Abstract
The disease-resistant transgenic peanut cv ICG 13942 plants were developed by using Tcchitinase-I gene. Agrobacterium tumefaciens strain LBA4404 harboring the binary vector (pBinAR) contains the chitinase (Tcchitinase-I) gene and neomycin phosphotransferase resistance (nptII) gene. The transformed shoots were developed on selection medium (MMS + 0.5 mg/L IAA + 15 mg/L TDZ + 100 mg/L Kan + 250 mg/L Cefotaxime) from deembryonated cotyledon (DC) explants. Established plantlets were screened for the presence of Tcchitinase-I and nptII genes. Stable integration and expression of the transgenes (T0) were confirmed by using PCR, RT-PCR and Southern blot analyses. The transformation frequency 63.34% was recorded. All the transformed (T0) plants were found normal, flowered and set seeds. After selfing the T0 plants, a Mendelian inheritance pattern (3:1) for the transgene in T1 progeny is revealed. T1 transgenic peanut plants were evaluated for resistance against Cercospora arachidicola, C. personatum and Puccinia arachidis by infection with the microspores using detached leaf assay. These T1 plants have shown longer incubation, latent period and lower infection frequencies in comparison to non-transformed (WT) plants. The Tcchitinase-I gene expression in resistant transgenic plants was compared to that of a susceptible control. A significant negative correlation was recorded between chitinase activity and the frequency of infection to the three tested disease causing agents.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Peanut or groundnut (Arachis hypogaea L.) is an important oilseed crop grown in the tropical and sub-tropical regions of the world. Due to various biotic stresses such as insect pests, bacterial and fungal diseases, the yield in peanut is decreased. Among the major fungal diseases, early leaf spot (ELS) (Cercospora arachidicola), late leaf spot (LLS) (Cercospora personatum) and rust caused by Puccinia arachidis are more destructive (50–70%) in peanut (Subrahmanyam et al. 1984). Chitin is an important cell wall component of fungi and this is degraded by chitinases. Chitinases (E.C. 3.2.1.14) are poly (1,4-(N-acetyl-β-D)glucosaminide))-glycanohydrolases. They directly hydrolyze fungal cell wall, chitin the substrate for the enzyme, and by this action fungal hyphallysis and inhibition of fungal growth occur (Patil et al. 2000). Thus, plants use one of the many natural defense mechanisms to resist against pathogens and accumulate proteins (e.g., chitinases) active against disease causing organisms. Where this mechanism is too weak or appears too late to induce full protection against pathogen, engineering the expression of a defense protein can enhance the resistance to fungal diseases (Broglie et al. 1991; Grison et al. 1996).
The use of fungicide to control the disease is often ineffective because the pathogen spreads rapidly under favorable conditions. The crop production heavily relies on chemicals for protection which is not viable as these chemicals provide ephemeral benefits often with adverse side effects (Kumar et al. 2008). The major destructive fungi, on the other hand, develop tolerance to most classes of fungicides and these can cause environmental pollution (Moham et al. 2003). In view of this, genetic engineering with chitinase is a powerful tool to improve the fungal disease resistance in plants.
Genetic engineering technology plays a great role in transfer of gene(s) of interest for developing disease resistance and improving quality and crop yield. By using this technology, the chitinase gene from different origins has been introduced into various crop plants for developing enhanced fungal resistance: tobacco (Zhu et al. 1994), rice (Lin et al. 1995), cucumber (Kishimoto et al. 2002), Italian ryegrass (Takahashi et al. 2005), cotton (Ganesan et al. 2009), banana (Sreeramanan et al. 2009) and peanut (Chu et al. 2008, 2013).
Though enhanced fungal resistance has been developed in peanut by using tobacco chitinase (Rohini and Rao 2001), barley oxalate oxidase (Livingstone et al. 2005), mustard defensin (Anuradha et al. 2008) and tobacco β-1,3-glucanase (Sundaresha et al. 2009), there is no report on Tcchitinase-I gene in peanut cvs. Hence, in the present study, we have developed the transgenic peanut cv ICG 13942 plants by using deembryonated cotyledon (DC) explants through Agrobacterium-mediated genetic transformation by using Tcchitinase-I gene for resistance to leaf spot (ELS, LLS) and rust diseases.
Materials and methods
Plant material
The mature seeds of peanut cv ICG 13942, obtained from the germplasm bank of ICRISAT, Patancheru, Hyderabad, Telangana, India were used. This variety is susceptible to leaf spot and rust fungal diseases.
Mature groundnut seeds of peanut cv ICG 13942 were washed under running tap water for 10–15 min followed by treating with liquid detergent Tween-20 (5%-v/v) for 5 min and it was repeated twice followed by rinsing in sterile distilled water thoroughly. Later the seeds were surface sterilized with 0.1% (w/v) HgCl2 for 8 min followed by rinsing in sterilized distilled water for 3–4 times under aseptic conditions and soaked for 24 h in sterile distilled water. These soaked seeds were dried on sterile tissue paper, dissected aseptically and removed the zygotic embryo. Now, these deembryonated cotyledons (DC) were cut longitudinally and precultured on shoot induction medium (SIM) containing Modified Murashige and Skoog’s (1962) (MMS) medium + 0.5 mg/L IAA + 15 mg/L TDZ for 3 days.
Gene constructs
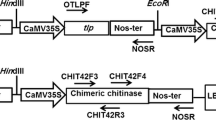
Agrobacterium tumefaciens strain LBA4404 harboring the binary plasmid pBinAR (13.7 Kb) was used for genetic transformation of groundnut cv ICG 13942. The binary vector pBinAR carrying Theobroma cacao chitinase-I (Tcchitinase-I) gene with a nptII selectable marker gene was used. The T-DNA portion of pBinAR having nos-nptII cassette in RB and 770 bp EcoRI/Hind III fragment contains the CaMV 35S promoter, a partial pUC18 polylinker and the OCS terminator in LB and selectable marker gene (nptII) driven by the NOS promoter and PNOS terminator sequences, respectively. A 1.2Kb SmaI-XbaI fragement of T. cacao class I chitnase was taken out from pGH00.0126 vector and cloned intocorresponding sites of binary vector pBinAR (Fig. 1).
Transformation procedure
The precultured DC (deembryonated cotyledon) explants were infected with A. tumefaciens LBA 4404 harboring binary vector pBinAR containing Tcchitinase-I gene and nptII as selectable marker gene and cocultivated on SIM (shoot induction medium) containing MMS (modified MS medium) + 0.5 mg/L IAA + 15 mg/L TDZ for 4 days. After cocultivation, these explants were shifted onto selection medium containing SIM + 100 mg/L Kan + 250 mg/L Cefotaxime. After 2 weeks of incubation, the explants with KanR shoots were cultured on SIM + 50 mg/L Kan for further proliferation of microshoots. Subsequently, the shoots were elongated, rooted and established the plantlets by following our earlier study (Rajinikanth and Rama Swamy 2018). The putative transformants (T0) were obtained within 4 months of culture initiation (Fig. 2a–e). The plants were regenerated from non-transformed explants and established in the greenhouse as control. T0 transgenic plants were maintained in the greenhouse and seeds were harvested to obtain the T1, T2 generations. The transgenic plants in T0, T1and T2 generations were analyzed using standard procedures.
Agrobacterium-mediated genetic transformation in deembryonated cotyledon (DC) explants of peanut cv ICG 13942 by using binary vector pBinAR. a Infected DC explants on SIM for cocultivation. b, c Induction of KanR shoots on selection medium after 4 & 6 weeks of incubation, respectively. d In vitro rooting of elongated microshoots on RIM augmented with 1.0 mg/L NAA + 50 mg/L Kan. e T0 plants are shifted to plastic pots containing soil mix and maintained in the green house
PCR and RT-PCR analysis of the transformants
The genomic DNA was isolated from randomly selected putative transgenic plants and one non-transformed plant (control) according earlier method (Sharma et al. 2000) and subjected to PCR amplification using the Tcchitinase-I gene-specific primers:(F)5′-GGAAAATGGTTGCCAGAGTCAGTGC-3′, (R)5′-GCTACATTGAGTCCACCGAGGGT-3′ and nptII gene-specific primers: (F) 5′-GCTTGGGTGGAGAGGGCTATT-3′, (R) 5′-AGAACTCGTCAAGAA GGCGA-3′. The PCR analysis for Tcchitinase-I gene was carried out by initial denaturation at 94 °C for 5 min followed by 35 cycles of 94 °C for 1 min, 58 °C for 1.30 min and 72 °C for 2 min and final extension at 72 °C for 10 min and nptII gene was carried out by initial denaturation at 94 °C for 5 min followed by 35 cycles of 94 °C for 1 min, 54 °C for 1 min and 72 °C for 1.30 min and final extension at 72 °C for 10 min. The amplified products were subjected to electrophoresis on 1.2% agarose gel and visualized under gel documentation system, Biorad, USA (Fig. 3a, b). The randomly selected PCR-positive transgenic plants were used for RT-PCR analysis. Total RNA was isolated from leaf tissue of the putative transformants using the TRIzol reagent according to the manufacture’s protocol and RT-PCR analysis of the putative transformants was carried out using the Thermoscript RT-PCR system for 35–40 cycles using Tcchitinase-I gene-specific primers for carrying out RT-PCR. One sample of RNA subjected directly to PCR without reverse transcription served as the negative control and plasmid DNA from pBinAR-chitinase-I served as the positive control. The amplified fragments were separated on 1.2% agarose gel, photographed under ultraviolet light (Fig. 3c).
Molecular analysis of the transformed peanut cv ICG 13942 plants. a PCR amplification of genomic DNA showing amplification of a 1200 bp fragment of the Tcchitinase-I gene, Lanes: 1–12: carry genomic DNA from T0 putative transformants. b PCR amplification of genomic DNA showing amplification of a 750 bp fragment of the nptII gene, Lanes: 1–5: carry genomic DNA from T0 putative transformants. c RT-PCR of the cDNA showing amplification of a 1200 bp fragment of the Tcchitinase-I gene, Lanes: 1–6: carry cDNA from T0 putative transformants. M: molecular size marker (1 Kb ladder), B: blank, C: non-transformed control plant DNA (-ve control-WT), P: plasmid pBinAR (+ ve control)
Southern blot analysis
30 µg of genomic DNA from the putatively transformed and non-transformed (control) plants was digested with the enzyme EcoRI to restrict the genomic DNA which cuts at restriction site within the plasmid DNA to determine the copy number of the Tcchitinase-I gene. The digested DNA was separated by electrophoresis through a 1% agarose gel and transferred onto a Nylon N + membrane (Amersham Biosciences, UK) according to the manufacturer’s instructions. The 1.2 kb Tcchitinase-I coding sequence fragment with a non-radioactively labeled (Alkphos Direct Labeling and Detection system of Amersham Biosciences) was used as a probe (Fig. 4).
Southern blot analysis of the genomic DNA from leaves of transgenics obtained through Agrobacterium-mediated genetic transformation. The genomic DNA of peanut transgenics was digested with EcoR1 to check the copy number of the integrated gene. Lanes: 1: carry EcoRI-restricted genomic DNA from event TC-20 showed two copy numbers, 2: carry EcoRI-restricted genomic DNA from event TC-9 showed one copy number, 3–7: carry EcoRI-restricted genomic DNA from events TC-1, TC-7, TC-12, TC-14, TC-18 showed three copy numbers, respectively, C: EcoRI-restricted genomic DNA from control plants, P: EcoRI-restricted plasmid pBinAR: Tcchitinase-I, M: molecular weight marker
Segregation analysis
Inheritance of the transgene was studied by using the PCR screening of Tcchitinase-I gene in T1 and T2 generations. PCR + ve and –ve plants were identified and chi-square test was performed to validate the data for 3:1 segregation.
Chitinase assay
A colorimetric assay was performed with the leaves of 45-day-old transformed and non-transformed (WT) control peanut plants following the method of Mauch et al. (1984).
Detached leaf assay for ELS, LLS and rust diseases
Disease evaluation of the transgenic peanut plants for ELS, LLS and rust pathogens was conducted in T1 generation plants by detached leaf assay technique as reported earlier.
When the plants (T1) were 40 days old, the quadrifoliate leaves from either second or third fully expanded leaf of 10 selected PCR + ve transgenic plants per event were excised from pulvinus region and arranged in randomized block design in plastic trays containing sterile river sand. The leaves were immediately dipped in distilled water and were taken for planting in plastic trays having dimensions 39.5 cm × 29 cm × 7 cm. The trays were filled with a layer (approximately 1.5 cm thick) of sterile sand. The sand was kept moist with distilled water. Holes were made with the help of plastic droppers to place the leaves. In each hole, a leaf was planted and the lower portion of the rachis was covered with sand. Simultaneously, ten leaves of susceptible control plants were also planted in a separate tray. The Hoagland’s nutrition solution (Hoagland and Arnon 1950) was supplied to leaves, which provided the essential nutrients. The trays were covered with clear polythene cover and incubated before inoculation for 24 h in a growth chamber for acclimatization. The day and night temperature in the growth chamber was maintained at 23 °C with relative humidity of 60% and illuminated with white light. The leaves were allowed to get acclimatized for 24 h after transferring to plastic trays filled with sterile sand and then challenged with ELS, LLS and rust disease pathogens separately by using the method of Prasad et al. (2013).
The experimental trays were examined daily beginning 6 days after inoculation (DAI) for incubation period (IP) and latent period (LP). After 28 days of inoculation, observations on lesions or pustules per leaf (LN, PN), percentage leaf area damaged (LAD%) and infection frequency (IF, defined as number of lesions or pustules/cm2 leaf area) were recorded.
Data analysis
The data collected on chitinase activity, ELS, LLS and rust infection were subjected to analysis of variance (ANOVA) where the mean values in each treatment was compared using LSD at the 5% level of significance (P = 0.05). The values were means of ten replicates per event. The correlation analysis was done using Pearson correlation coefficient at 5% level of significance among the transgenics and non-transformed control plants for infection frequency of three tested pathogens with chitinase activity.
Results
Genetic transformation of peanut with Tcchitinase-I gene
Agrobacterium tumefaciens-mediated genetic transformation was carried out by using the binary vector pBinAR-Tcchitinase-I with DC explants of groundnut cv ICG 13942. A total of 38 primary transformants (T0) were regenerated from 60 precultured DC explants following cocultivation with A. tumefaciens harboring the binary vector pBinAR-Tcchitinase-I with 63.34% of transformation efficiency. The regenerated plants (KanR) exhibited normal growth under greenhouse conditions and produced morphologically normal flowers and pods that contained viable seeds (Fig. 2a–e).
Molecular analysis of transgenics
Integration and expression of transgenes
The insertion of the Tcchitinase-I gene into the peanut through Agrobacterium gene transformation was initially screened by PCR analysis. The presence of 1200 bp region of the Tcchitinase-I gene was detected in 12 of the 38 transgenic plants produced with the binary plasmid pBinAR:Tcchitinase-I with the transformation efficiency is 63.34% (Fig. 3a). Randomly selected transformants of 5 of the 38 transgenic plants also showed amplification of 750 bp fragment of the nptII gene (Fig. 3b). Expression of the introduced gene was analyzed by RT-PCR from the randomly selected 6 T1 and T2 PCR-positive plants. The expected 1200 bp amplified fragment corresponding to the Tcchitinase-I gene was detected in all the plants that were selected for analysis (Fig. 3c). Randomly selected PCR- and RT-PCR-positive events were analyzed by Southern blot hybridization for copy number (EcoRI digested DNA) using 1.2 Kb fragment as probe (Fig. 4). The Southern analysis indicated the presence of three copies of the transgene in event number TC-1, TC-7, TC-12, TC-14, TC-18 (Lanes 3–7), while the event TC-20 (Lane 1) showed two copies of the transgene and event TC-9 (Lane 2) showed single copy of the transgene, whereas no transgene insertion was detected in non-transformed control plant DNA (Lane C). The segregation pattern of PCR tested transgenic plants and their progeny showed the Mendelian ratio (3:1 ratio) at p = 0.05 in all the events in T1 and T2 progenies.
Chitinase activity in the transgenic plants
The chitinase activity varied among the transgenic events expressing Tcchitinase-I gene, where 6.5-fold increase in the chitinase activity (0.29–1.30 Umg−1 protein) was recorded as compared to the non-transformed control plants (0.20 Umg−1 protein) (Fig. 5). Of the 20 T1 transgenic events tested, 6 transgenic events (TC-1, TC-6, TC-14, TC-15 and TC-18) had significantly higher chitinase activity than the rest, which sustained in the T2 progeny of four of these six events (TC-1-7, TC-6-7, TC-12-4 and TC-18-6).
Evaluation of peanut transgenics for resistance against early leaf spots (ELS), late leaf spot (LLS) and rust diseases
The progenies of twenty T1 transgenic events were tested and they showed significant differences for all the components of resistance to ELS, LLS and rust diseases in detached leaf bioassay (Table 1).
For evaluation of ELS, the event TC-18 and TC-06 showed longer incubation period (18 and 15 days), longer latent period (21 and 20 days) and less number of lesions per leaf (27 and 30) in T1 transformed plants compared to non-transformed (control) plants (10 days IP, 16 days LP and 40 lesions). Most of the transgenic events (TC-1, TC-06, TC-07, TC-09 and TC-18) showed less LAD (5.62–11.18%), less number of lesions per leaf (27–34) and less IF (2.13–4.63 cm2) compared to non-transformed counter parts (LAD 14.32%, lesions 40, IF 4.84 cm2). According to our observations, the event TC-18 showed better performance in all the recorded resistance parameters for ELS disease. They are longer IP (18 days), longer LP (21 days), lesser number of lesions per leaf (27), less LAD (5.62%) and lower IF (2.13 cm2) in comparison to other events in T1 plants (Table 1).
T1 transgenic events evaluated for LLS showed significant genotypic difference for all the components. Most of the transgenic events 01,06,10,12 and 18 showed longer IP (17 to 23 days) and the events 9,10,12,14 and 18 showed longer LP (21 to 24 days) than the non-transformed plants (13 days IP, 19 days LP). The event no. 14 showed lesser number of lesions per leaf (22 lesions), less leaf area damage (6.20%) and lower IF (2.50 cm2) in comparison to the control plants (35 lesions, 18.10% LAD, 5.43 cm2 IF). Thus, the event TC-14 was found to be performed better in all the parameters (longer IP-21 days, longer LP-26 days, lesser number of lesions per leaf-22, less LAD-6.20% and lower IF-2.50 cm2) tested compared to other events screened for LLS disease resistance (Table 1).
For evaluation of rust, the event no. 7 showed longer incubation (23 days), longer latent periods (31 days) and less no. of lesions (22 lesions) than their control plants (15 days IP, 21 days LP and 31 lesions). All transgenic events except event 10, 12 showed less LAD (3.23–11.10%) than the control plants (12.08%). Similarly, most of the transgenic events showed lower Infection frequencies (3.68 to 7.84 cm2) except transgenic event TC-9 (9.15 cm2) than the control plants (8.16 cm2). Thus, the event TC-7 showed the best results for all the resistance parameters tested for rust disease even compared to all other T1 transgenic plants (Table 1).
According to our observation, the three transgenic events of TC-18, TC-14 and TC-7 displayed significantly higher resistance to C. arachidicola, C. personatum and P. arachidis pathogens in T1 plants, respectively (Fig. 6a–d).
Correlation between chitinase activity and disease resistance
Disease severity correlated well with the chitinase activity and the infection frequency of ELS, LLS and rust in the T1 transgenic plants with the Pearson correlation coefficients ranging from − 0.7226 (P = 0.05), − 0.8036 (P = 0.05) and − 0.8475 (P = 0.05), respectively. These results indicated that the transgenic events with high chitinase activity showed lower disease incidence and vice versa.
Discussion
Fungal diseases constitute a major challenge to the millions of peanut growing farmers throughout the tropical regions. A large proportion of the potential peanut crop is lost yearly to several major stresses despite the efforts at transgenics for resistance. Analysis of transgenic plants provides a powerful tool for functional studies of defense genes in peanut.
As there are no reports on Tcchitinase-I gene expression in peanut, we report the transgenic peanut events expressing the Tcchitinase-I gene and also evaluated for their tolerance to leaf spot (ELS, LLS) and rust diseases.
The T. cacao chitinase gene used in the present study is a class I chitinase and belongs to PR3 family, having high chitinase activity due to the presence of chitin binding domain (CBD) (Sela-Buurlage et al. 1993). Deletion of chitin binding domain (ChBDTob) from tobacco class I chitinase has been reported to cause a threefold reduction of activation energy and antifungal activity due to lack of its binding capacity to chitin. While a CBD is not required for chitinolytic or antifungal activities, it increases both, perhaps by anchoring to the substrate and increasing its effective concentration for hydrolysis (Iseli et al. 1993). Interestingly, all other classes of chitinase have either no or lower antifungal activity as compared to class I chitinases (Sela-Buurlage et al. 1993).
Engineering for disease resistance in legumes has been considered important in the recent years. However, very little progress is seen in the improvement of legumes through the transgenic approach and more so with peanut because of the recalcitrancy in regeneration of the crop. There are reports of regenerability and Agrobacterium-mediated genetic transformation efficiency in the peanut, but less frequency of transformation 55%, 31% in JL 24 (Sharma and Anjaiah 2000; Anuradha et al. 2006), 55% in ICGV 86031 (Prasad et al. 2013), 40% in ICGV 89104 (Prasad et al. 2013) and 34% in K6 (Mehta et al. 2013). Survival rate of the in vitro regenerated plantlets was over 75% in cv. Golden and 49% in cv. Bari-2000, while healthy putatively transgenic (T0) plants with over 41% transformation frequency in cv. Golden and 32% in cv. Bari-2000 were produced through Agrobacterium-mediated gene transfer of the rice chitinase gene and all the plants flowered and set seed normally (Iqbal et al. 2012). Whereas in the present investigations, we have developed the transgenic T0 peanut plants with high transformation efficiency of 63.63% when compared to earlier reports.
The integration of the transgene was confirmed by PCR and Southern blot analyses. Segregation studies showed Mendelian ratio of 3:1 of the Tcchitinase-I gene in T1 and T2 generation transgenic peanut plants. Similarly, Anuradha et al. (2006) have reported that the inheritance of a promoter less gus:nptII bifunctional fusion gene in groundnut through Chi-square analysis showed that the segregation of fusion gene followed the Mendelian 3:1 ratio. Tiwari et al. (2008) obtained 3:1 segregation ratio for cry1 EC gene in transgenic groundnut as we have observed in the present study. Thus, the inheritance and stable expression of transgenes is important in crop improvement through gene manipulations. In the present study, the expression of the transgenes was also confirmed by RT-PCR.
In the present study, 6.5-fold increase in the chitinase activity (0.29–1.30 Umg−1 protein) was recorded in transgenic plants as compared to 0.20 Umg−1 protein in the non-transformed control plants. The enhanced chitinase activity in the transgenic plants compared to their non-transformed controls confirmed the expression of Tcchitinase-I. Several reports on intensified chitinase activity have been observed in the transgenic plants expressing the other type of chitinase genes (Lin et al. 1995; Datta et al. 2000; Nandakumar et al. 2007). Over 14-fold increase in chitinase activity over controls was reported in the leaves of peanut transformants (Prasad et al. 2013), while in transgenic rice it has been reported up to 14 times (Lin et al. 1995; Nandakumar et al. 2007). A 5-fold increase in chitinase activity in transgenic peanut plants transformed with rice chitinase gene was observed (Iqbalet al. 2012).
Although in our study some of the transgenic events showed increased Tc-chitinase-I activity, these differed in their level of resistance to ELS, LLS and rust diseases. This variation may be explained by differences in the biochemical composition and structure of the fungal cell wall, tissue and cellular localization of the recombinant chitinase, concordance in chitinase expression kinetics and the period of infection, and the type of interaction between the plant and the pathogen (Grison et al. 1996; Datta et al. 2001; Pasonen et al. 2004). Nevertheless, most of the transgenic plants showed reduced infection than their non-transformed control plants confirming the antimicrobial property of the expressed Tcchitinase-I against these pathogens.
In the present study, correlation analysis showed a significant trend towards decreased disease severity in the transgenics with the increasing chitinase activity that confirmed that the inhibition observed was due to the presence of over expressed Tcchitinase-I protein. A positive correlation between increased chitinase activity and resistance to ELS has also been shown earlier (Rohini and Rao 2001). Similar correlations have also been observed in various studies on different crop species (Lin et al. 1995; Tabei et al. 1998; Zhu et al. 1998; Carstens et al. 2003; Itoh et al. 2003; Liang et al. 2005; Nandakumar et al. 2007).
For the evaluation of transgenic resistance against ELS, LLS and rust diseases, a total of 20 T1 events were tested. According to our observation, the events of TC-18, TC-14 and TC-7 performed better in all the parameters tested compared to other events screened for ELS, LLS and rust disease resistance in T1 transgenic plants, respectively. The level of resistance to ELS, LLS and rust in these transgenic peanut plants was comparable or higher than that identified in the cultivated peanut showing 2–5 disease scores on a 1–9 scale (Reddy et al. 1992, 1996; Pensuk et al. 2003; Hossain et al. 2007; Badigannavar et al. 2005).
Recent development of transgenic plants using bacterial chitinase (Bchit) and rice chitinase (RCG-3) genes showed higher expression of enzyme activity conjoined with varied levels of resistance to C. Arachidicola (Iqbalet al. 2011, 2012). The variability of pathogen resistance between transgenic events may be due to the localization of chitinase enzymes at the tissue and cellular levels (Leeuwenet al.2001). Further use of rice chitinase (Rchit) in peanut transgenics displayed longer incubation and latent periods, lower infection rating, fewer lesions against late leaf spot (LLS) and rust diseases (Prasad et al. 2013) as we have observed in the present investigations.
Thus, we conclude that the resistance against C. arachidicola C. Personatum and P. arachidis appeared to be enhanced in those lines which were exhibiting 6.5-fold increase in chitinase enzyme activity by using Tcchitinase-I gene for the first time in peanut cv ICG 13942. The lines showing less enzyme activity did not played a role in resistance against pathogen. The resistant lines will go in breeding cycle in next generations to fix the character in character-deficient cultivar. Once this character proved to be inheritable, these lines could be used as good genetic source of disease resistance breeding material. We expect that the combination of this transgenic strategy based on the use of Tcchitinase-I gene and traditional breeding will provide durable fungal disease-resistant peanut lines with good agronomic phenotypes.
References
Anuradha TS, Divya K, Jami SK, Kirti PB (2008) Transgenic tobacco and peanut plants expressing a mustard defensin show resistance to fungal pathogens. Plant Cell Rep 27:1777–1786
Anuradha TS, Jami SK, Dalta RS, Kirti PB (2006) Genetic transformation of peanut (Arachis hypogaea L.) using cotyledonary node as explant and a promoterlessgus: nptII fusion gene based vector. J Biosci 31(2):235–246
Badigannavar AM, Kale DM, Mondal S, Murty GSS (2005) Trombay groundnut recombinants resistant to foliar diseases. Mutat Breed Newsl Rev 1:11–12
Broglie K, Chet I, Holliday M, Cressman R, Biddle P, Knowlton S (1991) Transgenic plants with enhanced resistance to the fungal pathogen Rizoctoniasolani. Science 254:1194–1197
Carstens M, Vivier MA, Pretorius IS (2003) The Saccharomyces cerevisiaechitinase, encoded by the CTS1-2 gene, confers antifungal activity against Botrytis cinerea to transgenic tobacco. Transgenic Res 2:497–508
Chu Y, Bhattacharya A, Wu C, Knoll JE, Ozias-Akins P (2013) Improvement of peanut (Arachis hypogaea L.) transformation efficiency and determination of transgene copy number by relative uantitative real-time PCR. Vitro Cell Dev Biol Plant 49:266–275
Chu Y, Deng XY, Faustinelli P, Ozias-Akins P (2008) Bcl-xL transformed peanut (Arachis hypogaea L.) exhibits paraquat tolerance. Plant Cell Rep 27:85–92
Datta K, Koukolikova-Nicola Z, Baisakh N, Oliva N, Datta SK (2000) Agrobacterium-mediated engineering for sheath blight resistance of indica rice cultivars from different ecosystems. Theor Appl Genet 100:832–839
Datta K, Tu J, Oliva N, Ona I, Velazhahan R, Mew TW, Muthukrishnan S, Datta SK (2001) Enhanced resistance to sheath blight by constitutive expression of infection-related rice chitinase in transgenic elite indica rice cultivars. Plant Sci 160:405–414
Ganesan M, Bhanumathi P, Ganesh Kumar K, Lakshmi Prabha A, Pill-Soon S, Jayabalan N (2009) Transgenic Indian cotton (Gossypiumhirutum) harboring rice chitinase gene (Chi II) confers resistance to two pathogens. Am J Biochem Biotechnol 5:63–74
Grison R, Grezes-Besset B, Schneider M, Lucante N, Olsen L, Leguay JJ (1996) Field tolerance to fungal pathogens of Brassicanapus constitutively expressing a chimeric chitinase gene. Nat Biotechnol 14:643–646
Hoagland DR, Arnon DI (1950) The water culture method for growing plant without soil. California AgrExp Sta. Cir No 347. University of California Berkley Press, California, pp. 1–32.
Hossain MD, Rahman MZ, Abeda K, Rahman MM (2007) Screening of groundnut genotypes for leaf spots and rust resistance. Int J Sustain Crop Prod 2:7–10
Iqbal MM, Nazir F, Ali S, Asif MA, Zafar Y, Iqbal J, Ali GM (2012) Over expression of rice chitinase gene in transgenic peanut (Arachis hypogaea L.) improves resistance against leaf spot. Mol Biotechnol 50:129–136
Iqbal MM, Zafar Y, Zazir F, Ali S, Iqbal J, Asif MA, Rasid O, Ali GM (2011) Over expression of bacterial chitinase gene in Pakistani peanut (Arachis hypogaea L.) cultivar golden. Afr J Biotechnol 10(31):5838–5844
Iseli B, Boller T, Neuhaus JM (1993) The N-terminal cysteine-rich domain of tobacco class I chitinase is essential for chitin binding but not for catalytic or antifungal activity. Plant Physiol 103(1):221–226
Itoh Y, Takahashi K, Takizawa H, Nikaidou N, Tanaka H, Nishihashi H, Watanabe T, Nishizawa Y (2003) Family 19 chitinase of Streptomyces griseus HUT6037 increases plant resistance to the fungal disease. Biosci Biotechnol Biochem 67:847–855
Kishimoto K, Nishizawa Y, Tabei Y, Hibi T, Nakajima A, Akutsu K (2002) Detailed analysis of rice chitinase gene expression in transgenic cucumber plants showing different levels of disease resistance to graymold (Botrytis cinerea). Plant Sci 162:655–662
Kumar S, Kumar S, Nagi SP, Kanwar JK (2008) In vitro selection and regeneration of chrysanthemum (Dendranthema grandiflorum Tzelev) plants resistant to culture filtrate of SeptoriaobeseSyd. Vitro Cell Dev Biol Plant 44:474–479
Leeuwen VW, Ruttink T, Borst-Vrenssen AW, Van der Plas LH, Van der Krol AR (2001) Characterization of position induced spatial and temporal regulation of transgene promoter activity in plants. J Exp Bot 52:949–959
Liang XQ, Holbmok CC, Lynch RE, Guo BZ (2005) β -1, 3- Glucanase activity in peanut seed (Arachis hypognea) is induced by inoculation with Aspergillus flavus and copurifies with a conglutin-like protein. Phytopathol 95:506–511
Lin W, Anuratha CS, Datta K, Potrykus I, Muthukrishnan S, Datta SK (1995) Genetic engineering of rice for resistance to sheath blight. Nat Biotechnol 13:686–691
Livingstone DM, Hampton JL, Phipps PM, Grabau EA (2005) Enhancing resistance to Sclerotinia minor in peanut by expressing a barely oxalate gene. Plant Physiol 137:1354–1362
Mauch F, Hadwiger LA, Boller T (1984) Ethylene: symptom, not signal for the induction of chitinase and β- 1,3 glucanase in pea pods by pathogens and elicitors. Plant Physiol 76:607–611
Mehta R, Radhakrishnan T, Kumar A, Yadav R, Dobaria JR, Thirumalaisamy PP, Jain RK, Chigurupati P (2013) Coat protein-mediated transgenic resistance of peanut (Arachis hypogaea L.) to peanut stem necrosis disease through Agrobacterium-mediated genetic transformation. Indian J Virol 24:205–213
Moham BR, Sajeena A, Setharaman K, Reddy MS (2003) Advances in genetically engineered (transgenic) plants in pest management-an overview. Crop Prot 22:1071–1086
Murashige T, Skoog I (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Plant Physiol 15:473–497
Nandakumar R, Babu S, Kalpana K, Raguchander T, Balasubramanian P, Samiyappan R (2007) Agrobacterium- mediated transformation of indica rice with chitinase gene for enhanced sheath blight resistance. Biol Plant 51:142–148
Pasonen HL, Seppanen SK, Degefu Y, Rytkonen A, Weissenberg K, Pappinen A (2004) Field performance of chitinase transgenic silver birches (Betulapendula): resistance to fungal diseases. Theor Appl Genet 109:562–570
Patil RS, Ghormade V, Deshpande MV (2000) Chitinolytic enzymes: an exploration. Enzyme Microb Technol 26:473–483
Pensuk V, Patanothai A, Jogloy S, Wongkaew S, Akkasaeng C, Vorasoot N (2003) Reaction of peanut cultivars to late leaf spot and rust. Songklanakarin J Sci Technol 25:289–295
Prasad K, Bhatnagar-Mathur P, Waliyar F, Sharma KK (2013) Over expression of a chitinase gene in transgenic peanut confers enhanced resistance to major soil borne and foliar fungal pathogens. J Plant Biochem Biotechnol 22:222–233
Rajinikanth M, Rama Swamy N (2018) Optimization of factors affecting Agrobacterium-mediated genetic transformation in groundnut (Arachis hypogaea L.). Adv Plants Agric Res 8(3):275–282
Reddy LJ, Nigam SN, Moss JP, Singh AK, Subrahmanyam P, McDonald D, Reddy AGS (1996) Registration of ICGV 86699 peanut germplasm line with multiple disease and insect resistance. Crop Sci 36:821
Reddy LJ, Nigam SN, Subrahmanyam P, Reddy AGS, McDonald D, Gibbons RW, Pentaiah V (1992) Registration of ‘ICGV 87160’ peanut cultivar. Crop Sci 32:1075
Rohini VK, Rao KS (2001) Transformation of peanut (Arachis hypogaea L.) with tobacco chitinase gene: variable response of transformants to leaf spot disease. Plant Sci 160:889–898
Sela-Buurlage MB, Ponstein AS, Bres-Vloemans SA, Melchers LS, Van Den Elzen PJM, Cornelissen BJC (1993) Only specific tobacco (Nicotianatobacum) chitinases and β-1, 3-glucanases exhibit antifungal activity. Plant physiol 101:857–863
Sharma KK, Anjaiah V (2000) An efficient method for the production of transgenic plants of peanut (Arachis hypogaea L.) through Agrobacterium tumefaciens mediated genetic transformation. Plant Sci 159:7–19
Sharma KK, Lavanya M, Anjaiah V (2000) Method for the isolation and purification of genomic DNA from peanut that is suitable for analytical applications. Plant Mol Biol Rep 18:393–400
Sreeramanan S, Maziah M, Xavier R (2009) A protocol for Agrobacterim-mediated transformation of banana with a rice chitinase gene. Emirates J Food Agric 21:18–33
Subrahmanyam P, Williams JH, McDonald D, Gibbons RW (1984) The influence of foliar disease and their control by selective fungicide on a range of groundnut (Arachis hypogaea L.) genotypes. Ann Appl Biol 104:467–476
Sundaresha S, Manoj Kumar A, Rohini S, Math SA, Keshsamma E, Chadrashekar SC, Udayakumar M (2009) Enhanced protection against two major fungal pathogens of groundnut, Cercospora arachidicola and Aspergillus flavus in transgenic groundnut over-expressing a tobacco β 1–3 glucanase. Eur J Plant Pathol 126:497–508
Tabei Y, Kitade S, Nishizawa Y, Kikuchi N, Kayano T, Hibi T, Akutsu K (1998) Transgenic cucumber plants harboring a rice chitinase gene exhibit enhanced resistance to graymold (Botrytis cinerea). Plant Cell Rep 17:159–164
Takahashi W, Fujimori M, Miura Y, Komatsu T, Nishizawa Y, Hibi T, Takamizo T (2005) Increased resistance to crown rust disease in transgenic Italian ryegrass (Lolium multiflorum Lam) expressing the rice chitinase gene. Plant Cell Rep 23:811–818
Tiwari S, Mishra DK, Singh A, Singh PK, Tuli R (2008) Expression of asynthetic cry1EC gene for resistance against Spodoptera litura in transgenic peanut (Arachis hypogeae L.). Plant Cell Rep 27:1017–1025
Zhu H, Muthukrishnan S, Krishnaveni S, Wilde G, Jeoung JM, Liang GH (1998) Biolistic transformation of sorghum using a rice chitinase gene. J Genet Plant Breed 52:243–252
Zhu Q, Maher EA, Masoud S, Dixon RA, Lamb CJ (1994) Enhanced protection against fungal attack by constitutive co-expression of chitinase and glucanase genes in transgenic tobacco. Nat Biotechnol 12:807–812
Acknowledgements
We thank the University Grants Commission, New Delhi for financial assistance as Fellow under Basic Science Research-Research Fellowship in Science for Meritorious Students (BSR–RFSMSF.4-1/2006(BSR)/7-211/2009(BSR) dt.26-02-2013.). We acknowledge Dr. HD. Upadhyaya ICRISAT, Patancheru, Hyderabad (TS), India for providing the germplasm (seeds) of cv ICG 13942 and Prof. N. Jayabalan Department of Plant Science, Bharatidasan University, Thiruchurappally, Tamil Nadu, India, for the providing binary plasmid pBinAR.
Author information
Authors and Affiliations
Contributions
RM, the first author, has performed experiments and wrote the manuscript. RSN has designed the experiments and also corrected the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Marka, R., Nanna, R. Expression of Tcchitinase-I gene in transgenic peanut (Arachis hypogaea L.) confers enhanced resistance against leaf spot and rust diseases. Plant Growth Regul 93, 53–63 (2021). https://doi.org/10.1007/s10725-020-00663-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-020-00663-8