Abstract
Defensins are small positively charged, antimicrobial peptides (~5 kDa in size) and some of them exhibit potent antifungal activity. We have cloned the complete cDNA containing an ORF of 243 bp of a defensin of mustard. The deduced amino acid sequence of the peptide showed more than 90% identity to the amino acid sequence of the well-characterized defensins, RsAFP-1 and RsAFP-2 of Raphanus sativus. We have generated and characterized transgenic tobacco and peanut plants constitutively expressing the mustard defensin. Transgenic tobacco plants were resistant to the fungal pathogens, Fusarium moniliforme and Phytophthora parasitica pv. nicotianae. Transgenic peanut plants showed enhanced resistance against the pathogens, Pheaoisariopsis personata and Cercospora arachidicola, which jointly cause serious late leaf spot disease. These observations indicate that the mustard defensin gene can be deployed for deriving fungal disease resistance in transgenic crops.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the innate immunity response, only the defensins are a class of peptides that are conserved in plants, invertebrates and vertebrates (Thomma et al. 2002). Although they share common chemical elements and 3-D structures, the plant defensin family is quite diverse in amino acid composition and biological activity (Lay and Anderson 2005; Thomma et al. 2002). Defensins have been found to display antimicrobial activity not only against plant and insect pathogens, but also against human fungal pathogens including Candida and Aspergillus sps, and they are employed as novel leads in antifungal therapeutics (Thevissen et al. 2007). The antifungal activity of SPE10, a dimeric defensin isolated from Pachyrhizus erosus was assessed on three fungal species and the antifungal activity depended on the test fungus (Song et al. 2004, 2005). In vitro antifungal activity of a defensin from Trigonella foenum graecum was tested against some fungal pathogens (Olli and Kirti 2006).
Peanut is a crop of high commercial value and its cultivation is hampered by the attack from several diseases. Leaf spot disease, jointly caused by Phaeosariopsis personata and Cercospora arachidicola, is a very serious problem affecting peanut cultivation in various countries. This disease predominates in the tropics and subtropics, and is reported to be the one of the major diseases of peanut causing high yield losses. The genus Cercospora belongs to the class of Fungi Imperfecti and the pathogen survives in the form of conidia, perithecia and mycelia in the soil. Symptoms first appear on the leaves as small brown flecks on both sides of the leaves; early leaf spots caused by C. arachidicola appear on the adaxial surface of the leaflets, where as the late leaf spots caused by P. personata appear on the abaxial surface and the lesion diameter increases to >2 mm within 10–15 days. If not controlled, the leaf spots reduce yields of typical peanut cultivars up to 90% (Austin Hagan, http://www.aces.edu/pubs/docs/A/ANR-0369/).
Plant growth promoting rhizobacteria (PGPR), which were earlier reported as elicitors in induced systemic resistance (ISR) in several crop species, did not provide significant protection in peanut against late leaf spot pathogen (Zhang et al. 2001). Other biological/chemical strategies include the application of Serratia marcescens (GPSS5), which was reported to provide an improved control of late leaf spot disease of peanut (Kishore et al. 2005). However, these strategies need repeated application on the plants during the growth phase to be effective. Hence, the genetic transformation for transferring suitable candidate genes to peanut offers a more effective solution for developing transgenic varieties with enhanced disease resistance.
Expression of genes for suitable pathogenesis related proteins and defensins offers a suitable approach for controlling diseases that affect crop productivity. Overexpression of pepper pathogen induced genes CAPIP2, CASAR82A and RAV1 in transgenic plants resulted in disease resistance and abiotic stress tolerance (Lee and Hwang 2006; Lee et al. 2006; Sohn et al. 2006). The pathogenesis related proteins, that have been shown to have potent antifungal activities, have also been implicated in abiotic stress tolerance. Sarowar et al. (2005) showed that the overexpression of a pepper basic pathogenesis related protein-1 in transgenic tobacco plants could be correlated with disease resistance and enhanced heavy metal tolerance, whereas the same protein conferred oxidative stress tolerance and resistance to Pseudomonas syringae in Arabidopsis (Hong and Hwang 2005).
There were very few reports on transforming peanut for introduction of genes for fungal disease resistance. Transgenic peanut expressing a tobacco chitinase gene was shown to possess enhanced resistance to the late leaf spot causing organism P. personata (Rohini and Sankara Rao 2001). However, the analysis of resistance in this study was limited to detached leaf assay only. Chenault et al. (2005) expressed a rice chitinase and an alfalfa glucanase in transgenic peanut and observed enhanced resistance against Slerotinia blight in the transgenic plants. Expression of a barley oxalate oxidase in transgenic peanut also enhanced resistance to Sclerotinia minor (Livingstone et al. 2005). These observations indicate that the introduction of suitable genes for resistance through genetic transformation is a feasible option in combating the diseases in peanut.
In the present investigation, we have shown that a mustard defensin confers resistance to fungal pathogens, Phytophthora parasitica pv. nicotianae, Fusarium moniliforme in transgenic tobacco and against pathogens causing the leaf spot disease in peanut.
Materials and methods
Plasmid construction and Agrobacterium transformation
The ORF of the cDNA of the mustard defenisn (BjD) was cloned by RT-PCR (Swathi Anuradha et al., unpublished) of the total RNA extracted from leaves of 1-month-old mustard (Brassica juncea cv. Pusa Jai Kisan) plants using TRI-reagent (Sigma, USA) using the forward and reverse primers (OD′F′:5′GGG TAC CAT GGC TAA GGT TGA TTC CATC 3′, OD′R: 5′GGG ATC CTT AAC AAG GGA AGT AGC AGA 3′) designed on the basis of sequence information available for a Brassica oleracea defensin (NCBI Accession No. CAC37558). The open reading frame of BjD was cloned as an NcoI–BamHI fragment into a plant expression cassette containing vector pRT100. The entire defensin cassette along with 5′ CaMV 35S promoter and 3′ Nos terminator was cloned as a PstI fragment in the binary vector pCAMBIA2300, which was mobilized into a super virulent Agrobacterium tumefaciens strain EHA105 by the freeze thaw method. A fresh overnight culture of Agrobacterium was obtained by inoculating a single colony and the culture with bacterial OD 0.6–0.8 was used in leaf disc transformation of tobacco, Nicotiana tabacum cv. Petit Havana following the method of Horsch et al. (1985). A peanut cultivar, JL-24 was used in transformations experiments for expressing the mustard defensin. Agrobacterium mediated transformation of peanut, selection and regeneration were carried out following the procedures detailed out earlier using embryo axis explants (Swathi Anuradha et al. 2008). Putative transgenic plants were transferred to the glass house, seed collected and the T1 generation plants were characterized further using Southern and northern analyses.
Molecular analysis of the transformants
DNA isolation, southern blotting, hybridization and washing were performed according to Swathi Anuradha et al. (2006). RNA was isolated from the young leaves frozen in liquid nitrogen using the TRI-Reagent (Sigma–Aldrich, USA) following the manufacturer’s instructions in molecular analyses. Radiolabelled probes for BjD and neomycin phosphotransferase (nptII) were used in the nucleic acid hybridization reactions. Standard protocols were applied in molecular analyses (Sambrook et al. 1989).
Evaluation of transgenic tobacco plants for resistance against fungal pathogens
Detached leaf assays were conducted on transgenic tobacco plants in T1 generation, which were characterized for the expression of BjD. The oomycete pathogen Phytophthora parasitica pv. nicotianae was cultured on PDA medium (potato 200 g/l, sucrose 20 g/l, agar 15 g/l, pH 6.5) at 24°C for 5–7 days. Fusarium moniliforme was grown on PDA medium at 37°C for 3–4 days. For the detached leaf assay, fully expanded leaves of 2-month-old plants (Control, T1 progeny of tobacco transgenic plants 8 and 10) were used for inoculation. A small plug of the mycelium was placed at the center of the adaxial leaf surface. The inoculated leaves were then placed on two layers of filter paper saturated with sterilized double distilled water in a Petri dish at a 16 h light/8 h dark photoperiod in a BOD incubator at 24°C and symptoms were evaluated 10 days after inoculation.
Evaluation of peanut transgenic plants expressing the mustard defensin
For resistance studies against P. personata and C. arachidicola, the causal agents of leaf spot disease (Tikka disease) in peanut, conidia collected from infected leaf samples were allowed to germinate in sterile double distilled water and sucrose at a concentration of 2.0% (w/v) was added to aqueous spore suspension for improving conidial germination. The suspension consisted mostly of conidia of P. personata (~90%) with conidia belonging to C arachidicola; conidia of both the pathogens could be very easily identified by observing the morphology microscopically. The concentration of conidia was adjusted to 5 × 104/ml using a haemocytomer and plant assays were carried out using the conidial suspension with 0.1% of Triton X-100 (v/v) as a wetting agent. One-month-old transgenic plants growing in glass house were inoculated with conidial suspension using an atomizer and high humidity was provided to the plants before and after inoculation of spores. The plants were covered with polythene bags to maintain ≥95% RH for 48 h. After this treatment, the bags were removed and the plants were monitored regularly for disease development. Data was scored after 28 days of inoculation and plants were assessed for leaf spots based on a 9-point disease scale developed at ICRISAT (Subrahmanyam et al. 1982).
Detached leaf assays on peanut transgenics
For detached leaf assays, the same spore suspension was painted on the surface of the leaflets and ≥95% RH was maintained by placing them on filter papers inside 110 mm Petri dishes, sealed with parafilm. The number and average diameter of the lesions were scored after 21 and 28 days of inoculation on control and transgenic leaves. All these experiments were carried out thrice and the data obtained is an average of three independent experiments.
Results
Mustard defensin and its characterization
The cDNA of a defensin (BjD) with an open reading frame of 243 bp was cloned by RT-PCR by using the primers designed on the sequence of Brassica oleracea defensin (Swathi Anuradha et al., unpublished). PCR amplification of the genomic DNA with the same primers produced the genomic clone, which showed a 91 bp intron interrupting the coding region (NCBI accession numbers DQ191751, DQ191752). RACE has resulted in cloning a 5′-untranslated region (UTR) of 32 bp upstream to the start codon and a 3′-UTR, which was 185 bp long downstream of the stop codon (Swathi Anuradha et al., unpublished).
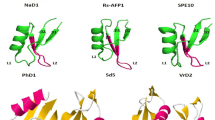
A NCBI database search with BLAST and the multiple alignment of the amino acid sequence of the present defensin with other defensins showed that BjD had considerable homology to known Crucifer defensins. It showed 98% amino acid sequence identity with RsAFP-1, 90% with the well-characterized defensin, RsAFP-2 with antifungal activity, and 95% with the defensin from B. oleracea. The alignment of the deduced mature polypeptide sequences of BjD and plant defensins from several Crucifer species is shown in Fig. 1.
Bj: Brassica juncea, Sa: Sinapis alba, Rs: Raphanus sativus, Bo: Brassica oleracea, Ah: Arabidopsis halleri, Br: Brassica rapa, Ew: Eutrema wasabi, At: Arabidopsis thaliana and Lm: Lepidium meyenii. The protein sequence shared maximum identity (only the mature part of the peptide from different accession has been indicated here) of 98% to Rs AFP1 of Raphanus sativus (P69241), 95% to Bo of Brassica oleracea (CAC37558), 91% to Sinapis alba (AAY15221), 90% to Rs AFP2 of Raphanus sativus (P30230) and Ah2 of Arabidopsis halleri (AAY27736). The BjD protein also showed significant sequence identity with other defensins like Br of Brassica rapa (AAQ92328), Ew of Eutrema wasabi (BAB19054), At of Arabidopsis thaliana (NP_199256) and Lm of Lepidium meyenii (AAV85992)
Tobacco transformation and molecular analysis
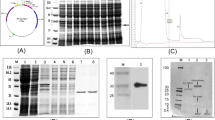
Agrobacterium strain EHA105, harboring the binary vector pCAMBIA2300 carrying the BjD expressed under the CaMV 35S promoter (Fig. 2), was used in the genetic transformation of tobacco using the leaf disc method. This has resulted in the production of several putative transgenic plants. However, molecular characterization was carried out for 14 T0 plants obtained from leaf disc transformation of tobacco. These plants were acclimatized under culture room conditions, transferred to pots, grown in the glass house, and confirmed through Southern hybridization for copy number. The hybridization pattern indicated that they carried independent integration events for the transgenes and some of them (Plants 7, 8, 10, 13) are single copy carrying transgenic plants (Fig. 3).
The expression of the transgene BjD in the T0 transgenic plants was analyzed by northern hybridization using the PCR amplified fragment as a probe. Out of 14 T0 plants that were analyzed, 12 plants exhibited varied levels of expression of BjD mRNA. The presence of ~250 bp band in the transgenic plants indicated the expression of BjD in the T0 tobacco plants. Transgenic plants 8 and 10 represented the primary transgenics carrying single copy insertion with relatively high expression level of the introduced BjD. The plants 2, 5 and 6 showed relatively low-level expression of BjD (Fig. 4a). The progeny of plants 8 and 10 were analyzed by Northern hybridization to detect the stability of expression of BjD mRNA level. Progeny plants of the transgenic plants that did not carry the transgene due to segregation also did not show BjD expression (Fig. 4b).
a Northern analysis of tobacco transgenic plants in T0 generation using the PCR amplified mustard defensin as a probe. Plants 4, 8, 9, 10, 14 represent relatively high expression of mustard defensin. b Confirmation of defensin gene expression in T1 generation plants of the transgenics 8 and 10 and identification of transgene containing progeny plants and segregants without transgenes for resistance analysis
Genetic transformation and Southern blot analysis of peanut transgenic plants
By employing pCAMBIA2300 harboring BjD and the same Agrobacterium strain for transformation, we have generated transgenic peanut plants in the cultivar JL-24. Overall, 17 putative transformants were obtained from six independent transformation experiments using embryo axis explants. Out of these, 13 survived under the glass house conditions, 7 of which were found to be positive in a preliminary PCR analysis. We collected seeds from 11 T0 plants and the Southern analysis was done with PCR positive T1 plants. The genomic DNA of transgenic plants was digested with enzyme EcoR1, which has a single site on the T-DNA between the nptII and defensin expression cassette, and the Southern blot was probed with the PCR amplified nptII fragment as probe. The varying length of hybridizing bands indicated that at least three of these primary transgenic plants represented single copy containing independent transformation events. Transgene integration was evident in the progeny of five plants out of eleven T0 plants that survived the transfer to the glass house (Fig. 5).
Analysis of constitutive expression of mustard defensin in peanut T1 plants
Northern hybridization analysis on total RNA isolated from the leaves of T1 transgenic peanut plants confirmed the constitutive expression of BjD (Fig. 6). The expression levels varied among individual plants and hybridization signals corresponding to the expected transcript size of ~250 nt for defensin were detected in transgenic plants. High level expression was evident in the progeny of transgenic plants 3 and 4. No hybridization signals were observed in the control plant. Progeny plants of the transgenic plants that did not carry the transgene due to segregation also did not show BjD expression.
Enhanced disease resistance in transgenic tobacco against Phytophthora parasitica pv. nicotianae and Fusarium moniliforme
Phytophthora parasitica pv. nicotianae causes a serious disease called black shank disease in tobacco. To determine whether the constitutive expression of mustard defensin conferred resistance against this pathogen, a detached leaf assay was performed with the leaves of BjD overexpressing tobacco T1 plants. This experiment was carried out to correlate the expression of the BjD and resistance against pathogen invasion as the transgenes segregate in a Mendelian fashion from T0 to the T1 generation. This results in transgenic segregants expressing the transgene and non-transgenic segregants lacking the transgene. Randomly selected T1 plants in the greenhouse were checked for the presence of transgenes using PCR for nptII and BjD (data not shown). Subsequently, the selected plants carrying the transgenes and their non-transgenic counterparts were analyzed for the expression of the defensin through northern analysis and fungal resistance. Disease symptoms started appearing after three days of inoculation and they were scored after 10 days of inoculation. Control plants showed browning of entire leaves with necrotic lesions all over the leaves. T1 plants (8-1, 8-3, 10-2, 10-3 and 10-5), which did not carry the transgene due to segregation (as shown in the northern analysis for the expression of the mustard defensin, Fig. 4b), also did not show resistance to the pathogen infection like the control plants (Fig. 7), whereas the T1 transgenic plants carrying stably inherited transgene (8-2, 8-4, 8-5, 10-1 and 10-4) showed significantly enhanced resistance to the pathogen. On these plants, symptoms were not detected even after 15 days post-inoculation and the leaves remained fresh without any sign of disease. In some of the plants, lesions formed, but lesion size did not increase further.
Analysis of tobacco transgenic plants with the fungus Phytophthora parasitica pv. nicotianae. Note the development of disease on control and segregants that did not express mustard defensin (8-1, 8-3, 8-5, 10-2, 10-3 and 10-5) and the control plants as shown in the northern analysis in Fig 4b. The transgene expressing segregants resisted the infection
Fusarium moniliforme attacks young seedlings causing vein clearing, wilting, chlorosis and necrosis in many crop plants including tobacco. Plants were affected at later stages also. Transgenic plants were tested for their ability to resist this pathogen infection. Detached leaf assays were performed and symptoms were observed after 1 week. Vein clearing was evident in the detached control leaves, which was not evident in transgenic leaves (Fig. 8).
The above results suggest that transgenic plants expressing the mustard defensin constitutively exhibited enhanced resistance to the fungal pathogen infection.
Evaluation of the transgenic peanut plants for resistance to tikka or late leaf spot disease-whole plant assay
The Southern confirmed peanut transgenic plants were assayed for their ability to resist infection from the leaf spot causing pathogens in whole plant analysis using conidial spray. The disease symptoms started appearing on control plants after 10–12 days as small specks, which later developed into lesions. Data was scored after 28 days of inoculation and plants were assessed for leaf spots. On control plants the numbers of spots were high in number and the lesion size also increased within 10–15 days, whereas the size and number of lesions on resistant transgenic plants did not increase even after 28 days. When the total number of intact leaflets (four leaflets per leaf) and the number of diseased leaves on the main stem were counted at the glass house level, the control plants were observed to have a disease severity of 6–7 out of 9 against the transgenic plants, which showed a reaction of 1–3 in the scale (Fig. 9).
Detached leaf assay
We have carried out detached leaf assay to further confirm our results on the analysis in the glass house and the data obtained was consistent with our data on the whole plant assay. Disease symptoms started appearing after 15 days as brown color specks on these leaves. All the transgenic lines showed increased resistance to P. personata as measured by the average number of lesions and a reduction in lesion area (Fig. 10). Some of the lesions observed on these leaves belonged to the early leaf spots caused by Cercospora arachidicola.
The average number of lesions was found to be less in transgenic plants compared to the controls (Fig. 11a). The decrease in the number of lesions was found to be significantly lower in transgenics (P ≥ 0.05). The type of lesion also varied between transgenic and control lines. Controls showed large, continuous, black colored lesions; whereas the transgenic lines typically exhibited smaller lesions resembling a hypersensitive response. The lesion size was found to be significantly less in transgenics (P ≥ 0.05). The lesion frequency has further decreased to 90.91% in BjD-3-1 and 92.73 in BjD-7-2. In the same set of plants, the lesion size also decreased to 85.72% in BjD-10-2 and 88.58% in BjD-7-2 (Fig. 11b). Though the lesion frequency was high in BjD10-2, the size of the lesions did not grow even after continued incubation of leaves.
a Analysis of lesion number in control and transgenic plants after conidial spray in detached leaf assays. There is a significant difference in the number of lesions on transgenic plants and the control (P ≥ 0.05). b Analysis of lesion diameter in control and transgenic plants in detached leaf assay. Note the significant reduction in lesion diameter in the transgenic plants (P ≥ 0.05)
The estimation of spore/conidial germination or growth using crude protein extracts from plants is also commonly used in determining the antifungal activity. We performed MTT assay to check the germination of conidia of the late leaf spot fungus in plant protein crude extracts of both control and transgenic plants. We observed that 50 μg of crude protein extracts of transgenic plants BjD 4-2 was able to inhibit 18% of the spore germination. Hence, this concentration can be considered as minimal inhibitory concentration (MIC). Crude protein (100 μg) was able to inhibit 50% of spore germination and this concentration was considered to be lethal for spore germination. Spore germination was not inhibited in control plant protein extracts. This assay showed that transgenic plant protein extracts inhibited fungal spore germination considerably (data not shown).
These results showed that the constitutive expression of BjD in peanut confers enhanced resistance against late leaf spot pathogen also.
Discussion
Transgenic expression of plant defensins has been reported to enhance protection in vegetative tissues against pathogen attack. Constitutive expression of RsAFP-2 enhanced resistance of tobacco plants to the leaf pathogen Alternaria longipes (Terras et al. 1995) and tomato to Alternaria solani (Parashina et al. 2000). Canola expressing a pea defensin showed enhanced resistance against blackleg disease caused by Leptosphaeria maculans (Wang et al. 1999). The constitutive expression of an alfalfa defensin in potato provided robust resistance against the agronomically important fungus Verticillium dahliae under field conditions (Gao et al. 2000). Overexpression of BSD1 (stamen specific defensin) in transgenic tobacco plants enhanced their tolerance against the pathogen Phytophthora parasitica (Park et al. 2002). Expression of a defensin gene along with a glucanase gene has been shown to enhance tomato resistance against Ralstonia solanacearum (Chen et al. 2006). More recently, Aerts et al. (2007) have reported that the expression of human beta defensin-2 in Arabidopsis resulted in enhanced resistance against Botrytis cinerea demonstrating the functional homology of defensins across Kingdoms.
We have cloned a defensin from mustard using the sequence information available for a B. oleracea defensin and it had high amino acid sequence identity to the B. oleracea defensin as well as the well-characterized defensins RsAFP-1 and 2 of Raphanus sativus. RsAFP-2 has been shown to possess antifungal activity (Terras et al. 1995). We have generated transgenic tobacco and peanut plants expressing the mustard defensin constitutively. Transgenic tobacco plants expressing mustard defensin showed enhanced resistance against two fungal pathogens, Fusarium moniliforme and Phytophthora parasitica pv. nicotianae in detached leaf assays showing the efficacy of the mustard defensin in imparting disease resistance in tobacco.
Peanut transgenic plants also displayed substantially increased levels of leaf spot disease resistance in detached assay and green house test using the conidial suspension mixture of P. personata and C. arachidicola. The T1 progeny plants, segregating for the transgene, were evaluated for resistance against the leaf spot disease and they exhibited varied resistance as indicated by a range of 1–7 on the 9 point scale (Subrahmanyam et al. 1982); those progeny plants that received the transgene showed a reaction 1–2 as against the non-transgenic segregants, which behaved like the wild type control plants. All the control plants and the segregants from the transgenic plants that did not receive the transgene showed extensive disease symptoms leading to defoliation and severe necrosis. Highest percent reduction in lesion frequency was noticed on the transgenic plant BjD-7-2. Transgenic plants expressing mustard defensin consistently showed higher levels of resistance.
Our results correlated well with previously published reports in other plants. Lesion size resulting from inoculation with Sclerotinia minor was reduced to 97% in transgenic peanut plants over expressing a barley oxalate oxidase gene (Livingstone et al. 2005). Similarly, the lesion size was reduced by 63% in poplar expressing a wheat oxalate oxidase gene (Liang et al. 2001). Oxalate decarboxylase was shown to be produced by the resistant host plants to degrade oxalic acid produced in some plant pathogen attacks and the expression of the gene for this enzyme decreased the lesion size approximately by 89% in transgenic tomato plants inoculated with S. sclerotiarum (Kesarwani et al. 2000). Transgenic rice plants overexpressing a wasabi defensin also showed enhanced resistance to Magnaporthe grisea and the average size of disease lesions was reduced to about half of that in the non-transformed plants (Kanzaki et al. 2002). The lesion size decreased significantly in grapevine transgenic plants inoculated with anthracnose fungus (Yamamoto et al. 2000). Transgenic tobacco plants expressing a magainin analog MYP30 were evaluated for resistance to Peronospora tabacina infection and significant reduction in sporulation and lesion size were observed (Li et al. 2001). Resistance to Alternaria brassisicola in transgenic broccoli expressing a Trichoderma harzianum endochitinase gene was assessed by a detached leaf assay of T0 plants inoculated with Alternaria brassisicola and the lesion size showed a negative correlation with endochitinase levels (Mora et al. 2001).
The present investigation shows that the expression of the mustard defensin in transgenic tobacco and peanut plants also conferred enhanced resistance against various pathogens amply demonstrating the efficacy of defensins in imparting disease resistance in crop plants and indicating that they are good candidate genes in crop improvement.
References
Aerts MGM, Thevissen K, Bresseleers SM, Sels J, Wouters P, Cammue BPA, Francois IEJA (2007) Arabidopsis thaliana plants overexpressing human beta defensin-2 are more resistant to fungal attack: functional homology between plant and human defensins. Plant Cell Rep 26:1391–1398
Chen S-C, Lin A-R, Zou Z-R (2006) Overexpression of a glucanase gene and a defensin gene in transgenic tomato enhances resistance to Ralstonia solanacearum. Russ J Plant Physiol 53:671–677
Chenault KD, Melouk HA, Payton ME (2005) Field reaction to Sclerotinia blight among transgenic peanut lines containing antifungal genes. Crop Sci 45:511–515
Gao AG, Hakimi SM, Mittanck CA, Wu Y, Woerner BM, Stark DM, Shah DM, Liang J, Rommens CM (2000) Fungal pathogen protection in potato by expression of a plant defensin peptide. Nature Biotechnol 18:1307–1310
Hong JK, Hwang BK (2005) Induction enhanced disease resistance and oxidative stress tolerance by overexpression of a pepper basic PR1 gene in Arabidopsis. Physiol Plant 124:267–277
Horsch RB, Fry JE, Hoffman NL, Eichholtz D, Rogers SG, Fraley RT (1985) A simple and general method for transferring genes into plants. Science 227:1229–1231
Kanzaki H, Nirasawa S, Saitoh H, Ito M, Nishihara M, Terauchi R, Nakamura I (2002) Overexpression of the wasabi defensin gene confers enhanced resistance to blast fungus (Magnaporthe grisea) in transgenic rice. Theor Appl Genet 105:809–814
Kesarwani M, Azam M, Natarajan K, Mehta A, Datta A (2000) Oxalate decarboxylase from Collybia velutipes: Molecular cloning and its overexpression to confer resistance to fungal infection in transgenic tobacco and tomato. J Biol Chem 275:7230–7238
Kishore GK, Pande S, Podile AR (2005) Chitin-supplemented foliar application of Serratia marcescens GPS 5 improves control of late leaf spot disease of groundnut by activating defence-related enzymes. J Phytopathol 153:169–175
Lay FT, Anderson MA (2005) Defensins-Components of innate immunity in plants. Curr Prot Pept Sci 6:85–101
Lee SC, Hwang BK (2006) CASAR82A, a pathogen induced pepper SAR8.2 exhibits antifungal activity and its overexpression enhances disease resistance and stress tolerance. Plant Mol Biol 61:95–109
Lee SC, Kim SH, An SH, Yi SY, Hwang BK (2006) Identification and functional expression of the pepper pathogen induced gene, CAPIP2, involved in disease resistance and drought and salt tolerance. Plant Mol Biol 62:151–164
Li QS, Lawrence CB, Xing H-Y, Babbitt RA, Bass WT, Maiti IB, Everett NP (2001) Enhanced disease resistance conferred by expression of an antimicrobial magainin analog in transgenic tobacco. Planta 212:635–639
Liang H, Maynard CA, Allen RD, Powell WA (2001) Increased Septoria musiva resistance in transgenic hybrid poplar leaves expressing a wheat oxalate oxidase gene. Plant Mol Biol 45:619–629
Livingstone DM, Hampton JL, Phipps PM, Grabau EA (2005) Enhancing resistance to Sclerotinia minor in peanut by expressing a barley oxalate oxidase Gene. Plant Physiol 137:1354–1362
Mora AA, Earle ED (2001) Resistance to Alternaria brassicicola in transgenic broccoli expressing a Trichoderma harzianum endochitinase gene. Mol Breed 8:1–9
Olli S, Kirti PB (2006) Cloning, characterization and antifungal activity of defensin Tfgd1 from Trigonella foenum-graecum L. J Biochem Mol Biol 39:278–283
Parashina EV, Serdobinskii LA, Kalle EG, Lavorova NV, Avetisov VA, Lunin VG, Naroditskii BS (2000) Genetic engineering of oilseed rape and tomato plants expressing a radish defensin gene. Russ J Plant Physiol 47:417–423
Park HC, Kang YH, Chun HJ, Koo JC, Cheong YH, Kim CY, Kim MC, Chung WS, Kim JC, Yoo JH, Koo YD, Koo SC, Lim CO, Lee SY, Cho MJ (2002) Characterization of a stamen-specific cDNA encoding a novel plant defensin in Chinese cabbage. Plant Mol Biol 50:57–68
Rohini VK, Sankara Rao K (2001) Transformation of peanut (Arachis hypogaea L.) with tobacco chitinase gene: variable response of transformants to leaf spot disease. Plant Sci 160:883–892
Sambrook J, Fritsch EF, Maniatis T (1989) Molecular cloning: a laboratory manual, 2nd edn. Cold Spring Harbor Laboratory Press, New York
Sarowar S, Kim JJ, Kim EN, Kim KD, Hwang BK, Islam R, Shin JH (2005) Overexpression of a pepper basic pathogenesis related protein 1 gene in tobacco plants enhances resistance to heavy metal and pathogen stresses. Plant Cell Rep 24:216–224
Sohn KH, Lee SC, Jung HW, Hong JK, Hwang BK (2006) Expression and functional roles of the pepper pathogen induced transcription factor RAV1 bacterial disease resistance and drought and salt tolerance. Plant Mol Biol 61:897–915
Song X, Wang J, Wu F, Li X, Teng M, Gong W (2005) cDNA cloning, functional expression and antifungal activities of a dimeric plant defensin SPE10 from Pachyrrhizus erosus seeds. Plant Mol Biol 57:13–20
Song X, Zhaocai Z, Wang J, Wu F, Gong W (2004) Purification, characterization and preliminary crystallographic studies of a novel plant defensin from Pachyrrhizus erosus seeds. Acta Crystallographica D 60:1121
Subrahmanyam P, McDonald D, Gibbons RW, Nigam SN, Nevill DJ (1982) Resistance to rust and late leaf spot diseases in some genotypes of Arachis hypogaea. Peanut Sci 9:6–10
Swathi Anuradha T, Jami SK, Datla RS, Kirti PB (2006) Genetic transformation of peanut (Arachis hypogaea L) using cotyledonary node as explant and a promoterless gus::nptII fusion vector. J Biosci 31:1–12
Swathi Anuradha T, Jami SK, Beena MR, Kirti PB (2008) Cotyledonary node and embryo axes as explants in legume transformation with special reference to peanut. In: Kirti PB (ed) Handbook of new technologies for genetic improvement of legumes. CRC Press, Boca Raton, pp 253–271
Terras FR, Eggermont K, Kovalega V, Raikhel NV, Osborn RW, Kester A, Rees SB, Torrekens S, Van Leuven F, Vanderleyden J (1995) Small cysteine-rich antifungal proteins from radish: their role in host defense. Plant Cell 7:573–588
Thevissen K, Kristensen HK, Thomma BPHJ, Cammue BPA, François IEJA (2007) Therapeutic potential of antifungal plant and insect defensins. Drug Discovery Today 12:966–971
Thomma BPHJ, Cammue BPA, Thevissen K (2002) Plant defensins. Planta 216:193–202
Wang YP, Nowak G, Culley D, Hadwiger LA, Fristensky B (1999) Constitutive expression of pea defense gene DRR206 confers resistance to blackleg (Leptosphaeria maculans) disease in transgenic canola (Brassica napus). Mol Plant Microbe Interact 12:410–418
Yamamoto T, Iketani H, Ieki H, Nishizawa Y, Notsuka K, Hibi T, Hayashi T, Matsuta N (2000) Transgenic grapevine plants expressing a rice chitinase with enhanced resistance to fungal pathogens. Plant Cell Rep 19:639–646
Zhang S, Reddy MS, Kokalis-Burelle N, Wells LW, Nightingale SP, Kloepper JW (2001) Lack of induced systemic resistance in peanut to late leaf spot disease by plant growth promoting rhizobacteria and chemical elicitors. Plant Disease 85:879–884
Acknowledgments
This work was supported by a research grant from the Andhra Pradesh-Netherlands Biotechnology Program administered by the Institute of Public Enterprise, Osmania University Campus, Hyderabad. Thanks are due to the Head, Department of Plant Sciences for facilities provided by the UGC-SAP, DST-FIST, and COSIST etc. SKJ and KD are grateful to the Council of Scientific and Industrial Research, Government of India for Research Fellowships. SKJ and KD received the Research Fellowships from the Council of Scientific and Industrial Research, Government of India.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by A. Atanassov.
Rights and permissions
About this article
Cite this article
Swathi Anuradha, T., Divya, K., Jami, S.K. et al. Transgenic tobacco and peanut plants expressing a mustard defensin show resistance to fungal pathogens. Plant Cell Rep 27, 1777–1786 (2008). https://doi.org/10.1007/s00299-008-0596-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-008-0596-8