Abstract
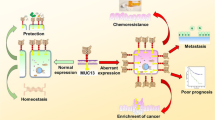
Mucins are high-molecular-weight glycoproteins dysregulated in aggressive cancers. The role of mucins in disease progression, tumor proliferation, and chemotherapy resistance has been studied extensively. This article provides a comprehensive review of mucin’s function as a physical barrier and the implication of mucin overexpression in impeded drug delivery to solid tumors. Mucins regulate the epithelial to mesenchymal transition (EMT) of cancer cells via several canonical and non-canonical oncogenic signaling pathways. Furthermore, mucins play an extensive role in enriching and maintaining the cancer stem cell (CSC) population, thereby sustaining the self-renewing and chemoresistant cellular pool in the bulk tumor. It has recently been demonstrated that mucins regulate the metabolic reprogramming during oncogenesis and cancer progression, which account for tumor cell survival, proliferation, and drug-resistance. This review article focuses on delineating mucin’s role in oncogenic signaling and aberrant regulation of gene expressions, culminating in CSC maintenance, metabolic rewiring, and development of chemoresistance, tumor progression, and metastasis.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Background
Body cavities like the gastrointestinal tract and respiratory tract are lined by mucosal surfaces that protect them against pathogens and prevent dehydration. The mucosa is lined by epithelial cells, which express a class of highly glycosylated proteins called mucins. These gel-forming mucins are produced by goblet cells and constitute a viscous mucus layer that covers mucosal surfaces and protects the mucosa from bacterial penetration [1]. Mucins are also expressed by epithelial cells of the pancreas, gall bladder, liver, eyes, kidney, salivary glands, and lacrimal glands [2]. The mucin family of glycoproteins is responsible for mucoadhesiveness, hydrophobicity, and viscoelasticity of mucus, which allows the mucus to protect the epithelium from chemical, enzymatic, and mechanical damage. Mucins are comprised of amino (N) and carboxy-terminal protein regions with tandem repeats of identical sequences rich in proline, threonine, and serine residues with O-linked or N-linked oligosaccharides [3, 4]. The mucoadhesive quality of mucins aids them in adhering to other substances via hydrogen bonds, hydrophobic bonds, and electrostatic interactions, which lead to the creation of gel aggregates [5, 6].
The mucin family can be broadly classified into two categories, based on their structural characteristics and domain organization: transmembrane mucins and secreted mucins. Transmembrane mucins consist of several members that vary in molecular size and composition of cytosolic signal-transducing and membrane-tethered domains. The following transmembrane mucins have been identified in humans: MUC1, MUC3A, MUC3B, MUC4, MUC12, MUC13, MUC14, MUC15, MUC16, MUC17, MUC20, MUC21, and MUC22. Secreted mucins are categorized into gel-forming mucins (MUC2, MUC5AC, MUC5B, MUC6, and MUC19) and non-gel-forming mucins (MUC7, MUC8, and MUC9). While the non-gel-forming mucins are noticed as monomers and cannot oligomerize [5], gel-forming mucins, characteristically secreted by goblet cells, form a viscoelastic mucus layer over most mucosal surfaces and hence contribute to the lubricating property of mucus [5, 7].
Tumor cells harness the physiological properties of these highly glycosylated huge glycoproteins to create rheologic barriers and dynamic interactome of the cancer epithelia during oncogenesis, cancer progression, and metastasis [7]. Over the past two decades, it has been increasingly recognized that mucins are associated with the pathogenesis of multiple cancers [3, 8,9,10]. Specifically, deregulated expression of mucins not only provides an essential bridge between inflammation and cancer [11] but have also been observed to contribute towards the carcinogenesis and metastatic cascade of carcinoma cells of epithelial origin, as observed in the case of breast, prostate, lung, ovarian, and pancreas [1, 11]. Under the extended tandem-repeats of transmembrane mucins and polymeric gel of secreted mucins, epithelial cells can transduce extracellular signals from the environment. This renders mucins capable of functioning as cell-surface receptors and sensors to conduct oncogenic signals and coordinate cellular responses against external biochemical stimuli, thereby facilitating enhanced tumorigenicity, invasiveness, metastasis, immune modulation, metabolic shift, and drug resistance [3].
In promoting discussion of the crucial involvement of mucins and the associated aberrant glycosylation in regulating epithelial to mesenchymal transition, CSC maintenance, chemoresistance, and metabolic reprogramming in cancer cells, we have further emphasized the impact of mucin inhibition in blocking tumor progression and metastasis.
2 Mucins promote extrinsic and intrinsic resistance of cancer cells to therapeutics
2.1 Mucins form a physical barrier on epithelial surfaces
It was previously believed that the primary function of mucins was protection and lubrication of epithelial surfaces. Lately, several studies have associated it with other functions like growth, fetal development, epithelial renewal, differentiation, oncogenesis, immune system evasion, metastasis, and chemotherapeutic resistance [12, 13]. The treatment of mucin-expressing cancer cells with drugs is difficult because of the resistance mechanism [12]. One of the major reasons for chemoresistance and poor prognosis is the mucus barrier [14].
Membrane-bound mucins (MUC1, MUC4, and MUC16) and secreted mucins (MUC2, MUC5AC, and MUC5B) form an outstanding web, depending on the mucus network porosity. Diseases like chronic inflammatory cystic fibrosis, chronic obstructive pulmonary disease, chronic respiratory disease, and cancer usually have dense mucus [15, 16]. Based on the porosity of the mucus network, they function as size filters and hydrophobic or electrostatic forces. Mucins limit the effectiveness of cytotoxic drugs by restricting the accessibility of drugs to plasma-membrane and by hampering the intracellular uptake [17,18,19,20]. These reduce the drug efficacy of antibody-based therapies and chemotherapies, leading to drug-resistance [18, 21]. The high density of mucin present at the membrane surface creates a structural mesh, which restricts the interaction of tumor cell epitopes to immune cells [22, 23]. It also limits the cytotoxic response generated by antibody-based therapy and the immune cell-mediated killing of tumor cells [21, 24, 25]. Though several studies have shown the chemoresistance activity of mucins via the extrinsic route of forming physical barriers, mucins are also implicated in modulating a plethora of intrinsic resistance mechanisms, including cell survival, resistance to apoptosis, drug metabolism, enrichment of stemness, and EMT.
2.2 Mucins reprogram cell-survival and resistance to apoptosis
The primary objective of cytotoxic therapy is to induce cell apoptosis. A major reason for chemoresistance is the ability of cancer cells to surpass programmed cell death, wherein mucin expression reduces their susceptibility to genotoxic drugs by decreasing the DNA damage or endowing survival mechanisms to overpower the physiological stress [21]. Most cancer cells harbor dysregulated apoptotic pathways, because of which anticancer treatments are rendered ineffective. It has been demonstrated that cancer cells avoid cell death with mucins that block intrinsic apoptotic pathway activation [26]. In human pancreatic cancer, an elevated amount of MUC1 correlates to enhanced resistance to chemotherapeutic drugs (gemcitabine and etoposide) compared to the low MUC1-expressing cells. In pancreatic cancer cells, MUC1 upregulates MRP1 through the PI3K/Akt pathway [27]. MUC1-CT stimulates the PI3K/Akt pathway, which in turn increases MUC1 expression. The MUC1 CT translocates to the nucleus and binds to the promoter of the Abcc1 gene, thus operates as a part of the transcriptional complex to stimulate the expression of multidrug resistance (MDR) genes, including ABCC1, ABCC3, ABCC5, and ABCB1 [27].
In various cancer types like breast, it has been reported that the expression of MUC1 plays a role in therapy resistance [28]. In cancer cells that achieve chemoresistance after prolonged Paclitaxel (PTX) treatment, the expression of MUC1 is significantly upregulated. Chemotherapeutic drugs activate the expression of MUC1 in cancer cells, which leads to the activation and nuclear distribution of EGFR. EGFR and MUC1 work together to upregulate ABCB1 transcriptionally and results in the attainment of chemoresistance [29]. MUC1-CT plays a vital role in the induction of stemness and PTX resistance in human NSCLC A549 cells. Along with MUC1-CT, oncoproteins such as PI3K/Akt and β-catenin were also found to be considerably high in A549/PTX cells [30], suggesting that MUC1-CT, PI3K/AKT, and β-catenin may work through the same pathway to generate drug resistance in lung cancer cells. Further, stemness-related factors such as Nanog, Oct4, Sox2, CXCR4, and ALDH1 were also shown to be overexpressed in PTX-resistant A549 cells [30] (Fig. 1).
Schematic diagram showing the mucins role in cancer stemness and chemoresistance. MUC1-C promotes the activation of PI3K/Akt phosphorylation and the regulation of stemness genes such as Nanog, Oct3/4, Sox2, CXCR, and ALDH1, which are responsible for chemoresistance in lung cancer cells. This leads to MUC1-C mediated PTX resistance. The nuclear translocation of MUC1 stimulates chemotherapy resistance via the upregulation of ABCB1 in an EGFR-dependent manner. MUC1 provides the resistance to drugs (PTX) by directly regulating ABCC1, ABCB1 gene expression, dependent on the PI3K/Akt pathway. Moreover, the stimulation of PI3K/Akt pathway by MUC1-Cter, in turn, increases the expression of MUC1. The MUC4/NF-κB pathway downregulates the expression of the hCNT1 transporter and regulates the apoptotic proteins Bax and Bcl-xL; thus, it aids the pancreatic cancer cells in resisting gemcitabine. Similarly, MUC4 also activates NF-κB through the ERK1/2 and JNK pathways. MUC4 alters the stability of Her2 protein, and auranofin downregulates the Her2 protein. When the expression of MUC4 is attenuated, it aids auranofin in promoting FOXO3 translocation from the cytoplasm into the nucleus to regulate the pro-apoptotic and anti-apoptotic proteins. The upregulation of the secretory mucin MUC5AC provides resistance to 5-FU via up-regulation of β-catenin, CD44, and Lgr5 and downregulation of p53 and p21 in colorectal cancer. MUC13/NFkB signaling pathway promotes drug resistance and prevents apoptosis by stimulating the anti-apoptotic proteins, Bcl-xL, survivin in sorafenib, or sunitinib-treated renal cancer cells. MUC16/JAK2/STAT3/TSPYL5 signaling axis downregulates p53, which leads to chemoresistance (cisplatin and gemcitabine) in lung cancer cells. Ectopic expression of MUC16-Cter confers CSC phenotype via the Activation of JAK2, which phosphorylates histone-3 and upregulates stemness-specific genes like LMO2, NANOG, and ALDH activity in pancreatic cancer cells. Abbreviations of mucin-mediated signaling pathways, their downstream target genes associated with chemoresistance, and anti-apoptosis upregulated genes are indicated in red-colored arrow and downregulated genes using the green colored arrow. ALDH, aldehyde dehydrogenase; ABCB1, ATP-binding cassette subfamily B member 1; ABCC1, ATP-binding cassette subfamily C member 1; EGFR, epidermal growth factor receptor; FOXO3, forkhead box O-3; hCNT1, human concentrative nucleoside transporter; Lgr5, leucine-rich repeat-containing G protein-coupled receptor 5; LIMO2, LIM domain only 2; LRP6, low-density lipoprotein receptor protein 6; PTX, paclitaxel; 5FU, 5-Fluorouracil
MUC4 negatively regulates hCNT1 transporter expression through the NF-κB pathway, helping pancreatic cancer cells gain resistance against gemcitabine [31]. It is already known that the NF-κB pathway is highly activated in PC when compared to the normal pancreas [32]. The NF-κB pathway is associated with chemoresistance to gemcitabine, regulation of apoptosis, proliferation, and angiogenesis [33]. MUC4 protects pancreatic cancer cells from gemcitabine-induced apoptosis through HER2/ERK-dependent phosphorylation and inactivation of the pro-apoptotic protein Bad [11]. Skrypek et al. have illustrated the involvement of MUC4-dependent drug-resistance development, where MUC4 either reduces the hCNT1 nucleoside transporter expression through the NF-κB pathway or modulates the Bax/Bcl ratio by upregulating Bcl and downregulating Bax expression [34]. The activation of ErbB2 (HER2) by MUC4 is achieved by stimulating Her2 dimerization with other ErbB receptors [35, 36]. The MUC4/HER2 complex in several tumors and cancer cell lines has been identified in various studies [37, 38]. These studies have revealed that silencing of MUC4 expression leads to the proteasomal degradation of HER2, reducing HER2 stability and expression. Next, it has been demonstrated that knockdown of MUC4 with concurrent auranofin treatment in SKOV3 cells increased the proteasomal degradation of HER2. Auranofin is an FDA-approved drug for the treatment of rheumatoid arthritis, and it induces apoptosis in SKOV3 cells through the regulation of the IKKβ/FOXO3 pathway [39]. The coordinated anticancer activity of the combination is achieved by the downregulation of Her2 expression and phosphorylation of Akt, thus translocating FOXO3 from the cytosol to nucleus and initiating the transcriptional activity of FOXO3, which induces caspase-3-mediated apoptosis and expression of Bcl-2 interacting mediator of cell death (BimEL)[40].
Ramesh et al. showed that MUC5AC and CD44 levels are elevated when colorectal cancer cells were treated with 5-FU. This upregulation of MUC5AC provided resistance to 5-FU by the downregulation of p53 and its target gene p21 and upregulation of β-catenin and its target genes CD44 and Lgr5. The findings of this study prove that MUC5AC promotes 5-FU resistance via the β-catenin/p53/p21 axis in colorectal cancer [41]. An increased expression of MUC13 is strongly associated with increased tumor grade and poor prognosis in renal cell carcinoma (RCC) [42, 43]. Moreover, prolonged exposure to sunitinib or sorafenib facilitates cell growth in RCC and drug resistance development, which is related to higher expression of MUC13. MUC13 promotes the activation of the NF-kB signaling, which plays a vital role in the behavior of cancer cells in various types of adenocarcinoma and RCC. The activation of several target genes like cyclin D1, Bcl-xL, and survivin by NF-kB leads to epithelial cancer cell growth and survival, thus facilitating proliferation and blocking apoptosis [42]. The silencing of MUC13 solely or when combined with sorafenib or sunitinib reduces the expression of phosphorylated NF-kB p65 and Bcl-xL. Additionally, ABCB1 protein, an efflux pump responsible for multidrug resistance, was upregulated by chronic sorafenib and sunitinib treatments, dependent on MUC13 expression. The findings of these studies indicate that MUC13 is a crucial mediator of drug resistance through the NFkB signaling pathway [42].
Overexpression of MUC16 was observed in human lung adenocarcinoma and lung cancer tissues of genetically engineered mice [44]. Furthermore, the knockdown of MUC16 in human and mouse tumor cells led to increased sensitivity to cisplatin and gemcitabine, wherein cells that overexpressed MUC16-Cter showed higher resistance to chemotherapy. This study further illustrated that MUC16 upregulates TSPYL5 via JAK2/STAT3/GR axis and suppresses p53 activity in lung cancer [44]. MUC16-Cter overexpressed pancreatic cancer cells increased resistance to apoptosis and proliferative potential [45], and another study has shown that ectopically expressed MUC16-Cter in SKOV3 cells increases cisplatin resistance in ovarian cancer cells [46] (Fig. 1).
3 Mucins in stem cells, glycosylation, and EMT-programing
Stem cells are undifferentiated cells that can transform into differentiated and specialized cell types. Pluripotency and self-renewal are the most significant features of stem cells [47]. The expression of specific markers, which may be cell surface or intracellular proteins, transcription factors, enzymes, etc., are used to identify and sort stem cells [48]. Besides, glycosylation has been found to play a significant role in embryonic development. Yan et al. demonstrated that the O-fucosylation of Notch receptors controls blood lineage commitment [49]. Another study has shown that core O-fucosylation of apolipoprotein B is necessary for proper midline patterning in zebrafish development by modulating the sonic hedgehog signaling [50]. These studies demonstrate the importance of glycosylation in mediating stem cell maintenance and fate determination during the embryonic development process.
3.1 Mucins bolster cancer stem cell maintenance
Cancer stem cells (CSC) are a small subset of the stem cell population within tumors and are characterized by an extensive self-renewing capacity. Many studies have indicated that CSCs represent the chemoresistant pool of cells in the bulk tumor, and they mostly account for tumor relapse post-therapy [51]. Chemotherapy and various other anticancer therapies provide a strong selection for CSC survival and proliferation [52].
MUC1-CT has been shown to upregulate breast cancer stem cell marker aldehyde dehydrogenase 1A1 (ALDH1A1) through ERK1 and C/EBPß by engaging in a transcriptional activating complex on the ALDH1A1 gene promoter [53]. MUC1 is expressed by a majority of side population (SP) cells, which are maintained in MUC1+ tumors in vivo. MUC1, in SP cells, is hypoglycosylated and strongly sialylated [54].
MUC1 expression has also been identified in CD34+/CD38- acute myeloid leukemia (AML) cells, associated with leukemia stem cells (LSC), suggesting that MUC1 is a potential marker in the AML stem cell population [55]. MUC1 is one of the cell surface antigens in colorectal cancer stem cells (CCSC). In the CT26 mouse model, mice that were vaccinated with MUC1 knockin CCSC displayed anti-colorectal cancer immunity, demonstrated by enhanced innate and adaptive immune responses and immune memory. The anti-tumor effect of the CCSC vaccine decreased partially when the antibody against MUC1 was neutralized; this potentiates MUC1 as the major antigen for the CCSC vaccine. CCSC vaccine with high expression of MUC1 is a novel prophylactic vaccine for CRC [56, 57]. A study identified the expression and role of MUC1 in pancreatic cancer stem cells. MUC1 levels were present in two populations of CSC, namely CD44 + CD24 + EpCAM+ (Triple+ cells) and CD133+ cells, detecting greater than 95% of CSC population. In this study, an anti MUC1 antibody was used to detect the shed MUC1 fragment in the serum of pancreatic cancer patients and speculate the disease stage progression [58].
MUC4 overexpression causes an increase in CD133+ positive cancer stem cells in ovarian cancer [59]. In a small subpopulation of pancreatic epithelial cells, stem cell-like marker CD133 has been observed along with MUC4 expression in the basal compartment of non-malignant pancreatic tissue specimens [60]. Mimeault et al. have proposed that MUC4 downregulation may partly reverse the CD133 + -associated resistance property of the cancer-initiating cells after gemcitabine treatment [60]. This is particularly significant since gemcitabine treatment of pancreatic tumor xenografts promotes the selective enrichment of subpopulations of cells expressing stem cell markers such as ALDH, CD24, and efflux pumps such as ABCB1, ABCG2 [61, 62]. Thus, targeting MUC4 oncoprotein may present a promising therapeutic strategy to reduce the cancer stem cell population in a tumor, thereby preventing disease relapse [60].
A significant decline in side population cells was detected upon MUC5AC knockout in CRC cells. MUC5AC and CD44 were overexpressed in the isolated stem cell population [41]. High throughput RNA-sequencing analysis of pancreatic tissues from an autochthonous murine model of pancreatic cancer demonstrated that Muc5ac-knock out mice had a substantial reduction in cancer stem cell markers Aldh1a1, Klf4, EpCAM, and CD133. Biochemical experiments in this study delineated that MUC5AC/integrin avß5 interaction results in pSrc (Y416)/ pSTAT3 (Y705) signaling axis, which further upregulates Klf4 expression, resulting in the enrichment of the self-renewing CSC population in pancreatic cancer [63] (Fig. 1).
Zhang et al. demonstrated that while CA125-positive cells could form tumors upon orthotopic implantation into the ovary, a similar number of CA125-negative cells failed to develop any cancer. It was also observed that the primary cause of tumor recurrence in ovarian cancer is CA125-positive cells, which were ovarian cancer stem cells [64]. MUC16-Cter enriched ALDH+ cancer stem cells in pancreatic cancer cells, thereby contributing to gemcitabine and cisplatin resistance. MUC16-Cter initiates the nuclear translocation of JAK2 and upregulates stemness-specific genes such as LMO2 and NANOG [45] (Fig. 1).
3.2 Mucins induce the epithelial-mesenchymal transition (EMT) for metastasis of cancer
Epithelial-mesenchymal transition (EMT) is a reversible pathophysiological process associated with loss of cell polarity, decreased surface expression of epithelial markers, and increased expression of mesenchymal markers. EMT inducers include transcription factors like Snail, Slug, and Twist1/2. EMT is a crucial step contributing towards metastatic tumor progression; promotion of drug resistance, selective maintenance of cancer stem cells, and ultimately disease recurrence [65, 66]. Several studies have demonstrated the molecular events of EMT, triggered by MUC1, MUC4, MUC5AC, and MUC16 [67,68,69,70] [71].
MUC1 promotes EMT by increasing the expression of Vimentin and Slug and downregulating E-cadherin expression to stimulate the invasion and metastasis [69]. Studies have shown that in triple-negative breast cancer cells, the oncogenic MUC1 cytoplasmic tail protein is expressed aberrantly. Furthermore, MUC1-CT stimulates the phosphorylation of STAT3 and upregulates the TWIST1 gene. Moreover, MUC1-CT binds directly to TWIST1, and this MUC1-CT/TWIST1 complex is adequate for stimulating the ZEB1 gene expression in TNBC [72]. Besides occupying the ZEB1 promoter with NF-κB/p65, MUC1-CT promotes ZEB1 transcription in breast cancer [73]. Tumor progression and metastasis are promoted by MUC1 signaling via its cytoplasmic tail (MUC1-CT) and interaction with other oncogenic signaling molecules. The induction of EMT independent of SMAD4 is regulated through the interaction of MUC1-CT with the TGF-β signaling pathway. The overexpression of MUC1 in pancreatic cancer cells induces EMT transition, leading to increased invasiveness in response to exogenous TGF-β [74].
In ovarian cancer, MUC4 stimulates the activation of FAK directly or by interacting with HER2, which leads to the activation of MKK7, JNK1/2, and c-Jun signaling, resulting in the upregulation of N-cadherin. The over-expression of MUC4 results in decreased expression of epithelial markers and induction of mesenchymal markers, such as Twist1, Twist2, and Snail [70]. A study has shown that MUC4 mediates N-cadherin upregulation in part through the Akt pathway and partly by the JNK1/2 pathway in pancreatic cancer cells. Silencing of MUC4 has been shown to downregulate pAkt, Twist, Slug, and Zeb1 in PC cells, namely Capan1 and BXPC3 [75].
The overexpression of MUC5AC in lung cancer patient tissues was associated with poor survival [71]. The interaction of MUC5AC with integrin β4 resulted in activation of FAK at Y397; this contributed to lung cancer cell migration. In this study, Lakshmanan et al. showed that MUC5AC knockdown resulted in considerably less migratory capacity. Moreover, in MUC5AC knockdown cells, mesenchymal cell markers N-cadherin and Vimentin were decreased, whereas epithelial marker Cytokeratin 18 and E-cadherin were increased. [71].
The interaction of MUC16 with FAK resulted in the activation of Akt and ERK/MAPK signaling, which further facilitated tumorigenesis and metastasis of PC cells. The loss of MUC16 reduced FAK activation, downregulated mesenchymal cell markers such as N-cadherin, and ZEB-1 and upregulated epithelial-specific markers such as E-cadherin and Cytokeratin 18. Therefore, this study demonstrated that MUC16-FAK interaction promotes the EMT process via Akt and ERK/MAPK signaling axis in PC [76] (Fig. 2).
Mucin’s role in epithelial to mesenchymal transition (EMT). Mucins regulate the EMT, a reversible pathophysiological process like loss of cell polarity, decreased surface expression of epithelial markers, and increased expression of mesenchymal markers. MUC1 phosphorylates STAT3, which regulates TWIST1, also MUC1 directly regulates the expression of Slug, Vimentin, and E-cadherin proteins in TNBC. Moreover, MUC1 promotes the transcription of ZEB1 via the activation/nuclear translocation of NF-κB p65. MUC1 promotes the induction of EMT independent of SMAD4, and it is regulated by the interaction of MUC1-CT with the TGF-β signaling pathway in pancreatic cancer. MUC4 activates FAK directly or by interacting with HER2. This leads to the activation of MKK7, JNK1/2, and Akt pathways and decreased expression of epithelial markers and induction of mesenchymal markers. MUC5AC/integrin-FAK dependent signaling pathway upregulates Vimentin and N-Cadherin and downregulates the epithelial markers Cytokeratin 18 and E-cadherin in lung cancer. MUC16-FAK interaction induces the EMT process through the Akt and ERK/MAPK signaling axis in PDAC. EMT markers upregulated genes are indicated in red-colored arrow and downregulated genes using the green colored arrow. MUC1, MUC4, MUC5AC, and MUC16 mediated signaling pathways and their downstream target genes associated with EMT markers. FAK, focal adhesion kinase; HER2, human epidermal growth factor receptor 2; JNK1/2, c-Jun N-terminal kinases 1/2; MKK7, mitogen-activated protein kinase kinases 7; NFκB, nuclear factor κB; STAT3, signal transducer and activator of transcription 3; TGFβ, transforming growth factor-beta; TGFβR I/II, transforming growth factor-beta receptor I/II; TNBC, triple-negative breast cancer; ZEB1, zinc finger E-box-binding homeobox 1
4 Mucins rewire metabolic pathways of cancer cells
Altered metabolism is one of the pivotal hallmarks of oncogenesis and tumor progression. The cancer cells exhibit aberrant expression of various genes, including glucose, amino acid, or nucleotide transporters, and harness a plethora of dysregulated signaling pathways for enhanced utilization of the limited nutrient reserve [77, 78]. This is particularly true for solid tumors where the tumor cores are highly hypoxic and less transfused with blood vessels. The tumor cells are often involved in the metabolic crosstalk with other tumor microenvironment cells and often exploit the host system acting as “metabolic parasites” [79].
4.1 Mucins in altered carbohydrate metabolism
One of the critical traits that enable cancer cell survival and proliferation is altered glucose metabolism. The cytoplasmic tail of MUC1, which is overexpressed in various carcinomas, including PC, triggers multiple signaling pathways through interaction with several transcription factors/co-regulators at the promoter elements of various genes. The study by Nina et al. demonstrated that MUC1 functions as a modulator of the hypoxic response in PC cells by regulating the expression, stability, and activity of hypoxia-inducible factor-1a (HIF-1α). In PC, MUC1 physically interacts with HIF-1α and p300 and controls several metabolite intermediates such as HK2, PFKFB2, ENO1, PGK1, PGM2, and LDHA [80].
Several studies show that the overexpression of the MUC1 oncoprotein in 3Y1 fibroblasts results in glycolysis regulation and the activation of the PI3K/Akt pathway [81, 82]. MUC1 controls the glycolytic flux by triggering various signaling pathways like PI3K/Akt/mTOR, p53, and HIF-1α, which regulate the transcription of glycolytic genes [83]. Furthermore, MUC1 interacts with Receptor Tyrosine Kinases (RTKs) to enhance their stability on the membranes, which further aids the downstream signaling of RTKs to promote PKM2 activity and the transcriptional activation of glycolytic genes [83]. Chaika et al. showed that MUC1 enhances glucose uptake and glycolysis via HIF-1α modulation, and the cytoplasmic tail of MUC1 is recruited at the promoters of several glycolytic genes in PC. The expression of MUC1 also enhances the expression of specific glycolytic genes like PFKFB2 in a hypoxia/HIF-1α-independent manner. Even in normoxic conditions, MUC1 occupies ENO1 and PGM2 glycolytic gene promoters and triggers their expression [83].
Breast cancer type 1 susceptibility gene (BRCA1) is a breast cancer suppressor gene, which has a vital role in DNA double-strand break repair [84]. Also, BRCA1 restricts glycolysis by downregulating glycolytic enzymes in breast and ovarian cancer [85, 86]. A study has shown that the mRNA and protein levels of BRCA1 increase considerably in MUC1 KO cells, which implies that MUC1 inhibits the transcription of the BRCA1 gene. Also, increased BRCA1 level results in decreased glycolysis in MUC1 KO cells. On the other hand, MUC1 KO CFPAC-1 cells displayed enhanced glucose uptake and lactate release post BRCA1 inhibitor treatment [87]. In comparison to WT cells, MUC1 KO cells showed reduced expression of metabolic intermediates such as SLC2A1, HK2, PFKFB2, PGI, LDHA, ENO1, PGK1, and PKM. This study demonstrates that MUC1 promotes glycolysis by preventing BRCA1 expression [87].
The cytoplasmic domain of MUC1-C harbors a CQC motif, essential for its homodimerization and function [88, 89]. Therefore, to prevent the activation of various MUC1-C-mediated pathways, cell-penetrating peptides GO-203 have been developed to target the MUC1-C CQC motif [90]. In immunohistochemistry results, it shows that targeting MUC1-C with GO-203 resulted in the downregulation of TP53-induced glycolysis and apoptosis regulator (TIGAR) expression in xenograft tissue; this confirms the relationship between MUC1-C and TIGAR in human ESCC [91].
In pancreatic tumors, overexpression of MUC13 leads to tumor progression through the regulation of HER2 receptor tyrosine kinase activity [92]. Sonam et al. showed that the expression of MUC13 resulted in the activation/nuclear translocation of NF-κB/p65 and phosphorylation of IκB, which increases the expression of genes associated with glucose metabolisms like Glut-1, HIF-1α, and c-Myc. MUC13 interacts with and stabilizes Glut-1, which is accountable for the increased glucose uptake in PC cells. The glucose uptake and lactate secretion in PC cells were regulated by altering the expression of MUC13 and the associated molecular mechanisms [92].
The primary tumor and metastatic lesions exhibit high expression of MUC16 in PC patients [93]. A recent study shows that the PC cells in which MUC16 has been knocked down display decreased glucose uptake and lactate efflux, with a concurrent decrease in migration and invasion potential [94]. MUC16 knockdown also reduced GLUT1 and HK2, reduced phosphorylation of Akt and mTORC1 target proteins p70S6K and 4EBP1, and lowered the expression of its downstream target c-Myc, which in turn is a key player in cellular growth, proliferation, and metabolism [94]. Interestingly, various aspects of cancer cells, including survival, proliferation, cell cycle, motility, and cellular metabolism regulation, are regulated by the PI3K-Akt-mTOR pathway [95, 96]. These studies hence illustrate that MUC16 bolsters the glycolytic propensity of PC cells (Fig. 3).
Schematic diagram showing the role of mucins in cancer metabolism. Mucins regulate metabolism through the activation of multiple signaling pathways. MUC1 physically interacts with HIF-1α and p300 and controls several metabolite intermediates in pancreatic cancer cells. BRCA1 prevents glycolysis by downregulating glycolytic enzymes in cancer. MUC1 promotes glycolysis by inhibiting the expression of BRCA1 in pancreatic cancer. MUC1 regulates PI3K/AKT/mTOR, p53, TIGAR, and HIF-1α, which regulate the transcription of glycolytic genes. MUC1 interacts with RTKs and increases the stability of the RTK membrane. As a result, the MUC1/RTK complex promotes glycolytic genes. MUC13 activates the phosphorylation of IκB and activation/nuclear translocation of NF-κB/p65. This, in turn, leads to increased expression of genes linked with glucose metabolism. MUC16 regulates glycolysis by activating the PI3K-AKT-mTOR/c-Myc pathway in pancreatic cancer. Metabolic pathways upregulated by mucins are written in red color. Abbreviations of metabolic genes regulated by mucins: Acetyl-CoA, acetyl-coenzyme A; ENO1, enolase 1; G-1-P, glucose -1- phosphate; GLUT1, glucose transporter1; GLU, glutamate; GSL1, glutaminase 1; GLN, glutamine; GF, growth factor; HK2, hexokinase 2; HIF-1α, hypoxia-inducible factor 1, alpha; LDHA, lactate dehydrogenase A; mTORC1, mammalian target of rapamycin complex 1; PFK PFKFB2, phosphofructokinases; PGI, phosphoglucoisomerase; PGM, phosphoglucomutase; PGK1, phosphoglycerate kinase 1; PDH, pyruvate dehydrogenase; PDK1, pyruvate dehydrogenase kinase 1; PKM2, pyruvate kinase isozyme M2; RTK, receptor tyrosine kinase; S6K1, ribosomal protein S6 kinase 1; SLC1A5, solute carrier family 1 member 5; TIGAR, TP53-induced glycolysis and apoptosis regulator; TCA cycle, tricarboxylic acid cycle; α-KG, α-Ketoglutarate
4.2 Mucins in altered lipid metabolism
Energy metabolism and lipid metabolism are quite interrelated. Therefore, changes in one of these will alter the course of other pathways. Altered lipid metabolism is implicated in the pathogenesis of cancer [97]. Many studies have shown that most tumors increase de novo fatty acid synthesis, regardless of available lipid resources [98]. This helps tumors expand their cells rapidly by providing them with more building blocks, crucial signaling molecules, and valuable energy sources in response to limited glucose availability. Consequently, besides synthesis, the beta-oxidation pathway is modified to manage the changes in cellular energetics of cancer cells [99].
Most tumors overexpress enzymes and factors involved in lipid biosynthesis, such as acetyl-CoA carboxylase (ACC), ATP citrate lyase (ACL), fatty acid synthase (FAS), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), and sterol regulatory element-binding protein 1 (SREBP1) [100,101,102]. As expected, these enzymes are downstream to many tumor suppresser genes and oncogenes, including PI3K, MAPK, p53, c-Myc, and EGFR [103], which, as mentioned above, are modulated by various mucins.
A study by Pitroda et al. reported that the genes associated with MUC1-induced tumorigenesis are also involved in lipid metabolism regulation. This study established that MUC1 induces a set of lipid metabolic genes, named MUC1-induced lipid metabolism signature (MLMS), in breast cancer cells [28]. MLMS consists of a set of 38 genes, which come under lipid-metabolizing enzymes and transporters. Notably, the list contains the most commonly upregulated genes in various malignancies and transformed phenotypes, such as ACL, FASN, and SERBF1 (sterol regulatory element-binding transcription factor 1). ACL helps synthesize acetyl-CoA, a precursor for fatty acid and cholesterol synthesis. SREBP1 controls the genes involved in fatty acid, cholesterol, and triglyceride synthesis. This process also includes FAS, which catalyzes the synthesis of palmitate from acetyl-CoA and malonyl-CoA in the presence of NADPH. MLMS, which represents the alterations in MUC1-induced lipid metabolism, predicts cancer recurrence and metastasis in tamoxifen-treated breast cancer patients [28]. Moreover, MUC1-mediated HIF-1α stabilization/activation can promote transcription of FASN and SREBP, while excessive lactate secretion-induced acidosis can promote FAS induction through epigenetic mechanisms, as reported in breast cancer cells [104].
5 Conclusions and future perspectives on therapeutic strategies
The membrane-bound and secretory mucins MUC1, MUC4, MUC16, and MUC5AC are promising diagnostic and therapeutic targets because of their differential expression and functional involvement in pathogenesis. Currently, methods to exploit the overexpression and glycosylation of MUC1 in various carcinomas and the probability of MUC1-based immunotherapy is being explored [105]. In the development of MUC1-based immunotherapy, a major point to be considered is MUC1-N (highly glycosylated), and MUC1-C components role in immune evasion by cancer cells. Recently, Govind et al. generated mAb 3D1 against the non-shed oncogenic MUC1 C-terminal subunit. mAb 3D1 binds to the surface of multiple non-small cell lung cancer and breast cancer cell lines. Human Ab-3D1 monomethyl auristatin E (MMAE) antibody-drug conjugate (ADC)-mediated killing of cancer cells could be effective in reversing the suppressive immune microenvironment of tumors, thus enhancing the activity of other immuno-therapeutic agents, including antibodies used for the blockade of the PD-1/PD-L1 axis [106]. The MUC4-ErbB2 interaction at the biophysical level proves that the presence of three EGF domains of MUC4 is adequate to provide effective interaction. Future work may help if directed towards identifying small-molecule inhibitors to decrease the binding affinities of the MUC4-ErbB2 complex for drug discovery development in cancer [107]. Emerging evidence confirms that MUC4 can be a potential immunotherapy target [10]. Therapeutic strategies can use MUC4-specific antitumor immune response using vaccines or chemotherapeutic agents with anti-MUC4 antibodies in combination. The cleavage of MUC16-Cter takes place in the juxta-membrane ectodomain stretch of twelve amino acids that create approximately 17 kDa cleaved product [108]. A functional cleaved MUC16 provides tumorigenic, metastatic, and drug-resistant properties to pancreatic cancer cells in part by enriching ALDH+ CSCs, which depends on nuclear JAK2 mediated upregulation of stemness specific genes LMO2 and NANOG. Therefore, therapeutic strategies targeted against MUC16-Cter will be crucial in treating MUC16-expressing pancreatic cancer patients. The study has shown that when a rapid and reversible block in the secretory pathway is induced by brefeldin A (BFA), it prevents the cleavage of MUC16 [45]. One of the major barriers to effectively target MUC16-expressing cancers is cleavage and shedding of the extracellular domain of MUC16. Hence coordinated targeting of the non-cleaved cell surface retained portion of MUC16 may be beneficial. This can be achieved by using antibody-based therapeutics against the carboxyl-terminal of MUC16. A recent study demonstrated a panel of monoclonal antibodies against the carboxy-terminal (CT) portion of MUC16 [109]. MUC5AC, a polymeric secreted mucin, has been shown to harbor a protumorigenic role in several malignancies, including pancreatic [63], colon[41], and lung [71] tumors. While the expression of MUC5AC significantly correlates with the poor prognosis of cancer patients, the biochemical isoforms of this mucin and their mechanistic involvement in disease pathologies are still less understood. Our group has delineated that MUC5AC promotes the enrichment of CSCs and drives chemoresistance. Recently, MUC5AC has been demonstrated to harbor therapeutic utility. The monoclonal antibody NEO-102 (NPC-1C, ensitumab) targets a cancer-associated variant of MUC5AC that is specific to CRC [110]. The mechanism of action of NEO-102 is via antibody-dependent cellular cytotoxicity (ADCC). Understanding the pathologies of cancer-specific molecular variants of MUC5AC will pave the way for better diagnosis, patient stratification, and designing a therapeutic regimen for cancer patients.
Data availability
Not applicable
Code availability
Not applicable
Abbreviations
- ALDH:
-
aldehyde dehydrogenase
- AML:
-
acute myeloid leukemia
- CSC:
-
cancer stem cells
- EGFR:
-
epidermal growth factor receptor
- EMT:
-
epithelial-mesenchymal transition
- FAK:
-
focal adhesion kinase
- FOXO3:
-
forkhead box O-3
- hCNT1:
-
human concentrative nucleoside transporter 1
- HER2:
-
human epidermal growth factor receptor 2
- HIF-1α:
-
hypoxia-inducible factor 1, alpha
- JNK1/2:
-
c-Jun N-terminal kinases
- Lgr5:
-
leucine-rich-repeat-containing G protein-coupled receptor 5
- LIMO2:
-
LIM domain only 2
- LRP6:
-
low density lipoprotein receptor protein 6
- LSC:
-
leukemia stem cells
- MDR:
-
multiple drug resistance
- MKK7:
-
mitogen-activated protein kinase kinases 7
- mTORC1:
-
mammalian target of rapamycin complex 1
- NFkB:
-
nuclear factor-kappa B
- PTX:
-
paclitaxel
- RCC:
-
renal cell carcinoma
- SP:
-
side population
- STAT3:
-
signal transducer and activator of transcription 3
- TGFβ:
-
transforming growth factor-beta
- TGFβR I/ II:
-
transforming growth factor-beta type I/II receptor
- TIGAR:
-
TP53-induced glycolysis and apoptosis regulator
- TNBC:
-
triple negative breast cancer
- 5FU:
-
5-Fluorouracil
References
van Putten, J. P. M., & Strijbis, K. (2017). Transmembrane mucins: Signaling receptors at the intersection of inflammation and cancer. Journal of Innate Immunity, 9(3), 281–299. https://doi.org/10.1159/000453594.
Forstner, J. F. (1978). Intestinal mucins in health and disease. Digestion, 17(3), 234–263. https://doi.org/10.1159/000198115.
Kaur, S., Kumar, S., Momi, N., Sasson, A. R., & Batra, S. K. (2013). Mucins in pancreatic cancer and its microenvironment. Nature Reviews. Gastroenterology & Hepatology, 10(10), 607–620. https://doi.org/10.1038/nrgastro.2013.120.
Chaturvedi, P., Singh, A. P., & Batra, S. K. (2008). Structure, evolution, and biology of the MUC4 mucin. The FASEB Journal, 22(4), 966–981. https://doi.org/10.1096/fj.07-9673rev.
Dhanisha, S. S., Guruvayoorappan, C., Drishya, S., & Abeesh, P. (2018). Mucins: Structural diversity, biosynthesis, its role in pathogenesis and as possible therapeutic targets. Critical Reviews in Oncology/Hematology, 122, 98–122. https://doi.org/10.1016/j.critrevonc.2017.12.006.
Krishn, S. R., Ganguly, K., Kaur, S., & Batra, S. K. (2018). Ramifications of secreted mucin MUC5AC in malignant journey: A holistic view. Carcinogenesis, 39(5), 633–651. https://doi.org/10.1093/carcin/bgy019.
Ganguly, K., Rauth, S., Marimuthu, S., Kumar, S., & Batra, S. K. (2020). Unraveling mucin domains in cancer and metastasis: When protectors become predators. Cancer Metastasis Reviews, 39(3), 647–659. https://doi.org/10.1007/s10555-020-09896-5.
Lakshmanan, I., Ponnusamy, M. P., Macha, M. A., Haridas, D., Majhi, P. D., Kaur, S., Jain, M., Batra, S. K., & Ganti, A. K. (2015). Mucins in lung cancer: Diagnostic, prognostic, and therapeutic implications. Journal of Thoracic Oncology, 10(1), 19–27. https://doi.org/10.1097/jto.0000000000000404.
Pothuraju, R., Krishn, S. R., Gautam, S. K., Pai, P., Ganguly, K., Chaudhary, S., Rachagani, S., Kaur, S., & Batra, S. K. (2020). Mechanistic and functional shades of mucins and associated glycans in colon cancer. Cancers (Basel), 12(3). https://doi.org/10.3390/cancers12030649.
Gautam, S. K., Kumar, S., Dam, V., Ghersi, D., Jain, M., & Batra, S. K. (2020). MUCIN-4 (MUC4) is a novel tumor antigen in pancreatic cancer immunotherapy. Seminars in Immunology, 47, 101391. https://doi.org/10.1016/j.smim.2020.101391.
Bafna, S., Kaur, S., & Batra, S. K. (2010). Membrane-bound mucins: the mechanistic basis for alterations in the growth and survival of cancer cells. Oncogene, 29(20), 2893–2904. https://doi.org/10.1038/onc.2010.87.
Rao, C. V., Janakiram, N. B., & Mohammed, A. (2017). Molecular pathways: Mucins and drug delivery in cancer. Clinical Cancer Research, 23(6), 1373–1378. https://doi.org/10.1158/1078-0432.Ccr-16-0862.
Moniaux, N., Escande, F., Porchet, N., Aubert, J. P., & Batra, S. K. (2001). Structural organization and classification of the human mucin genes. Frontiers in Bioscience, 6, D1192–D1206. https://doi.org/10.2741/moniaux.
Messager, M., Lefevre, J. H., Pichot-Delahaye, V., Souadka, A., Piessen, G., & Mariette, C. (2011). The impact of perioperative chemotherapy on survival in patients with gastric signet ring cell adenocarcinoma: A multicenter comparative study. Annals of Surgery, 254(5), 684–693; discussion 693. https://doi.org/10.1097/SLA.0b013e3182352647.
Leal, J., Smyth, H. D. C., & Ghosh, D. (2017). Physicochemical properties of mucus and their impact on transmucosal drug delivery. International Journal of Pharmaceutics, 532(1), 555–572. https://doi.org/10.1016/j.ijpharm.2017.09.018.
Bhat, P. G., Flanagan, D. R., & Donovan, M. D. (1996). Drug diffusion through cystic fibrotic mucus: Steady-state permeation, rheologic properties, and glycoprotein morphology. Journal of Pharmaceutical Sciences, 85(6), 624–630. https://doi.org/10.1021/js950381s.
Suh, H., Pillai, K., & Morris, D. L. (2017). Mucins in pancreatic cancer: Biological role, implications in carcinogenesis and applications in diagnosis and therapy. American Journal of Cancer Research, 7(6), 1372–1383.
Kalra, A. V., & Campbell, R. B. (2009). Mucin overexpression limits the effectiveness of 5-FU by reducing intracellular drug uptake and antineoplastic drug effects in pancreatic tumours. European Journal of Cancer, 45(1), 164–173. https://doi.org/10.1016/j.ejca.2008.10.008.
Shaw, L. R., Irwin, W. J., Grattan, T. J., & Conway, B. R. (2005). The influence of excipients on the diffusion of ibuprofen and paracetamol in gastric mucus. International Journal of Pharmaceutics, 290(1-2), 145–154. https://doi.org/10.1016/j.ijpharm.2004.11.028.
Sigurdsson, H. H., Kirch, J., & Lehr, C. M. (2013). Mucus as a barrier to lipophilic drugs. International Journal of Pharmaceutics, 453(1), 56–64. https://doi.org/10.1016/j.ijpharm.2013.05.040.
Jonckheere, N., Skrypek, N., & Van Seuningen, I. (2014). Mucins and tumor resistance to chemotherapeutic drugs. Biochimica et Biophysica Acta, 1846(1), 142–151. https://doi.org/10.1016/j.bbcan.2014.04.008.
Jentoft, N. (1990). Why are proteins O-glycosylated? Trends in Biochemical Sciences, 15(8), 291–294. https://doi.org/10.1016/0968-0004(90)90014-3.
van de Wiel-van Kemenade, E., Ligtenberg, M. J., de Boer, A. J., Buijs, F., Vos, H. L., Melief, C. J., Hilkens, J., & Figdor, C. G. (1993). Episialin (MUC1) inhibits cytotoxic lymphocyte-target cell interaction. Journal of Immunology, 151(2), 767–776.
Bhatia, R., Gautam, S. K., Cannon, A., Thompson, C., Hall, B. R., Aithal, A., Banerjee, K., Jain, M., Solheim, J. C., Kumar, S., & Batra, S. K. (2019). Cancer-associated mucins: role in immune modulation and metastasis. Cancer Metastasis Reviews, 38(1-2), 223–236. https://doi.org/10.1007/s10555-018-09775-0.
Hollingsworth, M. A., & Swanson, B. J. (2004). Mucins in cancer: Protection and control of the cell surface. Nature Reviews. Cancer, 4(1), 45–60. https://doi.org/10.1038/nrc1251.
Nath, S., & Mukherjee, P. (2014). MUC1: A multifaceted oncoprotein with a key role in cancer progression. Trends in Molecular Medicine, 20(6), 332–342. https://doi.org/10.1016/j.molmed.2014.02.007.
Nath, S., Daneshvar, K., Roy, L. D., Grover, P., Kidiyoor, A., Mosley, L., Sahraei, M., & Mukherjee, P. (2013). MUC1 induces drug resistance in pancreatic cancer cells via upregulation of multidrug resistance genes. Oncogenesis, 2(6), e51. https://doi.org/10.1038/oncsis.2013.16.
Pitroda, S. P., Khodarev, N. N., Beckett, M. A., Kufe, D. W., & Weichselbaum, R. R. (2009). MUC1-induced alterations in a lipid metabolic gene network predict response of human breast cancers to tamoxifen treatment. Proceedings of the National Academy of Sciences of the United States of America, 106(14), 5837–5841. https://doi.org/10.1073/pnas.0812029106.
Jin, W., Liao, X., Lv, Y., Pang, Z., Wang, Y., Li, Q., Liao, Y., Ye, Q., Chen, G., Zhao, K., & Huang, L. (2017). MUC1 induces acquired chemoresistance by upregulating ABCB1 in EGFR-dependent manner. Cell Death & Disease, 8(8), e2980. https://doi.org/10.1038/cddis.2017.378.
Ham, S. Y., Kwon, T., Bak, Y., Yu, J. H., Hong, J., Lee, S. K., Yu, D. Y., & Yoon, D. Y. (2016). Mucin 1-mediated chemoresistance in lung cancer cells. Oncogenesis, 5(1), e185. https://doi.org/10.1038/oncsis.2015.47.
Hagmann, W., Jesnowski, R., & Löhr, J. M. (2010). Interdependence of gemcitabine treatment, transporter expression, and resistance in human pancreatic carcinoma cells. Neoplasia, 12(9), 740–747. https://doi.org/10.1593/neo.10576.
Wang, W., Abbruzzese, J. L., Evans, D. B., Larry, L., Cleary, K. R., & Chiao, P. J. (1999). The nuclear factor-kappa B RelA transcription factor is constitutively activated in human pancreatic adenocarcinoma cells. Clinical Cancer Research, 5(1), 119–127.
Arlt, A., Gehrz, A., Müerköster, S., Vorndamm, J., Kruse, M. L., Fölsch, U. R., & Schäfer, H. (2003). Role of NF-kappaB and Akt/PI3K in the resistance of pancreatic carcinoma cell lines against gemcitabine-induced cell death. Oncogene, 22(21), 3243–3251. https://doi.org/10.1038/sj.onc.1206390.
Skrypek, N., Duchêne, B., Hebbar, M., Leteurtre, E., van Seuningen, I., & Jonckheere, N. (2013). The MUC4 mucin mediates gemcitabine resistance of human pancreatic cancer cells via the Concentrative Nucleoside Transporter family. Oncogene, 32(13), 1714–1723. https://doi.org/10.1038/onc.2012.179.
Singh, A. P., Moniaux, N., Chauhan, S. C., Meza, J. L., & Batra, S. K. (2004). Inhibition of MUC4 expression suppresses pancreatic tumor cell growth and metastasis. Cancer Research, 64(2), 622–630. https://doi.org/10.1158/0008-5472.can-03-2636.
Carraway, K. L., Perez, A., Idris, N., Jepson, S., Arango, M., Komatsu, M., Haq, B., Price-Schiavi A., Zhang, J., & Carraway, C. (2002). Muc4/sialomucin complex, the intramembrane ErbB2 ligand, in cancer and epithelia: To protect and to survive. Progress in Nucleic Acid Research and Molecular Biology, 71, 149–185. https://doi.org/10.1016/s0079-6603(02)71043-x.
Chaturvedi, P., Singh, A. P., Chakraborty, S., Chauhan, S. C., Bafna, S., Meza, J. L., Singh, P. K., Hollingsworth, M. A., Mehta, P. P., & Batra, S. K. (2008). MUC4 mucin interacts with and stabilizes the HER2 oncoprotein in human pancreatic cancer cells. Cancer Research, 68(7), 2065–2070. https://doi.org/10.1158/0008-5472.Can-07-6041.
Ramsauer, V. P., Pino, V., Farooq, A., Carothers Carraway, C. A., Salas, P. J., & Carraway, K. L. (2006). Muc4-ErbB2 complex formation and signaling in polarized CACO-2 epithelial cells indicate that Muc4 acts as an unorthodox ligand for ErbB2. Molecular Biology of the Cell, 17(7), 2931–2941. https://doi.org/10.1091/mbc.e05-09-0895.
Park, S. H., Lee, J. H., Berek, J. S., & Hu, M. C. (2014). Auranofin displays anticancer activity against ovarian cancer cells through FOXO3 activation independent of p53. International Journal of Oncology, 45(4), 1691–1698. https://doi.org/10.3892/ijo.2014.2579.
Bae, J. S., Lee, J., Park, Y., Park, K., Kim, J. R., Cho, D. H., Jang, K. Y., & Park, S. H. (2017). Attenuation of MUC4 potentiates the anticancer activity of auranofin via regulation of the Her2/Akt/FOXO3 pathway in ovarian cancer cells. Oncology Reports, 38(4), 2417–2425. https://doi.org/10.3892/or.2017.5853.
Pothuraju, R., Rachagani, S., Krishn, S. R., Chaudhary, S., Nimmakayala, R. K., Siddiqui, J. A., Ganguly, K., Lakshmanan, I., Cox, J. L., Mallya, K., Kaur, S., & Batra, S. K. (2020). Molecular implications of MUC5AC-CD44 axis in colorectal cancer progression and chemoresistance. Molecular Cancer, 19(1), 37. https://doi.org/10.1186/s12943-020-01156-y.
Sheng, Y., Ng, C. P., Lourie, R., Shah, E. T., He, Y., Wong, K. Y., Seim, I., Oancea, I., Morais, C., Jeffery, P. L., Hooper, J., Gobe, G. C., & McGuckin, M. A. (2017). MUC13 overexpression in renal cell carcinoma plays a central role in tumor progression and drug resistance. International Journal of Cancer, 140(10), 2351–2363. https://doi.org/10.1002/ijc.30651.
Xu, Z., Liu, Y., Yang, Y., Wang, J., Zhang, G., Liu, Z., Fu, H., Wang, Z., Liu, H., & Xu, J. (2017). High expression of Mucin13 associates with grimmer postoperative prognosis of patients with non-metastatic clear-cell renal cell carcinoma. Oncotarget, 8(5), 7548–7558. https://doi.org/10.18632/oncotarget.13692.
Lakshmanan, I., Salfity, S., Seshacharyulu, P., Rachagani, S., Thomas, A., Das, S., Majhi, P. D., Nimmakayala, R. K., Vengoji, R., Lele, S. M., Ponnusamy, M. P., Batra, S. K., & Ganti, A. K. (2017). MUC16 regulates TSPYL5 for lung cancer cell growth and chemoresistance by suppressing p53. Clinical Cancer Research, 23(14), 3906–3917. https://doi.org/10.1158/1078-0432.Ccr-16-2530.
Das, S., Rachagani, S., Torres-Gonzalez, M. P., Lakshmanan, I., Majhi, P. D., Smith, L. M., Wagner, K., & Batra, S. K. (2015). Carboxyl-terminal domain of MUC16 imparts tumorigenic and metastatic functions through nuclear translocation of JAK2 to pancreatic cancer cells. Oncotarget, 6(8), 5772–5787. https://doi.org/10.18632/oncotarget.3308.
Boivin, M., Lane, D., Piché, A., & Rancourt, C. (2009). CA125 (MUC16) tumor antigen selectively modulates the sensitivity of ovarian cancer cells to genotoxic drug-induced apoptosis. Gynecologic Oncology, 115(3), 407–413. https://doi.org/10.1016/j.ygyno.2009.08.007.
Weissman, I. L. (2000). Stem cells: units of development, units of regeneration, and units in evolution. Cell, 100(1), 157–168. https://doi.org/10.1016/s0092-8674(00)81692-x.
Zhao, W., Ji, X., Zhang, F., Li, L., & Ma, L. (2012). Embryonic stem cell markers. Molecules, 17(6), 6196–6236. https://doi.org/10.3390/molecules17066196.
Yan, Q., Yao, D., Wei, L. L., Huang, Y., Myers, J., Zhang, L., Xin, W., Shim, J., Man, Y., Petryniak, B., Gerson, S., Lowe, J. B., & Zhou, L. (2010). O-fucose modulates Notch-controlled blood lineage commitment. The American Journal of Pathology, 176(6), 2921–2934. https://doi.org/10.2353/ajpath.2010.090702.
Seth, A., Machingo, Q. J., Fritz, A., & Shur, B. D. (2010). Core fucosylation is required for midline patterning during zebrafish development. Developmental Dynamics, 239(12), 3380–3390. https://doi.org/10.1002/dvdy.22475.
Greaves, M., & Maley, C. C. (2012). Clonal evolution in cancer. Nature, 481(7381), 306–313. https://doi.org/10.1038/nature10762.
Gupta, P. B., Onder, T. T., Jiang, G., Tao, K., Kuperwasser, C., Weinberg, R. A., & Lander, E. S. (2009). Identification of selective inhibitors of cancer stem cells by high-throughput screening. Cell, 138(4), 645–659. https://doi.org/10.1016/j.cell.2009.06.034.
Alam, M., Ahmad, R., Rajabi, H., Kharbanda, A., & Kufe, D. (2013). MUC1-C oncoprotein activates ERK → C/EBPβ signaling and induction of aldehyde dehydrogenase 1A1 in breast cancer cells. The Journal of Biological Chemistry, 288(43), 30892–30903. https://doi.org/10.1074/jbc.M113.477158.
Engelmann, K., Shen, H., & Finn, O. J. (2008). MCF7 side population cells with characteristics of cancer stem/progenitor cells express the tumor antigen MUC1. Cancer Research, 68(7), 2419–2426. https://doi.org/10.1158/0008-5472.Can-07-2249.
Stroopinsky, D., Rosenblatt, J., Ito, K., Mills, H., Yin, L., Rajabi, H., Vasir, B., Kufe, T., Luptakova, K., Arnason, J., Nardella, C., Levine, J. D., Joyce, R. M., Galinsky, I., Reiter, Y., Stone, R. M., Pandolfi, P. P., Kufe, D., & Avigan, D. (2013). MUC1 is a potential target for the treatment of acute myeloid leukemia stem cells. Cancer Research, 73(17), 5569–5579. https://doi.org/10.1158/0008-5472.Can-13-0677.
Guo, M., Luo, B., Pan, M., Li, M., Xu, H., Zhao, F., & Dou, J. (2020). Colorectal cancer stem cell vaccine with high expression of MUC1 serves as a novel prophylactic vaccine for colorectal cancer. International Immunopharmacology, 88, 106850. https://doi.org/10.1016/j.intimp.2020.106850.
Guo, M., You, C., Dong, W., Luo, B., Wu, Y., Chen, Y., Li, J., Pan, M., Li, M., Zhao, F., & Dou, J. (2020). The surface dominant antigen MUC1 is required for colorectal cancer stem cell vaccine to exert anti-tumor efficacy. Biomedicine & Pharmacotherapy, 132, 110804. https://doi.org/10.1016/j.biopha.2020.110804.
Curry, J. M., Thompson, K. J., Rao, S. G., Besmer, D. M., Murphy, A. M., Grdzelishvili, V. Z., Ahrens, W. A., McKillop, I. H., Sindram, D., Iannitti, D. A., Martinie, J. B., & Mukherjee, P. (2013). The use of a novel MUC1 antibody to identify cancer stem cells and circulating MUC1 in mice and patients with pancreatic cancer. Journal of Surgical Oncology, 107(7), 713–722. https://doi.org/10.1002/jso.23316.
Ponnusamy, M. P., Seshacharyulu, P., Vaz, A., Dey, P., & Batra, S. K. (2011). MUC4 stabilizes HER2 expression and maintains the cancer stem cell population in ovarian cancer cells. Journal of Ovarian Research, 4(1), 7. https://doi.org/10.1186/1757-2215-4-7.
Mimeault, M., Johansson, S. L., Senapati, S., Momi, N., Chakraborty, S., & Batra, S. K. (2010). MUC4 down-regulation reverses chemoresistance of pancreatic cancer stem/progenitor cells and their progenies. Cancer Letters, 295(1), 69–84. https://doi.org/10.1016/j.canlet.2010.02.015.
Jimeno, A., Feldmann, G., Suárez-Gauthier, A., Rasheed, Z., Solomon, A., Zou, G. M., Rubio-Viqueira, B., García-García, E., López-Ríos, F., Matsui, W., Maitra, A., & Hidalgo, M. (2009). A direct pancreatic cancer xenograft model as a platform for cancer stem cell therapeutic development. Molecular Cancer Therapeutics, 8(2), 310–314. https://doi.org/10.1158/1535-7163.Mct-08-0924.
Zhou, J., Wang, C. Y., Liu, T., Wu, B., Zhou, F., Xiong, J. X., Wu, H. S., Tao, J., Zhao, G., Yang, M., & Gou, S. M. (2008). Persistence of side population cells with high drug efflux capacity in pancreatic cancer. World Journal of Gastroenterology, 14(6), 925–930. https://doi.org/10.3748/wjg.14.925.
Ganguly, K., Krishn, S. R., Rachagani, S., Jahan, R., Shah, A., Nallasamy, P., Rauth, S., Atri, P., Cox, J. L., Pothuraju, R., Smith, L. M., Ayala, S., Evans, C., Ponusamy, M. P., Kumar, S., Kaur, S., & Batra, S. K. (2020). Secretory mucin 5 AC promotes neoplastic progression by augmenting KLF4-mediated pancreatic cancer cell stemness. Cancer Research. https://doi.org/10.1158/0008-5472.Can-20-1293.
Zhang, H., Yang, Y., Wang, Y., Gao, X., Wang, W., Liu, H., He, H., Liang, Y., Pan, K., Wu, H., Shi, J., Xue, H., Liang, L., Cai, Z., Fan, Y., & Zhang, Y. (2015). Relationship of tumor marker CA125 and ovarian tumor stem cells: Preliminary identification. Journal of Ovarian Research, 8, 19. https://doi.org/10.1186/s13048-015-0132-8.
Lim, S., Becker, A., Zimmer, A., Lu, J., Buettner, R., & Kirfel, J. (2013). SNAI1-mediated epithelial-mesenchymal transition confers chemoresistance and cellular plasticity by regulating genes involved in cell death and stem cell maintenance. PLoS One, 8(6), e66558. https://doi.org/10.1371/journal.pone.0066558.
Yamada, S., Fuchs, B. C., Fujii, T., Shimoyama, Y., Sugimoto, H., Nomoto, S., Takeda, S., Tanabe, K. K., Kodera, Y., & Nakao, A. (2013). Epithelial-to-mesenchymal transition predicts prognosis of pancreatic cancer. Surgery, 154(5), 946–954. https://doi.org/10.1016/j.surg.2013.05.004.
Comamala, M., Pinard, M., Thériault, C., Matte, I., Albert, A., Boivin, M., Beaudin, J., Piché, A., & Rancourt, C. (2011). Downregulation of cell surface CA125/MUC16 induces epithelial-to-mesenchymal transition and restores EGFR signalling in NIH:OVCAR3 ovarian carcinoma cells. British Journal of Cancer, 104(6), 989–999. https://doi.org/10.1038/bjc.2011.34.
Horn, G., Gaziel, A., Wreschner, D. H., Smorodinsky, N. I., & Ehrlich, M. (2009). ERK and PI3K regulate different aspects of the epithelial to mesenchymal transition of mammary tumor cells induced by truncated MUC1. Experimental Cell Research, 315(8), 1490–1504. https://doi.org/10.1016/j.yexcr.2009.02.011.
Roy, L. D., Sahraei, M., Subramani, D. B., Besmer, D., Nath, S., Tinder, T. L., Bajaj, E., Shanmugam, K., Lee, Y. Y., Hwang, S. I. L., Gendler, S. J., & Mukherjee, P. (2011). MUC1 enhances invasiveness of pancreatic cancer cells by inducing epithelial to mesenchymal transition. Oncogene, 30(12), 1449–1459. https://doi.org/10.1038/onc.2010.526.
Ponnusamy, M. P., Lakshmanan, I., Jain, M., Das, S., Chakraborty, S., Dey, P., & Batra, S. K. (2010). MUC4 mucin-induced epithelial to mesenchymal transition: A novel mechanism for metastasis of human ovarian cancer cells. Oncogene, 29(42), 5741–5754. https://doi.org/10.1038/onc.2010.309.
Lakshmanan, I., Rachagani, S., Hauke, R., Krishn, S. R., Paknikar, S., Seshacharyulu, P., Karmakar, S., Nimmakayala, R. K., Kaushik, G., Johansson, S. L., Carey, G. B., Ponnusamy, M. P., Kaur, S., Batra, S. K., & Ganti, A. K. (2016). MUC5AC interactions with integrin β4 enhances the migration of lung cancer cells through FAK signaling. Oncogene, 35(31), 4112–4121. https://doi.org/10.1038/onc.2015.478.
Hata, T., Rajabi, H., Yamamoto, M., Jin, C., Ahmad, R., Zhang, Y., Kui, L., Li, W., Yasumizu, Y., Hong, D., Miyo, M., Hiraki, M., Maeda, T., Suzuki, Y., Takahashi, H., Samur, M., & Kufe, D. (2019). Targeting MUC1-C inhibits TWIST1 signaling in triple-negative breast cancer. Molecular Cancer Therapeutics, 18(10), 1744–1754. https://doi.org/10.1158/1535-7163.Mct-19-0156.
Rajabi, H., Alam, M., Takahashi, H., Kharbanda, A., Guha, M., Ahmad, R., & Kufe, D. (2014). MUC1-C oncoprotein activates the ZEB1/miR-200c regulatory loop and epithelial-mesenchymal transition. Oncogene, 33(13), 1680–1689. https://doi.org/10.1038/onc.2013.114.
Grover, P., Nath, S., Nye, M. D., Zhou, R., Ahmad, M., & Mukherjee, P. (2018). SMAD4-independent activation of TGF-β signaling by MUC1 in a human pancreatic cancer cell line. Oncotarget, 9(6), 6897–6910. https://doi.org/10.18632/oncotarget.23966.
Rachagani, S., Macha, M. A., Ponnusamy, M. P., Haridas, D., Kaur, S., Jain, M., & Batra, S. K. (2012). MUC4 potentiates invasion and metastasis of pancreatic cancer cells through stabilization of fibroblast growth factor receptor 1. Carcinogenesis, 33(10), 1953–1964. https://doi.org/10.1093/carcin/bgs225.
Muniyan, S., Haridas, D., Chugh, S., Rachagani, S., Lakshmanan, I., Gupta, S., Seshacharyulu, P., Smith, L. M., Ponnusamy, M. P., & Batra S. K. (2016). MUC16 contributes to the metastasis of pancreatic ductal adenocarcinoma through focal adhesion mediated signaling mechanism. Genes & Cancer, 7(3–4), 110–124. https://doi.org/10.18632/genesandcancer.104.
Hanahan, D., & Weinberg, R. A. (2011). Hallmarks of cancer: The next generation. Cell, 144(5), 646–674. https://doi.org/10.1016/j.cell.2011.02.013.
Frezza, C. (2020). Metabolism and cancer: The future is now. British Journal of Cancer, 122(2), 133–135. https://doi.org/10.1038/s41416-019-0667-3.
Jung, J. G., & Le, A. (2018). Targeting metabolic cross talk between cancer cells and cancer-associated fibroblasts. Advances in Experimental Medicine and Biology, 1063, 167–178. https://doi.org/10.1007/978-3-319-77736-8_12.
Chaika, N. V., Gebregiworgis, T., Lewallen, M. E., Purohit, V., Radhakrishnan, P., Liu, X., Zhang, B., Mehla, K., Brown, R. B., Caffrey, T., Yu, F., Johnson, K. R., Powers, R., Hollingsworth, M. A., & Singh, P. K. (2012). MUC1 mucin stabilizes and activates hypoxia-inducible factor 1 alpha to regulate metabolism in pancreatic cancer. Proceedings of the National Academy of Sciences of the United States of America, 109(34), 13787–13792. https://doi.org/10.1073/pnas.1203339109.
Kosugi, M., Ahmad, R., Alam, M., Uchida, Y., & Kufe, D. (2011). MUC1-C oncoprotein regulates glycolysis and pyruvate kinase M2 activity in cancer cells. PLoS One, 6(11), e28234. https://doi.org/10.1371/journal.pone.0028234.
Raina, D., Kharbanda, S., & Kufe, D. (2004). The MUC1 oncoprotein activates the anti-apoptotic phosphoinositide 3-kinase/Akt and Bcl-xL pathways in rat 3Y1 fibroblasts. The Journal of Biological Chemistry, 279(20), 20607–20612. https://doi.org/10.1074/jbc.M310538200.
Mehla, K., & Singh, P. K. (2014). MUC1: A novel metabolic master regulator. Biochimica et Biophysica Acta, 1845(2), 126–135. https://doi.org/10.1016/j.bbcan.2014.01.001.
Yoshida, K., & Miki, Y. (2004). Role of BRCA1 and BRCA2 as regulators of DNA repair, transcription, and cell cycle in response to DNA damage. Cancer Science, 95(11), 866–871. https://doi.org/10.1111/j.1349-7006.2004.tb02195.x.
Gunda, V., Souchek, J., Abrego, J., Shukla, S. K., Goode, G. D., Vernucci, E., Dasgupta, A., Chaika, N. V., King, R. J., Li, S., Wang, S., Yu, F., Bessho, T., Lin, C., & Singh, P. K. (2017). MUC1-mediated metabolic alterations regulate response to radiotherapy in pancreatic cancer. Clinical Cancer Research, 23(19), 5881–5891. https://doi.org/10.1158/1078-0432.Ccr-17-1151.
Chiyoda, T., Hart, P. C., Eckert, M. A., McGregor, S. M., Lastra, R. R., Hamamoto, R., Nakamura, Y., Yamada, S. D., Olopade, O. I., Lengyel, E., & Romero, I. L. (2017). Loss of BRCA1 in the cells of origin of ovarian cancer induces glycolysis: A window of opportunity for ovarian cancer Chemoprevention. Cancer Prevention Research (Philadelphia, Pa.), 10(4), 255–266. https://doi.org/10.1158/1940-6207.Capr-16-0281.
Fu, X., Tang, N., Xie, W. Q., Mao, L., & Qiu, Y. D. (2020). MUC1 promotes glycolysis through inhibiting BRCA1 expression in pancreatic cancer. Chinese Journal of Natural Medicines, 18(3), 178–185. https://doi.org/10.1016/s1875-5364(20)30019-4.
Leng, Y., Cao, C., Ren, J., Huang, L., Chen, D., Ito, M., & Kufe, D. (2007). Nuclear import of the MUC1-C oncoprotein is mediated by nucleoporin Nup62. The Journal of Biological Chemistry, 282(27), 19321–19330. https://doi.org/10.1074/jbc.M703222200.
Raina, D., Ahmad, R., Rajabi, H., Panchamoorthy, G., Kharbanda, S., & Kufe, D. (2012). Targeting cysteine-mediated dimerization of the MUC1-C oncoprotein in human cancer cells. International Journal of Oncology, 40(5), 1643–1649. https://doi.org/10.3892/ijo.2011.1308.
Raina, D., Kosugi, M., Ahmad, R., Panchamoorthy, G., Rajabi, H., Alam, M., Shimamura, T., Shapiro, G. I., Supko, J., Kharbanda, S., & Kufe, D. (2011). Dependence on the MUC1-C oncoprotein in non-small cell lung cancer cells. Molecular Cancer Therapeutics, 10(5), 806–816. https://doi.org/10.1158/1535-7163.Mct-10-1050.
GongSun, X., Zhao, Y., Jiang, B., Xin, Z., Shi, M., Song, L., Qin, Q. M., Wang, Q., & Liu, X. Y. (2019). Inhibition of MUC1-C regulates metabolism by AKT pathway in esophageal squamous cell carcinoma. Journal of Cellular Physiology, 234(7), 12019–12028. https://doi.org/10.1002/jcp.27863.
Kumari, S., Khan, S., Gupta, S. C., Kashyap, V. K., Yallapu, M. M., Chauhan, S. C., & Jaggi, M. (2018). MUC13 contributes to rewiring of glucose metabolism in pancreatic cancer. Oncogenesis, 7(2), 19. https://doi.org/10.1038/s41389-018-0031-0.
Haridas, D., Chakraborty, S., Ponnusamy, M. P., Lakshmanan, I., Rachagani, S., Cruz, E., Kumar, S., Das, S., Lele, S. M., Anderson, J. M., Wittel, U. A., Hollingsworth, M. A., & Batra, S. K. (2011). Pathobiological implications of MUC16 expression in pancreatic cancer. PLoS One, 6(10), e26839. https://doi.org/10.1371/journal.pone.0026839.
Shukla, S. K., Gunda, V., Abrego, J., Haridas, D., Mishra, A., Souchek, J., Chaika, N. V., Yu, F., Sasson, A. R., Lazenby, A. J., Batra, S. K., & Singh, P. K. (2015). MUC16-mediated activation of mTOR and c-Myc reprograms pancreatic cancer metabolism. Oncotarget, 6(22), 19118–19131. https://doi.org/10.18632/oncotarget.4078.
Shimobayashi, M., & Hall, M. N. (2014). Making new contacts: The mTOR network in metabolism and signalling crosstalk. Nature Reviews. Molecular Cell Biology, 15(3), 155–162. https://doi.org/10.1038/nrm3757.
Düvel, K., Yecies, J. L., Menon, S., Raman, P., Lipovsky, A. I., Souza, A. L., Triantafellow, E., Ma, Q., Gorski, R., Cleaver, S., Vander Heiden, M. G., MacKeigan, J. P., Finan, P. M., Clish, C. B., Murphy, L. O., & Manning, B. D. (2010). Activation of a metabolic gene regulatory network downstream of mTOR complex 1. Molecular Cell, 39(2), 171–183. https://doi.org/10.1016/j.molcel.2010.06.022.
Nomura, D. K., & Cravatt, B. F. (2013). Lipid metabolism in cancer. Biochimica et Biophysica Acta, 1831(10), 1497–1498. https://doi.org/10.1016/j.bbalip.2013.08.001.
Mashima, T., Seimiya, H., & Tsuruo, T. (2009). De novo fatty-acid synthesis and related pathways as molecular targets for cancer therapy. British Journal of Cancer, 100(9), 1369–1372. https://doi.org/10.1038/sj.bjc.6605007.
Liu, Y. (2006). Fatty acid oxidation is a dominant bioenergetic pathway in prostate cancer. Prostate Cancer and Prostatic Diseases, 9(3), 230–234. https://doi.org/10.1038/sj.pcan.4500879.
Mukherjee, A., Wu, J., Barbour, S., & Fang, X. (2012). Lysophosphatidic acid activates lipogenic pathways and de novo lipid synthesis in ovarian cancer cells. The Journal of Biological Chemistry, 287(30), 24990–25000. https://doi.org/10.1074/jbc.M112.340083.
Martel, P. M., Bingham, C. M., McGraw, C. J., Baker, C. L., Morganelli, P. M., Meng, M. L., Armstrong, J. M., Moncur, J. T., & Kinlaw, W. B., (2006). S14 protein in breast cancer cells: Direct evidence of regulation by SREBP-1c, superinduction with progestin, and effects on cell growth. Experimental Cell Research, 312(3), 278–288. https://doi.org/10.1016/j.yexcr.2005.10.022.
Migita, T., Narita, T., Nomura, K., Miyagi, E., Inazuka, F., Matsuura, M., Ushijima, M., Mashima, T., Seimiya, H., Satoh, Y., Okumura, S., Nakagawa, K., & Ishikawa, Y. (2008). ATP citrate lyase: Activation and therapeutic implications in non-small cell lung cancer. Cancer Research, 68(20), 8547–8554. https://doi.org/10.1158/0008-5472.Can-08-1235.
Menendez, J. A., & Lupu, R. (2007). Fatty acid synthase and the lipogenic phenotype in cancer pathogenesis. Nature Reviews. Cancer, 7(10), 763–777. https://doi.org/10.1038/nrc2222.
Furuta, E., Pai, S. K., Zhan, R., Bandyopadhyay, S., Watabe, M., Mo, Y. Y., Hirota, S., Hosobe, S., Tsukada, T., Miura, K., Kamada, S., Saito, K., Iiizumi, M., Liu, W., Ericsson, J., & Watabe, K. (2008). Fatty acid synthase gene is upregulated by hypoxia via activation of Akt and sterol regulatory element binding protein-1. Cancer Research, 68(4), 1003–1011. https://doi.org/10.1158/0008-5472.Can-07-2489.
Taylor-Papadimitriou, J., Burchell, J. M., Graham, R., & Beatson, R. (2018). Latest developments in MUC1 immunotherapy. Biochemical Society Transactions, 46(3), 659–668. https://doi.org/10.1042/bst20170400.
Panchamoorthy, G., Jin, C., Raina, D., Bharti, A., Yamamoto, M., Adeebge, D., Zhao, Q., Bronson, R., Jiang, S., Li, L., Suzuki, Y., Tagde, A., Ghoroghchian, P. P., Wong, K. K., Kharbanda, S., & Kufe, D. (2018). Targeting the human MUC1-C oncoprotein with an antibody-drug conjugate. JCI Insight, 3(12). https://doi.org/10.1172/jci.insight.99880.
Liberelle, M., Magnez, R., Thuru, X., Bencheikh, Y., Ravez, S., Quenon, C., Drucbert, A. S., Foulon, C., Melnyk, P., van Seuningen, I., & Lebègue, N. (2019). MUC4-ErbB2 oncogenic complex: Binding studies using microscale thermophoresis. Scientific Reports, 9(1), 16678. https://doi.org/10.1038/s41598-019-53099-0.
Das, S., & Batra, S. K. (2015). Understanding the unique attributes of MUC16 (CA125): Potential implications in targeted therapy. Cancer Research, 75(22), 4669–4674. https://doi.org/10.1158/0008-5472.Can-15-1050.
Aithal, A., Rauth, S., Kshirsagar, P., Shah, A., Lakshmanan, I., Junker, W. M., Jain, M., Ponnusamy, M. P., & Batra, S. K. (2018). MUC16 as a novel target for cancer therapy. Expert Opinion on Therapeutic Targets, 22(8), 675–686. https://doi.org/10.1080/14728222.2018.1498845.
Patel, S. P., Bristol, A., Saric, O., Wang, X. P., Dubeykovskiy, A., Arlen, P. M., & Morse, M. A. (2013). Anti-tumor activity of a novel monoclonal antibody, NPC-1C, optimized for recognition of tumor antigen MUC5AC variant in preclinical models. Cancer Immunology, Immunotherapy, 62(6), 1011–1019. https://doi.org/10.1007/s00262-013-1420-z.
Acknowledgements
We thank Kavita Mallya and Corinn E Grabow for their support.
Funding
The authors in this article were supported primarily by the following grants from the National Institutes of Health P01 CA217798, R01 CA210637, R01 CA183459, R01 CA195586, R01 CA201444, R01 CA228524, F99 CA234962, U01 CA200466, and U01 CA210240, and the Nebraska Department of Health and Human Services LB595.
Author information
Authors and Affiliations
Contributions
SM, SR, CZ, and KG, collected the related literature and drafted the manuscript. SM, SR, CZ, KG, IL, MPP, and SKB participated in the manuscript’s draft. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Ethics approval
Not applicable
Consent to participate
Not applicable
Consent for publication
Not applicable
Conflict of interest/Competing interests
SKB is one of the co-founders of Sanguine Diagnostics and Therapeutics, Inc. The other authors disclosed no potential conflicts of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Marimuthu, S., Rauth, S., Ganguly, K. et al. Mucins reprogram stemness, metabolism and promote chemoresistance during cancer progression. Cancer Metastasis Rev 40, 575–588 (2021). https://doi.org/10.1007/s10555-021-09959-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-021-09959-1