Abstract
In order to well understand the molecular basis of heterosis in soybean, the methylation-sensitive amplification polymorphism (MSAP) method based on capillary electrophoresis was used to estimate levels and patterns of cytosine methylation in 15-day post-emergence leaves of four parental lines [Jilin 47 (no. 19), EXP (no. 12), Jilin 38 (no. 3) and Yi 3 (no. 6)] and 12 hybrids [Jilin 38 × Yi 3(3 × 6), Jilin 38 × EXP(3 × 12), Jilin 38 × Jilin 47(3 × 19), Yi 3 × Jilin 38(6 × 3), Yi 3 × EXP(6 × 12), Yi 3 × Jilin 47(6 × 19), EXP × Jilin 38(12 × 3), EXP × Yi 3(12 × 6), EXP × Jilin 47(12 × 19), Jilin 47 × Jilin 38(19 × 3), Jilin 47 × Yi 3(19 × 6), Jilin 47 × EXP(19 × 12)]. In addition, 12 traits of the hybrids and their parents were also analyzed to understand the relationship between DNA methylation variation and heterosis. MSAP results showed that the total relative methylation level of all hybrids was lower than the corresponding middle parent value, indicating that the methylation degree was decreasing. And may express a variety of genes related to the phenotypic variation of hybridization. Moreover, the hemi-methylation levels of Jilin 38 × Jilin 47 and Yi 3 × Jilin 47 hybrids and full-methylation levels of EXP × Yi 3 and EXP × Jilin 47 hybrids was significant higher than the corresponding mid-parent values. In addition, the heredity of methylation from parents in hybrids is more than the variations, in which there were four types appeared great higher: A1, B4, B8, and D2. Furthermore, the results of relationship between genetic variation in DNA methylation and heterosis showed that the hypo-methylation had a promoting effect to increase node number, and the hype-methylation of hybrids was helpful to add to stem thick. Our results may provide new insights into well understanding the molecular mechanisms of heterosis at the epigenetic level in soybean.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In the post-genomics era, cytosine DNA methylation has become one of the most important hotspots to research chromatin modification. Recent research has demonstrated that the disturbance of DNA methylation patterns may consequently have functional consequences in organisms with this epigenetic code (Dyachenko et al. 2014; Kumari et al. 2013). Moreover, numerous studies found that cytosine DNA methylation plays a significant role in various cellular activities, including plant response to the environmental stresses (Wang et al. 2016a), embryonic development (Zhang et al. 2016), cell differentiation (Hassan-Zadeh et al. 2017), inactivation of chromatin (Keown et al. 2017), and plant growth and development (Yang et al. 2015).
Heterosis is an extremely common phenomenon that refers to the superior average performance of hybrids over their parents with respect to various agronomic traits, first submitted by Shull (1908, 1952). Although this phenomenon has been widely exploited to increase agronomic production with ensuing economic and societal benefits for well over a century, the molecular mechanisms underlying heterosis remain poorly understood and mainly focused on dominance or over-dominance hypothesis (Birchler et al. 2010; Hochholdinger and Hoecker 2007). Recently, numerous studies have obtained lots of valuable research results about the relationship between DNA methylation and heterosis (Sun et al. 2015; Kawanabe et al. 2016). In addition, some papers also reported that the DNA methylation pattern of F1 offspring experienced big changes or adjustments to coordinate the expression of heterogeneous genes derived from parents, then made some genes efficiently transcript (Wang et al. 2016a, b; Li et al. 2013).
With the rapid development of biotechnology, there are many techniques to detect the DNA methylation level, such as bisulfite sequencing (Hernandez et al. 2013), HPLC (Cappetta et al. 2015), and Bisulfite Genomic Sequencing (Garg et al. 2015). However, these methods may accomplish higher costs or lower detection efficiency of DNA methylation sites. The methylation-sensitive amplification polymorphism (MSAP) technique is base on digestion with methylation-sensitive restriction endonucleases followed by amplification of restriction fragments and has been applied in various topics, including biotic and abiotic stress (Luo et al. 2016; Wang et al. 2016a), development (Wang et al. 2016b), differentiation of ecotypes (Xia et al. 2016) and somaclonal variability (Baránek et al. 2016), which can not only reduce costs but also generate broader coverage to discover key methylated sites. In addition, this method has also been used in many plant genomes and obtained lots of valuable results, such as Arabidopsis thaliana (Li et al. 2015), wheat (Venetsky et al. 2015), cotton (Wang et al. 2016a), Sorghum bicolor (Zhang et al. 2011), rice (Xia et al. 2016), and maize (Sun et al. 2015). Although the MSAP method has been investigated in many common crops, few studies have focused on heterosis in soybean.
In this study, the MSAP technique based on capillary electrophoresis was used to compare differences in cytosine methylation patterns and levels of 12 reciprocal soybean hybrids and their parents based on leaves of 15-day emergence. A total of 1239 fragments were detected in each sample on average, and the DNA cytosine methylated level of all reciprocal hybrids was remarkably lower than mid-parent heterosis (MPH). In addition, correlation coefficients between 12 traits and 16 subgroups methylation pattern were calculated to further analyze the relationship between DNA methylation variation and heterosis. These results obtained in our study would provide more theoretical basis for soybean genetic breeding.
Materials and methods
Plant materials
To determine the molecular mechanisms of heterosis at the epigenetic level in soybean, a total of four cultivars [Jilin 47 (no. 19) and Jilin 48 (no. 3) were cultivated by Jilin Academy of Agricultural Sciences in China; EXP (no. 12) and Yi 3 (no. 6) were imported from abroad] were used here. Meanwhile, the four cultivars were designated as females or males in accordance with complete diallel cross to generate 12 F1 hybrids in 2013, including Jilin 38 × Yi 3(3 × 6), Jilin 38 × EXP(3 × 12), Jilin 38 × Jilin 47(3 × 19), Yi 3 × Jilin 38(6 × 3), Yi 3 × EXP(6 × 12), Yi 3 × Jilin 47(6 × 19), EXP × Jilin 38(12 × 3), EXP × Yi 3(12 × 6), EXP × Jilin 47(12 × 19), Jilin 47 × Jilin 38(19 × 3), Jilin 47 × Yi 3(19 × 6), Jilin 47 × EXP(19 × 12). In 2014 spring, four parents and 12 hybrids were all sown in the Jilin Agricultural University fields, with 2 m long and 2 rows per plot with 3 replicates.
Agronomic and quality traits analysis
A total of five plants every plot were randomly taken out from the parents and hybrids harvested in 2014 autumn to analyze the agronomic traits, which included plant height (cm), node number, branch number, height of low pod (cm), pod number per plant, grain weight per plant (g), insect food grain rate (%), hundred-grain weight (g), grain number per plant, stems thick (mm). Moreover, the quality of traits, protein (%) and fat (%), were analyzed by Near Infrared Spectroscope (Model N 500, BUCHI, Swiss). Then, the estimation of heterosis was obtained by calculating the mid-parent heterosis (MPH) and over-parent heterosis (BPH). However, it was noteworthy that the insect grain rate should be taken negative over-parent heterosis.
Genomic DNA isolation
After 15-day emergence, the leaves of 16 accessions were collected with 3 replicates. Then the DNA of 48 samples was extracted respectively by a modified CTAB method (Kidwell and Osborn 1992). The DNA was purified by phenol extractions, and checked for quality and quantity by agarose gel electrophoresis and spectrometric measurement (Supplementary file 1A). In order to analysis the uniformity or variation of methylation alterations among different individuals, genomic DNA was isolated from the same stage.
MSAP analysis of DNA methylation
The methylation sensitive amplified polymorphism (MSAP) analysis method was performed as reported (Sun et al. 2015; Salmon et al. 2005). Two combinations of restriction enzymes were used by mixing EcoRI with each of the isoschizomers, HpaII and MspI, which can recognize the same sequence (5′-CCGG) and cut with differential sensitivity to DNA methylation of internal or external cytosine. HpaII can recognize the hemi-methylated external cytosine sites, while MspI can recognize full-methylated internal cytosine sites. Therefore, if HpaII can cut while MspI cannot cut for same sequence (5′-CCGG), recorded (1, 0), on the contrary we recorded (0, 1); if HpaII and MspI can cut at the same sites, recorded (1, 1). In another case, when the hemi-methylated external cytosine sites, or the full-methylated cytosine sites throughout the inside and outside existed, HpaII and MspI had no bands were showed, recorded (0,0). Because this situation is more complex, uncertain, this type of methylation band information was ignored.
A total of 50 μl reaction liquid was digested in 37 °C incubator for 2 h, which contains 5 μl T4 10× reaction buffer, 10 ng BSA, 2 U EcoRI, 2 U HpaII/MspI, 150 ng DNA, and ddH2O. The effect of digestion was detected by the electrophoresis (Supplementary file 1B). Subsequently, one pair adaptor (HpaII/MspI adaptor and EcoRI adaptor) were used in ligation reactions, which was consisted of 1 μl EcoRI adaptor, 1 μl H/M adaptor, and 0.1 μl T4 ligase incubated in 16 °C for 4 h. A total of one pair pre-selective primers and 20 pairs of selective primers were used for amplification and the sequences of adaptor and primers were all list in Supplementary file 2. The restriction enzymes EcoRI, HpaII and MspI were purchased from the Takara Biotech companies in Japan.
A total of 5 μl of each ligated sample, diluted 5-fold with sterilized distilled water, was used for the pre-amplification reactions. The PCR reactions conditions were: 94 °C for 45 s, 56 °C for 45 s and 72 °C for 1 min for 30 cycles. The pre-amplified products were displayed in the Supplementary file 1C. Selective amplification reactions were done with 5 μl of the pre-amplified cDNA that had been diluted 20-fold, using the following touchdown PCR conditions: 94 °C for 5 min, 94 °C for 30 s, 65 °C for 30 s (− 1 °C per cycle) and 72 °C for 1 min for 10 cycles, followed by 94 °C for 30 s, 56 °C for 30 s, and 72 °C for 1 min for 35 cycles. In order to confirm the successful of the capillary electrophoresis, the products of selective amplification was also detected by agarose gel electrophoresis (Supplementary file 1D).
Capillary electrophoresis analysis
Products of the selective amplification were denatured and analyzed on the Applied Biosystems 3730 XL Genetic Analyzer (Thermo Fisher, US) equipped with a 50 cm, 96 capillary electrophoresis. Reactions were carried out in 96-well reaction plates (Applied Bio-Systems), which contained 1 μl diluted sample, 8.5 μl Hi-Di™ formamide (Applied Bio-Systems) and 0.5 μl gene scan marker. The samples were denatured 3 min at 95 °C, then injected for 5 s at 3 V and electrophoresed for 50 min in Performance Optimized Polymer (POP-7™) at 60 °C. After that, ABI Foundation Data Collection Software version 3.0 was used to collect data, and analyzed by the GeneMapper version 4.0 (Applied Bio-Systems).
Results and discussion
Heterosis analysis of 12 hybrids
In order to analysis the heterosis, 12 traits of hybrids and parents were detected to analyze mid-parent heterosis (MPH) and over-parent heterosis (BPH). The results showed that different traits in different combinations displayed different heterosis. The MPH value of 12 traits was showed in Table 1, in which the MPH value of low pod height, insect rate, and fat were negative in all combinations; protein content in Jilin 38 × Jilin 47(3 × 19) hybrid offspring displayed negative; and the other traits in all combinations were positive. In addition, the MPH value of pod number, grain weight and number of per plant was more than 100%, in which grain weight in EXP × Yi 3(12 × 6) hybrid offspring displayed significant higher, reached 534.44%. In summary, we concluded that a total of four hybrids [EXP × Yi 3(12 × 6), Jilin 38 × Jilin 47(3 × 19), Yi 3 × EXP(6 × 12), and Yi 3 × Jilin 38(6 × 3)] have significant MPH value according to the Table 1.
Moreover, the BPH value of 12 traits was showed in Table 2, in which 3 traits (low pod height, insect rate, and fat content) of BPH were negative; five traits (branch no., pod no., grain no., grain weight, and stems thick) of BPH in all combinations were positive; and there was both negative and positive in the other traits. A total of four traits (branch no., pod no., grain no., and grain weight) of BPH in some combinations were more than 100%, in which grain weight in EXP × Yi 3 (12 × 6) hybrid offspring displayed significant higher, reached 497.84%. Concluded above results, four hybrids [EXP × Yi 3(12 × 6), Jilin 38 × Jilin 47(3 × 19), Yi 3 × EXP(6 × 12), and Yi 3 × Jilin 47(6 × 19)] have significant BPH.
MSAP analysis in cytosine methylation levels, patterns among hybrids and their parental lines
Recently, various studies have evidenced that the epigenetic inheritance may vary in plant hybrids, and may be accompanied by extensive modifications in DNA methylation (Sun et al. 2015; Sanetomo and Hosaka 2011; He et al. 2010). The results of these studies suggest that detailed investigation of epigenetic regulation of critical loci in hybrid genomes may lead to a better understanding of the mechanisms underlying hybrid vigor. In addition, heterosis is manifested at the early seedling stage in hybrids (Joel and Zhang 2001), thus MSAP profiles were generated for 12 hybrids and their parents from leaves after 15-day post-emergence.
Differences in cytosine methylation levels between the hybrids and their parents
A total of 20 selective primer combinations were used to compare the status of cytosine methylation in 12 hybrids and their parents. The results showed that approximately 1239 fragments were detected in each sample on average, and the number of non-methylated sites, hemi-methylated external cytosines and full-methylated internal cytosines were calculated (Table 3) according to the results of capillary electrophoresis (Fig. 1). Base on the MSAP profiles, total relative methylation levels of all samples were 41.33–58.89%, of which 16.9–25.43% involved external cytosine hemi-methylation, and 21.35–35.97% corresponded to full-methylated internal cytosines in 5′-CCGG recognition sites. In addition, the DNA cytosine methylated level of all reciprocal hybrids were remarkably lower than MPH, which indicated that the methylation level of hybrids was significantly decreased corresponding to parental lines, and this result was in accordance with the findings of previous studies (Li et al. 2013; Zhao et al. 2007; Zhang et al. 2007). Many studies have also reported that DNA demethylation can reactivate gene expression (Zhu et al. 2015; Hsieh et al. 2009). Thus, we can conclude that the trend of decreased methylation in the hybrids compared with their parents may enable de-repression and possibly expression of many genes associated with phenotypic variation observed in the hybrids.
Moreover, the differences of DNA cytosine methylation levels in the reciprocal hybrids were 0.65–6.86%, of which the combination Jilin 38(3) and EXP (12), Jilin 38(3) and Jilin 47(19), EXP(12) and Jilin 47(19) displayed significantly high, 6.86, 4.01, and 4.44%, respectively, while other combinations were very low. Such significant differences of DNA methylation levels in different hybrids may be due to differences in plant materials.
Cytosine methylation patterns between the hybrids and their parents
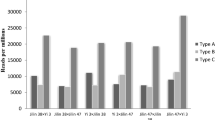
To further analysis cytosine methylation variation, the different cytosine methylation patterns between hybrids and their parental lines were observed, and then divided into four major classes (Table 4) according to previous reported (Sun et al. 2015; Zhao et al. 2008). Class A is the methylation pattern of hybrids completely inherited in their parents; class B is the methylation pattern of hybrids inherited in one of their parents, followed the Mendelian inheritance; class C is the methylation level of hybrids increased compared to their parents, called hyper-methylation pattern; class D is the methylation level of hybrids decreased compared to their parents, called hypo-methylation pattern. Four classes were further divided into 16 subgroups based on epigenetic inheritance patterns and alteration in cytosine methylation between parents and hybrids (Table 4).
A comparative analysis revealed that the number of class B was significantly higher than other classes in all hybrids comparing their parents. The second was the class A. The number of class C was the lowest, but there was a special hybrid [Jilin 38 × Jilin 47(3 × 19)] that the number of class C was higher than class D. Therefore, the results revealed that the inheritance of methylation (class A and class B) was higher than variation of methylation (class C and class D). Meanwhile, inheritance of methylation from one of parents (class B) was higher than both parents (class A); hypo-methylation (class D) was higher than hyper-methylation (class C). Liu and Wendel have reported that DNA methylation pattern of hybrids may be altered, but it will generally be genetic from parent to offspring when they researched on the epigenetic evolution of plant allopolyploids (Liu and Wendel 2003). Meanwhile, Wang reported that the inherited and altered methylation patterns were also present in cotton hybrid, and the inherited patterns was significantly higher than altered pattern (Wang et al. 2016a, b), which results are basically the same as our results. Therefore, our results indicated that methylation sites of hybrids inherited in the parents may be associated with some traits for plant growth and development, which was also consistent with other study (Sakthivel et al. 2010). In addition, the higher level hypo-methylation (class D) in our study may suggest that it is benefit for explaining hybrid-specific gene expression, as also indicated by Sakthivel et al. (2010).
The correlationship between heterosis and DNA methylation variation
In order to further analyze the relationship between heterosis and DNA methylation in soybean, the data of Tables 1, 2 and 3 was integrated for obtaining Table 5. The results showed that hemi-methylation level of two hybrids [Jilin 38 × Jilin 47(3 × 19), Yi 3 × Jilin 47(6 × 19)] that used 19 as father and full-methylation level of two hybrids [EXP × Yi 3(12 × 6), EXP × Jilin 47(12 × 19)] that used 12 as mother was higher than MPH (Tables 3, 5). Moreover, the four hybrids displayed great heterosis (Table 5). In addition, we also found that the total relative methylation levels of hybrids that performed significant heterosis showed a downward trend, but had different methylation patterns. Three patterns were observed: (1) hemi-methylation and full-methylation level all decreased, such as Yi 3 × EXP(6 × 12) and Yi 3 × Jilin 38(6 × 3); (2) hemi-methylation level decreased, and full-methylation level increased, such as EXP × Yi 3(12 × 6) and EXP × Jilin 47(12 × 19); (3) hemi-methylation level increased, and full-methylation level decreased, such as Jilin 38 × Jilin 47(3 × 19) and Yi 3 × Jilin 47(6 × 19).
In addition, correlation coefficients between heterosis and DNA methylation patterns were calculated (Tables 6, 7). The results of correlation analysis in MPH showed that there was only six correlation coefficients displayed significant correlation (P < 0.05) (Table 6), for instance, node number and the bottom pod height showed a significant negative correlation with C2 (− 0.62* and − 0.58*, respectively); and the node number displayed a significant correlation with D4 (0.69*); the hundred-grain weight performed a significant correlation with B1 (0.60*); the stem thick showed a significant correlation with C1 (0.66*), and a significant negative correlation with D1 (− 0.58*). In addition, the results of correlation analysis in BPH showed that there was only 3 correlation coefficients displayed significant correlation (P < 0.05) (Table 7), including the height of plants correlated to B5 (− 0.60*), the stem thick correlated to C1 (0.68*) and D1 (− 0.60*). These results in our study indicated that the hypo-methylation play an enhancement role in increasing node number in hybrid, while hyper-methylation play a promotion role for increasing stem thick in hybrid. Therefore, we can preliminary infer that there is an important relationship between DNA methylation variation of leaves and heterosis in soybean hybrid. However, to completely understand this relationship, a comprehensive analysis of the genetic and epigenetic regulation of 234 gene expression variation in hybrid should be further researched just as suggested by He et al. (2013).
References
Baránek M, Čechová J, Kovacs T, Eichmeier A, Wan S, Raddová J, Nečas T, Ye X (2016) Use of combined MSAP and NGS techniques to identify differentially methylated regions in somaclones: a case study of two stable somatic wheat mutants. PLoS ONE 11(10):e0165749
Birchler JA, Yao H, Chudalayandi S, Vaiman D, Veitia RA (2010) Heterosis. Plant Cell 22:2105–2112
Cappetta M, Berdasco M, Hochmann J, Bonilla C, Sans M, Hidalgo PC, Artagaveytia N, Kittles R, Martinez M, Esteller M, Bertoni B (2015) Effect of genetic ancestry on leukocyte global DNA methylation in cancer patients. BMC Cancer 15(1):434
Dyachenko OV, Tarlachkov SV, Marinitch DV, Shevchuk TV, Buryanov YI (2014) Expression of exogenous DNA methyltransferases: application in molecular and cell biology. Biochemistry 79(2):77–87
Garg R, Chevala VN, Shankar R, Jain M (2015) Divergent DNA methylation patterns associated with gene expression in rice cultivars with contrasting drought and salinity stress response. Sci Rep 5:144922. https://doi.org/10.1038/srep14922
Hassan-Zadeh V, Rugg-Gunn P, Bazett-Jones DP (2017) DNA methylation is dispensable for changes in global chromatin architecture but required for chromocentre formation in early stem cell differentiation. Chromosoma 126(5):605–614. https://doi.org/10.1007/s00412-017-0625-x
He GM, Zhu XP, Elling AA, Chen LB, Wang XF, Guo L, Liang MZ, He H, Zhang HY, Chen FF, Qi YJ, Chen RS, Deng XW (2010) Global epigenetic and transcriptional trends among two rice subspecies and their reciprocal hybrids. Plant Cell 22:17–33
He GM, He H, Deng XW (2013) Epigenetic variations in plant hybrids and their potential roles in heterosis. J Genet Genomics 40:205–210
Hernandez HG, Tse MY, Pang SC, Arboleda H, Forero DA (2013) Optimizing methodologies for PCR-based DNA methylation analysis. Biotechniques 55(4):181–197
Hochholdinger F, Hoecker N (2007) Towards the molecular basis of heterosis. Trends Plant Sci 12:427–432
Hsieh TF, Ibarra CA, Silva P, Zemach A, Eshed-Williams L, Fischer RL, Zilberman D (2009) Genome-wide demethylation of Arabidopsis endosperm. Science 324(5933):1451–1454
Joel, Zhang (2001) Immune related genetic polymorphisms and schizophrenia among the Chinese. Hum Immunol 62(7):714–724
Kawanabe T, Ishikura S, Miyaji N, Sasaki T, Wu LM, Itabashi E, Takada S, Shimizu M, Takasa-Yasuda T, Osabe K, Peacock WJ (2016) Role of DNA methylation in hybrid vigor in Arabidopsis thaliana. Proc Natl Acad Sci USA 113(43):E6704–E6711
Keown CL, Berletch JB, Castanon R, Nery JR, Disteche CM, Ecker JR, Mukamel EA (2017) Allele-specific non-CG DNA methylation marks domains of active chromatin in female mouse brain. Proc Natl Acad Sci USA 114(14):E2882–E2890
Kidwell KK, Osborn TC (1992) Simple plant DNA isolation procedures. In: Beckman JS, Osborn TC (eds) Plant genomes: methods for genetic and physical mapping. Kluwer, Dordrecht, pp 1–13
Kumari R, Yadav G, Sharma V, Sharma V, Kumar S (2013) Cytosine hypomethylation at CHG and CHH sites in the pleiotropic mutants of Mendelian inheritance in Catharanthus roseus. J. Genet. 92(3):499–511
Li A, Song WQ, Chen CB, Zhou YN, Qi LW, Wang CG (2013) DNA methylation status is associated with the formation of heterosis in Larix kaempferi intraspecific hybrids. Mol Breed 31(2):463–475
Li Z, Liu Z, Chen R, Li X, Tai P, Gong Z, Jia C, Liu W (2015) DNA damage and genetic methylation changes caused by Cd in Arabidopsis thaliana seedlings. Environ Toxicol Chem 34(9):2095–2103
Liu B, Wendel JF (2003) Epigenetic phenomena and the evolution of plant allopolyploids. Mol Phylogenet Evol 29(3):365–379
Luo JY, Pan XL, Peng TC, Chen YY, Zhao H, Mu L, Peng Y, He R, Tang H (2016) DNA methylation patterns of banana leaves in response to Fusarium oxysporum f. sp. cubense tropical race 4. J Integr Agric 15(12):2736–2744
Sakthivel K, Girishkumar K, Ramkumar G, Shenoy VV, Kajjidoni ST, Salimath PM (2010) Alterations in inheritance pattern and level of cytosine DNA methylation, and their relationship with heterosis in rice. Euphytica 175:303–314
Salmon A, Ainouche ML, Wendel JF (2005) Genetic and epigenetic consequences of recent hybridization and polyploidy in Spartina (Poaceae). Mol Ecol 14(4):1163–1175
Sanetomo R, Hosaka K (2011) Reciprocal differences in DNA sequence and methylation status of the pollen DNA between F1 hybrids of Solanum tuberosum × S. demissum. Euphytica 182:219–229
Shull GH (1908) The composition of a field of maize. Am Breed Assoc. Rep. 4:296–301
Shull GH (1952) Beginnings of the heterosis concept. In: Gowen JW (ed) Heterosis. A record of researches directed toward explaining and utilizing the vigor of hybrids. Iowa State College Press, Ames, pp 14–48
Sun LF, Liu TJ, Liu HK, Shan XH, Su SZ, Li SP, Yuan YP, Zhang J (2015) Analysis of DNA cytosine methylation patterns in maize hybrids and their parents. Biol Plantarum 59(2):266–272
Venetsky A, Levy-Zamir A, Khasdan V, Domb K, Kashkush K (2015) Structure and extent of DNA methylation-based epigenetic variation in wild emmer wheat (T. turgidum ssp. dicoccoides) populations. BMC Plant Biol 15(1):200
Wang B, Zhang M, Fu R, Qian X, Rong P, Zhang Y, Jiang P, Wang JJ, Lu XK, Wang D, Ye W, Zhu XY (2016a) Epigenetic mechanisms of salt tolerance and heterosis in Upland cotton (Gossypium hirsutum L.) revealed by methylation-sensitive amplified polymorphism analysis. Euphytica 208(3):477–491
Wang Y, Rong H, Xie T, Jiang J, Wu J (2016b) Comparison of DNA methylation in the developing seeds of yellow-and black-seeded Brassica napus through MSAP analysis. Euphytica 209(1):157–169
Xia H, Huang W, Xiong J, Tao T, Zheng X, Wei H, Yue Y, Chen L, Luo L (2016) Adaptive epigenetic differentiation between upland and lowland rice ecotypes revealed by methylation-sensitive amplified polymorphism. PLoS ONE 11(7):e0157810
Yang H, Chang F, You C, Cui J, Zhu G, Wang L, Zheng Y, Qi J, Ma H (2015) Whole-genome DNA methylation patterns and complex associations with gene structure and expression during flower development in Arabidopsis. Plant J 81(2):268–281
Zhang MS, Yan HY, Zhao N et al (2007) Endosperm-specific hypomethylation, and meiotic inheritance and variation of DNA methylation level and pattern in sorghum (Sorghum bicolor L.) inter-strain hybrids. Theor Appl Genet 115:195–207
Zhang M, Xu C, von Wettstein D, Liu B (2011) Tissue-specific differences in cytosine methylation and their association with differential gene expression in sorghum. Plant Physiol 156(4):1955–1966
Zhang T, Termanis A, Özkan B, Bao XX, Culley J, de Lima Alves F, Rappsilber J, Ramsahoye B, Stancheva I (2016) G9a/GLP complex maintains imprinted DNA methylation in embryonic stem cells. Cell Rep 15(1):77–85
Zhao XX, Chai Y, Liu B (2007) Epigenetic inheritance and variation of DNA methylation level and pattern in maize intra-specific hybrids. Plant Sci 172:930–938
Zhao X, Ruan Y, Wei C-L (2008) Tackling the epigenome in the pluripotent stem cells. J Genetics Genom 35(7):403-412
Zhu N, Cheng S, Liu X, Du H, Dai M, Zhou D-X, Yang W, Zhao Y (2015) The R2R3-type MYB gene OsMYB91 has a function in coordinating plant growth and salt stress tolerance in rice. Plant Sci 236:146–156
Acknowledgements
This study was supported by the Jilin Provincial Science Foundation of China (20140101015JC) and Jilin Provincial Key Scientific and Technological Project of China (20150204004NY).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: Jun Zhang and Piwu Wang. Performed the experiments: Yuanqian, Kaixin Zhang, Xiao Han and Shujie Fan. Analyzed the data: Lifang Sun, Xueying Li and Yiwei Qu. Contributed reagents/materials/analysis tools: Dan Yao. Wrote the paper: Kaixin Zhang, Yuanqiang Wang and Lifang Sun. All authors read and approved the final version of the manuscript.
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, Y., Zhang, K., Sun, L. et al. Study on the relationship between genetic variation of DNA methylation and heterosis in soybean leaves. Euphytica 214, 85 (2018). https://doi.org/10.1007/s10681-018-2161-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-018-2161-z