sprules WG, O’Gorman R (2009) Shifts in the diet of Lake Ontario alewife in response to ecosystem change. J Great Lakes Res 35:241–249
Stokstad E (2007) Feared quagga mussel turns up in western United States. Science 315:453
Strayer DL, Malcolm HM (2006) Long-term demography of a zebra mussel (Dreissena polymorpha) population. Freshw Biol 51:117–130
Tepolt CK, Blum MJ, Lee VA, Hanson ED (2007) Genetic analysis of the Chinese mitten crab (Eriocher sinensis) introduced to the North American Great Lakes and St. Lawrence Seaway. J Great Lakes Res 33:658–667
Therriault TW, Orlova MI, Docker MF, MacIsaac HJ, Heath DD (2005) Invasion genetics of a freshwater mussel (Dreissena rostriformis bugensis) in eastern Europe: high gene flow and multiple introductions. Heredity 95:16–23
van Oosterhout C, Hutchison WF, Wills DPM, Shipley P (2004) MICROCHECKER: software for identifying and correcting genotyping errors in microsatellite data. Mol Ecol Notes 4:535–538
van Oosterhout C, Hutchison WF, Wills DPM, Shipley P (2006) MICROCHECKER. www.microchecker.hull.ac.uk
Vanderploeg HA, Nalepa TF, Jude DJ, Mills EL, Holeck KT, Liebig JR, Grigorovich IA, Ojaveer H (2002) Dispersal and emerging ecological impacts of Ponto-Caspian species in the Laurentian Great Lakes. Can J Fish Aquat Sci 59:1209–1228
Viard F, Ellien C, Dupont L (2006) Dispersal ability and invasion success of Crepidula fornicata in a single gulf: insights from genetic markers and larval-dispersal model. Helgol Mar Res 60:144–152
Weir BS, Cockerham CC (1984) Estimating F-statistics for the analysis of population structure. Evolution 38:1358–1370
Whitney KD, Gabler CA (2008) Rapid evolution in introduced species, ‘invasive traits’ and recipient communities: challenges for predicting invasive potential. Divers Distrib 14:569–580
Williams DA, Overholt WA, Cuda JP, Hughes CR (2005) Chloroplast and microsatellite DNA diversities reveal the introduction history of Brazilian peppertree (Schinus terebinthifolius) in Florida. Mol Ecol 14:3643–3656
Wilson AB, Boulding EG, Naish KA (1999) Characterization of tri- and tetranucleotide microsatellite loci in the invasive mollusk Dreissena bugensis. Mol Ecol 8:692–693
Wilson KA, Howell ET, Jackson DA (2006) Replacement of zebra mussels by quagga mussels in the Canadian nearshore of Lake Ontario: the importance of substrate, round goby abundance, and upwelling frequency. J Great Lakes Res 32:11–28
Zenger KR, Richardson BJ, Vachot-Griffin AM (2003) A rapid population expansion retains genetic diversity within European rabbits in Australia. Mol Ecol 12:789–794
Zhu B, Fitzgerald DG, Mayer CM, Rudstam LG, Mills EL (2006) Alteration of ecosystem function by zebra mussels in Oneida Lake: impacts on submerged macrophytes. Ecosystems 9:1017–1028
Acknowledgments
This project was funded by Ohio Sea Grant R/LR-009-PD and University of Toledo start-up funds to CAS. JEB was supported in 2009 by an Ohio Sea Grant NOAA Knauss fellowship, and by previous teaching and research fellowships from the Department of Environmental Sciences of the University of Toledo. Samples generously were provided by W. Baldwin, C. Balogh, B. Bodamer, C. Bowen, D. Clapp, N. Claudi, T. Crail, R. Dermott, K. DeVanna, D. Garton, J. Goeckler, I. Grigorovich, J. Gunderson, J. Hageman, S. Hogan, K. Holeck, H. Jenner, D. Jensen, T. Johnson, T. Kakareko, Y. Kvach, E. Laney, Lee, E. Marsden, B. May, C. Mayer, B. Moore, C. Munté i Carrollo, M. Neilson, T. Nalepa, D. Pavlov, J. Postberg, A. Ricciardi, G. Rosenberg, J. Ross, M. Sapota, J. Schooley, B. Sket, A. Spidle, K. Stewart, D. Strayer, M. Thomas, L. Voronina, and M. Watson. We thank P. Uzmann of the Lake Erie Center for administrative assistance and J. Banda, J. Bossenbroek, L. Corkum, A. Haponski, R. Lohner, C. Mayer, J. Miner, D. Murphy, M. Neilsen, L. Pierce, and O.J. Sepulveda Villet for valuable comments on the manuscript. This is publication 2010-004 from the Lake Erie Research Center.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Brown, J.E., Stepien, C.A. Population genetic history of the dreissenid mussel invasions: expansion patterns across North America. Biol Invasions 12, 3687–3710 (2010). https://doi.org/10.1007/s10530-010-9763-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-010-9763-2