Abstract
Powdery mildew resistance from Thinopyrum intermedium was introgressed into common wheat (Triticum aestivum L.). Genetic analysis of the F1, F2, F3 and BC1 populations from powdery mildew resistant line CH5025 revealed that resistance was controlled by a single dominant allele. The gene responsible for powdery mildew resistance was mapped by the linkage analysis of a segregating F2 population. The resistance gene was linked to five co-dominant genomic SSR markers (Xcfd233, Xwmc41, Xbarc11, Xgwm539 and Xwmc175) and their most likely order was Xcfd233–Xwmc41–Pm43–Xbarc11–Xgwm539–Xwmc175 at 2.6, 2.3, 4.2, 3.5 and 7.0 cM, respectively. Using the Chinese Spring nullisomic-tetrasomic and ditelosomic lines, the polymorphic markers and the resistance gene were assigned to chromosome 2DL. As no powdery mildew resistance gene was previously assigned to chromosome 2DL, this new resistance gene was designated Pm43. Pm43, together with the identified closely linked markers, could be useful in marker-assisted selection for pyramiding powdery mildew resistance genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Powdery mildew, caused by Blumeria graminis f. sp. tritici, is a globally important disease of wheat (Triticum aestivum L.). The most feasible means of controlling the disease and reducing yield loss is the breeding of resistant varieties. To date, 54 formally designated Pm resistance genes including alleles have been identified, mapped to 40 loci (Pm1–Pm42), and assigned to specific chromosomes or chromosome arms. This includes three recessive alleles (Pm5, Pm9, Pm26) (Huang and Röder 2004; Zhu et al. 2005; Miranda et al. 2006, 2007; Blanco et al. 2008; Perugini et al. 2008; Spielmeyer et al. 2008; Lillemo et al. 2008; McIntosh et al. 2008, personal communication). Of these loci, 26 genes were transferred from wild relatives of wheat including T. turgidum var. dicoccoides and var. dicoccum, T. timopheevii, T. monococcum, Aegilops squarrosa, Ae. speltoides, Ae. longissima, Ae. ovata, and from more distant species, including Secale cereale and Dasypyrum villosum. Unfortunately, major host resistance alleles often become ineffective because of frequent changes in the pathogen population, especially when a single resistance gene is deployed over a wide area. Therefore, effective and durable resistance requires new sources of resistance and their use in combinations.
Thinopyrum intermedium (Host) Barkworth and Dewey (2n = 6x = 42, JJsS) has been hybridized extensively with wheat and proven to be a valuable source of genes for disease resistance. Partial wheat-Th. intermedium amphiploids are potential tertiary gene pools for wheat improvement, even possessing biotic stress resistances that are not common in wheat, such as resistances to wheat streak mosaic virus (WSMV), its vector the wheat curl mite (WCM) (Aceria tosichella Keifer) and barley yellow dwarf virus (BYDV) (Friebe et al. 1996; Chen 2005; Fedak and Han 2005). Excellent resistance to Fusarium head blight was found on a Th. intermedium chromosome that was substituted for wheat chromosome 2D (Han et al. 2003). Genes for resistance to leaf rust and stem rust were incorporated into wheat and tagged with molecular markers (Autrique et al. 1995; Fedak 1999). Recently, resistance to powdery mildew was found in a partial amphiploid (Liu et al. 2005) and a substitution line (Liu and Wang 2005) derived from Th. intermedium, but no translocation from Thinopyrum to wheat chromosomes was reported.
Molecular markers, including RAPD, RFLP, SSR, and RGAP, are useful tools for gene mapping in wheat. Microsatellite or SSR linkage maps developed for wheat provide the extensive genome coverage that is required for marker-assisted breeding strategies (Röder et al. 1998; Stephenson et al. 1998; Gupta et al. 1999, 2002; Pestova et al. 2000; Paillard et al. 2003; Somers et al. 2004). Linked microsatellite markers have already been found for Pm1e (Singrün et al. 2003), Pm3g (Bougot et al. 2002), Pm3h, Pm3i, Pm3j (Huang et al. 2004), Pm4a (Ma et al. 2004), Pm5e (Huang et al. 2003), Pm16 (Chen et al. 2005), Pm24 (Huang et al. 2000), Pm27 (Järve et al. 2000), Pm30 (Liu et al. 2002), Pm31 (Xie et al. 2003), Pm33 (Zhu et al. 2005), Pm34 (Miranda et al. 2006), Pm35 (Miranda et al. 2007), Pm36 (Blanco et al. 2008), Pm37 (Perugini et al. 2008), Pm38 (Spielmeyer et al. 2008) and Pm39 (Lillemo et al. 2008). Some of these markers have been successfully used in map-based cloning (Yahiaoui et al. 2004), and marker-assisted selection and pyramiding of the resistance genes (Liu et al. 2000), as well as understanding the relationships between different genes (Singrün et al. 2003).
A program for introgression of alien resistance genes into wheat was initiated at the Institute of Crop Genetics, Shanxi Academy of Agricultural Sciences, Yaiyuan. The program aimed to transfer resistances to powdery mildew and yellow rust into wheat from Th. intermedium and Th. ponticum. To date, many multi-resistance lines have been developed by crossing susceptible wheat cultivars with resistant partial amphiploids as donor parents. The objectives of this study were to determine the inheritance, chromosomal location, and linkage to molecular markers, of gene(s) for resistance to powdery mildew in the Th. intermedium-derived line CH5025.
Materials and methods
Plant materials
The materials used in this study were Th. intermedium (accession Z1141) with the genomic formula E1E2St (Wang et al. 1994) or JJsS (Chen et al. 1998); two partial amphiploids, TAI7045 and 78829, derived from accession Z1141; wheat genotypes ‘CH5025’, ‘76216–96’, ‘Jing 411’, ‘Jinchun 5’, ‘Taiyuan 768’ and ‘Chinese Spring’; and various ‘Chinese Spring’ nullisomic-tetrasomic (NT) stocks. CH5025 and CH5065 are homogeneous BC2F4-derived wheat lines derived from the cross 76216-96/TAI7045//2*Jing 411. CH5025 is resistant to powdery mildew whereas CH5065 is susceptible. TAI7045, the resistance donor of CH5025, was derived from the cross of Taiyuan 768/Z1141//Jinchun 5.
To investigate the inheritance of powdery mildew resistance introgressed from Th. intermedium, CH5025 was crossed to susceptible cultivars CH5065 and Jintai 170 to yield segregating populations. The F2, F3 and BC1 were tested for segregation of powdery mildew resistance. An F2 population from CH5025/CH5065 was further used for microsatellite screening and gene mapping. The F2 plants were self-pollinated to produce F2-derived F3 families. The mapping population consisted of 177 F2 plants and the derived F3 families, the difference being due to random loss of eight F2 plants.
Evaluation of powdery mildew responses
Seedlings of Th. intermedium, two partial amphiploids and seven wheat cultivars/lines (Table 1), were evaluated in the greenhouse using four B. graminis f. sp. tritici (Bgt) isolates (E09, E20, E21 and E26) and methods described by Xiang et al. (1994). E09, a prevalent pathotype in the Beijing area, is virulent to Pm1, Pm3a, Pm3c, Pm5, Pm7, Pm8, Pm17, and Pm19 (Zhou et al. 2005). E20 and E21 are the most widely virulent pathotypes in China and are virulent to most of the Pm genes including Pm4a, Pm4b, PmPs5A and Pm33. E26 is virulent to Pm4b and Pm33, but avirulent to Pm4a and PmPs5A (Wang et al. 2005; Zhou et al. 2005).
Six plants of each cultivar and differential line were grown in 70 × 45 × 18 cm flat plastic trays. The highly susceptible cv. Jingshuang 16 was used as a control. Inoculation with each of the four isolates was performed in separate trays when the first leaves were fully expanded. Host reactions were scored 7–10 days after inoculation, when the susceptible checks were heavily infected. A 0–4 infection type scale (Shi et al. 1987) was used to describe host responses to infection, viz., 0 = no visible symptoms; 0; = hypersensitive necrotic flecks; 1 = minute colonies with few conidia produced; 2 = colonies with moderately developed hyphae, but few conidia; 3 = colonies with well-developed hyphae and abundant conidia, but unjoined colonies; and 4 = colonies with well-developed hyphae and abundant conidia, with mostly joined colonies. Scores of 0–2 were classified as resistant and 3–4 as susceptible.
Isolate E09, which is avirulent to both CH5025 seedlings and adults, but virulent to CH5065 and Jintai 170, was used to test F1, F2, and BC1 populations derived from CH5025/CH5065 and Jintai 170/CH5025 (Table 2). Seeds from the parents, F1, F2, F3 and BC1 populations were planted in the greenhouse. Twenty seeds for each parent and F1, 200 seeds of the F2, 100 seeds of the BC1 and 15 seeds for each of the F2-derived F3 families were planted in a randomized design with 15–20 plants in a 1.2 m row, 25 cm apart. Susceptible spreaders of cv. Jingshuang 16 and SY95-71 were planted in every tenth row. Seedlings at the two-leaf stage were inoculated with Bgt race E09, and evaluated at the seedling and heading stages. The infection types (ITs) were rated using the 0–4 scale (Shi et al. 1987). Adult plant reactions were scored twice, at the ear emergence stage and at the milky ripe stage, using a modified 0–9 scale (Sheng et al. 1986). To determine the genotypes of F2 plants from CH5025/CH5065, the F2-derived F3 families were tested with the same race as used in the F2 tests.
Bulk segregant analysis
Total DNA was extracted as in Sharp et al. (1988), from healthy leaves of the parents and F2 plants of the CH5025/CH5065 cross. For bulk segregant analysis (BSA) (Michelmore et al. 1991), the resistant (Br) and susceptible (Bs) bulks were made from equal amounts of DNA from 10 resistant and 10 susceptible F2 segregants, respectively. The bulk pools were used with parent samples to identify markers that showed polymorphisms between the four samples. These were used to further analyze the F2 population to determine linkages between SSR markers and the resistance gene in CH5025.
Microsatellite marker analysis
Wheat microsatellite markers evenly distributed across the A, B and D genomes designated as either Xgwm for Gatersleben (Germany) wheat microsatellite (Röder et al. 1998), or Xwmc for Wheat Microsatellite Consortium (Gupta et al. 2002) were used to detect polymorphism between parents and resistant and susceptible bulks. The resulting polymorphism markers were subsequently genotyped in the F2 individuals to determine the genetic linkage between the powdery mildew resistance gene(s) and markers. Additional markers BARC (Beltsville Agriculture Research Centre) and CFD (Pierre Sourdille) markers located on 2D, showing polymorphism between resistant and susceptible bulks, were also tested on the F2 mapping population.
PCR and product analysis
The polymerase chain reaction (PCR) for each SSR marker was performed in a PTC200 Peltier Thermal Cycler (Bio-Rad Inc., USA) in a total volume of 20 μl containing 10 mM Tris–HCl, pH 8.3, 50 mM KCl, 1.5 mM MgCl2, 0.2 mM dNTPs, 1U Taq DNA polymerase, 50 ng each primer, and 80–100 ng total DNA. PCR was performed at 94°C for 5 min, followed by 35 cycles of 94°C for 1 min; 50, 55, or 60°C (based on primer annealing temperature) for 1 min; and 72°C for 1 min, with final incubation at 72°C for 10 min before cooling to 4°C. Each PCR product was denatured with 8 μL loading buffer (98% formamide, 10 mM EDTA, pH 8.0, 0.25% bromophenol blue, and 0.25% xylene cyanol), heating for 5 min at 95°C, and chilling on ice before loading 4–6 μL on a 6% polyacyamide gel (19:1 acrylamide–bis) with 8 mol urea/L and 1 × TBE (90 mM Tris–borate, pH 8.3, 2 mM EDTA). Samples were run at 2,000 V, 30 mA, and 50 W for approximately 1 h, and visualized by silver staining.
Chromosome assignment and linkage analysis
Chromosomal locations of the linked microsatellite markers were confirmed using Chinese Spring nullisomic 2D-tetrasomic 2A (N2DT2A) and ditelosomic 2DL (Dt2DL) lines kindly provided by the Wheat Genetics Resource Centre, Kansas State University. Genomic DNA from N2DT2A, Dt2DL, euploid Chinese Spring, CH5025 and CH5065 were used in PCR of microsatellite markers putatively linked to the CH5025 resistance gene. As positive controls, the reactions included DNA from N2DT2A and Dt2DL amplified with a primer pair that amplify sequences in the A genome.
Chi-squared (χ2) tests for goodness-of-fit were used to test for deviations of observed data from theoretically expected segregations. Linkages between DNA markers and the resistance gene were established with MAPMAKER/Exp version 3.0b software (Lincoln et al. 1993) with a LOD threshold >3.0. Map distances were determined using the Kosambi mapping function (Kosambi 1944) and loci were ordered using the ‘sequence’ and ‘compare’ commands, with an LOD threshold score 3.0.
Results
Origin of the powdery mildew resistance
CH5025, TAI7045 and the Th. intermedium parent were resistant to Bgt isolates E09, E20, E21 and E26, whereas the wheat parents or lines CH5065, and Taiyuan 768, Jinchun 5, 76216-96 and Jing 411, were susceptible (Table 1). These results demonstrated that CH5025 conferred resistance to powdery mildew similar to its donor TAI7045, and the parent Th. intermedium. Partial amphiploid 78829 was susceptible (Table 1). Although derived from the same accession of Th. intermedium as 78829, TAI7045 had a different chromosome composition.
Inheritance of the powdery mildew resistance
The infection types of the F1s, and the segregation patterns of the F2, and BC1 populations are shown in Table 2. When inoculated with isolate E09 at the seedling and adult stages, F1 plants from all crosses showed infection types similar to the resistant parent, indicating that the resistance was dominant. Segregations of the F2 populations from the resistant/susceptible crosses and the BC1 population (Table 2) were consistent with those expected for segregations at a single locus. When tested with the same race, the F3 lines from CH5025/CH5065 were scored 48 homozygous resistant (RR):91 segregating (Rr):38 homozygous susceptible (rr), confirming the single gene segregation (χ21:2:1 = 1.03, P 2df > 0.5) (Table 3). In addition the pooled segregation for the F3 segregating lines was 867 resistant:309 susceptible (χ23:1 = 0.95, P 1df > 0.25).
Mapping of the powdery mildew resistance gene
Identification of Microsatellite markers linked to Pm43
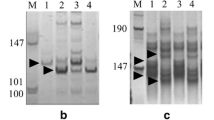
The F2 population of CH5025/CH5065 was used for mapping the resistance gene. About 129 (37.0%) of the 349 SSR markers chosen for the initial primer screening were polymorphic between CH5025 and CH5065. Among the polymorphic markers, only Xwmc41 produced a polymorphism between the contrasting bulks, and a fragment of about 190 bp was associated with the CH5025 allele (Fig. 1).
Relationship between F3 line responses (F2 genotypes) of CH5025/CH5065 and the bands amplifies by SSR primer pair WMC41. R homozygous resistant, Se segregating, S homozygous susceptible, Se* recombinant genotype, M 100-bp DNA ladder. Arrow on the left side indicates the fragment linked to the resistance gene
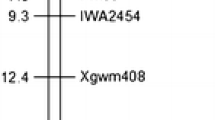
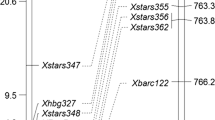
Because Xwmc41 was previously mapped to the long arm of chromosome 2D (Somers et al. 2004), 54 additional SSR primer pairs on the same chromosome arm were tested. Four polymorphic markers, Xcfd233, Xbarc11, Xgwm539 and Xwmc175, were associated with resistance in both the bulk segregant pools and the parents, and were used to genotype all the 177 plants of the F2 population. Linkage analyses confirmed the genetic associations of the five SSR markers with powdery mildew resistance. The F2 population segregated 1:2:1 for all five markers (Table 3). Analyses with MAPMAKER/EXP also showed linkage between the markers and Pm43; Xcfd233 and Xwmc41 were proximal to the resistance gene with genetic distances of 2.6 and 2.3 cM respectively, and Xbarc11, Xgwm539, and Xwmc175 were distal with respective genetic distances of 4.2, 3.5 and 7.0 cM. The most likely order is shown in Fig. 2.
Chromosomal assignment
Based on the reported chromosomal locations of the five linked microsatellite markers (Somers et al. 2004), Pm43 was putatively assigned to the long arm of chromosome 2D. However, microsatellite markers are not always chromosome-specific (Plaschke et al. 1996). Of the marker loci linked to Pm43, Xwmc41, Xcfd233, and Xgwm539 were all assigned to the chromosome arm 2DL, whereas Xwmc175 was assigned to 2DL and 2BL, and Xbarc11 to 2DL and 5BL (http://wheat.pw.usda.gov/cgi-bin/graingenes). Therefore, the locations of the linked microsatellite loci were verified using CS nulli-tetrasomic and ditelosomic lines. Four of the five microsatellite primer pairs, WMC41, CFD233, BARC11 and GWM529, amplified products of the expected size in Chinese Spring and the CS Dt2DL line, but no PCR product was observed in the nulli-tetrasomic N2DT2A line for any of them (Fig. 3). The absence of PCR products in the N2DT2A line and their presence in the Dt2DL line further confirmed the assignment of the linked microsatellite markers to the long arm of chromosome 2D. Based on its origin and map location the dominant allele from Th. intermedium was apparently new and was therefore designated Pm43.
Discussion
Wild relatives and related species have been widely used as genetic resources for introgression of useful traits into crops. Th. intermedium (2n = 6x = 42, JJsS) is an important perennial Triticeae species with considerable potential for wheat improvement. As they are readily crossed with wheat, Th. intermedium-derived partial amphiploids have been widely used for attempted introgressions of useful traits into wheat, including resistance to viral diseases. Resistances to BYDV, WSMV and WCM were found in partial amphiploids and addition lines derived from Th. intermedium (Larkin et al. 1995; Friebe et al. 1996; Chen et al. 2003), and some genes were incorporated into wheat and tagged with molecular markers (Ayala et al. 2001; Qi et al. 2007). Excellent resistance to Fusarium head blight was found on a Th. intermedium chromosome that was substituted for chromosome 2D in wheat (Han et al. 2003).
Th. intermedium is immune to wheat powdery mildew. A resistance gene was recently found in partial amphiploids and in the substitution line 2J(2D), in which a J-chromosome of Th. intermedium was substituted for chromosome 2D in wheat (Liu and Wang 2005; Liu et al. 2005). However, there is no published report of transfer of powdery mildew resistance from this species to a wheat chromosome. In the present study, a novel powdery mildew resistance gene was transferred from Th. intermedium into common wheat, using a resistant partial amphiploid as a bridging parent in crosses with susceptible wheat lines. After selection in the BC1F2 for fertility and resistance to powdery mildew, followed by a second backcross with a susceptible wheat cultivar, and further selfing and selection of resistant plants, stable hexaploid introgression lines with good agronomic appearance and resistance to powdery mildew were obtained. There are no reports of resistance to powdery mildew in well known partial amphiploids, including the Zhong 1 to Zhong 5 series (Sun 1981), 78829 (Li et al. 1985) and TAF46 (Cauderon et al. 1973). CH5025 was produced by crossing the resistant partial amphiploid TAI7045 with a susceptible wheat cultivar, but no signal was observed in in situ hybridization experiments when using genomic DNA from either Th. intermedium or Pseudoroegneria strigosa (S genome, 2n = 2x = 14) as a probe (data not shown). This indicated that the introgressed Th. intermedium segment in CH5025 was very small and cytologically undetectable. In fact, some recent studies had demonstrated that some traits of interest were transferred to recipient genotypes without detectable cytological or genetic changes (Dong et al. 2004; Kuraparthy et al. 2007). In our powdery mildew resistance testing, the donor TAI7045 was resistant to all Bgt isolates, whereas 78829 was susceptible (Table 1). Although TAI7045 was derived from the same accession of Th. intermedium as 78829 and the Zhong series, the chromosomal compositions of their alien genome was different. Of the partial amphiploids mentioned above, only 78829 was reported to contain a complete alien S genome (Zhang et al. 1996). TAI7045 had 14 alien chromosomes including eight S-genome chromosomes, four Js-genome chromosomes, and one pair of S-Js translocation chromosomes (Chang 1999; Chang et al. 2001), suggesting that, when TAI7045 was the donor, resistance to powdery mildew was on the Js-genome chromosome.
The transfer of Thinopyrum chromatin specifying resistance to BYDV, WSMV, leaf rust, stem rust, FHB, and powdery mildew into wheat has shown that, in compensating substitution or translocation lines, chromosomes of Thinopyrum and the D genome were often involved (Friebe et al. 1996; Han et al. 2003; Liu et al. 2005; Qi et al. 2007). This is possibly because the Thinopyrum genome shares more homology with the D, than with the A or B genomes. In addition, pairing between T. aestivum and Th. intermedium chromosomes occurred only with J or Js chromosomes, demonstrating that the relationships between common wheat genomes and the Th. intermedium J and Js genomes are much closer than with the S genome (Chen et al. 2001). The low levels of chromosome pairing in wheat crossed with Th. intermedium could be due to partial homology between wheat and alien chromosomes. Based on these results and our own, it appears that a possible mechanism for the introgression of the alien resistance gene in CH5025 was is through homeologous chromosome pairing and recombination between the Js genome of Th. intermedium and the D genome. However, such spontaneous transfer may have gone undetected because of the limitations of alien introgression research often based on cytological methods, which needs to be further confirmed by molecular characterization.
To investigate the inheritance of the powdery mildew resistance, we developed segregating populations by crossing the resistant line CH5025 with susceptible wheat lines. The segregation patterns of all populations confirmed that the resistance was controlled by a single dominant allele. The establishment of linkage between molecular markers and the resistance gene not only confirmed a precise chromosomal placement, but will also be helped in judicious deployment of resistance genes through marker-aided selection. Five co-dominant microsatellite markers were linked to Pm43. Since microsatellite markers are not always chromosome-specific (Plaschke et al. 1996), the locations of four of the linked microsatellite loci were verified with Chinese Spring nulli-tetrasomic and ditelosomic lines (Fig. 3). Although a ditelosomic 2DS line was not available, the presence of PCR products of the same size in euploid Chinese Spring and CS Dt2DL and their absence in CS N2DT2A confirmed the locations of the markers on 2DL.
Molecular markers closely linked to resistance genes provide powerful tools for marker-assisted breeding programs to enable transfer of resistance genes without performing disease tests. Introgression of disease resistance genes from related species or genera into wheat has become crucial to the development of resistant genotypes. Because resistance to powdery mildew in many Chinese cultivars has gradually decreased, the introgression of Pm43 into wheat and the identification of closely flanking markers may be beneficial for increasing the overall diversity of available resistance genes with potential to provide more comprehensive and durable protection against the disease.
References
Autrique E, Singh RP, Tanksley SD, Sorrells ME (1995) Molecular markers for four leaf rust resistance genes introgressed into wheat from wild relatives. Genome 38:75–83
Ayala L, Henry M, Gonzalez-de-Leon D, van Ginkel M, Mujeeb-Kazi A, Keller B, Khairallah M (2001) A diagnostic molecular marker allowing the study of Th. intermedium-derived resistance to BYDV in bread wheat segregating populations. Theor Appl Genet 102:942–949
Blanco A, Gadaleta A, Cenci A, Carluccio AV, Abdelbacki AMM, Simeone R (2008) Molecular mapping of the novel powdery mildew resistance gene Pm36 introgressed from Triticum turgidum var. dicoccoides in durum wheat. Theor Appl Genet 117:135–142
Bougot Y, Lemoine J, Pavoine MT, Barloy D, Doussinault G (2002) Identification of a microsatellite associated with Pm3 resistance alleles to powdery mildew in wheat. Plant Breed 121:325–329
Cauderon Y, Saigne B, Dauge M (1973) The resistance to wheat rusts of Agropyron intermedium and its use in wheat improvement. In: Sears ER, Sears LMS (eds) Proc 4th Int Wheat Genet Symp. University of Missouri Agricutural Research Station, Columbia, pp 401–407
Chang ZJ (1999) Production and molecular cytogenetic characterization of several Thinopyrum intermedium-derived wheat germplasm lines. PhD Dissertation, Sichuan Agricultural University, China
Chang ZJ, Yuan ZY, Guo XR, Yang ZJ, Ren ZL (2001) Production and genome analysis of a new 56-chromosome line derived from wheat × Agropyron intermedium. Proc Int Wheat Genetics and Breeding Symp, May 9–11, 2001, Zhengzhou, Henan. China Agricultural Scientech Press, Beijing, 237–241
Chen Q (2005) Detection of alien chromatin introgression from Thinopyrum into wheat using S genomic DNA as a probe-a landmark approach for Thinopyrum genome research. Cytogenet Genome Res 109:350–359
Chen Q, Conner RL, Laroche A, Thomas JB (1998) Genome analysis of Thinopyrum intermedium and Th. ponticum using genomic in situ hybridization. Genome 41:580–586
Chen Q, Conner RL, Laroche A, Ahmad F (2001) Molecular cytogenetic evidence for a high level of chromosome pairing among different genomes in Triticum aestivum-Thinopyrum intermedium hybrids. Theor Appl Genet 102:847–852
Chen Q, Conner RL, Li HJ, Sun SC, Ahmad F, Laroche A, Graf RF (2003) Molecular cytogenetic discrimination and reaction to wheat streak mosaic virus and the wheat curl mite in the Zhong series of wheat-Thinopyrum intermedium partial amphiploids. Genome 46:135–145
Chen XM, Luo YH, Xia XC, Xia LQ, Chen X, Ren ZL, He ZH, Jia JZ (2005) Chromosomal location of powdery mildew resistance gene Pm16 in wheat using SSR marker analysis. Plant Breed 124:225–228
Dong YS, Bu XL, Luan YS, He MY, Liu B (2004) Molecular characterization of a cryptic wheat–Thinopyrum intermedium translocation line: evidence for genomic instability in nascent allopolyploid and aneuploid lines. Genet Mole Bio 27:237–241
Fedak G (1999) Molecular aids for integration of alien chromatin through wide crosses. Genome 42:584–591
Fedak G, Han F (2005) Characterization of derivatives from wheat-Thinopyrum wide crosses. Cytogenet Genome Res 109:350–359
Friebe B, Jiang J, Raupp WJ, McIntosh RA, Gill BS (1996) Characterization of wheat-alien translocations conferring resistance to diseases and pests: current status. Euphytica 91:59–87
Gupta PK, Varshney RK, Sharma PC, Ramesh B (1999) Molecular markers and their applications in wheat breeding. Plant Breed 118:369–390
Gupta PK, Balyan HS, Edwards KJ, Isaac P, Korzun V, Röder M, Gautier MF, Joudrier P, Schlatter AR, Dubcovsky J, De la Pena RC, Khairallah M, Penner G, Hayden MJ, Sharp P, Keller B, Wang RCC, Hardouin JP, Jack P, Leroy P (2002) Genetic mapping of 66 new microsatellite (SSR) loci in bread wheat. Theor Appl Genet 105:413–422
Han FP, Fedak G, Benabdelmouna A, Armstrong KC, Ouellet T (2003) Characterization of six wheat × Thinopyrum intermedium derivatives by GISH, RFLP and multicolor GISH. Genome 46:490–495
Huang XQ, Röder MS (2004) Molecular mapping of powdery mildew resistance genes in wheat: a review. Euphytica 137:203–223
Huang XQ, Hsam SLK, Zeller FJ, Wenzel G, Mohler V (2000) Molecular mapping of wheat powdery mildew resistance gene Pm24 and marker validation for molecular breeding. Theor Appl Genet 101:407–414
Huang XQ, Wang LX, Xu MX, Röder MS (2003) Microsatellite mapping of the powdery mildew resistance gene Pm5e in common wheat (Triticum aestivum L.). Theor Appl Genet 106:858–865
Huang XQ, Hsam SLK, Mohler V, Röder MS, Zeller F (2004) Genetic mapping of three alleles at the Pm3 locus conferring powdery mildew resistance in common wheat (Triticum aestivum L.). Genome 47:1130–1136
Järve K, Peusha HO, Tsybalova J, Tamm S, Devos KM, Enno TM (2000) Chromosomal location of a Triticum timopheevi derived powdery mildew resistance gene transferred to common wheat. Genome 43:377–381
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Kuraparthy V, Sood S, Chhuneja P, Dhaliwal HS, Kaur S, Bowder RL, Gill BS (2007) A cryptic wheat–Aegilops triuncialis translocation with leaf rust resistance gene Lr58. Crop Sci 47:1995–2003
Larkin PJ, Banks PM, Lagudah ES, Appels R, Chen X, Xin XY, Ohm HW, McIntosh RA (1995) Disomic Thinopyrum intermedium addition lines in wheat with barley yellow dwarf virus resistance and with rust resistance. Genome 38:385–394
Li ZS, Rong S, Chen SY, Zhong GC, Mu SM (1985) Wheat wide hybridization. Chinese Scientific Press, Beijing, pp 52–58
Lillemo M, Asalf B, Singh RP, Huerta-Espino J, Chen XM, He ZH, Bjørnstad Å (2008) The adult plant rust resistance loci Lr34/Yr18 and Lr46/Yr29 are important determinants of partial resistance to powdery mildew in bread wheat line Saar. Theor Appl Genet 116:1155–1166
Lincoln SE, Daly MJ, Lander ES (1993) Constructing linkage maps with MAPMAKER/Exp Version 3.0. A tutorial reference manual, 3rd edn. Whitehead Institute for Medical Res, Cambridge
Liu SB, Wang HG (2005) Characterization of a wheat–Thinopyron intermedium substitution line with resistance to powdery mildew. Euphytica 143:229–233
Liu J, Liu D, Tao W, Li W, Wang S, Chen P, Cheng S, Gao D (2000) Molecular marker-facilitated pyramiding of different genes for powdery mildew resistance in wheat. Plant Breed 119:21–24
Liu ZY, Sun QX, Ni ZF, Nevo E, Yang TM (2002) Molecular characterization of a novel powdery mildew resistance gene Pm30 in wheat originating from wild emmer. Euphytica 123:21–29
Liu SB, Wang HG, Zhang XY, Li XF, Li DY, Duan XY, Zhou YL (2005) Molecular Cytogenetic identification of a wheat-Thinopyron intermedium (Host) Barkworth and DR Dewey partial amphiploid resistant to powdery mildew. J Integr Plant Biol 47:726–733
Ma ZQ, Wei JB, Chen SH (2004) PCR based markers for the powdery mildew resistance gene Pm4a in wheat. Theor Appl Genet 109:140–145
McIntosh RA, Yamazaki Y, Dubcovsky J, Rogers J, Morris C, Somers DJ, Appels R, Devos KM (2008) Catalogue of Gene Symbols for Wheat. Proc 11th Int Wheat Genet Symp, University of Sydney Press, Australia. http://wheat.pw.usda.gov/GG2/Triticum/wgc/2008/
Michelmore RM, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci USA 88:9828–9832
Miranda LM, Murphy JP, Leath S, Marshall DS (2006) Pm34: a new powdery mildew resistance gene transferred from Aegilops tauschii Coss. to common wheat (Triticum aestivum L.). Theor Appl Genet 113:1497–1504
Miranda LM, Murphy JP, Marshall DS, Cowger C, Leath S (2007) Chromosomal location of Pm35, a novel Aegilops tauschii derived powdery mildew resistance gene introgressed into common wheat (Triticum aestivum L.). Theor Appl Genet 114:1451–1456
Paillard S, Schnurbusch T, Winzeler M, Messmer M, Sourdille P, Abderhalden O, Keller B, Schachermayr G (2003) An integrative genetic linkage map of wheat. Theor Appl Genet 107:1235–1242
Perugini LD, Murphy JP, Marshall D, Brown-Guedira G (2008) Pm37, a new broadly effective powdery mildew resistance gene from Triticum timopheevii. Theor Appl Genet 116:417–425
Pestova E, Ganal MW, Röder MS (2000) Isolation and mapping of microsatellite markers specific for the D genome of bread wheat. Genome 43:689–697
Plaschke JB, Börner A, Wendehake K, Ganal MW, Röder MS (1996) The use of aneuploids for the chromosomal assignment of microsatellite loci. Euphytica 89:33–40
Qi L, Friebe B, Zhang P, Gill BS (2007) Homoeologous recombination, chromosome engineering and crop improvement. Chromosome Res 15:3–19
Röder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Sharp PJ, Kreis M, Shewry PR, Gale MD (1988) Location of β-amylase sequence in wheat and its relatives. Theor Appl Genet 75:289–290
Sheng BQ, Duan XY, Zhang XX (1986) The improved adult resistance scales of wheat powdery mildew. Plant Prot Sin 3:44–45
Shi QM, Zhang XX, Duan XY (1987) Identification of isolates of Blumeria graminis f. sp. tritici. Sci Agric Sin 20:64–70
Singrün CH, Hsam SL, Zeller FJ, Mohler V (2003) Powdery mildew resistance gene Pm22 is a member of the complex Pm1 locus in common wheat (Triticum aestivum L). Theor Appl Genet 106:1420–1424
Somers DJ, Isaac P, Edwards K (2004) A high density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Spielmeyer W, Singh RP, McFadden H, Wellings CR, Huerta-Espino J, Kong X, Appels R, Lagudah ES (2008) Fine scale genetic and physical mapping using interstitial deletion mutants of Lr34/Yr18: a disease resistance locus effective against multiple pathogens in wheat. Theor Appl Genet 116:481–490
Stephenson P, Bryan G, Kirby J, Collins A, Devos K, Busso C, Gale M (1998) Fifty new microsatellite loci for the wheat genetic map. Theor Appl Genet 100:564–568
Sun SC (1981) The approach and methods of breeding new varieties and new species from Agrotriticum hybrids. Acta Agron Sin 7:51–58
Wang RR-C, van Bothmer R, Dvórak R, Fedak G, Linde-Laursen I, Muramatsu M (1994) Genome symbols in the Triticeae (Poaceae). In: Wang RR-C, Jensen KB, Jaussi C (eds) Proc 2nd Int Triticeae Symp. Utah State University Press, Logan, pp 29–34
Wang ZL, Li LH, He ZH, Duan XY, Zhou YL, Chen XM, Lillemo M, Singh RP, Wan H, Xia XC (2005) Seedling and adult plant resistance to powdery mildew in Chinese bread wheat cultivars and lines. Plant Dis 89:457–463
Xiang QJ, Sheng BQ, Zhou YL, Duan XY, Zhang KC (1994) Analyses of resistance genes of three differential varieties to the isolates of Blumeria graminis f. sp. tritici in wheat. Acta Agric Boreali-Sin 9:94–97
Xie CJ, Sun QX, Ni ZF, Yang ZM, Nevo E, Fahima T (2003) Chromosomal location of a Triticum dicoccoides-derived powdery mildew resistance gene in common wheat by using microsatellite markers. Theor Appl Genet 106:341
Yahiaoui N, Srichumpa P, Dudler R, Keller B (2004) Genome analysis at different ploidy levels allows cloning of the powdery mildew resistance Pm3b from hexaploid wheat. Plant J 34:528–538
Zhang XY, Koul A, Petroski R, Ouellet T, Fedak G, Dong YS, Wang RR-C (1996) Molecular verification and characterization of BYDV-resistant germplasms derived from hybrids of wheat with Thinopyrum ponticum and Th. intermedium. Theor Appl Genet 93:1033–1039
Zhou R, Zhu Z, Kong X, Huo N, Tian Q, Li C, Jin P, Dong Y, Jia J (2005) Development of wheat near-isogenic lines for powdery mildew resistance. Theor Appl Genet 110:640–648
Zhu ZD, Zhou RH, Kong XY, Dong YC, Jia JZ (2005) Microsatellite markers linked to two genes conferring resistance to powdery mildew in common wheat introgressed from Triticum carthlicum accession PS5. Genome 48:585–590
Acknowledgments
The authors are grateful to Drs. Bernd Friebe, Chengdao Li and Robert McIntosh for critical reviews of this manuscript, and to Dr. Shubing Liu for technical guidance in the SSR analyses. This project was funded by National Natural Science Foundation (30671299 and 39870398) and Shanxi Key Technologies R and D Program of China.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by E. Guiderdoni.
Runli He and Zhijian Chang contributed equally to this work.
Rights and permissions
About this article
Cite this article
He, R., Chang, Z., Yang, Z. et al. Inheritance and mapping of powdery mildew resistance gene Pm43 introgressed from Thinopyrum intermedium into wheat. Theor Appl Genet 118, 1173–1180 (2009). https://doi.org/10.1007/s00122-009-0971-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-009-0971-z