Abstract
Thauera is the most widely found dominant denitrifying genus in wastewater. In earlier study, MBBR augmented with a specially developed denitrifying five-membered bacterial consortium (DC5) where Thauera was found to be the most abundant and persistent genus. Therefore, to check the functional potential of Thauera in the removal of nitrate-containing wastewater in the present study Thauera sp.V14 one of the member of the consortium DC5 was used as the model organism. Thauera sp.V14 exhibited strong hydrophobicity, auto-aggregation ability, biofilm formation and denitrification ability, which indicated its robust adaptability short colonization and nitrate removal efficiency. Continuous reactor studies with Thauera sp.V14 in 10 L dMBBR showed 91% of denitrification efficiency with an initial nitrate concentration of 620 mg L−1 within 3 h of HRT. Thus, it revealed that Thauera can be employed as an effective microorganism for nitrate removal from wastewater based on its performance in the present studies.
Graphical abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Moving bed biofilm reactor (MBBR) has been considered the most efficient and widely used biofilm-based wastewater treatment process [1]. It has attracted greater attention due to its simple operation, cost effectiveness and minimal waste generation. However, due to the increased emphasis on reducing the cost of treatment and minimizing organic carbon demand in wastewater treatment, the usage of MBBR has increased worldwide [2]. Moreover, MBBR has been widely used reactors to enable the growth of higher biomass. Bioaugmentation of denitrifying bacterial culture is necessary when the native bacterial community does not remove sufficient nitrate from wastewater. Bioaugmentation of microorganisms is one of the cost-effective and environmentally friendly approaches for the removal of nitrate from wastewater. Bioaugmentation of specific bacterial strains with high adaptability and specialized function can be added into the biologic treatment process to enhance specific functional species, alter the composition of the microbial community, and increase the effectiveness of the removal of contaminants [3,4,5]. Various denitrifying bacteria have been isolated and shown to reduce nitrate in wastewater. Additionally, few recent reports on bioaugmentation have also suggested that strain of Zobellella B307, Acinetobacter sp. TAC-1 Pseudomonas stutzeri strain HK13, Hydrogenophaga sp. H7 and the special microbial seed of biofilm forming denitrifying bacteria enhanced nitrate and nitrogen removal of wastewater [6,7,8,9,10]. Among many denitrifying bacteria, Thauera was the most commonly found organism in wastewater treatment plants, polluted freshwater, wet soil, and water contaminated with aromatic or aliphatic organic compounds [11]. Thauera is a Gram-negative bacterium that can denitrify nitrogenous oxides under anoxic conditions [12]. It contributes to waste treatment by targeting recalcitrant compounds, humic acids, metal ions and detoxifying them into more easily utilizable compounds and serves as nutrients for other microorganisms [4, 5]. Its degradative abilities toward nitrate and other recalcitrant pollutants make them potential organism for bioremediation. It has been the most widely used organism for wastewater treatment applications due to its adaptable metabolism and ability to grow in a variety of environmental conditions. Thauera is well-known for denitrification [13], aerobic denitrification [14], partial denitrification [15], hydrogen-oxidizing autotrophic denitrification [16] and Hydrogen-based denitrification [17]. In the literature, different species of Thauera have been reported to be major contributors in a variety of reactors, such as Thauera mechernichensis was the major contributor in the treatment of resin wastewater in the aerobic nitrifying-denitrifying membrane bioreactor [18], Thauera aminoaromatica in hypoxic quinoline denitrifying sequencing batch reactor [19], Thauera sp. SND5 removed nitrogen and phosphorus from wastewater via simultaneous nitrification and denitrification [20]. It was also dominant source of functional genes in aerobic granular sludge [21]. Moreover, it has diverse functional metabolic pathways like carbon, nitrogen, sulphur and phosphorus etc. [20]. The presence of these diverse metabolic pathways made Thauera more competitive and one of the potential microorganism for the treatment of wastewater [22]. Therefore, the study aimed to assess the functional potential of Thauera sp. V14, a member of the DC5 consortia, in the removal of nitrate from wastewater. Metagenomic data suggested that MBBR inoculated with DC5 showed Thauera spp. as the most dominant and persistent microorganism. Thus, to explore the potential application of Thauera spp. in the denitrification process, Thauera sp. V14 was chosen as the model organism because of its significant properties such as strong hydrophobicity, auto-aggregation ability, biofilm formation and denitrification competence, implying its adaptability for efficient colonization and nitrate removal. Consequently, the findings collectively highlight Thauera sp. V14 as a microorganism with significant promise for novel and efficient nitrate removal from wastewater.
Materials and methods
Microorganism, media and growth condition
Thauera sp. V14, a member of the DC5 consortium was used in dMBBR studies. DC5 consortium was prepared as reported in our earlier published work [10]. Thauera sp.V14 was isolated from an activated sludge sample, using CPNA (Congo red peptone nitrate agar) medium (0.5 g peptone, 0.3 g beef extract, 1 g potassium nitrate, 2 g agar, 0.005 g Congo red in 100 mL of distilled water). Synthetic wastewater used in this study was prepared as follows: 0.2 g MgSO4 0.7H2O, 0.2 g K2HPO4, 0.05 g FeSO4.7H20, 0.02 g CaCl2.2H2O, 0.002 g MnCl2.4H2O, 0.001 g NaMoO4.2H2O, 1 g KNO3, 0.6 g sodium acetate, 0.5 g yeast extract were added per liter of distilled water in the flask. Total 50 mL MM2 (minimal media) medium was prepared. All the chemicals/media used in the experiments were purchased from HiMedia Laboratories Pvt. Ltd (Mumbai, India).
Phylogenetic tree
For phylogenetic analysis of sequences of Thauera sp. V14, Thauera MZ1T and Thauera humireducens (spp. obtained in metagenomic analysis) were done in MEGA 11 using the neighbor joining method in MEGA X.
Auto-aggregation ability of Thauera sp.V14
Thauera sp. V14 was grown in PNB for 18–24 h and centrifuged at 8000 rpm for 7 min. Harvested cells were washed twice with PBS (phosphate buffer solution) and then resuspended in PBS with absorbance set at 0.6 OD600nm. This suspension was inoculated in 50 mL of MM2 medium and allowed to stand for 8 h to allow aggregation and the OD600nm was measured again (At). The auto-aggregation index was calculated by Eq. (1) to evaluate the auto-aggregation capacity of Thauera sp. V14 [23].
where At and A0 are OD600nm at time t = 1, 2, 3, 4, 5, 6, 7 and 8 and at time t = 0 time respectively.
Hydrophobicity of Thauera sp.V14
The hydrophobicity of cells was determined by measuring bacterial adhesion. Thauera sp.V14 was grown in PNB for 18–24 h and centrifuged at 8000 rpm for 7 min. Harvested cells were washed twice with PBS and resuspended in PBS with absorbance set at 0.3 OD546nm and inoculated in 50 mL of MM2 medium. Aliquots were drawn at every 24 h intervals for 120 h. To check the hydrophobicity, 4 mL of cell suspension with 1 mL of hexadecane was vortexed for 2 min and the two phases were allowed to separate for 15 min. The aqueous phase was removed carefully, and its OD546nm was taken. The cell surface hydrophobicity was calculated using Eq. (2) [23].
Confirmation of denitrifying ability of Thauera sp. V14
Cells of the Thauera sp.V14 were grown for 24 h in PNB and were harvested by centrifugation at 10,000 rpm for 5 min, washed twice with PBS and resuspended in PBS. 500 µL of this was inoculated into peptone nitrate broth (PNB) and the tube was sealed with suba-seal rubber stopper and incubated for 48 h. After 48 h, 1 µL gaseous sample collected and was injected into the GC. Dinitrogen (N2) content was determined by gas chromatography (GC) using Sigma Instruments Ltd (India) equipped with Flame Ionization Detector (FID). The temperature of the column, the injector port, and the FID were 250 °C, 260 °C and 260 °C, respectively. CR-624 column with a mesh size of 3 µm was used, and nitrogen gas was used as a carrier gas.
Resting cells kinetics using Thauera sp.V14
Nitrate and nitrite reduction rates by resting cell suspension of the Thauera sp.V14 were performed as follows. Cells grown for 24 h in PNB were harvested by centrifugation at 10,000 rpm for 5 min, washed twice with PBS and again resuspended in PBS. Sodium acetate and potassium nitrate were added as electron donors and acceptors respectively and the reduction of nitrate and formation of nitrite were estimated for 48 h after every 12 h time intervals.
Flask level denitrification efficiency of Thauera sp.V14
Denitrification studies were carried out with Thauera sp.V14 with an initial nitrate concentration of 765 mg L−1 in a 250 mL Erlenmeyer flask. It was grown in PNB for 18–24 h and centrifuged at 8000 rpm for 7 min. The cell pellet was washed twice with PBS and resuspended in PBS with absorbance set at 0.5 OD600nm. 1% of this was inoculated in a flask containing 100 mL of MM2 medium. Experimental flasks were incubated at 37 °C for 48 h in static conditions; aliquots were drawn at 24 h intervals and assayed for nitrate, nitrite and ammonia. Denitrification efficiency was calculated by the following formula:
Inoculum preparation for continuous dMBBR studies
Thauera sp.V14 was grown in PNB for 24 h. The absorbance of 0.5 OD600nm was set. The cell pellet obtained after centrifugation at 8000 rpm for 7 min was washed twice with PBS and resuspended in 80 mL of PBS. 80 mL of this suspension was then added to 800 mL of MM2 medium and incubated at 37 °C under the static condition for 24 h. This was used as an inoculum.
Bench-scale dMBBR developed with Thauera sp.V14
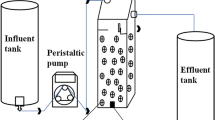
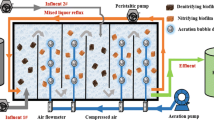
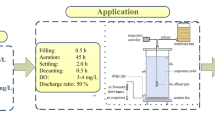
Continuous dMBBR studies were carried out with Thauera sp.V14 in a 10 L reactor. For continuous reactor studies, 10 L of synthetic effluent (MM2 medium) was continuously fed from the inlet tank to the reactor with the peristaltic pump (Masterflex®). 8% of inoculum was prepared as mentioned in the above section was added in the 10 L reactor containing carriers and biofilm was allowed to form on the carriers for 10 days. Here, 620, 744, 930, 1116, 1500 and 2400 mg L−1 nitrate loading and COD concentrations were 186, 223, 279, 335, 450 and 750 mg L−1, respectively. Studies were carried out with C/N ratio 0.3, HRT of 3 h and filling ratio 20% of pall ring carrier [10]. pH-8, dissolved oxygen (DO) – 0.1–0.8 mg L−1, and turbidity were 15–350 Nephelometric Turbidity Unit (NTU) maintained in the dMBBR.
Analytical methods
Treated synthetic effluent was collected in the outlet tank and assayed for nitrate, nitrite, ammonia, pH, turbidity, biomass and DO at the temperature of 37 °C. Biomass from the carriers was quantified by drying carrier material at 105 °C for 1 h. Nitrate, nitrite, ammonia and chemical oxygen demand (COD) estimation methods were performed according to APHA1998. DO was measured using a DO probe (Thermo Fisher Scientific, India) and the turbidity was checked using a turbidity meter (Hanna Instruments, India).
Functional potential of genus Thauera
The functional potential of genus Thauera was checked using MG RAST software was used to filter genus level data and to check its functional potential [24].
Measurement of the abundance of Thauera sp.V14
Biofilm samples were taken from different parts of the dMBBR. Genomic DNA from biofilm samples was extracted using the modified CTAB (cetyltrimethylammonium bromide) method. Biofilm developed on the carriers of dMBBR was scrapped and resuspended in 10 mL of PBS. Resuspended biomass was centrifuged at 8000 rpm for 5 min. The pellet was resuspended in 500 μL of Tris–EDTA-Sucrose of buffer (25 mM TrisCl (pH 8), 25 mM EDTA (pH 8) and 300 mM Sucrose). Then, 8 μL of lysozyme (10 mg/mL) was added to the system and incubated at 37 °C for 1 h. After incubation 10% SDS was added and samples were mixed properly, and kept in a water bath at 60 °C for 1 h. Then 200 μL of 5 M NaCl and 80 μL of 10% CTAB were added and incubated at 65 °C for 10 min. An equal volume of phenol:chloroform:isoamylalcohol (25:24:1) was added and centrifuged at 10,000 rpm for 10 min. This step was repeated twice. The upper aqueous phase was collected and an equal volume of chloroform:isoamylalcohol (24:1) was added and centrifuged at 10,000 rpm for 10 min. Aqueous phase was extracted and 1/10th volume of 3 M chilled sodium acetate was added. Then double volume of absolute alcohol was added and incubated at chilled temperature for 24 h. After incubation centrifugation was done at 10,000 rpm for 10 min, the supernatant was discarded and the pellet was air dried and resuspended in sterile Milli-Q water. Resuspended DNA was treated with 3 μL of 0.01% RNase added to it incubated at 65 °C for 10 min and stored at 4 °C which was preserved in cold conditions. The quality and quantity of extracted DNA were assessed using agarose gel electrophoresis and nanospectrophotometer, respectively. From this isolated gDNA abundance of Thauera in the biofilm developed inside dMBBR was quantified using Real-Time PCR (Applied Biosystems StepOne™ Real-Time PCR System, India). For this Thauera specific primers were used Forward: TGCATTGCTGCTCCGAAC and Reverse: CGCTCGTTG CGGGACTTAACC and its abundance was quantified as per [25].
Observation of carrier-associated biofilm
SEM was used to characterize biofilm morphology on carriers in dMBBR. Carriers before biofilm development and continuous nitrate removal studies (after biofilm development) were collected from the reactor without disruption. After collection carriers with biofilm were allowed to dry in the hot air oven at 100 °C for 1 h then carriers were cut into small pieces. Further for SEM analysis they were coated with gold and examined under JEOL JSM-6380 LV SEM, accelerating voltage of 15 kV.
Qualitative biochemical analysis of Thauera sp. V14 biofilm
FTIR analysis was used to characterize the major components of the Thauera sp. V14 biofilm. Carriers with developed biofilm were collected after continuous denitrification studies from the dMBBR and dried at 70 °C for 1 h, and then biomass of biofilm was scrapped off from the carriers and it was analyzed using FTIR Spectrometer (FTIR Spectrometer: ALPHA, India).
Results and discussions
In our earlier investigations, Thauera spp. emerged as the most dominant and persistent microorganisms in the MBBR augmented with consortium DC5, comprising Diaphorobacter sp. R4, Pannonibacter sp. V5, Thauera sp. V9, Pseudomonas sp. V11 and Thauera sp. V14. After 300 days of continuous operation, whole-genome metagenomic studies confirmed Thauera spp.'s dominance as the key contributor to the denitrification of nitrate-containing synthetic wastewater [10]. These findings led to an in-depth investigation into the functional potential of Thauera spp. specifically investigating the metabolic pathways present in the genus. To investigate the practical application of Thauera spp. in denitrification processes, Thauera sp. V14 was chosen as the model organism due to its distinctive attributes.
Metabolic potential analysis of genus Thauera in dMBBR bioaugmented with consortium DC5
Metabolic potential of the microorganisms is directly related to the bioremediation process [26]. Functional metabolic potential of microorganism is generally categorized into two groups (i) The housekeeping genes that are included in the first group encode essential enzymes involved in the metabolism of carbohydrates, proteins, amino acids, lipids, fatty acids, DNA and RNA, cofactors and vitamins, cell walls and capsules, respiration, membrane transport, cell division and cell cycle, cell signaling, motility, and chemotaxis (ii) Second group of genes are those that are involved in specific function such as stress response, virulence, toxic compound resistance, xenobiotic substance metabolism and degradation.
Thauera contains high abundance of metabolic functions (57.44%) followed by genetic information processing (17.96%), environmental information processing (18.24%), cellular processes (6.11%) and organismal systems (0.26%) were identified at KO level 1 (KEGG Orthology level 1) (Fig. 1a). Genes responsible for cell growth and death (1.58%), cell motility (3.56%), transport and catabolism (1.10%), membrane transport (9.12%), signal transduction (8.30%), folding, sorting and degradation (3.31%), replication and repair (5.72%), transcription (1.64%), translation (7.61%), amino acid metabolism (20.49%), biosynthesis of other secondary metabolites (0.18%), carbohydrate metabolism (11.30%), energy metabolism (6.52%), glycan biosynthesis and metabolism (2.65%), lipid metabolism (2.45%), metabolism of cofactors and vitamins (6.68%), metabolism of other amino acids (1.06%), metabolism of terpenoids and polyketides (1.13%), nucleotide metabolism (4.67%), xenobiotics and biodegradation and metabolism (0.65%), environmental adaptation (0.25%) were identified at KO level 2 in genus Thauera (Fig. 1b). In genus Thauera, high abundance of genes were involved in amino acid metabolism i.e., 20.49% which might be providing carbon and energy source for bacterial metabolism [27]. High abundance of genes was also involved in carbohydrate metabolism i.e. 11.30% were attributed to a decomposition of complex organics into more easily degradable matter [28]. High abundance of amino acid metabolism and carbohydrate metabolism in Thauera may act as electron donors and carbon sources for denitrification [27, 29]. These carbohydrate metabolism genes might be involved in the central carbon metabolism (glycolysis, citrate cycle, and pentose phosphate pathway), producing ATP for energy or NADPH for reducing power, which is necessary for cell metabolism. Moreover, it also can decompose complex organics into more easily degradable matter [28]. Genes involved in energy metabolism (6.52%) mainly include oxidative phosphorylation and nitrogen metabolism. Genes involved in oxidative phosphorylation and nitrogen metabolism might be related to the degradation of phosphate and nitrogen containing pollutants [30]. Genes involved in xenobiotics biodegradation (0.65%) and genes of biosynthesis of secondary metabolites (0.18%) play important role in defense against other microorganisms and harmful stresses such as toxins or UV exposure, were also identified in the genus Thauera.
Genes and functions associated with organic contaminant degradation in Thauera spp. developed inside dMBBR
Denitrification process is often associated with the degradation of organic compounds under anaerobic or anoxic conditions. These organic compounds provide electrons that are subsequently used by different electron acceptors (such as sulfate, nitrate and CO2) which are further converted into harmless gases such as sulfide, nitrogen gas and biomethane gas under anaerobic or anoxic conditions [31]. Therefore, the presence of genes associated with organic contaminant degradation and denitrification was also checked in the genus Thauera.
Relative abundance of genes involved in nitrogen metabolism
The biologic process of nitrogen removal in wastewater takes a specific path, depending on the microbial communities involved and the substrates available [40, 41]. The major pathway in nitrogen metabolism is the denitrification process which could be linked to carbon catabolic pathways including amino acid, fatty acid and carbohydrate degradation and could provide electron donors for nitrate reduction and efficient nitrogen removal as well as it could also be linked to COD removal [32]. According to KO annotation level-2 in genus Thauera, genes involved in the denitrification process were most abundant than other enzymes involved the nitrogen metabolism. In nitrogen metabolism pathways 42.06% of genes nitrate reductase catalytic subunit [EC: 1.7.99.4], NapA [EC: 1.7.99.4], NapB, NapC, NapD, NapF, NapG, NapH were involved in the conversion of nitrate to nitrite.6.2% of nirB; [EC: 1.7.1.4] and nirD; [EC: 1.7.1.4] genes were involved in the conversion of nitrite to nitric oxide. 11.8% of norB [EC: 1.7.2.5], norC [EC: 1.7.99.7], norD [EC: 1.7.99.7] and norF [EC: 1.7.99.7] of genes were responsible for the conversion of nitric oxide to nitrous oxide and 1.18% nosZ [EC: 1.7.2.4] involved in the conversion of nitrous oxide to nitrogen gas. Figure 2a shows the relative abundance of genes involved in nitrogen metabolism.
Relative abundance of genes involved in sulfur metabolism in genus Thauera
Sulfur metabolism plays an important role in central biochemistry as a carbon carrier and stable redox center [33]. In the absence of molecular oxygen and nitrate microbes often prefer sulfur compounds as electron donors or acceptors for energy transformation and metabolism [17,18,19]. Among different metabolic pathways present in the genus Thauera genes involved in Assimilatory sulfate Reduction (ASR) pathway were most abundantly present. These genes involved in ASR have been found to contribute to sulfate removal in various wastewater treatments [19]. In sulfur metabolism 32.25% of cys [EC:2.7.7.4] genes convert sulfate to adenylylsulfate, 5.49% cysH [EC:1.8.4.8] genes for the conversion of 3 phosphoadenylylsulfate (PAPS) to sulfite, 4.52% cysI gene converts sulfite to H2S and total 57.73% of cysN/cysNC [EC:2.7.7.4;2.7.1.25] converts adenylsulfate(APS) to 3 phosphoadenylsulfate (PAPS) (Fig. 2b).
Relative abundance of genes involved in methane metabolism
As reported by [34] in biofilm processes under anaerobic conditions methanogenesis and denitrification process co-exist. The anaerobic methane-driven denitrification process directly utilizes methane as electron donor that is abundant in biogas produced from the anaerobic digestion of organic waste. In anoxic environments in the presence of inorganic electron acceptors (e.g., sulfate or nitrate), methanogenesis plays an important role in the biodegradation of organic matter. Figure 2c shows the relative abundance of genes involved in methane metabolism in the genus Thauera. In methane metabolism, the hydrogenotrophic methanogenic pathway was present that reduces hydrogen gas to formate or CO2 to methane. In this pathway, 72.84% of genes were hydrogen dehydrogenase [EC: 1.12.1.2] which reversibly interconvert protons and electrons to molecular hydrogen, 24.43% of metF gene [EC: 1.5.1.20] catalyzes the reversible conversion of 5, 10- methylenetetrahydrofolate to 5-methyltetrahydrofolate and 2.73% of frmB, ESD fghA [EC: 3.1.2.12] hydrolyses to formate which can be converted to carbon dioxide.
Xenobiotic biodegradation and metabolism genes in genus Thauera
Presence of genes related to xenobiotics biodegradation and metabolism in Thauera suggest that it has a well-developed mechanism to neutralize the harmful effect of xenobiotic compounds, which is also important for the removal of nitrate containing organic pollutants. Xenobiotic compounds are chemicals that are foreign to microorganisms. Biotransformation is the metabolic modification of the molecular structure of a compound and involves the breakdown of the organic compound into less complex ones. Chlorocyclohexane and chlorobenzene degradation, Benzoate degradation and Nitrotoluene degradation were the most abundantly present pathways in the genus Thauera that were involved in the xenobiotic biodegradation and metabolic pathways. 46.55% of the genes were involved in the benzoate degradation [PATH: ko00362] (Fig. 2d) followed by 38.62% chlorocyclohexane and chlorobenzene degradation [PATH: ko00361] and 14.83% of genes that were involved in the nitrotoluene degradation [PATH: ko00633]. Benzoate pathways are the common intermediate compound of all aerobic and anaerobic metabolic pathways of aromatic compounds (phenolics, polycyclic aromatic hydrocarbons). Ubiquitous degradation pathways for benzoate and interrelated compounds are the central pathways for xenobiotics mineralization and detoxification by microbial communities [35].
Phylogenetic analysis of Thauera spp.
Phylogenetic analysis of Thauera sp. V14, Thauera MZ1T and Thauera humireducens were performed using 16S rRNA gene sequences. 16s rRNA sequences of Thauera MZ1T and Thauera humireducens were obtained from NCBI. Our strain Thauera sp.V14 showed 99.38% sequence similarity with Thauera mechernichensis which was closely related to Thauera humireducens and Thauera sp. MZ1T which are similar in genetic makeup [10]. Therefore, our further reactor studies were carried out with Thauera sp. V14 as a model organism to check its potential role in the treatment of nitrate containing wastewater (Fig. 3).
Characterization of Thauera sp.V14
Denitrification efficiency, auto-aggregation and hydrophobicity
Biofilm forming denitrifying Thauera sp.V14 showed 99% sequence similarity with Thauera sp. [10]. Figure 4a showed that Thauera sp. V14 showed 100% denitrification efficiency within 72 h with an initial nitrate concentration of 765.89 mg L−1. No nitrite and ammonia were detected in the medium. It indicates that Thauera sp.V14 exhibited complete denitrification efficiency without accumulation of intermediates like nitrite and ammonia that have harmful effects on human and environment and potential to contribute to water pollution.
Denitrification efficiency of Thauera sp.V14 showed 100% denitrification efficiency with 765.89 mg L−1 of nitrate which was higher compared to other organisms reported in the literature such as Acinetobacter haemolyticus ZYL removed 100% at 36 h with the initial concentration of 443 mg L−1 [36]. In addition, Pseudomonas stutzeri strain XL-2 removed 97.9% of nitrate within 24 h with an initial concentration of 443 mg L−1 [37]. Pannonibacter phragmitetus B1 reduced 98.77% nitrate within 18 h with an initial nitrate concentration of 65.16 mg L−1 [38]. Pseudomonas sp. JQ-H3 was able to reduce 438.6 mg L−1 of nitrate within 72 h [39]. Zoogloea sp. N299 reduced 75.42% nitrate within 72 h [40]. It also suggested that it was also one of the most efficient denitrifying microorganism that was reported in the literature.
A high auto-aggregation (93%) index of Thauera indicates a strong tendency of cells to cluster and it is a primary step in the development of biofilm [41]. Auto-aggregation increased gradually from 34% at 1 h to 93% at 8 h (Fig. 4b) and hydrophobicity from 7.7% on day 1 to 83.8% on day 5 (Fig. 4c). High denitrification efficiency, auto-aggregation and hydrophobicity suggest Thauera as a suitable organism for denitrification in biofilm reactor.
Thauera sp.V14 also showed 93% auto-aggregation capacity and 83.8% hydrophobicity, which is favorable for the formation of biofilm in dMBBR. A high auto-aggregation index of Thauera sp.V14 indicates a strong tendency for cells to cluster and hydrophobicity of isolate leads to better auto-aggregation ability. Auto-aggregation ability and hydrophobicity of bacteria are closely related to the aggregation and adhesion between the bacterial surface and between other bacteria [23]. Bioaugmentation of bacteria with these properties reduces the loss of bacteria from the wastewater treatment system [42]. Auto-aggregation ability and hydrophobicity of Thauera sp.V14 were higher than Enterobacter sp. strain FL which showed 54.3% at 48 h [42] and Methylobacterium gregans DC-1, showed 38.7% at 72 h [43] suggested Thauera as a suitable microorganism for biofilm reactor i.e., dMBBR.
Kinetic analysis of nitrate removal by Thauera sp.V14
Nitrogen gas production by Thauera sp.V14 was confirmed by gas chromatography. Production of nitrogen gas was confirmed by comparing the peak at 1.373 with the standard of nitrogen gas. In PNB media after 48 h of incubation Thauera sp.V14 showed 64.68% of nitrogen gas production (Fig. 5a).
The nitrate removal rate by Thauera sp.V14 was investigated using the zero-order kinetic model [44]. The model can be described as follows:
where Ci represents the initial NO3 concentration (mg L −1), Ct (mg L −1) is the remaining NO3 concentration at time (t), t is the time (h) and K0 is the rate constant for zero order kinetic. K0 constant was obtained from the slopes of the plots of Ct versus t for Eq. (3), and the regression coefficients (R2). R2 value for Thauera sp.V14 was 0.92 respectively suggesting a better fit of zero-order kinetic model (Fig. 5b). Nitrate removal rate constant for Thauera sp.V14 was 4.6 mg L−1 h−1.
Further, relative rates (RR) of nitrate reduction were calculated using the following formula:
In the case of no nitrite build up RR would be equal to 1, and KNO2 = 0. Whereas RR ≥ 1 signifies the buildup of nitrite [44]. Relative rate of Thauera sp.V14 was 1 suggesting no nitrite build up by Thauera sp.V14. The results of kinetics studies and GC analysis suggested Thauera sp.V14 as an efficient denitrifying organism which is able to reduce nitrate without accumulation of nitrite, which is harmful.
Nitrate removal studies with Thauera sp.V14 in continuous 10L dMBBR
To check the denitrification ability of the Thauera sp.V14 in the dMBBR different concentrations of nitrate viz. 620, 744, 930, 1500 and 2400 mg L−1 were added in the influent of synthetic wastewater. As shown in Fig. 6 denitrification efficiency in the dMBBR was 91%, 90%, 76%, 66% and 60% at 620, 744, 930,1500 and 2400 mg L−1 of NO3 concentration and every time COD reduction was below the stipulated permissible range i.e. 250 mg L−1. Notably, no nitrite and ammonia were detected inside dMBBR which was important for the complete denitrification process. It was also observed that as the nitrate concentration increases denitrification efficiency in the dMBBR was decreased. This could be due to the saturation of denitrifying enzymes. Because as the denitrifying enzymes saturated their denitrification efficiency decreases. Hence, it can be suggested that Thauera sp.V14 was able to reduce nitrate up to 2400 mg L−1 with 60% efficiency. In the context of nitrate contamination in India, where concentration in the groundwater and surface water was in the range of 139 to 557 mg L−1 in wastewater [45,46,47,48,49,50,51,52,53], and our isolate Thauera sp. V14 showed high denitrification efficiency i.e., 90% till 744 mg L−1, suggested that it can be used for the treatment of high nitrate containing wastewater. Moreover, nitrate removal efficiency of Thauera in dMBBR was also higher than other reported denitrifying organisms such as Pseudomonas mendocina IHB602 in sequencing batch biofilm reactor [23], Acinetobacter sp.TAC-1in MBBR [7] and Diaphorobacter polyhydroxybutyrativorans in the denitrifying reactor [54]. These findings suggested Thauera sp.V14 as the most efficient microorganism and can be used as a potential organism for denitrification of nitrate containing wastewater. Furthermore, abundance of Thauera sp.V14 in the biofilm of dMBBR was checked i.e. 2 × 108 copy number/µL of biomass as per RTPCR suggesting that dMBBR was fully conditioned with Thauera and this organism may be contributing majorly in the denitrification. Moreover, High abundance of Thauera inside the developed biofilm of dMBBR suggested its successful bioaugmentation in the dMBBR.
Characterization of Thauera biofilm developed inside dMBBR
Surface topology and EPS composition of the biofilm developed in the dMBBR with Thauera sp.V14
The biofilm morphology developed on the pall ring carrier was examined using SEM images. Figure 7a represents the SEM images of the carriers before biofilm formation and after nitrate removal studies and biofilm formation on Pall ring carriers. Biomass on the carrier material was 30 ± 5 mg/carrier. Carriers after biofilm formation showed high bacterial density and biofilm seemed to be dominated by rod shaped bacteria. These results were similar to [55] who also showed rod-shaped bacteria as the most predominant bacteria in the denitrifying bioreactors. Successful colonization of bacteria on the pall ring carriers and the carrier surface provided a good porous structure so that bacteria did not easily wash away in dMBBR.
Further, FTIR spectroscopy was employed to analyze the composition and functional group characteristics present in the developed biofilm of the dMBBR. The FTIR profiles of different functional groups are illustrated in Fig. 7b. Results of the FTIR spectrum (Fig. 7b) revealed that the spectrum in Fig. 7b displayed a broad absorption region between 3418 cm−1 assigned to the O–H bond in hydroxyl functional groups [56]. This confirmed the presence of the hydrogen bond of amines and alcohols (or phenols) in the biofilm. Small peaks of 2851 and 2921 cm−1 were due to CH2 stretching. The peak at 2519 cm−1 was due to O–H and C–N stretching and bending of carboxylic acids [57]. The peak at 860 cm−1 (aromatic C–H bending vibration) [58] and, peaks in the region of 600–800 cm−1 are due to aromatic compounds [59], which could be an indicator of humic substrates. The broad peak at around 1426 cm−1 was mainly derived from amides I and II [56]. In addition, a broad peak at 1072 cm−1 exhibits the character of carbohydrates or carbohydrate-like substances, which indicates that carbohydrates were present in the EPS [60].
Conclusion
Overall, the results of this study suggested that the denitrifying bacterium Thauera sp.V14 has great potential in the denitrification of nitrate containing wastewaters. Bioaugmentation of dMBBR with Thauera sp.V14 successfully enhanced denitrification efficiency as well as biofilm formation in the reactor. Thauera sp.V14 could efficiently remove nitrate and COD from wastewater without accumulation of NO2 and NH4 and form biofilms. Its hydrophobicity, auto-aggregation ability, biofilm formation ability, and denitrification competence indicate its adaptability for efficient colonization and nitrate removal. These characteristics emphasize potential of Thauera sp. V14 as an effective bacterium for the removal of nitrate from wastewater.
Availability of data and materials
Not applicable.
Code availability
Not applicable.
References
Barwal A, Chaudhary R (2016) Application of response surface methodology to optimize the operational parameters for enhanced removal efficiency of organic matter and nitrogen: moving bed biofilm reactor. Environ Sci Pollut Res 23:9944–9955. https://doi.org/10.1007/s11356-016-6250-z
Zekker I, Rikmann E, Mandel A et al (2016) Step-wise temperature decreasing cultivates a biofilm with high nitrogen removal rates at 9°C in short-term anammox biofilm tests. Environ Technol 37:1933–1946. https://doi.org/10.1080/09593330.2015.1135995
Ikram M, Naeem M, Zahoor M et al (2022) Biological degradation of the azo dye basic orange 2 by Escherichia coli: a sustainable and ecofriendly approach for the treatment of textile wastewater. Water 14:2063. https://doi.org/10.3390/w14132063
Khan AU, Rehman MU, Zahoor M et al (2021) Biodegradation of brown 706 dye by bacterial strain Pseudomonas aeruginosa. Water 13:2959. https://doi.org/10.3390/w13212959
Zekker I, Rikmann E, Tenno T et al (2015) Start-up of low-temperature anammox in UASB from mesophilic yeast factory anaerobic tank inoculum. Environ Technol 36:214–225. https://doi.org/10.1080/09593330.2014.941946
Xiang Z, Chen X, Bai J et al (2023) Bioaugmentation performance for moving bed biofilm reactor (MBBR) treating mariculture wastewater by an isolated novel halophilic heterotrophic nitrification aerobic denitrification (HNAD) strain (Zobellella B307). J Environ Manage 325:116566. https://doi.org/10.1016/j.jenvman.2022.116566
Chen X, Yuan C, Zhu Y et al (2022) Bioaugmentation with Acinetobacter sp. TAC-1 to enhance nitrogen removal in swine wastewater by moving bed biofilm reactor inoculated with bacteria. Bioresour Technol 359:127506. https://doi.org/10.1016/j.biortech.2022.127506
Ding L, Han B, Zhou J (2022) Characterization of the facultative anaerobic Pseudomonas stutzeri strain HK13 to achieve efficient nitrate and nitrite removal. Process Biochem 118:236–242. https://doi.org/10.1016/j.procbio.2022.04.021
Fan X, Nie L, Chen Z, et al (2023) Simultaneous removal of nitrogen and arsenite by heterotrophic nitrification and aerobic denitrification bacterium Hydrogenophaga sp. H7. Front Microbiol 13:1103913. https://doi.org/10.3389/fmicb.2022.1103913
Patel RJ, Patel UD, Nerurkar AS (2021) Moving bed biofilm reactor developed with special microbial seed for denitrification of high nitrate containing wastewater. World J Microbiol Biotechnol 37:1–13. https://doi.org/10.1007/s11274-021-03035-0
Heider J, Fuchs G (2015) Thauera. Bergey’s Man Syst Archaea Bact. https://doi.org/10.1002/9781118960608.gbm01004
Macy JM, Rech S, Auling G et al (1993) Subclass of Proteobacteria with a novel type of anaerobic respiration. Int J Syst Bacteriol 43:135–142. https://doi.org/10.1099/00207713-43-1-135
Wei Q, Zhang J, Luo F et al (2022) Molecular mechanisms through which different carbon sources affect denitrification by Thauera linaloolentis : electron generation, transfer, and competition. Environ Int 170:107598. https://doi.org/10.1016/j.envint.2022.107598
Ji B, Yang K, Zhu L et al (2015) Aerobic denitrification: a review of important advances of the last 30 years. Biotechnol Bioprocess Eng 20:643–651. https://doi.org/10.1007/s12257-015-0009-0
Du R, Cao S, Li B et al (2019) Step-feeding organic carbon enhances high-strength nitrate and ammonia removal via DEAMOX process. Chem Eng J 360:501–510. https://doi.org/10.1016/j.cej.2018.12.011
Mao Y, Xia Y, Zhang T (2013) Characterization of Thauera -dominated hydrogen-oxidizing autotrophic denitrifying microbial communities by using high-throughput sequencing. Bioresour Technol 128:703–710. https://doi.org/10.1016/j.biortech.2012.10.106
Rujakom S, Kamei T, Kazama F (2022) Thauera sp. in hydrogen-based denitrification: effects of plentiful bicarbonate supplementation on powerful nitrite reducer. Sustain 15:277. https://doi.org/10.3390/su15010277
Chang C, Tanong K, Xu J, Shon H (2011) Microbial community analysis of an aerobic nitrifying-denitrifying MBR treating ABS resin wastewater. Bioresour Technol 102:5337–5344. https://doi.org/10.1016/j.biortech.2010.12.045
Wu X, Wu X, Li J et al (2020) Cross-feeding between Thauera aminoaromatica and Rhodococcus pyridinivorans drove quinoline biodegradation in a denitrifying bioreactor. BioRxiv 2020:01. https://doi.org/10.1101/2020.01.31.929745
Wang Q, He J (2020) Complete nitrogen removal via simultaneous nitrification and denitrification by a novel phosphate accumulating Thauera sp. strain SND5. Water Res 185:116300. https://doi.org/10.1016/j.watres.2020.116300
Xiong W, Wang S, Jin Y et al (2023) Insights into nitrogen and phosphorus metabolic mechanisms of algal-bacterial aerobic granular sludge via metagenomics: performance, microbial community and functional genes. Bioresour Technol 369:128442. https://doi.org/10.1016/j.biortech.2022.128442
Ren T, Chi Y, Wang Y et al (2021) Diversified metabolism makes novel Thauera strain highly competitive in low carbon wastewater treatment. Water Res 206:117742. https://doi.org/10.1016/j.watres.2021.117742
Hong P, Wu X, Shu Y et al (2020) Bioaugmentation treatment of nitrogen-rich wastewater with a denitrifier with biofilm-formation and nitrogen-removal capacities in a sequencing batch biofilm reactor. Bioresour Technol 303:122905. https://doi.org/10.1016/j.biortech.2020.122905
Wilke A, Gerlach W, Harrison T, Paczian T, Trimble WL, Meyer F (2017) MG-RAST manual for version 4, revision 3. Argonne National Laboratory, Lemont
Loy A, Schulz C, Lücker S, Schöpfer-Wendels A, Stoecker K, Baranyi C, Wagner M (2005) 16S rRNA gene-based oligonucleotide microarray for environmental monitoring of the betaproteobacterial order “Rhodocyclales.” Appl Environ Microbiol 71(3):1373–1386. https://doi.org/10.1128/AEM.71.3.1373-1386.2005
Szulc A et al (2014) The in fl uence of bioaugmentation and biosurfactant addition on bioremediation ef fi ciency of diesel-oil contaminated soil : feasibility during field studies. J Environ Manage 132:121–128. https://doi.org/10.1016/j.jenvman.2013.11.006
López MJ, Jurado MM, Moreno J (2015) Bioresource technology dynamics of bacterial microbiota during lignocellulosic waste composting : studies upon its structure, functionality and biodiversity. Bioresour Technol 175:406–416. https://doi.org/10.1016/j.biortech.2014.10.123
Wei H, Wang L, Hassan M, Xie B (2018) Succession of the functional microbial communities and the metabolic functions in maize straw composting process. Bioresour Technol 256:333–341. https://doi.org/10.1016/j.biortech.2018.02.050
Cui YX, Guo G, Biswal BK et al (2019) Investigation on sulfide-oxidizing autotrophic denitrification in moving-bed biofilm reactors: an innovative approach and mechanism for the process start-up. Int Biodeterior Biodegrad 140:90–98. https://doi.org/10.1016/j.ibiod.2019.03.016
Yan W, Wang N, Wei D et al (2021) Bacterial community compositions and nitrogen metabolism function in a cattle farm wastewater treatment plant revealed by Illumina high-throughput sequencing. Environ Sci Pollut Res 28:40895–40907. https://doi.org/10.1007/s11356-021-13570-w
Nzila A (2018) Biodegradation of high-molecular-weight polycyclic aromatic hydrocarbons under anaerobic conditions: Overview of studies, proposed pathways and future perspectives. Environ Pollut 239:788–802. https://doi.org/10.1016/j.envpol.2018.04.074
Sul W-J, Kim I-S, Ekpeghere KI et al (2016) Metagenomic insight of nitrogen metabolism in a tannery wastewater treatment plant bioaugmented with the microbial consortium BM-S-1. J Environ Sci Heal Part A 51:1164–1172. https://doi.org/10.1080/10934529.2016.1206387
Klotz MG, Bryant DA, Hanson TE (2011) The microbial sulfur cycle. Front Microbiol 2:00241. https://doi.org/10.3389/fmicb.2011.00241
Andalib M, Nakhla G, McIntee E, Zhu J (2011) Simultaneous denitrification and methanogenesis (SDM): review of two decades of research. Desalination 279:1–14. https://doi.org/10.1016/j.desal.2011.06.018
Cheng X-Y, Tian X-L, Wang Y-S et al (2013) Metagenomic analysis of the pinewood nematode microbiome reveals a symbiotic relationship critical for xenobiotics degradation. Sci Rep 3:1869. https://doi.org/10.1038/srep01869
Wang Y, Zou YL, Chen H, Lv YK (2021) Nitrate removal performances of a new aerobic denitrifier, Acinetobacter haemolyticus ZYL, isolated from domestic wastewater. Bioprocess Biosyst Eng 44:391–401. https://doi.org/10.1007/s00449-020-02451-0
Zhao B, Cheng DY, Tan P et al (2018) Characterization of an aerobic denitrifier Pseudomonas stutzeri strain XL-2 to achieve efficient nitrate removal. Bioresour Technol 250:564–573. https://doi.org/10.1016/j.biortech.2017.11.038
Bai H, Liao S, Wang A et al (2019) High-efficiency inorganic nitrogen removal by newly isolated Pannonibacter phragmitetus B1. Bioresour Technol 271:91–99. https://doi.org/10.1016/j.biortech.2018.09.090
Wang X, Wang W, Zhang Y et al (2019) Simultaneous nitrification and denitrification by a novel isolated Pseudomonas sp. JQ-H3 using polycaprolactone as carbon source. Bioresour Technol. https://doi.org/10.1016/j.biortech.2019.121506
Huang TL, Zhou SL, Zhang HH et al (2015) Nitrogen removal characteristics of a newly isolated indigenous aerobic denitrifier from oligotrophic drinking water reservoir, Zoogloea sp. N299. Int J Mol Sci 16:10038–10060. https://doi.org/10.3390/ijms160510038
Malik A, Sakamoto M, Hanazaki S et al (2003) Coaggregation among nonflocculating bacteria isolated from activated sludge. Appl Environ Microbiol 69:6056–6063. https://doi.org/10.1128/AEM.69.10.6056
Wang X, An Q, Zhao B et al (2018) Auto-aggregation properties of a novel aerobic denitrifier Enterobacter sp. strain FL. Appl Microbiol Biotechnol 102:2019–2030. https://doi.org/10.1007/s00253-017-8720-8
Hong P, Shu Y, Wu X et al (2019) Efficacy of zero nitrous oxide emitting aerobic denitrifying bacterium, Methylobacterium gregans DC-1 in nitrate removal with strong auto-aggregation property. Bioresour Technol 293:122083. https://doi.org/10.1016/j.biortech.2019.122083
Dhamole PB, Nair RR, D’Souza SF, Lele SS (2007) Denitrification of high strength nitrate waste. Bioresour Technol 98:247–252. https://doi.org/10.1016/j.biortech.2006.01.019
Adimalla N, Qian H (2021) Ecotoxicology and Environmental Safety Groundwater chemistry, distribution and potential health risk appraisal of nitrate enriched groundwater : a case study from the semi-urban region of. Ecotoxicol Environ Saf 207:111277. https://doi.org/10.1016/j.ecoenv.2020.111277
Vaiphei SP, Kurakalva RM (2021) Ecotoxicology and environmental safety hydrochemical characteristics and nitrate health risk assessment of groundwater through seasonal variations from an intensive agricultural region of upper Krishna River basin, Telangana, India. Ecotoxicol Environ Saf 213:112073. https://doi.org/10.1016/j.ecoenv.2021.112073
Aravinthasamy DKP, Subramani MDT (2021) Groundwater pollution and human health risks in an industrialized region of Southern India: impacts of the COVID-19 lockdown and the monsoon seasonal cycles. Arch Environ Contam Toxicol 80:259–276. https://doi.org/10.1007/s00244-020-00797-w
Nawale VP, Malpe DB, Marghade D, Yenkie R (2021) Ecotoxicology and Environmental Safety Non-carcinogenic health risk assessment with source identification of nitrate and fluoride polluted groundwater of Wardha sub-basin, central India. Ecotoxicol Environ Saf 208:111548. https://doi.org/10.1016/j.ecoenv.2020.111548
Karunanidhi D, Aravinthasamy P, Subramani T, Kumar M (2021) Human health risks associated with multipath exposure of groundwater nitrate and environmental friendly actions for quality improvement and sustainable management: a case study from Texvalley (Tiruppur region) of India. Chemosphere 265:129083. https://doi.org/10.1016/j.chemosphere.2020.129083
Suvarna B, Sunitha V, Reddy YS, Reddy NR (2020) Data health risk assessment of nitrate contamination in groundwater of rural region in the Yerraguntla Mandal. South India Data Br 30:105374. https://doi.org/10.1016/j.dib.2020.105374
Singh S, Hariteja N, Renuka Prasad TJ et al (2020) Impact assessment of faecal sludge on groundwater and river water quality in Lucknow environs, Uttar Pradesh, India. Groundw Sustain Dev 11:100461. https://doi.org/10.1016/j.gsd.2020.100461
Maurya J, Pradhan SN, Seema GAK (2020) Evaluation of ground water quality and health risk assessment due to nitrate and fluoride in the Middle Indo-Gangetic plains of India. Hum Ecol Risk Assess 27:1349–1365. https://doi.org/10.1080/10807039.2020.1844559
Panneerselvam B, Muniraj K, Duraisamy K et al (2023) An integrated approach to explore the suitability of nitrate-contaminated groundwater for drinking purposes in a semiarid region of India. Environ Geochem Health 45:647–663. https://doi.org/10.1007/s10653-022-01237-5
Zhang S, Sun X, Wang X et al (2018) Bioaugmentation with Diaphorobacter polyhydroxybutyrativorans to enhance nitrate removal in a poly (3-hydroxybutyrate-co-3-hydroxyvalerate)-supported denitrification reactor. Bioresour Technol 263:499–507. https://doi.org/10.1016/j.biortech.2018.04.115
Shen Z, Zhou Y, Hu J, Wang J (2013) Denitrification performance and microbial diversity in a packed-bed bioreactor using biodegradable polymer as carbon source and biofilm support. J Hazard Mater 250–251:431–438. https://doi.org/10.1016/j.jhazmat.2013.02.026
Wang Z, Wu Z, Tang S (2009) Extracellular polymeric substances ( EPS ) properties and their effects on membrane fouling in a submerged membrane bioreactor. Water Res 43:2504–2512. https://doi.org/10.1016/j.watres.2009.02.026
Islam MA, Benhouria A, Asif M, Hameed BH (2015) Journal of the Taiwan Institute of Chemical Engineers Methylene blue adsorption on factory-rejected tea activated carbon prepared by conjunction of hydrothermal carbonization and sodium hydroxide activation processes. J Taiwan Inst Chem Eng 52:57–64. https://doi.org/10.1016/j.jtice.2015.02.010
Huang H, Ren H, Ding L et al (2014) Aging biofilm from a full-scale moving bed biofilm reactor: characterization and enzymatic treatment study. Bioresour Technol 154:122–130. https://doi.org/10.1016/j.biortech.2013.12.031
Tran T, Bolto B, Gray S et al (2007) An autopsy study of a fouled reverse osmosis membrane element used in a brackish water treatment plant. Water Res 41:3915–3923. https://doi.org/10.1016/j.watres.2007.06.008
Fan Y, Su J, Zheng Z et al (2021) Denitrification performance and mechanism of a novel isolated Acinetobacter sp FYF8 in oligotrophic ecosystem. Bioresour Technol 320:124280. https://doi.org/10.1016/j.biortech.2020.124280
Acknowledgements
This work was supported by the Department of Biotechnology (DBT), (Grant No. BT/PR15531 /BCE/8/1156/2016) GOI for fellowship and Department of Science and Technology, GOI, FIST programme for instrumentation facility.
Funding
The authors acknowledge the Department of Biotechnology (DBT), (Grant No.BT/PR15531/BCE/8/1156/2016) GOI for fellowship, and Department of Science and Technology, GOI, FIST programme for instrumentation facility.
Author information
Authors and Affiliations
Contributions
Conceptualization and design of experiments: RJP and ASN; performance of experiments: RJP; reagents/materials/equipment: ASN; manuscript writing, analysis and discussion: RJP and ASN.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to declare that are relevant to the content of this article.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Patel, R.J., Nerurkar, A.S. Thauera sp. for efficient nitrate removal in continuous denitrifying moving bed biofilm reactor. Bioprocess Biosyst Eng 47, 429–442 (2024). https://doi.org/10.1007/s00449-024-02977-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-024-02977-7