Abstract
There has been a surge of interest in recent years in understanding the intricate mechanisms underlying cancer progression and treatment resistance. One molecule that has recently emerged in these mechanisms is MUC13 mucin, a transmembrane glycoprotein. Researchers have begun to unravel the molecular complexity of MUC13 and its impact on cancer biology. Studies have shown that MUC13 overexpression can disrupt normal cellular polarity, leading to the acquisition of malignant traits. Furthermore, MUC13 has been associated with increased cancer plasticity, allowing cells to undergo epithelial-mesenchymal transition (EMT) and metastasize. Notably, MUC13 has also been implicated in the development of chemoresistance, rendering cancer cells less responsive to traditional treatment options. Understanding the precise role of MUC13 in cellular plasticity, and chemoresistance could pave the way for the development of targeted therapies to combat cancer progression and enhance treatment efficacy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Cancer is one of the leading causes of mortality throughout the world, and it is projected to account for 10 million fatalities in the year 2020 [1]. The main cancer treatments that are currently available are targeted therapy, radiation, laser, immunotherapy, gene therapy, hormone therapy, and chemotherapy. The most common and promising cancer treatment is chemotherapy [2,3,4]. Chemotherapeutic drug resistance is the leading cause of relapses and low survival rates in cancer patients, despite recent breakthroughs in cancer therapy. Most of the drugs that are used to treat the severe forms of cancer have been demonstrated to be resistant to chemotherapy. Metastases cancer cell resistant to chemotherapy causes 90% of failed cancer treatments, making it a major obstacle to cancer treatment [5]. For instance, a relapse of breast cancer caused by resistance to doxorubicin, paclitaxel, 5-fluorouracil, cyclophosphamide, and carboplatin is linked with a worse prognosis and shorter survival time [6,7,8,9,10,11,12,13].
Studies have shown that treatment resistance is either directly or indirectly responsible for between 80 and 90% of cancer patients’ deaths [14]. It is widely held that therapeutic medications can only destroy actively dividing cancer cells; however, quiescent cancer cells, also called as cancer stem cells (CSCs), are believed to be resistant to these treatments and continue to thrive. According to the findings of studies, these CSCs are to blame for cancer’s ability to recur, to relapse, to metastasize, and to become resistant to treatment medicines [15,16,17]. After seeing the histological similarities that exist between cancers and embryonic tissues, the notion of CSCs was originally described for the very first time in the early nineties, which is more than a century ago. On the other hand, it was demonstrated in 1937 that a cell transplanted from a mouse tumor can result in the development of a new tumor in the recipient mouse [18, 19]. It has been shown that CSCs may survive chemotherapy and even regenerate following treatment, despite the fact that the majority of chemotherapy medicines are effective to kill the tumor cells [20]. For instance, use of temozolomide results in the death of differentiated GBM cells but leads to an increase in the number of glioma stem cells in patient-derived cell lines [21]. The methods by which CSCs avoid the effects of chemotherapy are not well known. CSCs are responsible for orchestrating chemoresistance by regulating a number of different processes, including EMT, ROS production, hypoxia, and ABC transporters [22]. CSCs have been shown to express several different transporter proteins, including the multidrug resistance protein, phospho-glycoprotein, and breast cancer resistant protein (BCRP) [23, 24]. During cancer progression, tumors adopt several changes including accumulation of CSCs, metabolism, stromal-tumor cell interactions, and epithelial mesenchymal transition (EMT) to offer plasticity to harboring tumor cells. This comprehensive review will focus on deciphering the significance of MUC13 mucin in cancer cell plasticity and drug resistance to provide valuable insights for the generation of innovative therapeutic methods.
2 Role of mucins in cancer biology
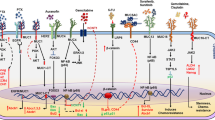
Mucins, a type of glycoproteins, have been found to play a significant role in cancers, particularly in their development and progression [25, 26]. Mucins are normally responsible for maintaining the mucus layer in different organs, such as the digestive, respiratory, and reproductive systems [25, 27,28,29,30,31,32]. In cancers, however, aberrant mucin expression and altered glycosylation patterns have been observed [33]. These changes can lead to the formation of a dense mucin-rich extracellular matrix, which promotes tumor growth, invasion, and metastasis (Fig. 1) [33,34,35,36,37,38,39]. Mucins have also been linked resistance to chemotherapy and immune evasion, further leads to the aggressiveness and treatment challenges posed by many cancers [40, 41]. Understanding the specific roles and mechanisms of mucins in different cancer types holds great potential for the development of targeted therapies and improved cancer management strategies [33,34,35,36,37,38,39].
Schematic representation of function of normal and aberrant MUC13 expression. The role of normal MUC13 is to provide protection from environmental toxin, pathogenic microbe insults, and homeostasis. If aberrant, MUC13 induces chemoresistance, metastasis, poor prognosis, and enrichment of cancer cell stemness
There is an increasing body of research that points to a relationship between mucins and the plasticity of cancer cells. Mucins have been shown to influence the epithelial-mesenchymal transition (EMT), a process often associated with increased cancer cell plasticity and metastatic potential [42]. Certain mucin proteins can induce or suppress EMT, thereby influencing cancer cell behavior [43]. Moreover, mucins can interact with various signaling pathways involved in cell plasticity regulation, such as the transforming growth factor-beta (TGF-β) pathway. It is widely recognized that TGF-β signaling stimulates the epithelial to mesenchymal transition and cancer cell plasticity. Understanding the relationship between mucins and cancer cell plasticity is important for designing specific therapies and improving patient outcomes. By deciphering the mechanisms underlying mucin-mediated plasticity, researchers can potentially identify new therapeutic targets and strategies to better control tumor progression and metastasis. This comprehensive review primarily focusses on MUC13, an emerging oncogene that plays crucial role in cancer cell plasticity and drug resistance, particularly in the context of its interaction with CSCs, provides valuable insights for the development of novel therapeutic approaches.
3 The structural features of MUC13
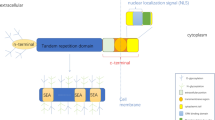
Mucins are broadly classified into two categories: secreted mucins and membrane-bound mucins. Secreted mucins, such as MUC2, MUC5AC, MUC5B, MUC6, and MUC19, primarily comprise viscoelastic gels. They form a protective network that defends the epithelial cells against various aggressors like inflammation, bacteria, viruses, pollutants, and pH changes [40, 44,45,46]. On the other hand, membrane-bound mucins are a group of protein anchored to the cell membrane. They include MUC1, MUC3, MUC4, MUC12, MUC13, MUC15, MUC16, MUC17, MUC20, and MUC21. These mucins play roles in cell-to-cell and cell-to-extracellular matrix interactions, as well as in cell signaling processes, owing to their specific structure and localization on the cell surface [47,48,49]. MUC13 is most notably characterized for protective functions in epithelial tissues and its large molecular weight (Fig. 2). MUC13 plays a crucial role in the intestinal epithelium by regulating tight junctions [50]. MUC13, like other mucins, has a tandem repeat (TR) domain rich in serine and threonine glycosylation sites. It has a cytoplasmic domain with putative phosphorylation sites and three epidermal growth factor domains. MUC13 has been found in the esophagus, trachea, kidney, small intestine, gastric epithelium, and large intestine under normal physiologic conditions [51]. According to its protective and lubricating role on the mucosal surface, MUC13 protein is generally found on the apical surface of epithelial cells. Mucin production, both secreted and linked with the cell membrane, offers a barrier against the colonization of harmful bacteria. Recent research has demonstrated the importance of altered MUC13 activity in variety of malignancies [51,52,53,54,55,56,57] (Table 1). Additionally, Filippou and colleagues conducted significant research on MUC13’s ability as a marker for cancer and other disorders [58]. Therefore, the role of MUC13 is beginning to be redefined through its expression in various cancers from a protective glycoprotein to a potential oncogene. In our recent article, the various isoforms of MUC13 were discussed. A total of five transcripts of MUC13 were identified and among all, two were protein coding transcripts. These two protein coding isoforms further structurally identified as short form MUC13 (non-tumorigenic), and long form (tumorigenic). Multiple sequence alignment (MSA) was performed for better sequential elucidation of these forms of MUC13. The short of MUC13 having three functional domains while long form having five functional domains.
The presence of cytoplasmic domain in long form of MUC13 could be a main reason of tumorigenic effect. These structural and sequential features confirm the significance of MUC13 as a potential oncogene [79].
4 Oncogenic expression of MUC13 in various cancers
Prior research has suggested the significance of MUC13 overexpression in a number of malignancies [41,42,43,44,45,46,47, 80] (Table 1). MUC13 has gained attention for its potential use in prognostic applications. Studies have utilized datasets from The Cancer Genome Atlas (TCGA) to investigate MUC13 correlation with patient survival in different types of cancers. Higher levels of MUC13 expression in tumor tissues have been linked to poorer patient outcomes, such as reduced overall survival or disease-free survival rates in stomach, esophageal, and lung adenocarcinoma [80]. Dai and colleagues explored the protein expression profile of MUC13 in liver cancer (HCC) and examined its relationship with cell survival pathways [54]. They ultimately observed that MUC13 is significantly overexpressed in HCC tissues. They illustrated that the overexpression of MUC13 was strongly correlated with the size of the tumor, stage, invasion, and encapsulation. From this, it is evident that MUC13 interacts with several cell survivals signaling pathways to promote the proliferation and invasion in various cancers [41,42,43,44,45,46,47]. In a similar manner, functional assays have shown that MUC13 possesses strong oncogenic activity in liver cancer. Interestingly, Axin2 levels are modulated with MUC13 overexpression. It is assumed the cytoplasmic tail of MUC13 is involved in β-catenin nuclear localization. The clinical significance of MUC13 in liver cancer was also analyzed. It was observed that out of 119 patient samples, 51 samples had high MUC13 expression (42.9%). Furthermore, the expression of MUC13 has been associated to the number of copies of HBV and poor overall survival. A conducted ROC curve showed that in liver cancer, the connection of MUC13 upregulation and localized β-catenin has a stronger prognostic potential than alone. Emerging research for the potential of MUC13 in biomarker studies was performed by Fillppou and colleagues [52]. By employing ELISA assay, they performed an unprecedented quantification of serum MUC13 in healthy and liver cancer patients and observed a significantly elevated level in patients with liver cancer. These findings suggest that MUC13 holds promise for its use in prognostic applications. Moreover, further study is needed to determine the correlation of MUC13 in stage specific pathogenesis of various cancers.
4.1 MUC13 in pancreatic cancer
The prevalence of pancreatic cancer is abundant and is associated with extremely poor prognosis. Early diagnosis of this disease can be rather difficult to fulfill due to common misdiagnosis and lack of early representation [61]. In pancreatic cancer, MUC13 is aberrantly expressed and involved in tumor progression through various mechanisms such as HER2, PAK1, ERK, and Akt [57, 59] (Fig. 3). Recent studies suggest that upregulation of MUC13 increases cell growth, invasion, and migration of pancreatic cancer cells, which can be further enhanced by the addition of lactate [57, 59, 60]. Furthermore, MUC13 enhances glucose metabolism in cancer cells [60]. In another study, the tumor suppressor miR-145 was found to be associated with MUC13 upregulation in pancreatic cancer (Fig. 3). In short, the expression of miR-145 leads to the downregulation of MUC13, resulting in moderate growth of tumor cells [56]. MUC13 caused significant biophysical alterations in pancreatic cancer cells, and these cells displayed the distinctive phenotypic changes observed in prior research conducted from our lab. This study sheds light on the potential utility of biophysical measurements in cancer diagnosis and monitoring, as well as in elucidating the mechanistic consequences of genetic modification [61]. MUC13 expression was found in 94.6% of pancreatic ductal adenocarcinoma samples and 100% of pancreatic intraepithelial neoplasia (PanIN) lesions. In contrast, MUC13 expression was found to be low in tumor-adjacent tissues and to be faint or absent in normal pancreatic tissues. Positive correlations were found between nuclear MUC13 expression and nodular metastasis (P < 0.05), malignant intrusion of peripheral tissues (P < 0.5), and poor survival of patient (P < 0.05; prognostic AUC = 0.9) [59]. Intraductal papillary mucinous neoplasms (IPMN) were found to have higher MUC13 activity than non-mucinous cysts. This study indicates that MUC13 could serve as a diagnostic marker for distinguishing between low and high risk IPMNs [63]. Additionally, our recent integrative big transcriptomic analytic data suggests important function for MUC13 in pancreatic cancer. In a cohort of 179 tumors and 171 normal samples, MUC13 was observed to be a highly expressed gene with 3.73 log2 fold change and 92 ranked out of all upregulated genes [79]. The pairwise gene correlation analysis of MUC13, MUC1, and S100A4 shows a strong positive correlation. This analysis indicated that the MUC13 isoform analysis contained a total of 05 transcripts. Only two of the five transcripts—long and short—are protein-coding transcripts. It is intriguing to note that the long form of MUC13 was much more common in tumors than the short form [79]. The prognostic implications of MUC13 in cancer have sparked controversy, as various studies present contrasting findings. Thompson et al. [81] reported a study showing that higher expression of a specific splice variant of MUC13 is associated with significantly longer survival in pancreatic cancer. Contrary to these findings, other research groups have demonstrated that MUC13 expression is linked to a more aggressive phenotype in pancreatic ductal adenocarcinoma (PDAC) cell line models. The disparate findings may be attributed to various factors, including unique post-translational modifications such as glycosylation on the MUC13 protein or cytoplasmic tail-mediated mechanisms. These factors could influence the functional status of the MUC13 protein during tumorigenesis, leading to different outcomes in terms of survival and disease progression.
Functional interaction of MUC13 glycoprotein with cell survival signaling pathways. HER2, human epidermal growth factor receptor 2; TNFR1, tumor necrosis factor-alpha receptor 1; TYK2, tyrosine kinase 2; JAK1, Janus kinase 1; TRADD, tumor necrosis factor receptor 1-associated death domain protein; RIP1, receptor-interacting protein kinase 1; TRAF6, TNF receptor Associated Factor 6; MAPK, mitogen-activated protein kinase; PAK1, p21-activated kinase 1; GSK-3β, glycogen synthase kinase-3β; APC, adenomatous polyposis coli protein; IK-β, inhibitor of nuclear factor kappa B; NF-κB, nuclear factor kappa-light-chain-enhancer of activated B cells; ATM, ataxia-telangiectasia; PARP1, poly[ADP-ribose]polymerase 1; MEMO, mediator of cell motility; PIA, polymorphism interaction analysis; LEF-1/TCF, lymphoid enhancer factor 1/T cell factor; miR, microRNA
4.2 MUC13 in colon cancer
Less than 20% of colorectal cancer patients survive five years [82]. Studies suggest that the overexpression of mucins can lead to catastrophic diseases such as cancer [83, 84]. As discussed, mucins play a significant and invasive role in many cancers, and colon cancer has shown to be one of these [83, 85]. In colorectal cancer, MUC13 is frequently upregulated and shows abnormal localization [86]. This overexpression of MUC13 increases tumorigenesis and metastasis [55]. Moreover, the upregulation and aberrant localization of MUC13 may prevent cancer cells from adhering to the cell’s extracellular matrix and promote cell motility and invasion; therefore, indicating that MUC13 showing a dissatisfying involvement in colorectal tumorigenesis and metastasis [86, 87]. Furthermore, we recently reported the broad significance of MUC13 and YAP1 in colorectal cancer metastasis by targeting YAP1. Anchorage-independent circulating tumor cells gain a significant survival advantage via a novel molecular mechanism that is MUC13-YAP1 driven, resulting in efficient extravasation and aggressive cancer metastatic spread at unique peripheral sites [88].
4.3 MUC13 in stomach/gastric cancer
Gastric cancer, the fourth most leading cause of cancer deaths worldwide, also exhibits poor prognosis [89]. The 5-year survival rate for patients diagnosed with this disease is 33% since the representation of stomach cancer can easily be mistaken for indigestion or heartburn. In result, stomach cancer is often diagnosed in its advanced stages leading to catastrophic outcomes [90]. The expression level of MUC13 in gastric cancer tissues is notably higher than in normal tissues. Research demonstrates that MUC13 expression in gastric cancer tissues is considerably upregulated; however, results also indicate that decrease of miR-132 leads to an increase of MUC13 expression and elevated levels of HER2 and phosphorylation of ERK and Akt in gastric cancer cells [74]. Moreover, the overexpression of MUC13 enhances cell invasion, migration, HER2 levels, and phosphorylation of ERK and Akt.
4.4 MUC13 in liver cancer
Liver cancer, a fast spreading disease that continues to affect many individuals throughout the USA annually, seems to have an ever-rising mortality rate [91]. It has been noted that the rate of liver cancer-related mortality is chronically transcending other cancers [92]. Hepatocellular Carcinoma (HCC), as briefly mentioned, is the main subtype of liver cancer responsible for many deaths in the USA [93]. Studies have shown that MUC13 is significantly overexpressed in HCC tissues [69]. The overexpression of MUC13 is significantly involved in various assays such as tumor growth, vascular invasion, and encapsulation. Furthermore, MUC13 also plays a pivotal role in oncogenic activity thus encouraging cell proliferation, migration, and invasion in HCC tissues [69]. While it has been suggested that MUC13 overexpression is a major factor in the onset and spread of hepatocellular carcinoma (HCC), its exact function in this process is still unknown [69]. Another study indicates that abnormal hypermethylation-induced downregulation of miR-212-3p leads to an increase in the expression of MUC13 in intrahepatic cholangiocarcinoma (iCCA). This upregulation in MUC13 activates the EGFR/PI3K/AKT signaling pathway, thereby promoting metastasis [68]. Nevertheless, an adequate amount of research is needed to better apprehend the role of MUC13 in the pathological processes of liver cancer.
4.5 MUC13 in ovarian cancer
Ovarian cancer, specifically epithelial ovarian cancer, is another type of cancer associated with poor diagnosis, with 5-year survival rates of less than 45%, making it the eighth most prevalent cancer-related mortality and seventh-most frequent cancer in women [94]. To challenge this, studies are striving to find methods in which to identify ovarian cancers in their beginning stages. As such, the identification of a biomarker would aid in achieving this goal. MUC13, as explored with multiple other cancers, has been identified to show a positively correlated trend with the expression of ovarian cancers [95]. To explore this, our team produced clone PPZ0020, a monoclonal antibody, to verify the general expression and chemical characteristics of MUC13 in ovarian cancer. Utilizing tissue samples of ovarian cancer, epithelial ovarian cancer, benign tumor growths, and normally functioning tissue, the study found the expression of MUC13 to be notably greater in the tissues affected with cancer [54]. Furthermore, the expression of MUC13 was especially prevalent in epithelial ovarian cancer, further supporting the initial notion that MUC13 shares a positive trend with ovarian cancer. Interestingly, the addition of MUC13 into SK-OV-3, a void ovarian cancer cell line, initiated molecular characteristic changes, such as the separation of cells through a depletion of cell–cell adhesion molecules. This resulting data suggests the functional effects of MUC13 in ovarian cancer and SK-OV-3 has great potential in being a successful indicator of ovarian cancer [54].
4.6 MUC13 in glioblastoma
A recent study exploring the role of MUC13 in glioblastoma, an extremely deadly cancer with very poor prognosis, supports the notion that the overexpression of MUC13 relates to cancerous activity. More specifically, it was observed that its overexpression resulted in the increase in glioblastoma stem cell migratory capabilities. As a result, the metastasis of glioblastoma was greatly magnified. Additionally, it was noted that the expression of MUC13 heightened the phosphorylation of protein kinase B and P38 mitogen-activated protein (MAP) kinase. This idea was further supported by the censorship of MUC13 expression in the system, which resulted in the suppression of protein kinase B and P38 MAP kinase phosphorylation. The levels of MUC13 expression were regulated through the presence and absence of upstream stimulatory factor 1 (USF1), which binds to the MUC13 promoter region. The relationship between MUC13 and USF1 was noted to be positively correlated, which explains the increased expression of MUC13 upon the addition of USF1. Overall, this study further adds to and supports the novel concept that MUC13 expression is directly related to cancerous activity [77].
4.7 MUC13 in renal cell carcinoma
A recent study perceiving the role of MUC13 in renal cell carcinoma (RCC), a cancer that affects many individuals annually with poor prognosis and chemoresistance, proclaims that MUC13 plays a potential role in therapeutics and prognosis of RCC. Although early presentation of this disease is unlikely, due to late presentation and screenings, activation of MUC13 may have a potential role in the detection and prognosis of RCC and clear cell RCC (ccRCC)—a main subtype that affects more than 75% of the population annually. Through clinical trials, they observed that high expressions of MUC13 are related to worse overall survival than lower MUC13 expressions [71]. Furthermore, high expressions of MUC13 are present in stage 1 tumors with even more expression in stages 2–4 tumors; however, stage 1 patients who express more MUC13 show a considerably lower survival rate than those who express less MUC13. Overall, they concluded that lower survival rates are associated with upregulation of MUC13. Patients with high MUC13 expression had a 5-year survival rate of 51%, while those with low MUC13 expression had an overall survival rate of 86% [71].
4.8 MUC13 in lung cancer
Lung cancer is a highly aggressive malignancy and the leading cause of cancer-related deaths worldwide [96, 97]. In a study conducted by Yao Pang et al. in 2021 [70], the expression levels of MUC13 were investigated in lung tumor tissues and adjacent normal tissues. The researchers utilized RT-qPCR to analyze the expression levels of MUC13, and reported that MUC13 levels were significantly higher in the lung tumor tissues compared to the adjacent normal tissues. Further examination through Western blotting confirmed that the protein expression of MUC13 was also increased in the lung tumor tissues compared to the adjacent normal tissues. To validate these findings, MUC13 expression was assessed in 526 cancer samples and 59 normal samples obtained from patients with lung adenocarcinoma using datasets. Immunohistochemistry (IHC) analysis revealed an upregulation of MUC13 expression in the lung tumor tissues compared to the adjacent normal tissues. These findings suggest that MUC13 expression levels are abnormally elevated in lung cancer.
5 Unraveling influence of MUC13 on drug/chemo-resistance: a barrier to effective cancer treatment
Chemotherapy resistance occurs when cancer cells can endure and even thrive despite of chemotherapy exposure. Tumor-specific features in each patient influence this failure, ultimately dictating resistance and cancer growth in unanticipated. Chemotherapeutic drug resistance is the leading cause of relapses and low survival rates in cancer patients, despite recent breakthroughs in cancer therapy. Many of the drugs that are used to treat the severe forms of cancer have been demonstrated to have developed resistant to therapy. Resistance of metastasis cancer to chemotherapeutic compound is responsible for 90% of all failed cancer treatments, making it a significant barrier to cancer care [5]. For instance, it has been proven that breast cancer re-occurrence is driven by resistance to treatment with doxorubicin, paclitaxel, 5-fluorouracil, cyclophosphamide, and carboplatin, which is linked with a poor outcome and shortened survival time [6,7,8,9,10,11,12,13].
5.1 Intrinsic and acquired resistance
Multiple clinically relevant factors and their mechanisms of action have been described (Fig. 2). Intrinsic resistance and acquired resistance are the two main types of resistance. When tumor cells already have a resistance to chemotherapy medications before a patient starts treatment, this is called intrinsic chemoresistance [98]. Numerous elements affect intrinsic chemoresistance. To begin, chemoresistance may be caused by a tumor’s innate genetic alterations. The KEAP1-NFE2L2 pathway is one such example; it controls redox and metabolic balance. Overexpression of NFE2L2 may come from mutations in these genes, which can cause resistance to chemotherapeutic treatments. Lung, breast, bladder, ovarian, and liver cancers are only some of the many forms of malignancy that have been linked to these alterations [99,100,101,102]. Also, contributing to intrinsic chemoresistance is the fact that tumor cell populations tend to be rather diverse. Cancer stem cells (CSCs) that make up a tiny fraction of cancer cells are notoriously difficult to kill with chemotherapy. Recurrence of cancer and poor clinical outcomes may result from the enrichment of these CSCs after therapy [103,104,105,106,107]. Thirdly, chemoresistance might be facilitated by the stimulation of intrinsic mechanisms in tumor cells [108]. Multiple oncogenic signaling pathways, including mitogen-activated protein kinase (MAPK), hedgehog, PI3K/Akt, nuclear factor kappa B (NFkB), and notch, have been linked to the development of treatment resistance in malignant tumors. Angiogenesis, cell proliferation, drug distribution, apoptosis, and survival are all impacted by their activation [109]. Finally, pharmacological factors might play a part in the emergence of chemoresistant conditions. Chemotherapy efficacy may be diminished by a number of factors [110]. These include inadequate amounts of drugs at the tumor location, changed target molucules, and modifications to drug absorption, distribution, metabolism, or excretion. Drug resistance may develop when certain drug transporters, such as MRP1 and LRP, actively remove medicines from tumor cells, resulting in reduced intracellular drug concentrations [111]. In order to optimize treatment options and reduce hazardous side effects, knowing the causes of intrinsic chemoresistance is crucial for early detection of drug resistance. Personalized therapy techniques may enhance patient outcomes by identifying the individual resistance mechanisms in a patient’s tumor cells [112].
Acquired chemoresistance refers to the development of resistance to chemotherapy drugs after treatment has been initiated. This type of resistance is characterized via emergence of treatment resistant cell populations and a reduction in the effectiveness of the anticancer therapy. Several mechanisms contribute to acquired chemoresistance. One mechanism is the mutation of drug targets. Changes in the genes encoding the targets of chemotherapy drugs can render them less susceptible to the drugs, reducing their effectiveness. Proto-oncogene stimulation is another process which may result in abnormal growth and proliferation of cells. When these genes are turned on, it results in the emergence of chemotherapy drug resistance [4, 14]. Alterations in the microenvironment of the tumor may also contribute to acquired chemoresistance. The microenvironment can provide a protective niche for cancer cells, allowing them to evade the effects of chemotherapy drugs. Furthermore, histone modification, DNA methylation and miRNA expression, epigenetic modifications can contribute to acquired chemoresistance. These changes can affect the activity of genes associated with drug response, either by activating oncogenes or inhibiting tumor suppressor genes [113]. Chemoresistance can also result through modifications to DNA repair processes, disruptions to apoptosis, and changes in the cell cycle and its checkpoints [114]. Acquired chemoresistance is also linked to non-coding RNAs, particularly lncRNAs and miRNAs. Dysregulation of these RNA molecules may impact the synthesis of proteins involved in drug resistance [114,115,116,117,118,119,120].
5.2 Significance of MUC13 in key chemoresistance signaling pathways
CSCs’ carcinogenic characteristics necessitate several developmental pathways that were previously connected to the regulating of normal stem cells [121]. Among these, EMT and CSCs exhibit dysregulation of the WNT pathway [122]. Although colorectal malignancies were the first to show mutations in WNT signaling molecules such as β-catenin APC, AXIN, and WNT ligands, but these changes have been discovered in other varieties of solid tumors [123,124,125,126,127,128,129] and hematologic cancers, like multiple myeloma and leukemia [130,131,132]. To defend themselves from and resist chemotherapy treatments, cancer stem cells evolve drug resistance mechanisms. [121, 133]. In addition to the ways of understanding tumor heterogeneity, two distinct approaches accounted for the development of cancer chemotherapy resistance. Stem cells, including CSC, demonstrate high levels of ABCB1 and ABCG2 expression, the two major MDR genes [133]. Moreover, the human ABCB1 gene promoter has many TCF4/LEF binding domains, rendering it a targeted gene for the β-catenin and TCF4 transcriptional factors. The WNT/β-catenin circuit is linked to chemoresistance, as evidenced by the fact that β-catenin stimulation elevates ABCB1 expression.
In addition, a number of studies have shown that there is a relationship between this and the development of a multidrug resistant phenotype in tumor cells [134, 135]. This protein, along with other members of the family, has a physiological function in the blood–brain barrier via transporting substances through the placenta, gut, and another sites [136]. Additionally, these kinds of transporters effectively efflux xenobiotics to protect the cells from harmful compounds [133, 137]. Several solid tumors and hematological cancers develop chemoresistance due to their ability to expel drugs from cancer cell [122, 138, 139]. ABCB1 enables the development of multi drug resistance character because it may entangle and expel wide variety of medicines, hence causing treatment futility and tumor recurrence. Additionally, these proteins promote cell growth, invasion, and dysregulation of the pathways involved in apoptosis or complement-mediated cell toxicity [138,139,140,141,142,143] as well as transport ABC proteins to intracellular or extracellular regions to enhance product retention to enhance drug retention [144, 145].
Accumulating studies suggest that MUC13 expression has been linked to tumor differentiation. For example, MUC13 is especially upregulated in pancreatic neoplasms, in comparison to undetectable or poor expression in the normal pancreas [52]. In a similar vein, comparable expression, and localization trends of MUC13 were also seen in malignant and non-malignant colon cancer samples [65]. MUC13’s cytoplasmic/nuclear localization may contribute to colon cancer’s growth, progression, and spread to distant sites. MUC13 expression was shown to vary between the linked colorectal cancer cells SW620 and SW480. Two types of cells were taken from a diabetic patient: non-metastatic SW480 cells and metastasized SW620 cells from the patient’s lymph node metastatic lesion 1 year later [146]. This strongly suggests that MUC13 may contribute to the colon cancer growth. These findings are consistent with our lab reports demonstrating that MUC13 is expressed more often and is localized abnormally in colon cancer tissues [65].
Our laboratory has demonstrated the molecular signaling pathways that control MUC13 regulation, such as the JAK2/STAT5 pathway [55] (Fig. 3). In renal cancer, high expression of MUC13 was shown to have a positive correlation with increased Fuhrman grade, a pathological grading of renal cell cancers, and with reduced overall survival. Following surgery, increased MUC13 activation was identified as a novel distinct risk factor for the clinical prognosis of non-metastatic clear cell renal cell carcinoma [147]. Additionally, silencing of MUC13 in combination with multi-kinase inhibitors, sorafenib, and sunitinib has showed enhanced sensitization of renal carcinoma cells to respond to the therapy [71], thus indicating the function of MUC13 in chemosensitivity. Moreover, research conducted in our laboratory has proven that upregulation of MUC13 is related with increased activity of HER2 in pancreatic cancer [52]. This was elucidated further in our follow-up investigation, which indicated that MUC13 functions as a ligand for HER2 and activates key oncogenic signaling pathways in pancreatic cancer [57]. In a different investigation, our lab identified that miR-145 plays a tumor suppressor role in a tumor-promoting network that also contains MUC13 [148]. Therefore, our lab further developed a novel formulation of miR-145 that restores miR-145 levels in cells and inhibits MUC13 [149]. These studies affirm that MUC13 is a very important oncogenic target in pancreatic cancer, the therapeutic strategies against which warrants suppressing tumorigenic and metastatic phenotypes of pancreatic cancer cells. MUC13 overexpression has been also observed in hepatocellular carcinoma and associated with poor outcomes. An interesting study shows that c-myc and cyclin D, the downstream effector of Wnt/β-catenin signaling cascade, were found positively correlated with MUC13 levels. High β-catenin nuclear transition was found correlating with enhanced MUC13 overexpression, while the regulatory downstream targets of β-catenin decreased following MUC13 silencing/inhibition [69]. In addition, we and other researchers have defined how MUC13 works in ovarian cancer. Expression and functional studies conducted by our laboratory on MUC13 in ovarian cancer have demonstrated its direct correlation with this disease. Moreover, we found that its overexpression significantly impacts cellular characteristics [54]. All the studies discussed above are consistent with other studies that suggest the vital aspect of MUC13 upregulation in tumors development, including ovary [54, 150], gastric [73, 151], liver [69], colon [55], and pancreatic cancer [56, 57, 149, 152].
6 Implications of MUC13 in cancer stem cell plasticity in maintaining tumor heterogeneity
Cancer stem cells, also known as CSCs, play an important part in the development of intrinsic chemoresistance in a variety of cancer types. These cells make up a small subset of the cancer cells that have undergone terminal differentiation. Following therapeutic treatment, this particular subset of cells becomes predominant in a wide variety of malignancies. For instance, breast cancer stem cells exhibit characteristics such as quiescence, increased DNA repair, overexpression of drug efflux transporters, self-renewal and differentiation, and immune system evasion. These characteristics facilitate cancer invasion, metastasis, and recurrence in patients following chemotherapy, which ultimately accounts for the unfavorable clinical results, increased mortality, and chemoresistance in patients with breast cancer [106, 107]. Cell plasticity describes how a cell can be reprogrammed to develop an alternative fate in response to internal or external influences [153, 154]. Stem cells’ plasticity allows them to self-renew and transform into a variety of cell lineages. The ability of terminally differentiated cells like fibroblasts to revert to pluripotency reveals that plasticity is not limited to stem cells [155, 156]. Cancer cells exhibit plasticity through dedifferentiation, trans-differentiation, and EMT, when epithelial cells shift to a mesenchymal phenotype and lose features including cell–cell junctions and polarity [157] (Fig. 3). Plasticity and therapy resistance are two significant challenges in cancer treatment. They pose a major obstacle to successful treatment of cancer, especially in cases when it has spread (metastatic setting). These challenges are accountable for 90% of treatment failures [5]. There are two basic types of drug resistance: inherent, which occurs even before a drug is given to the patient, and acquired, which develops during therapy. Therapeutic treatments are assumed to only eliminate cancer cells that are actively proliferating, but CSCs stay impotent and persist. Resistance to therapy, tumorigenesis, tumor heterogeneity, tumor recurrence, and metastasis have all been linked to cancer stem cells (CSCs) [158, 159]. The mechanisms that allow CSCs to elude chemotherapy are not fully understood currently. This has been revealed that while most chemotherapy drugs destroy the majority of tumor cells, CSCs typically persist and develop following chemotherapy [20]. For instance, treatment of temozolomide enhances glioma stem cells in patient-derived cell lines but kills differentiated GBM cells [160]. Studies have shown that MUC13 expression enhances the CD133 + /CD44 + self-renewing CSC populations. It is notable that CSC populations have a vital impact in inducing chemoresistance and tumor recurrence [161]. Of note, MUC13 knockdown decreased the CD133 + /CD44 + population’s relative abundance and, more specifically, the enrichment of CSC that occurs after 5-FU therapy. Difference in CSC survival between MUC13-expressing and MUC13-deficient cells is paving a new way for combination therapy for involving the MUC13 siRNA along with available chemotherapy [71]
7 Role of MUC13 and epithelial to mesenchymal transitions (EMT)
Trans differentiation processes known as epithelial to mesenchymal transitions (EMT) is necessary for tissue morphogenesis in the developing embryo. According to recent research, EMT can trigger the formation of CSCs by sending signals to tumor stromal components. During the growth of tumors, oncogenically altered cells have the ability to override these developmental pathways [162]. Importantly, EMT has the unique ability to cause reversal to a CSC-like phenotype [163, 164], demonstrating a connection between therapy resistance, CSCs, and EMT. EMT is a developmental process that has been conserved throughout the process of evolution [165]. Through the process of EMT induction, pathophysiological events like tissue injury or cancer can cause differentiated cells to adopt on the characteristics of multipotent stem cells. This could reflect developmentally controlled EMT signaling pathways, which promote normal and CSC formation and regeneration [166,167,168]. Based on particular studies, cancer cells that have undergone EMT and have spread to distant sites may also have the characteristics of a CSC [169]. It is interesting to note that CD44 is a promising candidate for both β-catenin and TCF-4, which lends support to the notion that the EMT-associated Wnt pathway linked in the upkeep of CSCs [170]. Mesenchymal-like cells with the CD133 + chemokine receptor predominate in the intrusive front of pancreatic cancers, which may ready them for metastatic dissemination [171]. There are reports suggesting that alterations in cell adhesion properties caused by MUC13 can promote the invasion and spread of cancer [52, 54]. The extracellular tandem repeat domain of MUC13 is highly glycosylated and contribute to anti-adhesiveness property of cells [34, 51]. Notably, MUC13 overexpression in pancreatic carcinoma reduced cell–cell and cell–matrix adhesion.
Our findings provide valuable information about how MUC13 influences the growth of pancreatic cancer by modulation of PAK1 and S100A4. PAK1 upregulation promotes cell migration and controls the actin cytoskeleton during motility in HCC, breast, and colon cancer cells [172, 173]. Of note, S100A4 has also been linked to metastasis and invasion [174, 175]. Similarly, MUC13 overexpression in ovarian cancer decreased cell–cell interaction, increased cell migration, and facilitated F-actin reconfiguration. In order to migrate, a cell must form cell membrane projections with actin filaments and a continuous phase of actin polymerization near its prominent surfaces [176]. MUC13 localization in the basal area and along the basement membrane in ovarian cancer cells has been confirmed to promote ovarian cancer cell detachment from the primary location, as well as tumor cell invasion into ovarian stromal tissue [54]. These results indicate a clear correlation between MUC13 and enhanced cellular motility.
8 Role of MUC13 in tumor cell metabolism
Accumulating studies suggest that the impact of Warburg effect, occurs when cancer cells increase their aerobic glycolysis, establishes an acidic micro-environment that encourages invasion of tumor cells in the surroundings area [177]. Furthermore, the Warburg effect is primarily influenced by glucose, and Glut-1 increases the cellular glucose transport essential to drive anaerobic metabolism in developing tumor cells. Enhanced lactate synthesis, on the other hand, increases tumorigenic characteristics including cell invasion, proliferation, which metastasis, and is linked to tumor recurrence [178]. Importantly, cancer cells develop an oncogene addiction, making them extremely reliant on an oncogene’s action for survival and proliferation [179]. We have reported MUC13’s novel function in pancreatic cancer metabolic reprogramming [60]. We demonstrated that MUC13 alters cancer cells’ metabolism and the fundamental molecular pathways that underlie the tumorigenic features. Additionally, we discovered for the first time a substantial relationship between MUC13 and Glut-1 in patient tissue samples (Figs. 3 and 4). This indicates that MUC13 plays an important role in the tumor microenvironment, which promotes the growth of a more violent and invasive phenotype. Though more research is needed, we anticipate that a MUC13 cytoplasmic domain interacts to Glut-1 since the MUC13 cytoplasmic domain is believed to have a function in cell signaling. Low survival rates and poor outcomes are linked to hypoxia in the tumor environment, which typically occurs in later stage of cancer [180, 181]. A growing body of research has revealed that hypoxia can trigger the invasion, metastasis, and epithelial-mesenchymal transition of cancer cells, hence fostering stem-like properties in these cells [182, 183]. HIF-1 is essential for regulating CSCs through controlling the transcription of specific genes. It also contributes to tumor growth, evasion of the immune system, metabolic reprogramming, and resistance to drugs [181, 183, 184]. According to studies, HIF-1 has been linked with the development of CSCs. It has been reported that the production of various stem cell markers, such as Krüppel-like factor 4, OCT4, SOX2, and NANOG, can promoted by HIF-1 [185,186,187]. Additionally, by inhibiting the production of stem cell markers, HIF-1 silencing can slow the spread of cancer. Low survival rates and poor outcomes are linked to hypoxia in the tumor environment, which typically occurs in later stage of cancer [188]. It has been discovered that HIF-1 directly binds to the CD47 promoter in order to enhance gene transcription. This serves in avoiding macrophage phagocytosis and retaining the stem phenotype of breast CSCs [184, 189]. The promoter of CD24 is recruited by endogenous HIF-1, which enhances CD24 activity and also tumor development and metastasis [190]. HIF-1α interacts with the CD133 promoter, activating CD133 + glioma, colorectal, and pancreas CSCs through OCT4 and SOX2 [188, 191,192,193,194]. Angiogenesis, tumorigenesis, immunological response, cancer metastasis, and recurrence, as well as EMT progression and CSC formation, are all controlled by hypoxia [188, 189, 195]. Determining the methods via CSCs develop and sustain stemness may assist in overcoming the limitations of current cancer treatments.
9 Role of MUC13 and cell PI3K/Akt signaling pathways
The mTOR and PI3K/Akt pathways are essential for numerous physiologic and pathologic situations, such as, cell growth, differentiation, metabolism, angiogenesis, and survival [196]. In most cases of human cancer, the activation of mTOR is often regulated incorrectly. Among ovarian malignancies, for instance, the PI3K/Akt/mTOR pathway is active in around 70% of cases [197]. In addition, Tapia and colleagues [198] discovered that the PI3K/Akt/mTOR signaling is active in the tissues of individuals who had progressed gastric cancer, in contrast to the non-tumor mucosa [198]. Aside from the focus on cancer cells, an increasing number of recent studies have demonstrated the connections involving PI3K/Akt/mTOR signaling and CSCs [199, 200]. Number of studies showed the role of mTOR pathway in CSC maintenance. Prostate cancer radio resistance has been linked to EMT and increased cancer stem cells characteristics through the signaling pathway of PI3K/Akt/mTOR [201]. Induction of the mTOR pathway is necessary for colony-forming ability and tumor growth in breast cancer stem-like cells [202]. mTOR inhibition may diminish the activity of ALDH1, an indicator of colon cancer stem cells [203, 204]. When mTORC2 was inhibited, the activity of liver CSC marker decreased, and hepatocellular cancer stem cells demonstrated minimal or no ability to form tumors [205]. According to Sunayama et al., cancer stem-like cells’ ability to self-renew and proliferate is maintained through cross-inhibitory modulation between the MEK/ERK and PI3K/mTOR pathways in glioblastoma [206]. It has been discovered that in glioma tumor stem-like cells, Akt modulates the activity of ATP-binding cassette transporters (ABCG2) but does not control its downstream target mTOR [207]. The PI3K/Akt/mTOR pathway has a clear relationship with cancer stem cells, and its components are strong candidates for potential treatment. MUC13 interacts with cell-survival signaling pathways both actively and implicitly, leading to cell survival and proliferation. Ectopic expression of MUC13 was shown to have a greater capacity for colony formation and increased cellular proliferation, as well as a shorter doubling period of cells, as opposed to control cells. As a result, there were more cells in the S phase of the cell cycle. Furthermore, it was observed that increase in the proliferation was mediated by involvement of p38 MAPK pathways. Of note, ectopic expression of MUC13 leads to an upregulation of PAK1 and HER2, suggesting having a role in stabilization of EGFRs. Interestingly, it was reported that MUC13 expression also modulates the cell morphology as evident in ovarian cancer cells [54].
10 Conclusion
In conclusion, the study of MUC13 and its multifaceted functions in drug/chemo-resistance and cancer cell plasticity has offered valuable insights into the complex nature of cancer progression. It is evident that MUC13 overexpression disrupts normal cellular polarity, promoting the acquisition of malignant traits and facilitating metastasis through epithelial-mesenchymal transition (EMT). Additionally, MUC13 has been found to contribute to chemoresistance, presenting a significant challenge in cancer treatment. The unraveling of the molecular complexity surrounding MUC13 opens new opportunities for targeted therapies that address these mechanisms and offer improved outcomes for patients. By understanding the intricate biology of MUC13, researchers can develop strategies to restore cellular polarity, inhibit plasticity, and overcome chemoresistance. Further research is needed to explore this molecule’s potential as a therapeutic target, while considering its intricate network of interactions within the cancer microenvironment. Ultimately, unraveling the mysteries of MUC13’s role in cancer biology holds great promise for advancing our ability to combat cancer progression and enhance the efficacy of treatment strategies.
References
Sung, H., Ferlay, J., Siegel, R. L., Laversanne, M., Soerjomataram, I., Jemal, A., & Bray, F. (2021). Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians, 71, 209–249. https://doi.org/10.3322/caac.21660
Urruticoechea, A., Alemany, R., Balart, J., Villanueva, A., Viñals, F., & Capellá, G. (2010). Recent advances in cancer therapy: An overview. Current Pharmaceutical Design, 16, 3–10. https://doi.org/10.2174/138161210789941847
Baskar, R., Lee, K. A., Yeo, R., & Yeoh, K. W. (2012). Cancer and radiation therapy: Current advances and future directions. International Journal of Medical Sciences, 9, 193–199. https://doi.org/10.7150/ijms.3635
Wang, X., Zhang, H., & Chen, X. (2019). Drug resistance and combating drug resistance in cancer. Cancer Drug Resist, 2, 141–160. https://doi.org/10.20517/cdr.2019.10
Longley, D. B., & Johnston, P. G. (2005). Molecular mechanisms of drug resistance. The Journal of Pathology, 205, 275–292. https://doi.org/10.1002/path.1706
Smith, L., Watson, M. B., O’Kane, S. L., Drew, P. J., Lind, M. J., & Cawkwell, L. (2006). The analysis of doxorubicin resistance in human breast cancer cells using antibody microarrays. Molecular Cancer Therapeutics, 5, 2115–2120. https://doi.org/10.1158/1535-7163.Mct-06-0190
Harris, L. N., Broadwater, G., Lin, N. U., Miron, A., Schnitt, S. J., Cowan, D., Lara, J., Bleiweiss, I., Berry, D., Ellis, M., et al. (2006). Molecular subtypes of breast cancer in relation to paclitaxel response and outcomes in women with metastatic disease: Results from CALGB 9342. Breast Cancer Research, 8, R66. https://doi.org/10.1186/bcr1622
Murray, S., Briasoulis, E., Linardou, H., Bafaloukos, D., & Papadimitriou, C. (2012). Taxane resistance in breast cancer: Mechanisms, predictive biomarkers and circumvention strategies. Cancer Treatment Reviews, 38, 890–903. https://doi.org/10.1016/j.ctrv.2012.02.011
Szakács, G., Paterson, J. K., Ludwig, J. A., Booth-Genthe, C., & Gottesman, M. M. (2006). Targeting multidrug resistance in cancer. Nature Reviews Drug Discovery, 5, 219–234. https://doi.org/10.1038/nrd1984
Vulsteke, C., Pfeil, A. M., Schwenkglenks, M., Pettengell, R., Szucs, T. D., Lambrechts, D., Peeters, M., van Dam, P., Dieudonné, A. S., Hatse, S., et al. (2014). Impact of genetic variability and treatment-related factors on outcome in early breast cancer patients receiving (neo-) adjuvant chemotherapy with 5-fluorouracil, epirubicin and cyclophosphamide, and docetaxel. Breast Cancer Research and Treatment, 147, 557–570. https://doi.org/10.1007/s10549-014-3105-5
Porkka, K., Blomqvist, C., Rissanen, P., Elomaa, I., & Pyrhönen, S. (1994). Salvage therapies in women who fail to respond to first-line treatment with fluorouracil, epirubicin, and cyclophosphamide for advanced breast cancer. Journal of Clinical Oncology, 12, 1639–1647. https://doi.org/10.1200/jco.1994.12.8.1639
Sládek, N. E., Kollander, R., Sreerama, L., & Kiang, D. T. (2002). Cellular levels of aldehyde dehydrogenases (ALDH1A1 and ALDH3A1) as predictors of therapeutic responses to cyclophosphamide-based chemotherapy of breast cancer: A retrospective study. Rational individualization of oxazaphosphorine-based cancer chemotherapeutic regimens. Cancer Chemotherapy and Pharmacology, 49, 309–321. https://doi.org/10.1007/s00280-001-0412-4
Galluzzi, L., Vitale, I., Michels, J., Brenner, C., Szabadkai, G., Harel-Bellan, A., Castedo, M., & Kroemer, G. (2014). Systems biology of cisplatin resistance: past, present and future. Cell Death Dis, 5, e1257. https://doi.org/10.1038/cddis.2013.428
Mansoori, B., Mohammadi, A., Davudian, S., Shirjang, S., & Baradaran, B. (2017). The different mechanisms of cancer drug resistance: A brief review. Advanced Pharmaceutical Bulletin, 7, 339–348. https://doi.org/10.15171/apb.2017.041
Chang, J. C. (2016). Cancer stem cells: Role in tumor growth, recurrence, metastasis, and treatment resistance. Medicine (Baltimore), 95, S20-s25. https://doi.org/10.1097/md.0000000000004766
Chen, K., Huang, Y. H., & Chen, J. L. (2013). Understanding and targeting cancer stem cells: Therapeutic implications and challenges. Acta Pharmacologica Sinica, 34, 732–740. https://doi.org/10.1038/aps.2013.27
Prasad, S., Ramachandran, S., Gupta, N., Kaushik, I., & Srivastava, S. K. (2020). Cancer cells stemness: A doorstep to targeted therapy. Biochimica et Biophysica Acta, Molecular Basis of Disease, 1866, 165424. https://doi.org/10.1016/j.bbadis.2019.02.019
Furth, J., Kahn, M. C., & Breedis, C. (1937). The transmission of leukemia of mice with a single cell. The American Journal of Cancer, 31, 276–282.
Dick, J. E. (2008). Stem cell concepts renew cancer research. Blood, The Journal of the American Society of Hematology, 112, 4793–4807.
Visvader, J. E., & Lindeman, G. J. (2008). Cancer stem cells in solid tumours: Accumulating evidence and unresolved questions. Nature Reviews Cancer, 8, 755–768.
Iwata, M. (1968). Study of serum lipids in infants. 1. Serum lipids in healthy infants. Nihon Shonika Gakkai Zasshi Acta Paediatrica Japonica, 72, 1075–1081.
Phi, L.T.H., Sari, I.N., Yang, Y.-G., Lee, S.-H., Jun, N., Kim, K.S., Lee, Y.K., Kwon, H.Y. (2018). Cancer stem cells (CSCs) in drug resistance and their therapeutic implications in cancer treatment. Stem Cells International, 2018, 5416923. https://doi.org/10.1155/2018/5416923
Shervington, A., & Lu, C. (2008). Expression of multidrug resistance genes in normal and cancer stem cells. Cancer Investigation, 26, 535–542.
Vinogradov, S., & Wei, X. (2012). Cancer stem cells and drug resistance: The potential of nanomedicine. Nanomedicine, 7, 597–615.
Rachagani, S., Torres, M. P., Moniaux, N., & Batra, S. K. (2009). Current status of mucins in the diagnosis and therapy of cancer. BioFactors, 35, 509–527. https://doi.org/10.1002/biof.64
Reynolds, I. S., Fichtner, M., McNamara, D. A., Kay, E. W., Prehn, J. H. M., & Burke, J. P. (2019). Mucin glycoproteins block apoptosis; promote invasion, proliferation, and migration; and cause chemoresistance through diverse pathways in epithelial cancers. Cancer and Metastasis Reviews, 38, 237–257. https://doi.org/10.1007/s10555-019-09781-w
Dhanisha, S. S., Guruvayoorappan, C., Drishya, S., & Abeesh, P. (2018). Mucins: Structural diversity, biosynthesis, its role in pathogenesis and as possible therapeutic targets. Critical Reviews in Oncology Hematology, 122, 98–122. https://doi.org/10.1016/j.critrevonc.2017.12.006
Brockhausen, I. (2003). Glycodynamics of mucin biosynthesis in gastrointestinal tumor cells. Advances in Experimental Medicine and Biology, 535, 163–188. https://doi.org/10.1007/978-1-4615-0065-0_11
Yonezawa, S., Higashi, M., Yamada, N., Yokoyama, S., Kitamoto, S., Kitajima, S., & Goto, M. (2011). Mucins in human neoplasms: Clinical pathology, gene expression and diagnostic application. Pathology International, 61, 697–716. https://doi.org/10.1111/j.1440-1827.2011.02734.x
Nath, S., & Mukherjee, P. (2014). MUC1: A multifaceted oncoprotein with a key role in cancer progression. Trends in Molecular Medicine, 20, 332–342. https://doi.org/10.1016/j.molmed.2014.02.007
Moniaux, N., Escande, F., Porchet, N., Aubert, J. P., & Batra, S. K. (2001). Structural organization and classification of the human mucin genes. Frontiers in Bioscience, 6, D1192-1206. https://doi.org/10.2741/moniaux
van Putten, J. P. M., & Strijbis, K. (2017). Transmembrane mucins: Signaling receptors at the intersection of inflammation and cancer. Journal of Innate Immunity, 9, 281–299. https://doi.org/10.1159/000453594
Chauhan, S. C., Singh, A. P., Ruiz, F., Johansson, S. L., Jain, M., Smith, L. M., Moniaux, N., & Batra, S. K. (2006). Aberrant expression of MUC4 in ovarian carcinoma: Diagnostic significance alone and in combination with MUC1 and MUC16 (CA125). Modern Pathology, 19, 1386–1394. https://doi.org/10.1038/modpathol.3800646
Hollingsworth, M. A., & Swanson, B. J. (2004). Mucins in cancer: Protection and control of the cell surface. Nature Reviews Cancer, 4, 45–60. https://doi.org/10.1038/nrc1251
Andrianifahanana, M., Moniaux, N., Schmied, B. M., Ringel, J., Friess, H., Hollingsworth, M. A., Büchler, M. W., Aubert, J. P., & Batra, S. K. (2001). Mucin (MUC) gene expression in human pancreatic adenocarcinoma and chronic pancreatitis: A potential role of MUC4 as a tumor marker of diagnostic significance. Clinical Cancer Research, 7, 4033–4040.
Balagué, C., Audié, J. P., Porchet, N., & Real, F. X. (1995). In situ hybridization shows distinct patterns of mucin gene expression in normal, benign, and malignant pancreas tissues. Gastroenterology, 109, 953–964. https://doi.org/10.1016/0016-5085(95)90406-9
Dong, Y., Walsh, M. D., Cummings, M. C., Wright, R. G., Khoo, S. K., Parsons, P. G., & McGuckin, M. A. (1997). Expression of MUC1 and MUC2 mucins in epithelial ovarian tumours. The Journal of Pathology, 183, 311–317. https://doi.org/10.1002/(sici)1096-9896(199711)183:3%3c311::Aid-path917%3e3.0.Co;2-2
Giuntoli, R. L., Rodriguez, G. C., Whitaker, R. S., Dodge, R., & Voynow, J. A. (1998). Mucin gene expression in ovarian cancers. Cancer Research, 58, 5546–5550.
Maher, D. M., Gupta, B. K., Nagata, S., Jaggi, M., & Chauhan, S. C. (2011). Mucin 13: Structure, function, and potential roles in cancer pathogenesis. Molecular Cancer Research, 9, 531–537. https://doi.org/10.1158/1541-7786.Mcr-10-0443
Jonckheere, N., Skrypek, N., & Van Seuningen, I. (2014). Mucins and tumor resistance to chemotherapeutic drugs. Biochimica et Biophysica Acta, 1846, 142–151. https://doi.org/10.1016/j.bbcan.2014.04.008
Lee, D.H., Choi, S., Park, Y., Jin, H.S. (2021). Mucin1 and Mucin16: Therapeutic targets for cancer therapy. Pharmaceuticals (Basel), 14. https://doi.org/10.3390/ph14101053.
Ponnusamy, M. P., Seshacharyulu, P., Lakshmanan, I., Vaz, A. P., Chugh, S., & Batra, S. K. (2013). Emerging role of mucins in epithelial to mesenchymal transition. Current Cancer Drug Targets, 13, 945–956. https://doi.org/10.2174/15680096113136660100
Marimuthu, S., Rauth, S., Ganguly, K., Zhang, C., Lakshmanan, I., Batra, S. K., & Ponnusamy, M. P. (2021). Mucins reprogram stemness, metabolism and promote chemoresistance during cancer progression. Cancer and Metastasis Reviews, 40, 575–588. https://doi.org/10.1007/s10555-021-09959-1
Gendler, S. J., & Spicer, A. P. (1995). Epithelial mucin genes. Annual Review of Physiology, 57, 607–634. https://doi.org/10.1146/annurev.ph.57.030195.003135
Thornton, D. J., Rousseau, K., & McGuckin, M. A. (2008). Structure and function of the polymeric mucins in airways mucus. Annual Review of Physiology, 70, 459–486. https://doi.org/10.1146/annurev.physiol.70.113006.100702
Desseyn, J.-L., Gouyer, V., Tetaert, T. (2008). Architecture of the gel-forming mucins. The Epithelial Mucins: Structure/Function. Roles in Cancer and Inflammatory Diseases, Isabelle Van Seuningen/Research Signpost, 1–16, 978-81-308-0256-5. ⟨hal-02340996⟩.
Hattrup, C. L., & Gendler, S. J. (2008). Structure and function of the cell surface (tethered) mucins. Annual Review of Physiology, 70, 431–457. https://doi.org/10.1146/annurev.physiol.70.113006.100659
Jonckheere, N., Skrypek, N., Frénois, F., & Van Seuningen, I. (2013). Membrane-bound mucin modular domains: From structure to function. Biochimie, 95, 1077–1086. https://doi.org/10.1016/j.biochi.2012.11.005
Jonckheere, N., & Van Seuningen, I. (2008). The membrane-bound mucins: How large O-glycoproteins play key roles in epithelial cancers and hold promise as biological tools for gene-based and immunotherapies. Critical Reviews in Oncogenesis, 14, 177–196. https://doi.org/10.1615/critrevoncog.v14.i2-3.30
Segui-Perez, C., Stapels, D.A.C., Ma, Z., Su, J., Passchier, E., Westendorp, B., Wu, W., Putten, J.P.M.V., Strijbis, K. (2022). MUC13 negatively regulates tight junction proteins and intestinal epithelial barrier integrity via Protein Kinase C. bioRxiv,. https://doi.org/10.1101/2022.10.27.51398
Williams, S. J., Wreschner, D. H., Tran, M., Eyre, H. J., Sutherland, G. R., & McGuckin, M. A. (2001). Muc13, a novel human cell surface mucin expressed by epithelial and hemopoietic cells. Journal of Biological Chemistry, 276, 18327–18336. https://doi.org/10.1074/jbc.M008850200
Chauhan, S. C., Ebeling, M. C., Maher, D. M., Koch, M. D., Watanabe, A., Aburatani, H., Lio, Y., & Jaggi, M. (2012). MUC13 mucin augments pancreatic tumorigenesis. Molecular Cancer Therapeutics, 11, 24–33. https://doi.org/10.1158/1535-7163.Mct-11-0598
Chauhan, S. C., Kumar, D., & Jaggi, M. (2009). Mucins in ovarian cancer diagnosis and therapy. Journal of Ovarian Research, 2, 21. https://doi.org/10.1186/1757-2215-2-21
Chauhan, S. C., Vannatta, K., Ebeling, M. C., Vinayek, N., Watanabe, A., Pandey, K. K., Bell, M. C., Koch, M. D., Aburatani, H., Lio, Y., et al. (2009). Expression and functions of transmembrane mucin MUC13 in ovarian cancer. Cancer Research, 69, 765–774. https://doi.org/10.1158/0008-5472.Can-08-0587
Gupta, B. K., Maher, D. M., Ebeling, M. C., Stephenson, P. D., Puumala, S. E., Koch, M. R., Aburatani, H., Jaggi, M., & Chauhan, S. C. (2014). Functions and regulation of MUC13 mucin in colon cancer cells. Journal of Gastroenterology, 49, 1378–1391. https://doi.org/10.1007/s00535-013-0885-z
Khan, S., Ebeling, M. C., Zaman, M. S., Sikander, M., Yallapu, M. M., Chauhan, N., Yacoubian, A. M., Behrman, S. W., Zafar, N., Kumar, D., et al. (2014). MicroRNA-145 targets MUC13 and suppresses growth and invasion of pancreatic cancer. Oncotarget, 5, 7599–7609. https://doi.org/10.18632/oncotarget.2281
Khan, S., Sikander, M., Ebeling, M. C., Ganju, A., Kumari, S., Yallapu, M. M., Hafeez, B. B., Ise, T., Nagata, S., Zafar, N., et al. (2017). MUC13 interaction with receptor tyrosine kinase HER2 drives pancreatic ductal adenocarcinoma progression. Oncogene, 36, 491–500. https://doi.org/10.1038/onc.2016.218
Filippou, P. S., Ren, A. H., Korbakis, D., Dimitrakopoulos, L., Soosaipillai, A., Barak, V., Frenkel, S., Pe’er, J., Lotem, M., Merims, S., et al. (2018). Exploring the potential of mucin 13 (MUC13) as a biomarker for carcinomas and other diseases. Clinical Chemistry and Laboratory Medicine, 56, 1945–1953. https://doi.org/10.1515/cclm-2018-0139
Khan, S., Zafar, N., Khan, S. S., Setua, S., Behrman, S. W., Stiles, Z. E., Yallapu, M. M., Sahay, P., Ghimire, H., Ise, T., et al. (2018). Clinical significance of MUC13 in pancreatic ductal adenocarcinoma. HPB: The Official Journal of the International Hepato Pancreato Biliary Association, 20, 563–572. https://doi.org/10.1016/j.hpb.2017.12.003
Kumari, S., Khan, S., Gupta, S. C., Kashyap, V. K., Yallapu, M. M., Chauhan, S. C., & Jaggi, M. (2018). MUC13 contributes to rewiring of glucose metabolism in pancreatic cancer. Oncogenesis, 7, 19. https://doi.org/10.1038/s41389-018-0031-0
Massey, A. E., Doxtater, K. A., Yallapu, M. M., & Chauhan, S. C. (2020). Biophysical changes caused by altered MUC13 expression in pancreatic cancer cells. Micron, 130, 102822. https://doi.org/10.1016/j.micron.2019.102822
Nishii, Y., Yamaguchi, M., Kimura, Y., Hasegawa, T., Aburatani, H., Uchida, H., Hirata, K., & Sakuma, Y. (2015). A newly developed anti-Mucin 13 monoclonal antibody targets pancreatic ductal adenocarcinoma cells. International Journal of Oncology, 46, 1781–1787. https://doi.org/10.3892/ijo.2015.2880
Stiles, Z. E., Khan, S., Patton, K. T., Jaggi, M., Behrman, S. W., & Chauhan, S. C. (2019). Transmembrane mucin MUC13 distinguishes intraductal papillary mucinous neoplasms from non-mucinous cysts and is associated with high-risk lesions. HPB: The Official Journal of the International Hepato Pancreato Biliary Association, 21, 87–95. https://doi.org/10.1016/j.hpb.2018.07.009
Mito, K., Saito, M., Morita, K., Maetani, I., Sata, N., Mieno, M., & Fukushima, N. (2018). Clinicopathological and prognostic significance of MUC13 and AGR2 expression in intraductal papillary mucinous neoplasms of the pancreas. Pancreatology, 18, 407–412. https://doi.org/10.1016/j.pan.2018.04.003
Gupta, B. K., Maher, D. M., Ebeling, M. C., Sundram, V., Koch, M. D., Lynch, D. W., Bohlmeyer, T., Watanabe, A., Aburatani, H., Puumala, S. E., et al. (2012). Increased expression and aberrant localization of mucin 13 in metastatic colon cancer. Journal of Histochemistry and Cytochemistry, 60, 822–831. https://doi.org/10.1369/0022155412460678
Walsh, M. D., Young, J. P., Leggett, B. A., Williams, S. H., Jass, J. R., & McGuckin, M. A. (2007). The MUC13 cell surface mucin is highly expressed by human colorectal carcinomas. Human Pathology, 38, 883–892. https://doi.org/10.1016/j.humpath.2006.11.020
Kasprzak, A., Adamek, A. (2019). Mucins: The Old, the new and the promising factors in hepatobiliary carcinogenesis. International Journal of Molecular Sciences, 20. https://doi.org/10.3390/ijms20061288
Tiemin, P., Fanzheng, M., Peng, X., Jihua, H., Ruipeng, S., Yaliang, L., Yan, W., Junlin, X., Qingfu, L., Zhefeng, H., et al. (2020). MUC13 promotes intrahepatic cholangiocarcinoma progression via EGFR/PI3K/AKT pathways. Journal of Hepatology, 72, 761–773. https://doi.org/10.1016/j.jhep.2019.11.021
Dai, Y., Liu, L., Zeng, T., Liang, J. Z., Song, Y., Chen, K., Li, Y., Chen, L., Zhu, Y. H., Li, J., et al. (2018). Overexpression of MUC13, a poor prognostic predictor, promotes cell growth by activating wnt signaling in hepatocellular carcinoma. American Journal of Pathology, 188, 378–391. https://doi.org/10.1016/j.ajpath.2017.10.016
Pang, Y., Zhang, Y., Zhang, H. Y., Wang, W. H., Jin, G., Liu, J. W., & Zhu, Z. J. (2022). MUC13 promotes lung cancer development and progression by activating ERK signaling. Oncology Letters, 23, 37. https://doi.org/10.3892/ol.2021.13155
Sheng, Y., Ng, C. P., Lourie, R., Shah, E. T., He, Y., Wong, K. Y., Seim, I., Oancea, I., Morais, C., Jeffery, P. L., et al. (2017). MUC13 overexpression in renal cell carcinoma plays a central role in tumor progression and drug resistance. International Journal of Cancer, 140, 2351–2363. https://doi.org/10.1002/ijc.30651
Xu, Z., Liu, Y., Yang, Y., Wang, J., Zhang, G., Liu, Z., Fu, H., Wang, Z., Liu, H., & Xu, J. (2017). High expression of Mucin13 associates with grimmer postoperative prognosis of patients with non-metastatic clear-cell renal cell carcinoma. Oncotarget, 8, 7548–7558. https://doi.org/10.18632/oncotarget.13692
Shimamura, T., Ito, H., Shibahara, J., Watanabe, A., Hippo, Y., Taniguchi, H., Chen, Y., Kashima, T., Ohtomo, T., Tanioka, F., et al. (2005). Overexpression of MUC13 is associated with intestinal-type gastric cancer. Cancer Science, 96, 265–273. https://doi.org/10.1111/j.1349-7006.2005.00043.x
He, L., Qu, L., Wei, L., Chen, Y., & Suo, J. (2017). Reduction of miR-132-3p contributes to gastric cancer proliferation by targeting MUC13. Molecular Medicine Reports, 15, 3055–3061. https://doi.org/10.3892/mmr.2017.6347
Zhao, Z. T., Li, Y., Yuan, H. Y., Ma, F. H., Song, Y. M., & Tian, Y. T. (2020). Identification of key genes and pathways in gastric signet ring cell carcinoma based on transcriptome analysis. World J Clin Cases, 8, 658–669. https://doi.org/10.12998/wjcc.v8.i4.658
Wang, H., Shen, L., Lin, Y., Shi, Q., Yang, Y., & Chen, K. (2015). The expression and prognostic significance of Mucin 13 and Mucin 20 in esophageal squamous cell carcinoma. Journal of Cancer Research and Therapeutics, 11(Suppl 1), C74-79. https://doi.org/10.4103/0973-1482.163846
Li, P., Wang, H., Hou, M., Li, D., & Bai, H. (2017). Upstream stimulating factor1 (USF1) enhances the proliferation of glioblastoma stem cells mainly by activating the transcription of mucin13 (MUC13). Die Pharmazie, 72, 98–102. https://doi.org/10.1691/ph.2017.6788
Sung, H. Y., Park, A. K., Ju, W., & Ahn, J. H. (2014). Overexpression of mucin 13 due to promoter methylation promotes aggressive behavior in ovarian cancer cells. Yonsei Medical Journal, 55, 1206–1213. https://doi.org/10.3349/ymj.2014.55.5.1206
Dhasmana, A., Dhasmana, S., Agarwal, S., Khan, S., Haque, S., Jaggi, M., Yallapu, M. M., & Chauhan, S. C. (2023). Integrative big transcriptomics data analysis implicates crucial role of MUC13 in pancreatic cancer. Computational and Structural Biotechnology Journal, 21, 2845–2857. https://doi.org/10.1016/j.csbj.2023.04.029
Jonckheere, N., Vincent, A., Neve, B., & Van Seuningen, I. (2021). Mucin expression, epigenetic regulation and patient survival: A toolkit of prognostic biomarkers in epithelial cancers. Biochimica et Biophysica Acta - Reviews on Cancer, 1876, 188538. https://doi.org/10.1016/j.bbcan.2021.188538
Thompson, C. M., Cannon, A., West, S., Ghersi, D., Atri, P., Bhatia, R., Smith, L., Rachagani, S., Wichman, C., Kumar, S., et al. (2021). Mucin expression and splicing determine novel subtypes and patient mortality in pancreatic ductal adenocarcinoma. Clinical Cancer Research, 27, 6787–6799. https://doi.org/10.1158/1078-0432.Ccr-21-1591
Fabregas, J. C., Ramnaraign, B., & George, T. J. (2022). Clinical updates for colon cancer care in 2022. Clinical Colorectal Cancer, 21, 198–203. https://doi.org/10.1016/j.clcc.2022.05.006
Byrd, J. C., & Bresalier, R. S. (2004). Mucins and mucin binding proteins in colorectal cancer. Cancer and Metastasis Reviews, 23, 77–99. https://doi.org/10.1023/a:1025815113599
Jonckheere, N., & Van Seuningen, I. (2010). The membrane-bound mucins: From cell signalling to transcriptional regulation and expression in epithelial cancers. Biochimie, 92, 1–11. https://doi.org/10.1016/j.biochi.2009.09.018
Cox, K.E., Liu, S., Lwin, T.M., Hoffman, R.M., Batra, S.K., Bouvet, M. (2023). The mucin family of proteins: Candidates as potential biomarkers for colon cancer. Cancers (Basel), 15. https://doi.org/10.3390/cancers15051491.
Sheng, Y. H., He, Y., Hasnain, S. Z., Wang, R., Tong, H., Clarke, D. T., Lourie, R., Oancea, I., Wong, K. Y., Lumley, J. W., et al. (2017). MUC13 protects colorectal cancer cells from death by activating the NF-κB pathway and is a potential therapeutic target. Oncogene, 36, 700–713. https://doi.org/10.1038/onc.2016.241
Sheng, Y. H., Wong, K. Y., Seim, I., Wang, R., He, Y., Wu, A., Patrick, M., Lourie, R., Schreiber, V., Giri, R., et al. (2019). MUC13 promotes the development of colitis-associated colorectal tumors via β-catenin activity. Oncogene, 38, 7294–7310. https://doi.org/10.1038/s41388-019-0951-y
Doxtater, K., Tripathi, M. K., Sekhri, R., Hafeez, B. B., Khan, S., Zafar, N., Behrman, S. W., Yallapu, M. M., Jaggi, M., & Chauhan, S. C. (2023). MUC13 drives cancer aggressiveness and metastasis through the YAP1-dependent pathway. Life Sci Alliance, 6(12). https://doi.org/10.26508/lsa.202301975
Lei, Z. N., Teng, Q. X., Tian, Q., Chen, W., Xie, Y., Wu, K., Zeng, Q., Zeng, L., Pan, Y., Chen, Z. S., et al. (2022). Signaling pathways and therapeutic interventions in gastric cancer. Signal Transduction and Targeted Therapy, 7, 358. https://doi.org/10.1038/s41392-022-01190-w
Stomach Cancer Survival Rates. The American Cancer Society medical and editorial content team 2023. https://www.cancer.org/cancer/types/stomach-cancer/detection-diagnosis-staging/survival-rates.html
Hepatocellular Carcinoma (HCC). Cleveland Clinic medical professional 2021. https://my.clevelandclinic.org/health/diseases/21709-hepatocellular-carcinoma-hcc
Toh, M. R., Wong, E. Y. T., Wong, S. H., Ng, A. W. T., Loo, L. H., Chow, P. K., & Ngeow, J. (2023). Global Epidemiology and genetics of hepatocellular carcinoma. Gastroenterology, 164, 766–782. https://doi.org/10.1053/j.gastro.2023.01.033
Ganesan, P., & Kulik, L. M. (2023). Hepatocellular carcinoma: New developments. Clinics in Liver Disease, 27, 85–102. https://doi.org/10.1016/j.cld.2022.08.004
Webb, P. M., & Jordan, S. J. (2017). Epidemiology of epithelial ovarian cancer. Best Practice & Research Clinical Obstetrics & Gynaecology, 41, 3–14. https://doi.org/10.1016/j.bpobgyn.2016.08.006
Ren, A. H., Filippou, P. S., Soosaipillai, A., Dimitrakopoulos, L., Korbakis, D., Leung, F., Kulasingam, V., Bernardini, M. Q., & Diamandis, E. P. (2023). Mucin 13 (MUC13) as a candidate biomarker for ovarian cancer detection: Potential to complement CA125 in detecting non-serous subtypes. Clinical Chemistry and Laboratory Medicine, 61, 464–472. https://doi.org/10.1515/cclm-2022-0491
Bade, B. C., & Dela Cruz, C. S. (2020). Lung cancer 2020: Epidemiology, etiology, and prevention. Clinics in Chest Medicine, 41, 1–24. https://doi.org/10.1016/j.ccm.2019.10.001
Yang, D., Liu, Y., Bai, C., Wang, X., & Powell, C. A. (2020). Epidemiology of lung cancer and lung cancer screening programs in China and the United States. Cancer Letters, 468, 82–87. https://doi.org/10.1016/j.canlet.2019.10.009
Vaidya, F. U., Sufiyan Chhipa, A., Mishra, V., Gupta, V. K., Rawat, S. G., Kumar, A., & Pathak, C. (2022). Molecular and cellular paradigms of multidrug resistance in cancer. Cancer Rep (Hoboken), 5, e1291. https://doi.org/10.1002/cnr2.1291
Barrera-Rodríguez, R. (2018). Importance of the Keap1-Nrf2 pathway in NSCLC: Is it a possible biomarker? Biomedical Reports, 9, 375–382. https://doi.org/10.3892/br.2018.1143
Hammerman, P. S., Lawrence, M. S., Voet, D., Jing, R., Cibulskis, K., Sivachenko, A., Stojanov, P., McKenna, A., Lander, E. S., Gabriel, S., Getz, G., Sougnez, C., Imielinski, M., Helman, E., Hernandez, B., Pho, N. H., Meyerson, M., Chu, A., Chun, H.-J. E., et al. Comprehensive genomic characterization of squamous cell lung cancers. PubMed. nih.gov
Giordano, T. J. (2014). The cancer genome atlas research network: A sight to behold. Endocrine Pathology, 25, 362–365. https://doi.org/10.1007/s12022-014-9345-4
Collisson, E. A., Campbell, J. D., Brooks, A. N., Berger, A. H., Lee, W., Chmielecki, J., Beer, D. G., Cope, L., Creighton, C. J., Danilova, L., Ding, L., Getz, G., Hammerman, P. S., Neil Hayes, D., Hernandez, B., Herman, J. G., Heymach, J. V., Jurisica, I., Kucherlapati, R., … John Flynn, H. (2014). Comprehensive molecular profiling of lung adenocarcinoma. Nature, 511(7511), 543–550. https://doi.org/10.1038/nature13385
Begicevic, R.R., Falasca, M. (2017). ABC transporters in cancer stem cells: Beyond chemoresistance. International Journal of Molecular Sciences, 18. https://doi.org/10.3390/ijms18112362.
Zhao, J. (2016). Cancer stem cells and chemoresistance: The smartest survives the raid. Pharmacology & Therapeutics, 160, 145–158. https://doi.org/10.1016/j.pharmthera.2016.02.008
Hasan, S., Taha, R., & Omri, H. E. (2018). Current opinions on chemoresistance: An overview. Bioinformation, 14, 80–85. https://doi.org/10.6026/97320630014080
Chen, W., Qin, Y., & Liu, S. (2018). Cytokines, breast cancer stem cells (BCSCs) and chemoresistance. Clinical and Translational Medicine, 7, 27. https://doi.org/10.1186/s40169-018-0205-6
Samuel, S.M., Varghese, E., Koklesová, L., Líšková, A., Kubatka, P., Büsselberg, D. (2020). Counteracting chemoresistance with metformin in breast cancers: Targeting cancer stem cells. Cancers (Basel), 12. https://doi.org/10.3390/cancers12092482.
Nussinov, R., Tsai, C. J., & Jang, H. (2017). A new view of pathway-driven drug resistance in tumor proliferation. Trends in Pharmacological Sciences, 38, 427–437. https://doi.org/10.1016/j.tips.2017.02.001
Rajabpour, A., Rajaei, F., & Teimoori-Toolabi, L. (2017). Molecular alterations contributing to pancreatic cancer chemoresistance. Pancreatology, 17, 310–320. https://doi.org/10.1016/j.pan.2016.12.013
Alfarouk, K. O., Stock, C. M., Taylor, S., Walsh, M., Muddathir, A. K., Verduzco, D., Bashir, A. H., Mohammed, O. Y., Elhassan, G. O., Harguindey, S., et al. (2015). Resistance to cancer chemotherapy: Failure in drug response from ADME to P-gp. Cancer Cell International, 15, 71. https://doi.org/10.1186/s12935-015-0221-1
Moulder, S. (2010). Intrinsic resistance to chemotherapy in breast cancer. Womens Health (Lond), 6, 821–830. https://doi.org/10.2217/whe.10.60
Lippert, T. H., Ruoff, H. J., & Volm, M. (2008). Intrinsic and acquired drug resistance in malignant tumors. The main reason for therapeutic failure. Arzneimittelforschung, 58, 261–264. https://doi.org/10.1055/s-0031-1296504
Schwarzenbach, H., & Gahan, P. B. (2019). Resistance to cis- and carboplatin initiated by epigenetic changes in ovarian cancer patients. Cancer Drug Resist, 2, 271–296. https://doi.org/10.20517/cdr.2019.010
Chen, Y., Song, Y., Mi, Y., Jin, H., Cao, J., Li, H., Han, L., Huang, T., Zhang, X., Ren, S., et al. (2020). microRNA-499a promotes the progression and chemoresistance of cervical cancer cells by targeting SOX6. Apoptosis, 25, 205–216. https://doi.org/10.1007/s10495-019-01588-y
Wang, H., Guan, Z., He, K., Qian, J., Cao, J., & Teng, L. (2017). LncRNA UCA1 in anti-cancer drug resistance. Oncotarget, 8, 64638–64650. https://doi.org/10.18632/oncotarget.18344
Ebrahimi Ghahnavieh, L., Tabatabaeian, H., Ebrahimi Ghahnavieh, Z., Honardoost, M. A., Azadeh, M., Moazeni Bistgani, M., & Ghaedi, K. (2020). Fluctuating expression of miR-584 in primary and high-grade gastric cancer. BMC Cancer, 20, 621. https://doi.org/10.1186/s12885-020-07116-5
Lim, S. K., Tabatabaeian, H., Lu, S. Y., Kang, S. A., Sundaram, G. M., Sampath, P., Chan, S. W., Hong, W. J., & Lim, Y. P. (2020). Hippo/MST blocks breast cancer by downregulating WBP2 oncogene expression via miRNA processor Dicer. Cell Death & Disease, 11, 669. https://doi.org/10.1038/s41419-020-02901-3
Adami, B., Tabatabaeian, H., Ghaedi, K., Talebi, A., Azadeh, M., & Dehdashtian, E. (2019). miR-146a is deregulated in gastric cancer. Journal of Cancer Research and Therapeutics, 15, 108–114. https://doi.org/10.4103/jcrt.JCRT_855_17
Chan, J. J., Tay, Y. (2018). Noncoding RNA:RNA regulatory networks in cancer. The International Journal of Molecular Science, 19. https://doi.org/10.3390/ijms19051310.
Anastasiadou, E., Jacob, L. S., & Slack, F. J. (2018). Non-coding RNA networks in cancer. Nature Reviews Cancer, 18, 5–18. https://doi.org/10.1038/nrc.2017.99
Ramos, E. K., Hoffmann, A. D., Gerson, S. L., & Liu, H. (2017). New opportunities and challenges to defeat cancer stem cells. Trends in cancer, 3, 780–796.
Singh, A., & Settleman, J. (2010). EMT, cancer stem cells and drug resistance: An emerging axis of evil in the war on cancer. Oncogene, 29, 4741–4751.
Flanagan, D. J., Barker, N., Costanzo, N. S. D., Mason, E. A., Gurney, A., Meniel, V. S., Koushyar, S., Austin, C. R., Ernst, M., & Pearson, H. B. (2019). Frizzled-7 is required for Wnt signaling in gastric tumors with and without Apc mutations. Cancer Research, 79, 970–981.
Zhang, Y., Morris, J. P., IV., Yan, W., Schofield, H. K., Gurney, A., Simeone, D. M., Millar, S. E., Hoey, T., Hebrok, M., & Pasca di Magliano, M. (2013). Canonical wnt signaling is required for pancreatic carcinogenesis. Cancer Research, 73, 4909–4922.
Sinnberg, T., Levesque, M. P., Krochmann, J., Cheng, P. F., Ikenberg, K., Meraz-Torres, F., Niessner, H., Garbe, C., & Busch, C. (2018). Wnt-signaling enhances neural crest migration of melanoma cells and induces an invasive phenotype. Molecular Cancer, 17, 1–19.
Liu, Y., Chang, Y., Lu, S., & Xiang, Y. Y. (2019). Downregulation of long noncoding RNA DGCR5 contributes to the proliferation, migration, and invasion of cervical cancer by activating Wnt signaling pathway. Journal of Cellular Physiology, 234, 11662–11669.
Veeck, J., Bektas, N., Hartmann, A., Kristiansen, G., Heindrichs, U., Knüchel, R., & Dahl, E. (2008). Wnt signalling in human breast cancer: Expression of the putative Wnt inhibitor Dickkopf-3 (DKK3) is frequently suppressed by promoter hypermethylation in mammary tumours. Breast Cancer Research, 10, 1–11.
Murillo-Garzón, V., & Kypta, R. (2017). WNT signalling in prostate cancer. Nature Reviews Urology, 14, 683–696.
McCord, M., Mukouyama, Y.-S., Gilbert, M. R., & Jackson, S. (2017). Targeting WNT signaling for multifaceted glioblastoma therapy. Frontiers in cellular neuroscience, 11, 318.
van Andel, H., Kocemba, K. A., Spaargaren, M., & Pals, S. T. (2019). Aberrant Wnt signaling in multiple myeloma: Molecular mechanisms and targeting options. Leukemia, 33, 1063–1075.
Ashihara, E., Takada, T., & Maekawa, T. (2015). Targeting the canonical Wnt/β-catenin pathway in hematological malignancies. Cancer Science, 106, 665–671.
Yadav, A. K., & Desai, N. S. (2019). Cancer stem cells: Acquisition, characteristics, therapeutic implications, targeting strategies and future prospects. Stem Cell Reviews and Reports, 15, 331–355.
Steinbichler, T. B., Dudás, J., Skvortsov, S., Ganswindt, U., Riechelmann, H., & Skvortsova, I. I. (2018). Therapy resistance mediated by cancer stem cells. Seminars in Cancer Biology, 53, 156–167. https://doi.org/10.1016/j.semcancer.2018.11.006
Jaiswal, R., Luk, F., Dalla, P. V., Grau, G. E. R., & Bebawy, M. (2013). Breast cancer-derived microparticles display tissue selectivity in the transfer of resistance proteins to cells. PLoS ONE, 8, e61515.
Vesel, M., Rapp, J., Feller, D., Kiss, E., Jaromi, L., Meggyes, M., Miskei, G., Duga, B., Smuk, G., & Laszlo, T. (2017). ABCB1 and ABCG2 drug transporters are differentially expressed in non-small cell lung cancers (NSCLC) and expression is modified by cisplatin treatment via altered Wnt signaling. Respiratory Research, 18, 1–11.
Schinkel, A. H., Smit, J. J., van Tellingen, O., Beijnen, J. H., Wagenaar, E., van Deemter, L., Mol, C. A., van der Valk, M. A., Robanus-Maandag, E. C., te Riele, H. P., et al. (1994). Disruption of the mouse mdr1a P-glycoprotein gene leads to a deficiency in the blood-brain barrier and to increased sensitivity to drugs. Cell, 77, 491–502. https://doi.org/10.1016/0092-8674(94)90212-7
Johnson, W. W. (2002). P-glycoprotein-mediated efflux as a major factor in the variance of absorption and distribution of drugs: Modulation of chemotherapy resistance. Methods and Findings in Experimental and Clinical Pharmacology, 24, 501–514. https://doi.org/10.1358/mf.2002.24.8.705071
Cerezo, D., Lencina, M., Ruiz-Alcaraz, A. J., Ferragut, J. A., Saceda, M., Sanchez, M., Cánovas, M., García-Peñarrubia, P., & Martín-Orozco, E. (2012). Acquisition of MDR phenotype by leukemic cells is associated with increased caspase-3 activity and a collateral sensitivity to cold stress. Journal of Cellular Biochemistry, 113, 1416–1425. https://doi.org/10.1002/jcb.24016
Cerezo, D., Ruiz-Alcaraz, A. J., Lencina-Guardiola, M., Cánovas, M., García-Peñarrubia, P., Martínez-López, I., & Martín-Orozco, E. (2017). Attenuated JNK signaling in multidrug-resistant leukemic cells Dual role of MAPK in cell survival. Cell Signal, 30, 162–170. https://doi.org/10.1016/j.cellsig.2016.12.003
Pallis, M., & Russell, N. (2000). P-glycoprotein plays a drug-efflux-independent role in augmenting cell survival in acute myeloblastic leukemia and is associated with modulation of a sphingomyelin-ceramide apoptotic pathway. Blood, 95, 2897–2904.
Weisburg, J. H., Roepe, P. D., Dzekunov, S., & Scheinberg, D. A. (1999). Intracellular pH and multidrug resistance regulate complement-mediated cytotoxicity of nucleated human cells. Journal of Biological Chemistry, 274, 10877–10888. https://doi.org/10.1074/jbc.274.16.10877
Ding, S., Chamberlain, M., McLaren, A., Goh, L., Duncan, I., & Wolf, C. R. (2001). Cross-talk between signalling pathways and the multidrug resistant protein MDR-1. British Journal of Cancer, 85, 1175–1184. https://doi.org/10.1054/bjoc.2001.2044
Cerezo, D., Cánovas, M., García-Peñarrubia, P., & Martín-Orozco, E. (2015). Collateral sensitivity to cold stress and differential BCL-2 family expression in new daunomycin-resistant lymphoblastoid cell lines. Experimental Cell Research, 331, 11–20. https://doi.org/10.1016/j.yexcr.2014.11.017
Goler-Baron, V., & Assaraf, Y. G. (2011). Structure and function of ABCG2-rich extracellular vesicles mediating multidrug resistance. PLoS ONE, 6, e16007. https://doi.org/10.1371/journal.pone.0016007
Horio, M., Gottesman, M. M., & Pastan, I. (1988). ATP-dependent transport of vinblastine in vesicles from human multidrug-resistant cells. Proceedings of the National Academy of Sciences of the United States of America, 85, 3580–3584. https://doi.org/10.1073/pnas.85.10.3580
Leibovitz, A., Stinson, J. C., McCombs, W. B., 3rd., McCoy, C. E., Mazur, K. C., & Mabry Mabry, N. D. (1976). Classification of human colorectal adenocarcinoma cell lines. Cancer Research, 36, 4562–4569.
Xu, Z., Liu, Y., Yang, Y., Wang, J., Zhang, G., Liu1, Z., Fu, H., Wang, Z., Liu, H., & Xu, J. (2016). High expression of Mucin13 associates with grimmer postoperative prognosis of patients with non-metastatic clear-cell renal cell carcinoma. Oncotarget, 8(5), 7548–7558.
Khan, S., Ebeling, M. C., Zaman, M. S., Sikander, M., Yallapu, M. M., Chauhan, N., Yacoubian, A. M., Behrman, S. W., Zafar, N., Kumar, D., et al. (2014). MicroRNA-145 targets MUC13 and suppresses growth and invasion of pancreatic cancer. Oncotarget, 5(17), 7599–7609.
Setua, S., Khan, S., Yallapu, M. M., Behrman, S. W., Sikander, M., Khan, S. S., Jaggi, M., & Chauhan, S. C. (2017). Restitution of tumor suppressor MicroRNA-145 using magnetic nanoformulation for pancreatic cancer therapy. Journal of Gastrointestinal Surgery, 21, 94–105. https://doi.org/10.1007/s11605-016-3222-z
Sikander, M. (2012). Mucins: Potential for ovarian cancer biomarkers. International Journal of Biomedical and Advance Research, 3(6). https://doi.org/10.7439/ijbar.v3i6.526.
Gupta, B. K., Sikander, M., & Jaggi, M. J. (2016). Gastroenterol Dig Dis 1(2). Link: Overview of mucin (MUC13) in gastrointestinal cancers. (https://www.alliedacademies.org/).
Sikander, M., Malik, S., Khan, S., Kumari, S., Chauhan, N., Khan, P., Halaweish, F.T., Chauhan, B., Yallapu, M. M., Jaggi, M., et al. (2019). Novel mechanistic insight into the anticancer activity of cucurbitacin D against pancreatic cancer (Cuc D attenuates pancreatic cancer). Cells, 9. https://doi.org/10.3390/cells9010103.
Mills, J. C., Stanger, B. Z., & Sander, M. (2019). Nomenclature for cellular plasticity: Are the terms as plastic as the cells themselves? EMBO Journal, 38, e103148. https://doi.org/10.15252/embj.2019103148
Yuan, S., Norgard, R. J., & Stanger, B. Z. (2019). Cellular plasticity in cancer. Cancer Discov, 9, 837–851. https://doi.org/10.1158/2159-8290.Cd-19-0015
Gurdon, J. B. (1962). The developmental capacity of nuclei taken from intestinal epithelium cells of feeding tadpoles. Journal of Embryology and Experimental Morphology, 10, 622–640.
Takahashi, K., & Yamanaka, S. (2006). Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell, 126, 663–676. https://doi.org/10.1016/j.cell.2006.07.024
Nieto, M. A., Huang, R. Y., Jackson, R. A., & Thiery, J. P. E. M. T. (2016). Cell, 2016(166), 21–45. https://doi.org/10.1016/j.cell.2016.06.028
Shibata, M., Hoque, M.O. (2019). Targeting cancer stem cells: A strategy for effective eradication of cancer. Cancers (Basel), 11 https://doi.org/10.3390/cancers11050732.
Khan, A.Q., Ahmed, E.I., Elareer, N. R., Junejo, K., Steinhoff, M., Uddin, S. (2019). Role of miRNA-regulated cancer stem cells in the pathogenesis of human malignancies. Cells, 8. https://doi.org/10.3390/cells8080840.
Iwata, M. (1968). Study of serum lipids in infants. 1. Serum lipids in healthy infants].Nihon Shonika Gakkai Zasshi, 72, 1075–1081.
Jordan, C. T., Guzman, M. L., & Noble, M. (2006). Cancer stem cells. New England Journal of Medicine, 355, 1253–1261. https://doi.org/10.1056/NEJMra061808
Thiery, J. P., Acloque, H., Huang, R. Y., & Nieto, M. A. (2009). Epithelial-mesenchymal transitions in development and disease. Cell, 139, 871–890. https://doi.org/10.1016/j.cell.2009.11.007
Mani, S. A., Guo, W., Liao, M. J., Eaton, E. N., Ayyanan, A., Zhou, A. Y., Brooks, M., Reinhard, F., Zhang, C. C., Shipitsin, M., et al. (2008). The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell, 133, 704–715. https://doi.org/10.1016/j.cell.2008.03.027
Polyak, K., & Weinberg, R. A. (2009). Transitions between epithelial and mesenchymal states: Acquisition of malignant and stem cell traits. Nature Reviews Cancer, 9, 265–273. https://doi.org/10.1038/nrc2620
Thiery, J. P. (2003). Epithelial-mesenchymal transitions in development and pathologies. Current Opinion in Cell Biology, 15, 740–746. https://doi.org/10.1016/j.ceb.2003.10.006
Huber, M. A., Kraut, N., & Beug, H. (2005). Molecular requirements for epithelial-mesenchymal transition during tumor progression. Current Opinion in Cell Biology, 17, 548–558. https://doi.org/10.1016/j.ceb.2005.08.001
Malanchi, I., Peinado, H., Kassen, D., Hussenet, T., Metzger, D., Chambon, P., Huber, M., Hohl, D., Cano, A., Birchmeier, W., et al. (2008). Cutaneous cancer stem cell maintenance is dependent on beta-catenin signalling. Nature, 452, 650–653. https://doi.org/10.1038/nature06835
Peacock, C. D., & Watkins, D. N. (2008). Cancer stem cells and the ontogeny of lung cancer. Journal of Clinical Oncology, 26, 2883–2889. https://doi.org/10.1200/jco.2007.15.2702
Al-Hajj, M., Wicha, M. S., Benito-Hernandez, A., Morrison, S. J., & Clarke, M. F. (2003). Prospective identification of tumorigenic breast cancer cells. Proceedings of the National Academy of Sciences of the United States of America, 100, 3983–3988. https://doi.org/10.1073/pnas.0530291100
Wielenga, V. J., Smits, R., Korinek, V., Smit, L., Kielman, M., Fodde, R., Clevers, H., & Pals, S. T. (1999). Expression of CD44 in Apc and Tcf mutant mice implies regulation by the WNT pathway. American Journal of Pathology, 154, 515–523. https://doi.org/10.1016/s0002-9440(10)65297-2
Hermann, P. C., Huber, S. L., Herrler, T., Aicher, A., Ellwart, J. W., Guba, M., Bruns, C. J., & Heeschen, C. (2007). Distinct populations of cancer stem cells determine tumor growth and metastatic activity in human pancreatic cancer. Cell Stem Cell, 1, 313–323. https://doi.org/10.1016/j.stem.2007.06.002
Kumar, R., Gururaj, A. E., & Barnes, C. J. (2006). p21-activated kinases in cancer. Nature Reviews Cancer, 6, 459–471. https://doi.org/10.1038/nrc1892
Coniglio, S. J., Zavarella, S., & Symons, M. H. (2008). Pak1 and Pak2 mediate tumor cell invasion through distinct signaling mechanisms. Molecular and Cellular Biology, 28, 4162–4172. https://doi.org/10.1128/mcb.01532-07
Boye, K., & Maelandsmo, G. M. (2010). S100A4 and metastasis: A small actor playing many roles. American Journal of Pathology, 176, 528–535. https://doi.org/10.2353/ajpath.2010.090526
Sherbet, G. V., & Lakshmi, M. S. (1998). S100A4 (MTS1) calcium binding protein in cancer growth, invasion and metastasis. Anticancer Research, 18, 2415–2421.
Feldner, J. C., & Brandt, B. H. (2002). Cancer cell motility–on the road from c-erbB-2 receptor steered signaling to actin reorganization. Experimental Cell Research, 272, 93–108. https://doi.org/10.1006/excr.2001.5385
Estrella, V., Chen, T., Lloyd, M., Wojtkowiak, J., Cornnell, H. H., Ibrahim-Hashim, A., Bailey, K., Balagurunathan, Y., Rothberg, J. M., Sloane, B. F., et al. (2013). Acidity generated by the tumor microenvironment drives local invasion. Cancer Research, 73, 1524–1535. https://doi.org/10.1158/0008-5472.Can-12-2796
Walenta, S., Wetterling, M., Lehrke, M., Schwickert, G., Sundfør, K., Rofstad, E. K., & Mueller-Klieser, W. (2000). High lactate levels predict likelihood of metastases, tumor recurrence, and restricted patient survival in human cervical cancers. Cancer Research, 60, 916–921.
Weinstein, I. B., & Joe, A. (2008). Oncogene addiction. Cancer Research, 68, 3077–3080. https://doi.org/10.1158/0008-5472.Can-07-3293
Sureshbabu, S. K., Chaukar, D., & Chiplunkar, S. V. (2020). Hypoxia regulates the differentiation and anti-tumor effector functions of γδT cells in oral cancer. Clinical and Experimental Immunology, 201, 40–57. https://doi.org/10.1111/cei.13436
Zhang, C., Samanta, D., Lu, H., Bullen, J. W., Zhang, H., Chen, I., He, X., & Semenza, G. L. (2016). Hypoxia induces the breast cancer stem cell phenotype by HIF-dependent and ALKBH5-mediated m6A-demethylation of NANOG mRNA. Proceedings of the National Academy of Sciences of the United States of America, 113, E2047-2056. https://doi.org/10.1073/pnas.1602883113
Cao, Z., Zheng, X., Yang, H., Li, S., Xu, F., Yang, X., & Wang, Y. (2020). Association of obesity status and metabolic syndrome with site-specific cancers: A population-based cohort study. British Journal of Cancer, 123, 1336–1344. https://doi.org/10.1038/s41416-020-1012-6
Ajduković, J. (2016). HIF-1–a big chapter in the cancer tale. Experimental Oncology, 38, 9–12.
Schito, L., & Semenza, G. L. (2016). Hypoxia-inducible factors: Master regulators of cancer progression. Trends Cancer, 2, 758–770. https://doi.org/10.1016/j.trecan.2016.10.016
Mathieu, J., Zhang, Z., Zhou, W., Wang, A. J., Heddleston, J. M., Pinna, C. M., Hubaud, A., Stadler, B., Choi, M., Bar, M., et al. (2011). HIF induces human embryonic stem cell markers in cancer cells. Cancer Research, 71, 4640–4652. https://doi.org/10.1158/0008-5472.Can-10-3320
Lin, Y. T., & Wu, K. J. (2020). Epigenetic regulation of epithelial-mesenchymal transition: Focusing on hypoxia and TGF-β signaling. Journal of Biomedical Science, 27, 39. https://doi.org/10.1186/s12929-020-00632-3
Rankin, E. B., Nam, J. M., & Giaccia, A. J. (2016). Hypoxia: Signaling the metastatic cascade. Trends Cancer, 2, 295–304. https://doi.org/10.1016/j.trecan.2016.05.006
Hajizadeh, F., Okoye, I., Esmaily, M., Ghasemi Chaleshtari, M., Masjedi, A., Azizi, G., Irandoust, M., Ghalamfarsa, G., & Jadidi-Niaragh, F. (2019). Hypoxia inducible factors in the tumor microenvironment as therapeutic targets of cancer stem cells. Life Sciences, 237, 116952. https://doi.org/10.1016/j.lfs.2019.116952
Zhang, H., Lu, H., Xiang, L., Bullen, J. W., Zhang, C., Samanta, D., Gilkes, D. M., He, J., & Semenza, G. L. (2015). HIF-1 regulates CD47 expression in breast cancer cells to promote evasion of phagocytosis and maintenance of cancer stem cells. Proceedings of the National Academy of Sciences of the United States of America, 112, E6215-6223. https://doi.org/10.1073/pnas.1520032112
Thomas, S., Harding, M. A., Smith, S. C., Overdevest, J. B., Nitz, M. D., Frierson, H. F., Tomlins, S. A., Kristiansen, G., & Theodorescu, D. (2012). CD24 is an effector of HIF-1-driven primary tumor growth and metastasis. Cancer Research, 72, 5600–5612. https://doi.org/10.1158/0008-5472.Can-11-3666
Ohnishi, S., Maehara, O., Nakagawa, K., Kameya, A., Otaki, K., Fujita, H., Higashi, R., Takagi, K., Asaka, M., Sakamoto, N., et al. (2013). hypoxia-inducible factors activate CD133 promoter through ETS family transcription factors. PLoS ONE, 8, e66255. https://doi.org/10.1371/journal.pone.0066255
Hashimoto, O., Shimizu, K., Semba, S., Chiba, S., Ku, Y., Yokozaki, H., & Hori, Y. (2011). Hypoxia induces tumor aggressiveness and the expansion of CD133-positive cells in a hypoxia-inducible factor-1α-dependent manner in pancreatic cancer cells. Pathobiology, 78, 181–192. https://doi.org/10.1159/000325538
Chiu, D. K., Zhang, M. S., Tse, A. P., & Wong, C. C. (2019). Assessment of stabilization and activity of the HIFs important for hypoxia-induced signalling in cancer cells. Methods in Molecular Biology, 1928, 77–99. https://doi.org/10.1007/978-1-4939-9027-6_6
Soeda, A., Park, M., Lee, D., Mintz, A., Androutsellis-Theotokis, A., McKay, R. D., Engh, J., Iwama, T., Kunisada, T., Kassam, A. B., et al. (2009). Hypoxia promotes expansion of the CD133-positive glioma stem cells through activation of HIF-1alpha. Oncogene, 28, 3949–3959. https://doi.org/10.1038/onc.2009.252
Xiang, L., & Semenza, G. L. (2019). Hypoxia-inducible factors promote breast cancer stem cell specification and maintenance in response to hypoxia or cytotoxic chemotherapy. Advances in Cancer Research, 141, 175–212. https://doi.org/10.1016/bs.acr.2018.11.001
Porta, C., Paglino, C., & Mosca, A. (2014). Targeting PI3K/Akt/mTOR signaling in cancer. Frontiers in Oncology, 4, 64. https://doi.org/10.3389/fonc.2014.00064
Li, H., Zeng, J., & Shen, K. (2014). PI3K/AKT/mTOR signaling pathway as a therapeutic target for ovarian cancer. Archives of Gynecology and Obstetrics, 290, 1067–1078. https://doi.org/10.1007/s00404-014-3377-3
Tapia, O., Riquelme, I., Leal, P., Sandoval, A., Aedo, S., Weber, H., Letelier, P., Bellolio, E., Villaseca, M., Garcia, P., et al. (2014). The PI3K/AKT/mTOR pathway is activated in gastric cancer with potential prognostic and predictive significance. Virchows Archiv, 465, 25–33. https://doi.org/10.1007/s00428-014-1588-4
Honjo, S., Ajani, J. A., Scott, A. W., Chen, Q., Skinner, H. D., Stroehlein, J., Johnson, R. L., & Song, S. (2014). Metformin sensitizes chemotherapy by targeting cancer stem cells and the mTOR pathway in esophageal cancer. International Journal of Oncology, 45, 567–574. https://doi.org/10.3892/ijo.2014.2450
Mohammed, A., Janakiram, N. B., Brewer, M., Ritchie, R. L., Marya, A., Lightfoot, S., Steele, V. E., & Rao, C. V. (2013). Antidiabetic drug metformin prevents progression of pancreatic cancer by targeting in part cancer stem cells and mTOR signaling. Transl Oncol, 6, 649–659. https://doi.org/10.1593/tlo.13556
Chang, L., Graham, P. H., Hao, J., Ni, J., Bucci, J., Cozzi, P. J., Kearsley, J. H., & Li, Y. (2013). Acquisition of epithelial-mesenchymal transition and cancer stem cell phenotypes is associated with activation of the PI3K/Akt/mTOR pathway in prostate cancer radioresistance. Cell Death & Disease, 4, e875. https://doi.org/10.1038/cddis.2013.407
Zhou, J., Wulfkuhle, J., Zhang, H., Gu, P., Yang, Y., Deng, J., Margolick, J. B., Liotta, L. A., Petricoin, E., 3rd., & Zhang, Y. (2007). Activation of the PTEN/mTOR/STAT3 pathway in breast cancer stem-like cells is required for viability and maintenance. Proceedings of the National Academy of Sciences of the United States of America, 104, 16158–16163. https://doi.org/10.1073/pnas.0702596104
Douville, J., Beaulieu, R., & Balicki, D. (2009). ALDH1 as a functional marker of cancer stem and progenitor cells. Stem Cells Dev, 18, 17–25. https://doi.org/10.1089/scd.2008.0055
Huang, E. H., Hynes, M. J., Zhang, T., Ginestier, C., Dontu, G., Appelman, H., Fields, J. Z., Wicha, M. S., & Boman, B. M. (2009). Aldehyde dehydrogenase 1 is a marker for normal and malignant human colonic stem cells (SC) and tracks SC overpopulation during colon tumorigenesis. Cancer Research, 69, 3382–3389. https://doi.org/10.1158/0008-5472.Can-08-4418
Nishitani, S., Horie, M., Ishizaki, S., & Yano, H. (2013). Branched chain amino acid suppresses hepatocellular cancer stem cells through the activation of mammalian target of rapamycin. PLoS ONE, 8, e82346. https://doi.org/10.1371/journal.pone.0082346
Sunayama, J., Matsuda, K., Sato, A., Tachibana, K., Suzuki, K., Narita, Y., Shibui, S., Sakurada, K., Kayama, T., Tomiyama, A., et al. (2010). Crosstalk between the PI3K/mTOR and MEK/ERK pathways involved in the maintenance of self-renewal and tumorigenicity of glioblastoma stem-like cells. Stem Cells, 28, 1930–1939. https://doi.org/10.1002/stem.521
Bleau, A. M., Hambardzumyan, D., Ozawa, T., Fomchenko, E. I., Huse, J. T., Brennan, C. W., & Holland, E. C. (2009). PTEN/PI3K/Akt pathway regulates the side population phenotype and ABCG2 activity in glioma tumor stem-like cells. Cell Stem Cell, 4, 226–235. https://doi.org/10.1016/j.stem.2009.01.007
Acknowledgements
This work was parcially supported by Start-up from Department of Immunology and Microbiology, School of Medicine, University of Texas Rio Grande Valley, and NIH grants (SC1GM139727, R01 CA210192, and R01 CA206069). Authors thank the CPRIT (RP210180 and RP230419) and UT-System Start Award facilities. Authors thank Ms. Anyssa Rodriguez and Ms. Molly K. Vela for proofreading this manuscript.
Author information
Authors and Affiliations
Contributions
Conceived the idea: S.M., M.S1. and S.C.C; Writing Original draft preparation: S.M., and M.S1; Supervision: S.C.C; Proofreading and editing: S.M, M.S1., S.C.C., S.K., A.D., M.M.Y., M.W., E.C., M.J., and M.S2. All authors have reviewed and given their approval to the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Shabnam Malik and Mohammed Sikander contributed equally to this manuscript.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Malik, S., Sikander, M., Wahid, M. et al. Deciphering cellular and molecular mechanism of MUC13 mucin involved in cancer cell plasticity and drug resistance. Cancer Metastasis Rev 43, 981–999 (2024). https://doi.org/10.1007/s10555-024-10177-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10555-024-10177-8