Abstract
To isolate endophytic bacterium with the ability to specifically convert ginsenoside Rc from Panax quinquefolius. An endophytic bacterium G9y was isolated from Panax quinquefolius and indentified as Bacillus sp. based on 16s rDNA gene sequence. Ginsenoside Rc was effectively converted to Rd by G9y, which was confirmed by thin-layer chromatography and high performance liquid chromatography (HPLC) analysis. The biotransformation conditions were further optimized as follows: inoculum amount 5%, converting temperature 45 °C, medium beef extract peptone broth at pH of 7, and the time of Rc addition was 4 h after bacterium G9y growth, under which ginsenoside Rc was completely converted to Rd by bacterium G9y within 25 h after inoculation. A strain of G9y with the ability to convert ginsenoside Rc into Rd was screened from endophytic bacteria isolated from P. quinquefolius. The results provide a new microbial resource for preparing ginsenoside Rd via biotransformation, and explore a pathway for Rc utilization, which has great potential application value.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Ginsenosides are mainly found in plants of ginseng species (Ginsheng, Panax notoginsheng, Panax quinqueflius, etc.) involving in multiple physiological functions. There are three categories of ginsenosides based on their ginsenogenin structure: protopanaxadiol, protopanaxatriol and oleanolic acid. Ginsenoside Rd acts as a protective agent in nervous system, cardiovascular system and endocrine system (Shi et al. 2005; Nabavi et al. 2015; Fan et al. 2020; Zhang et al. 2020), also exerts a high pharmacologically active in anti-tumor and anti-inflammatory (Zhang et al. 2013; Tian et al. 2020).
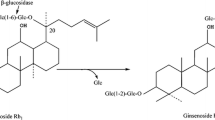
Ginsenoside Rd exhibit the best in physiological activity than that of other protopanaxadiol ginsenosides (Rb1, Rb2, Rc), because it is shorter sugar chain and has higher permeability across the cell membran, ginsenoside Rd also an important intermediate compound that can be transformed into other deglycosylated ginsenosides (Shin and Oh 2015; Fu et al. 2016), thus, it is valuable to prepare ginsenoside Rd from glycosylated ginsenoside. Ginsenoside Rb1, Rb2 and Rc can be metabolized to Rd by microorganisms or enzymes, which is responsible for their structure that possess an additional sugar molecule at the end of the C-20 sugar chain. Most studies have focused on the preparation of ginsenoside Rd from ginsenoside Rb1 through biotransformation (Cui et al. 2016; Senthil et al. 2011; Shahina and Amdadul 2018). Ginsenoside Rc accounted for 7–22% of the total ginsenosides, however, there are few studies on the preparation of ginsenoside Rd from ginsenoside Rc by microbial transformation, besides, previous studies have shown that the transformation pathways of ginsenoside Rc followes the sequence of Rc → C-Mc (Cui et al. 2016; Ten et al. 2014), which limits the utilization of the ginsenoside Rc (Fig. 1).
Endophytes can be obtained from virtually all terrestrial plants by surface sterilization technique in sterile environment (Hardoim et al. 2008; White et al. 2019). Nowdays, many species endophytes have been detected in different medicinal plants (Vendan et al. 2010; Liu et al. 2017), which can secrete extracellular metabolic substances to present strongest capabilities for host, such as growth promotion and bioantagonism. Besides, a variety of enzymes produced by bacteria in the fermentation process that play key roles in transforming active substances in the host, so as to improve the content and medicinal value of the active substances (Cui et al. 2016). Therefore, endophytes pay a significant role for developing and utilization of medicinal plant resources.
In this study, a strain of Bacillus sp. G9y with the ability to convert ginsenoside Rc into Rd was screened from endophytic bacteria isolated from Panax quinquefolius. The optimal conditions of biotransformation of ginsenoside Rc by strain G9y and the changes of ginsenoside in transformation process were preliminarily investigated.
Materials and methods
Materials
Panax quinquefolius roots were purchased from the Changbaishan planting base in Jilin Province, China. Protopanaxadiol ginsenosides standards (Rb1, Rb2, Rc, Rd) and protopanaxatriol ginsenosides standards (Re, Rg1) were from Shanghai Yuanye Bio-Technology Co., Ltd., China. Silica gel G Pre-coated Plate for thin-layer chromatography (TLC) was from Qingdao Haiyang Chemical Co., Ltd., China.
Isolation and purification of endophytic bacteria from Panax quinquefolius
The healthy Panax quinquefolius roots were selected and washed with running water to remove the surface soil. On a clean worktable, the materials were washed three times using sterile water and dried by sterilized filter papers, then immersed in alcohol (70% v/v) for 3 min and sodium hypochlorite solution (4% w/v) for 5 min, rinsed thoroughly with sterile water. Small pieces of 1 cm2 in size were separated from the surface-disinfected roots by a sterile knife and incubated on Nutrient Agar (NA) medium at 37 °C for two days to release endophytic bacteria from Panax quinquefolius. After colony growth, the colonies were picked and streaked onto fresh NA medium using inoculation loops and cultured at 37 °C. This step was repeated for several times to obtain pure bacteria colonies.
Screening of saponin‐transforming strains
After overnight growing on nutrient agar medium at 37 °C, the isolated strains were inoculated into beef extract peptone broth (peptone 10 g/L, beef extract 3 g/L, Sodium chloride 5 g/L). The suspension of the strains that incubated in a shaking incubator (30 °C, 180 rpm) for 12 h were mixed with 1 mg/mL ginsenoside (Rb1, Rb2, Rc, Re, Rg1) at 2:1 (V:V) respectively, continued to culture for 3 days under the same conditions. The reaction mixture was extracted by the addition of an equal volume of aqueous saturated n-butanol and then centrifuged with 10,000 rpm for 5 min, the extracted organic phase was used to select transformation ability of the strains by thin-layer chromatography (TLC) analysis, and then dried followed by dissolving in methanol for high performance liquid chromatography (HPLC) analysis.
Optimization of biotransformation conditions of ginsenoside Rc
The different single-factor effects of the biotransformation conditions for ginsenoside Rc by strain G9y were optimized. The strain suspension reaching logarithmic growth phase was mixed with 1 mg/mL ginsenoside Rc at 2:1 (V:V) continued to culture for 12 h. The different medium pH (4.0, 5.0, 6.0, 7.0, 8.0 and 9.0), incubation temperature (30 °C, 35 °C, 37 °C, 40 °C, 45 °C and 50 °C), inoculation amount (1%, 5%, 10%, 15%, 20%) and time of ginsenoside addition (0, 4, 6, 9, 12 h) were vestigated. The biotransformation process of saponin Rc was further monitored by HPLC.
Analytic methods
The samples (n-butanol extracts) were applied onto Silica gel G Pre-coated Plate with CHCl3, ethanol, water (10:6:1, v/v/v) as the solvent for TLC analysis. The spots on the TLC plates were sprayed with 10% (v/v) H2SO4 in ethanol followed by heating at 105 °C until visualization. HPLC analysis was carried out following the method described by Cui et al. (2016), except that our analysis was performed on Agilent 1200 and using Promesil C18-BIO (250 mm × 4.6 mm) with column temperature of 30 °C. The 16S rRNA gene sequence of the strain (identified by Sangon Biotech Co., Ltd. Shanghai) was compared with the similar sequences which searched on the EzTaxon-e server (http://www.eztaxon.org/), and then the phylogenetic tree was constructed in MEGA software by Neighbor-Joining (NJ) methods. The resulting 16S rRNA gene sequence of strain G9y has been deposited in GenBank under the accession number of MW474824.
Results
Screening of saponin‐transforming strains
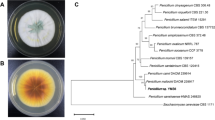
A strain G9y with efficient transformation function of ginsenoside was screened from the endophytic bacteria of Panax quinquefolius by the spots on TLC plate, which converted ginsenoside Rc to ginsenoside Rd, and it was found that the product ginsenoside Rd was not further converted into other saponins. It is clear that as shown in Fig. 2, the staining spots of ginsenoside Rc changed and coverted into a product with the same Rf values of ginsenoside standards Rd, indicating that strain G9y can convert ginsenoside Rc based on Rf values. the staining spots of ginsenoside Rc was disappeared completely after in 3 days, and the staining spots of ginsenoside producted did not change after 7 days, indicating that ginsenoside Rd was the final product. As seen in Fig. 3, after transformation by G9y strain, the Rc peak disappeared and a new peak appeared. The retention time of the product was consistent with that of ginsenoside Rd standard. Therefore, the product was confirmed to be ginsenoside Rd using TLC plate. Furthermore, the result was confirmed by HPLC analysis.
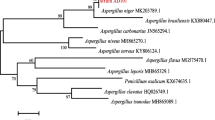
Phylogenetic analysis
The 16S rRNA gene sequence of strain G9y (1484 bp) was compared with the similar sequences which searched on the EzTaxone database, and the results showed that strain G9y exhibited high sequence similarities with B. velezensis CR-502T (100%), B. nematocida B-16T (99.8%), B. subtilis NCIB 3610T (99.7%), B. siamensis KCTC 13613T (99.7%), and B. amyloliquefaciens DSM 7T (99.6%). Figure 4 reveals that the strain G9y was a member of the genus Bacillus with 97% bootstrap support, clustered with B. amyloliquefaciens DSM 7T, B. nematocida B-16T, B. siamensis KCTC 13613T, and B. velezensis CR-502T.
Optimization of biotransformation conditions of ginsenoside Rc
According to a growth curve of this strain, the growth effect in beef extract peptone broth showed the best, followed by Luria–Bertani broth, and then potato dextrose broth (Fig. 5). Appropriate medium pH and culture temperature played the key factors in convertion, as shown in Fig. 6a and b, which represent the transformation of ginsenoside Rc to Rd at different incubation temperatures and medium pH. It was found that the temperature affected the biotransformation significantly, and the strain G9y exhibited the best transformation ability at 45 °C in medium with a pH of 7.
The optimal inoculation amount was studied as shown in Fig. 6c, in which the effect of inoculation amount on ginsenoside transformation was not significant, but it can be observed the maximum degree of transformation when the inoculation amount was at 5%. To enhance the biotransformation, the effect of time of ginsenoside addition on biotransformation was analyzed. Finally, as show in Fig. 6d, strain G9y showed its best transformation ability when the time of ginsenoside addition was at 4 h. It may be because the proliferation of the strain was affected when Rc added too early, and the reaction time was reduced when too late, which caused lower biotransformation efficiency of Rc.
Time‐course of ginsenoside conversion after optimization
Based on the optimal ginsenoside bioconversion conditions, the peak area changes of ginsenoside Rc at different times were analyzed by HPLC. As shown in Fig. 7, transformation product of Rd started generating at 3 h after Rc was added, and the generation rate was accelerated between 6 and 15 h. After 21 h of reaction, ginsenoside Rc was completely transformed into Rd and the conversion rate of ginsenoside Rc was 98%.
Discussion
The method of biotransformation has the advantages of mild reaction conditions, low cost, regional and stereo-specificity, etc. (Anbu et al. 2017; Sultana 2018). Recently, endophytes have been used to improve the content of ginsenoside and transform it into highly bioactive ginsenoside, which has great economic value. Many reports has focused on converting ginsenoside Rb1 to other deglycosylated ginsenoside by endophytic. However, there have been few studies on the transformation of ginsenoside Rc into ginsenoside Rd. An endophytic bacterium Burkholderia sp. GE 17-7 isolated from ginseng that could convert ginsenoside Rb1 into ginsenoside Rg3, F2 or C-K (Fu et al. 2017). Flavobacterium sp. GE32 transformed ginsenoside Rb1 into ginsenosides Rg3 via Rd (Fu 2019; Cui et al. 2016). In our study, an endophytic Bacillus sp. G9y from Panax quinquefolius was screened, which can specifically transform ginsenoside Rc into the target product Rd, may act as a potential microbial source for converting ginsenoside Rc.
Studies have demonstrated that ginsenoside Rd has a variety of biological activities, especially in neuroprotection (Nabavi et al. 2015; Ye et al. 2011). Our work found an endophytic bacterium Bacillus sp. G9y can transform ginsenoside Rc into Rd without other by-products, providing a method for preparing ginsenoside Rd via biotransformation. We have carried out preliminary investigation on the conditions for ginsenoside Rd production, further studies are still necessary on biotransformation conditions for high efficiency preparation of ginsenoside Rd. The transformation mechanism may be that the strain produces α-l-arabinofuranosidase which specifically hydrolyzes the terminal arabinofuranosyl moieties at the 20-position sugar chain of ginsenoside Rc (Xie et al. 2016; Lee et al. 2011). Therefore, the structure and properties of the enzyme is also a topic worth of further research.
References
Anbu P, Gopinath SCB, Chaulagain BP, Lakshmipriya T (2017) Microbial enzymes and their applications in industries and medicine 2016(Editorial). BioMed Res Int
Cui L, Wu SQ, Zhao CA, Yin CR (2016) Microbial conversion of major ginsenosides in ginseng total saponins by Platycodon grandiflorum endophytes. J Ginseng Res 40:366–374
Fan W, Huang Y, Zheng H, Li S, Sun J (2020) Ginsenosides for the treatment of metabolic syndrome and cardiovascular diseases: pharmacology and mechanisms. Biomed Pharmacother 132:110915
Fu Y (2019) Biotransformation of ginsenoside Rb1 to Gyp-XVII and minor ginsenoside Rg3 by endophytic bacterium Flavobacterium sp. GE 32 isolated from Panax ginseng. Lett Appl Microbiol 68:134–141
Fu Y, Yin Z, Wu L, Yin C (2016) Biotransformation of ginsenoside Rb1 to ginsenoside C-K by endophytic fungus Arthrinium sp. GE 17-18 isolated from Panax ginseng. Lett Appl Microbiol 63:196–201
Fu Y, Yin ZH, Yin CY (2017) Biotransformation of ginsenoside Rb1 to ginsenoside Rg3 by endophytic bacterium Burkholderia sp. GE 17-7 isolated from Panax ginseng. J Appl Microbiol 122:1579–1585
Hardoim PR, van Overbeek LS, van Elsas JD (2008) Properties of bacterial endophytes and their proposed role in plant growth. Trends Microbiol 16:471
Lee JH, Hyun YJ, Kim DH (2011) Cloning and characterization of α-l-arabinofuranosidase and bifunctional α-l-arabinopyranosidase/β-d-galactopyranosidase from Bifidobacterium longum H-1. J Appl Microbiol 111:1097–1107
Liu S, Li D, Cui X, Chen L, Nian H (2017) Community analysis of endophytic bacteria from the seeds of the medicinal plant Panax notoginseng. J Agric Sci Camb 9:37
Nabavi SF, Sureda A, Habtemariam S, Nabavi SM (2015) Ginsenoside Rd and ischemic stroke; a short review of literatures. J Ginseng Res 39:299–303
Senthil K, Veena V, Mahalakshmi M, Pulla R, Parvatham R (2011) Microbial conversion of major ginsenoside Rb1 to minor ginsenoside Rd by Indian fermented food bacteria. Afr J Biotechnol 8:6961–6966
Shahina A, Amdadul HM (2018) Biological synthesis of ginsenoside Rd using Paenibacillus horti sp. nov. Isolated from vegetable garden. Curr Microbiol 75:1566–1573
Shi Q, Hao Q, Bouissac J, Lu Y, Tian S et al (2005) Ginsenoside Rd from Panax notoginseng enhances astrocyte differentiation from neural stem cells. Life Sci 76:983–995
Shin KC, Oh DK (2015) Classification of glycosidases that hydrolyze the specific positions and types of sugar moieties in ginsenosides. Crit Rev Biotechnol 36:1036–1049
Sultana N (2018) Microbial biotransformation of bioactive and clinically useful steroids and some salient features of steroids and biotransformation. Steroids 136:76–92
Ten LN, Chae SM, Yoo SA (2014) Biotransformation of the principal ginsenosides of panax ginseng into minor glycosides through the action of bacterium Paenibacillus sp. BG134. Chem Nat Compd 50:691–696
Tian YZ, Liu YP, Tian SC, Ge SY, Zhang BL (2020) Antitumor activity of ginsenoside Rd in gastric cancer via up-regulation of Caspase-3 and Caspase-9. Pharmazie 75:147–150
Vendan RT, Yu YJ, Lee SH, Rhee YH (2010) Diversity of endophytic bacteria in ginseng and their potential for plant growth promotion. J Microbiol 48:559–565
White JF, Kingsley KL, Zhang Q, Verma R, Obi N et al (2019) Review: Endophytic microbes and their potential applications in crop management. Pest Manag Sci 75:2558–2565
Xie J, Zhao D, Zhao L, Pei J, Xiao W et al (2016) Characterization of a novel arabinose-tolerant α-l-arabinofuranosidase with high ginsenoside Rc to ginsenoside Rd bioconversion productivity. J Appl Microbiol 120:647–660
Ye R, Kong X, Yang Q, Zhang Y, Han J et al (2011) Ginsenoside Rd in experimental stroke: superior neuroprotective efficacy with a wide therapeutic window. Neurotherapeutics 8:515–525
Zhang YX, Wang L, Xiao EL, Li SJ, Chen JJ et al (2013) Ginsenoside Rd exhibits anti-inflammatory activities through elevation of antioxidant enzyme activities and inhibition of JNK and ERK activation in vivo. Int Immunopharmacol 17:1094–1100
Zhang C, Liu X, Xu H, Hu G, Shi M (2020) Protopanaxadiol ginsenoside Rd protects against NMDA receptor-mediated excitotoxicity by attenuating calcineurin-regulated DAPK1 activity. Sci Rep 10:1–14
Acknowledgements
This research was financially supported by the Henan Provincial Department of Education Fund (19A180031).
Author information
Authors and Affiliations
Contributions
DZ designed experiments; CZ carried out experiments and wrote the manuscript; YX analyzed experimental results; MG and ZL assisted with high performance liquid chromatography (HPLC) analysis; JZ and QZ inspection experiments.
Corresponding author
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhang, C., Xu, Y., Gu, M. et al. Biotransformation of ginsenoside Rc to Rd by endophytic bacterium Bacillus sp. G9y isolated from Panax quinquefolius. Antonie van Leeuwenhoek 114, 437–444 (2021). https://doi.org/10.1007/s10482-021-01529-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-021-01529-3