Abstract
Chitin, the world’s second most abundant biopolymer after cellulose, is composed of β-1,4-N-acetylglucosamine (GlcNAc) residues. It is the key structural component of many organisms, including crustaceans, mollusks, marine invertebrates, algae, fungi, insects, and nematodes. There has been a significant increase in the generation of chitinous waste from seafood businesses, resulting in a big amount of scrap. Although several organisms, such as plants, crustaceans, insects, nematodes, and animals, produce chitinases, microorganisms are promising candidates and a sustainable option that mediates chitin degradation. Fungi are the dominant group of chitinase producers among microorganisms. In fungi, chitinases are involved in morphogenesis, cell division, autolysis, chitin acquisition for nutritional purposes, and mycoparasitism. Many efficient chitinolytic fungi with potential applications have been identified in a variety of environments, including soil, water, marine wastes, and plants. The current review highlights the key sources of chitinolytic fungi and the characterization of fungal chitinases. It also discusses the applications of fungal chitinases and the cloning of fungal chitinase genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Annually, about 1012–1014 tonnes of chitin are produced in the biosphere (Dhillon et al. 2013), with the shrimp and crab shell wastes being the main source of extraction. Chitin degradation is an important phase in the environmental nutrient cycle process. The majority of these chitinous wastes are processed at ocean dumps, incinerated, or landfilled, resulting in natural resource wastage, economic loss, and environmental pollution. Under high-temperature conditions, chemical conversion of chitinous wastes to chitin requires steps such as demineralization, deproteinization, and deacetylation using strong acids and bases such as NaOH and HCl (Hahn et al. 2022). These methods impact the intrinsic qualities of pure chitin by affecting polymer parameters such as molecular weight, viscosity, purity, and degree of acetylation (Ploydee and Chaiyanan 2014). Furthermore, the purification of chitin by chemical methods is dangerous, energy-intensive, costly, and damaging to human health and ecological systems (Kim and Park 2015). Enzymatic deproteination and microbial-mediated fermentation are two biological approaches for chitin extraction (Kaczmarek et al. 2019). Chitin breakdown is predominantly carried out by microbial chitinases in nature. Chitinase breaks down chitin into oligosaccharides and monosaccharides, which flow into the microbial loop and are absorbed by other species. Microbial chitinases are critical for maintaining a balance between a high amount of carbon and nitrogen trapped as insoluble chitin in the biomass (Poria et al. 2021). Because of their huge abundance, cheap availability of raw material for their cultivation, low cost, variability in catalytic activity, and higher stability, microorganisms are considered the preferable source of chitinases in nature above plants and animals (Thakur et al. 2022). Chitinolytic microorganisms thrive on chitin-rich substrates. Considering the relevance of chitinous biomass bioconversion and the role of chitinases in the development of value-added products, the current review focuses on the diversity of chitinolytic fungi and applications of chitinases in a range of industries such as food, medicines, waste management, and agriculture. The prospect of boosting native catalytic efficacy with a recombinant biotechnological tool has also been addressed.
Chitin: substrate for chitinase production
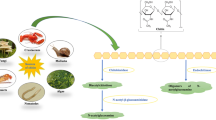
In 1811, Henri Braconnot, who was the director of the Botanical Gardens at the Academy of Sciences in Nancy, France, discovered chitin as an alkaline-insoluble fraction from mushrooms after treating them with dilute warm alkali and named it fongine/fungine (Braconnot 1811). Antonie Odier obtained a comparable alkaline-insoluble fraction from insect cuticles by repeatedly treating them with hot hydroxide solutions in 1823 (Odier 1823). The alkaline-insoluble fraction was given the term chitin which means “tunic” or “envelope.” In 1843, Jean Louis Lassaigne demonstrated the existence of nitrogen in chitin while studying the exoskeleton of the silkworm butterfly Bombyx mori (Lassaigne 1843). Glucosamine and acetic acid were recognized as structural components of chitin by Ledderhose (1876). Gilson (1894) confirmed glucosamine as the repeating unit of chitin. The final chemical composition of chitin was elucidated by Purchase and Braun (1946). Chitin is a nitrogenous polymer that is rigid, inelastic, and hydrophobic, making it insoluble in water and most organic solvents. Chitin is structurally similar to cellulose, but instead of a hydroxyl group at the C-2 position inside the glucose unit, it has an acetamide group (Das et al. 2012). Chitin is found in insects, nematodes, fungal cell walls, mollusks, marine diatoms, and crustacean shells (Merzendorfer and Zimoch 2003; Adams 2004; Furuhashi et al. 2009; Rahman and Halfar 2014; Casadidio et al. 2019). Figure 1 presents the different sources of chitin in nature.
Chitin can be found in three different crystallographic forms α, β, and γ (Rudall 1969). The organization of polypeptide chains in these three polymorphic forms differs. The dominant and more stable form is α-chitin which is composed of antiparallel chitin microfibrils with strong intermolecular hydrogen bonding, whereas the β form is composed of parallel chains with weaker intermolecular forces, resulting in a more flexible and less stable molecule (Gardner and Blackwell 1975). The α-chitin can be found in fungal cell walls (Hassainia et al. 2018), insect cuticles (Sajomsang and Gonil 2010), and crustacean shells (Ifuku et al. 2009). The extracellular spines of the euryhaline diatoms Thalassiosira fluviatilis and Cyclotella cryptica were shown to contain a highly crystalline form of β-chitin (Blackwell et al. 1967). The β-chitin was also extracted from vestimentiferan Tevnia jerichonana (Gaill et al. 1992) and squid pens like Todarodes pacifica and Loligo chenesis (Youn et al. 2013; Cuong et al. 2016). The γ-chitin form has polymeric chains that are arbitrarily organized, with two parallel chains and one antiparallel chain forming the polymeric structure. The cocoon of the Orgyia dubia moth has γ crystallographic structure (Kaya et al. 2017).
Chitinases: a diverse category of enzymes that break down chitin
Bernard (1911) identified chitinase as a thermosensitive and diffusible antifungal component from orchid bulbs and later Karrer and Hofmann (1929) discovered this enzyme in snails. Chitinases (EC 3.2.1.14) are glycosyl hydrolases that may hydrolyze chitin into its oligomeric, dimeric, and monomeric components by cleaving the β-1,4 linkage between GlcNAc. Chitinase has two structural domains: the chitin-binding domain (CBD) for chitin attachment and the chitin-catalytic domain for chitin digestion (Arakane et al. 2003). CBD is essential for improving the efficacy of chitinases in degrading the ubiquitous biopolymer chitin or binding to a chitin-containing external surface (Frederiksen et al. 2013). Accessory domains such as fibronectin type III (FnIII) or cadherin-like domains link these two domains. Chitinases can be divided into two types based on their cleavage patterns: endochitinases and exochitinases. By randomly cleaving β-1,4-glycosidic bonds at internal locations in chitin, endochitinases (EC 3.2.1.14) produce soluble, low molecular weight oligomers. Exochitinases can be divided into two sub-categories. Chitobiosidases (EC 3.2.1.29) catalyze the progressive release of diacetylchitobiose from the non-reducing end of the chitin chain, whereas N-acetyl-glucosaminidases (EC 3.2.1.52) hydrolyzes the chitobiose to its monomeric unit, GlcNAc. Endochitinases are non-processive and have a shallow and open substrate-binding groove, whereas exochitinases are processive and tend to have a tunnel-shaped substrate-binding cleft (Horn et al. 2006; Sikorski et al. 2006). Chitinases mainly belong to four glycoside hydrolase (GH) families, i.e., GH18, GH19, GH23, and GH48. Family 18 chitinases use substrate-assisted catalysis to maintain the product’s anomeric conformation, whereas family 19 chitinases use a general acid–base mechanism to invert the anomeric configuration of the hydrolyzed GlcNAc residue. The glycosidic linkages cleaved by GH48 chitinases use an inverting method of configuration. Chitinases from a wide range of species, including bacteria, fungi, archaea, mammals, plants, and insects, are found in Family 18 (Iseli et al. 1996; Gao et al. 2003; Arakane and Muthukrishnan 2010). Glutamate is used as a catalytic proton donor by chitinases in the GH23 family.
Chitin-binding proteins (CBPs) play a fundamental role in increasing the efficiency of chitinases in the degradation of the ubiquitous biopolymer chitin or to bind to an environmental surface containing chitin (Frederiksen et al. 2013). CBPs refer to a group of proteins that include one or more CBDs, but their binding affinity is not limited to chitin; it may also extend to a variety of complex glycoconjugates that contain GlcNAc or N-acetyl-D-neuraminic acid (NeuNAc) as building blocks (Raikhel et al. 1993). CBPs are categorized into three families CBM14, CBM18, and CBM33 of carbohydrate-binding modules. Families 14 and 18 CBPs are found in insects, fungi, and yeast, whereas family 33 CBPs are found mostly in bacteria (Vaaje-Kolstad et al. 2005, 2009).
Function of chitinases in diverse organisms
In nature, chitinases have a broad spectrum of distribution, including crustaceans (Wang et al. 2015), insects (Qu et al. 2021), nematodes (Dravid et al. 2015), plants (Ali et al. 2020), bacteria (Jenifer et al. 2021), actinobacteria (Brzezinska et al. 2019), fungi (Xie et al. 2021), archaea (Staufenberger et al. 2012; Hanazono et al. 2016), viruses (Rao et al. 2004; Wang et al. 2013), and animals (Hu et al. 2021). Chitinases play a variety of physiological and ecological roles in these organisms. Chitinases are involved in the degradation of the chitinous cuticle during molting in crustaceans, which is necessary for metamorphosis, development, and reproduction as well as digestion of chitin-containing foods and pathogen protection (Huang et al. 2010). During the molting process in insects, chitinases are shown to be involved in the digestion of chitin contained in cuticles and stomach lining (Muthukrishnan et al. 2020). In nematodes, chitinases are involved in several physiological processes, including egg hatching, larval molting, and reproduction (Chen et al. 2021). In plants, chitinases are part of the plant’s defense mechanisms against pathogens (Karasuda et al. 2003). In bacteria, chitinases play an important role in parasitism, nutrition, and the recycling of chitin in nature (Itoh and Kimoto 2019). Human chitinases have been shown to protect against chitin-containing infections by degrading chitin found in pathogen cell walls (Kumar and Zhang 2019).
Chitinases are involved in different aspects of the life cycle of fungi, including (a) cell wall remodeling during spore germination and constriction, hyphal development, branching, and autolysis; (b) degradation of exogenous chitin present in the hyphal cell wall or the exoskeleton of arthropods for nutrition; and (c) competition with and defense against other fungi or arthropods in an aggressive pattern by killing the fungal prey first and then feeding on the dead cell contents (Seidl 2008; Wang et al. 2021).
Production of chitinases from fungi
In nature, fungi are the primary mediators of chitin degradation. Gooday et al. (1992) observed chitinase production in all stages of active growth of filamentous fungi, i.e., during spore germination, exponential growth, and mycelial development. Chitinases are involved in the autolysis associated with the release of spores from fruiting bodies of Coprinus lagopus (Iten and Matile 1970) and Aspergillus nidulans (Reyes et al. 1989). Chitinase also plays an essential role in cell separation during growth in Saccharomyces cerevisiae (Kuranda and Robbins 1991) and Ustilago maydis (Langner et al. 2015). GH18 chitinases are found in fungi; however, Nosema bombycis chitinase was the first GH19 chitinase discovered (Han et al. 2016). Fungal chitinases are classified phylogenetically into three groups: A, B, and C, each with its own domain architecture. Group A, the processive chitinases, contains a catalytic domain with a deep substrate binding site but no carbohydrate-binding modules (CBMs). They have a molecular mass of 40–50 kDa on average. Chitinases from group A are found in all fungal genomes. On the C-terminal of their catalytic domain, group B chitinases have a CBM or a serine/threonine-rich domain. Most non-processive chitinases belong to group B. The size, domain structure, and molecular weights of these chitinases differ. Small group B chitinases (30–45 kDa) that contain CBMs and large proteins (90 kDa) attached to the plasma membrane are two types of group B chitinases. Due to their deep substrate binding site, group C fungal chitinases are also processive, with a CBM 18 on the N-terminal of the catalytic domain. The presence of numerous lysine motifs (LysM) in fungal chitinase C, which are now classed as CBM family 50 in the carbohydrate active enzymes database (CAZy) database, is a distinguishing trait. The molecular mass of group C chitinases is typically 140–170 kDa. Hypocrea atroviridis (Trichoderma atroviride) Chi18-10 was the first chitinase described from group C (Seidl et al. 2005). Wang et al. (2021) speculated that group A chitinases play a major role in the growth and development of species, while group B chitinases are related to the mycoparasitic and entomopathogenic abilities of the fungi, and group C chitinases seem to be correlated with the host range broadening of some plant-pathogenic fungi in Sordariomycetes.
Fungal cell walls play a key role in maintaining mechanic stability during cell division and growth. Chitin is an important scaffolding substance found in fungal cell walls, where it offers mechanical stability (Brown et al. 2019). Polymeric chitin requires ongoing remodeling to preserve its plasticity, which is accomplished by chitinolytic enzymes. Chitinases can play a general role in cell wall plasticization or act more specifically during cell separation, nutritional chitin acquisition, or competitive interactions with other fungi (Langner and Gohre 2016). As the fungal cell wall is made of chitin, it has to protect itself from self-lysis. Gruber and Seidl-Seiboth (2012) hypothesized that the regulation of self and non-self fungal cell wall breakdown is not caused by chitinase speciation but is rather governed by substrate accessibility in healthy hyphae due to cell wall protection vs. deprotection during the mycoparasitic attack, hyphal aging, and autolysis. Fungi produce hydrophobic cell wall proteins like QID74 and carbohydrate-binding proteins to protect their cell walls from hydrolytic enzymes. QID74, a gene from Trichoderma harzianum CECT 2413, encodes a 74 kDa cell wall protein that plays a key function in mycelium protection in addition to its role in adhesion to hydrophobic surfaces (Rosado et al. 2007). Carbohydrate binding proteins bind to short oligosaccharides and chitin, preventing them from being degraded.
Trichoderma is one of the most extensively studied chitinolytic microorganisms. Diverse species of Trichoderma such as Trichoderma harzianum (De Marco et al. 2000; El-Katatny et al. 2001), T. longibrachiatum (Kovacs et al. 2004), T. atroviride (Harighi et al. 2007; Matroudi et al. 2008), T. virens (Ekundayo et al. 2016), T. erinaceum (Herath et al. 2015), T. lixii (Pasqualetti et al. 2019), and T. asperellum (Loc et al. 2020) are efficient chitin degraders. Aspergillus is also prominent chitinolytic fungal genera, including species such as Aspergillus carneus (Sherief et al. 1991), A. terreus (Krishnaveni and Ragunathan 2014; Farag et al. 2016), A. niger, A. fumigatus (Jenin et al. 2016), A. griseoaurantiacus (Shehata et al. 2018), A. flavus (Rawway et al. 2018), and A. niveus (Alves et al. 2018). Other chitinolytic fungi belong to the genera Acremonium (Gunaratna and Balasubramanian 1994), Penicillium (Rodriguez et al. 1995; Atalla et al. 2020), Fusarium (Mathivanan et al. 1998), Alternaria (Sharaf 2005), Chaetomium (Wang and Yang 2009), Basidiobolous (Mishra et al. 2012), Paecilomyces (Homthong et al. 2016), Rhizopus (Yanai et al. 1992; Sonawane et al. 2016), Acremonium (Chung et al. 2019), and Clonostachys (Pasqualetti et al. 2022). Several entomopathogenic fungi (EPF) such as Nomuraea rileyi (Wattanalai et al. 2004), Isaria fumosorosea (Ali et al. 2010), Metarhizium anisopliae (Staats et al. 2013), Beauveria bassiana (Svedese et al. 2013), and Verticillium lecanii (Yu et al. 2015) have been found to produce chitinases. Nematophagous fungi such as Verticillium chlamydosporium, Verticillium suchlasporium (Tikhonov et al. 2002), and Monacrosporium thaumasium (de Freitas Soares et al. 2015) have also been used as biopesticides against nematode eggs and larvae. The biochemical characteristics of fungal chitinases are depicted in Table 1.
Sources of chitinolytic fungi
Chitinolytic fungi have been reported to exist in a variety of environments, including shore soil, marine sediments, and seawater. Tweddell et al. (1994) isolated chitinolytic Stachybotrys elegans from the soil in Izmir, Turkey. S. elegans was shown to have strong antagonistic activity against Rhizoctonia solani. Patidar et al. (2005) isolated Penicillium chrysogenum (PPCS 1 and PPCS 2), Aspergillus flavus PAFS 3, and Aspergillus niger PANS 6 from the chitin-rich soils of shrimp drying fields. Isolate PPCS 1 yielded maximum chitinase (3809 U g−1 initial dry substrate) at pH 5.0, inoculum size 1 × 106 spore ml−1, and 80% initial moisture content. Eight fungal species, viz. Aspergillus flavus, A. niger, A. foetidus, A. ungius, Alternaria alternata, Cladosporium herbarum, Fusarium equisitum, and Dendryphiella vinosa characterized by chitinolytic activity were isolated from Egyptian black sand collected from the Rosetta coast. A. alternata was the most promising species for chitinase production. The crude A. alternata chitinase showed 82% mortality of the larvae of the fruit fly (Sharaf 2005). Another chitinolytic fungus, Lecanicillium psalliotae was isolated from soil samples in Yunnan Province and deposited as the strain code CGMCC1312 in the China General Microbiological Culture Collection Center (Gan et al. 2007). Paecilomyces javanicus, an EPF of coleopteran and lepidopteran insects, was isolated from infected pupae of casuarina (Lymantria xylina) (Chen et al. 2007). Lee et al. (2009) procured chitinolytic fungi Penicillium sp. LYG 0704 from the soil of the crop field at Chonnam National University. The molecular mass of chitinase was estimated to be 47 kDa. Plectosphaerella sp. strain MF-1 was isolated from marine substrates such as calcareous shells, plant materials, and wood blocks. SDS-PAGE revealed that purified chitinase has a molecular mass of 67 kDa. Among the metal ions tested, Ag+, Hg2+, and Pb2+ strongly inhibited enzyme activity, whereas Mg2+ and Fe2+ had minimal inhibition. “DNISQTGEHARYXPMVWFIKL” was found to be the N-terminal amino acid sequence (Velmurugan et al. 2011). Krishnaveni and Ragunathan (2014) isolated Aspergillus terreus CBNRKR KF529976 from marine soils in Pichavaram, Tamil Nadu. The strain successfully biodegraded four different marine wastes, i.e., crab shell, snail shell, shrimp shell, and fish scales, resulting in the production of highly active chitinase. Chitinolytic fungal strains, viz. Paecilomyces sp., Gongronella sp., and Fusarium sp. were isolated from soil in Thailand. Paecilomyces sp. showed higher shrimp shell digestion ability than Gongronella sp. and Fusarium sp. (Homthong et al. 2016).
Jenin et al. (2016) isolated a total of 10 fungi (J1 to J10) from an infected Artemia parthenogenetica sample collected from saltpan of Puthalam, Kanyakumari District, Tamil Nadu. In this study, 5 fungi (J1, J3, J4, J5, and J8) showed chitinolytic activity, and 2 best strains (J1 and J5) were selected for further study. Fungi J1 and J5 were identified as Aspergillus niger and A. fumigatus, respectively. In another study, Trichoderma harzianum, T. viride, and T. hamatum were isolated from soil sampled from agricultural fields and the rhizosphere of plantation crops in the north Gujarat region of India. T. viride was found to be a more promising isolate for the production of chitinase and showed antagonistic activity against fungal phytopathogens such as Aspergillus niger, Fusarium oxysporum, and Sclerotium rolfsii (Khatri et al. 2017). Shehata et al. (2018) isolated Aspergillus griseoaurantiacus KX010988 from algae and decayed salty wood samples collected from Egypt’s Port Said Governorate. The chitinase activity was enhanced by Mn2+ and Zn2+ ions, while Fe2+ and Cu2+ ions strongly inhibited the chitinase activity. Chitinase also showed antifungal activity against the pathogenic fungus Fusarium solani. Pasqualetti et al. (2019) screened twenty-eight fungal strains for the production of chitinolytic activity isolated from different natural marine substrates. The best chitinase producers were the halotolerant marine fungus Trichoderma lixii IG127 and the halophilic marine fungus Clonostachys rosea IG119. Acremonium sp. YS2-2, a chitinolytic marine-derived fungal strain, was isolated from seawater. The chitinolytic activity of the extracellular crude enzyme of YS2-2 was highest at pH 6.0–7.6 and a temperature of 23–45 °C (Chung et al. 2019). In another study, Atalla et al. (2020) isolated the chitinolytic marine fungus Penicillium chrysogenum MH745129 from red seawater. The optimum reaction mixture conditions for partially purified chitinase activity were obtained using 60% acetone at 40 °C, pH 6.0 at 40 min, and it was stable at 50 °C for 60 min. The molecular weight of chitinase was discovered to be 42 kDa. The linear mycelial development of both Penicillium digitatum and P. italicum was significantly reduced by chitinase. Twenty fungal strains were isolated from the tomato rhizosphere of Senegal. Among these 20 strains, Trichoderma asperellum TG4 showed the maximum chitinase activity at a temperature of 30 °C and a pH of 6.0 (Gueye et al. 2020).
Applications of fungal chitinases
The non-toxicity, biodegradability, biocompatibility, antimicrobial, and antiallergenic properties of chitin make it useful for a wide range of applications. The applications of fungal chitinases include biocontrol of phytopathogens and pests, generation of protoplasts, production of single cell proteins (SCPs), and production of chitooligosaccharides (COS) (Fig. 2).
Biocontrol of phytopathogens
Agricultural production has increased in recent decades, and farmers have become increasingly reliant on chemical fertilizers as a reliable form of crop protection approach. The increased use of chemical inputs, on the other hand, has presented a huge danger to food production and ecological stability around the world. Biological control is a potent and viable alternative to chemical control in agriculture. Mycolytic enzymes like chitinase, which operate as bioshields against phytopathogens, are produced by microbial communities. Chitinolytic fungi have been reported to affect fungal growth through the lysis of the cell wall (Tominaga and Tsujisaka 1976) and germ tube (Gunaratna and Balasubramanian 1994). Table 2 depicts the variety of chitinolytic fungi that have been reported as potential biocontrol agents against several plant pathogens.
Biocontrol of pests
Inappropriate use of pesticides can have detrimental repercussions for the environment and human health. There has been an upsurge in demand for biopesticides in agriculture due to the lower risk to human health and minimal detrimental impact on the environment (Kumar et al. 2021). Chitin is present in the cuticles of the epidermis, the trachea, and peritrophic matrices lining the gut epithelium of insects and the eggshell of nematodes. Fungal chitinases enzymatically split the chitin and cause the perforations that lead to illness and the death of the larvae. Elegant demonstrations of the involvement of chitinolytic fungi as an alternative to chemical pesticides are presented in Table 3.
Formation of protoplasts
Protoplasts are plasma membrane enclosed naked cells without a cell wall. The cell wall is removed mechanically or enzymatically to produce protoplasts. The formation of protoplasts from Mulbranchea sulfurea IMI 337,352 mycelia using chitinase and laminarinase produced by Paecilomyces varioti IMI 334,593 was reported by Gautam et al. (1996). They also reported a 34.6% frequency of protoplast regeneration. Using a chitinolytic enzyme produced by Enterobacter sp. NRG4, Trichoderma reesei, Pleurotus florida, Agaricus bisporus, and Aspergillus niger were reported to release protoplast (Dahiya et al. 2005). A purified 61 kDa chitinase from Cellulosimicrobium cellulans strain 191 was capable of protoplast formation from Rhizopus oligosporus and Penicillium sp. (Fleuri et al. 2009). With a 60% regeneration capacity, Penicillium ochrochloron MTCC 517 chitinase was extremely effective in the formation of protoplasts from Aspergillus niger (Patil et al. 2013). Sonawane et al. (2016) found that protoplasts of Aspergillus niger, A. oryzae, Trichoderma viride, and Fusarium moniliforme were efficiently produced using crude chitinase preparation from Rhizopus stolonifer NCIM 880. The protoplasts generated by A. niger and T. viride, respectively, showed 70 and 66% regeneration frequencies.
Production of single cell proteins
Production of single cell proteins (SCPs) from chitinous waste is a viable option for biomass production where chitinous waste is employed as a carbon and nutritional source. Fungal chitinase has the potential for producing SCPs, which might be used as a cheaper protein source. The bioconversion of chitin to SCP from shellfish waste by Pichia kudriuvzevii chitinolytic enzyme has been reported by Revah-Moiseev and Carroad (1981). Vyas and Deshpande (1991) reported chitin enzymatic hydrolysis with the culture filtrate of Myrothecium verrucaria NCIM 903. Further, they reported the utilization of chitin hydrolysate by Saccharomyces cerevisiae NCIM 3052 for the production of SCP. SCP was found to contain 9.5 g/l biomass with 61% of total protein and 3.1% of nucleic acids. In another study, Patil and Jadhav (2014) reported that the chitinolytic enzyme of Penicillium ochrochloron degraded chitin to GlcNAc which was then employed as a substrate for SCP production by Yerrowia lipolytica NCIM 3450. SCP contained 65% total protein and 2.9% nucleic acid content with a biomass of 9.4 g/l.
Chitooligosaccharides production
Chitooligosaccharides (COS) are produced by the degradation of either chitin or chitosan via physical hydrolysis, acid hydrolysis, and enzymatic hydrolysis. COS has been the center of attention in the pharmaceutical, food, and agriculture sectors, mainly because of its higher water solubility, low viscosity, low molecular weight, biocompatibility, biodegradability, and nontoxicity. COS interacts readily with various cell receptors in living organisms, resulting in anticancer (Luo et al. 2016), antibacterial (Choi et al. 2001), antifungal (Mei et al. 2015), anti-HIV-1 (Artan et al. 2010), antihypertensive (Hong et al. 1998), antihyperlipidemic (Jiang et al. 2018), and antiobesity effects (Choi et al. 2012). COS also acts as a therapeutic agent against oxidative stress (Qu and Han 2016), diabetes (Ju et al. 2010), and inflammation (Kunanusornchai et al. 2016). COS also acts as an excellent preservative in food and enhances its shelf life. The use of COS inhibited Lactobacillus brevis contamination in the brewing process of beer, acting as an excellent preservative (Zhao et al. 2016). COS is also reported to stimulate several defense responses in plants (Lan et al. 2016).
Molecular cloning and expression of chitinase genes
The use of recombinant DNA (rDNA) technology as a tool for the development of genetically engineered microbial strains with selected characteristics for enzyme production is essential. Recombinant chitinases have shown greater characteristics than the native enzymes, which can be employed in the fermentation industry. Chitinase genes from a variety of fungal strains have been cloned and produced in suitable hosts for a variety of purposes. A chitinase gene, Lpchi1, was cloned from Lecanicillium psalliotae in Pichia pastoris GS115. P. pastoris expressed a recombinant chitinase of 45 kDa with optimal activity at a pH of 7.0 and a temperature of 37.6 °C. The purified chitinase also degraded chitinous components of the eggs of Meloidogyne incognita (Gan et al. 2007). Ramli et al. (2011) cloned a gene encoding a cold-adapted chitinase gene (CHI II) from Glaciozyma antarctica PI12 in P. pastoris GS115. The strain showed a strong affinity for colloidal chitin and little effect on glycol chitosan. The molecular weights of the purified recombinant chitinase were approximately 39 and 50 kDa, respectively. The enzyme was stable between pH 3.0–4.5 and could maintain its activity at temperatures ranging from 5 to 25 °C, with an optimal pH of 4.0 and a temperature of 15 °C. Prasad and Palanivelu (2012) cloned a chitinase gene from Thermomyces lanuginosus ATCC 44,008 in Saccharomyces cerevisiae SEY 2101. The molecular mass of the recombinant chitinase was 42 kDa, with optimum activity at a pH of 6.5 and a temperature of 60 °C. The recombinant chitinase showed excellent thermostability, retaining more than 60% of enzyme activity after 6 h at 50 °C. The cloning of fungal chitinase genes and their expression in microbial strains are demonstrated in Table 4.
Conclusion
Since chitinous waste is produced in large quantities, its valorization is essential to prevent environmental pollution. In this context, chitinolytic enzymes are critical toolboxes for chitin waste management and the generation of value-added products. Several organisms, including microorganisms, crustaceans, plants, insects, and animals, produce chitinases that mediate chitin degradation. Chitinases have a wide range of applications; hence, finding potent chitinases is critical for dealing with the growing problem of marine trash loaded with chitin compounds. Continued research into chitin breakdown will necessitate the development of novel enzymes with better activity and high-throughput chitinase tests. Further research is required to improve the market demand for chitinolytic enzymes.
References
Adams DJ (2004) Fungal cell wall chitinases and glucanases. Microbiol 150:2029–2035. https://doi.org/10.1099/mic.0.26980-0
Al Abboud MA, Al-Rajhi AMH, Shater ARM, Alawlaqi MM, Mashraqi A, Selim S, Al Jaouni SK, Abdel Ghany TM (2022) Halostability and thermostability of chitinase produced by fungi isolated from salt marsh soil in subtropical region of Saudi Arabia. BioResources 17:4763–4780. https://doi.org/10.15376/biores.17.3.4763-4780
Ali M, Li QH, Zou T, Wei AM, Gombojav G, Lu G, Gong ZH (2020) Chitinase gene positively regulates hypersensitive and defense responses of pepper to Colletotrichum acutatum infection. Int J Mol Sci. https://doi.org/10.3390/ijms21186624
Ali S, Wu J, Huang Z, Ren SX (2010) Production and regulation of extracellular chitinase from the entomopathogenic fungus Isaria fumosorosea. Biocontrol Sci Technol 20:723–738. https://doi.org/10.1080/09583151003714091
Al-Rashed SAA, Bakar FDA, Said M, Hassan O, Rabu A, Illias RM, Murad AMA (2010) Expression and characterization of the recombinant Trichoderma virens endochitinase Cht2. Afr J Microbiol Res 4:1758–1767
Alves TB, de Oliveira Ornela PH, de Oliveira AHC, Jorge JA, Guimaraes LHS (2018) Production and characterization of a thermostable antifungal chitinase secreted by the filamentous fungus Aspergillus niveus under submerged fermentation. 3 Biotech. https://doi.org/10.1007/s13205-018-1397-6
Arakane Y, Muthukrishnan S (2010) Insect chitinase and chitinase-like proteins. Cell Mol Life Sci 67:201–216. https://doi.org/10.1007/s00018-009-0161-9
Arakane Y, Zhu Q, Matsumiya M, Muthukrishnan S, Kramer KJ (2003) Properties of catalytic, linker and chitin-binding domains of insect chitinase. Insect Biochem Mol Biol 33:631–648. https://doi.org/10.1016/s0965-1748(03)00049-3
Artan M, Karadeniz F, Karagozlu MZ, Kim MM, Kim SK (2010) Anti-HIV-1 activity of low molecular weight sulfated chitooligosaccharides. Carbohydr Res 345:656–662. https://doi.org/10.1016/j.carres.2009.12.017
Atalla SMM, Gamal EL, NG, Awad HM, (2020) Chitinase of marine Penicillium chrysogenum MH745129: isolation, identification, production and characterization as controller for citrus fruits postharvest pathogens. Jordan J Biol Sci 13:19–28
Awad GEA, Abdel Wahab WA, Hussein M, El-Diwany A, Esawy MA (2017) Sequential optimizations of Aspergillus awamori EM66 exochitinase and its application as biopesticide. J App Pharm Sci 7:67–75. https://doi.org/10.7324/JAPS.2017.70208
Baratto CM, da Silva MV, Santi L, Passaglia L, Schrank IS, Vainstein MH, Schrank A (2003) Expression and characterization of the 42 kDa chitinase of the biocontrol fungus Metarhizium anisopliae in Escherichia coli. Can J Microbiol. https://doi.org/10.1139/w03-085
Beltagy EA, Rawway M, Abdul-Raouf UM, Elshenawy MA, Kelany MS (2018) Purification and characterization of theromohalophilic chitinase producing by halophilic Aspergillus flavus isolated from Suez Gulf. Egypt J Aquat Res 44:227-232. https://doi.org/10.1016/j.ejar.2018.08.002
Berini F, Caccia S, Franzetti E, Congiu T, Marinelli F, Casartelli M, Tettamanti G (2016) Effects of Trichoderma viride chitinases on the peritrophic matrix of Lepidoptera. Pest Manag Sci 72:980–989. https://doi.org/10.1002/ps.4078
Bernard N (1911) Sur la fonction fungicide des bulbes d’Ophrydeae. Ann Sci Nat Bot Biol 14:221–234
Binod P, Sukumaran RK, Shirke SV, Rajput JC, Pandey A (2007) Evaluation of fungal culture filtrate containing chitinase as a biocontrol agent against Helicoverpa armigera. J Appl Microbiol 103:1845–1852. https://doi.org/10.1111/j.1365-2672.2007.03428.x
Blackwell J, Parker KD, Rudall KM (1967) Chitin fibers of the diatoms Thalassiosira fluviatilis and Cyclotella cryptica. J Mol Biol 28:383–385. https://doi.org/10.1016/s0022-2836(67)80018-4
Braconnot H (1811) De la fongine, ou analyse des champignons. Journal de Physique, de Chimie, d’Histoire Naturelle et des Arts. Paris: chez Courcier, Imprimeur-Libraire pour les mathématiques, tome LXXIII 130–135
Brown HE, Esher SK, Alspaugh JA (2019) Chitin: a "hidden figure" in the fungal cell wall. In: Latge JP (ed) The fungal cell wall: an armour and a weapon for human fungal pathogens. 425. Springer, Switzerland 83–111
Brzezinska MS, Jankiewicz U (2012) Production of antifungal chitinase by Aspergillus niger LOCK 62 and its potential role in the biological control. Curr Microbiol 65:666–672. https://doi.org/10.1007/s00284-012-0208-2
Brzezinska MS, Jankiewicz U, Kalwasinska A, Swiatczak J, Zero K (2019) Characterization of chitinase from Streptomyces luridiscabiei U05 and its antagonist potential against fungal plant pathogens. J Phytopathol 167:404–412. https://doi.org/10.1111/jph.12809
Carsolio C, Gutierrez A, Jimenez B, Van Montagu M, Herrera-Estrella A (1994) Characterization of ech-42, a Trichoderma harzianum endochitinase gene expressed during mycoparasitism. Proc Natl Acad Sci USA 91:10903–10907. https://doi.org/10.1073/pnas.91.23.10903
Casadidio C, Peregrina DV, Gigliobianco MR, Deng S, Censi R, Di Martino P (2019) Chitin and chitosans: characteristics, eco-friendly processes, and applications in cosmetic science. Mar Drugs 17:369. https://doi.org/10.3390/md17060369
Chen CC, Kumar HGA, Kumar S, Tzean SS, Yeh KW (2007) Molecular cloning, characterization, and expression of a chitinase from the entomopathogenic fungus Paecilomyces javanicus. Curr Microbiol 55:8–13. https://doi.org/10.1007/s00284-006-0405-y
Chen W, Chen Q, Kumar A, Jiang X, Zhang KYJ, Yang Q (2021) Structure-based virtual screening of highly potent inhibitors of the nematode chitinase CeCht1. J Enzyme Inhib Med Chem 36:1198–1204. https://doi.org/10.1080/14756366.2021.1931862
Choi BK, Kim KY, Yoo YJ, Oh SJ, Choi JH, Kim CY (2001) In vitro antimicrobial activity of a chitooligosaccharide mixture against Actinobacillus actinomycetemcomitans and Streptococcus mutans. Int J Antimicrob Agents 18:553–557. https://doi.org/10.1016/s0924-8579(01)00434-4
Choi EH, Yang HP, Chun HS (2012) Chitooligosaccharide ameliorates diet-induced obesity in mice and affects adipose gene expression involved in adipogenesis and inflammation. Nutr Res 32:218–228. https://doi.org/10.1016/j.nutres.2012.02.004
Chung D, Baek K, Bae SS, Jung J (2019) Identification and characterization of a marine-derived chitinolytic fungus, Acremonium sp. YS2-2. J Microbiol 57:1–9. https://doi.org/10.1007/s12275-019-8469-0
Chung D, Kwon YM, Lim JY, Bae SS, Choi G, Lee DS (2022) Characterization of chitinolytic and antifungal activities in marine-derived Trichoderma bissettii strains. Mycobiol 50:244–253. https://doi.org/10.1080/12298093.2022.2105509
Cuong HN, Minh NC, Hoa NV, Trung TS (2016) Preparation and characterization of high purity β-chitin from squid pens (Loligo chenisis). Int J Biol Macromol 93:442–447. https://doi.org/10.1016/j.ijbiomac.2016.08.085
Dahiya N, Tewari R, Tiwari RP, Hoondal GS (2005) Production of an antifungal chitinase from Enerobacter sp. NRG4 and its application in protoplast production. World J Microbiol Biotechnol 21:1611–1616. https://doi.org/10.1007/s11274-005-8343-6
Das SN, Neeraja C, Sharma PVSRN, Prakash JM, Purushotham P, Kaur M, Dutta S, Podile AR (2012) Microbial chitinases for chitin waste management. In: Satyanarayana T, Johri BN, Prakash A (eds) Microorganisms in environmental management: microbes and environment. Springer, Dordrecht 135–150
de Freitas Soares FE, de Queiroz JH, de Araujo JV, Queiroz PV, de Souza GA, Hiura E, Braga FR (2015) Nematicidal action of chitinases produced by the fungus Monacrosporium thaumasium under laboratorial conditions. Biocontrol Sci Technol 25:337–344. https://doi.org/10.1080/09583157.2014.979133
De Marco JL, Lima LHC, de Sousa MV, Felix CR (2000) A Trichoderma harzianum chitinase destroys the cell wall of the phytopathogen Crinipellis perniciosa, the causal agent of witches’ broom disease of cocoa. World J Microbiol Biotechnol 16:383–386. https://doi.org/10.1023/A:1008964324425
Deng JJ, Shi D, Mao HH, Li ZW, Liang S, Ke Y, Luo XC (2019) Heterologous expression and characterization of an antifungal chitinase (Chit46) from Trichoderma harzianum GIM 3.442 and its application in colloidal chitin conversion. Int J Biol Macromol 134:113–121. https://doi.org/10.1016/j.ijbiomac.2019.04.177
Dhillon GS, Kaur S, Brar SK, Verma M (2013) Green synthesis approach: extraction of chitosan from fungus mycelia. Crit Rev Biotechnol 33:379–403. https://doi.org/10.3109/07388551.2012.717217
Draborg H, Christgau S, Halkier T, Rasmussen G, Dalboge H, Kauppinen S (1996) Secretion of an enzymatically active Trichoderma harzianum endochitinase by Saccharomyces cerevisiae. Curr Genet 29:404–409. https://doi.org/10.1007/BF02208622
Dravid P, Kaushal DC, Saxena JK, Kaushal NA (2015) Isolation and characterization of endochitinase and exochitinase of Setaria cervi. Parasitol Int 64:579–586. https://doi.org/10.1016/j.parint.2015.08.007
Ekundayo EA, Ekundayo FO, Bamidele F (2016) Production, partial purification and optimization of a chitinase produced from Trichoderma viride, an isolate of maize cob. Mycosphere 7:786–793. https://doi.org/10.5943/mycosphere/7/6/9
El-Katatny MH, Gudelj M, Robra KH, Elnaghy MA, Gubitz GM (2001) Characterization of a chitinase and an endo-β-1,3-glucanase from Trichoderma harzianum Rifai T24 involved in control of the phytopathogen Sclerotium rolfsii. Appl Microbiol Biotechnol 56:137–143. https://doi.org/10.1007/s002530100646
Farag AM, Abd-Elnabey HM, Ibrahim HAH, El-Shenawy M (2016) Purification, characterization and antimicrobial activity of chitinase from marine-derived Aspergillus terreus. Egyptian J Aquat Res 42:185–192. https://doi.org/10.1016/j.ejar.2016.04.004
Fenice M, Selbmann L, Di Giambattista R, Federici F (1998) Chitinolytic activity at low temperature of an Antarctic strain (A3) of Verticillium lecanii. Res Micrbiol 149:289–300. https://doi.org/10.1016/s0923-2508(98)80304-5
Fleuri LF, Kawaguti HY, Sato HH (2009) Production, purification and application of extracellular chitinase from Cellulosimicrobium cellulans 191. Braz J Microbiol 40:623–630. https://doi.org/10.1590/S1517-838220090003000026
Frederiksen RF, Paspaliari DK, Larsen T, Storgaard BG, Larsen MH, Ingmer H, Palcic MM, Leisner JJ (2013) Bacterial chitinases and chitin-binding proteins as virulence factors. Microbiol 159:833–847. https://doi.org/10.1099/mic.0.051839-0
Furuhashi T, Schwarzinger C, Miksik I, Smrz M, Beran A (2009) Molluscan shell evolution with review of shell calcification hypothesis. Comp Biochem Physiol Part B 154:351–371. https://doi.org/10.1016/j.cbpb.2009.07.011
Gaill F, Persson J, Sugiyama J, Vuong R, Chanzy H (1992) The chitin system in the tubes of deep sea hydrothermal vent worms. J Struct Biol 109:116–128. https://doi.org/10.1016/1047-8477(92)90043-a
Gan Z, Yang J, Tao N, Liang L, Mi Q, Li J, Zhang KQ (2007) Cloning of the gene Lecanicillium psalliotae chitinase Lpchi1 and identification of its potential role in the biocontrol of root-knot nematode Meloidogyne incognita. Appl Microbiol Biotechnol 76:1309–1317. https://doi.org/10.1007/s00253-007-1111-9
Gao J, Bauer MW, Shockley KR, Pysz MA, Kelly RM (2003) Growth of hyperthermophilic archaeon Pyrococcus furiosus on chitin involves two family 18 chitinases. Appl Environ Microbiol 69:3119–3128. https://doi.org/10.1128/AEM.69.6.3119-3128.2003
Gardner KH, Blackwell J (1975) Refinement of the structure of β-chitin. Biopolymers 14:1581–1595. https://doi.org/10.1002/bip.1975.360140804
Gautam SP, Gupta AK, Shrivastava R, Awasthi M (1996) Protoplast formation from the thermophilic fungus Malbranchea sulfurea, using the thermostable chitinase and laminarinase of Paecilomyces varioti. World J Microbiol Biotechnol 12:99–100. https://doi.org/10.1007/BF00327811
Gilson E (1894) Recherches chimiques sur la membrane cellulaire des champignons. Bull Soc Chim France 3:1099–1102
Gooday GW, Zhu WY, O’Donnell RW (1992) What are the roles of chitinases in the growing fungus? FEMS Microbiol Lett 100:387–392
Govindsamy V, Gunaratna KR, Balasubramanian R (1998) Properties of extracellular chitinase from Myrothecium verrucaria, an antagonist to the groundnut rust Puccinia arachidis. Can J Plant Pathol 20:62–68. https://doi.org/10.1080/07060669809500446
Gruber S, Seidl-Seiboth V (2012) Self versus non-self: fungal cell wall degradation in Trichoderma. Microbiol 158:26–34. https://doi.org/10.1099/mic.0.052613-0
Gueye N, Kumar GK, Ndiaye M, Sall SYD, Ndiaye MAF, Diop TA, Ram MR (2020) Factors affecting the chitinase activity of Trichoderma asperellum isolated from agriculture field soils. J App Biol Biotech 8:41–44. https://doi.org/10.7324/JABB.2020.80207
Gunaratna KR, Balasubramanian R (1994) Partial purification and properties of extracellular chitinase produced by Acremonium obclavatum, an antagonist to the groundnut rust, Puccinia arachidis. World J Microbiol Biotechnol 10:342–345. https://doi.org/10.1007/BF00414876
Hahn T, Tafi E, von Seggern N, Falabella P, Salvia R, Thoma J, Febel E, Fijalkowska M, Schmitt E, Stegbauer L, Zibek S (2022) Purification of chitin from pupal exuviae of the black soldier fly. Waste Biomass Valor 13:1993–2008. https://doi.org/10.1007/s12649-021-01645-1
Han B, Zhou K, Li Z, Sun B, Ni Q, Meng X, Pan G, Li C, Long M, Li T, Zhou C, Li W, Zhou Z (2016) Characterization of the first fungal glycosyl hydrolase family 19 chitinase (NbchiA) from Nosema bombycis (Nb). J Eukaryot Microbiol 63:37–45. https://doi.org/10.1111/jeu.12246
Hanazono Y, Takeda K, Niwa S, Hibi M, Takahashi N, Kanai T, Atomi H, Miki K (2016) Crystal structures of chitin binding domains of chitinase from Thermococcus kodakarensis KOD1. FEBS Lett 590:298–304. https://doi.org/10.1002/1873-3468.12055
Harighi MJ, Zamani MR, Motallebi M (2007) Evaluation of antifungal activity of purified chitinase 42 from Trichoderma atroviride PTCC5220. Biotechnol 6:28–33. https://doi.org/10.3923/biotech.2007.28.33
Hassainia A, Satha H, Boufi S (2018) Chitin from Agaricus bisporus: extraction and characterization. Int J Biol Macromol 117:1334–1342. https://doi.org/10.1016/j.ijbiomac.2017.11.172
He B, Yang L, Yang D, Jiang M, Ling C, Chen H, Ji F, Pan L (2022) Biochemical purification and characterization of a truncated acidic, thermostable chitinase from marine fungus for N-acetylglucosamine production. Front Bioeng Biotechnol 10:1013313. https://doi.org/10.3389/fbioe.2022.1013313
Herath HHMAU, Wijesundera RLC, Chandrasekharan NV, Wijesundera WSS, Kathriarachchi HS (2015) Isolation and characterization of Trichoderma erinaceum for antagonistic activity against plant pathogenic fungi. CREAM 5:120–127. https://doi.org/10.5943/cream/5/2
Homthong M, Kubera A, Srihuttagum M, Hongtrakul V (2016) Isolation and characterization of chitinase from soil fungi, Paecilomyces sp. Agric Nat Resour 50:232–242. https://doi.org/10.1016/j.anres.2015.09.005
Hong SP, Kim MH, Oh SW, Han CK, Kim YH (1998) ACE inhibitory and antihypertensive effect of chitosan oligosaccharides in SHR. Korean J Food Sci Technol 30:1476–1479
Horn SJ, Sorlie M, Vaaje-Kolstad G, Norberg AL, Synstad B, Varum KM, Eijsink VGH (2006) Comparative studies of chitinases A, B and C from Serratia marcescens. Biocatal Biotransfor 24:39–53. https://doi.org/10.1080/10242420500518482
Hu C, Ma Z, Zhu J, Fan Y, Tuo B, Li T, Liu X (2021) Physiological and pathophysiological roles of acidic mammalian chitinase (CHIA) in multiple organs. Biomed Pharmacother 138:111465. https://doi.org/10.1016/j.biopha.2021.111465
Huang QS, Yan JH, Tang JY, Tao YM, Xie XL, Wang Y, Wei XQ, Yan QH, Chen QX (2010) Cloning and tissue expressions of seven chitinase family genes in Litopenaeus vannamei. Fish Shellfish Immunol 29:75–81. https://doi.org/10.1016/j.fsi.2010.02.014
Ifuku S, Nogi M, Abe K, Yoshioka M, Morimoto M, Saimoto H, Yano H (2009) Preparation of chitin nanofibers with a uniform width as α-chitin from crab shells. Biomacromol 10:1584–1588. https://doi.org/10.1021/bm900163d
Iseli B, Armand S, Boller T, Neuhaus JM, Henrissat B (1996) Plant chitinases use two different hydrolytic mechanisms. FEBS Lett 382:186–188. https://doi.org/10.1016/0014-5793(96)00174-3
Iten W, Matile P (1970) Role of chitinase and other lysosomal enzymes of Coprinus zagopus in the autolysis of fruiting bodies. J Gen Microbiol 61:301–309
Itoh T, Kimoto H (2019) Bacterial chitinase system as a model of chitin biodegradation. In: Yang Q, Fukamizo T (eds) Targeting chitin-containing organisms. Springer, Singapore 131–151
Jenifer SG, Marimuthu G, Raghuram H (2021) Isolation and characterization of chitinolytic bacterium, Escherichia fergusonii AMC01 from insectivorous bat, Taphozous melanopogon. J Basic Microbiol 61:940–946. https://doi.org/10.1002/jobm.202100271
Jenin GA, Babu MM, Murugan M, Murugan T (2016) Isolation and identification of chitinase producing native fungi from saltpan of Puthalam, Kanyakumari District, Tamil Nadu, India. J Appl Biol Biotechnol 4:1–5. https://doi.org/10.7324/JABB.2016.40301
Jiang Y, Fu C, Liu G, Guo J, Su Z (2018) Cholesterol-lowering effects and potential mechanisms of chitooligosaccharide capsules in hyperlipidemic rats. Food Nutr Res. https://doi.org/10.29219/fnr.v62.1446
Ju C, Yue W, Yang Z, Zhang Q, Yang X, Liu Z, Zhang F (2010) Antidiabetic effect and mechanism of chitooligosaccharides. Biol Pharm Bull 33:1511–1516. https://doi.org/10.1248/bpb.33.1511
Kaczmarek MB, Struszczyk-Swita K, Li X, Szczesna-Antczak M, Daroch M (2019) Enzymatic modifications of chitin, chitosan, and chitooligosaccharides. Front Bioeng Biotechnol 7:243. https://doi.org/10.3389/fbioe.2019.00243
Karasuda S, Tanaka S, Kajihara H, Yamamoto Y, Koga D (2003) Plant chitinase as a possible biocontrol agent for use instead of chemical fungicides. Biosci Biotechnol Biochem 67:221–224. https://doi.org/10.1271/bbb.67.221
Karrer P, Hofmann A (1929) Uber den enzymatischen Abbau von chitin und chitosan. Helv Chim Acta 12:616–637
Kaya M, Mujtaba M, Ehrlich H, Salaberria AM, Baran T, Amemiya CT, Galli R, Akyuz L, Sargin I, Labidi J (2017) On chemistry of γ-chitin. Carbohydr Polym 176:177–186. https://doi.org/10.1016/j.carbpol.2017.08.076
Khan A, Williams KL, Nevalainen HKM (2004) Effects of Paecilomyces lilacinus protease and chitinase on the eggshell structures and hatching of Meloidogyne javanica juveniles. Biol Control 31:346–352. https://doi.org/10.1016/j.biocontrol.2004.07.011
Khatri DK, Tiwari DN, Bariya HS (2017) Chitinolytic efficacy and secretion of cell wall-degrading enzymes from Trichoderma spp. in response to phytopathological fungi. J Appl Biol Biotechnol 5:1–8. https://doi.org/10.7324/JABB.2017.50601
Kim Y, Park RD (2015) Progress in bioextraction processes of chitin from crustacean biowastes. J Korean Soc Appl Biol Chem 58:545–554. https://doi.org/10.1007/s13765-015-0080-4
Kovacs K, Szakacs G, Pusztahelyi T, Pandey A (2004) Production of chitinolytic enzymes with Trichoderma longibrachiatum IMI 92027 in solid substrate fermentation. Appl Biochem Biotechnol 118:189–204. https://doi.org/10.1385/abab:118:1-3:189
Krishnaveni B, Ragunathan R (2014) Chitinase production from marine wastes by Aspergillus terreus and its application in degradation studies. Int J Curr Microbiol App Sci 3:76–82
Kumar A, Zhang KYJ (2019) Human chitinases: structure, function, and inhibitor discovery. In: Yang Q, Fukamizo T (eds) Targeting chitin-containing organisms. Springer, Singapore 221–251
Kumar J, Ramlal A, Mallick D, Mishra V (2021) An overview of some biopesticides and their importance in plant protection for commercial acceptance. Plants 10:1185. https://doi.org/10.3390/plants10061185
Kumar M, Brar A, Vivekanand V, Pareek N (2018) Process optimization, purification and characterization of a novel acidic, thermostable chitinase from Humicola grisea. Int J Biol Macromol. https://doi.org/10.1016/j.ijbiomac.2018.05.125
Kunanusornchai W, Witoonpanich B, Pichyangkura R, Chatsudthipong V, Muanprasat C (2016) Chitosan oligosaccharide suppresses synovial inflammation via AMPK activation: an in vitro and in vivo study. Pharmacol Res 113:458–467. https://doi.org/10.1016/j.phrs.2016.09.016
Kuranda MJ, Robbins PW (1991) Chitinase is required for cell separation during growth of Saccharomyces cerevisiae. J Biol Chem 266:19758–19767
Lan W, Wang W, Yu Z, Qin Y, Luan J, Li X (2016) Enhanced germination of barley (Hordeum vulgare L.) using chitooligosaccharide as an elicitor in seed priming to improve malt quality. Biotechnol Lett 38:1935–1940. https://doi.org/10.1007/s10529-016-2181-5
Langner T, Gohre V (2016) Fungal chitinases: function, regulation, and potential roles in plant/pathogen interactions. Curr Genet 62:243–254. https://doi.org/10.1007/s00294-015-0530-x
Langner T, Ozturk M, Hartmann S, Cord-Landwehr S, Moerschbacher B, Walton JD, Gohre V (2015) Chitinases are essential for cell separation in Ustilago maydis. Eukaryot Cell 14:846–857. https://doi.org/10.1128/EC.00022-15
Lassaigne JL (1843) Mémoire sur un procédé simple pour constater la présence d’azote dans des quantités minimes de matière organique. Compte Rendus Des Séances Hebdomadaires De L’académie Des Sciences 16:387–391
Ledderhose G (1876) Ueber salzsaures glycosamin. Ber Dtsch Chem Ges 9:1200–1201
Lee YG, Chung KC, Wi SG, Lee JC, Bae HJ (2009) Purification and properties of a chitinase from Penicillium sp. LYG 0704. Protein Expr Purif 65:244–250. https://doi.org/10.1016/j.pep.2008.12.004
Li AN, Yu K, Liu HQ, Zhang J, Li H, Li DC (2010) Two novel thermostable chitinase genes from thermophilic fungi: cloning, expression and characterization. Biores Technol 101:5546–5551. https://doi.org/10.1016/j.biortech.2010.02.058
Lima LHC, Ulhoa CJ, Fernandes AP, Felix CR (1997) Purification of a chitinase from Trichoderma sp. and its action on Sclerotium rolfsii and Rhizoctonia solani cell walls. J Gen Appl Microbiol 43:31–37. https://doi.org/10.2323/jgam.43.31
Liu T, Han H, Wang D, Guo X, Zhou Y, Fukamizo T, Yang Q (2020) A potent fungal chitinase for the bioconversion of mycelial waste. J Agric Food Chem 68:5384–5390. https://doi.org/10.1021/acs.jafc.0c01342
Liu ZH, Yang Q, Hu S, Zhang JD, Ma J (2008) Cloning and characterization of a novel chitinase gene (chi46) from Chaetomium globosum and identification of its biological activity. Appl Microbiol Biotechnol 80:241–252. https://doi.org/10.1007/s00253-008-1543-x
Loc NH, Hoa NTQ, Cuc PTK, Quang HT (2013) Expression of chitinase (chi42) gene from Trichoderma asperellum in Saccharomyces cerevisiae. Ann Biol Res 4:15–19
Loc NH, Huy ND, Quang HT, Lan TT, Ha TTT (2020) Characterisation and antifungal activity of extracellular chitinase from a biocontrol fungus, Trichoderma asperellum PQ34. Mycol 11:38–48. https://doi.org/10.1080/21501203.2019.1703839
Luo Y, Deng L, Deng QJ, Wen L (2016) Comparative study of the chitooligosaccharides effect on the proliferation inhibition and radiosensitization of three types of human gastric cancer cell line. Asian Pac J Trop Med 9:601–605. https://doi.org/10.1016/j.apjtm.2016.04.014
Ma GZ, Gao HL, Zhang YH, Li SD, Xie BY, Wu SJ (2012) Purification and characterization of chitinase from Gliocladium catenulatum strain HL-1-1. Afr J Microbiol Res 6:4377–4383. https://doi.org/10.5897/AJMR12.605
Mansour SH, Abdel-Fattah AM, Esawy MA, Ahmed EF, Haroun AA, Hussein MA, Hassanien NM, Khater HM (2019) Immobilization, thermodynamic studies and application of chitinase enzyme from Penicillium chrysogenum. Egypt J Aquat Biol Fish 23:527–544. https://doi.org/10.21608/EJABF.2019.48698
Mathivanan N, Kabilan V, Murugesan K (1998) Purification, characterization, and antifungal activity of chitinase from Fusarium chlamydosporum, a mycoparasite to groundnut rust, Puccinia arachidis. Can J Microbiol 44:646–651
Matroudi S, Zamani MR, Motallebi M (2008) Molecular cloning of chitinase 33 (chit33) gene from Trichoderma atroviride. Braz J Microbiol 39:433–437. https://doi.org/10.1590/S1517-83822008000300005
Mei YX, Dai XY, Yang W, Xu XW, Liang YX (2015) Antifungal activity of chitooligosaccharides against the dermatophyte Trichophyton rubrum. Int J Biol Macromol 77:330–335. https://doi.org/10.1016/j.ijbiomac.2015.03.042
Mellor KJ, Nicholas RO, Adams DJ (1994) Purification and characterization of chitinase from Candida albicans. FEMS Microbiol Lett 119:111–118. https://doi.org/10.1111/j.1574-6968.1994.tb06876.x
Mendrofa A, Munir E, Yurnaliza Y, Lutfia A, Hartanto A (2021) Chitinase and antifungal activity of endophytic fungi isolated from Hedychium coronarium. J Koenig Agric Conspec Sci 86:131–137
Meng H, Wang Z, Meng X, Xie L, Huang B (2015) Cloning and expression analysis of the chitinase gene Ifu-chit2 from Isaria fumosorosea. Genet Mol Biol 38:381–389. https://doi.org/10.1590/S1415-475738320150003
Merzendorfer H, Zimoch L (2003) Chitin metabolism in insects: structure, function and regulation of chitin synthases and chitinases. J Exp Biol 206:4393–4412. https://doi.org/10.1242/jeb.00709
Mishra P, Kshirsagar PR, Nilegaonkar SS, Singh SK (2012) Statistical optimization of medium components for production of extracellular chitinase by Basidiobolus ranarum: a novel biocontrol agent against plant pathogenic fungi. J Basic Microbiol 52:1–10. https://doi.org/10.1002/jobm.201100446
Muthukrishnan S, Mun S, Noh MY, Geisbrecht ER, Arakane Y (2020) Insect cuticular chitin contributes to form and function. Curr Pharm Des 26:3530–3545. https://doi.org/10.2174/1381612826666200523175409
Nguyen NV, Kim YJ, Oh KT, Jung WJ, Park RD (2007) The role of chitinase from Lecanicillium antillanum B-3 in parasitism to root-knot nematode Meloidogyne incognita eggs. Biocontrol Sci Technol 17:1047–1058. https://doi.org/10.1080/09583150701668658
Odier A (1823) Mémoire sur la composition chimique des parties cornées des insectes. Mémoires de la Société d’Histoire Naturelle de Paris. Paris: Baudouin Frères Libraires-Éditeurs, tome premier 29–42
Pasqualetti M, Barghini P, Giovannini V, Fenice M (2019) High production of chitinolytic activity in halophilic conditions by a new marine strain of Clonostachys rosea. Molecules. https://doi.org/10.3390/molecules24101880
Pasqualetti M, Gorrasi S, Giovannini V, Braconcini M, Fenice M (2022) Polyextremophilic chitinolytic activity by a marine strain (IG119) of Clonostachys rosea. Molecules 27:688. https://doi.org/10.3390/molecules27030688
Patidar P, Agrawal D, Banerjee T, Patil S (2005) Optimisation of process parameters for chitinase production by soil isolates of Penicillium chrysogenum under solid substrate fermentation. Process Biochem 40:2962–2967. https://doi.org/10.1016/j.procbio.2005.01.013
Patil NS, Jadhav JP (2014) Single cell protein production using Penicillium ochrochloron chitinase and its evaluation in fish meal formulations. J Microbial Biochem Technol. https://doi.org/10.4172/1948-5948.S4-005
Patil NS, Waghmare SR, Jadhav JP (2013) Purification and characterization of an extracellular antifungal chitinase from Penicillium ochrochloron MTCC 517 and its application in protoplast formation. Process Biochem 48:176–183. https://doi.org/10.1016/j.procbio.2012.11.017
Perez-Martinez AS, De Leon-Rodriguez A, Harris LJ, Herrera-Estrella A, de la Rosa APB (2007) Overexpression, purification and characterization of the Trichoderma atroviride endochitinase, Ech42, in Pichia pastoris. Protein Expr Purif 55:183–188. https://doi.org/10.1016/j.pep.2007.05.009
Ploydee E, Chaiyanan S (2014) Production of high viscosity chitosan from biologically purified chitin isolated by microbial fermentation and deproteinization. Int J Polym Sci 2014:1–8
Poria V, Rana A, Kumari A, Grewal J, Pranaw K, Singh S (2021) Current perspectives on chitinolytic enzymes and their agro-industrial applications. Biol 10:1319. https://doi.org/10.3390/biology10121319
Prasad M, Palanivelu P (2012) Overexpression of a chitinase gene from the thermophilic fungus, Thermomyces lanuginosus in Saccharomyces cerevisiae and characterization of the recombinant chitinase. J Microb Biochem Technol. https://doi.org/10.4172/1948-5948.1000076
Purchase ER, Braun CE (1946) D-glucosamine hydrochloride. Org Syntheses 26:36–37
Qu D, Han J (2016) Investigation of the antioxidant activity of chitooligosaccharides on mice with high-fat diet. R Bras Zootec 45:661–666. https://doi.org/10.1590/S1806-92902016001100004
Qu MB, Sun SP, Liu YS, Deng XR, Yang J, Yang Q (2021) Insect group II chitinase of ChtII promotes chitin degradation during larva-pupa molting. Insect Sci 28:692–704. https://doi.org/10.1111/1744-7917.12791
Rahman MA, Halfar J (2014) First evidence of chitin in calcified coralline algae: new insights into the calcification process of Clathromorphum compactum. Sci Rep. https://doi.org/10.1038/srep06162
Raikhel NV, Lee HI, Broekaert WF (1993) Structure and function of chitin-binding proteins. Annu Rev Plant Physiol Plant Mol Biol 44:591–615. https://doi.org/10.1146/annurev.pp.44.060193.003111
Ramli ANM, Mahadi NM, Rabu A, Murad AMA, Bakar FDA, Illias RM (2011) Molecular cloning, expression and biochemical characterisation of a cold-adapted novel recombinant chitinase from Glaciozyma antarctica PI12. Microb Cell Fact 10:94. https://doi.org/10.1186/1475-2859-10-94
Rao R, Fiandra L, Giordana B, de Eguileor M, Congiu T, Burlini N, Arciello S, Corrado G, Pennacchio F (2004) AcMNPV ChiA protein disrupts the peritrophic membrane and alters midgut physiology of Bombyx mori larvae. Insect Biochem Mol Biol 34:1205–1213. https://doi.org/10.1016/j.ibmb.2004.08.002
Rawway M, Beltagy EA, Abdul-Raouf UM, Elshenawy MA, Kelany MS (2018) Optimization of process parameters for chitinase production by a marine Aspergillus flavus MK20. J Eco Heal Env 6:1–8. https://doi.org/10.18576/jehe/060101
Revah-Moiseev S, Carroad PA (1981) Conversion of the enzymatic hydrolysate of shellfish waste chitin to single-cell protein. Biotechnol Bioeng 23:1067–1078. https://doi.org/10.1002/bit.260230514
Reyes F, Calatayud J, Martinez MJ (1989) Endochitinase from Aspergillus nidulans implicated in the autolysis of its cell wall. FEMS Microbiol Lett 60:119–124. https://doi.org/10.1016/0378-1097(89)90088-8
Rodriguez J, Copa-Patlno JL, Perez-Leblic MI (1995) Purification and properties of a chitinase from Penicillium oxalicum autolysates. Lett Appl Microbiol 20:46–49. https://doi.org/10.1111/j.1472-765x.1995.tb00404.x
Rosado IV, Rey M, Codon AC, Govantes J, Moreno-Mateos MA, Benitez T (2007) QID74 cell wall protein of Trichoderma harzianum is involved in cell protection and adherence to hydrophobic surfaces. Fungal Genet Biol 44:950–964. https://doi.org/10.1016/j.fgb.2007.01.001
Rudall KM (1969) Chitin and its association with other molecules. J Polymer Sci Part C 28:83–102. https://doi.org/10.1002/polc.5070280110
Sajomsang W, Gonil P (2010) Preparation and characterization of α-chitin from cicada sloughs. Mater Sci Eng C 30:357–363. https://doi.org/10.1016/j.msec.2009.11.014
Sakurada M, Morgavi DP, Komatani K, Tomita Y, Onodera R (1997) Purification and characteristics of an autolytic chitinase of Piromyces communis OTS1 from culture medium. Curr Microbiol 35:48–51. https://doi.org/10.1007/s002849900210
Seidl V (2008) Chitinases of filamentous fungi: a large group of diverse proteins with multiple physiological functions. Fungal Biol Rev 22:36–42. https://doi.org/10.1016/j.fbr.2008.03.002
Seidl V, Huemer B, Seiboth B, Kubicek CP (2005) A complete survey of Trichoderma chitinases reveals three distinct subgroups of family 18 chitinases. FEBS J 272:5923–5939. https://doi.org/10.1111/j.1742-4658.2005.04994.x
Sharaf EF (2005) A potent chitinolytic activity of Alternaria alternata isolated from Egyptian black sand. Pol J Microbiol 54:145–151
Shehata AN, Abd El Aty AA, Darwish DA, Abdel Wahab WA, Mostafa FA (2018) Purification, physicochemical and thermodynamic studies of antifungal chitinase with production of bioactive chitosan-oligosaccharide from newly isolated Aspergillus griseoaurantiacus KX010988. Int J Biol Macromol 107:990–999. https://doi.org/10.1016/j.ijbiomac.2017.09.071
Sherief AA, El-Sawah MMA, El-Naby MAA (1991) Some properties of chitinase produced by a potent Aspergillus carneus strain. Appl Microbiol Biotechnol 35:228–230
Sikorski P, Sorbotten A, Horn SJ, Eijsink VGH, Varum KM (2006) Serratia marcescens chitinases with tunnel-shaped substrate-binding grooves show endo activity and different degrees of processivity during enzymatic hydrolysis of chitosan. Biochem 45:9566–9574. https://doi.org/10.1021/bi0603701
Sonawane KD, Dandagal NR, Naikwadi AG, Gurav PT, Anapat SV, Nadaf NH, Jadhav DB, Waghmare SR (2016) Intergeneric fusant development using chitinase preparation of Rhizopus stolonifer NCIM 880. AMB Expr 6:114. https://doi.org/10.1186/s13568-016-0287-8
Staats CC, Kmetzsch L, Lubeck I, Junges A, Vainstein MH, Schrank A (2013) Metarhizium anisopliae chitinase CHIT30 is involved in heat-shock stress and contributes to virulence against Dysdercus peruvianus. Fungal Biol 117:137–144. https://doi.org/10.1016/j.funbio.2012.12.006
Staufenberger T, Imhoff JF, Labes A (2012) First crenarchaeal chitinase found in Sulfolobus tokodaii. Microbiol Res 167:262–269. https://doi.org/10.1016/j.micres.2011.11.001
Suryawanshi N, Eswari JS (2022) Purification and characterization of chitinase produced by thermophilic fungi Thermomyces lanuginosus. Prep Biochem Biotechnol 52:1087–1095. https://doi.org/10.1080/10826068.2022.2028639
Svedese VM, Tiago PV, Bezerra JDP, Paiva LM, de Luna Alves Lima EA, Porto ALF (2013) Pathogenicity of Beauveria bassiana and production of cuticle-degrading enzymes in the presence of Diatraea saccharalis cuticle. Afr J Biotechnol 12:6491–6497. https://doi.org/10.5897/AJB2013.11972
Thakur D, Chauhan A, Jhilta P, Kaushal R, Dipta B (2022) Microbial chitinases and their relevance in various industries. Folia Microbiol. https://doi.org/10.1007/s12223-022-00999-w
Tikhonov VE, Lopez-Llorca LV, Salinas J, Jansson HB (2002) Purification and characterization of chitinases from the nematophagous fungi Verticillium chlamydosporium and V. suchlasporium. Fungal Genet Biol 35:67–78. https://doi.org/10.1006/fgbi.2001.1312
Tominaga Y, Tsujisaka Y (1976) Purifications and some properties of two chitinases from Streptomyces orientalis which lyse Rhizopus cell wall. Agr Biol Chem 40:2325–2333. https://doi.org/10.1080/00021369.1976.10862407
Tweddell RJ, Jabaji-Hare SH, Charest PM (1994) Production of chitinases and β-1,3-glucanases by Stachybotrys elegans, a mycoparasite of Rhizoctonia solani. Appl Environ Microbiol 60:489–495
Vaaje-Kolstad G, Bunaes AC, Mathiesen G, Eijsink VGH (2009) The chitinolytic system of Lactococcus lactis ssp lactis comprises a nonprocessive chitinase and a chitin-binding protein that promotes the degradation of α- and β-chitin. FEBS J 276:2402–2415. https://doi.org/10.1111/j.1742-4658.2009.06972.x
Vaaje-Kolstad G, Houston DR, Riemen AHK, Eijsink VGH, van Aalten DMF (2005) Crystal structure and binding properties of the Serratia marcescens chitin-binding protein CBP21. J Biol Chem 280:11313–11319. https://doi.org/10.1074/jbc.M407175200
Velmurugan N, Kalpana D, Han JH, Cha HJ, Lee YS (2011) A novel low temperature chitinase from the marine fungus Plectosphaerella sp. strain MF-1. Bot Mar 54:75–81. https://doi.org/10.1515/BOT.2010.068
Viterbo A, Haran S, Friesem D, Ramot O, Chet I (2001) Antifungal activity of a novel endochitinase gene (chit36) from Trichoderma harzianum Rifai TM. FEMS Microbiol Lett 200:169–174. https://doi.org/10.1111/j.1574-6968.2001.tb10710.x
Vyas P, Deshpande M (1991) Enzymatic hydrolysis of chitin by Myrothecium verrucaria chitinase complex and its utilization to produce SCP. J Gen Appl Microbiol 37:267–275. https://doi.org/10.2323/jgam.37.267
Wang C, Zeng ZQ, Zhuang WY (2021) Comparative molecular evolution of chitinases in ascomycota with emphasis on mycoparasitism lifestyle. Microbial Genomics 7:000646. https://doi.org/10.1099/mgen.0.000646
Wang J, Zhang J, Song F, Gui T, Xiang J (2015) Purification and characterization of chitinases from ridgetail white prawn Exopalaemon carinicauda. Molecules 20:1955–1967. https://doi.org/10.3390/molecules20021955
Wang Q, Qu L, Zhang Z, Wang Y, Zhang Y (2013) Characterization of a novel chitinase, DkChi, from Dendrolimus kikuchii nucleopolyhedrovirus. Arch Virol 158:2523–2530. https://doi.org/10.1007/s00705-013-1775-7
Wang SL, Hsiao WJ, Chang WT (2002) Purification and characterization of an antimicrobial chitinase extracellularly produced by Monascus purpureus CCRC31499 in a shrimp and crab shell powder medium. J Agric Food Chem 50:2249–2255. https://doi.org/10.1021/jf011076x
Wang YJ, Yang Q (2009) Cloning and expression of a novel chitinase chi58 from Chaetomium cupreum in Pichia pastoris. Biochem Genet 47:547–558. https://doi.org/10.1007/s10528-009-9251-5
Wattanalai R, Wiwat C, Boucias DG, Tartar A (2004) Chitinase gene of the dimorphic mycopathogen, Nomuraea rileyi. J Invertebr Pathol 85:54–57. https://doi.org/10.1016/j.jip.2003.12.007
Wu JH, Ali S, Ren SX (2010) Evaluation of chitinase from Metarhizium anisopliae as biopesticide against Plutella xylostella. Pakistan J Zool 42:521–528
Xia G, Jin C, Zhou J, Yang S, Zhang S, Jin C (2001) A novel chitinase having a unique mode of action from Aspergillus fumigatus YJ-407. Eur J Biochem 268:4079–4085. https://doi.org/10.1046/j.1432-1327.2001.02323.x
Xie XH, Fu X, Yan XY, Peng WF, Kang LX (2021) A broad-specificity chitinase from Penicillium oxalicum k10 exhibits antifungal activity and biodegradation properties of chitin. Mar Drugs. https://doi.org/10.3390/md19070356
Yanai K, Takaya N, Kojima N, Horiuchi H, Ohta A, Takagi M (1992) Purification of two chitinases from Rhizopus oligosporus and isolation and sequencing of the encoding genes. J Bacteriol 174:7398–7406. https://doi.org/10.1128/jb.174.22.7398-7406.1992
Youn DK, No HK, Prinyawiwatkul W (2013) Preparation and characteristics of squid pen β-chitin prepared under optimal deproteinisation and demineralisation condition. Int J Food Sci Technol 48:571–577. https://doi.org/10.1111/ijfs.12001
Yu G, Xie LQ, Li JT, Sun XH, Zhang H, Du Q, Li QY, Zhang SH, Pan HY (2015) Isolation, partial characterization, and cloning of an extracellular chitinase from the entomopathogenic fungus Verticillium lecanii. Genet Mol Res 14:2275–2289. https://doi.org/10.4238/2015.March.27.13
Zhao X, Yu Z, Wang T, Guo X, Luan J, Sun Y, Li X (2016) The use of chitooligosaccharide in beer brewing for protection against beer-spoilage bacteria and its influence on beer performance. Biotechnol Lett 38:629–635. https://doi.org/10.1007/s10529-015-2013-z
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Handling Editor: Sonia Malik
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Thakur, D., Bairwa, A., Dipta, B. et al. An overview of fungal chitinases and their potential applications. Protoplasma 260, 1031–1046 (2023). https://doi.org/10.1007/s00709-023-01839-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00709-023-01839-5