Abstract
To select genotypes with stable growth in diverse environments, tree breeders use multisite trials to evaluate genotypic stability and adaptation. Since genotype-by-environment (G × E) interaction effects vary with age, most multisite trials focus only on site effects, ignoring age effects. Liriodendron tulipifera trees are valuable due to their rapid growth and high-quality wood. Currently, multisite trials involving L. tulipifera plants are rare and the lack of data on G × E interaction effects impedes its selection. In this study, to explore the better performance and G × E pattern of L. tulipifera across ages, the growth traits (tree height, H; and diameter at breast height, DBH) of trees that were of five consecutive ages and grown on progeny-testing plantations were studied for 27 open-pollinated families at three sites. The results showed that the heritability of DBH was greater than that of H at almost all ages, and the individual breeding value ranking differed across sites and ages. The additive genetic correlations (rA) between different site pairs were relatively small and varied with age, indicating an age trend for G × E, and showed a difference in traits. It was found that the absolute differences in some monthly average climatic indicators correlated with the G × E. Based on a comprehensive analysis considering stability and productivity, four elite families were identified. These results could aid in selecting stable, adaptable L. tulipifera genotypes and provide a reference for evaluating G × E interaction effects in multiage, multisite trials of other tree species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
With global climate change and the frequent occurrence of extreme climate change events, forest ecosystems will be greatly disturbed in the future, which poses a potential threat to the security of timber production. Breeding plant varieties with broad adaptability and high productivity is an effective approach to coping with changing environments (White et al. 2007).
Tree growth is influenced by many environmental factors (Apiolaza 2012; Burdon et al. 2017). A genotype that performs well at one site may not perform well at another site. In addition to being controlled by genetic factors, tree growth is also affected by interactions between the genotype and the environment (G × E) (White et al. 2007; Des Marais et al. 2013). To date, G × E have been reported for many tree species, including Pinus taeda (Lauer et al. 2021), Pinus massoniana (Yuan et al. 2021), Cunninghamia lanceolata (Bian et al. 2014), Betula platyphylla (Zhao et al. 2015), and Picea glauca (Rweyongeza 2011), indicating that G × E interactions are a widespread phenomenon.
G × E interactions will complicate genotype variance and affect estimates of genetic parameters, leading to biases in family or individual rankings. As a result, genotype performance across multiple environments is unpredictable, making it difficult to identify the best-performing genotypes for a given site (Li et al. 2017). Various analytical methods have been proposed to improve the accuracy of evaluations of G × E interaction effects, including stability analysis based on joint regression (Eberhart and Russell 1966), Shukla’s stability (1972), type-B genetic correlations (Burdon 1977), additive main effect and multiplicative interaction (AMMI) analysis (Gauch and Zobel 1996), and factor analysis (FA) (Cullis et al. 2014) as well as the use of genotype main effect plus G × E (GGE) biplots (Yan 2001) and the use of GGE biplots based on best linear unbiased prediction (BLUP) analysis (BLUP-GGE) (Zhang et al. 2018). GGE biplots can visually display G × E results and can be used to analyze the representativeness and classification of the test site; however, similar to AMMI, this method is limited to fixed-effect models and requires balanced data as well as environmental error homogeneity (Yan et al. 2007). BLUP-GGE analysis, which combines GGE with BLUP analysis, can compensate for the above shortcomings and can be used to obtain more reliable results than raw data. Additionally, the harmonic mean of the relative performance of genotypic values (HMRPGV) approach proposed by Resende (2007), which is based on a mixed model, has been used in many studies because it allows unbalanced data to be analyzed and can address adaptability, stability, and productivity simultaneously (Colombari et al. 2013; de Souza et al. 2020; Evangelista et al. 2021; Yuan et al. 2021).
Generally, a single-site test often inaccurately estimates genetic variance and overestimates heritability by neglecting to consider the G × E variance, whereas a considerable G × E interaction effect across multiple sites decreases the estimated heritability (Sierra-Lucero et al. 2003; Li et al. 2017) and reduces the accuracy of genetic gain predictions (Diaz et al. 2011). Multisite tests are commonly applied in plant breeding to analyze G × E and evaluate the relative performance of the target genotypes, and they can provide higher predictive accuracy than can single-site trials when phenotypes at diverse sites are assessed (El-Dien et al. 2015). However, in the forestry trials, there is clear environmental heterogeneity at tree test sites, resulting in unbalanced data in most cases, and the heterogeneous variance of genotypes at different sites always exhibits a certain degree of correlation or covariance, i.e., a stronger G × E effect (Isik et al. 2017). Additionally considering the challenge of dealing with large amounts of complex, imbalanced data (Möhring and Piepho 2009; Piepho et al. 2012) and the long periodicity of multiage trials, most previous multisite tests in trees have been carried out within a single year or at a certain growth stage (Zhao et al. 2015; Yuan et al. 2021) and have focused mainly on the influence of the site effect, whereas few reports have focused on the age effect.
However, in addition to environmental or site effects, age effects cannot be ignored, as they can inflate the genotypic variance to the magnitude of the genotype × age variance, leading to potential bias in genotype performance assessments (Arief et al. 2019). Previous studies in P. taeda and Pseudotsuga menziesii revealed that the importance of G × E appeared to decline with age, that early growth data might not be reliable for evaluating G × E at maturity (Zas et al. 2003; Roth et al. 2007; Li et al. 2017) and that a multiyear data analysis would provide better estimates of genotype performance (Arief et al. 2015). In P. glauca, Rweyongeza (2011) found that the G × E fluctuated with age and that the type-B correlations with heights (H) were always stronger than those with diameters at breast height (DBH), thus revealing the G × E pattern in the different traits. Therefore, the age effect should receive more attention during multisite trials, because it is important for accurate evaluations of genotypic stability. Furthermore, different studies have identified many different environmental factors that affect tree growth as the main potential factors driving G × E interactions that influence selection (Raymond 2011; Rweyongeza 2011; Chen et al. 2017). Therefore, assessing the extent of and variation patterns in G × E interaction effects, especially across ages, and obtaining a better understanding of the potential key drivers of G × E interactions are very important for tree breeding and deployment purposes.
The L. tulipifera belongs to the Liriodendron genus and is a tree species that exhibits rapid growth and produces high-quality wood and is therefore planted widely in many regions around the world for afforestation and timber production for its high economic and ecological value (Chen et al. 2019). Previous studies on superior genotype selection in Liriodendron have been concentrated mainly on provenance, open-pollinated families, and hybrid combinations (Li and Wang 2001; Wang 2003; Li et al. 2005). However, in terms of genotypic stability, few studies have employed multisite tests of L. tulipifera, especially to analyze multisite data collected over many years. In this study, 27 open-pollinated L. tulipifera families from progeny test plantations at three sites were used to compose an experimental population. We measured their growth traits at five successive ages and analyzed the additive genetic correlation of genotypes among sites and their genotypic stability. The aims of this study were as follows: (1) to explore the age trends in G × E interaction effects on growth traits and (2) to select genotypes with excellent growth performance and stability across ages and sites. This work will help to improve the productivity of L. tulipifera plantations and promote the extensive application of improved varieties.
Materials and method
Experimental sites, materials and design
The progeny tests were conducted at three sites, namely Xiashu town (XS), Jingdezhen city (JDZ), and Jingxian County (JX), which are located in East China (Table 1). The experimental population comprised a total of 27 L. tulipifera open-pollinated families (marked as Lt1-27, average of 23 progenies in each family per site), which were derived from the same batch of open-pollinated seeds collected in 2005 from parental trees during a provenance trial related to growth characteristics and flowering and fruiting performance. The provenance trial plantation was established in 1993 and is located at the Xiashu Forest Farm affiliated with Nanjing Forestry University (latitude and longitude: 31°59′ N, 119°14′ E). More details about this provenance trial were presented in our previous study (Xia et al. 2021). The seeds were sown in 2005, and the seedlings were grown at the nursery on the campus of Nanjing Forestry University. During the next year, a large proportion of the seedlings were used to establish progeny test plantations at the XS and JDZ sites. In 2006, because of the inappropriate conditions at the target test site, another group of seedlings was temporarily transferred to a nursery located in Siyang County, Suqian city, Jiangsu Province, China (33°23′ N, 118°20′ E) for 1 year. Then, the progeny test plantation was established at the JX site in 2007. All the management measures carried out at each location were similar to ensure uniform growth conditions. A randomized complete block design was applied to the three test sites, and JDZ, JX and XS had 4, 2, and 3 blocks, respectively. There were 10 plants per plot at these three sites, with a plant spacing of 4 × 4 m. However, due to natural mortality, missing manual records and other factors, there were unequal numbers of plants per family and observed years (unbalanced data).
Growth trait measurements and climatic data collection
At each test site, when the trees were 4, 5, 6, 7, and 8 years old, tree H and DBH were measured at the end of the vegetative period each year. The seedling height (SH) and ground diameter of the seedlings (SGD) were also measured during the year (age 2) when the progeny test plantation was established, and they were employed as the fixed covariates to eliminate the effects of inconsistent growth at the seedling stage during the subsequent model analysis. In total, an average of approximately 473–1020 trees were measured. The daily climatic data for the cities where the test sites were located were provided by the China Meteorological Data Service Centre (http://data.cma.cn/en). We obtained some daily climatic indexes for the years when the trees were 5 to 8 years old, and then, the average values for the daily temperature (T, °C), daily maximum temperature (TM, °C), daily minimum temperature (Tm, °C), daily relative humidity (RH, %) and daily total rain precipitation (PP, mm) were calculated separately at the annual and monthly levels.
Statistical analysis of growth traits
All statistical analyses were conducted in R software version 3.6.0 (Team R Core 2019). The stem volume index (VI) was calculated according to the formula of VI = H × DBH2 (Liu et al. 1991). The average survival ratio (SR) of one site and phenotypic coefficient of variation (PCV) was calculated as follows: PCV = SD/u and SR = nob/Nto, where u is the mean value of the trait (H or DBH) at one age, SD is the standard deviation, and nob and Nto are the average numbers of the observed individuals across ages and the total number across each site, respectively. Pairwise comparisons of growth traits between different locations were performed using the nonparametric Wilcoxon test with the Wilcox function in R. The heatmaps in this study were generated with the ComplexHeatmap package (Gu et al. 2016).
Variance component and genetic parameter estimates
The variance components and genetic parameters were estimated using the ASReml-R 3.0 package (Gilmour et al. 2008) via the BLUP approach with the restricted maximum likelihood (REML) method. To estimate the estimated breeding value (EBV) of each family at each site across ages, we conducted a single-site-single-age (Eq. 1) analysis to study the performance differences in families at different ages and sites by following a general linear mixed model (LMM) as follows:
To explore the age trend of the G × E effects and genotypic stability across tree ages, an LMM was constructed for the analysis of multisite-single-age (Eq. 2) as follows:
Moreover, to evaluate the growth stability and identify the adaptable genotype for each site, we also implement a BLUP-GGE biplot analysis based on the multiage-single-site (Eq. 3) model:
In the above equation, yijksy is the observation of the k-th individual of the i-th family in the j-th block at the s-th site and y-th age and u is the overall mean; xij is the growth traits of SH or SGD for i-th family in j-th block and treated as a fixed covariate, and β is the slope associated with covariate. Yy and Ss are the fixed effects of the y-th age and s-th site, respectively. Bj, B(S)js and B(Y)jy are the random effects of the j-th block, and that within the s-th site and within y-th age, respectively. Fi, FBij, F(S)is and F(Y)iy are the random effects of the i-th family, interaction of i-th family and j-th block, i-th family within s-th site, and i-th family within s-th age, respectively. eijk is the random residual effect of the k-th individual from the i-th family in the j-th block (eijks, at the s-th site or eijky, at the y-th age).
In Eq. (1), we assume that the error effect of e and random effect of F and FB had normal distribution with zero mean and identical and independent variance—NID (0, R) and NID (0, G)—where \(R = \sigma_{e}^{2} I_{n}\); \(G = \sigma_{F}^{2} I_{i} \oplus \sigma_{{{\text{FB}}}}^{2} I_{ij}\); \(\sigma_{e}^{2}\) is the error variance, I is the identity matrix and n, i and ij are the number of observed individuals, families, and blocks times families in each single-site-single-age trial, respectively; and \(\oplus\) is the direct sum. Given the differences in random effects such as genotype and error effect in different sites or ages, that is, variance heterogeneity, it is necessary to decompose them; thus, in Eq. (2), the variance of families at different sites is assumed to be heterogeneous and to exhibit covariance with each other, while the covariances of block and residual across sites are assumed to be zero, although also with the heterogeneous variance, namely \(G_{F(S)} = \left[ {\begin{array}{*{20}c} {\sigma_{F(s1)}^{2} } & {\sigma_{F(s1)F(s2)} } & {\sigma_{F(s1)F(s3)} } \\ {\sigma_{F(s1)F(s2)} } & {\sigma_{F(s2)}^{2} } & {\sigma_{F(s2)F(s3)} } \\ {\sigma_{F(s1)F(s3)} } & {\sigma_{F(s2)F(s3)} } & {\sigma_{F(s3)}^{2} } \\ \end{array} } \right] \otimes I_{i}\), \(G_{B(S)} = \sigma_{B(s1)}^{2} I_{s1} \oplus \sigma_{B(s2)}^{2} I_{s2} \oplus \sigma_{B(s3)}^{2} I_{s3}\), and \(R = \sigma_{e1}^{2} I_{s1} \oplus \sigma_{e2}^{2} I_{s2} \oplus \sigma_{e3}^{2} I_{s3}\), where s1, s2 and s3 are the number of individuals at three different sites and \(\otimes\) is the direct product. In Eq. (3), we assume that the family effect variances are heterogeneous at different ages but with the same identical correlation between each pair of ages, and the block and error effect at different ages is also assumed to be heterogeneous and with covariance of zero. The variance of R and G are as follows:
\(G_{B(Y)} = \sigma_{B(y1)}^{2} I_{y1} \oplus \sigma_{B(y2)}^{2} I_{y2} \oplus \sigma_{B(y3)}^{2} I_{y3} \oplus \sigma_{B(y4)}^{2} I_{y4} \oplus \sigma_{B(y5)}^{2} I_{y5}\), and \(R = \sigma_{e(y1)}^{2} I_{y1} \oplus \sigma_{e(y2)}^{2} I_{y2} \oplus \sigma_{e(y3)}^{2} I_{y3} \oplus \sigma_{e(y4)}^{2} I_{y4} \oplus \sigma_{e(y5)}^{2} I_{y5}\), where y1, y2, y3, y4 and y5 are the number of individuals at each age and \(\rho\) is the correlation between each pair of ages.
For multisite-single-age analysis, the heritability was estimated with reference to Isik et al. (2017) using the following formula, and its standard errors were estimated using the delta method (Holland et al. 2010; Isik et al. 2017):
where \(h_{i}^{2}\) and \(h_{f}^{2}\) are the narrow-sense heritability and heritability of family mean, respectively. \(\sigma_{F(s)}^{2}\), \(\sigma_{F(ss^{\prime})}^{{}}\), and \(\sigma_{e(s)}^{2}\) are the variance components of the family within site s, the covariance between sites s and s’, and the residual error variance within site s, respectively. \(\overline{{\sigma_{F(ss^{\prime})}^{{}} }}\) and \(\overline{{\sigma_{e(s)}^{2} }}\) are the average values of their corresponding variance. The rhs is the repeated harmonic mean of each family at each site, \(S_{f}^{2}\) is the number of sites for tested family f, and other parameters are the same as described above. The significance of the variance component was inferred from the estimated standard error (Isik et al. 2017).
HMRPGV and additive genetic correlation
The HMRPGV method, which can simultaneously account for adaptability, stability, and productivity, was used to evaluate the genetic stability of the different genotypes at the various sites. The applied formula was as follows (Resende 2007; Colombari et al. 2013; Olivoto et al. 2020):
where \({\text{RPGV}}_{i} = 1/S(\sum\nolimits_{s = 1}^{S} {{\text{GV}}_{is} /{\text{mu}}_{s} } )\) and \({\text{GV}}_{is} = {\text{mu}}_{s} + g_{is}\), GVis is the genetic value of genotype i at site s (estimated from multisite-single-age model), mus is the overall mean value of the trait at site s, and gis is the genotypic value or EBV value of the i-th genotype at the s-th site, and S is the number of sites. We used the additive genetic correlation (rA) to express the size of the G × E interaction effects for each of the two sites at the same age. Generally, when this value is lower, suggesting there is higher G × E interaction effect. The rA was calculated with the following expression (Burdon 1977; Isik et al. 2017):
where \(\sigma_{F(s)}^{2}\) and \(\sigma_{F(s^{\prime})}^{2}\) are the genotype variance within site s and s’, respectively, and \(\sigma_{F(ss^{\prime})}^{{}}\) is the additive genetic covariance of the same trait between the sites s and s’. The Pearson correlations between rA and the absolute value of the difference in monthly and annual climatic indicators were obtained using the cor function; then, the stepwise regression was used to screen the important and significant variables predicting rA, and their explained variation was obtained with multivariate regression.
GGE biplot analysis
A GGE biplot was constructed based on the average EBV value (according to Eq. (3)) for each family at each site using the following model (Yan et al. 2007; Zhang et al. 2018):
where yis is the adjusted mean EBV value for the i-th family at the s-th site, βs is the mean EBV value for all families at the s-th site, λ1 and λ2 are the singular values for the first two principal components, γi1 and γi2 are the scores for family i for these two principal components, δs1 and δs2 are the corresponding scores at site s, and \(\varepsilon_{{i{\text{s}}}}\) is the residual error. The GGE biplot was generated using the GGEBiplotGUI package (Frutos et al. 2014) in R, with the parameters being set at 0 (nonstandardized) for scaling, G + GE for center, and symmetric for singular-value partitioning (SVP) (Zhang et al. 2018), resulting to three specific plot types. In “Which Won Where/What”, the genotype located at the vertex of the polygon was the one with the best performance in the site of the region; in “Discriminativeness vs. representativeness”, the length of the dotted line from the origin represented the discrimination of each site; in “Mean vs. Stability”, the length of the dotted line represented stability, and the shorter the line, the more stable it was.
Results
Growth trait performance and climatic indicators
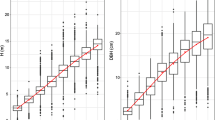
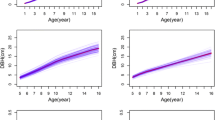
There was an approximately 74.7% average survival rate across the different ages and sites (Table 1), and the box plot (Fig. 1a, b) showed that, except for the difference in H between the JX and XS sites at age 7, which was not significant, there were significant differences (P < 0.05) among the three sites at other ages. The descriptive statistics table (Table 2) shows that across sites and ages, the total average H, DBH and VI ranged from 0.78 to 10.45 m, 1.40 to 14.66 cm and 0.002 to 0.2300 m3, respectively, and the PCVs ranged from 20.01 to 43.95% for H, 27.68 to 56.85% for DBH, and 29.90 to 54.90%. The detailed monthly average climatic indicators (from ages 5 to 8) at the three sites are shown in Table S1. The changing patterns in average growth traits at the three sites were different from those of the various annual average climatic indicators (Fig. 1c) and were the opposite of those of the T, Tm and RH. Taken together, these results indicate that these three climatic indicators may be important correlative factors affecting tree growth traits.
Variations in growth traits and climatic factors in different locations and ages. a and b: Box plots showing the differences between the three sites with pairwise comparisons by Wilcox test in the H (a) and DBH (b) groups. The whiskers mean 1.5 times the interquartile range above the upper quartile and below the lower quartile, but when there is no maximum or minimum value exceeding the upper and lower beard lines, the whiskers are the location of the maximum or minimum value. At each age, the same lowercase letter above the box plot means that there is no difference between two sites; otherwise, there is a difference at the 0.05 level. JDZ, JX and XS are the three tested sites in the text and are the same below. c: The heatmaps of different climatic indicators varied by age and site, and they included the annual average temperature (T), annual average daily maximum temperature (TM), average annual daily minimum temperature (Tm), average annual relative humidity (RH) and total annual rain precipitation (PP)
EBV values varied with ages and sites through single-site-single-age analysis
Based on single-site-single-age model, the EBV ranking of the H and DBH in each family differentially changed at different ages and sites (Fig. 2). For example, regardless of whether the trait was H or DBH, the Lt4 family performed well at sites JDZ and XS but performed poorly at site JX. Family Lt20 performed well at site JX, but its performance at sites JDZ and XS was moderate. In addition, even if the same family was located at the same site, there were also differences during different ages. For instance, the family Lt7 performed well in H at the age of 4 at site XS, but it performed moderately well at other ages. Of course, the extent of this difference varied among different traits. This observation showed that the age and site should be considered within the family selection.
Age trends of variance components, rA and heritability with multisite-single-age analysis
Various genetic parameters were estimated using a multisite-single-age model to display the age trend of them. As shown in Table 3, except for the age of 6 for trait H, the fixed site effects were significant (P < 0.05) or highly significant (P < 0.01) in H and DBH at all other ages, and those of SH and SGD were all highly significant. The family variance within sites JDZ, JX and XS for trait H ranged from 0.048 to 0.550, 0.056 to 1.623, and 0.043 to 0.688, respectively, while that for the trait DBH ranged from 0.097 to 0.908, 0.136 to 3.125, and 0.202 to 3.158, respectively. With age, the family effect variance all increased at all sites; however, the age trend of additive covariance differed among locations and was different, showing that the covariance fluctuated with age and even displayed negative correlations among sites at some age. The additive genetic covariance among sites ranged from -0182 to 0.312 in H and -0.069 to 0.567 in DBH, respectively; both traits had the maximum value between sites JDZ and JX, and minimum value between site JX and XS. Moreover, the covariance values were all negative between sites JX and XS across ages.
For H, the rA ranged from 0.063 to 0.212, −0.084 to 0.054, and −0.125 to −0.024 between sites JDZ and JX, sites JDZ and XS, and sites JX and XS, respectively. For DBH, the rA of these three pairs of sites ranged from 0.086 to 0.279, 0.053 to 0.241, and −0.119 to 0.213, respectively. The age trend of rA showed different patterns of temporality (Fig. 3). The rA of the three pairs of sites generally increased with age for DBH; while for H, only the age trend of rA between JDZ and XS showed an increase, whereas the rA in the other two pairs of sites fluctuated with age but showed a decreasing trend overall. Additionally, all estimated rA values were below 0.4, relatively small, or even negative at some ages.
The hi2 and hf2 of the H ranged from 0.008 to 0.055 and 0.030 to 0.211, respectively; those for the DBH were 0.006 to 0.124 and 0.030 to 0.405, respectively (also shown in Table 3). The maximum value of both heritabilities was at age 6 for H and the minimum was at age 7 for DBH. The heritability of the H and DBH all fluctuated with age, showing an increasing trend and then declining with age. These heritability values are all relatively low, and the heritability of the DBH was greater than that of the H aside from that at age 4.
Correlation analysis between rA and the differences in climatic indicators
To explore the possible environmental factors that may affect G × E, we used a correlation analysis between the rA and various climate factors at two sites of the same age. The results (Table 4) showed that the differences in the annual average T, TM, TM, RH and PP (DT, DTM, DTm, DRH and DPP) between the sites were not significantly correlated with the rA (−0.57 to 0.15 for H; −0.50 to 0.29 for DBH), but a further correlation analysis using the monthly average climatic indicator showed that the rA of H was significantly negatively correlated with the differences in the monthly average RH in January (DRH.Jan) and February (DRH.Feb) (P < 0.05), and the average T in July (DT.July) (P < 0.01) while that of DBH was significantly negatively correlated with the differences in the monthly average TM in April (DTM.Apr) and Tm in October (DTm.Oct) (P < 0.05). Thus, the greater the difference between these climatic indicators was, the lower the rA and the more significant the interaction between the genotype and the environment. What’s more, multivariate regression showed that (Table 5) DRH.Feb, DTM.Apr, and DT.July had a significant effect on rA of H, with a total of 74.21% explained variation, while that of DBH was only DTm.Oct found significantly with 30.66%.
Family selection for fast and stable growth based on HMRPGV
Given the interaction between genotype and site, we estimated the HMRPGV value of each family based on the multisite-single-age analysis, of which results showed that the HMRPGV ranking varied among families, and the detailed ranking information for the different ages is shown in Table S2. Under a selection rate of 30%, we used the HMRPGV ranking to select the H and DBH at the same time. However, some families were selected only at a certain age or only for H or DBH traits. For example, the Lt27 family was selected only at the age of 7 for the H, and family Lt6 was simultaneously selected at the ages of 5, 6 and 8 of DBH but not for H at any one age. The results (Fig. 4) showed that families Lt3, Lt4, Lt5, Lt7 and Lt20 were selected according to H and DBH concurrently for at least three ages, and their phenotypes were stable and excellent in terms of the traits and time scale, especially for the Lt3, Lt5 Lt7 and Lt20 families, which were selected for H and DBH at all five ages. Thus, these families could be used as elite genotypes for extensive planting.
Scatter plots of HMRPGV of DBH vs. HMRPGV of H across ages. H4, H5, H6, H7 and H8 represent the H at ages 4, 5, 6, 7 and 8, respectively. DBH4, DBH5, DBH6, DBH7 and DBH8 also represent the DBH at ages 4, 5, 6, 7 and 8, respectively. The two dashed lines, which are separately perpendicular to the X and Y axes, represent the selection intensity of 30%. The red points located in the upper right part of the intersection of the two lines represent the families selected on both H and DBH by the value of the HMRPGV
GGE biplot analysis
The GGE biplots were conducted based on EBV value that estimated by multiage-single-site analysis, and of which results (Fig. 5) showed that the sum of the variance explanation percentages of the first and second principal components (also named AXIS1 and AXIS2) were 82.23% and 89.13% in the H and DBH, respectively, indicating that the analysis was reliable.
GGE biplots of growth traits. The GGE biplots were created based on the adjusted phenotypic means by multiage-single-site analysis. AXIS1 and AXIS2 represent the first and second principal components (PCA 1 and PCA 2), respectively, and also indicate the variation explaining the proportion on each axis. The blue letters represent different sites, and the green letters represent the different families in each biplot. The blue and green lines with a small circle are average environmental vectors. a and d: “Which Won Where/What” biplots for H (a) and DBH (d). b and e: “Discriminativeness vs. representativeness” biplots for H (b) and DBH (e). c and f: “Mean vs. Stability” biplots for H (c) and DBH (f)
In the “which won where/what” analysis for the H and DBH, the family Lt4 performed best at the site XS for both traits, and the families Lt5 and Lt20 performed the best for the H at the sites JDZ and JX, respectively. In the “discriminativeness vs. representativeness” assessment, site JDZ had higher representativeness than the other two sites in terms of both H and DBH, but site JDZ had the worst performance, and the site JX performed best in terms of discriminativeness. In the “mean vs. stability” assessment, the Lt5 family had the highest mean values of H across different sites, followed by the Lt3 family, which had the strongest stability; for DBH, the Lt3 family had the highest productivity and stability, followed by the Lt5 and Lt7 families, but Lt7 had higher stability than Lt5. These results can be corroborated with the above HMRPGV analysis to screen excellent and reliable genotypes.
Discussion
Variations in growth traits across ages and sites
Genetic variation is the basis for genotype selection (White et al. 2007). In this study, the average PCVs of growth traits across different sites and ages were relatively high, which was conducive to selection (Lin et al. 2013). However, the PCV of V was the greatest at the overall level, followed by DBH that was greater than that of H, indicating that the variation in DBH is more abundant, which was different from the findings of Yuan et al. (2021) on Masson pine. Additionally, the differences in growth traits between any two sites were significant for almost all age groups, which indicated that the site effect was significant; these results were consistent with the subsequent Wald test for site effects. A similar highly significant site effect for H and DBH was also found on C. lanceolata by Bian et al. (2014) and the same substantial differences were also observed in the crown size and bark thickness among sites. Zhang et al. (2021) also found that the DBH, survival rate, and stand volume significantly differed among sites. These findings suggested that the site effect was significant and important. Moreover, the EBV values of families displayed a diverse change trend across age and site, suggesting that interactions might be present between the family and site or age. Yuan et al. (2021) also reported a significant effect of the site and interaction on families of P. massoniana. Even the performance of the same family during different years or ages is different; that is, the stability is different among sites or ages. These observed growth trait variations by site and age indicate that age and site effects should be taken into consideration in G × E interaction effects.
Age trends of G × E and heritability of growth traits
To date, G × E interaction effects have been reported in many tree species (Bian et al. 2014; Zhao et al. 2015; Lauer et al. 2021; Rweyongeza 2011; Yuan et al. 2021). For example, de Souza et al. (2020) reported that the variance of the G × E interaction effect was significantly correlated with the survival rate of C. citriodora according to a likelihood ratio test. Lauer et al. (2021) also observed the occurrence of G × E on the height and diameter revealed by rA among sites in loblolly pine, with genetic correlations among test sites. Yuan et al. (2021) also found, through a joint analysis of variance, that G × E interaction effects on the growth traits of Masson pine were significant. However, these G × E interaction effects were reported only at a certain age. In a study using multiyear breeding data for performance prediction, Arief et al. (2019) pointed out the presence of bias in genetic evaluations of single-year data and demonstrated the advantages of multiyear data. Rweyongeza (2011) also found a pattern of G × E interaction effects on different traits with age in P. glauca. Genetic correlations between trials are commonly used to assessment the amount of G × E interaction (Berlin et al. 2014; Chen et al. 2017). In this study, through a multisite test using a mixed model for many years of analysis, we found that the rA displayed different age trends in the growth traits, and the estimated rA in this study was relatively small, indicating that there was a certain large G × E effect, and the correlation between site XS and JX was negative, that is, the difference between these two sites was greater, with a more obvious G × E interaction. Similarly, large negative correlations were observed in P. contorta by Haleh et al. (2018), indicating significant differences between sites. The age trends of rA varied among different sites. For the trait DBH, the rA among these three site pairs showed an increasing trend overall, revealing a gradually decreasing G × E effect. However, for the trait H, except for sites JDZ and JX, this increasing trend was not obvious; on the contrary, it tended to decrease in general, indicating that trait H may be more vulnerable to the interaction effect between the genotype and environment. This fact was in agreement with the above-described trend of greater DBH heritability than H heritability, but in Picea glauca, Rweyongeza (2011) found that although the G × E interaction effects fluctuated with age, the type-B correlations of H were almost always higher than those of DBH, revealing the different G × E patterns in different traits.
Moreover, as many reports have pointed out, G × E will not only affect the family ranking but can also interfere with the accurate evaluation of genetic parameters, including heritability (Li et al. 2017; Zhang et al. 2021; Zhou et al. 2021). Heritability will be overestimated when the G × E interaction effect is not considered (Sierra-Lucero et al. 2003). Here, both hi2 and hf2 were estimated to be relatively low, and with a higher standard error, which might be related to the serious imbalance of data although we considered this in the calculation of heritability. de Souza et al. (2020) observed a similar result in C. citriodora; they inferred that it might have been related to the lower diversity of seed sources, the occurrence of natural selection processes, or the high environmental variability. Low cross-site heritability is caused mainly by high environmental residual variance or the loss of additive genetic variance (Stackpole et al. 2010; Isik et al. 2017). In this study, artificial or natural missing data might lead to imbalances in the data and affect the genetic parameter estimates. Bian et al. (2014) and Wu et al. (2007) also reported that heritability changes with age in radiata pine, which might be related to the impact of early planting, different sampling and measurement methods, and greater environmental impacts. With age, the heritability of both H and DBH fluctuated, but the DBH showed a larger value at both the individual and family levels at almost all ages, indicating that DBH was more strongly controlled by genetics, and similar results were also observed in many other tree species (Dieters et al. 1995; Hiraoka et al. 2019; Kusnandar et al. 1998).
Monthly average climate indicators correlated with G × E
Previous studies of many tree species have shown that the latitude, altitude, rainfall, precipitation, temperature, and other factors might be the driving factors underlying the G × E interaction effects that influence accurate selection, some of which have confirmed the importance of G × E interaction effects (Raymond 2011; Rweyongeza 2011; Cullis et al. 2014; Chen et al. 2017; Wu et al. 2021). In this study, due to the limitation in the number of tested sites, we conducted a correlation analysis on the differences in the rA and its corresponding climate factors between two sites in multiple years to make full use of the environmental information during different years. Although the rA correlation with the difference in annual average climate factors was not significant, it was surprising that the DTM between any two sites positively correlated with their rA, suggesting little impact of genotype and environment interactions, although Liriodendron is reportedly sensitive to low temperatures (Lu et al. 2015). This finding might be related to the fewer tested sites, large standard error of the rA estimation at some ages, and etc. Furthermore, the rA was significantly correlated with some monthly average climate factors, in which the DRH.Jan, DRH.Feb and DT.July were significantly negatively correlated with the rA of H, indicating that these three climatic factors were closely related to the G × E interaction effect on H. Otherwise, for DBH, the monthly average DTM in April( DTM.Apr) and DTm in October had a highly negative significant correlation with rA, which was different from that of trait H, suggesting that growth traits might respond distinctly to different environments, resulting in differences in their correlations with rA. Lauer et al. (2021) adopted multivariate regression to analyze the linear relationships between genetic correlations and differences in environmental factors in P. taeda; they also showed that H and DBH responded differently to the linear relationships between environmental factors at the test sites, and the difference in altitude could explain additional environmental factors that influence G × E interaction effects aside from temperature. Chen et al. (2017) also found that spring and autumn cold indices, annual average temperature, and altitude were all significantly Pearson correlated with genetic correlation between sites, but further stepwise regression showed that only the first two had significant effects, indicating that they were the main driving factors of G × E for H in Norway spruce. In our study, multivariate regression analysis showed that DRH.Feb, DTM.Apr and DT.July had a significant effect on rA in H, with explaining a greater total variation (74.21%); while in DBH, only DTm.Oct made the main contribution to rA, and the explained variation was 30.16%, which was all greater than 27.8% that in the study of Chen et al. (2017). Therefore, these significant variables with large variation might make the responses of different traits various, which further contributed to the understanding of G × E interaction effects on Liriodendron and should be given more attention in forestry promotion and planting efforts in the future.
Comprehensive selection for elite genotypes
In addition to the influence of climatic factors mentioned above, trees with diverse genotypes also respond differently to environmental conditions, resulting in differences in genotype performance and even changes in genotype ranking; these factors compromise breeding selection (Raymond 2011; Zhao et al. 2015). There are two main breeding strategies to address these factors: one is to select a genotype that is suitable for a specific location, and the other is to select a genotype with a wide range of adaptability.
GGE biplots and HMRPGV analysis have been applied in many studies investigating G × E interaction effects (Yan et al. 2007; Zhang et al. 2018; de Souza et al. 2020; Evangelista et al. 2021). HMRPGV considers adaptability, stability, and productivity simultaneously (Resende 2007), while GGE biplots provide a visual advantage but are limited by model requirements for balanced data (Yan et al. 2007). Forest tree test sites are relatively complicated, the environmental conditions are heterogeneous, and data imbalances are common in data collected from these sites. In view of these limitations to forestry applications, Zhang et al. (2018) proposed combining a spatial BLUP model with a GGE one to avoid the above constraints; they demonstrated that this approach was more reliable than direct GGE analysis. Thus, in this study, we referred to their method. This approach allowed us to consider age effects comprehensively and was more convenient and reliable than analyzing the average values from single-age or multiage trees (Arief et al. 2019).
In this study, the “which won where/what” assessment of the biplots showed that the most suitable genotypes for H and DBH, namely Lt4 families, were found at the XS site. This result suggested that genotype Lt4 was more specifically adapted to the environment at site XS than at sites JDZ and JX. However, at the JDZ site, the best-performing genotypes for the H and DBH were different, indicating that the traits had different adaptive responses to different environments and reflecting the necessity of comprehensive selection for multiple traits (Olivoto and Nardino 2020). Additionally, to the extent possible, an ideal test site should have strong discriminativeness and representativeness (Yan 2001). Among the three sites tested in this study, the JX site was more conducive to distinguishing the different genotypes by H and DBH, respectively, and the angle between site XS and JX in H was larger than 90°, indicating that they were strongly negatively correlated, which was also in agreement with their rA. Site JDZ was closer to the average environmental vector for both the H and DBH, and could therefore be used as test sites for selecting a genotype to adapt to a specific environment.
When selecting for productivity and stability simultaneously, we found that the GGE biplots of both the H and the DBH showed that, among all genotypes, those of the Lt3, Lt5, Lt7 and Lt20 families performed better. This result was similar to those from the HMRPGV ranking. Although the HMRPGV ranking of the families varied by different traits and ages, these four families were always selected as superior genotypes during the comprehensive consideration of these variations. Thus, the four common families (Lt3, Lt5, Lt7 and Lt20) were finally selected as the elite families. Similarly, de Souza et al. (2020) in C. citriodora, Evangelista et al. (2021) in soybean, and Yuan et al. (2021) in P. massoniana successfully identified superior stable genotypes through HMRPGV analysis. The excellent and stable L. tulipifera genotypes identified in this study can help increase the productivity of L. tulipifera plantations and these trees can be planted and popularized in regions similar to the test site in the future.
Notably, the number of sites tested in this study was limited; more quantitative and diverse genotypes should be included in future trials. Moreover, considering that G × E interactions can help to not only accurately evaluate genotypes in conventional breeding but also improve the accuracy of prediction through genome selection (Bajgain et al. 2020), which can in turn help to determine G × E interaction effects (Li et al. 2017), G × E interactions are highly important to accelerating the breeding process.
Conclusions
There were found substantial variations in growth traits at different sites and ages, resulting in differing EBV ranking. The rA among sites reflected the existence of G × E effects and displayed different age trends in different site pairs and traits; thus, the effects of these factors should not be ignored during genotype selection. Notably, the absolute difference in some monthly average climatic indicators correlated with rA and might affect the G × E interaction on growth traits; therefore, these indicators should be more carefully considered when considering the deployment of new varieties in the future. Based on a comprehensive evaluation, we identified four families (Lt3, Lt5, Lt7 and Lt20) that exhibited excellent performance in growth and adaptation. These four families could be used as elite genotypes for future deployment.
Availability of data and material
All relevant data are available from the corresponding author on reasonable request.
References
Apiolaza LA (2012) Basic density of radiata pine in New Zealand: genetic and environmental factors. Tree Genet Genome 8:87–96. https://doi.org/10.1007/s11295-011-0423-1
Arief VN, DeLacy IH, Jose C, Thomas P, Ravi S, Hans-J B, Tian T, Basford KE, Dieters MJ (2015) Evaluating testing strategies for plant breeding field trials: redesigning a CIMMYT international wheat nursery to provide extra genotype connection across cycles. Crop Sci 55:164–177. https://doi.org/10.2135/cropsci2014.06.0415
Arief VN, Desmae H, Hardner C, DeLacy IH, Gilmour A, Bull JK, Basford KE (2019) Utilization of multiyear plant breeding data to better predict genotype performance. Crop Sci 59:480–490. https://doi.org/10.2135/cropsci2018.03.0182
Bajgain P, Zhang X, Anderson JA (2020) Dominance and G×E interaction effects improve genomic prediction and genetic gain in intermediate wheatgrass (Thinopyrum intermedium). Plant Genome 13:e20012. https://doi.org/10.1002/tpg2.20012
Berlin M, Jansson G, Högberg K-A (2014) Genotype by environment interaction in the southern Swedish breeding population of Picea abies using new climatic indices. Scand J for Res 30:112–121. https://doi.org/10.1080/02827581.2014.978889
Bian L, Shi J, Zheng R, Chen J, Wu HX (2014) Genetic parameters and genotype–environment interactions of Chinese fir (Cunninghamia lanceolata) in Fujian Province. Can J for Res 44:582–592. https://doi.org/10.1139/cjfr-2013-0427
Burdon RD (1977) Genetic correlation as concept for studying genotype-environment interactions in forest breeding. Silvae Genet 26(5–6):168–175
Burdon RD, Li Y, Suontama M, Dungey HS (2017) Genotype × site × silviculture interactions in radiata pine: knowledge, working hypotheses and pointers for research. N Z J for Sci 47:6. https://doi.org/10.1186/s40490-017-0087-1
Chen ZQ, Harry BK, Wu HX (2017) Patterns of additive genotype-by-environment interaction in tree height of Norway spruce in southern and central Sweden. Tree Genet Genome 13:25. https://doi.org/10.1007/s11295-017-1103-6
Chen J, Hao Z, Guang X et al (2019) Liriodendron genome sheds light on angiosperm phylogeny and species-pair differentiation. Nature Plants 5:18–25. https://doi.org/10.1038/s41477-018-0323-6
Colombari FJ, de Resende MD, de Morais OP, de Castro AP, Guimarães ÉP, Pereira JA, Utumi MM, Breseghello F (2013) Upland rice breeding in Brazil: a simultaneous genotypic evaluation of stability, adaptability and grain yield. Euphytica 192:117–129. https://doi.org/10.1007/s10681-013-0922-2
Cullis BR, Jefferson P, Thompson R, Smith AB (2014) Factor analytic and reduced animal models for the investigation of additive genotype-by-environment interaction in outcrossing plant species with application to a Pinus radiata breeding programme. Theor Appl Genet 127:2193–2210. https://doi.org/10.1007/s00122-014-2373-0
de Souza BM, Freitas MLM, Sebbenn AM, Gezan SA, Zanatto B, Zulian DF, Lopes MTG, Longui EL, Guerrini IA, de Aguiar AV (2020) Genotype-by-environment interaction in Corymbia citriodora (Hook.) K.D. Hill, & L.A.S. Johnson progeny test in Luiz Antonio, Brazil. For Ecol Manage 460. https://doi.org/10.1016/j.foreco.2019.117855
Des Marais DL, Hernandez KM, Juenger TE (2013) Genotype-by-environment interaction and plasticity: exploring genomic responses of plants to the abiotic environment. Annu Re Ecol Evol Syst 44:5–29. https://doi.org/10.1146/annurev-ecolsys-110512-135806
Diaz SI, de Oliveira HN, Bezerra LAF, Lobo RB (2011) Genotype by environment interaction in Nelore cattle from five Brazilian states. Genet Mol Biol 34:435–442. https://doi.org/10.1590/S1415-47572011005000024
Dieters MJ, White TL, Hodge GR (1995) Genetic parameter estimates for volume from full–sib tests of slash pine (Pines elliottii). Can J for Res 25:1397–1408. https://doi.org/10.1139/x95-152
Eberhart SA, Russell WA (1966) Stability parameters for comparing varieties. Crop Sci 6:36–40
El-Dien OG, Ratcliffe B, Klápště J, Chen C, Porth I, El-Kassaby YA (2015) Prediction accuracies for growth and wood attributes of interior spruce in space using genotyping-by-sequencing. BMC Genomics 16:1–16. https://doi.org/10.1186/s12864-015-1597-y
Evangelista JSPC, Alves RS, Peixoto MA, Resende MDVD, Teodoro PE, Silva FLD, Bhering LL (2021) Soybean productivity, stability, and adaptability through mixed model methodology. Ciên Rural 51(2):e20200406. https://doi.org/10.1590/0103-8478cr20200406
Frutos E, Galindo MP, Leiva V (2014) An interactive biplot implementation in R for modeling genotype-by-environment interaction. Stoch Environ Res Risk Assess 28:1629–1641. https://doi.org/10.1007/s00477-013-0821-z
Gauch HG, Zobel RW (1996) AMMI analysis of yield trials. In: Kan MS, Gauch HG (ed) Genotype-by-environment interaction. CRC Press, Boca Raton, pp 85–122. https://doi.org/10.1201/9781420049374.ch4
Gilmour AR, Gogel BJ, Cullis BR, Thompson R (2008) ASReml user guide release 3.0. VSN International Ltd., Hemel Hempstead, UK. https://doi.org/www.vsni.co.uk
Gu Z, Roland E, Matthias S (2016) Complex heatmaps reveal patterns and correlations in multidimensional genomic data. Bioinformatics 32:2847. https://doi.org/10.1093/bioinformatics/btw313
Haleh H, Anders F, Johan K, Wu HX (2018) Estimation of genetic parameters, provenance performances, and genotype by environment interactions for growth and stiffness in lodgepole pine (Pinus contorta). Scand J for Res 34(5):1–11. https://doi.org/10.1080/02827581.2018.1542025
Hiraoka Y, Miura M, Fukatsu E, Iki T, Yamanobe T, Kurita M, Isoda K, Kubota M, Takahashi M (2019) Time trends of genetic parameters and genetic gains and optimum selection age for growth traits in sugi (Cryptomeria japonica) based on progeny tests conducted throughout Japan. J for Res 24:303–312. https://doi.org/10.1080/13416979.2019.1661068
Holland JB, Nyquist WE, Cervantes-Martinez CT (2010) Plant Breeding Reviews. John Wiley & Sons, Ltd. https://doi.org/10.1002/9780470650202.ch2
Isik F, Holland J, Maltecca C (2017) Genetic data analysis for plant and animal breeding. Springer International Publishing, Switzerland
Kusnandar D, Galwey NW, Hertzler GL, Butcher TB (1998) Age trends in variances and heritabilities for diameter and height in maritime pine (Pinus pinaster AIT.) in Western Australia. Silvae Genet 47(2–3):136–141
Lauer E, Sims A, McKeand S, Isik F (2021) Genetic parameters and genotype-by-environment interactions in regional progeny tests of Pinus taeda L. in the southern USA. For Sci 67(1):60–71. https://doi.org/10.1093/forsci/fxaa035
Li Z, Wang Z (2001) Variation and selection of interspecific hybrid in Liriodendron. J Northwest For Univ 16:5–9. https://doi.org/10.3969/j.issn.1001-7461.2001.02.002
Li H, Chen L, Liang C, Huang M (2005) A case study on provenance testing of tulip tree (Liriodendron spp.). China For Sci Technol 19:13–16. https://doi.org/10.3969/j.issn.1000-8101.2005.05.005
Li Y, Suontama M, Burdon RD, Dungey HS (2017) Genotype by environment interactions in forest tree breeding: review of methodology and perspectives on research and application. Tree Genet Genome 13:1–18. https://doi.org/10.1007/s11295-017-1144-x
Lin Y, Yang H, Ivković M, Gapare WJ, Matheson AC, Wu HX (2013) Effect of genotype by spacing interaction on radiata pine genetic parameters for height and diameter growth. For Ecol Manage 304:204–211. https://doi.org/10.1016/j.foreco.2013.05.015
Liu H, Shen X, Ceng Y (1991) Genetic variations in morphological and growth characteristics of Chinese, American tulip-tree and their hybrid. J Zhejiang For Sci Technol 11(5):18–22
Lu C, Li B, Zheng Y (2015) Analysis on differential expression of cold resistance related genes of Liriodendron chinense under low temperature stress. J Plant Resour Environ 24:25–31. https://doi.org/10.3969/j.issn.1674-7895.2015.03.04
Möhring J, Piepho HP (2009) Comparison of weighting in two-stage analysis in plant breeding trials. Crop Sci 49:1977–1988. https://doi.org/10.2135/cropsci2009.02.0083
Olivoto T, Nardino M (2020) MGIDI: towards an effective multivariate selection in biological experiments. Bioinformatics 37(10):1383–1389. https://doi.org/10.1093/bioinformatics/btaa981
Olivoto T, Lúcio ADC, Jarman S (2020) metan: an R package for multi-environment trial analysis. Methods Ecol Evol 11:783–789. https://doi.org/10.1111/2041-210X.13384
Piepho HP, Möhring J, Schulz-Streeck T, Ogutu JO (2012) A stage-wise approach for the analysis of multi environment trials. Biom J Biom Z 54:844–860. https://doi.org/10.1002/bimj.201100219
Raymond CA (2011) Genotype by environment interactions for Pinus radiata in New South Wales, Australia. Tree Genet Genomes 7:819–833. https://doi.org/10.1007/s11295-011-0376-4
Resende MD (2007) Matemática e estatística na análise de experimentos e no melhoramento genético. Embrapa Florestas, Colombo
Roth BE, Jokela EJ, Martin TA, Huber DA, White TL (2007) Genotype × environment interactions in selected loblolly and slash pine plantations in the southeastern United States. For Ecol Manage 238:175–188. https://doi.org/10.1016/j.foreco.2006.10.010
Rweyongeza DM (2011) Pattern of genotype-environment interaction in Picea glauca (Moench) Voss in Alberta, Canada. Ann for Sci 68:245–253. https://doi.org/10.1007/s13595-011-0032-z
Shukla GK (1972) Some statistical aspects of partitioning genotype-environmental components of variability. Heredity 29:237–245. https://doi.org/10.1038/hdy.1972.87
Sierra-Lucero V, Huber DA, McKeand SE, White TL, Rockwood DL (2003) Genotype-by-environment interaction and development considerations for families from Florida provenances of loblolly pine. For Genet 10(2):85–92
Stackpole DJ, Vaillancourt RE, de Aguigar M, Potts BM (2010) Age trends in genetic parameters for growth and wood density in Eucalyptus globulus. Tree Genet Genome 6:179–193. https://doi.org/10.1007/s11295-009-0239-4
Team R Core (2019) R: a language and environment for statistical computing. R Foundation for Statistical Computing. Vienna, Austria
Wang Z (2003) The review and outlook on hybridization in tulip tree breeding in China. J Nanjing for Univ (natural Sciences Edition) 27:76–78. https://doi.org/10.3969/j.issn.1000-2006.2003.03.019
White TL, Adams WT, Neale DB (2007) Forest genetics. CAB International, Cambridge
Wu HX, Eldridge KG, Matheson AC, Powell MP, McRae TA (2007) Achievement in forest tree improvement in Australia and New Zealand 8. Successful introduction and breeding of radiata pine to Australia. Aust for 70:215–225. https://doi.org/10.1080/00049158.2007.10675023
Wu HX, Ker R, Chen ZQ, Ivkovic M (2021) Balancing breeding for growth and fecundity in radiata pine (Pinus radiata D. Don) breeding programme. Evol Appl 14:834–846. https://doi.org/10.1111/eva.13164
Xia H, Si W, Hao Z, Zhong W, Zhu S, Tu Z, Zhang C, Li H (2021) Dynamic changes in the genetic parameters of growth traits with age and their associations with heterosis in hybrid Liriodendron. Tree Genet Genome 17. https://doi.org/10.1007/s11295-021-01504-z
Yan W (2001) GGEbiplot-a windows application for graphical analysis of multi-environment trial data and other types of two-way data. Agron J 93:1111–1118. https://doi.org/10.2134/agronj2001.9351111x
Yan W, Kang MS, Ma B, Woods S, Cornelius PL (2007) GGE biplot vs. AMMI analysis of genotype-by-environment data. Crop Sci 47:641–653. https://doi.org/10.2135/cropsci2006.06.0374
Yuan C, Zhang Z, Jin G, Zheng Y, Zhou Z, Sun L, Tong H (2021) Genetic parameters and genotype by environment interactions influencing growth and productivity in Masson pine in east and central China. For Ecol Manage 487:118991. https://doi.org/10.1016/j.foreco.2021.118991
Zas R, Merlo E, Díaz R, Fernández-López J (2003) Stability across sites of Douglas-fir provenances in northern Spain. For Genet 10(1):71–82
Zhang W, Hu J, Yang Y, Lin Y (2018) One compound approach combining factor-analytic model with AMMI and GGE biplot to improve multi-environment trials analysis. J for Res 31:123–130. https://doi.org/10.1007/s11676-018-0846-8
Zhang H, Zhou X, Gu W, Wang L, Li W, Gao Y, Wu L, Guo X, Tigabu M, Xia D, Chiang VL, Yang C, Zhao X (2021) Genetic stability of Larix olgensis provenances planted in different sites in northeast China. For Ecol Manage 485:118988. https://doi.org/10.1016/j.foreco.2021.118988
Zhao X, Xia H, Wang X, Wang C, Liang D, Li K, Liu G (2015) Variance and stability analyses of growth characters in half-sib Betula platyphylla families at three different sites in China. Euphytica 208:173–186. https://doi.org/10.1007/s10681-015-1617-7
Zhou M, Chen P, Shang X, Yang W, Fang S (2021) Genotype-environment interactions for tree growth and leaf phytochemical content of Cyclocarya paliurus (Batal.) Iljinskaja. Forests 12 (6):735. https://doi.org/10.3390/f12060735
Acknowledgements
We thank the National Natural Science Foundation of China (31770718, 31470660) and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
Author information
Authors and Affiliations
Contributions
HX, LCY, ZHT, CGZ, ZYH and WPZ participated in collecting and recording raw data. HX, LCY and ZHT arranged and reorganized the data. HX completed the data analysis and wrote the paper. HGL conceived the project; guided the experimental design and progeny tests; gave comments on data analysis; and revised the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Communicated by Tianjian Cao.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xia, H., Yang, L., Tu, Z. et al. Growth performance and G × E interactions of Liriodendron tulipifera half-sib families across ages in eastern China. Eur J Forest Res 141, 1089–1103 (2022). https://doi.org/10.1007/s10342-022-01494-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10342-022-01494-0