Abstract
Epiblast stem cells (EpiSCs), which are pluripotent cells isolated from early post-implantation mouse embryos (E5.5), show both similarities and differences compared to mouse embryonic stem cells (mESCs), isolated earlier from the inner cell mass (ICM) of the E3.5 embryo. Previously, we have observed that while chromatin is very dispersed in E3.5 ICM, compact chromatin domains and chromocentres appear in E5.5 epiblasts after embryo implantation. Given that the observed chromatin re-organization in E5.5 epiblasts coincides with an increase in DNA methylation, in this study, we aimed to examine the role of DNA methylation in chromatin re-organization during the in vitro conversion of ESCs to EpiSCs. The requirement for DNA methylation was determined by converting both wild-type and DNA methylation-deficient ESCs to EpiSCs, followed by structural analysis with electron spectroscopic imaging (ESI). We show that the chromatin re-organization which occurs in vivo can be re-capitulated in vitro during the ESC to EpiSC conversion. Indeed, after 7 days in EpiSC media, compact chromatin domains begin to appear throughout the nuclear volume, creating a chromatin organization similar to E5 epiblasts and embryo-derived EpiSCs. Our data demonstrate that DNA methylation is dispensable for this global chromatin re-organization but required for the compaction of pericentromeric chromatin into chromocentres.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mouse embryonic stem cells (ESCs) and epiblast stem cells (EpiSCs) are both pluripotent cells capable of self-renewal and differentiation into all cell types of the three germ layers upon receiving the appropriate signals. These pluripotent cells not only share similarities but also exhibit differences such as colony morphology, gene expression profile, epigenome, media requirement and signalling pathways involved in maintaining the pluripotent state. ESCs are isolated from the inner cell mass (ICM) at embryonic day 3.5 but EpiSCs are isolated later in development from the epiblast of the post-implantation embryo at day 5.5 (Brons et al. 2007; Tesar et al. 2007). EpiSCs, similar to ESCs, have a broad differentiation capacity in vitro and in vivo. They can make teratomas when injected into immunodeficient mice and give rise to derivatives of all three germ lines in vitro. However, unlike ESCs, chimera production from these pluripotent cells is impaired (Brons et al. 2007; Tesar et al. 2007).
Whereas mouse ESCs grow as small, compact and rounded colonies in vitro, EpiSC colonies are large, flat and monolayer (Brons et al. 2007; Tesar et al. 2007). Both express core pluripotency factors, namely Nanog, Sox2 and Oct4, but EpiSCs lose the expression of ESCs markers such as Rex1/Zfp42, Dppa3/Stella and Dax1 and instead, highly express genes such as Cer1, Sox17 and Fgf5 (Brons et al. 2007; Tesar et al. 2007). EpiSCs rely on activin/nodal and Fgf2 pathways to maintain the pluripotency state (Brons et al. 2007; Tesar et al. 2007), whereas ESCs require signalling mediated by leukaemia inhibitory factor (LIF) for pluripotency maintenance (Williams et al. 1988; Niwa et al. 1998). Murine ESCs and EpiSCs are inter-convertible in vitro (Bao et al. 2009), allowing for studies of the molecular and epigenetic changes correlated with and/or required for this transition.
We have demonstrated the power of electron spectroscopic imaging (ESI) for examining chromatin and nuclear architecture. ESI is an electron microscopy technique well suited to visualize chromatin fibres with high resolution and contrast without the requirement of heavy atom contrast agents (Bazett-Jones and Hendzel 1999; Dellaire et al. 2004; Bazett-Jones et al. 2008; Ahmed et al. 2009). Moreover, ESI can be used to study the spatial relationships of chromatin to protein- and ribonucleoprotein (RNP)-based structures by nitrogen and phosphorus mapping. With ESI, we have previously shown that there is a notable difference in chromatin organization between cells in the ICM and epiblast cells after embryo implantation in vivo. In the ICM, chromatin is uniformly dispersed, whereas in late epiblasts, small blocks of chromatin are visible within the nucleus and along the nuclear envelope. Interestingly, we also observed that embryo-derived ESCs and EpiSCs show a chromatin organization similar to ICM and epiblasts, respectively (Ahmed et al. 2010). In this study, we first asked whether this chromatin re-organization can be reproduced during early stem cell differentiation in vitro and whether maintenance and de novo DNA methylation play a role in the organization of the global and/or pericentromeric chromatin during this process.
In mammals, DNA methylation is mediated by three DNA methyltransferases, which are essential for embryo development (Li et al. 1992; Okano et al. 1999; Smith and Meissner 2013). Dnmt1, the first mammalian DNA methyltransferase to be discovered, has an affinity for hemimethylated DNA and maintains DNA methylation status during cell division (Bestor et al. 1988; Leonhardt et al. 1992; Pradhan et al. 1999). De novo methylation is carried out by the Dnmt3 family with two catalytically active members, Dnmt3a and Dnmt3b, which are expressed in ESCs and methylate unmethylated sites during differentiation (Lei et al. 1996; Okano et al. 1998; Hsieh 1999). During development, between embryonic days 4.5 and 6.5, a wave of de novo methylation occurs in the epiblasts (Borgel et al. 2010; Smith et al. 2012; Auclair et al. 2014) and this coincides temporally with chromatin re-organization (Ahmed et al. 2010), but the extent to which both processes are linked is unknown. In order to answer this question, wild-type and DNA methylation-deficient ESCs were used and converted to EpiSCs. Male ICM-derived ESCs cultured in serum are hypermethylated compared to the ICM cells (Habibi et al. 2013) and display a global level of DNA methylation similar to post-implantation epiblasts and EpiSCs (Senner et al. 2012; Habibi et al. 2013). However, when ESCs are converted to EpiSCs in vitro, there is a re-distribution of DNA methylation and an increased prevalence of hypermethylated promoters in EpiSCs has been observed (Veillard et al. 2014). Here, we aimed to determine whether this re-distribution of DNA methylation plays a role in the chromatin re-organization that occurs during ESC to EpiSC conversion. Our results show that EpiSC-like chromatin organization could be re-capitulated in vitro, and DNA methylation is dispensable for global chromatin organization during the transition of ESCs to EpiSCs. Interestingly, however, we observed that in the absence of DNA methylation, the organization of pericentromeric chromatin into chromocentres is affected.
Experimental procedures
ES cell culture and EpiSC conversion
Wild-type (J1) and mutant ESCs derived from the J1 background were cultured in DMEM high glucose (Invitrogen, 11965-092) media supplemented with 15% foetal bovine serum (ES cell qualified, Invitrogen, 10439-024), 1% penicillin-streptomycin-glutamine (Invitrogen, 10378-016), 2 mM l-glutamine (Invitrogen, 25,030–081), 0.1 mM MEM NEAA (Invitrogen, 11140-050), 1 mM sodium pyruvate (Invitrogen, 11,360–070), 0.1 mM mercaptoethanol (Invitrogen, 21985-023) and 1000 U/ml LIF (chemicon, ESG1107) on mouse embryo fibroblasts (MEFs). The double knockout [Dnmt3a −/− , Dnmt3b −/−] and the triple knockout [Dnmt1 −/− , Dnmt3a −/− , Dnmt3b −/−] ES cells were kindly provided by Dr. Masaki Okano (Okano et al. 1999; Tsumura et al. 2006).
After three passages in ES condition, cells were washed and then 1.5 × 105 cells were seeded on MEF-coated cover-slips in EpiSC media. EpiSC media contain 50% F-12 nutrient mixture (Invitrogen 31765-035), 50% Iscove’s modified Dulbecco’s medium (Invitrogen 31980-030), 5 mg/ml BSA fraction V (Sigma A1470), 450 μM 1-thioglycerol (Sigma M7522), 7 μg/ml insulin (Roche 11376497001), 15 μg/ml transferrin (Roche 10652202001), 20 ng/ml activin A and 12 ng/ml Fgf2 (Brons et al. 2007). Media was changed daily up to 7 days, and cells were then fixed.
Immunostaining, immunofluorescence microscopy and ESI
Cells were prepared for immunostaining as previously described by Ahmed et al. (2009). They were then fixed with 2% freshly diluted paraformaldehyde (Electron Microscopy Sciences), rinsed and permeabilized with 1% Triton X-100-PBS. Permeabilized cells were first immunolabelled with rabbit anti-H3K9me3 (1:500) and then with a donkey anti-rabbit antibody (1:300, Jackson Laboratories) combined with Cy3.
After immunolabelling, cells were fixed again with 1% glutaraldehyde (Electron Microscopy Sciences) and dehydrated gradually with 30, 50, 70, 90, and 100% ethanol. Dehydrated cells were embedded in Quetol 651 resin (Electron Microscopy Sciences) and sectioned using an ultramicrotome (Leica) 70 nm as described previously (Dellaire et al. 2004; Ahmed et al. 2009). Sections on grids were first imaged by a fluorescence microscope (leica), and then phosphorus and nitrogen images were generated using a transmission electron microscope (Tecnai 20, FEI) and collected by the digital micrograph software. Images were processed with Photoshop 7.0 (Adobe). Nitrogen signal level in chromatin structures was decreased to zero so the non-chromatin proteins are more visible. The phosphorus images were coloured in yellow, the processed nitrogen images were coloured in blue and both images were overlaid. Cell images taken by the transmission electron microscope were then overlaid with the same cell’s image taken by the fluorescence microscope. The correlative light and electron microscopic imaging approach involves many preparative, imaging and analysis steps and thus is not a high-throughput approach. However, we sampled as many cells and regions of interest as practically possible while guaranteeing sufficient statistical power. Wild-type (WT) ESC to EpiSCs differentiation experiment was carried out four times, double knockout (DKO) ESC to EpiSCs differentiation two times and triple knockout (TKO) ESC to EpiSCs differentiation was performed three times. For WT EpiSCs, nine sections; for DKO, EpiSCs five sections; and for TKO EpiSCs, eight sections were examined. In total, 10 WT ESCs, 8 DKO ESCs, 6 TKO ESCs, 41 WT ESC-derived EpiSCs, 14 DKO ESC-derived EpiSCs and 22 TKO ESC-derived EpiSCs were imaged. Nineteen WT ESC-derived EpiSCs and 9 TKO ESC-derived EpiSCs were labelled with antibodies against H3K9me3 marks.
Cluster analysis
The average size of the chromatin clusters was measured to provide a quantitative indication of the degree of compaction observed in the electron micrographs. Average cluster sizes were measured with ImageJ. Net phosphorus maps were binarized and measured using the particle size measure algorithm from 100 pixels to infinity. The lower cutoff of 100 prevented the inclusion of pixel noise or small RNP granules from being counted. Pixel size was converted to square nanometres.
DNA methylation analysis
One microgramme genomic DNA from WT, DKO and TKO ESCs and ESC-derived EpiSCs was digested with HpaII and MspI restriction enzymes. The digested DNA was analysed on 1% agarose gel.
Gene expression analysis
RNA extraction, cDNA synthesis and quantitative real-time PCR were carried out as described previously (Novo et al. 2016). The primer sequences are shown in the Supplementary Table S1. The results were normalized using Hmbs and displayed relative to WT ESCs.
Results
DNA methylation-deficient ESCs form EpiSC-like colonies in vitro
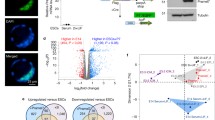
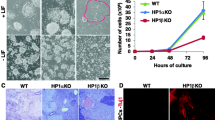
We first converted wild-type ESCs to EpiSCs by removing LIF and culturing cells in an EpiSC media for 7 days (see the “Experimental procedures” section). After 7 days in EpiSC media, colonies with a flattened, epithelial morphology appeared that is characteristic of EpiSC (Fig. 1a). We then converted de novo DNA methylation-deficient ESCs (Dnmt3a −/− , Dnmt3b −/−, DKO) and maintenance and de novo DNA methylation-deficient ESCs (Dnmt1 −/− , Dnmt3a −/− , Dnmt3b −/−, TKO) to EpiSCs (Fig. 1a). Both DNA methylation-deficient ES cell types formed EpiSC-like colonies similar to WT ESCs. Gene expression analyses show that after 7 days in EpiSC media, the transcript levels of ESC markers Klf2, Nanog, Nr5a2 and Zfp42/Rex1 are downregulated and the expression of EpiSC markers Cer1, Fgf5, Lefty1 and T are upregulated in the wild-type and DNA methylation-deficient ESC-derived EpiSCs compared to wild-type ESCs (Fig. 1b). To be certain that DNA methylation is indeed decreased in DKO and TKO ESC-derived EpiSCs, we digested genomic DNA from WT, DKO and TKO ESCs and ESC-derived EpiSCs with HpaII, an enzyme sensitive to CpG methylation, and MspI which is insensitive to DNA methylation. As expected, HpaII cuts the genomic DNA in the two DNA methylation-deficient ESC and EpiSC lines but its activity is prevented by CpG methyl-ation in the wild-type cells (Fig. 1c). These results show that, as expected, DNA methylation is reduced in DNA methylation-deficient ESCs and ESC-derived EpiSCs.
EpiSC-like colonies appear after 7 days in EpiSC media. a ESC and ESC-derived EpiSC colonies for WT, DKO (Dnmt3a −/− , Dnmt3b −/−) and TKO (Dnmt1 −/− , Dnmt3a −/− , Dnmt3b −/−) are displayed. b The transcript levels of the ESC and EpiSC marker Pou5f1/Oct-4, the ESC markers Klf2, Nanog, Nr5a2, and Zfp42/Rex1 and EpiSC markers Cer1, Fgf5, Lefty1 and T were quantified in WT and DNA methylation-deficient ESC-derived EpiSCs by qPCR. Quantitative polymerase chain reactions were performed in triplicates; values were normalized to Hmbs and displayed relative to WT ESC. Error bars represent standard deviation. ESC-derived EpiSCs show upregulation of EpiSC markers and downregulation of ESC markers. c DNA digestion profiles in WT, DKO and TKO ESCs (left panel) and ESC-derived EpiSCs (right -panel). Lanes 1, 4 and 7 are undigested WT, DKO and TKO genomic DNA, respectively. Lanes 2, 5 and 8 correspond to DNA digested by HpaII in WT, DKO and TKO cells, respectively. Lanes 3, 6 and 9 correspond to DNA digested by MspI. The minus sign denotes undigested DNA. ‘H’ and ‘M’ design DNA digested with HpaII and MspI, respectively
Chromatin is widely dispersed in both wild-type and DNA methylation-deficient ESCs
It has been shown that DNA methylation-deficient ESCs retain ES cell morphology, self-renew and remain able to express the pluripotency factor Oct4 (Okano et al. 1999; Tsumura et al. 2006). Tsumura et al. have shown by immunofluorescence microscopy that the absence of DNA methylation does not impede the formation of chromocentres (Tsumura et al. 2006). Here, we examined the chromatin organization of the DNA methylation-deficient ESCs with high resolution by ESI. Using wild type, the double (DKO) and triple knockout (TKO) ESC lines, we observe that the dispersed chromatin fibre architecture, typical of ESCs, is retained in the DKO and TKO ESC lines, indistinguishable from the wild-type ESCs (Fig. 2, compare upper panels in a, b and c). Besides the widely dispersed chromatin throughout the nucleoplasm, little compaction occurs along the nuclear envelope. The low degree of compaction is best illustrated with quantitative cluster analysis. This method measures the average size of the chromatin domains. The widely dispersed chromatin seen in ESCs would yield a lower average cluster size than that obtained from compact chromatin domains observed in EpiSCs. The cluster analysis results show the same degree of low compaction for all three ESC lines (1717, 1703 and 1548 nm2 for WT, DKO and TKO, respectively). We conclude that the loss of DNA methylation in the DKO and TKO lines leads to no structural consequences under these ESC growth conditions.
DNA methylation is dispensable for chromatin re-organization in ESC-derived EpiSCs. Low-magnification mass-sensitive ESI electron micrograph of a WT ESC and WT ESC-derived EpiSC (a), DKO ESC and DKO ESC-derived EpiSC (b) and TKO ESC and TKO ESC-derived EpiSC (c) are shown on the left. For all indicated fields, phosphorus maps representing nucleic acids (yellow) are merged with phosphorus subtracted nitrogen maps representing proteins (blue) and displayed on the right (see the Experimental procedures section). Scale bar represents 2 μm in the low-magnitude images and 500 nm in the element maps
Re-organization of global chromatin in wild-type EpiSCs
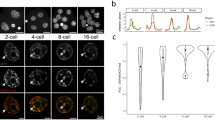
Pluripotent ESCs display a chromatin architecture of widely dispersed 10 nm fibres. Domains of compact chromatin are not observed, even along the nuclear envelope or in domains enriched for heterochromatin marks such as H3K9me3 (Ahmed et al. 2010; Fussner et al. 2010; Fussner et al. 2011). In contrast, the nuclear envelope and chromocentres represent regions of densely packed 10-nm chromatin fibres in differentiated cell types (Fussner et al. 2010; Fussner et al. 2011). We next examined the chromatin organization in WT ESC-derived EpiSCs. We observed a significant change in chromatin organization after 7 days in EpiSC media, a pattern similar to that observed in the E5 epiblasts and to embryo-derived EpiSCs (Ahmed et al. 2010). Instead of the uniformly dispersed chromatin, domains of compact chromatin are observed (Fig. 2a, lower panel). The average cluster size of the chromatin in WT ESC-derived EpiSCs is 8702 nm2. This value is five times the one observed in WT ESCs (1717 nm2). Hence, chromatin compaction taking place during implantation in vivo can be re-capitulated in vitro in 7 days.
DNA methylation is dispensable for global EpiSC-like chromatin organization
ESI imaging revealed that de novo methylation-deficient ESC-derived EpiSCs have a similar chromatin organization as seen in wild-type ESC-derived EpiSCs. Cluster analysis results show an average cluster size of 9421 nm2 for DKO ESC-derived EpiSCs, an increase of over 5-fold compared to ESCs and similar to that observed with the WT ESC-derived EpiSCs (8702 nm2, Fig. 2a, b). We conclude from this result that de novo DNA methyltransferases Dnmt3a and Dnmt3b are not required for chromatin re-organization during this transition. Double mutant EpiSCs still have the maintenance DNA methyltransferase Dnmt1. Next, we examined chromatin organization in triple knockout ESC-derived EpiSCs. Interestingly, we observed an EpiSC-like chromatin organization in TKO EpiSCs as well with an average cluster size of 10,009 nm2 (Fig. 2c), over 6-fold greater than that of the ESC line from which they were converted. This is in line with the values for the WT ESC-derived EpiSCs and DKO ESC-derived EpiSCs (Fig. 2a, b). We conclude that maintenance of DNA methylation is not a requirement for the global chromatin compaction that occurs with the ESC to EpiSC transition. Using a t test, the statistical significance of the greater than 5-fold increase in the cluster size distribution of the EpiSCs compared to the corresponding ESC lines is represented by a p value of 0.0001 (Supplementary Fig. S1).
DNA methylation is required for pericentromeric chromatin compaction
We then labelled the chromatin with H3K9me3 antibody to study the chromatin structure of particular domains such as chromocentres. Mouse chromocentres are clusters of pericentromeric repetitive DNA (major satellite DNA) characterized by constitutive heterochromatin (Guenatri et al. 2004; Almouzni and Probst 2011). They are enriched in H3K27me1, H3K9me3 and H4K20me3 chromatin marks (Peters et al. 2003; Schotta et al. 2004; Martens et al. 2005); heterochromatin protein 1 (HP1) (Lachner et al. 2001; Maison et al. 2002); and DNA methylation (Lehnertz et al. 2003). The correlation of fluorescence and ESI images allows us to focus on these structures and study their chromatin organization. In fibroblasts, using correlative light microscopy and ESI (LM/ESI), chromocentres appear as very compact chromatin, generally discrete or separated from other blocks of compact chromatin, usually with a high degree of radial symmetry and correspond to discrete foci of H3K9m3 (Rapkin et al. 2015). Here, we see that in labelled WT ESC-derived EpiSCs, some H3K9me3-enriched regions show characteristics typical of chromocentres, whereas others, while showing the high levels of H3K9me3, are not characterized by highly compact chromatin fibres. In fact, among 70 H3K9me3-enriched foci in 19 WT ESC-derived EpiSCs labelled with H3K9me3 antibody, 26 (37%) foci displayed morphological features characteristic of chromocentres (arrows in Fig. 3a), while 44 (63%) foci did not correlate with morphologically typical chromocentres (arrowheads in Fig. 3a; Supplementary Fig. S2). Interestingly, we observed that the absence of DNA methylation affects chromocentre integrity. In the labelled TKO ESC-derived EpiSCs, 62 H3K9me3-enriched regions were identified and only one region showed a chromocentre-like morphology (1.6%). The remaining 61 regions (98.4%) corresponded to dispersed chromatin (Fig. 3b; Supplementary Fig. S2). Chi-square test showed that the difference in terms of chromocentre morphology between the WT and DNA methylation-deficient ESC-derived EpiSCs was highly significant (χ 2 = 30.59, df = 1, p value < 0.005). We also failed to identify radially symmetric phosphorus-rich regions chara-cteristic of chromocentres when examining all the TKO ESC-derived EpiSC images.
Chromocentre formation is impaired in the absence of DNA methylation. Wild-type (a) and TKO (b) ESC-derived EpiSCs were labelled with H3K9me3 antibodies. Low-magnification mass-sensitive electron micrographs and fluorescence images, taken by ESI and a fluorescence microscope, respectively, were merged and displayed on the left (see the Experimental procedures section). Phosphorus (yellow) and nitrogen minus phosphorus maps (blue) of H3K9me3-enriched regions were merged and shown on the right. Arrows and arrowheads indicate compact and dispersed chromocentres, respectively. Scale bar represents 2 μm in the low-magnitude images and 500 nm in the element maps
Discussion
Embryonic stem cells and epiblast stem cells are transcriptionally and morphologically distinct and rely on different signalling pathways for pluripotency maintenance (Brons et al. 2007; Tesar et al. 2007). Recent data show that these two types of pluripotent cells are also epigenetically distinct (Jouneau et al. 2012; Sun et al. 2012; Murtha et al. 2015). We have previously shown that ESCs and embryo-derived EpiSCs differ in terms of global chromatin organization. Whereas chromatin is organized as highly dispersed 10 nm fibres in ESCs, numerous compact chromatin domains and chromocentre structures appear in embryo-derived EpiSCs similar to E5 epiblasts (Ahmed et al. 2010). Here, our imaging data reveals that the chromatin re-organization associated with this early differentiation can be re-capitulated in vitro. In fact, major alteration of chromatin architecture similar to E5 epiblasts and embryo-derived EpiSCs appear after 7 days in EpiSC media. The study of H3K9me3-enriched regions in wild-type ESC-derived EpiSCs shows that there are two types H3K9me3-enriched foci. Although in differentiated fibroblasts, all H3K9me3-enriched sites correspond to radially symmetrical isolated compact chromatin domains characteristic of chromocentres (Rapkin et al. 2015), while in the ESC-derived EpiSCs after 7 days in EpiSC media, only 37% of the labelled regions showed a compact chromatin state typical of chromocentres. The constitutive heterochromatin could be in a metastable state between decondensed and condensed, such that the sampling shows the two states at the instant that the cells are fixed. Alternatively, it may be that all chromocentres are not equivalent, perhaps depending on the chromosomes from which they are constituted. Some resist compaction longer in development than others.
After fertilization, the paternal and the maternal genomes undergo a wave of active and passive demethylation, respectively, which results in a hypomethylated genome in the ICM (Auclair and Weber 2012; Smith et al. 2012). Studies have shown that although the ICM-derived ESCs cultured in serum are hypermethylated compared to the ICM cells (Habibi et al. 2013), DNA methyltransferases are dispensable for their self-renewal (Lei et al. 1996; Okano et al. 1999; Jackson et al. 2004; Tsumura et al. 2006). We first asked whether the absence of DNA methylation affects the chromatin organization in self-renewing methylation-deficient ESCs. Tsumura et al. have previously shown by immunofluorescence analysis that TKO ES cells maintain DAPI- and H3K9me3-enriched regions similar to wild-type cells (Tsumura et al. 2006). However, this technique does not allow the visualization of the chromatin fibres at high resolution neither their distribution throughout the nucleus and around the nuclear envelop. Moreover, in contrast to ESI, the use of DAPI or even a heterochromatin histone mark does not reveal the state of compaction of the chromatin itself (Even-Faitelson et al. 2016). In this study, we compared the chromatin structure of the DKO and TKO ESCs with WT ESCs and could not observe any difference between these three cell lines confirming that DNA methylation is dispensable for ESC-like chromatin organization.
Around the time of implantation, a wave of de novo methylation occurs in vivo and the hypomethylated ICM genome becomes hypermethylated at distinct sites (Kafri et al. 1992; Borgel et al. 2010; Smith et al. 2012; Auclair et al. 2014). When ESCs are converted to EpiSCs in vitro, the same wave of de novo methylation takes place. Although, at a global level, DNA methylation is similar between male ESCs cultured in serum and EpiSCs (Senner et al. 2012; Habibi et al. 2013), there are differentially methylated regions. EpiSCs have more specific DNA methylated regions compared to ESCs, and many of these EpiSC-specific regions are promoters with high CpG content (Veillard et al. 2014). Veillard et al. showed that de novo methylation at promoters appears as early as 3 days after changing the culture media and correlates with the upregulation of Dnmt3b (Veillard et al. 2014). Here, we aimed to determine whether this re-distribution of DNA methylation plays a role in the chromatin re-organization that occurs during ESC to EpiSC conversion. Our results show that during ESC to EpiSC conversion, global chromatin re-organization can take place in the total absence of DNA methylation. The formation of chromocentres, however, is affected by DNA methylation deficiency. While numerous chromocentre-like structures are easily identifiable in WT ESC-derived EpiSCs, structures having the characteristic features of chromocentres are typically not observed in DNA methylation-deficient ESC-derived EpiSCs. In these cells, the majority of H3K9me3 foci (98.4%) correlate with very little chromatin compaction.
In general, DNA methylation exerts its effect in two ways. Cytosine methylation could hamper transcription factor binding at a target gene and prevent its activation. Alternatively, methylated CpGs could serve as docking sites for methyl DNA-binding proteins such as methyl CpG-binding protein 2 (MeCP2). In mouse cells, MeCP2 accumulates at chromocentres in a DNA methylation-dependent manner (Nan et al. 1996) and promotes pericentromeric heterochromatin clustering during differentiation (Brero et al. 2005; Bertulat et al. 2012). Interestingly, it has been shown that MeCP2 is able to bring chromatin fibres in close proximity and induce chromatin compaction (Georgel et al. 2003; Nikitina et al. 2007a; Nikitina et al. 2007b). Furthermore, the interaction of MeCP2 with various repressor factors such as the H3K9 trimethylating enzyme SUV39H1, HP1 and histone deacetylases re-inforce the state of chromatin repression (for review, see Della Ragione et al. 2016). One can hypothesize that in TKO ESC-derived EpiSCs, the absence of DNA methylation leads to MeCP2 depletion and dispersion of chromatin fibres at H3K9me3 foci. Further experiments are required to investigate whether MeCP2 is implicated in the observed absence of chromocentres.
Alternatively, changes in epigenetic marks at pericentromeric heterochromatin could account for the disappearance of morphologically characteristic chromocentres with high H3K9me3 enrichment in TKO ESC-derived EpiSCs. Indeed, Saksouk et al. have shown that in TKO ESCs, the loss of DNA methylation at pericentromeric heterochromatin opens the way for the recruitment of components of facultative heterochromatin such as polycomb repressive complex 2 (PRC2) and enrichment of H3K27me3 marks at these regions (Saksouk et al. 2014). One can imagine that part of the chromatin clusters we observe in TKO ESC-derived EpiSCs which are not labelled with H3K9me3 corresponds to clusters of major satellite marked by H3K27me3. A study by Gilbert et al. showed that the absence of de novo methyltransferases in mouse ESCs leads to increased clustering of major satellite DNA into large chromatin bodies (Gilbert et al. 2007), but in this study, the authors did not correlate H3K9me3 labelling with major satellite DNA clusters to clarify whether these bodies are bona fide chromocentres.
In summary, using ESI and correlative LM/ESI, we conclude that DNA methylation is required for chromocentre integrity but dispensable for global chromatin re-organization during early stem cell differentiation.
ESCs, embryonic stem cells; EpiSCs, epiblast stem cells; ICM, inner cell mass; ESI, electron spectroscopic imaging; DKO, double knockout; TKO, triple knockout
References
Ahmed K, Dehghani H, Rugg-Gunn P et al (2010) Global chromatin architecture reflects pluripotency and lineage commitment in the early mouse embryo. PLoS One 5:e10531. doi:10.1371/journal.pone.0010531
Ahmed K, Li R, Bazett-Jones DP (2009) Electron spectroscopic imaging of the nuclear landscape. Methods Mol Biol 464:415–423. doi:10.1007/978-1-60327-461-6_23
Almouzni G, Probst AV (2011) Heterochromatin maintenance and establishment: lessons from the mouse pericentromere. Nucleus 2:332–338. doi:10.4161/nucl.2.5.17707
Auclair G, Guibert S, Bender A, Weber M (2014) Ontogeny of CpG island methylation and specificity of DNMT3 methyltransferases during embryonic development in the mouse. Genome Biol 15:545–560. doi:10.1186/s13059-014-0545-5
Auclair G, Weber M (2012) Mechanisms of DNA methylation and demethylation in mammals. Biochimie 94:2202–2211. doi:10.1016/j.biochi.2012.05.016
Bao S, Tang F, Li X et al (2009) Epigenetic reversion of post-implantation epiblast to pluripotent embryonic stem cells. Nature 461:1292–1295. doi:10.1038/nature08534
Bazett-Jones DP, Hendzel MJ (1999) Electron spectroscopic imaging of chromatin. Methods 17:188–200. doi:10.1006/meth.1998.0729
Bazett-Jones DP, Li R, Fussner E et al (2008) Elucidating chromatin and nuclear domain architecture with electron spectroscopic imaging. Chromosom Res 16:397–412. doi:10.1007/s10577-008-1237-3
Bertulat B, De Bonis ML, Della Ragione F et al (2012) MeCP2 dependent heterochromatin reorganization during neural differentiation of a novel Mecp2-deficient embryonic stem cell reporter line. PLoS One 7:e47848. doi:10.1371/journal.pone.0047848
Bestor T, Laudano A, Mattaliano R, Ingram V (1988) Cloning and sequencing of a cDNA encoding DNA methyltransferase of mouse cells: the carboxyl-terminal domain of the mammalian enzymes is related to bacterial restriction methyltransferases. J Mol Biol 203:971–983
Borgel J, Guibert S, Li Y et al (2010) Targets and dynamics of promoter DNA methylation during early mouse development. Nat Genet 42:1093–1100. doi:10.1038/ng.708
Brero A, Easwaran HP, Nowak D et al (2005) Methyl CpG-binding proteins induce large-scale chromatin reorganization during terminal differentiation. J Cell Biol 169:733–743. doi:10.1083/jcb.200502062
Brons IG, Smithers LE, Trotter MW et al (2007) Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448:191–195. doi:10.1038/nature05950
Della Ragione F, Vacca M, Fioriniello S et al (2016) MECP2, a multi-talented modulator of chromatin architecture. Brief Funct Genomics 15:420–431. doi:10.1093/bfgp/elw023
Dellaire G, Nisman R, Bazett-Jones DP (2004) Correlative light and electron spectroscopic imaging of chromatin in situ. Methods Enzymol 375:456–478
Even-Faitelson L, Hassan-Zadeh V, Baghestani Z, Bazett-Jones DP (2016) Coming to terms with chromatin structure. Chromosoma 125:95–110. doi:10.1007/s00412-015-0534-9
Fussner E, Ahmed K, Dehghani H et al (2010) Changes in chromatin fiber density as a marker for pluripotency. Cold Spring Harb Symp Quant Biol 75:245–249. doi:10.1101/sqb.2010.75.012
Fussner E, Djuric U, Strauss M et al (2011) Constitutive heterochromatin reorganization during somatic cell reprogramming. EMBO J 30:1778–1789. doi:10.1038/emboj.2011.96
Georgel PT, Horowitz-Scherer RA, Adkins N et al (2003) Chromatin compaction by human MeCP2 assembly of novel secondary chromatin structures in the absence of DNA methylation. J Biol Chem 278:32181–32188. doi:10.1074/jbc.M305308200
Gilbert N, Thomson I, Boyle S et al (2007) DNA methylation affects nuclear organization, histone modifications, and linker histone binding but not chromatin compaction. J Cell Biol 177:401–411. doi:10.1083/jcb.200607133
Guenatri M, Bailly D, Maison C, Almouzni G (2004) Mouse centric and pericentric satellite repeats form distinct functional heterochromatin. J Cell Biol 166:493–505. doi:10.1083/jcb.200403109
Habibi E, Brinkman AB, Arand J et al (2013) Whole-genome bisulfite sequencing of two distinct interconvertible DNA methylomes of mouse embryonic stem cells. Cell Stem Cell 13:360–369. doi:10.1016/j.stem.2013.06.002
Hsieh CL (1999) In vivo activity of murine de novo methyltransferases, Dnmt3a and Dnmt3b. Mol Cell Biol 19:8211–8218
Jackson M, Krassowska A, Gilbert N et al (2004) Severe global DNA hypomethylation blocks differentiation and induces histone hyperacetylation in embryonic stem cells. Mol Cell Biol 24:8862–8871. doi:10.1128/MCB.24.20.8862-8871.2004
Jouneau A, Ciaudo C, Sismeiro O et al (2012) Naive and primed murine pluripotent stem cells have distinct miRNA expression profiles. RNA 18:253–264. doi:10.1261/rna.028878.111
Kafri T, Ariel M, Brandeis M et al (1992) Developmental pattern of gene-specific DNA methylation in the mouse embryo and germ line. Genes Dev 6:705–714
Lachner M, O’Carroll D, Rea S et al (2001) Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410:116–120. doi:10.1038/35065132
Lehnertz B, Ueda Y, Derijck AA et al (2003) Suv39h-mediated histone H3 lysine 9 methylation directs DNA methylation to major satellite repeats at pericentric heterochromatin. Curr Biol 13:1192–1200
Lei H, Oh SP, Okano M et al (1996) De novo DNA cytosine methyltransferase activities in mouse embryonic stem cells. Development 122:3195–3205
Leonhardt H, Page AW, Weier HU, Bestor TH (1992) A targeting sequence directs DNA methyltransferase to sites of DNA replication in mammalian nuclei. Cell 71:865–873
Li E, Bestor TH, Jaenisch R (1992) Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell 69:915–926
Maison C, Bailly D, Peters AH et al (2002) Higher-order structure in pericentric heterochromatin involves a distinct pattern of histone modification and an RNA component. Nat Genet 30:329–334. doi:10.1038/ng843
Martens JH, O’Sullivan RJ, Braunschweig U et al (2005) The profile of repeat-associated histone lysine methylation states in the mouse epigenome. EMBO J 24:800–812. doi:10.1038/sj.emboj.7600545
Murtha M, Strino F, Tokcaer-Keskin Z et al (2015) Comparative FAIRE-Seq analysis reveals distinguishing features of the chromatin structure of ground state-and primed pluripotent cells. Stem Cells 33:378–391. doi:10.1002/stem.1871
Nan X, Tate P, Li E, Bird A (1996) DNA methylation specifies chromosomal localization of MeCP2. Mol Cell Biol 16:414–421
Nikitina T, Ghosh RP, Horowitz-Scherer RA et al (2007a) MeCP2-chromatin interactions include the formation of chromatosome-like structures and are altered in mutations causing Rett syndrome. J Biol Chem 282:28237–28245. doi:10.1074/jbc.M704304200
Nikitina T, Shi X, Ghosh RP et al (2007b) Multiple modes of interaction between the methylated DNA binding protein MeCP2 and chromatin. Mol Cell Biol 27:864–877. doi:10.1128/MCB.01593-06
Niwa H, Burdon T, Chambers I, Smith A (1998) Self-renewal of pluripotent embryonic stem cells is mediated via activation of STAT3. Genes Dev 12:2048–2060
Novo CL, Tang C, Ahmed K et al (2016) The pluripotency factor Nanog regulates pericentromeric heterochromatin organization in mouse embryonic stem cells. Genes Dev 30:1101–1115. doi:10.1101/gad.275685.115
Okano M, Bell DW, Haber DA, Li E (1999) DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99:247–257
Okano M, Xie S, Li E (1998) Cloning and characterization of a family of novel mammalian DNA (cytosine-5) methyltransferases. Nat Genet 19:219–220. doi:10.1038/890
Peters AH, Kubicek S, Mechtler K et al (2003) Partitioning and plasticity of repressive histone methylation states in mammalian chromatin. Mol Cell 12:1577–1589
Pradhan S, Bacolla A, Wells RD, Roberts RJ (1999) Recombinant human DNA (cytosine-5) methyltransferase. I. Expression, purification, and comparison of de novo and maintenance methylation. J Biol Chem 274:33002–33010
Rapkin LM, Ahmed K, Dulev S et al (2015) The histone chaperone DAXX maintains the structural organization of heterochromatin domains. Epigenetics Chromatin 8:44–58. doi:10.1186/s13072-015-0036-2
Saksouk N, Barth TK, Ziegler-Birling C et al (2014) Redundant mechanisms to form silent chromatin at pericentromeric regions rely on BEND3 and DNA methylation. Mol Cell 56:580–594. doi:10.1016/j.molcel.2014.10.001
Schotta G, Lachner M, Sarma K et al (2004) A silencing pathway to induce H3-K9 and H4-K20 trimethylation at constitutive heterochromatin. Genes Dev 18:1251–1262. doi:10.1101/gad.300704
Senner CE, Krueger F, Oxley D et al (2012) DNA methylation profiles define stem cell identity and reveal a tight embryonic-extraembryonic lineage boundary. Stem Cells 30:2732–2745. doi:10.1002/stem.1249
Smith ZD, Chan MM, Mikkelsen TS et al (2012) A unique regulatory phase of DNA methylation in the early mammalian embryo. Nature 484:339–344. doi:10.1038/nature10960
Smith ZD, Meissner A (2013) DNA methylation: roles in mammalian development. Nat Rev Genet 14:204–220. doi:10.1038/nrg3354
Sun B, Ito M, Mendjan S et al (2012) Status of genomic imprinting in epigenetically distinct pluripotent stem cells. Stem Cells 30:161–168. doi:10.1002/stem.793
Tesar PJ, Chenoweth JG, Brook FA et al (2007) New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 448:196–199. doi:10.1038/nature05972
Tsumura A, Hayakawa T, Kumaki Y et al (2006) Maintenance of self-renewal ability of mouse embryonic stem cells in the absence of DNA methyltransferases Dnmt1, Dnmt3a and Dnmt3b. Genes Cells 11:805–814. doi:10.1111/j.1365-2443.2006.00984.x
Veillard AC, Marks H, Bernardo AS et al (2014) Stable methylation at promoters distinguishes epiblast stem cells from embryonic stem cells and the in vivo epiblasts. Stem Cells Dev 23:2014–2029. doi:10.1089/scd.2013.0639
Williams RL, Hilton DJ, Pease S et al (1988) Myeloid leukaemia inhibitory factor maintains the developmental potential of embryonic stem cells. Nature 336:684–687. doi:10.1038/336684a0
Acknowledgements
We thank Ren Li for assistance in preparing the sections for correlative fluorescence and electron spectroscopic imaging. The research was supported by the Canadian Institutes of Health Research. DPB-J holds the Canada Research Chair in Molecular and Cellular Imaging.
Author contribution
VH-Z helped design, carried out the experiments, helped to analyse the data, and wrote the paper. PR-G developed the method to convert ESCs to EpiSCs, provided data on converted EpiSCs, and helped to write the paper. DPB-J helped design the experiments, helped to analyse the data, and helped to write the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Funding
The funding received was the Operating Grant number 229881 from the Canadian Institutes of Health Research.
Availability of data and materials
The datasets supporting the conclusions of this article are included within the article.
Electronic supplementary material
Supplementary Table S1
Primer sequences used for gene expression analysis (DOCX 15 kb)
Fig. S1
The average cluster sizes in WT and DNA methylation-deficient ESC and ESC-derived EpiSCs. The average cluster size in WT, DKO and TKO ESC-derived EpiSCs are 8,702, 9,421 and 10,009 nm2 respectively. These values are five to six times that observed in WT and DNA methylation-deficient ESCs. Using a t-test, the statistical significance of the greater than 5-fold increase in the cluster size distribution of the EpiSCs compared to the corresponding ESC lines is represented by a p-value of 0.0001 (DOCX 46 kb)
Fig. S2
The number of characteristic chromocentres is significantly reduced in TKO ESC-derived EpiSCs. The number of H3K9me3-enriched foci with characteristic chromocentre morphology (red bars) and dispersed chromatin (blue bars) in WT and TKO ESC-derived EpiSCs is displayed. The chi-square test shows that the difference in chromocentre morphology between WT and TKO ESC-derived EpiSCs is highly significant (χ2 =30.59, df=1, p-value<0.005) (DOCX 17 kb)
Rights and permissions
About this article
Cite this article
Hassan-Zadeh, V., Rugg-Gunn, P. & Bazett-Jones, D.P. DNA methylation is dispensable for changes in global chromatin architecture but required for chromocentre formation in early stem cell differentiation. Chromosoma 126, 605–614 (2017). https://doi.org/10.1007/s00412-017-0625-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00412-017-0625-x