Abstract
NBS (nucleotide binding site) genes, one type of the most important disease-resistance genes in the plant kingdom, are usually found clustered in genome. In this study, a total of 2288 full-length NBS protein-coding sequences were isolated from the wheat (Triticum aestivum L.) genome, and 903 TaNBSs of which were found expressed in wheat. Meanwhile, 2203 microsatellite loci were detected within 1061 scaffolds containing TaNBS. The distribution of these microsatellite loci across wheat homologous groups (HG) is 20% HG2, 16% HG7, 15% HG1, 15% HG6, 12% HG4, 12% HG5 and 10% HG3. We developed 1830 NBS-related microsatellite (NRM) markers for the microsatellite loci on TaNBS-scaffold sequences.Among them, 342 NRM markers were developed for HG2 with the largest number of microsatellite loci, and 69 out of these markers were anchored to the wheat genetic map using mapping population. Then, a total of 26 2AS-NRM markers, nine 2BL-NRM markers and nine 2DL-NRM markers were integrated into the genetic maps carrying Yr69, Pm51 and Pm43, respectively. Finally, candidate sequences, within the gene clusters where Yr5 and Sr21 located, were analyzed according to the genomic position information of TaNBS and NRM markers. These NRM markers have clear chromosome locations and are correlated with potential disease resistance sequences, which can be manipulated to mapping or adding linkage markers of disease-resistance genes or QTLs, especially for those in the NBS gene clusters.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
NBS (nucleotide binding site) gene family is one of the largest and most important disease resistance gene families in plant. The NBS domain, encoded by NBS genes, contains several conserved motifs such as Kinase1, Kinase2, Kinase3 and HD residues (Meyers et al. 2003), which can directly or indirectly mediate the pathogen recognition via binding ATP or GTP and therefore participate in signal transduction (Elmore et al. 2011; Krattinger and Keller 2016). To date, more than 140 disease resistance (R) genes were cloned and 80% of them are NBS genes (Shao et al. 2016). With the publications of more and more draft genome sequence of different plants, NBS gene families of 19 dicots and 11 monocots have been isolated from respective genome (Urbach and Ausubel 2017; Morata and Puigdomènech 2017). In dicots, some NBS domains are usually linked to N-terminal TOLL/interleukin 1 receptor (TIR) or C-terminal leucine-rich repeat (LRR) that associated with pathogen recognition; while not the TIR but a coiled-coil domain (CC), which associated with protein–protein interactions, is present in the NBS domain of monocots (Lee and Yeom 2015; Urbach and Ausubel 2017).
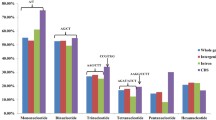
Bread wheat (Triticum aestivum L.) is the most widely cultivated food crop, which faces attacks from various pathogens during its growth. Of which, fungal diseases like powdery mildew and rust (stripe rust, leaf rust and stem rust) can cause severe damage to wheat production (Goutam et al. 2015). Extensively identifying and utilizing R genes for disease resistance breeding is the most cost-effective way to control wheat disease (Chen and Line 1995). So far, a total of 58 powdery mildew R genes (Wiersma et al. 2017), 76 stripe rust R genes (Xiang et al. 2016), 76 leaf rust R genes (Bansal et al. 2017) and 59 stem rust R genes (Rahmatov et al. 2016) were formally designated. However, only 24 out of them have been cloned, and 20 (83%) were NBS genes, namely: Pm2a (Sánchez-Martín et al. 2016), Pm3a-g (Yahiaoui et al. 2004; Tommasini et al. 2006), Pm8 (Hurni et al. 2014), Pm21 (Xing et al. 2017; He et al. 2017), Yr10 (Liu et al. 2014), Lr1 (Cloutier et al. 2007), Lr10 (Sela et al. 2012), Lr21 (Huang et al. 2003), Lr22 (Thind et al. 2017), Sr22 (Steuernagel et al. 2016), Sr33 (Periyannan et al. 2013), Sr35 (Saintenac et al. 2013), Sr45 (Steuernagel et al. 2016) and Sr50 (Mago et al. 2015). Moreover, Yr5 (Smith et al. 2007) and Sr21 (Chen et al. 2015) were mapped to the NBS gene clusters, and the expressed sequence tag (EST) of Yr5 was also reported to encode partial NBS domain (Smith et al. 2007). Except for this, a large number of resistances genes and QTLs, formally designated or not, were only preliminarily mapped. Most of the linked molecular markers are often unable to be effectively used for molecular breeding because of their distant locations from the target genes or QTLs. Therefore, isolation of wheat NBS family with clear position information and molecular markers is valuable for both fine mapping the target genes or QTLs mentioned above and the candidate sequences screening.
Using the 454 sequencing data of the common wheat cultivar Chinese Spring (Rachel et al. 2012), 580 and 986 complete NBS sequences were isolated respectively and their types and structures were analyzed (Bouktila et al. 2014, 2015; Gu et al. 2015); however, genomic positions of these sequences are unknown. In this study, the wheat genome sequence based on sequencing isolated chromosome arms (International Wheat Genome Sequencing Consortium 2014) was employed to isolate wheat NBS sequences with genomic location information. In addition, a set of NBS-related microsatellite (NRM) markers were developed according to the microsatellite loci adjacent to the NBS sequences; meanwhile, NRM markers in wheat homologous group (HG) 2, which harbors the most microsatellite loci, were used to construct the genetic map via mapping population and were analyzed their linkage with R genes. These TaNBS sequences and corresponding molecular markers can be further used for R genes or QTLs mapping.
Materials and methods
Plant materials
A RILs population of 194 lines derived from a cross between wheat CH7034 and SY95-71 were recruited to construct NRM marker map; An F2 population of 136 plants from CH5025 (carrying Pm43, He et al. 2009)/Taichang (TC) 29 and 92 F2:3 lines derived from CH7086 (carrying Pm51 and Yr69, Zhan et al. 2014; Hou et al. 2016)/TC 29 were used to test for marker-R gene association. DNA samples of the above mapping population and their parents as well as phenotype data (He et al. 2009; Zhan et al. 2014; Hou et al. 2016) were provided by Shanxi Key Laboratory of Crop Genetics and Molecular Improvement.
NBS sequences isolation and bioinformatics analysis
The wheat whole genome sequences and predicted protein sequences were downloaded from the URGI database (http://wheat-urgi.versailles.inra.fr/). The wheat predicted protein data were retrieved in HMMER 3.0 software (Mistry et al. 2013) using the NBS family Hidden Markov Model file (accession number PF00931, downloaded from http://pfam.xfam.org/); the search results were then examined for the conserved domains of NBS proteins via SMART (http://smart.embl-hei-delberg.de). The obtained NBS sequences were submitted to the CDD database in NCBI (http://www.ncbi.nlm.nih.gov/Structure/cdd) to further analyze characteristic domains such as CC, LRR that correlated with NBS. The corresponding coding sequences and scaffold sequences were extracted from the wheat genome data according to the protein accession number; and their position information was determined by retrieving the wheat genome; then, TaNBSs were assigned to corresponding chromosomes. Gene structures of TaNBS were determined by the GSDS2.0 (http://gsds.cbi.pku.edu.cn/). Expression profiles of TaNBSs were obtained from retrieving wheat transcriptome sequencing data (accession number PRJNA243835, downloaded from http://www.ncbi.nlm.nih.gov/sra/).
NRM markers development and PCR validation
SSRhurnter software (Li and Wan 2005) was used to search for the microsatellite loci within scaffolds where TaNBSs situated. The searching criteria were set as follows: the nucleotides per repeat unit were two to five, and the repeat times ≥ 5. Primers were designed using the Primer5 software (http://www.premierbiosoft.com/primerdesign/) for NRM loci.
To compare with the wheat molecular map constructed by wPt-DArT and SSR markers (Marone et al. 2013), 34 GBS-DArT markers, which located in the same loci (genomic distance < 10 kb) with the wPt-DArT markers in the HG2 of wheat, as well as 346 pairs of SSR markers were also recruited for NRM-map construction. PCR used for screening and mapping NRM and other SSR markers was performed in 15 μL reaction mixture containing 1 U Taq DNA polymerase (Takara Bio Inc. Dalian, China), 1.5 μL 10 × buffer, 0.2 mmol L−1 dNTPs, 0.25 μmol L−1 primers and 100 ng of genomic DNA. PCR products were separated in 8% non-polyacrylamide denaturing gel and visualized by silver staining. The GBS-DArT marker typing was performed by Diversity Arrays Technology (DArT) PL, Canberra, Australia. The linkage map was constructed using JoinMap 4.0 software (https://www.kyazma.nl/index.php/JoinMap/), and the parameters were set as follows: Node-population, Grouping-independence LOD and Mapping function-Kosambi’s.
Results and analysis
Distribution, types and expression of wheat NBS family
Conserved domain check was conducted on sequences isolated from the wheat database, and a total of 2288 complete protein sequences containing NBS domain were obtained. The distribution of these sequences in genome A, B and D is 34.1, 37.7 and 28.2%, respectively; among them, chromosome 4A contained the highest number of protein sequences, up to 227; while 4D contained the lowest, only 24 protein sequences (Fig. 1a). The length of TaNBS varies considerably, from the shortest sequence Ta1asLoc007418.1, only 48aa to the longest Ta4alLoc027793.2 for 1816aa (Table S1). Based on whether the CC and LRR domains are included, TaNBS sequences were classified into 4 types: CC-NBS-LRR (CNL), NBS-LRR (NL), CC-NBS (CN) and NBS (N); and the N-type was the largest, which comprises 1144 sequences, accounting for 50% (Fig. 1a). The gene length of TaNBSs range from 251 bp to 7762 bp and the introns contained varies from one to 13; a total of 903 (56.3%) TaNBSs expressed in wheat (Table S1); out of the 632 TaNBSs with genetic position information, 477 (75.5%) were clustered (Table S1).
TaNBS-related microsatellite loci analysis and marker development
In total, 2203 microsatellite loci were detected on 1061 scaffold sequences containing TaNBS, of which 1621 were dinucleotide repeats loci, accounting for 73.6%, while the five nucleotide repeats loci was the least, only six (Fig. 1b). The distribution of these microsatellite loci across the wheat HGs of is HG2 (20%), HG7 (16%), HG1 (15%), HG6 (15%), HG4 (12%), HG5 (12%) and HG3 (10%) (Fig. 1c). We totally developed 1830 pairs of NRM markers from TaNBS-scaffold sequences with microsatellite loci (Tables 1, S2). Among them, 342 pairs of NRM markers were developed on HG2 that contained the most microsatellite loci, including 49 2AS-NRM markers, 51 2AL-NRM markers, 71 2BS-NRM markers, 94 2BL-NRM markers, 34 2DS-NRM markers and 43 2DL-NRM markers (Table 1).
NRM markers map construction of the HG2 in wheat
The RILs population of CH7034/SY95-71 was amplified with 342 NRM markers of HG2, and 115 NRM markers showed polymorphism between the parents. Finally, 69 NRM markers, 20 SSR markers and 16 DArT markers were mapped to the genetic map (Fig. 2). The results showed that 31 NRM markers were assigned to chromosome 2A, 25 of them were on the short arm and mainly clustered in two regions; eight NRM markers were identified in the region of DArT marker 1088906-1138983, and this region may contain disease resistance genes, such as Yr69 (Hou et al. 2016) and Yr17/Lr37/Sr38 (Helguera et al. 2003). In addition, 22 NRM markers were assigned to chromosome 2B, and three 2BL-NRM markers were detected in the region of SSR markers Xgwm501-Xwmc332, which may contain Pm51 (Zhan et al. 2014), Yr5 (Smith et al. 2007; McGrann et al. 2014), Lr48 (Singh et al. 2011), Sr9a (Tsilo et al. 2007), Sr28 (Rouse et al. 2012) and QYraq.cau-2BL (Guo et al. 2008). Sixteen NRM markers were assigned to chromosome 2D, and four 2DL-NRM markers were in the region of DArT marker 1114628-1086188. There might be Sr6 (Tsilo et al. 2010) and QPm.caas-2DL (Lan et al. 2010) in this region.
Genetic map of NRM markers in homologous group 2 of wheat constructed by CH7034/SY95-71 RILs population. The NRM markers are labelled in blue, the SSR or DArT markers appearing in both linkage maps are labelled by pink, and the genes for the next enhancing map-density are labelled with red. (Color figure online)
Construction of more densely populated map with NRM markers
NRM markers were employed to enhance the map-density of three disease resistance genes Yr69 (2AS, Hou et al. 2016), Pm51 (2BL, Zhan et al. 2014) and Pm43 (2DL, He et al. 2009) in HG2, respectively. The results showed that 26, nine and nine NRM markers were integrated into the genetic maps where Yr69, Pm51 and Pm43 sited, respectively. Among them, eight NRM markers were integrated into the original X2AS33-1.9 cM-Yr69-3.1 cM-Xmag3807 region of Yr69, which narrowed the region to X2AS-NRM34-0.4 cM-Yr69-1.3 cM-X2AS-NRM31, inferring that Yr69 may locate in the NBS gene cluster (Fig. 3a). In addition, a marker X2DL-NRM05 (Fig. 3c), which is closer to Pm43, was obtained.
Identification of candidate sequences within gene clusters by TaNBS and NRM markers
Yr5 (Smith et al. 2007) and Sr21 (Chen et al. 2015) are two disease resistance genes that have been confirmed to be located in the NBS gene cluster. The EST (Genbank number JN631792) of Yr5 encoded partial NBS structure, and its co-segregation marker TaAffx.65234.1.S1_at as well as its flanking linkage marker S23M41-310 were also NBS sequences, in which S23M41-310 is orthologous to the rice NBS gene OsXa1 (Smith et al. 2007; McGrann et al. 2014). In this study, Yr5 was assigned to a gene cluster, consisting of 11 TaNBSs and 11 NRM markers, in the region 693,372,707-732,340,422 of chromosome 2B. The TaNBS sequence Ta2blLoc006115.1 includes the whole Yr5-EST (100% sequence similarity) and can express in wheat (Table S1); its co-segregation marker TaAffx.65234.1.S1_at and flanking marker S23M41-310 correspond to Ta2blLoc008215.1 and Ta2blLoc034091.1 in the gene cluster, respectively (Fig. 4).
In addition, we assigned seven linkage markers of Sr21 to chromosome 2A with the closest flanking markers corresponded to a 1.28 Mb region 709,765,601-711,049,068, which contained four TaNBS sequences and two NRM markers (Fig. 5). Multiple sequence alignment results showed that the TaNBS protein sequences Ta2blLoc019062.3, Ta2blLoc029872.3 and Ta2blLoc029875.1 exhibited highly similar motifs to those within the NBS domain of cloned disease resistance genes (Fig. S1), such as P-loop (GGxGKTT) for ATP/GTP binding, Kinase2 (LLVLDDxW), Kinase3 (GxxxLxTxR) and HD residues. Moreover, Ta2blLoc019062.3 expressed in wheat (Table S1), which means it may participate in the process against pathogens, this requires subsequent validation in the mapping population using its marker2AL-NRM05.
Discussion
Size of the wheat NBS family
NBS, the largest disease resistance gene family, has been surveyed in various plants, such as bryophytes, lycopodiums, gymnosperms and angiosperms (Elmore et al. 2011; Krattinger and Keller 2016; Shao et al. 2016; Urbach and Ausubel 2017; Lee and Yeom 2015). The fungal bloom during the Cretaceous-Paleogene boundary (~ 66 MYA) may trigger the intensive expansion of NBS genes in plants (Shao et al. 2016), which suggests the important role of NBS genes in the fight against fungal disease during evolution. In cotton, the NBS expansion in Gossypium raimondii enhanced its resistance to Verticillillm dahliae, while G. arboreum without NBS expansion was easily susceptible (Li et al. 2014). After undergoing three independent whole-genome duplications (WGD), however, only 117 NBS genes were included in banana genome, which may explain its susceptibility to pathogen attacks (D’Hont et al. 2012). Therefore, isolation and analysis of plant NBS gene families can help us better elucidate the underlying disease resistance mechanisms.
In this study, 2288 complete TaNBS sequences were isolated from the hexaploid wheat genome, which is significantly higher than that of other gramineous crops evolved from the same grass ancestor (50–70 MYA) (Salse et al. 2008), such as 535 of rice (Zhou et al. 2004), 420 of barley, 316 of Brachypodium distachyon (Gu et al. 2015), 274 of sorghum (Cheng et al. 2010) and 109 of maize (Cheng et al. 2012). We also isolated 463 (Liu et al. 2017) and 701 complete NBS sequences (the results were not listed) from Triticum urartu and Aegilops tauschii, respectively. It was hypothesized that the NBS families of three wild ancestral species of wheat were integrated into the genome of the hexaploid wheat after two polyploidization events (0.8 and 0.4 MYA, IWGSC 2014); then with the propagation of wheat, the TaNBS family expanded again to adapt to the infection of various pathogens in different planting areas, which eventually led to this gene family with large number of members. The majority of these TaNBS members (about 70%) are clustered in genome, which is also true for NBS family members in plant species like Arabidopsis thaliana (71.1%, Meyers et al. 2003), rice (76%, Zhou et al. 2004) and potato (73%, Jupe et al. 2012). The presence of the TaNBS gene clusters as well as the loss of some subgenomic copies in the evolutionary process (Comai 2005; Otto 2007) resulted in an uneven distribution of NBSs across chromosomes in HGs; For example, there were 248 TaNBS genes on chromosome 4A, while only 40 and 24 were on chromosome 4B and 4D, respectively. This may explain that the cloned disease resistance genes are often one copy rather than ‘triplet gene’ (Pfeifer et al. 2014).
TaNBSs provide reference for homology-based and map-based cloning
To date, some disease resistance-related NBS genes were cloned from wheat using a homology-based cloning strategy. In the case of resistance to powdery mildew, there are genes such as TmMla1 (Jordan et al. 2011), homologous to barley HvMla1 (78% sequence similarity), from diploid wheat; TdRGA-7Ba (Gong et al. 2013), a Pm3b homologue (sequence similarity > 90%), from durum wheat; and TaRGA from common wheat (Wang et al. 2016), which is homologous to multiple plant disease resistance genes. Many TaNBS isolated in this study exhibited high sequences similarity with those cloned disease resistance genes. For example, both Ta1bsLoc017427.1and Ta1bsLoc003202.1 shared over 70% identity with barley powdery resistance genes HvMla1 (Zhou et al. 2001), HvMla6 (Halterman et al. 2001) and HvMla13 (Halterman et al. 2003) at the protein level. These TaNBSs can provide candidate gene sequences for cloning wheat disease-resistance genes using homology-based method. Normally, genes with similar domains may possess similar functions. The 19 cloned wheat NBS-encoding protiens are all CNL-type, and the number of TaNBS proteins with CNL isolated from this study is 240 (Fig. 1). In addition, some TaNBSs contain other distinct domains, such as Ta2bsLoc003709 and Ta2bsLoc014737, in HG2, contian ABC (ATP-binding cassette) transport protein domain; Ta2alLoc020734 and Ta2dlLoc021209 contain Jacalin domain; while Ta2alLoc012596 and Ta2blLoc002392 contain zinc finger domian; these domains have been proved to play an important role in disease resistance process (Krattinger et al. 2009; Ma et al. 2013; Guo et al. 2013). ABC is also the function domains of Lr34 (Krattinger et al. 2009). Studies have shown that these domains may interact with the NBS domain and against pathogen invasion together (Deslandes et al. 2002). Hence, it is necessary to further explore these TaNBSs in depth.
Furthermore, TaNBSs can provide reference sequences for map-based cloning of disease-resistance genes that in the NBS gene cluster. It has been reported in previous studies that if the comparative genome analysis showed the region of a mapped wheat resistance gene corresponded to the NBS gene cluster of rice, B. distachyon or other model plants, then often this gene is in the wheat NBS gene cluster and is a TaNBS gene, like both Sr35 (Saintenac et al. 2013) and Sr50 (Mago et al. 2015) are TaNBS genes in the NBS gene cluster. In this study, we analyzed the genomic location of NBS gene cluster where Yr5 sited and identified a TaNBS sequence Ta2blLoc006115.1 containing Yr5-EST, which indicated the accuracy of the genomic location of TaNBS family. Then, Sr21, which is also in the NBS gene cluster, was analyzed and anchored its linkage marker to genomic map. Finally, four candidate TaNBS sequences were found in the target region.
NRM markers in gene mapping and molecular breeding
Extensively identification and cloning disease resistance genes are the foundations for wheat disease resistance breeding. So far, among nearly 300 powdery mildew and rust resistance genes were formally designated in wheat, only few of them can be used for wheat improvement and most of them are facing the risk of resistance loss due to pathogens variation before being used for breeding. This situation could be attributed to that the linkage marker of the R gene cannot be efficiently used for marker-assisted selection (MAS). Since the common wheat contains three sub-genomes and highly repetitive sequences (80%, IWGSC et al. 2014), most of the SSR markers, routinely used for R gene mapping, have a low distribution density across the wheat genome, which resulted in their often faraway locations from the gene of interest. Besides, it is not easy to develop markers in the linkage region due to the unclear genome location of these SSR markers. Thus, gene recombination may occur between linked markers and target genes and result in the marker missing, which leads to breeding failure.
With the rapid development of sequencing technology, markers with high density and precision like SNP, DArT have been developed, which greatly improved mapping accuracy and narrowed the distance between mark and gene. Meta analysis showed a number of DArT markers that in the same loci as the R genes or QTLs are NBS sequences, such as PmHNK54/wPt-5865, QPm.inra.2A/wPt-6064, Pm23/wPt-7024 and Pm42/wPt-2600 in wheat HG2 (Marone et al. 2013). This could be explained by the fact that many R genes are located in the NBS gene cluster, thus their linked DArT markers may also NBS-related sequences. However, relatively high chip scanning costs makes it unsuitable for large population screening in breeding process for now. Aiming at improving the effectiveness of routinely used molecular markers, we developed 1830 NRM markers, each of which lay in the same scaffold sequence with TaNBS. Of all the NRM markers is 7DL_NRM59 the farthest from TaNBS sequence Ta7dlLoc025447 with the longest genetic distance of 38589 bp, which is approximately equal to 0.007 cM based on the ratio of physical and genetic distance on chromosome 7DL (5.41 Mbp/cM; IWGSC, 2014). The remaining genetic distances between NRM markers and TaNBSs are below the value. Since most NRM markers were clustered in some regions with the NBS genes, the polymorphism of NRM markers is low in the genome regions that do not contain disease resistance loci. If R gene and multiple NRM markers are anchored to the same loci, this gene may locate in a NBS gene cluster; such as Yr69 in this study. Moreover, when a TaNBS is confirmed to be associated with disease resistance, the NRM marker(s) on its scaffold can be directly used as a co-segregation marker, which will improve the MAS efficiency in breeding process.
References
Bansal M, Kaur S, Dhaliwal HS, Bains NS, Bariana HS, Chhuneja P, Bansal UK (2017) Mapping of Aegilops umbellulata-derived leaf rust and stripe rust resistance loci in wheat. Plant Pathol 66(1):38–44. doi:10.1111/ppa.12549
Bouktila D, Habachi-Houimli Y, Khalfallah Y, Mezghani-Khemakhem M, Makni M, Makni H (2014) Characterization of novel wheat NBS domain-containing sequences and their utilization, in silico, for genome-scale R-gene mining. Mol Genet Genom 289(4):599–613. doi:10.1007/s00438-014-0834-4
Bouktila D, Khalfallah Y, Habachi Y, Mezghani M, Makni M, Makni H (2015) Full-genome identification and characterization of NBS-encoding disease resistance genes in wheat. Mol Genet Genom 290:257–271. doi:10.1007/s00438-014-0909-2
Chen XM, Line RF (1995) Gene-action in wheat cultivars for durable, high-temperature, adult-plant resistance and interaction with race-specific, seedling resistance to Puccinia striiformis. Phytopathology 85:567–572
Chen S, Rouse MN, Zhang W, Jin Y, Akhunov E, Wei Y, Dubcovsky J (2015) Fine mapping and characterization of Sr21, a temperature-sensitive diploid wheat resistance gene effective against the Puccinia graminis f. sp. tritici Ug99 race group. Theor Appl Genet 128(4):645–656. doi:10.1007/s00122-015-2460-x
Cheng X, Jiang H, Zhao Y, Qian Y, Zhu S, Cheng B (2010) A genomic analysis of disease-resistance genes encoding nucleotide binding sites in Sorghum bicolor. Genet Mol Biol 33(2):292–297. doi:10.1590/S1415-47572010005000036
Cheng Y, Li X, Jiang H, Ma W, Miao W, Yamada T, Zhang M (2012) Systematic analysis and comparison of nucleotide-binding site disease resistance genes in maize. FEBS J 279(13):2431–2443. doi:10.1111/j.1742-4658.2012.08621.x
Cloutier S, McCallum BD, Loutre C, Banks TW, Wicker T, Feuillet C, Keller B, Jordan MC (2007) Leaf rust resistance gene Lr1, isolated from bread wheat (Triticum aestivum L.) is a member of the large psr567 gene family. Plant Mol Biol 65:93–106. doi:10.1007/s11103-007-9201-8
Comai L (2005) The advantages and disadvantages of being polyploid. Nat Rev Genet 6:836–846. doi:10.1038/nrg1711
Deslandes L, Olivier J, Theulieres F, Hirsch J, Feng DX, Bittner-Eddy PD, Beynon J, Marco Y (2002) Resistance to Ralstonias olanacearum in Arabidopsis thaliana is conferred by the recessive RRS1-R gene, a member of a novel family of resistance genes. Proc Natl Acad Sci USA 99:2404–2409. doi:10.1073/pnas.032485099
D’Hont A, Denoeud F, Aury JM, Baurens FC, Carreel F, Garsmeur O, Noel B, Bocs S, Droc G, Rouard M, Da Silva C, Jabbari K, Cardi C, Poulain J, Souquet M, Labadie K, Jourda C, Lengellé J, Rodier-Goud M, Alberti A, Bernard M, Correa M, Ayyampalayam S, Mckain MR, Leebens-Mack J, Burgess D, Freeling M, Mbéguié-A-Mbéguié D, Chabannes M, Wicker T, Panaud O, Barbosa J, Hribova E, Heslop-Harrison P, Habas R, Rivallan R, Francois P, Poiron C, Kilian A, Burthia D, Jenny C, Bakry F, Brown S, Guignon V, Kema G, Dita M, Waalwijk C, Joseph S, Dievart A, Jaillon O, Leclercq J, Argout X, Lyons E, Almeida A, Jeridi M, Dolezel J, Roux N, Risterucci AM, Weissenbach J, Ruiz M, Glaszmann JC, Quétier F, Yahiaoui N, Wincker P (2012) The banana (Musa acuminata) genome and the evolution of monocotyledonous plants. Nature 488(7410):213–217. doi:10.1038/nature11241
Elmore JM, Lin ZJ, Coaker G (2011) Plant NB-LRR signaling: Upstreams and downstreams. Curr Opin Plant Biol 14:365–371. doi:10.1016/j.pbi.2011.03.011
Gong C, Cao S, Fan R, Wei B, Chen G, Wang X, Li Y, Zhang X (2013) Identification and phylogenetic analysis of a CC-NBS-LRR encoding gene assigned on chromosome 7B of wheat. Int J Mol Sci 14(8):15330–15347. doi:10.3390/ijms140815330
Goutam U, Kukreja S, Yadav R, Salaria N, Thakur K, Goyal AK (2015) Recent trends and perspectives of molecular markers against fungal diseases in wheat. Front Microbiol 6:861. doi:10.3389/fmicb.2015.00861
Gu L, Si W, Zhao L, Yang S, Zhang X (2015) Dynamic evolution of NBS-LRR genes in bread wheat and its progenitors. Mol Genet Genom 290(2):727–738. doi:10.1007/s00438-014-0948-8
Guo Q, Zhang ZJ, Xu YB, Li GH, Feng J, Zhou Y (2008) Quantitative trait loci for high-temperature adult-plant and slow-rusting resistance to Puccinia striiformis f. sp. tritici in wheat cultivars. Phytopathology 98(7):803–809. doi:10.1094/PHYTO-98-7-0803
Guo J, Bai P, Yang Q, Liu F, Wang X, Huang L, Kang Z (2013) Wheat zinc finger protein TaLSD1, a negative regulator of programmed cell death, is involved in wheat resistance against stripe rust fungus. Plant Physiol Biochem 71:164–172. doi:10.1016/j.plaphy.2013.07.009
Halterman D, Zhou F, Wei F, Wise RP, Schulze-Lefert P (2001) The MLA6 coiled-coil, NBS-LRR protein confers AvrMla6-dependent resistance specificity to Blumeria graminis f. sp. hordei in barley and wheat. Plant J 25(3):335–348. doi:10.1046/j.1365-313x.2001.00982.x
Halterman DA, Wei F, Wise RP (2003) Powdery mildew-induced Mla mRNAs are alternatively spliced and contain multiple upstream open reading frames. Plant Physiol 131(2):558–567. doi:10.1104/pp.014407
He R, Chang Z, Yang Z, Yuan Z, Zhan H, Zhang X, Liu J (2009) Inheritance and mapping of powdery mildew resistance gene Pm43 introgressed from Thinopyrum intermedium into wheat. Theor Appl Genet 118(6):1173–1180. doi:10.1007/s00122-009-0971-z
He HG, Zhu SY, Ji YY, Jiang ZN, Zhao RH, Bie TD (2017) Map-based cloning of the gene Pm21 that confers broad spectrum resistance to wheat powdery mildew. arXiv 1708.05475. doi:https://arxiv.org/abs/1708.05475
Helguera M, Khan IA, Kolmer J, Lijavetzky D, Zhong L, Dubcovsky J (2003) PCR assays for the Lr37-Yr17-Sr38 cluster of rust resistance genes and their use to develop isogenic hard red spring wheat lines. Crop Sci 43:1839–1847
Hou LY, Jia JQ, Zhang XJ, Li X, Yang ZJ, Jian Ma, Guo HJ, Zhan HX, Qiao LY, Chang ZJ (2016) Molecular mapping of the stripe rust resistance gene Yr69 on wheat chromosome 2AS. Plant Dis 100(8):1717–1724. doi:10.1094/PDIS-05-15-0555-RE
Huang L, Brooks SA, Li W, Fellers JP, Trick HN, Gill BS (2003) Map-based cloning of leaf rust resistance gene Lr21 from the large and polyploid genome of bread wheat. Genetics 164:655–664
Hurni S, Brunner S, Stirnweis D, Herren G, Peditto D, McIntosh RA, Keller B (2014) The powdery mildew resistance gene Pm8 derived from rye is suppressed by its wheat ortholog Pm3. Plant J 79(6):904–913. doi:10.1111/tpj.12593
International Wheat Genome Sequencing Consortium (IWGSC) (2014) A chromosome-based draft sequence of the hexaploid bread wheat (Triticum aestivum) genome. Science 345:1251788. doi:10.1126/science.1251788
Jordan T, Seeholzer S, Schwizer S, Töller A, Somssich IE, Keller B (2011) The wheat Mla homologue TmMla1 exhibits an evolutionarily conserved function against powdery mildew in both wheat and barley. Plant J 65(4):610–621. doi:10.1111/j.1365-313X.2010.04445.x
Jupe F, Pritchard L, Etherington GJ, Mackenzie K, Cock PJ, Wright F, Sharma SK, Bolser D, Bryan GJ, Jones JD, Hein I (2012) Identification and localisation of the NB-LRR gene family within the potato genome. BMC Genom 13:75. doi:10.1186/1471-2164-13-75
Krattinger SG, Keller B (2016) Molecular genetics and evolution of disease resistance in cereals. New Phytol 212(2):320–332. doi:10.1111/nph.14097
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Espino JH, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363. doi:10.1126/science.1166453
Lan CX, Ni XW, Yan J, Zhang Y, Xia XC, Chen XM, He ZH (2010) Quantitative trait loci mapping of adult-plant resistance to powdery mildew in Chinese wheat cultivar Lumai 21. Mol Breed 25:615–622. doi:10.1007/s11032-009-9358-8
Lee HA, Yeom SI (2015) Plant NB-LRR proteins: Tightly regulated sensors in a complex manner. Brief Funct Genom 14:233–242. doi:10.1093/bfgp/elv012
Li Q, Wan JM (2005) SSRHunter: development of a local searching software for SSR sites. Hereditas (Beijing) 27(5):808–810 [Article in Chinese]
Li F, Fan G, Wang K, Sun F, Yuan Y, Song G, Li Q, Ma Z, Lu C, Zou C, Chen W, Liang X, Shang H, Liu W, Shi C, Xiao G, Gou C, Ye W, Xu X, Zhang X, Wei H, Li Z, Zhang G, Wang J, Liu K, Kohel RJ, Percy RG, Yu JZ, Zhu YX, Wang J, Yu S (2014) Genome sequence of the cultivated cotton Gossypium arboreum. Nat Genet 46(6):567–572. doi:10.1038/ng.2987
Liu W, Frick M, Huel R, Nykiforuk CL, Wang X, Gaudet DA, Eudes F, Conner RL, Kuzyk A, Chen Q, Kang Z, Laroche A (2014) The stripe rust resistance gene Yr10 encodes an evolutionary-conserved and unique CC-NBS-LRR sequence in wheat. Mol Plant 7:1740–1755. doi:10.1093/mp/ssu112
Liu J, Qiao L, Zhang X, Li X, Zhan HX, Guo HJ, Zheng J, Chang ZJ (2017) Genome-wide identification and resistance expression analysis of the NBS gene family in Triticum urartu. Genes Genom 39:611–621. doi:10.1007/s13258-017-0526-7
Ma QH, Zhen WB, Liu YC (2013) Jacalin domain in wheat jasmonate-regulated protein Ta-JA1 confers agglutinating activity and pathogen resistance. Biochimie 95:359–365. doi:10.1016/j.biochi
Mago R, Zhang P, Vautrin S, Šimková H, Bansal U, Luo MC, Rouse M, Karaoglu H, Periyannan S, Kolmer J, Jin Y, Ayliffe MA, Bariana H, Park RF, McIntosh R, Doležel J, Bergès H, Spielmeyer W, Lagudah ES, Ellis JG, Dodds PN (2015) The wheat Sr50 gene reveals rich diversity at a cereal disease resistance locus. Nat Plants 1:15186. doi:10.1038/nplants.2015.186
Marone D, Russo MA, Laidò G, De Vita P, Papa R, Blanco A, Gadaleta A, Rubiales D, Mastrangelo AM (2013) Genetic basis of qualitative and quantitative resistance to powdery mildew in wheat: from consensus regions to candidate genes. BMC Genom 14:562. doi:10.1186/1471-2164-14-562
McGrann GRD, Smith PH, Burt C, Mateos GR, Chama TN, MacCormack R, Wessels E, Agenbag G, Boyd LA (2014) Genomic and genetic analysis of the wheat race-specific yellow rust resistance gene Yr5. J Plant Sci Mol Breed 3:2. doi:10.7243/2050-2389-3-2
Meyers BC, Kozik A, Griego A, Kuang H, Michelmore RW (2003) Genome-wide analysis of NBS-LRR-encoding genes in Arabidopsis. Plant Cell 15:809–834. doi:10.1105/tpc.009308
Mistry J, Finn RD, Eddy SR, Bateman A, Punta M (2013) Challenges in homology search: HMMER3 and convergent evolution of coiled-coil regions. Nucleic Acids Res 41(12):e121. doi:10.1093/nar/gkt263
Morata J, Puigdomènech P (2017) Variability among Cucurbitaceae species (melon, cucumber and watermelon) in a genomic region containing a cluster of NBS-LRR genes. BMC Genom 18(1):138. doi:10.1186/s12864-017-3529-5
Otto SP (2007) The evolutionary consequences of polyploidy. Cell 131:452–462. doi:10.1016/j.cell.2007.10.022
Periyannan S, Moore J, Ayliffe M, Bansal U, Wang X, Huang L, Deal K, Luo M, Kong X, Bariana H, Mago R, McIntosh R, Dodds P, Dvorak J, Lagudah E (2013) The gene Sr33, an ortholog of barley Mla genes, encodes resistance to wheat stem rust race Ug99. Science 341:786–788. doi:10.1126/science.1239028
Pfeifer M, Kugler KG, Sandve SR, Zhan B, Rudi H, Hvidsten TR, International Wheat Genome Sequencing Consortium, Mayer KF, Olsen OA (2014) Genome interplay in the grain transcriptome of hexaploid bread wheat. Science 345:1250091. doi:10.1126/science.1250091
Rachel B, Manuel S, Matthias P, Gary LAB, Rosalinda DA, Alexandra MA, Neil M, Melissa K, Arnaud K, Dan B, Suzanne K, Darren W, Martin T, Ian B, Gu Y, Huo NX, Luo MC, Sunish S, Bikram G, Sharyar K, Olin A, Paul K, Jan D, Richard M, Anthony H, Klaus FM, Keith JE, Michael WB, Hall N (2012) Analysis of the bread wheat genome using whole-genome shotgun sequencing. Nature 491:705–710. doi:10.1038/nature11650
Rahmatov M, Rouse MN, Nirmala J, Danilova T, Friebe B, Steffenson BJ, Johansson E (2016) A new 2DS 2RL Robertsonian translocation transfers stem rust resistance gene Sr59 into wheat. Theor Appl Genet 129(7):1383–1392. doi:10.1007/s00122-016-2710-6
Rouse MN, Nava IC, Chao S, Anderson JA, Jin Y (2012) Identification of markers linked to the race Ug99 effective stem rust resistance gene Sr28 in wheat (Triticum aestivum L.). Theor Appl Genet 125(5):877–885. doi:10.1007/s00122-012-1879-6
Saintenac C, Zhang W, Salcedo A, Rouse MN, Trick HN, Akhunov E, Dubcovsky J (2013) Identification of wheat gene Sr35 that confers resistance to Ug99 stem rust race group. Science 341:783–786. doi:10.1126/science.1239028
Salse J, Bolot S, Throude M, Jouffe V, Piegu B, Quraishi UM, Calcagno T, Cooke R, Delseny M, Feuillet C (2008) Identification and characterization of shared duplications between rice and wheat provide new insight into grass genome evolution. Plant Cell 20(1):11–24. doi:10.1105/tpc.107.056309
Sánchez-Martín J, Steuernagel B, Ghosh S, Herren G, Hurni S, Adamski N, Vrána J, Kubaláková M, Krattinger SG, Wicker T, Doležel J, Keller B, Wulff BB (2016) Rapid gene isolation in barley and wheat by mutant chromosome sequencing. Genome Biol 17(1):221. doi:10.1186/s13059-016-1082-1
Sela H, Spiridon LN, Petrescu AJ, Akerman M, Mandel-Gutfreund Y, Nevo E, Loutre C, Keller B, Schulman AH, Fahima T (2012) Ancient diversity of splicing motifs and protein surfaces in the wild emmer wheat (Triticum dicoccoides) LR10 coiled coil (CC) and leucine-rich repeat (LRR) domains. Mol Plant Pathol 13:276–287. doi:10.1111/j.1364-3703.2011.00744.x
Shao ZQ, Xue JY, Wu P, Zhang YM, Wu Y, Hang YY, Wang B, Chen JQ (2016) Large-scale analyses of angiosperm nucleotide-binding site-leucine-rich repeat genes reveal three anciently diverged classes with distinct evolutionary patterns. Plant Physiol 170(4):2095–2109. doi:10.1104/pp.15.01487
Singh A, Pallavi JK, Gupta P, Prabhu KV (2011) Identification of microsatellite markers linked to leaf rust adult plant resistance (APR) gene Lr48 in wheat. Plant Breed 130(1):31–34. doi:10.1111/j.1439-0523.2010.01820.x
Smith PH, Hadfield J, Hart NJ, Koebner RM, Boyd LA (2007) STS markers for the wheat yellow rust resistance gene Yr5 suggest a NBS-LRR-type resistance gene cluster. Genome 50:259–265. doi:10.1139/g07-004
Steuernagel B, Periyannan SK, Hernández-Pinzón I, Witek K, Rouse MN, Yu G, Hatta A, Ayliffe M, Bariana H, Jones JD, Lagudah ES, Wulff BB (2016) Rapid cloning of disease-resistance genes in plants using mutagenesis and sequence capture. Nat Biotechnol 34(6):652–655. doi:10.1038/nbt.3543
Thind AK, Wicker T, Šimková H, Fossati D, Moullet O, Brabant C, Vrána J, Doležel J, Krattinger SG (2017) Rapid cloning of genes in hexaploid wheat using cultivar-specific long-range chromosome assembly. Nat Biotechnol. doi:10.1038/nbt.3877 Published online
Tommasini L, Yahiaoui N, Srichumpa P, Keller B (2006) Development of functional markers specific for seven Pm3 resistance alleles and their validation in the bread wheat gene pool. TheorAppl Genet 114:165–175. doi:10.1007/s00122-006-0420-1
Tsilo TJ, Jin Y, Anderson JA (2007) Microsatellite markers linked to stem rust resistance allele Sr9a in wheat. Crop Sci 47:2013–2020. doi:10.2135/cropsci2007.02.0087
Tsilo TJ, Jin Y, Anderson JA (2010) Identification of flanking markers for the stem rust resistance gene Sr6 in wheat. Crop Sci 50:1967–1970. doi:10.2135/cropsci2009.11.0648
Urbach JM, Ausubel FM (2017) The NBS-LRR architectures of plant R-proteins and metazoan NLRs evolved in independent events. Proc Natl Acad Sci USA 114(5):1063–1068. doi:10.1073/pnas.1619730114
Wang D, Wang X, Mei Y, Dong H (2016) The wheat homolog of putative nucleotide-binding site-leucine-rich repeat resistance gene TaRGA contributes to resistance against powdery mildew. Funct Integr Genom 16(2):115–126. doi:10.1007/s10142-015-0471-y
Wiersma AT, Pulman JA, Brown LK, Cowger C, Olson EL (2017) Identification of Pm58 from Aegilops tauschii. Theor Appl Genet 130(6):1123–1133. doi:10.1007/s00122-017-2874-8
Xiang C, Feng JY, Wang MN, Chen XM, See DR, Wan AM, Wang T (2016) Molecular mapping of stripe rust resistance gene Yr76 in winter club wheat cultivar Tyee. Phytopathology 106(10):1186–1193
Xing LP, Hu P, Liu JQ, Cui CF, Wang H, Di ZC, Zhou S, Xu JF, Gao L, Huang ZP, Cao AZ (2017) NLR1-V, a CC-NBS-LRR encoding gene, is a potential candidate gene of the wheat powdery mildew resistance gene Pm21. bioRxiv. doi:10.1101/114058
Yahiaoui N, Srichumpa P, Dudler R, Keller B (2004) Genome analysis at different ploidy levels allows cloning of the powdery mildew resistance gene Pm3b from hexaploid wheat. Plant J 37:528–538. doi:10.1046/j.1365-313X.2003.01977.x
Zhan H, Li G, Zhang X, Li X, Guo H, Gong W, Jia J, Qiao L, Ren Y, Yang Z, Chang Z (2014) Chromosomal location and comparative genomics analysis of powdery mildew resistance gene Pm51 in a putative wheat-Thinopyrum ponticum introgression line. PLoS ONE 9(11):e113455. doi:10.1371/journal.pone.0113455
Zhou F, Kurth J, Wei F, Elliott C, Valè G, Yahiaoui N, Keller B, Somerville S, Wise R, Schulze-Lefert P (2001) Cell-autonomous expression of barley Mla1 confers race-specific resistance to the powdery mildew fungus via a Rar1-independent signaling pathway. Plant Cell 13(2):337–350. doi:10.1105/tpc.13.2.337
Zhou T, Wang Y, Chen JQ, Araki H, Jing Z, Jiang K, Shen J, Tian D (2004) Genome-wide identification of NBS genes in japonica rice reveals significant expansion of divergent non-TIR-NBS-LRR genes. Mol Genet Genomics 271(4):402–415. doi:10.1007/s00438-004-0990-z
Acknowledgements
This work was supported by the National Key Research and Development Plan of China (2016YFD0102004-07) and the Shanxi Province Science Foundation for Youths (2015021145).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Qiao, L., Zhang, X., Li, X. et al. Development of NBS-related microsatellite (NRM) markers in hexaploid wheat. Euphytica 213, 256 (2017). https://doi.org/10.1007/s10681-017-2039-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-017-2039-5